Convolutional Neural Network to Classify Infrared Thermal Images of Fractured Wrists in Pediatrics
Abstract
1. Introduction
1.1. Deep Learning
1.2. Convolutional Neural Network and Data Augmentation
- A CNN model devised to identify wrist fractures from IRT images. The ability of the CNN was leveraged to automatically extract features based on the convolution layers as previous methods used manual feature extraction methods.
- A proposed CNN model that was tailor-made for the specific input IRT image characteristics.
- An exploration of the effectiveness of image augmentation when developing DL models for IRT image classification was performed.
- The performance of CNN in identifying wrist fractures was compared with an earlier study that used a multilayer perceptron neural network, indicating improved accuracy for the CNN.
2. Related Studies
2.1. Infrared Thermal Imaging for Medical Diagnosis
2.2. Machine Learning and Deep Learning Techniques
3. Materials and Methods
3.1. Participants’ Recruitment and Data Recording
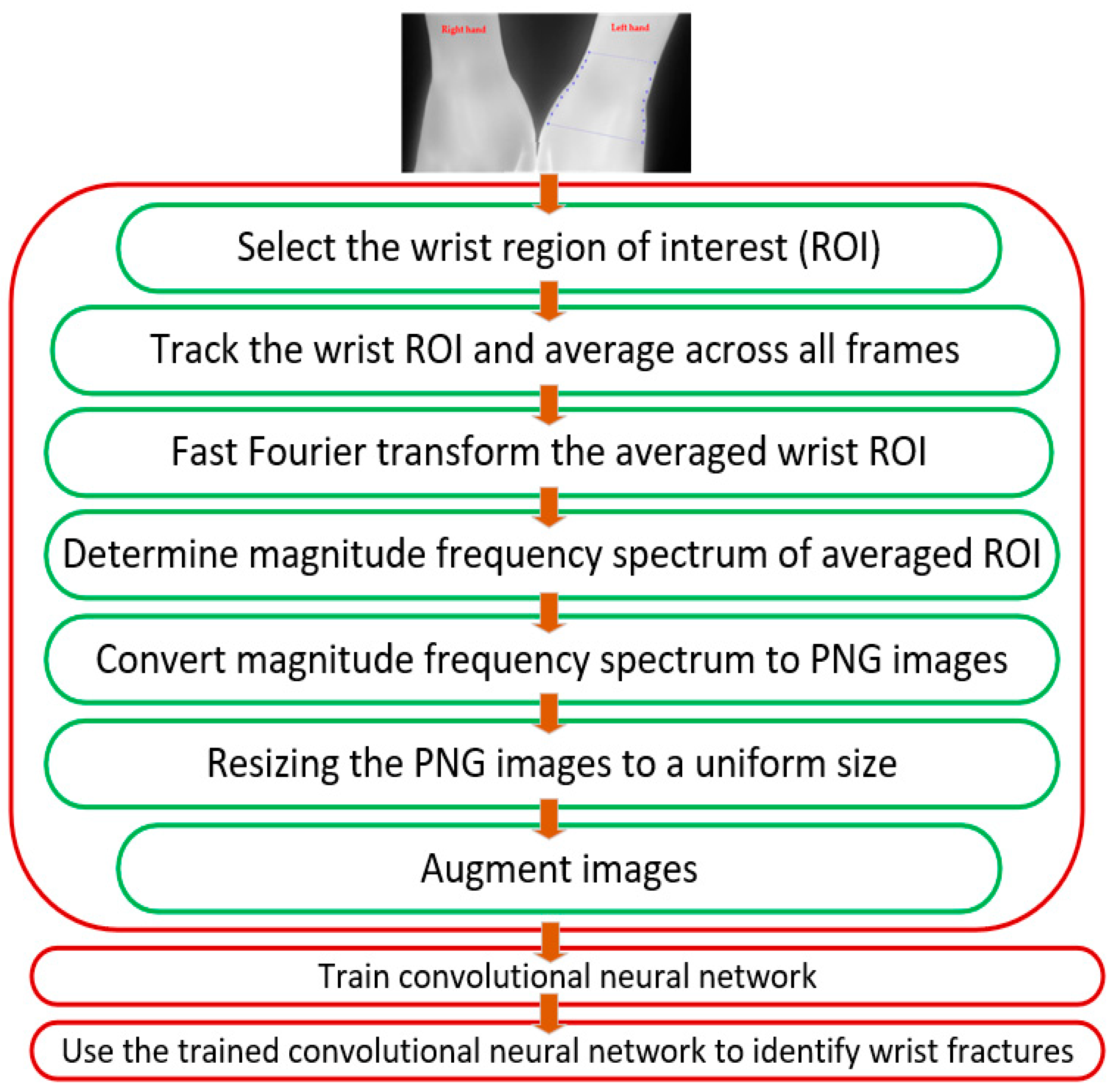
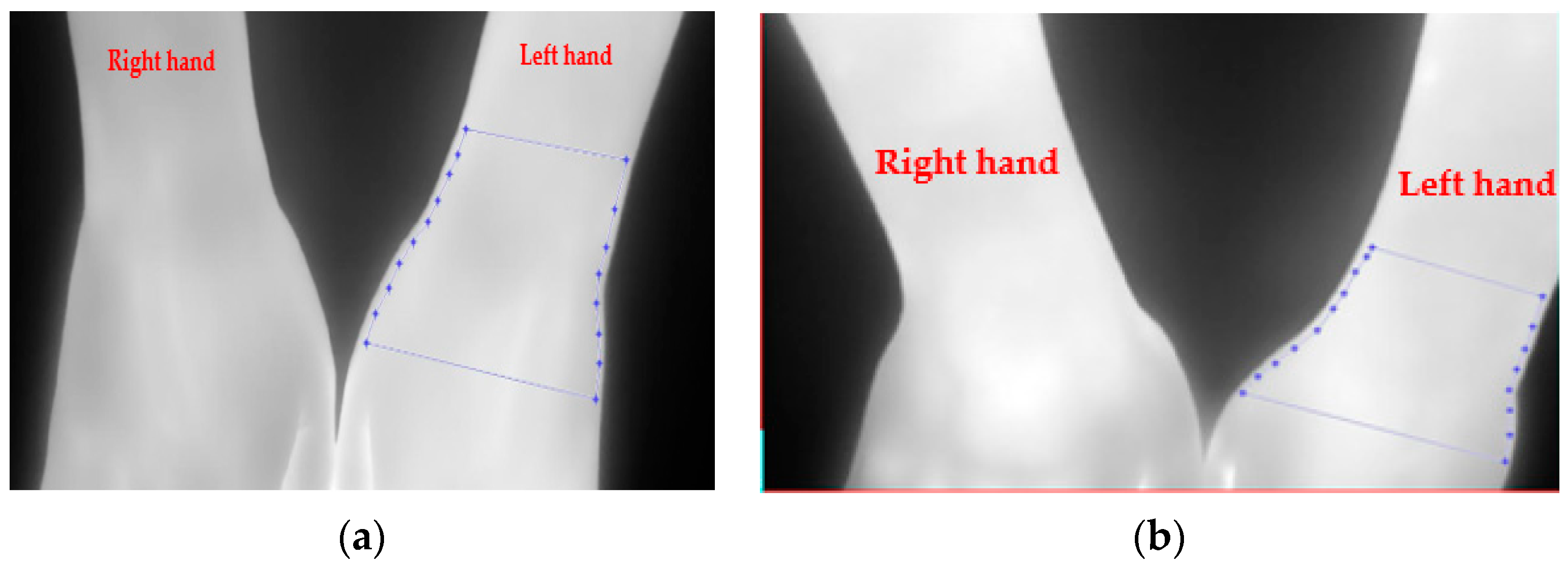
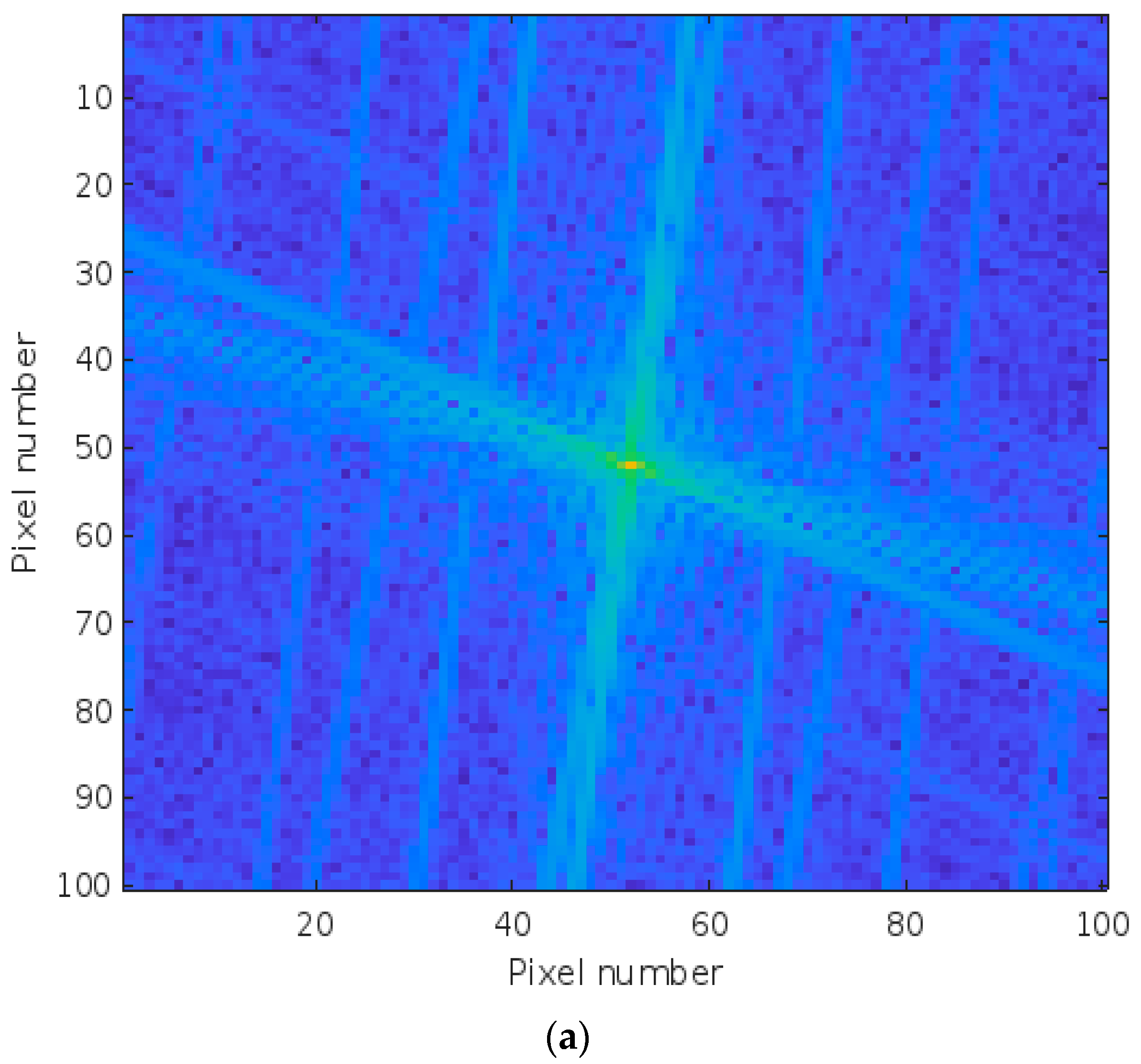
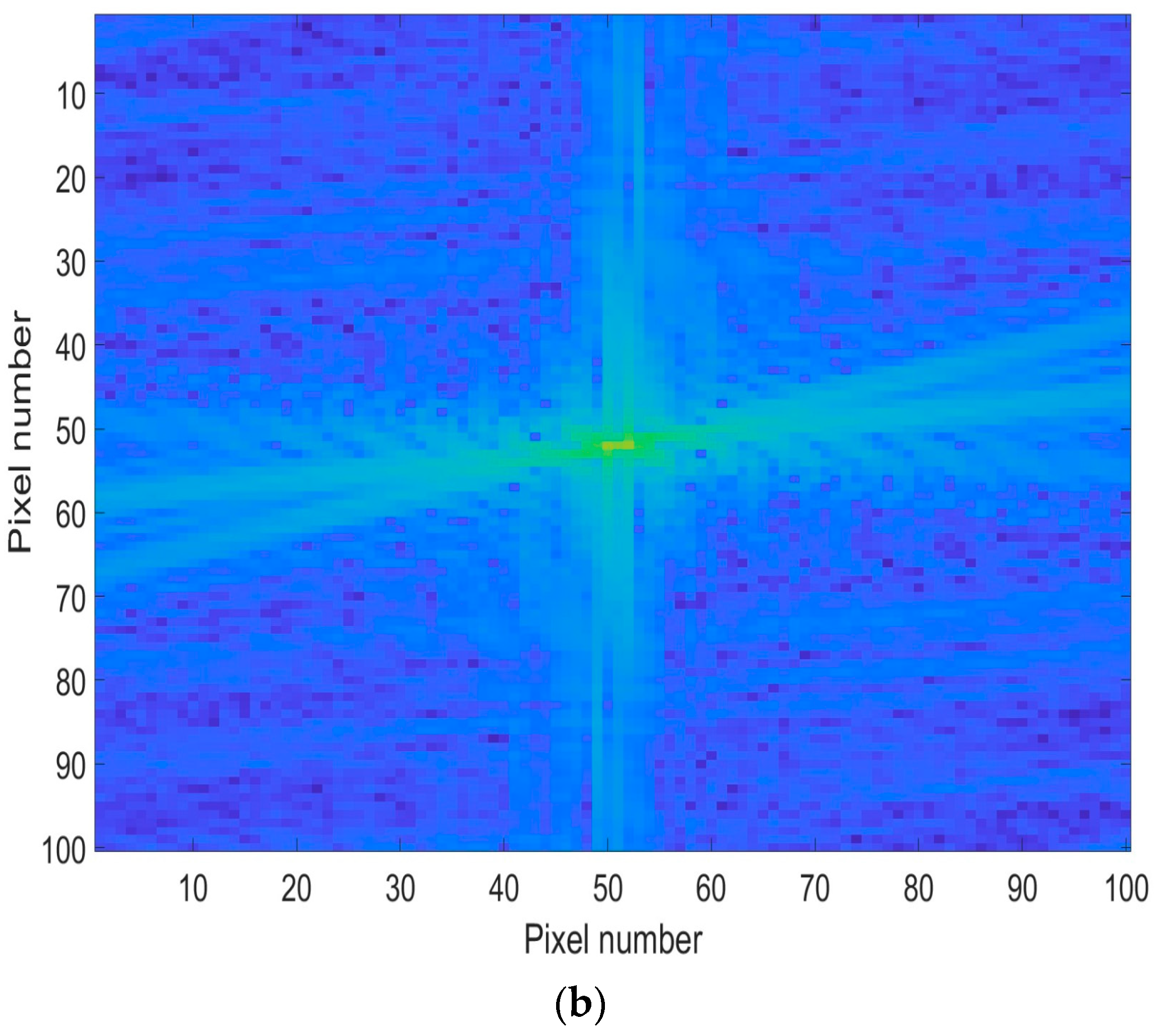
3.2. Image Pre-Processing
Image Augmentation
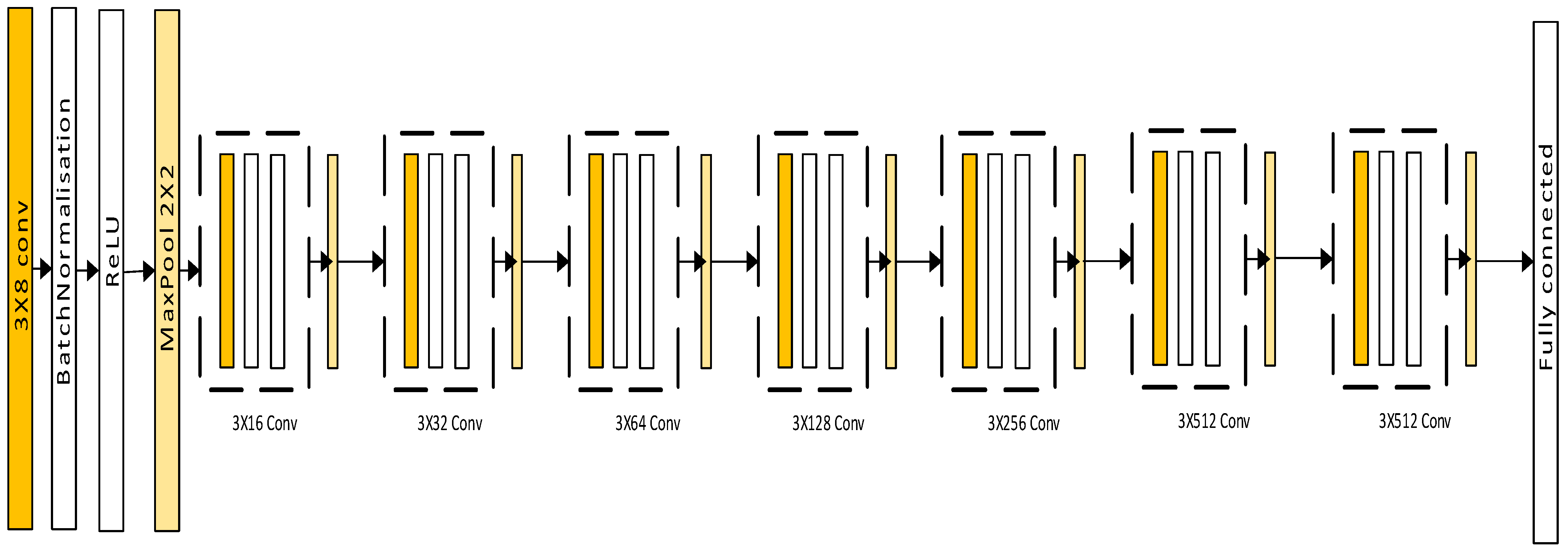
3.3. CNN-Based Deep Learning
3.4. Evaluation Metrics
4. Results
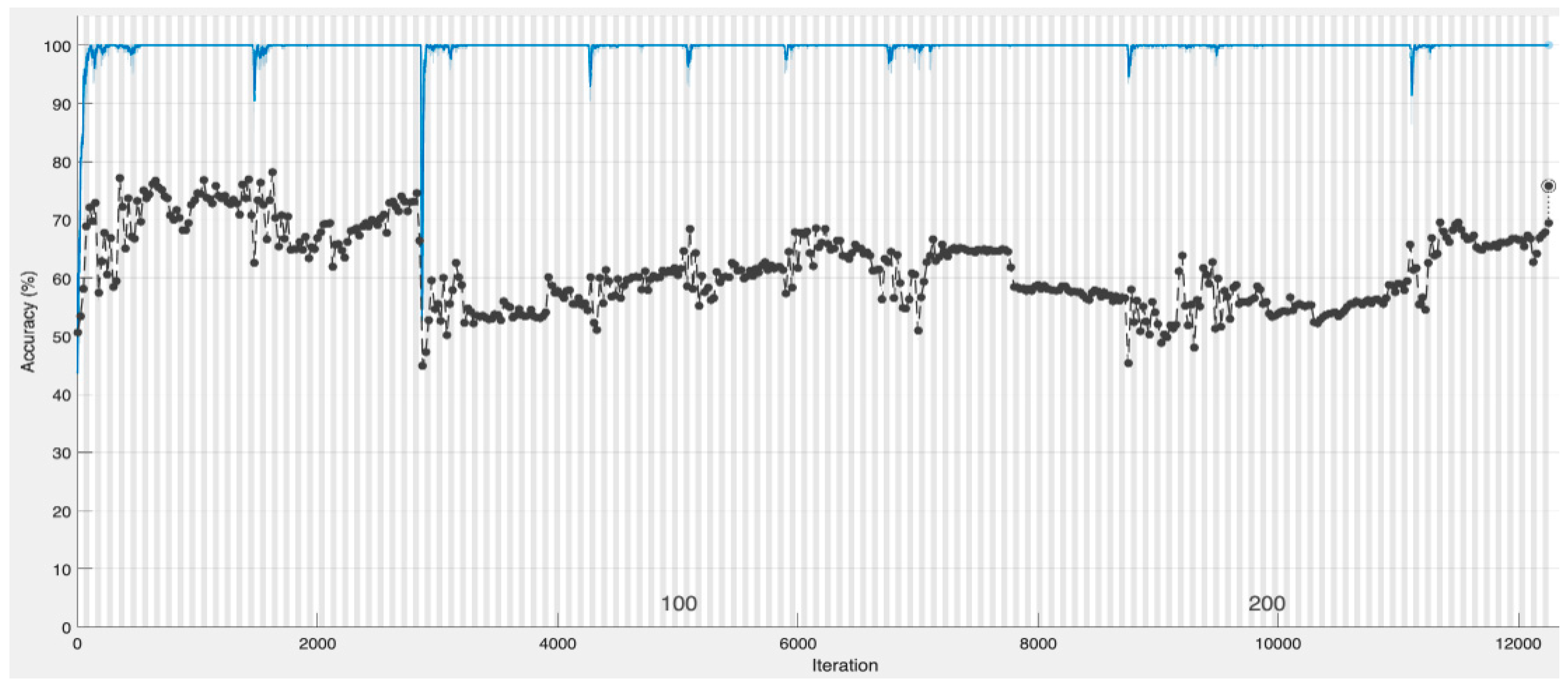
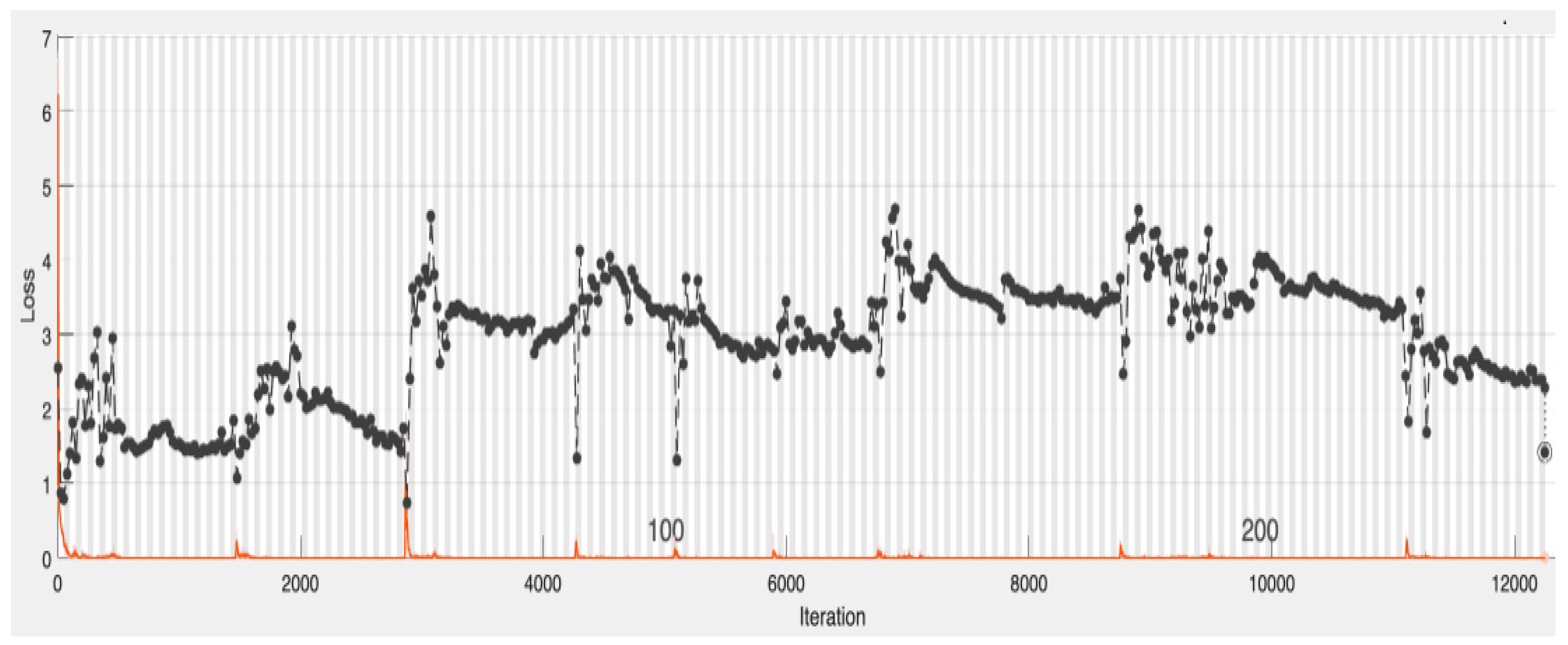
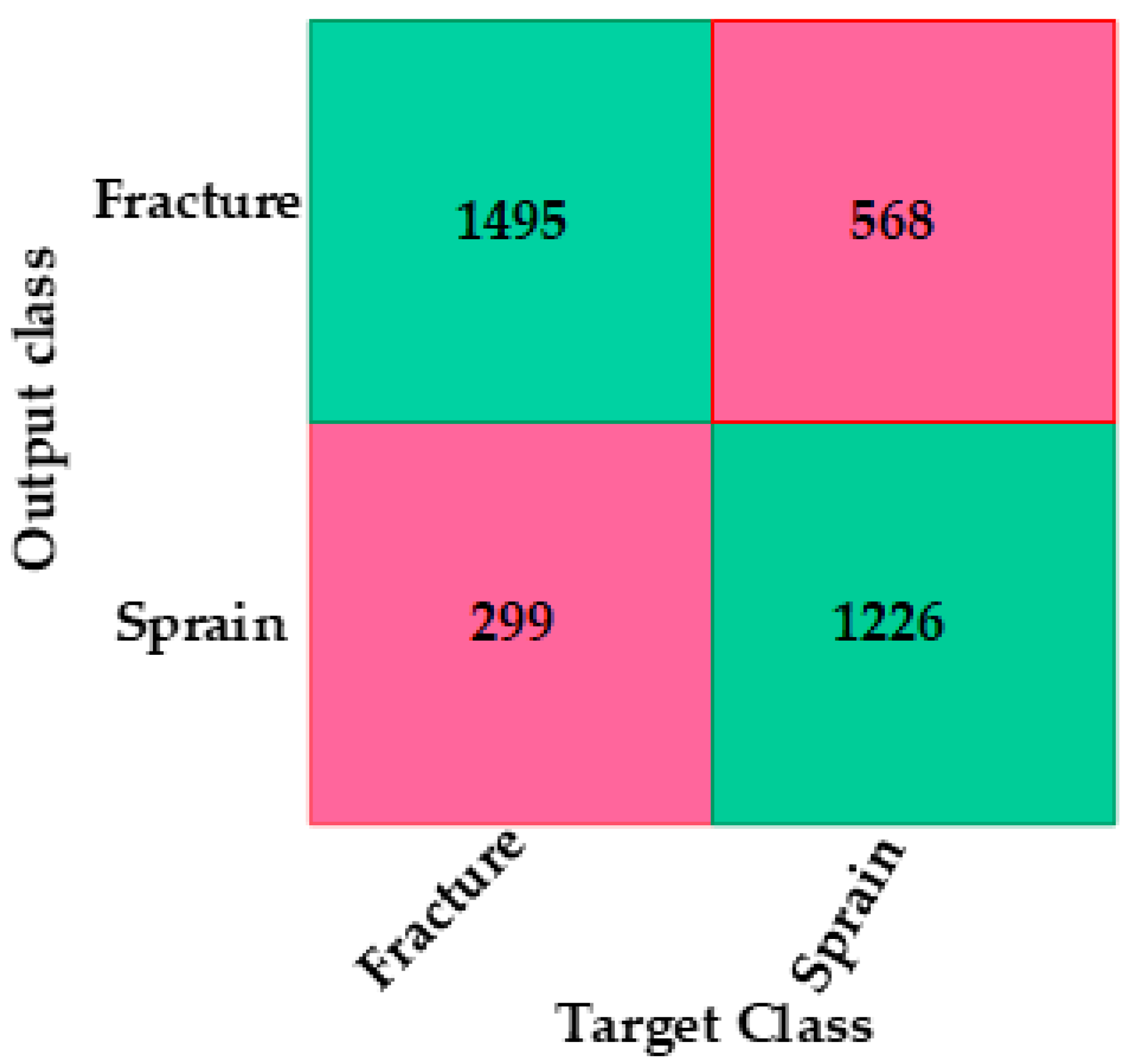
4.1. CNN-Based Deep Neural Network with Augmentation and without Dropout
4.2. Results for CNN-Based Deep Neural Network without Augmentation and with Dropout
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Reed, C.; Saatchi, R.; Burke, D.; Ramlakhan, S. Infrared thermal imaging as a screening tool for paediatric wrist fractures. Med. Biol. Eng. Comput. 2020, 58, 1549–1563. [Google Scholar] [CrossRef]
