Accelerating the Front End of Medicine: Three Digital Use Cases and HCI Implications
Abstract
1. Introduction
2. Conceptual Framework
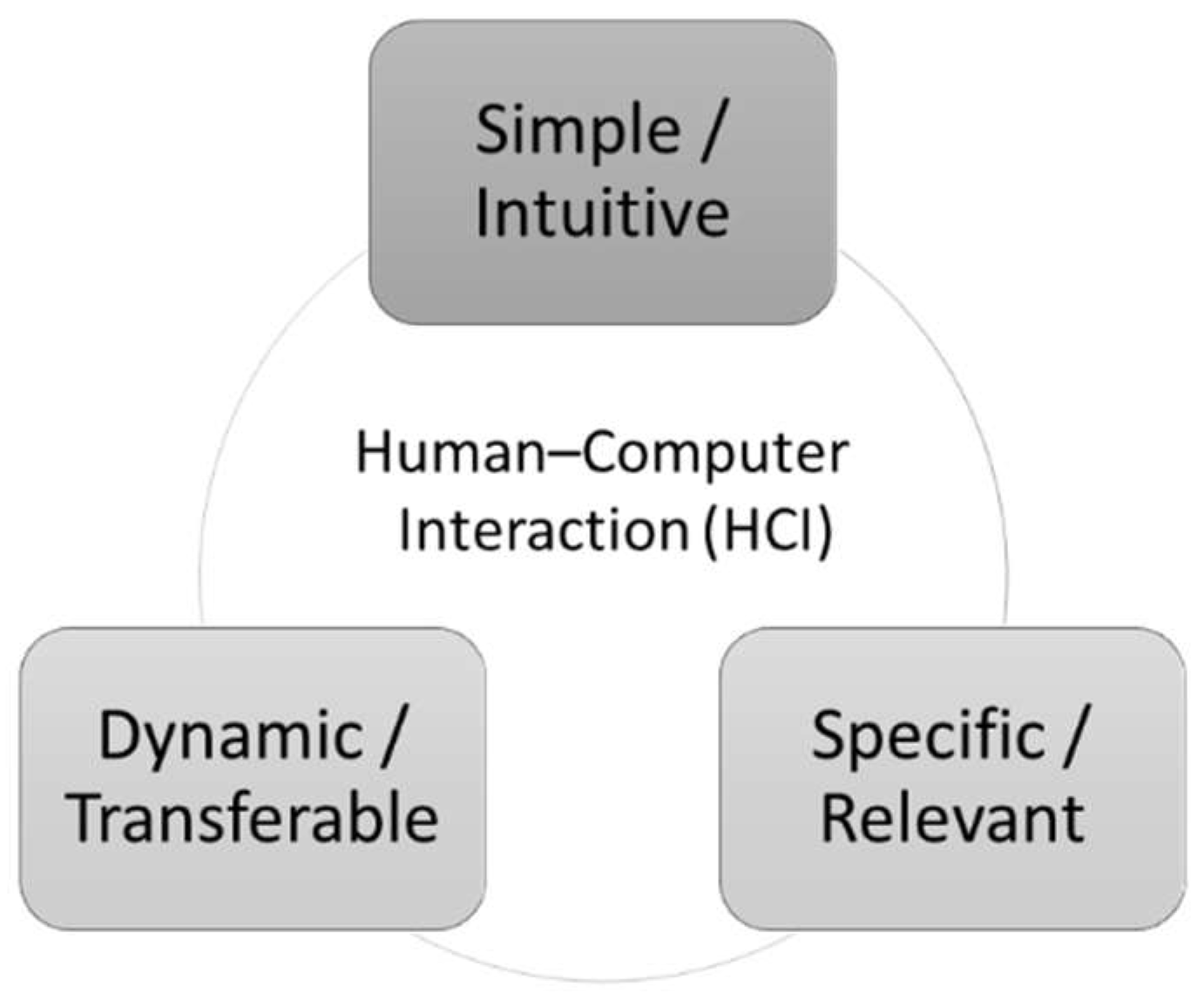
2.1. Human-Computer Interaction
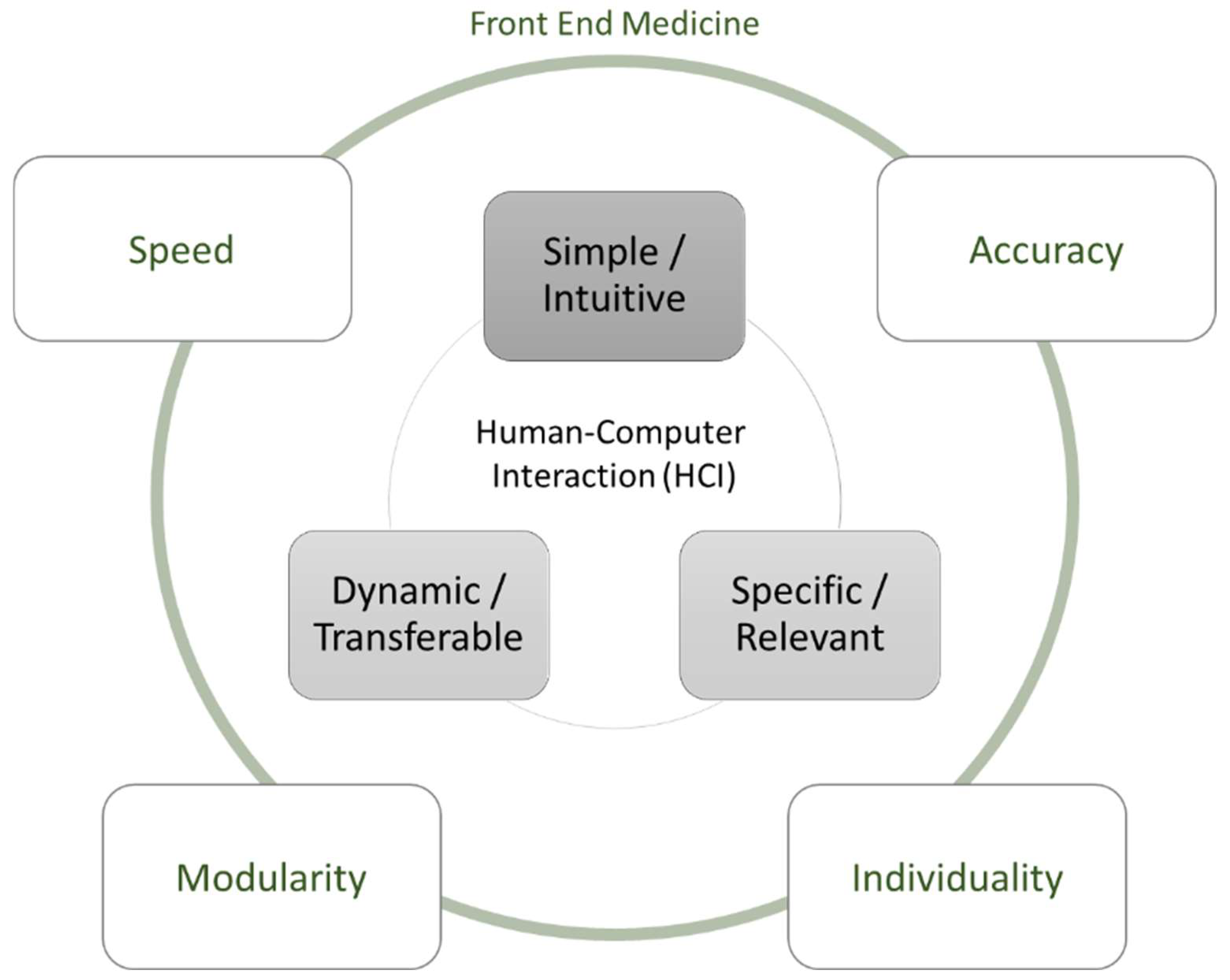
2.2. HCI Regarding the Front End
3. Use Case A: Digital Support for Emergency Medical Services (Münster)
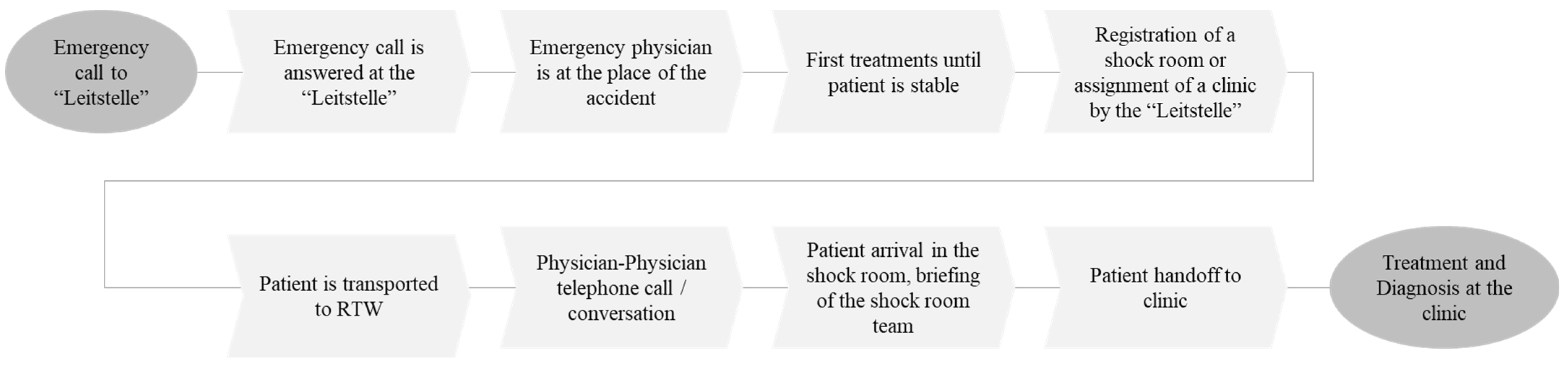
3.1. Background
3.2. Medical Requirements
3.3. Application Concept
3.4. Functional and Technical Requirements
- “As an emergency doctor, I would like the information about the patient and the emergency to be transmitted directly to the hospital without any loss of information in order to shorten the doctor–doctor consultation and avoid discussions.”
- “As the resuscitation room leader, I want the information to be exchanged between the emergency doctor and myself in a structured digital way so that as little irrelevant information as possible is exchanged and no errors occur.”
- “As an emergency doctor, I would like to be able to see the capacities of the surrounding clinics with a relevant competence profile in order to be able to decide which clinic is suitable for the transport decision.”
- “As an emergency doctor, I would like to be supported in the selection of the clinic by information such as the travel time to the clinic and the treatment focus of the clinic so that this decision is not made purely subjectively.”
