Chronic Training Induces Metabolic and Proteomic Response in Male and Female Basketball Players: Salivary Modifications during In-Season Training Programs
Abstract
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Saliva Samples Collection and Preparation
2.3. Two-Dimensional Polyacrylamide Gel Electrophoresis (2DE)
2.4. Protein Identification by Mass Spectrometry
2.4.1. In Situ Digestion of 2DE Spots
2.4.2. LC–MS/MS Analysis
2.5. Western Blot Analysis
2.6. Analysis of Salivary Samples by Gas Chromatography–Mass Spectrometry (GC–MS)
2.7. Image and Statistical Analysis
3. Results
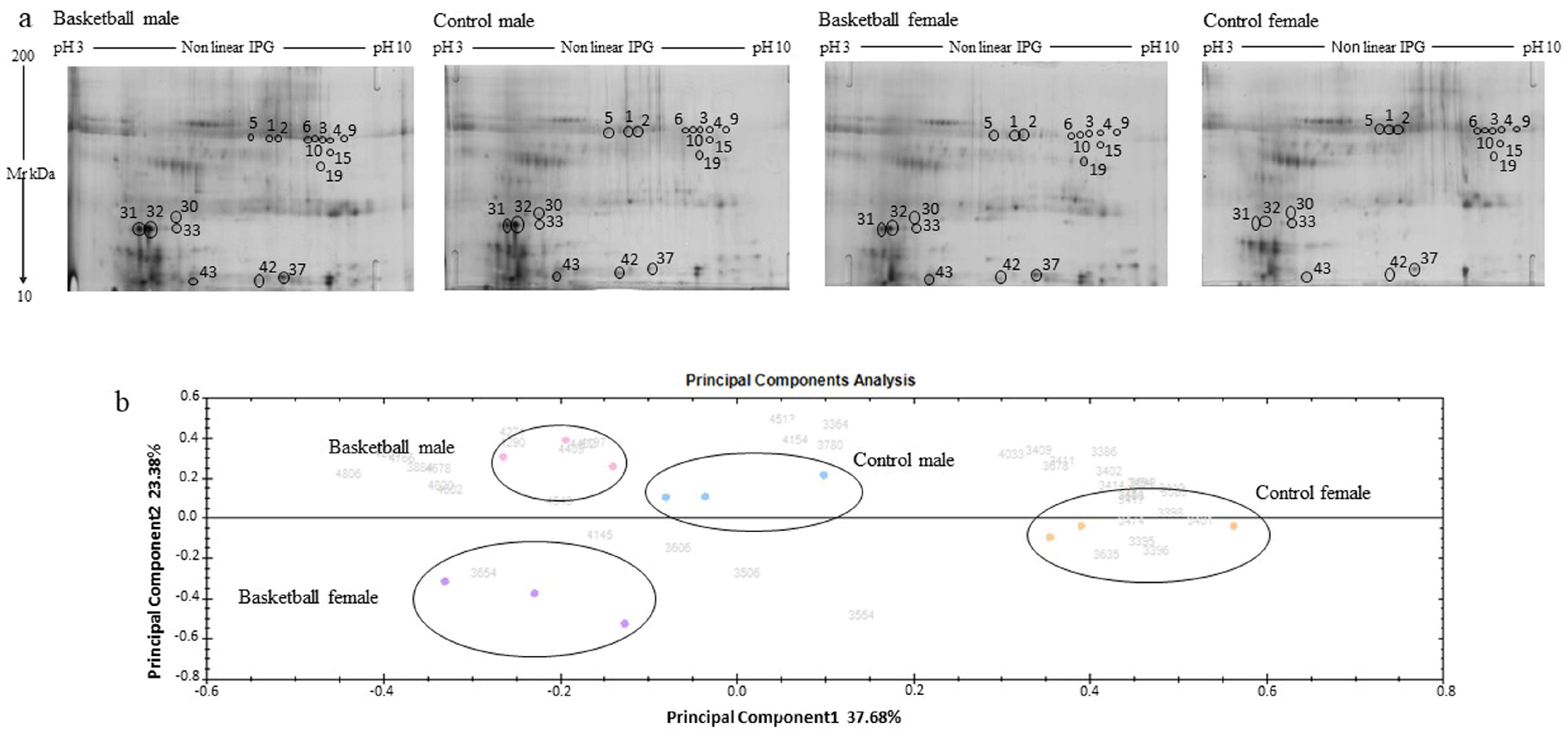
3.1. Overview of Saliva Protein Profiles
3.2. Principal Component Analysis
3.3. Proteins Differentially Modulated by Exercise and Sex
| Spot No. a | AC b | Gene Name | Protein Name | Score c | Protein Mass | Protein Cover d | Tukey’s Test e/Fold Change f | |||
|---|---|---|---|---|---|---|---|---|---|---|
| BM vs. CM | BF vs. CF | BM vs. BF | CM vs. CF | |||||||
| 1 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 791 | 56,484 | 52.2 | ns | */−2.6 | ns | ns |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha 1 | 243 | 38,486 | 22.9 | ns | */−2.6 | ns | ns | |
| P01877 | IGHA2 | Immunoglobulin heavy constant alpha 2 | 170 | 37,366 | 19 | ns | */−2.6 | ns | ns | |
| 2 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 628 | 56,484 | 31.9 | ns | */−2.6 | ns | ns |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha | 141 | 38,486 | 14.7 | ns | */−2.6 | ns | ns | |
| P01877 | IGHA2 | Immunoglobulin heavy constant alpha 2 | 121 | 37,366 | 13.9 | ns | */−2.6 | ns | ns | |
| 3 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 1120 | 56,484 | 52.8 | */−2.9 | **/−2.6 | ns | */−1.7 |
| P02452 | COL1A1 | Collagen alpha-1(I) chain | 117 | 37,077 | 11.3 | */−2.9 | **/−2.6 | ns | */−1.7 | |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha 1 | 223 | 38,486 | 20.2 | */−2.9 | **/−2.6 | ns | */−1.7 | |
| P01877 | IGHA2 | Immunoglobulin heavy constant alpha 2 | 176 | 37,366 | 16.2 | */−2.9 | **/−2.6 | ns | */−1.7 | |
| 4 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 1073 | 56,484 | 57.1 | ns | **/−3.3 | ns | ns |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha 1 | 194 | 38,486 | 16.6 | ns | **/−3.3 | ns | ns | |
| P01877 | IGHA2 | Immunoglobulin heavy constant alpha 2 | 143 | 37,366 | 12.6 | ns | **/−3.3 | ns | ns | |
| A0N4V7 | Tcr-alpha | putative T-cell receptor beta | 39 | 2269 | 38.1 | ns | **/−3.3 | ns | ns | |
| 5 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 478 | 56,484 | 30 | **/1.4 | **/−1.5 | ****/1.9 | ns |
| 6 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 1011 | 56,484 | 48.8 | ns | */−2.4 | ns | ns |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha 1 | 254 | 38,486 | 25.7 | ns | */−2.4 | ns | ns | |