- Zhang, L.; Guob, H.; Lic, Z. Application of medical infrared thermal imaging in the diagnosis of human internal focus. Infrared Phys. Technol. 2019, 101, 127–132. [Google Scholar] [CrossRef]
- Kastberger, G.; Stachl, R. Infrared imaging technology and biological applications. Behav. Res. Methods Instrum. Comput. 2003, 35, 429–439. [Google Scholar] [CrossRef]
- De Salis, A.F.; Saatchi, R.; Dimitri, P. Evaluation of high resolution thermal imaging to determine the effect of vertebral fractures on associated skin surface temperature in children with osteogenesis imperfecta. Med. Biol. Eng. Comput. 2018, 56, 1633–1643. [Google Scholar] [CrossRef] [PubMed]
- Mousavi, A.; Arefanjazi, H.; Sadeghi, M.; Ghahfarokhi, A.M.; Beheshtinejad, F.; Masouleh, M.M. Comparison of feature eExtraction with PCA and LTP methods and investigating the effect of dimensionality reduction in the Bat algorithm for face recognition. Int. J. Robot. Control Syst. 2023, 3, 500–509. [Google Scholar]
- Liu, X.; Song, L.; Liu, S.; Zhang, Y. A review of deep-learning-based medical image segmentation methods. Sustainability 2021, 13, 1224. [Google Scholar] [CrossRef]
- Cai, L.; Gao, J.; Zhao, D. A review of the application of deep learning in medical image classification and segmentation. Ann. Transl. Med. 2020, 8, 713. [Google Scholar] [CrossRef] [PubMed]
- Alzubaidi, L.; Zhang, J.; Humaidi, A.J.; Al-Dujaili, A.; Duan, Y.; Al-Shamma, O.; Santamaría, J.; Fadhel, M.A.; Al-Amidie, M.; Farhan, L. Review of deep learning: Concepts, CNN architectures, challenges, applications, future directions. J. Big Data 2021, 8, 53. [Google Scholar] [CrossRef]
- Affonso, C.; Rossi, A.L.; Vieira, F.H.; de Leon Ferreira, A.C. Deep learning for biological image classification. Expert Syst. Appl. 2017, 1, 114–122. [Google Scholar] [CrossRef]
- Wu, M.; Chen, L. Image recognition based on deep learning. Chin. Autom. Congr. (CAC) 2015, 27, 542–546. [Google Scholar]
