Genome-Wide Identification and Expression Profile Analysis of the NF-Y Transcription Factor Gene Family in Petunia hybrida
Abstract
1. Introduction
2. Results
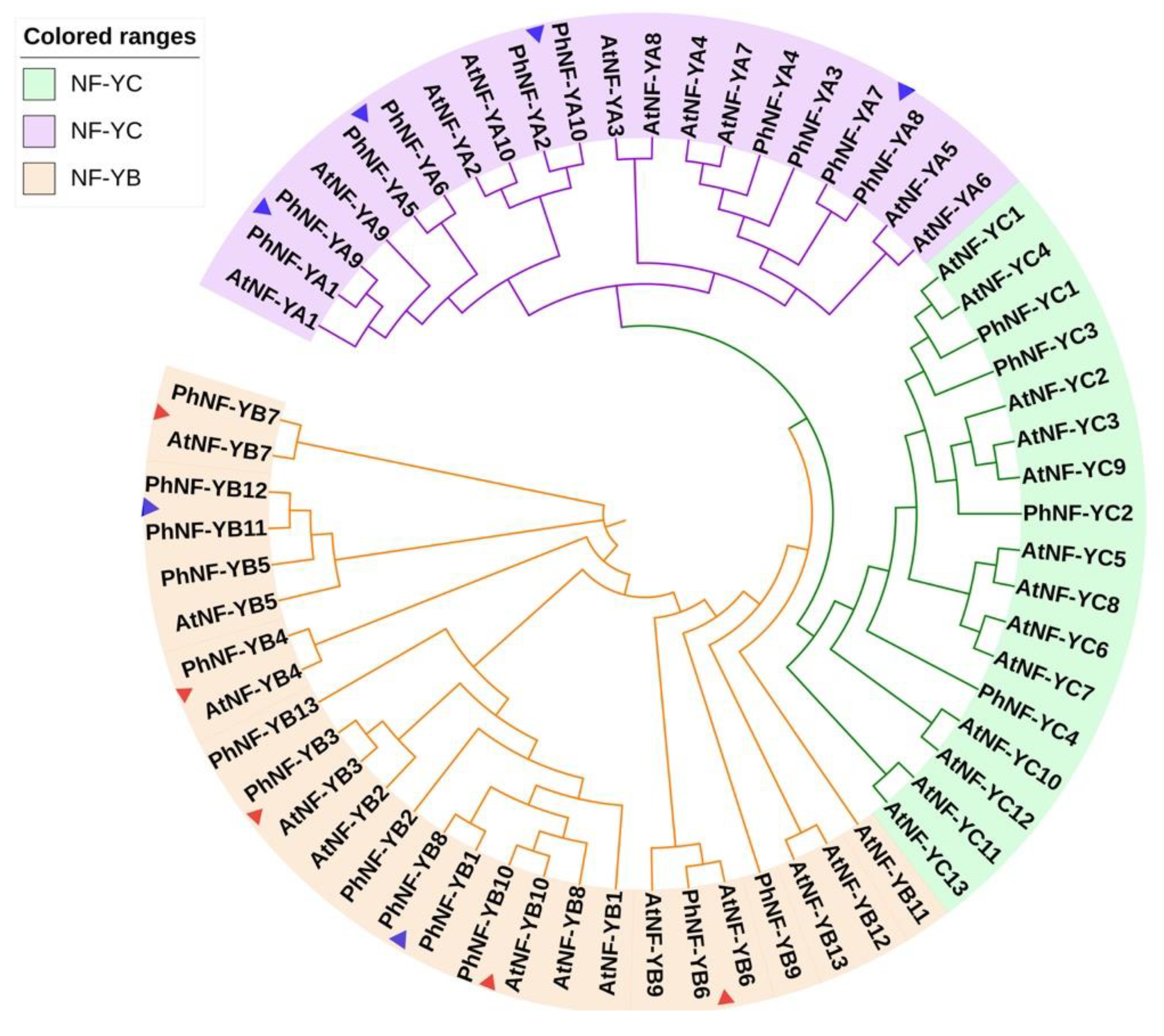
2.1. Identification of the NF-Y Family Members
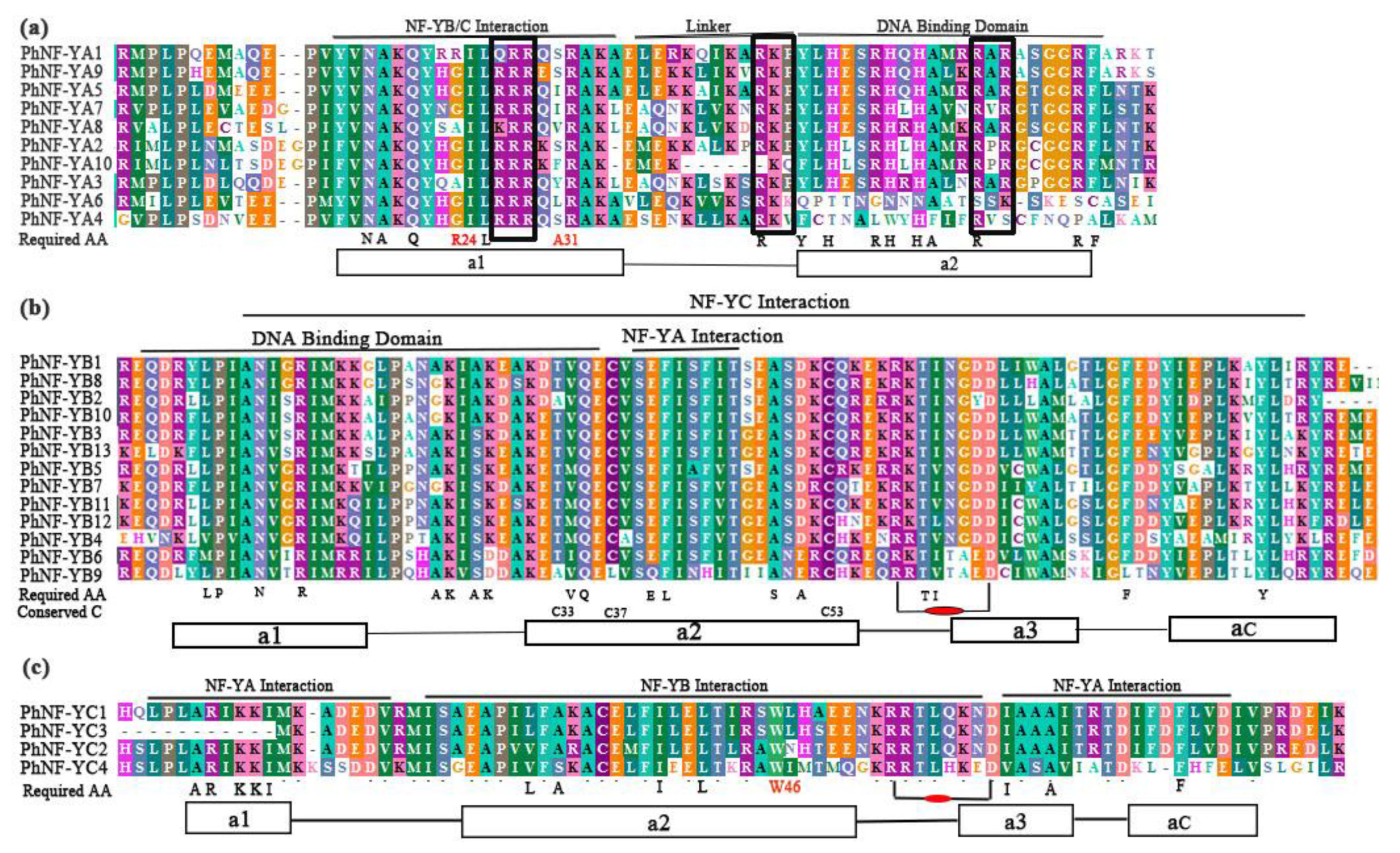
2.2. Multiple Alignment and Conserved Motifs of NF-Y Proteins