| P01877 | IGHA2 | Immunoglobulin heavy constant alpha 2 | 212 | 37,366 | 21.9 | ns | */−2.4 | ns | ns | |
| 9 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 904 | 58,415 | 43.2 | ns | */−3.2 | ns | ns |
| P68871 | HBB | Hemoglobin subunit beta | 147 | 16,102 | 35.4 | ns | */−3.2 | ns | ns | |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha 1 | 240 | 38,486 | 21.5 | ns | */−3.2 | ns | ns | |
| P01877 | IGHA2 | Immunoglobulin heavy constant alpha 2 | 179 | 37,366 | 16.2 | ns | */−3.2 | ns | ns | |
| 10 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 911 | 58,415 | 41.1 | */−2.3 | ns | ns | ns |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha 1 | 211 | 38,486 | 17.3 | */−2.3 | ns | ns | ns | |
| P01877 | IGHA2 | Immunoglobulin heavy constant alpha 2 | 175 | 37,366 | 16.2 | */−2.3 | ns | ns | ns | |
| 15 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 375 | 58,415 | 18 | ns | */−2.7 | ns | ns |
| P25311 | AZGP1 | Zinc-alpha-2-glycoprotein | 390 | 34,465 | 37.6 | ns | */−2.7 | ns | ns | |
| Q01469 | FABP5 | Fatty acid–binding protein 5 | 147 | 15,497 | 28.1 | ns | */−2.7 | ns | ns | |
| 19 | P0DTE8 | AMY1A | Alpha-Amylase 1A | 831 | 58,415 | 44.2 | ns | **/−2 | ns | **/−2.2 |
| 30 | P02768 | ALB | Albumin | 269 | 71,317 | 15.1 | ns | ns | ns | */2.1 |
| P07355 | ANXA2 | Annexin A2 | 249 | 38,808 | 20.6 | ns | ns | ns | */2.1 | |
| P81605 | DCD | Dermcidin | 208 | 11,391 | 38.2 | ns | ns | ns | */2.1 | |
| 31 | P01040 | CSTA | Cystatin A | 246 | 11,000 | 98 | ns | ns | ns | */3.8 |
| P81605 | DCD | Dermcidin | 177 | 11,391 | 35.5 | ns | ns | ns | */3.8 | |
| 32 | P01040 | CSTA | Cystatin A | 348 | 11,000 | 94.9 | ns | ns | ns | */4.3 |
| P81605 | DCD | Dermcidin | 73 | 11,391 | 30.9 | ns | ns | ns | */4.3 | |
| 33 | P02768 | ALB | Albumin | 675 | 71,317 | 28.1 | ns | ns | **/2.7 | ns |
| P81605 | DCD | Dermcidin | 150 | 11,391 | 30.9 | ns | ns | **/2.7 | ns | |
| P01876 | IGHA1 | Immunoglobulin heavy constant alpha 1 | 214 | 38,486 | 15.3 | ns | ns | **/2.7 | ns | |
| 42 | P02768 | ALB | Albumin | 190 | 68,408 | 20.5 | ns | ns | ns | */2.1 |
| P31151 | S100A7 | Psoriasin | 115 | 11,564 | 62.4 | ns | ns | ns | */2.1 | |
| 43 | P81605 | DCD | Dermcidin | 143 | 11,391 | 37.3 | ns | ns | ns | */3 |
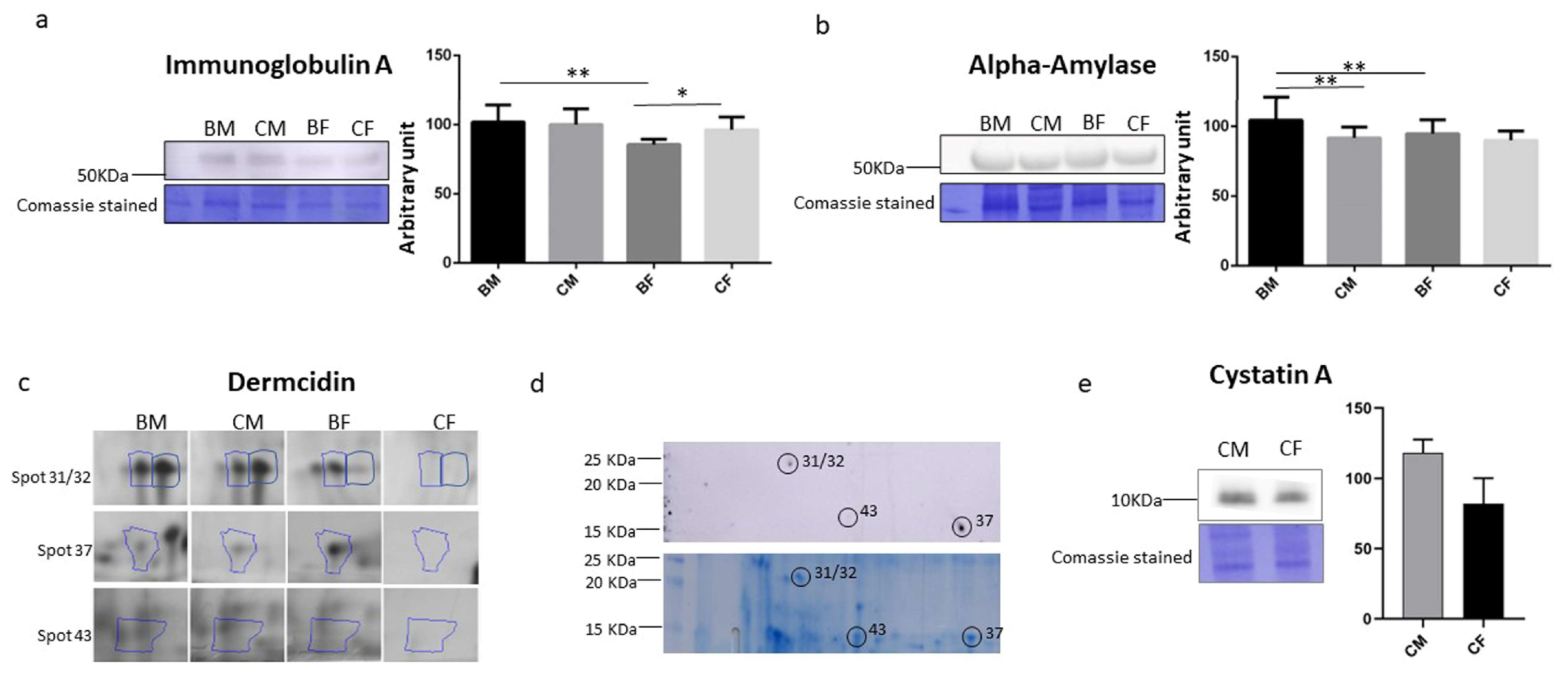
3.4. Saliva IgA, Alpha-Amylase, Dermcidin, and Cystatin a Determination
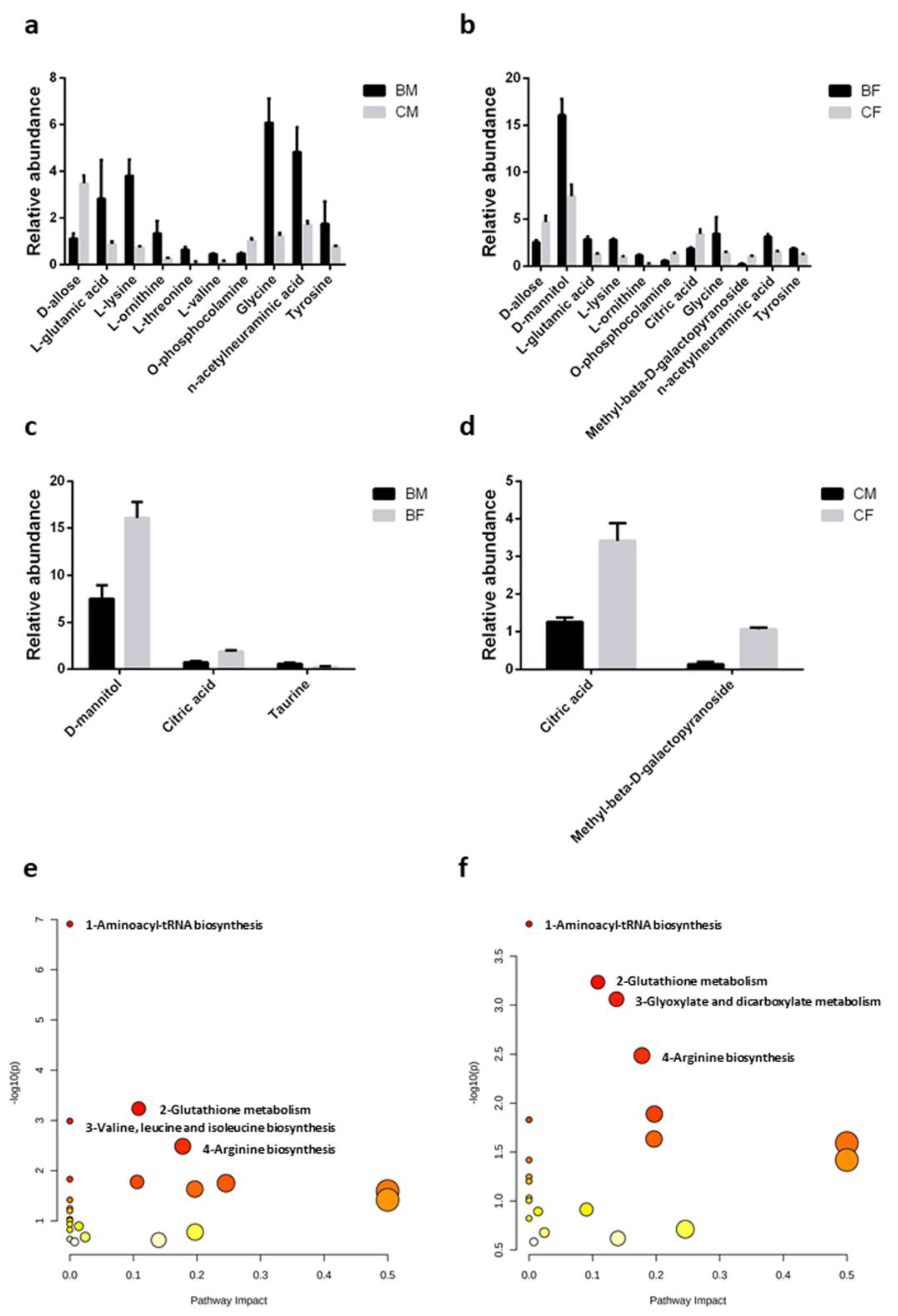
3.5. Metabolites Differentially Modulated by Exercise and by Sex
- (i)
- (ii)
- Glutathione metabolism—for both males and females, glycine, L-glutamate, and L-ornithine (number 2 in Figure 3e,f)
- (iii)
- Arginine biosynthesis; for both males and females, L-glutamate and L-ornithine (number 4 in Figure 3e,f).