- Su, D.; Chen, L.; Du, X.; Liu, M.; Jin, L. Constructing convolutional neural network by utilizing nematode connectome: A brain-inspired method. Appl. Soft Comput. 2023, 149, 110992. [Google Scholar] [CrossRef]
- Li, Z.; Liu, F.; Yang, W.; Peng, S.; Zhou, J. A survey of convolutional neural networks: Analysis, applications, and prospects. IEEE Trans. Neural Netw. Learn. Syst. 2022, 33, 6999–7019. [Google Scholar] [CrossRef] [PubMed]
- Morid, M.A.; Borjali, A.; Del Fiol, G. A scoping review of transfer learning research on medical image analysis using ImageNet. Comput. Biol. Med. 2021, 128, 104115. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Guedes, A.; Santana-Perez, I.; Arteaga-Marrero, N.; Fabelo, H.; Callico, G.M.; Ruiz-Alzola, J. Performance evaluation of deep learning models for image classification over small datasets: Diabetic foot case study. IEEE Access 2022, 10, 124373–124386. [Google Scholar] [CrossRef]
- Alomar, K.; Aysel, H.I.; Cai, X. Data augmentation in classification and segmentation: A survey and new strategies. J. Imaging 2023, 9, 46. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, G.; Talavera, E.; González-Prieto, Á.; Mozo, A.; Gómez-Canaval, S. Data augmentation techniques in time series domain: A survey and taxonomy. Neural Comput. Appl. 2023, 35, 10123–10145. [Google Scholar] [CrossRef]
- Nanni, L.; Paci, M.; Brahnam, S.; Lumini, A. Comparison of different image data augmentation approaches. J. Imaging 2021, 7, 254. [Google Scholar] [CrossRef] [PubMed]
- Mumuni, A.; Mumuni, F. Data augmentation: A comprehensive survey of modern approaches. Array 2022, 16, 100258. [Google Scholar] [CrossRef]
- Wu, M.; Wang, S.; Pan, S.; Terentis, A.C.; Strasswimmer, J.; Zhu, X. Deep learning data augmentation for Raman spectroscopy cancer tissue classification. Nature 2021, 11, 23842. [Google Scholar] [CrossRef]
- Li, L.; Hou, Y.; Che, W. Data augmentation approaches in natural language processing: A survey. AI Open 2022, 3, 71–90. [Google Scholar] [CrossRef]