3.5. Concept Outline
3.6. HCI Reflections
4. Use Case B: Inpatient Information Management with Automated Information Extraction (Göttingen)
4.1. Background
4.2. Functional and Technical Requirements
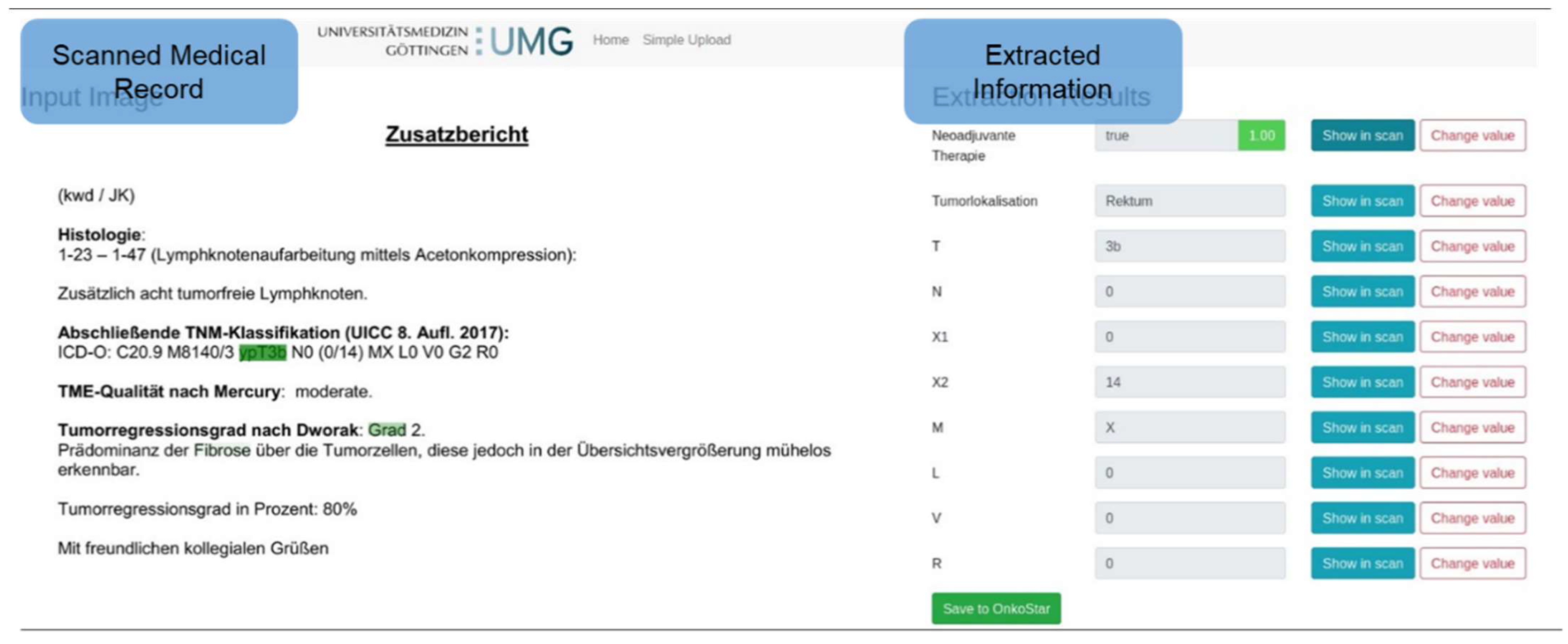
4.3. Concept Outline
4.4. Expert Interaction Module
4.5. Feature Extraction Module
4.6. Results
4.7. HCI Reflection
5. Use Case C: Cancer Diagnosis (Göttingen)
5.1. Background
5.2. Functional and Technical Requirements
- find the most important attributes for treatment decisions,
- predict an optimal treatment for a patient based on all available attributes, and
- estimate the prediction accuracy.
5.3. Concept Outline
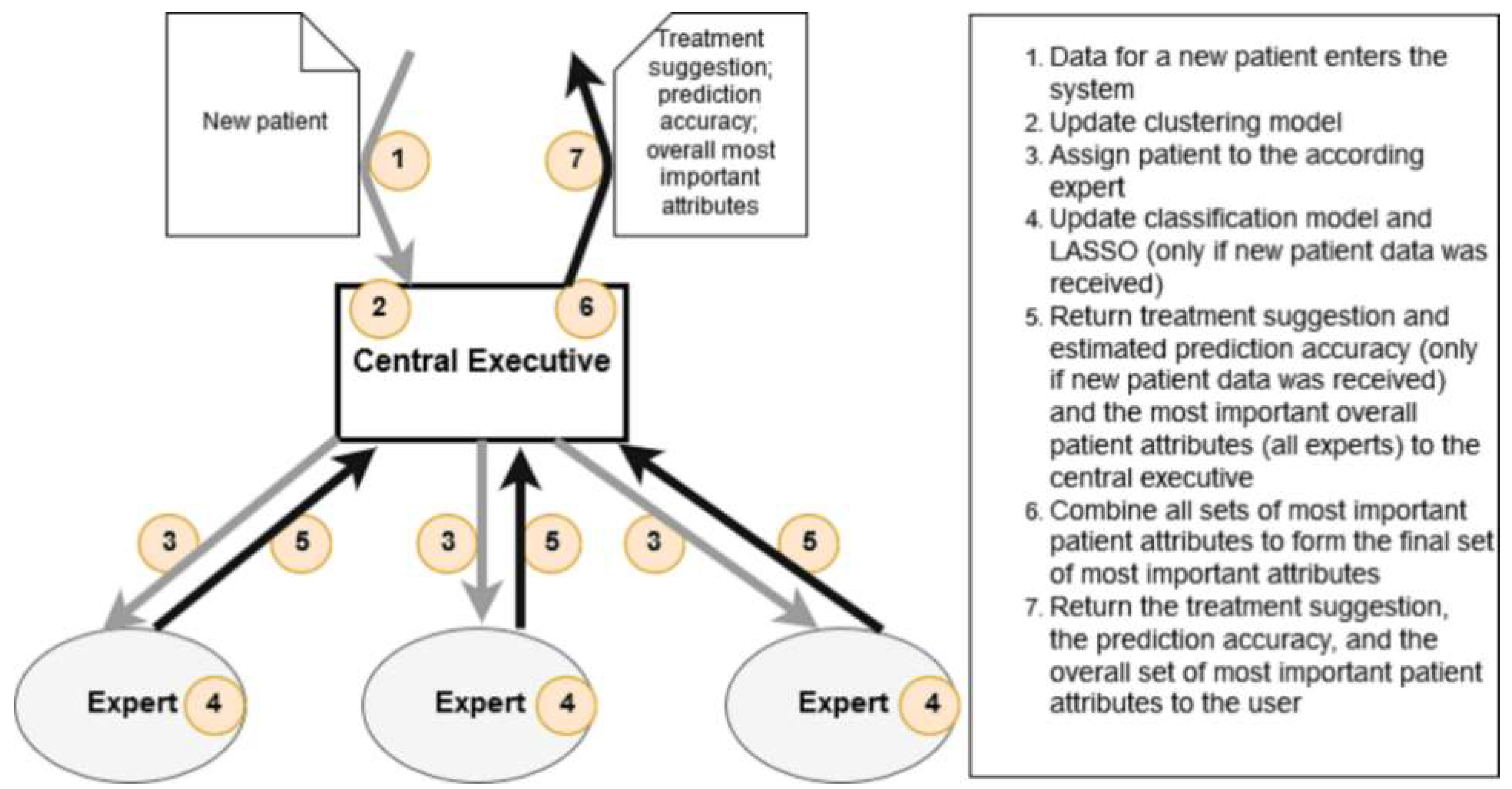
- Expert: We divide an Expert’s tasks into learning tasks, performed when the Expert receives new labeled data, and analysis tasks, performed when the Expert is queried for information. The learning tasks of an Expert are defined as (1) learning attribute importance ratings for each of the 6 treatment decisions and (2) modeling the relationship between the 160 features and the treatment decisions. Each time an Expert is provided with a data element, it updates its learning models for both tasks in an online manner and returns the suggested treatment decisions.