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Finsterer, J. Biomarkers of peripheral muscle fatigue during exercise. BMC Musculoskelet. Disord. 2012, 13, 218. [Google Scholar] [CrossRef] [PubMed]
- Williams, M.A.; Haskell, W.L.; Ades, P.A.; Amsterdam, E.A.; Bittner, V.; Franklin, B.A.; Gulanick, M.; Laing, S.T.; Stewart, K.J. Resistance Exercise in Individuals with and without Cardiovascular Disease: 2007 Update. Circulation 2007, 116, 572–584. [Google Scholar] [CrossRef] [PubMed]
- Luti, S.; Modesti, A.; Modesti, P.A. Inflammation, peripheral signals and redox homeostasis in athletes who practice different sports. Antioxidants 2020, 9, 1065. [Google Scholar] [CrossRef] [PubMed]
- Hwang, Y.; Park, J.; Lim, K. Effects of Pilates Exercise on Salivary Secretory Immunoglobulin A Levels in Older Women. J. Aging Phys. Act. 2016, 24, 399–406. [Google Scholar] [CrossRef]
- Lindsay, A.; Costello, J.T. Realising the Potential of Urine and Saliva as Diagnostic Tools in Sport and Exercise Medicine. Sport. Med. 2017, 47, 11–31. [Google Scholar] [CrossRef]
- Franco-Martínez, L.; González-Hernández, J.M.; Horvatić, A.; Guillemin, N.; Cerón, J.J.; Martínez-Subiela, S.; Sentandreu, M.Á.; Brkljačić, M.; Mrljak, V.; Tvarijonaviciute, A.; et al. Differences on salivary proteome at rest and in response to an acute exercise in men and women: A pilot study. J. Proteom. 2020, 214, 103629. [Google Scholar] [CrossRef]
- Papacosta, E.; Nassis, G.P. Saliva as a tool for monitoring steroid, peptide and immune markers in sport and exercise science. J. Sci. Med. Sport 2011, 14, 424–434. [Google Scholar] [CrossRef]
- Ntovas, P.; Loumprinis, N.; Maniatakos, P.; Margaritidi, L.; Rahiotis, C. The Effects of Physical Exercise on Saliva Composition: A Comprehensive Review. Dent. J. 2022, 10, 7. [Google Scholar] [CrossRef]
- Carpenter, G.H. The Secretion, Components, and Properties of Saliva. Annu. Rev. Food Sci. Technol. 2013, 4, 267–276. [Google Scholar] [CrossRef]
- Miller, C.S.; Foley, J.D.; Bailey, A.L.; Campell, C.L.; Humphries, R.L.; Christodoulides, N.; Floriano, P.N.; Simmons, G.; Bhagwandin, B.; Jacobson, J.W.; et al. Current developments in salivary diagnostics. Biomark. Med. 2010, 4, 171–189. [Google Scholar] [CrossRef]
- Kaufman, E.; Lamster, I.B. The diagnostic applications of saliva—A review. Crit. Rev. Oral Biol. Med. 2002, 13, 197–212. [Google Scholar] [CrossRef] [PubMed]
- Jasim, H.; Olausson, P.; Hedenberg-Magnusson, B.; Ernberg, M.; Ghafouri, B. The proteomic profile of whole and glandular saliva in healthy pain-free subjects. Sci. Rep. 2016, 6, 39073. [Google Scholar] [CrossRef] [PubMed]
- Fleissig, Y.; Reichenberg, E.; Redlich, M.; Zaks, B.; Deutsch, O.; Aframian, D.J.; Palmon, A. Comparative proteomic analysis of human oral fluids according to gender and age. Oral Dis. 2010, 16, 831–838. [Google Scholar] [CrossRef] [PubMed]