- Iwana, B.K.; Uchida, S. An empirical survey of data augmentation for time series classification with neural networks. PLoS ONE 2021, 17, e0254841. [Google Scholar] [CrossRef] [PubMed]
- Bayer, M.; Kaufhold, M.-A.; Reuter, C. A survey on data augmentation for text classification. ACM Comput. Surv. 2022, 55, 146. [Google Scholar] [CrossRef]
- Ma, J.; Hu, C.; Zhou, P.; Jin, F.; Wang, X.; Huang, H. Review of image augmentation used in deep learning-based material microscopic image segmentation. Appl. Sci. 2023, 13, 6478. [Google Scholar] [CrossRef]
- Zhang, R.; Zhou, B.; Lu, C.; Ma, M. The performance research of the data augmentation method for image classification. Math. Probl. Eng. 2022, 2022, 2964829. [Google Scholar] [CrossRef]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial networks. Commun. ACM 2020, 63, 139–144. [Google Scholar] [CrossRef]
- Brophy, E.; Wang, Z.; She, Q.; WARD, T. Generative adversarial networks in time Series: A systematic literature review. ACM Comput. Surv. 2023, 55, 199. [Google Scholar] [CrossRef]
- Chen, Y.; Yang, X.-H.; Wei, Z.; Heidari, A.A.; Zheng, N.; Li, Z.; Chen, H.; Hu, H.; Zhou, Q.; Guan, Q. Generative adversarial networks in medical image augmentation: A review. Comput. Biol. Med. 2022, 144, 105382. [Google Scholar] [CrossRef] [PubMed]
- Sandfort, V.; Yan, K.; Pickhardt, P.J.; Summers, R.M. Data augmentation using generative adversarial networks (CycleGAN) to improve generalizability in CT segmentation tasks. Nature 2019, 9, 16884. [Google Scholar] [CrossRef] [PubMed]
- Yahaya, M.S.M.; Teo, J. Data augmentation using generative adversarial networks for images and biomarkers in medicine and neuroscience. Front. Appl. Math. Stat. 2023, 9, 1162760. [Google Scholar] [CrossRef]
- Frid-Adar, M.; Diamant, I.; Klang, E.; Amitai, M.; Goldberger, J.; Greenspan, H. GAN-based synthetic medical image augmentation for increased CNN performance in liver lesion classification. Neurocomputing 2018, 321, 321–331. [Google Scholar] [CrossRef]