- Central Executive: Again, we divide the CE’s tasks into learning tasks and analysis tasks. The CE has a single learning task of performing patient segmentation and creating/updating an Expert for each segment. The analysis tasks involve (1) querying the most suitable Expert for a prediction and (2) identifying the most important attributes for decision making.
- Base Models: For our DoL system, we use of three kinds of ML models. First, we employ a feature selection method, namely, the least absolute shrinkage and selection operator (LASSO, [76]), to identify the most significant patient attributes. LASSO is a common method for feature selection that uses regularized regression to identify notable features (i.e., perform dimensionality reduction). Second, we use a binary classification model for the prediction of treatments. The last version of the DoL system employs a single type of classification. However, as part of our research, we try several classification models, both in a standalone manner and as a part of a DoL system, to find a model of high quality with respect to our measure of accuracy (i.e., the F_1 score). Third, for the purpose of task division, which is realized through patient segmentation, we make use of a clustering algorithm. Figure 6 illustrates our approach.
5.4. Results
5.5. HCI Reflection
6. Discussion of Specific HCI Challenges in the Front End of Medicine
7. Outlook
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Yousaf, K.; Mehmood, Z.; Awan, I.A.; Saba, T.; Alharbey, R.; Qadah, T.; Alrige, M.A. A comprehensive study of mobile-health based assistive technology for the healthcare of dementia and Alzheimer’s disease (AD). Health Care Manag. Sci. 2020, 23, 287–309. [Google Scholar] [CrossRef] [PubMed]
- Rejeb, O.; Pilet, C.; Hamana, S.; Xie, X.; Durand, T.; Aloui, S.; Doly, A.; Biron, P.; Perrier, L.; Augusto, V. Performance and cost evaluation of health information systems using micro-costing and discrete-event simulation. Health Care Manag. Sci. 2018, 21, 204–223. [Google Scholar] [CrossRef] [PubMed]
- Syed-Abdul, S.; Zhu, X.; Fernandez-Luque, L. Digital Health: Mobile and Wearable Devices for Participatory Health Applications; Elsevier: Amsterdam, The Netherlands, 2020; ISBN 9780128200773. [Google Scholar]
- Agnihothri, S.; Cui, L.; Delasay, M.; Rajan, B. The value of mHealth for managing chronic conditions. Health Care Manag. Sci. 2020, 23, 185–202. [Google Scholar] [CrossRef] [PubMed]
- Dohse, H. Patient perspective: Wearable and digital health tools to support managing our health during the COVID-19 pandemic and beyond. Cardiovasc. Digit. Health J. 2021, 2, 88. [Google Scholar] [CrossRef]
- Mahajan, S.; Lu, Y.; Spatz, E.S.; Nasir, K.; Krumholz, H.M. Trends and Predictors of Use of Digital Health Technology in the United States. Am. J. Med. 2021, 134, 129–134. [Google Scholar] [CrossRef]
- Gartner, D.; Zhang, Y.; Padman, R. Cognitive workload reduction in hospital information systems: Decision support for order set optimization. Health Care Manag. Sci. 2018, 21, 224–243. [Google Scholar] [CrossRef]
- van Essen, J.T.; Hurink, J.L.; Hartholt, W.; van den Akker, B.J. Decision support system for the operating room rescheduling problem. Health Care Manag. Sci. 2012, 15, 355–372. [Google Scholar] [CrossRef]
- Blandford, A. HCI for health and wellbeing: Challenges and opportunities. Int. J. Hum. -Comput. Stud. 2019, 131, 41–51. [Google Scholar] [CrossRef]
- Thieme, A.; Belgrave, D.; Doherty, G. Machine Learning in Mental Health. ACM Trans. Comput.-Hum. Interact. 2020, 27, 1–53. [Google Scholar] [CrossRef]
- Kjeldskov, J.; Paay, J. Indexicality. ACM Trans. Comput.-Hum. Interact. 2010, 17, 1–28. [Google Scholar] [CrossRef]
- Medhi, I.; Patnaik, S.; Brunskill, E.; Gautama, S.N.; Thies, W.; Toyama, K. Designing mobile interfaces for novice and low-literacy users. ACM Trans. Comput.-Hum. Interact. 2011, 18, 1–28. [Google Scholar] [CrossRef]
- Alnanih, R.; Ormandjieva, O. Mapping HCI Principles to Design Quality of Mobile User Interfaces in Healthcare Applications. Procedia Comput. Sci. 2016, 94, 75–82. [Google Scholar] [CrossRef][Green Version]
- Churchill, E.F. HCI and UX as translational research. Interactions 2020, 27, 22–23. [Google Scholar] [CrossRef]
- Velt, R.; Benford, S.; Reeves, S. Translations and Boundaries in the Gap Between HCI Theory and Design Practice. ACM Trans. Comput.-Hum. Interact. 2020, 27, 1–28. [Google Scholar] [CrossRef]
- Reynoso, J.M.G.; Romo, L.I.S. Measuring the Effectiveness of Designing End-User Interfaces Using Design Theories. Int. J. Inf. Technol. Syst. Approach 2020, 13, 54–72. [Google Scholar] [CrossRef]
- Hornbæk, K.; Hertzum, M. Technology Acceptance and User Experience. ACM Trans. Comput.-Hum. Interact. 2017, 24, 1–30. [Google Scholar] [CrossRef]