- Soares Nunes, L.A.; Mussavira, S.; Sukumaran Bindhu, O. Clinical and diagnostic utility of saliva as a non-invasive diagnostic fluid: A systematic review. Biochem. Med. 2015, 25, 177–192. [Google Scholar] [CrossRef]
- Pitti, E.; Petrella, G.; Di Marino, S.; Summa, V.; Perrone, M.; D’Ottavio, S.; Bernardini, A.; Cicero, D.O. Salivary Metabolome and Soccer Match: Challenges for Understanding Exercise induced Changes. Metabolites 2019, 9, 141. [Google Scholar] [CrossRef]
- Pinto, G.; Militello, R.; Amoresano, A.; Modesti, P.A.; Modesti, A.; Luti, S. Relationships between Sex and Adaptation to Physical Exercise in Young Athletes: A Pilot Study. Healthcare 2022, 10, 358. [Google Scholar] [CrossRef]
- Militello, R.; Pinto, G.; Illiano, A.; Luti, S.; Magherini, F.; Amoresano, A.; Modesti, P.A.; Modesti, A. Modulation of Plasma Proteomic Profile by Regular Training in Male and Female Basketball Players: A Preliminary Study. Front. Physiol. 2022, 13, 813447. [Google Scholar] [CrossRef]
- Costill, D.L.; Daniels, J.; Evans, W.; Fink, W.; Krahenbuhl, G.; Saltin, B. Skeletal muscle enzymes and fiber composition in male and female track athletes. J. Appl. Physiol. 1976, 40, 149–154. [Google Scholar] [CrossRef]
- Bongiovanni, T.; Pintus, R.; Dessì, A.; Noto, A.; Sardo, S.; Finco, G.; Corsello, G.; Fanos, V. Sportomics: Metabolomics applied to sports. The new revolution? Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 11011–11019. [Google Scholar] [CrossRef]
- Hagströmer, M.; Oja, P.; Sjöström, M. The International Physical Activity Questionnaire (IPAQ): A study of concurrent and construct validity. Public Health Nutr. 2006, 9, 755–762. [Google Scholar] [CrossRef]
- Guidi, F.; Magherini, F.; Gamberi, T.; Bini, L.; Puglia, M.; Marzocchini, R.; Ranaldi, F.; Modesti, P.A.; Gulisano, M.; Modesti, A. Plasma protein carbonylation and physical exercise. Mol. Biosyst. 2011, 7, 640–650. [Google Scholar] [CrossRef] [PubMed]
- Pietrovito, L.; Cano-Cortés, V.; Gamberi, T.; Magherini, F.; Bianchi, L.; Bini, L.; Sánchez-Martín, R.M.; Fasano, M.; Modesti, A. Cellular response to empty and palladium-conjugated amino-polystyrene nanospheres uptake: A proteomic study. Proteomics 2015, 15, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Luti, S.; Fiaschi, T.; Magherini, F.; Modesti, P.A.; Piomboni, P.; Governini, L.; Luddi, A.; Amoresano, A.; Illiano, A.; Pinto, G.; et al. Relationship between the metabolic and lipid profile in follicular fluid of women undergoing in vitro fertilization. Mol. Reprod. Dev. 2020, 87, 986–997. [Google Scholar] [CrossRef] [PubMed]
- Sumner, L.W.; Amberg, A.; Barrett, D.; Beale, M.H.; Beger, R.; Daykin, C.A.; Fan, T.W.-M.; Fiehn, O.; Goodacre, R.; Griffin, J.L.; et al. Proposed minimum reporting standards for chemical analysis. Metabolomics 2007, 3, 211–221. [Google Scholar] [CrossRef]