- Saatchi, R.; Ramlakhan, S. Infrared thermal imaging analysis in screening for toddler’s fracture: A proof-of-concept study. Appl. Sci. 2023, 13, 13299. [Google Scholar] [CrossRef]
- Sung, J.; Loughin, C.; Marino, D.; Leyva, F.; Dewey, C.; Umbaugh, S.; Lesser, M. Medical infrared thermal imaging of canine appendicular bone neoplasia. BMC Vet. Res. 2019, 15, 430. [Google Scholar] [CrossRef]
- Ćurković, S.; Antabak, A.; Halužan, D.; Luetić, T.; Prlić, I.; Šiško, J. Medical thermography (digital infrared thermal imaging–DITI) in paediatric forearm fractures—A pilot study. Injury 2015, 1, S36–S39. [Google Scholar] [CrossRef] [PubMed]
- Magas, V.; Abreu de Souza, M.; Borba Neves, E.; Nohama, P. Evaluation of thermal imaging for the diagnosis of repetitive strain injuries of the wrist and hand joints. Res. Biomed. Eng. 2019, 1, 57–64. [Google Scholar] [CrossRef]
- Sanchis-Sánchez, E.; Salvador-Palmer, R.; Codoñer-Franch, P.; Martín, J.; Vergara-Hernández, C.; Blasco, J.; Ballester, E.; Sanchis, E.; González-Peña, R.; Cibrián, R. Infrared thermography is useful for ruling out fractures in paediatric emergencies. Eur. J. Pediatr. 2015, 174, 493–499. [Google Scholar] [CrossRef] [PubMed]
- Snekhalatha, U.; Sangamithirai, K. Computer aided diagnosis of obesity based on thermal imaging using various convolutional neural networks. Biomed. Signal Process. Control 2021, 63, 102233. [Google Scholar]
- Ghayoumi, Z.H.; Haddadnia, J.; Hashemian, M.; Hassanpour, K. Diagnosis of breast cancer using a combination of genetic algorithm and artificial neural network in medical infrared thermal imaging. Iran. J. Med. Phys. 2012, 9, 265–274. [Google Scholar]
- Shobayo, O.; Saatchi, R.; Ramlakhan, S. Infrared thermal imaging and artificial neural networks to screen for wrist fractures in pediatrics. Technologies 2022, 10, 119. [Google Scholar] [CrossRef]