- Hornbæk, K.; Mottelson, A.; Knibbe, J.; Vogel, D. What Do We Mean by “Interaction”? An Analysis of 35 Years of CHI. ACM Trans. Comput.-Hum. Interact. 2019, 26, 1–30. [Google Scholar] [CrossRef]
- Hewett, T.; Baecker, R.; Card, S.; Carey, T.; Gasen, J.; Mantei, M.; Perlman, G.; Strong, G.; Verplank, W. ACM SIGCHI Curricula for Human-Computer Interaction; Association for Computing Machinery: New York, NY, USA, 1992; ISBN 0897914740. [Google Scholar]
- Li, L.; Zhang, P. The Intellectual Development of Human-Computer Interaction Research: A Critical Assessment of the MIS Literature (1990–2002). JAIS 2005, 6, 227–292. [Google Scholar] [CrossRef]
- Shneiderman, B. Designing the User Interface: Strategies for Effective Human-Computer Interaction, 3rd ed.; Addison-Wesley: Reading, MA, USA, 1998; ISBN 978-0-201-69497-0. [Google Scholar]
- Shneiderman, B.; Plaisant, C. Designing the User Interface: Strategies for Effective Human-Computer Interaction, 5th ed.; Addison-Wesley: Boston, MA, USA, 2010; ISBN 9780321537355. [Google Scholar]
- Carroll, J.M.; Rosson, M.B. Getting around the task-artifact cycle. ACM Trans. Inf. Syst. 1992, 10, 181–212. [Google Scholar] [CrossRef]
- Diederich, S.; Brendel, A.B.; Kolbe, L.M. Designing Anthropomorphic Enterprise Conversational Agents. Bus. Inf. Syst. Eng. 2020, 62, 193–209. [Google Scholar] [CrossRef]
- Goh, K.-Y.; Ping, J. Engaging Consumers with Advergames: An Experimental Evaluation of Interactivity, Fit and Expectancy. JAIS 2014, 15, 388–421. [Google Scholar] [CrossRef]
- Tilson, D.; Lyytinen, K.; Sørensen, C. Research Commentary —Digital Infrastructures: The Missing IS Research Agenda. Inf. Syst. Res. 2010, 21, 748–759. [Google Scholar] [CrossRef]
- Vodanovich, S.; Sundaram, D.; Myers, M. Research Commentary—Digital Natives and Ubiquitous Information Systems. Inf. Syst. Res. 2010, 21, 711–723. [Google Scholar] [CrossRef]
- Yoo, Y. Computing in Everyday Life: A Call for Research on Experiential Computing. MIS Q. 2010, 34, 213. [Google Scholar] [CrossRef]
- Lafky, D.B.; Tulu, B.; Horan, T.A. Information Systems and Health Care X: A User-Driven Approach to Personal Health Records. CAIS 2006, 17. [Google Scholar] [CrossRef]
- Du, K.; Yu, G.; Li, G.; Zhang, W. Applying Modular Design in Architecting Interorganizational Information Systems. MISQE 2019, 18, 175–189. [Google Scholar] [CrossRef]
- Kohli, R.; Tan, S.S.-L. Electronic Health Records: How Can IS Researchers Contribute to Transforming Healthcare? MISQ 2016, 40, 553–573. [Google Scholar] [CrossRef]
- Romanow, D.; Cho, S.; Straub, D. Editor’s Comments: Riding the Wave: Past Trends and Future Directions for Health IT Research. MIS Q. 2012, 36, III-A18. [Google Scholar] [CrossRef]
- Di, L.; Patrick, J.; Labeau, F. Estimating the waiting time of multi-priority emergency patients with downstream blocking. Health Care Manag. Sci. 2014, 17, 88–99. [Google Scholar] [CrossRef]
- Rachuba, S.; Salmon, A.; Zhelev, Z.; Pitt, M. Redesigning the diagnostic pathway for chest pain patients in emergency departments. Health Care Manag. Sci. 2018, 21, 177–191. [Google Scholar] [CrossRef]
- Kripalani, S.; LeFevre, F.; Phillips, C.O.; Williams, M.V.; Basaviah, P.; Baker, D.W. Deficits in communication and information transfer between hospital-based and primary care physicians: Implications for patient safety and continuity of care. JAMA 2007, 297, 831–841. [Google Scholar] [CrossRef] [PubMed]
- Meisel, Z.F.; Shea, J.A.; Peacock, N.J.; Dickinson, E.T.; Paciotti, B.; Bhatia, R.; Buharin, E.; Cannuscio, C.C. Optimizing the patient handoff between emergency medical services and the emergency department. Ann. Emerg. Med. 2015, 65, 310–317.e1. [Google Scholar] [CrossRef] [PubMed]
- Owen, C.; Hemmings, L.; Brown, T. Lost in translation: Maximizing handover effectiveness between paramedics and receiving staff in the emergency department. Emerg. Med. Australas. 2009, 21, 102–107. [Google Scholar] [CrossRef] [PubMed]
- Finkenzeller, C.; Burghofer, K.; Köhler, M.; Ruppert, M.; Stolpe, E.; Lackner, C.K. Verzögerungen im Prähospitalzeitintervall bei Luftrettungseinsätzen. Notarzt 2005, 21, 195–205. [Google Scholar] [CrossRef]
- Clarke, J.R.; Trooskin, S.Z.; Doshi, P.J.; Greenwald, L.; Mode, C.J. Time to laparotomy for intra-abdominal bleeding from trauma does affect survival for delays up to 90 minutes. J. Trauma 2002, 52, 420–425. [Google Scholar] [CrossRef]
- Fleet, R.; Lauzier, F.; Tounkara, F.K.; Turcotte, S.; Poitras, J.; Morris, J.; Ouimet, M.; Fortin, J.-P.; Plant, J.; Légaré, F.; et al. Profile of trauma mortality and trauma care resources at rural emergency departments and urban trauma centres in Quebec: A population-based, retrospective cohort study. BMJ Open 2019, 9, e028512. [Google Scholar] [CrossRef]