- Kind, T.; Wohlgemuth, G.; Lee, D.Y.; Lu, Y.; Palazoglu, M.; Shahbaz, S.; Fiehn, O. FiehnLib: Mass Spectral and Retention Index Libraries for Metabolomics Based on Quadrupole and Time-of-Flight Gas Chromatography/Mass Spectrometry. Anal. Chem. 2009, 81, 10038–10048. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Harb, J.; Wilson, B.S.; Hermouet, S. Editorial: Structure, Isotypes, Targets, and Post-translational Modifications of Immunoglobulins and Their Role in Infection, Inflammation and Autoimmunity. Front. Immunol. 2020, 11, 1761. [Google Scholar] [CrossRef]
- Hirtz, C.; Chevalier, F.; Centeno, D.; Rofidal, V.; Egea, J.-C.; Rossignol, M.; Sommerer, N.; Deville de Périère, D. MS characterization of multiple forms of alpha-amylase in human saliva. Proteomics 2005, 5, 4597–4607. [Google Scholar] [CrossRef]
- Bank, R.A.; Hettema, E.H.; Arwert, F.; Amerongen, A.V.N.; Pronk, J.C. Electrophoretic characterization of posttranslational modifications of human parotid salivary α-amylase. Electrophoresis 1991, 12, 74–79. [Google Scholar] [CrossRef]
- Schittek, B. The multiple facets of dermcidin in cell survival and host defense. J. Innate Immun. 2012, 4, 349–360. [Google Scholar] [CrossRef]
- Esposito, G.; Schiattarella, G.G.; Perrino, C.; Cattaneo, F.; Pironti, G.; Franzone, A.; Gargiulo, G.; Magliulo, F.; Serino, F.; Carotenuto, G.; et al. Dermcidin: A skeletal muscle myokine modulating cardiomyocyte survival and infarct size after coronary artery ligation. Cardiovasc. Res. 2015, 107, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Puigarnau, S.; Fernàndez, A.; Obis, E.; Jové, M.; Castañer, M.; Pamplona, R.; Portero-Otin, M.; Camerino, O. Metabolomics reveals that fittest trail runners show a better adaptation of bioenergetic pathways. J. Sci. Med. Sport 2022, 25, 425–431. [Google Scholar] [CrossRef] [PubMed]
- McKetney, J.; Jenkins, C.C.; Minogue, C.; Mach, P.M.; Hussey, E.K.; Glaros, T.G.; Coon, J.; Dhummakupt, E.S. Proteomic and metabolomic profiling of acute and chronic stress events associated with military exercises. Mol. Omi. 2022, 18, 279–295. [Google Scholar] [CrossRef]
- Tomasi, T.B.; Trudeau, F.B.; Czerwinski, D.; Erredge, S. Immune parameters in athletes before and after strenuous exercise. J. Clin. Immunol. 1982, 2, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Walsh, N.P.; Bishop, N.C.; Blackwell, J.; Wierzbicki, S.G.; Montague, J.C. Salivary IgA response to prolonged exercise in a cold environment in trained cyclists. Med. Sci. Sports Exerc. 2002, 34, 1632–1637. [Google Scholar] [CrossRef]
- McDowell, S.L.; Chaloa, K.; Housh, T.J.; Tharp, G.D.; Johnson, G.O. The effect of exercise intensity and duration on salivary immunoglobulin A. Eur. J. Appl. Physiol. Occup. Physiol. 1991, 63, 108–111. [Google Scholar] [CrossRef]