- Vardasca, R.; Vaz, L.; Magalhaes, C.; Seixas, A.; Mendes, J. Towards the diabetic foot ulcers classification with infrared thermal images. In Proceedings of the 14th Quantitative Infrared Thermography Conference, Berlin, Germany, 25–29 June 2018; pp. 25–29. [Google Scholar]
- Ghaderzadeh, M.; Hosseini, A.; Asadi, F.; Abolghasemi, H.; Bashash, D.; Roshanpoor, A. Automated detection model in classification of B-lymphoblast cells from normal B-lymphoid precursors in blood smear microscopic images based on the majority voting technique. Sci. Program. 2022, 2022, 4801671. [Google Scholar] [CrossRef]
- Gheisari, M.; Ghaderzadeh, M.; Li, H.; Taami, T.; Fernández-Campusano, C.; Sadeghsalehi, H.; Abbasi, A.A. Mobile apps for COVID-19 detection and diagnosis for future pandemic control: Multidimensional systematic review. JMIR Mhealth Uhealth 2024, 12, e44406. [Google Scholar] [CrossRef]
- Ghaderzadeh, M.; Asadi, F.; Ghorbani, N.R.; Almasi, S.; Taami, T. Toward artificial intelligence (AI) applications in the determination of COVID-19 infection severity: Considering AI as a disease control strategy in future pandemics. Iran. J. Blood Cancer 2023, 15, 93–111. [Google Scholar] [CrossRef]
- Özdil, A.; Yilmaz, B. Medical infrared thermal image based fatty liver classification using machine and deep learning. Quant. InfraRed Thermogr. J. 2024, 21, 102–119. [Google Scholar] [CrossRef]
- Hoffer, O.; Brzezinski, R.Y.; Ganim, A.; Shalom, P.; OvadiaBlechman, Z.; Ben-Baruch, L.; Lewis, N.; Peled, R.; Shimon, C.; Naftali-Shani, N.; et al. Smartphone-based detection of COVID-19 and associated pneumonia using thermal imaging and a transfer learning algorithm. J. Biophotonics 2024, 2024, e202300486. [Google Scholar] [CrossRef] [PubMed]
- Nagori, A.; Dhingra, L.S.; Bhatnagar, A.; Lodha, R.; Sethi, T. Predicting hemodynamic shock from thermal images using machine learning. Sci. Rep. 2019, 9, 91. [Google Scholar] [CrossRef] [PubMed]