- Book, M.; Gruhn, V.; Striemer, R. Erfolgreiche agile Projekte; Springer: Berlin/Heidelberg, Germany, 2017; ISBN 978-3-662-53329-1. [Google Scholar]
- Cohn, M. User Stories Applied: For Agile Software Development; Addison-Wesley: Boston, MA, USA, 2004; ISBN 978-0321205681. [Google Scholar]
- Bertram, N.; Püschner, F.; Gonçalves, A.S.O.; Binder, S.; Amelung, V.E. Einführung einer elektronischen Patientenakte in Deutschland vor dem Hintergrund der internationalen Erfahrungen. In Krankenhaus-Report 2019; Klauber, J., Geraedts, M., Friedrich, J., Wasem, J., Eds.; Springer: Berlin/Heidelberg, Germany, 2019; pp. 3–16. ISBN 978-3-662-58224-4. [Google Scholar]
- Yau, J.; Chan, A.; Eapen, T.; Oirourke, K.; Eapen, L. Accuracy of The Oncology Patients Information System in a regional cancer centre. Oncol. Rep. 2002, 2002, 167–169. [Google Scholar] [CrossRef]
- Brierley, J.D.; Gospodarowicz, M.K. TNM Classification of Malignant Tumours, 8th ed.; John Wiley & Sons Incorporated: Chicester, UK, 2016; ISBN 978-1-119-26357-9. [Google Scholar]
- Poissant, L.; Pereira, J.; Tamblyn, R.; Kawasumi, Y. The impact of electronic health records on time efficiency of physicians and nurses: A systematic review. J. Am. Med. Inform. Assoc. 2005, 12, 505–516. [Google Scholar] [CrossRef]
- Dalianis, H.; Hassel, M.; Velupillai, S. The Stockholm EPR Corpus@ Characteristics and Some Initial Findings. Proc. ISHIMR 2009, 2009, 243–249. [Google Scholar]
- Epstein, R.M. How doctors think. J. Clin. Invest. 2007, 117, 2738. [Google Scholar] [CrossRef][Green Version]
- Kruse, C.S.; Beane, A. Health Information Technology Continues to Show Positive Effect on Medical Outcomes: Systematic Review. J. Med. Internet Res. 2018, 20, e41. [Google Scholar] [CrossRef] [PubMed]
- Cowie, J.; Lehnert, W. Information extraction. Commun. ACM 1996, 39, 80–91. [Google Scholar] [CrossRef]
- Sarawagi, S. Information Extraction. FNT Databases 2007, 1, 261–377. [Google Scholar] [CrossRef]
- Small, S.G.; Medsker, L. Review of information extraction technologies and applications. Neural Comput. Appl. 2014, 25, 533–548. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, L.; Rastegar-Mojarad, M.; Moon, S.; Shen, F.; Afzal, N.; Liu, S.; Zeng, Y.; Mehrabi, S.; Sohn, S.; et al. Clinical information extraction applications: A literature review. J. Biomed. Inform. 2018, 77, 34–49. [Google Scholar] [CrossRef]
- D’Avolio, L.W.; Litwin, M.S.; Rogers, S.O.; Bui, A.A.T. Facilitating Clinical Outcomes Assessment through the automated identification of quality measures for prostate cancer surgery. J. Am. Med. Inform. Assoc. 2008, 15, 341–348. [Google Scholar] [CrossRef]
- Interactive Knowledge Discovery and Data Mining in Biomedical Informatics; Holzinger, A., Jurisica, I., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; ISBN 978-3-662-43967-8. [Google Scholar]
- Gregor, S.; Benbasat, I. Explanations from Intelligent Systems: Theoretical Foundations and Implications for Practice. MIS Q. 1999, 23, 497. [Google Scholar] [CrossRef]
- Sinha, R.; Swearingen, K. The role of transparency in recommender systems. In Proceedings of the CHI ‘02 Extended Abstracts on Human Factors in Computing Systems—CHI’02, CHI’02 Extended Abstracts, Minneapolis, MI, USA, 20–25 April 2002; Terveen, L., Wixon, D., Eds.; ACM Press: New York, NY, USA, 2002; p. 830, ISBN 1581134541. [Google Scholar]
- Cutillo, C.M.; Sharma, K.R.; Foschini, L.; Kundu, S.; Mackintosh, M.; Mandl, K.D. Machine intelligence in healthcare-perspectives on trustworthiness, explainability, usability, and transparency. NPJ Digit. Med. 2020, 3, 47. [Google Scholar] [CrossRef]
- Toepfer, M.; Corovic, H.; Fette, G.; Klügl, P.; Störk, S.; Puppe, F. Fine-grained information extraction from German transthoracic echocardiography reports. BMC Med. Inform. Decis. Mak. 2015, 15, 91. [Google Scholar] [CrossRef]
- Bologna, G.; Hayashi, Y. Characterization of Symbolic Rules Embedded in Deep DIMLP Networks: A Challenge to Transparency of Deep Learning. J. Artif. Intell. Soft Comput. Res. 2017, 7, 265–286. [Google Scholar] [CrossRef]
- Yimam, S.M.; Remus, S.; Panchenko, A.; Holzinger, A.; Biemann, C. Entity-Centric Information Access with the Human-in-the-Loop for the Biomedical Domains. In Proceedings of the Biomedical NLP Workshop, RANLP 2017—Biomedical NLP Workshop, Varna, Bulgaria, 8 September 2017; Incoma Ltd.: Shoumen, Bulgaria, 2017; pp. 42–48, ISBN 9789544520441. [Google Scholar]
- Böhrnsen, F.; Godek, F.; Kiesel, J.; Kramer, F.J.; Brockmeyer, P.; Schliephake, H. Influence of TGF-β1 on tumor transition in oral cancer cell and BMSC co-cultures. J. Craniomaxillofac. Surg. 2017, 45, 731–740. [Google Scholar] [CrossRef] [PubMed]
- Gibson, M.K.; Forastiere, A.A. Multidisciplinary approaches in the management of advanced head and neck tumors: State of the art. Curr. Opin. Oncol. 2004, 16, 220–224. [Google Scholar] [CrossRef] [PubMed]