- Allgrove, J.E.; Gomes, E.; Hough, J.; Gleeson, M. Effects of exercise intensity on salivary antimicrobial proteins and markers of stress in active men. J. Sports Sci. 2008, 26, 653–661. [Google Scholar] [CrossRef]
- Gleeson, M. Mucosal immunity and respiratory illness in elite athletes. Int. J. Sports Med. 2000, 21 (Suppl. S1), S33–S43. [Google Scholar] [CrossRef]
- Hackney, A.C.; Koltun, K.J. The immune system and overtraining in athletes: Clinical implications. Acta Clin. Croat. 2012, 51, 633–641. [Google Scholar]
- Nicolette, C.B.; Bishop, N.C.; Gleeson, M. Acute and chronic effects of exercise on markers of mucosal immunity. Front. Biosci. 2009, 14, 4444–4456. [Google Scholar] [CrossRef]
- Halson, S.L.; Jeukendrup, A.E. Does overtraining exist? An analysis of overreaching and overtraining research. Sports Med. 2004, 34, 967–981. [Google Scholar] [CrossRef] [PubMed]
- Kivlighan, K.T.; Granger, D.A. Salivary alpha-amylase response to competition: Relation to gender, previous experience, and attitudes. Psychoneuroendocrinology 2006, 31, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Alix-Sy, D.; Le Scanff, C.; Filaire, E. Psychophysiological responses in the pre-competition period in elite soccer players. J. Sports Sci. Med. 2008, 7, 446–454. [Google Scholar] [PubMed]
- Cadegiani, F.A.; Kater, C.E. Hormonal aspects of overtraining syndrome: A systematic review. BMC Sports Sci. Med. Rehabil. 2017, 9, 14. [Google Scholar] [CrossRef]
- de Oliveira, V.N.; Bessa, A.; Lamounier, R.P.M.S.; de Santana, M.G.; de Mello, M.T.; Espindola, F.S. Changes in the Salivary Biomarkers Induced by an Effort Test. Int. J. Sports Med. 2010, 31, 377–381. [Google Scholar] [CrossRef]
- Lee, D.Y.; Kim, E.; Choi, M.H. Technical and clinical aspects of cortisol as a biochemical marker of chronic stress. BMB Rep. 2015, 48, 209–216. [Google Scholar] [CrossRef]
- Nater, U.M.; Rohleder, N. Salivary alpha-amylase as a non-invasive biomarker for the sympathetic nervous system: Current state of research. Psychoneuroendocrinology 2009, 34, 486–496. [Google Scholar] [CrossRef]
- Sant’Anna, M.d.L.; Oliveira, L.T.; Gomes, D.V.; Marques, S.T.F.; Provance, D.W.; Sorenson, M.M.; Salerno, V.P. Physical exercise stimulates salivary secretion of cystatins. PLoS ONE 2019, 14, e0224147. [Google Scholar] [CrossRef]
- Jain, S.; Ahmad, Y.; Bhargava, K. Salivary proteome patterns of individuals exposed to High Altitude. Arch. Oral Biol. 2018, 96, 104–112. [Google Scholar] [CrossRef]
- Khodagholi, F.; Zareh Shahamati, S.; Maleki Chamgordani, M.; Mousavi, M.A.; Moslemi, M.; Salehpour, M.; Rafiei, S.; Foolad, F. Interval aerobic training improves bioenergetics state and mitochondrial dynamics of different brain regions in restraint stressed rats. Mol. Biol. Rep. 2021, 48, 2071–2082. [Google Scholar] [CrossRef]