- Jayanthi, T.; Anburajan, M. Model-based computer-aided method for diagnosis of cardiovascular disease using IR thermogram. Biomed. Res. 2019, 30, 95–101. [Google Scholar]
- Samant, P.; Agarwal, R. Machine learning techniques for medical diagnosis of diabetes using iris images. Comput. Methods Programs Biomed. 2018, 57, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, R.; Nishio, M.; Do, R.K.G.; Togashi, K. Convolutional neural networks: An overview and application in radiology. Insights Imaging 2018, 9, 611–629. [Google Scholar] [CrossRef] [PubMed]
- Selvathi, D.; Suganya, K. Support vector machine based method for automatic detection of diabetic eye disease using thermal images. In Proceedings of the 1st International Conference on Innovations in Information and Communication Technology (ICIICT), Chennai, India, 25–26 April 2019; pp. 1–6. [Google Scholar]
- Paulino, J.A.; da Silva Alves, A.V.; de Sousa, R.P.; Galdino, K.E.; Alves, P.M.; Oliveira, E.J. The use of hybrid classifier to assist in the diagnosis of lip lesions in thermal images: A preliminary study. IEEE Int. Conf. Bioinform. Biomed. (BIBM) 2019, 18, 1585–1588. [Google Scholar]
- Yadav, S.S.; Jadhav, S.M. Thermal infrared imaging based breast cancer diagnosis using machine learning techniques. Multimed. Tools Appl. 2022, 1, 13139–13157. [Google Scholar] [CrossRef]
- Wang, Y.; Jiang, X.; Yu, K.; Shi, F.; Qin, L.; Zhou, H.; Cai, F. Infrared thermal images classification for pressure injury prevention incorporating the convolutional neural networks. IEEE Access 2021, 9, 15181–15190. [Google Scholar] [CrossRef]
- Shorten, C.; Khoshgoftaar, T.M. A survey on image data augmentation for deep learning. J. Big Data 2019, 6, 60. [Google Scholar] [CrossRef]
- MathWorks Inc. Deep Learning Toolbox, Version: 23.2 (R2023b); The MathWorks Inc.: Natick, MA, USA, 2023; Available online: https://www.mathworks.com (accessed on 15 April 2024).