- Takeda, T.; Takeda, S.; Uryu, K.; Ichihashi, Y.; Harada, H.; Iwase, A.; Tamura, Y.; Hibino, M.; Horiuchi, S.; Kani, H. Multidisciplinary Lung Cancer Tumor Board Connecting Eight General Hospitals in Japan via a High-Security Communication Line. JCO Clin. Cancer Inform. 2019, 3, 1–7. [Google Scholar] [CrossRef]
- Müller, S.M.; Mueller, G.F.; Navarini, A.A.; Brandt, O. National Publication Productivity during the COVID-19 Pandemic-A Preliminary Exploratory Analysis of the 30 Countries Most Affected. Biology 2020, 9, 271. [Google Scholar] [CrossRef]
- Kehl, K.L.; Landrum, M.B.; Kahn, K.L.; Gray, S.W.; Chen, A.B.; Keating, N.L. Tumor board participation among physicians caring for patients with lung or colorectal cancer. J. Oncol. Pract. 2015, 11, e267–e278. [Google Scholar] [CrossRef] [PubMed]
- Tofangchi, S.; Hanelt, A.; Böhrnsen, F. Distributed Cognitive Expert Systems in Cancer Data Analytics: A Decision Support System for Oral and Maxillofacial Surgery. In Proceedings of the ICIS 2017, Transforming Society with Digital Innovation, Seoul, South Korea, 10–13 December 2017; Kim, Y.J., Agarwal, R., Lee, J.K., Eds.; ICIS: Seoul, South Korea, 2017; p. 35. [Google Scholar]
- Abernethy, A.P.; Etheredge, L.M.; Ganz, P.A.; Wallace, P.; German, R.R.; Neti, C.; Bach, P.B.; Murphy, S.B. Rapid-learning system for cancer care. J. Clin. Oncol. 2010, 28, 4268–4274. [Google Scholar] [CrossRef]
- Schilsky, R.L.; Michels, D.L.; Kearbey, A.H.; Yu, P.P.; Hudis, C.A. Building a rapid learning health care system for oncology: The regulatory framework of CancerLinQ. J. Clin. Oncol. 2014, 32, 2373–2379. [Google Scholar] [CrossRef]
- Mayo, R.M.; Summey, J.F.; Williams, J.E.; Spence, R.A.; Kim, S.; Jagsi, R. Qualitative Study of Oncologists’ Views on the CancerLinQ Rapid Learning System. J. Oncol. Pract. 2017, 13, e176–e184. [Google Scholar] [CrossRef]
- Valdes, G.; Solberg, T.D.; Heskel, M.; Ungar, L.; Simone, C.B. Using machine learning to predict radiation pneumonitis in patients with stage I non-small cell lung cancer treated with stereotactic body radiation therapy in Taiwan. Phys. Med. Biol. 2016, 61, 6105–6120. [Google Scholar] [CrossRef]
- Tofangchi, S.; Hanelt, A.; Kolbe, L.M. Towards Distributed Cognitive Expert Systems. In Designing the Digital Transformation; Maedche, A., vom Brocke, J., Hevner, A., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 145–159. ISBN 978-3-319-59143-8. [Google Scholar]
- Cox, D.R. The Regression Analysis of Binary Sequences. J. R. Stat. Soc. Ser. B (Methodol.) 1958, 20, 215–242. [Google Scholar] [CrossRef]
- Cox, D.R.; Snell, E.J. Analysis of Binary Data; Chapman & Hall: London, UK, 1989. [Google Scholar]
- Breiman, L.; Friedman, J.H.; Olshen, R.A.; Stone, C.J. Classification and Regression Trees; Wadsworth & Brooks/Cole Advanced Books & Software: Monterey, CA, USA, 1984. [Google Scholar]
- Cortes, C.; Vapnik, V. Support-Vector Networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Bengio, Y.; Lecun, Y. Scaling Learning Algorithms towards AI. In Large-Scale Kernel Machines; Bottou, L., Chapelle, O., Decoste, D., Weston, J., Eds.; MIT Press: Boston, MA, USA, 2007. [Google Scholar]
- Angst, A. Adoption of Electronic Health Records in the Presence of Privacy Concerns: The Elaboration Likelihood Model and Individual Persuasion. MIS Q. 2009, 33, 339. [Google Scholar] [CrossRef]
- Fichman, R.G.; Kohli, R.; Krishnan, R. Editorial Overview—The Role of Information Systems in Healthcare: Current Research and Future Trends. Inf. Syst. Res. 2011, 22, 419–428. [Google Scholar] [CrossRef]
- Zhuang, Z.Y.; Wilkin, C.L.; Ceglowski, A. A framework for an intelligent decision support system: A case in pathology test ordering. Decis. Support Syst. 2013, 55, 476–487. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression Shrinkage and Selection via the Lasso. J. R. Stat. Society. Ser. B (Methodol.) 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Gupta, A.; Sharda, R. Improving the science of healthcare delivery and informatics using modeling approaches. Decis. Support Syst. 2013, 55, 423–427. [Google Scholar] [CrossRef]
- Agarwal, R.; Gao, G.; DesRoches, C.; Jha, A. Research Commentary: The Digital Transformation of Healthcare: Current Status and the Road Ahead. Inf. Syst. Res. 2010, 21, 796–809. [Google Scholar] [CrossRef]
- Kruse, C.S.; Goswamy, R.; Raval, Y.; Marawi, S. Challenges and Opportunities of Big Data in Health Care: A Systematic Review. JMIR Med. Inform. 2016, 4, e38. [Google Scholar] [CrossRef]
- Phutela, N.; Chowdary, A.N.; Anchlia, S.; Jaisinghani, D.; Gabrani, G. Unlock Me: A Real-World Driven Smartphone Game to Stimulate COVID-19 Awareness. Int. J. Hum. -Comput. Stud. 2022, 164, 102818. [Google Scholar] [CrossRef]