- Sugino, T.; Shirai, T.; Kajimoto, Y.; Kajimoto, O. L-ornithine supplementation attenuates physical fatigue in healthy volunteers by modulating lipid and amino acid metabolism. Nutr. Res. 2008, 28, 738–743. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Carmona, Y.; Meijer, J.L.; Zhou, Y.; Jansen, E.C.; Perng, W.; Banker, M.; Song, P.X.K.; Téllez-Rojo, M.M.; Cantoral, A.; Peterson, K.E. Metabolomics reveals sex-specific pathways associated with changes in adiposity and muscle mass in a cohort of Mexican adolescents. Pediatr. Obes. 2022, 17, e12887. [Google Scholar] [CrossRef]
- Low, S.; Wang, J.; Moh, A.; Ang, S.F.; Ang, K.; Shao, Y.-M.; Ching, J.; Wee, H.N.; Lee, L.S.; Kovalik, J.-P.; et al. Amino acid profile of skeletal muscle loss in type 2 diabetes: Results from a 7-year longitudinal study in asians. Diabetes Res. Clin. Pract. 2022, 186, 109803. [Google Scholar] [CrossRef] [PubMed]
- Janssen, I.; Heymsfield, S.B.; Wang, Z.M.; Ross, R. Skeletal muscle mass and distribution in 468 men and women aged 18–88 yr. J. Appl. Physiol. 2000, 89, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, K.; Ishizu, Y.; Honda, T.; Ito, T.; Imai, N.; Nakamura, M.; Kawashima, H.; Kitaura, Y.; Ishigami, M.; Fujishiro, M. Patients with low muscle mass have characteristic microbiome with low potential for amino acid synthesis in chronic liver disease. Sci. Rep. 2022, 12, 3674. [Google Scholar] [CrossRef] [PubMed]
- McNulty, K.L.; Elliott-Sale, K.J.; Dolan, E.; Swinton, P.A.; Ansdell, P.; Goodall, S.; Thomas, K.; Hicks, K.M. The Effects of Menstrual Cycle Phase on Exercise Performance in Eumenorrheic Women: A Systematic Review and Meta-Analysis. Sport. Med. 2020, 50, 1813–1827. [Google Scholar] [CrossRef]
| Characteristics | Mean (SD) | Tukey’s Test a | ||||||
|---|---|---|---|---|---|---|---|---|
| BM | CM | BF | CF | BM vs. CM | BF vs. CF | BM vs. BF | CM vs. CF | |
| Age (year) | 21 ± 2.2 | 26.1 ± 4.1 | 25.1 ± 5.5 | 26.9 ± 2.2 | 0.01 * | 0.77 | 0.08 | 0.97 |
| Weight (kg) | 81.5 ± 10.2 | 73 ± 8.7 | 68.7 ± 11.9 | 58.7 ± 5.8 | 0.17 | 0.17 | 0.03 * | 0.02 * |
| Height (cm) | 186 ± 0.06 | 178.7 ± 0.06 | 175.6 ± 0.08 | 163.4 ± 0.06 | 0.06 | 0.003 ** | 0.006 ** | <0.0001 **** |
| BMI (kg/m2) | 23.6 ± 2.7 | 22.9 ± 2.9 | 22.1 ± 2.05 | 22 ± 2.3 | 0.64 | 1 | 0.18 | 0.72 |
| Spot No. a | Accession No. b | Description | Coverage (%) c | Unique Peptides d | Score Mascot e | Score Sequest HT f |
|---|---|---|---|---|---|---|
| 31–32 | P81605 | Dermcidin | 21 | 2 | 64 | 2.22 |
| 37 | P81605 | Dermcidin | 21 | 2 | 70 | 5.09 |
| 43 | P81605 | Dermcidin | 25 | 4 | 147 | 14.15 |
| Compound Name | CAS Number * | Formula | KEGG ID ° | Fold Change/Tukey’s Test | |||