- Venkatesan, R.; Li, B. Convolutional Neural Networks in Visual Computing: A Concise Guide; CRC Press: Boca Raton, FL, USA, 2017. [Google Scholar] [CrossRef]
- Nair, V.; Geoffrey, E.H. Rectified linear units improve restricted Boltzmann machines. In Proceedings of the 27th International Conference on Machine Learning (ICML-10), Haifa, Israel, 21–24 June 2010; pp. 807–814. [Google Scholar]
- Zhu, Q.; He, Z.; Zhang, T.; Cui, W. Improving classification performance of Softmax loss function based on scalable batch-normalization. Appl. Sci. 2020, 10, 2950. [Google Scholar] [CrossRef]
- Pham, T.C.; Luong, C.M.; Visani, M.; Hoang, V.D. Deep CNN and data augmentation for skin lesion classification. In Intelligent Information and Database Systems: 10th Asian Conference, ACIIDS 2018, Dong Hoi City, Vietnam, 19–21 March 2018; Proceedings, Part II 10; Springer International Publishing: Berlin/Heidelberg, Germany, 2018; pp. 573–582. [Google Scholar]
- Wu, H.; Gu, X. Towards dropout training for convolutional neural networks. Neural Netw. 2015, 71, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Obuchowski, N.A.; Bullen, J.A. Receiver operating characteristic (ROC) curves: Review of methods with applications in diagnostic medicine. Phys. Med. Biol. 2018, 63, 07TR01. [Google Scholar] [CrossRef] [PubMed]
- Han, K.; Wang, Y.; Chen, H.; Chen, X.; Guo, J.; Liu, Z.; Tang, Y.; Xiao, A.; Xu, C.; Xu, Y.; et al. A survey on vision transformer. IEEE Trans. Pattern Anal. Mach. Intell. 2023, 45, 87–110. [Google Scholar] [CrossRef] [PubMed]
- Majidpour, J.; Ahmed, A.M.; Hassan, B.A.; Abdalla, M.H.; Qader, S.M.; Tayfor, N.B.; Rashid, T.A. Thermal face image reidentification based on variational autoencoder, cascade object detector using lightweight architectures. In Practical Artificial Intelligence for Internet of Medical Things; CRC Press: Boca Raton, FL, USA, 2023; pp. 291–303. [Google Scholar]
- Creswell, A.; White, T.; Dumoulin, V.; Arulkumaran, K.; Sengupta, B.; Bharath, A.A. Generative adversarial networks: An overview. IEEE Signal Process. Mag. 2018, 35, 53–65. [Google Scholar] [CrossRef]
- Wang, K.; Gou, C.; Duan, Y.; Lin, Y.; Zheng, X. Generative adversarial networks: Introduction and outlook. IEEE/CAA J. Autom. Sin. 2017, 4, 588–598. [Google Scholar] [CrossRef]

| Study | Diagnosis | Number of Subjects | Region of Interest | Detected Temperature Difference (°C) |
|---|---|---|---|---|
| [32] | Bone neoplasia | 40 | Limb | 0.53 to 0.67 |
| [33] | Fracture | 19 | Forearm | 0.8 to 2 |
| [34] | Repetitive strain injuries | 33 | Wrist and hand Joints | 0.65 |
| [35] | Fracture/sprain | 133 | Various | 0.26 to 0.47 |
| [31] | Toddler’s fracture | 39 | Tibia | 1.1 |
| Study | Number of Subjects | Diagnosis | ML/AI Algorithm | Accuracy (%) |
|---|---|---|---|---|
| [49] | 283 | Diabetic eye disease | SVM | 86 |
| [50] | 11 | Lip lesions | SVM/KNN | 63–100 |
| [51] | 67 | Breast cancer | Various | 73–92 |
| [52] | 82 | Pressure injury | CNN | 95 |
| Layer Name | Parameters |
|---|---|
| 2D convolutional (Conv_1 (2D)) | 80 |
| Batch normalization (Batchnorm_1 (2D)) | 16 |
| 2D Convolutional (Conv_2 (2D)) | 1168 |
| Batch Normalization (Batchnorm_2 (2D)) | 32 |
| 2D Convolutional (Conv_3 (2D)) | 4640 |
| Batch Normalization (Batchnorm_3 (2D)) | 64 |
| 2D Convolutional (Conv_4 (2D)) | 18,496 |
| Batch Normalization (Batchnorm_4 (2D)) | 128 |
| 2D Convolutional (Conv_5 (2D)) | 73,856 |
| Batch normalization (Batchnorm_5 (2D)) | 256 |
| 2D Convolutional (Conv_6 (2D)) | 295,168 |
| Batch Normalization (Batchnorm_6 (2D)) | 512 |
| 2D Convolutional (Conv_7 (2D)) | 1,180,160 |
| Batch normalization (Batchnorm_7 (2D)) | 1024 |
| 2D Convolutional (Conv_8 (2D)) | 1,180,160 |
| Batch normalizations (Batchnorm_8 (2D)) | 1024 |
| Fully Connected (FC) | 1026 |
| Hyperparameters | Measures |
|---|---|
| Optimizer | Adam |
| Activation function | ReLU |
| Learning rate | 0.005 |
| Batch size | 170 |
| Maximum epoch | 250 (no early stopping) |
| Maximum iteration | 12,250 |
| Loss function | Cross-Entropy |
| Injury | Training | Validation |
|---|---|---|
| Fracture | 3887 | 1794 |
| Sprain | 4485 | 1794 |
| Injury Type | Sensitivity (%) | Specificity (%) | NPV (%) | PPV (%) | Accuracy (%) |
|---|---|---|---|---|---|
| Fracture versus sprain | 88.3 | 68.3 | 80.4 | 72.5 | 75.8 |
| Injury | Sensitivity (%) | Specificity (%) | NPV (%) | PPV (%) | Accuracy (%) |
|---|---|---|---|---|---|
| Fracture versus sprain | 50 | 66.7 | 60.0 | 57.1 | 58.3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shobayo, O.; Saatchi, R.; Ramlakhan, S. Convolutional Neural Network to Classify Infrared Thermal Images of Fractured Wrists in Pediatrics. Healthcare 2024, 12, 994. https://doi.org/10.3390/healthcare12100994
Shobayo O, Saatchi R, Ramlakhan S. Convolutional Neural Network to Classify Infrared Thermal Images of Fractured Wrists in Pediatrics. Healthcare. 2024; 12(10):994. https://doi.org/10.3390/healthcare12100994
Chicago/Turabian StyleShobayo, Olamilekan, Reza Saatchi, and Shammi Ramlakhan. 2024. "Convolutional Neural Network to Classify Infrared Thermal Images of Fractured Wrists in Pediatrics" Healthcare 12, no. 10: 994. https://doi.org/10.3390/healthcare12100994
APA StyleShobayo, O., Saatchi, R., & Ramlakhan, S. (2024). Convolutional Neural Network to Classify Infrared Thermal Images of Fractured Wrists in Pediatrics. Healthcare, 12(10), 994. https://doi.org/10.3390/healthcare12100994