| Feature | Manual Coding | Automated Transmission |
|---|---|---|
| Neo. therapy | 99.19% | 98.78% |
| T | 99.59% | 97.97% |
| N | 99.59% | 98.78% |
| X1/2 | 98.38% | 97.57% |
| M | 99.59% | 99.59% |
| L | 100% | 100% |
| V | 100% | 100% |
| R | 100% | 98.78% |
| Complete data | 97.57% | 93.52% |
| Extraction Method | True | False | Ambiguous | Total |
|---|---|---|---|---|
| Manual Coding | 241 (97.57%) | 6 | - | 247 |
| Automated Transmission | 233 (93.52%) | 5 | 9 | 247 |
| Most Important Patient Attributes for Predictions | |||||
|---|---|---|---|---|---|
| Decision Category | Attribute | Score | Decision Category | Attribute | Score |
| Therapy Intention | Lymph node-(N)-classification | 0.825 | Systemic Chemotherapy | Tumor-(T)-classification | 0.692 |
| Tumor-(T)-classification | 0.438 | Diagnosis admission status | 0.505 | ||
| Metastasis-(M)-classification | 0.332 | Clinical state | 0.441 | ||
| Grading of squamous cell carcinoma | 0.202 | Tumor number | 0.357 | ||
| Clinical state | 0.129 | Tumor type | 0.337 | ||
| Height | 0.0714 | Reason for visit | 0.278 | ||
| Tumor number | 0.0698 | Sex | 0.274 | ||
| Familial tumor predisposition | 0.0648 | Height | 0.211 | ||
| Reason for visit | 0.0568 | Familial tumor predisposition | 0.185 | ||
| Operation | Metastasis-(M)-classification | 0.276 | Amount of alcohol | 0.169 | |
| Grading of squamous cell carcinoma | 0.146 | Other Therapy | Clinical state | 0.906 | |
| Lymph node-(N)-classification | 0.145 | Reason for visit | 0.358 | ||
| Tumor type | 0.0592 | Duration of precancerous lesions | 0.109 | ||
| Tumor-(T)-classification | 0.0494 | Tumor type | 0.0811 | ||
| Radiation | Tumor-(T)-classification | 0.366 | Tumor-(T)-classification | 0.0550 | |
| Lymph node-(N)-classification | 0.322 | Overall | Tumor-(T)-classification | 0.323 | |
| Inflammatory response | 0.0789 | Clinical state | 0.299 | ||
| Degree of dysplasia | 0.0533 | Lymph node-(N)-classification | 0.273 | ||
| Existing precancerous lesions | 0.0430 | Reason for visit | 0.140 | ||
| Local Chemotherapy | Heavy heart disease | 0.0790 | Metastasis-(M)-classification | 0.120 | |
| Grading of squamous cell carcinoma | 0.0682 | ||||
| Lymph node-(N)-classification | 0.0675 | ||||
| Restricted lung functions | 0.0554 | ||||
| Existing precancerous lesions | 0.0454 | ||||
| Therapy Intention | Operation | Radiation | Local Chemo-therapy | Systemic Chemo-Therapy | Other Therapy | Overall | |
|---|---|---|---|---|---|---|---|
| Random Guesser | 0.120 | 0.610 | 0.315 | 0.00752 | 0.234 | 0.0153 | 0.218 |
| Prior Classifier | 0 | 0.961 | 0 | 0.286 | 0 | 0.571 | 0.308 |
| Prior Classifier DoL | 0 | 0.961 | 0 | 0.286 | 0 | 0.571 | 0.308 |
| LASSO | 0 | 0.966 | 0.48 | 0.213 | 0.240 | 0 | 0.320 |
| LASSO DoL | 0.434 | 0.958 | 0.430 | 0.259 | 0.259 | 0.286 | 0.439 |
| Logistic Regression | 0.452 | 0.958 | 0.458 | 0.285 | 0.285 | 0.571 | 0.500 |
| Logistic Regression DoL | 0.649 | 0.957 | 0.528 | 0.496 | 0.561 | 0.571 | 0.620 |
| LPM | 0.336 | 0.945 | 0.457 | 0.127 | 0.127 | 0.146 | 0.358 |
| LPM DoL | 0.526 | 0.946 | 0.454 | 0.139 | 0.139 | 0.146 | 0.392 |
| Decision Tree | 0.308 | 0.932 | 0.355 | 0.240 | 0.22 | 0.223 | 0.382 |
| Decision Tree DoL | 0.455 | 0.937 | 0.448 | 0.304 | 0.258 | 0.242 | 0.442 |
| SVM | 0.260 | 0.957 | 0.397 | 0.286 | 0.300 | 0.571 | 0.465 |
| SVM DoL | 0.41 | 0.959 | 0.513 | 0.476 | 0.517 | 0.571 | 0.577 |
| DNN | 0.0317 | 0.851 | 0.104 | 0.143 | 0.0395 | 0.571 | 0.316 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Klumpp, M.; Hanelt, A.; Greve, M.; Kolbe, L.M.; Tofangchi, S.; Böhrnsen, F.; Jakob, J.; Kaczmarek, S.; Börsting, I.; Ehmke, C.; et al. Accelerating the Front End of Medicine: Three Digital Use Cases and HCI Implications. Healthcare 2022, 10, 2176. https://doi.org/10.3390/healthcare10112176
Klumpp M, Hanelt A, Greve M, Kolbe LM, Tofangchi S, Böhrnsen F, Jakob J, Kaczmarek S, Börsting I, Ehmke C, et al. Accelerating the Front End of Medicine: Three Digital Use Cases and HCI Implications. Healthcare. 2022; 10(11):2176. https://doi.org/10.3390/healthcare10112176
Chicago/Turabian StyleKlumpp, Matthias, André Hanelt, Maike Greve, Lutz M. Kolbe, Schahin Tofangchi, Florian Böhrnsen, Jens Jakob, Sylvia Kaczmarek, Ingo Börsting, Christopher Ehmke, and et al. 2022. "Accelerating the Front End of Medicine: Three Digital Use Cases and HCI Implications" Healthcare 10, no. 11: 2176. https://doi.org/10.3390/healthcare10112176
APA StyleKlumpp, M., Hanelt, A., Greve, M., Kolbe, L. M., Tofangchi, S., Böhrnsen, F., Jakob, J., Kaczmarek, S., Börsting, I., Ehmke, C., Düsing, H., & Juhra, C. (2022). Accelerating the Front End of Medicine: Three Digital Use Cases and HCI Implications. Healthcare, 10(11), 2176. https://doi.org/10.3390/healthcare10112176