|---|---|---|---|---|---|---|---|
| BM vs. CM | BF vs. CF | BM vs. BF | CM vs. CF | ||||
| D-allose | 579-36-2 | C6H12O6 | C01487 | −3.1/*** | −1.8/* | ns | ns |
| D-mannitol | 87-78-5 | C6H12O6 | C00392 | ns | 2.4/*** | −2.1/*** | ns |
| L-glutamic acid | 56-86-0 | C5H9NO4 | C00025 | 3.2/**** | 2.2/*** | ns | ns |
| L-lysine | 56-87-1 | C6H14N2O2 | C00047 | 5.0/**** | 3.0/* | ns | ns |
| L-ornithine | 70-26-8 | C5H12N2O2 | C00077 | 5.2/** | 4.0/* | ns | ns |
| L-threonine | 72-19-5 | C4H9NO3 | C00188 | 8.9/** | ns | ns | ns |
| L-valine | 72-18-4 | C5HNO2 | C00183 | 5.0/* | ns | ns | ns |
| O-phosphocolamine | 1071-23-4 | C2H8NO4P | C00346 | −2.2/**** | −2.3/** | ns | ns |
| Citric acid | 5949-29-1 | C6H8O7 | C00158 | ns | −1.8/** | −2.5/* | −2.7/*** |
| Glycine | 56-40-6 | C2H5NO2 | C00037 | 4.9/* | 2.5/* | ns | ns |
| Methyl-beta-D-galactopyranoside | 1824-94-8 | C7H14O6 | C03619 | ns | −5.8/**** | ns | −8.2/**** |
| N-acetylneuraminic acid | 131-48-6 | C11H19NO9 | C00270 | 2.8/* | 2.0/* | ns | ns |
| Taurine | 107-35-7 | C2H7NO3S | C00245 | ns | ns | 2.6/* | ns |
| Tyrosine | 60-18-4 | C9H11NO3 | C00082 | 2.3/** | 1.6/* | ns | ns |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luti, S.; Militello, R.; Pinto, G.; Illiano, A.; Amoresano, A.; Chiappetta, G.; Marzocchini, R.; Modesti, P.A.; Pratesi, S.; Pazzagli, L.; et al. Chronic Training Induces Metabolic and Proteomic Response in Male and Female Basketball Players: Salivary Modifications during In-Season Training Programs. Healthcare 2023, 11, 241. https://doi.org/10.3390/healthcare11020241
Luti S, Militello R, Pinto G, Illiano A, Amoresano A, Chiappetta G, Marzocchini R, Modesti PA, Pratesi S, Pazzagli L, et al. Chronic Training Induces Metabolic and Proteomic Response in Male and Female Basketball Players: Salivary Modifications during In-Season Training Programs. Healthcare. 2023; 11(2):241. https://doi.org/10.3390/healthcare11020241
Chicago/Turabian StyleLuti, Simone, Rosamaria Militello, Gabriella Pinto, Anna Illiano, Angela Amoresano, Giovanni Chiappetta, Riccardo Marzocchini, Pietro Amedeo Modesti, Simone Pratesi, Luigia Pazzagli, and et al. 2023. "Chronic Training Induces Metabolic and Proteomic Response in Male and Female Basketball Players: Salivary Modifications during In-Season Training Programs" Healthcare 11, no. 2: 241. https://doi.org/10.3390/healthcare11020241
APA StyleLuti, S., Militello, R., Pinto, G., Illiano, A., Amoresano, A., Chiappetta, G., Marzocchini, R., Modesti, P. A., Pratesi, S., Pazzagli, L., Modesti, A., & Gamberi, T. (2023). Chronic Training Induces Metabolic and Proteomic Response in Male and Female Basketball Players: Salivary Modifications during In-Season Training Programs. Healthcare, 11(2), 241. https://doi.org/10.3390/healthcare11020241