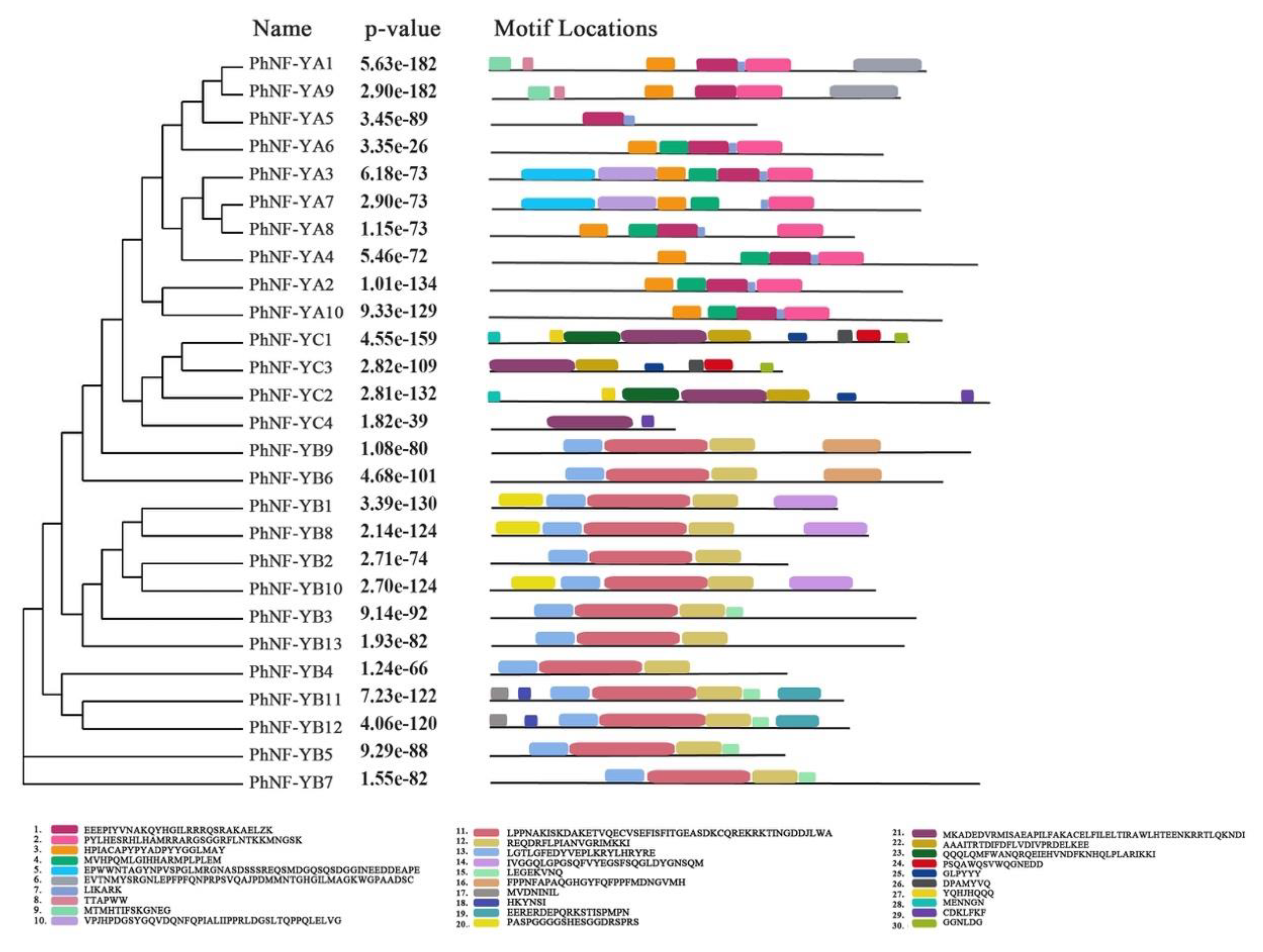
2.3. Gene Structures and Conserved Motifs of PhNF-Y Members
2.4. miRNA Target Site Prediction and Validation
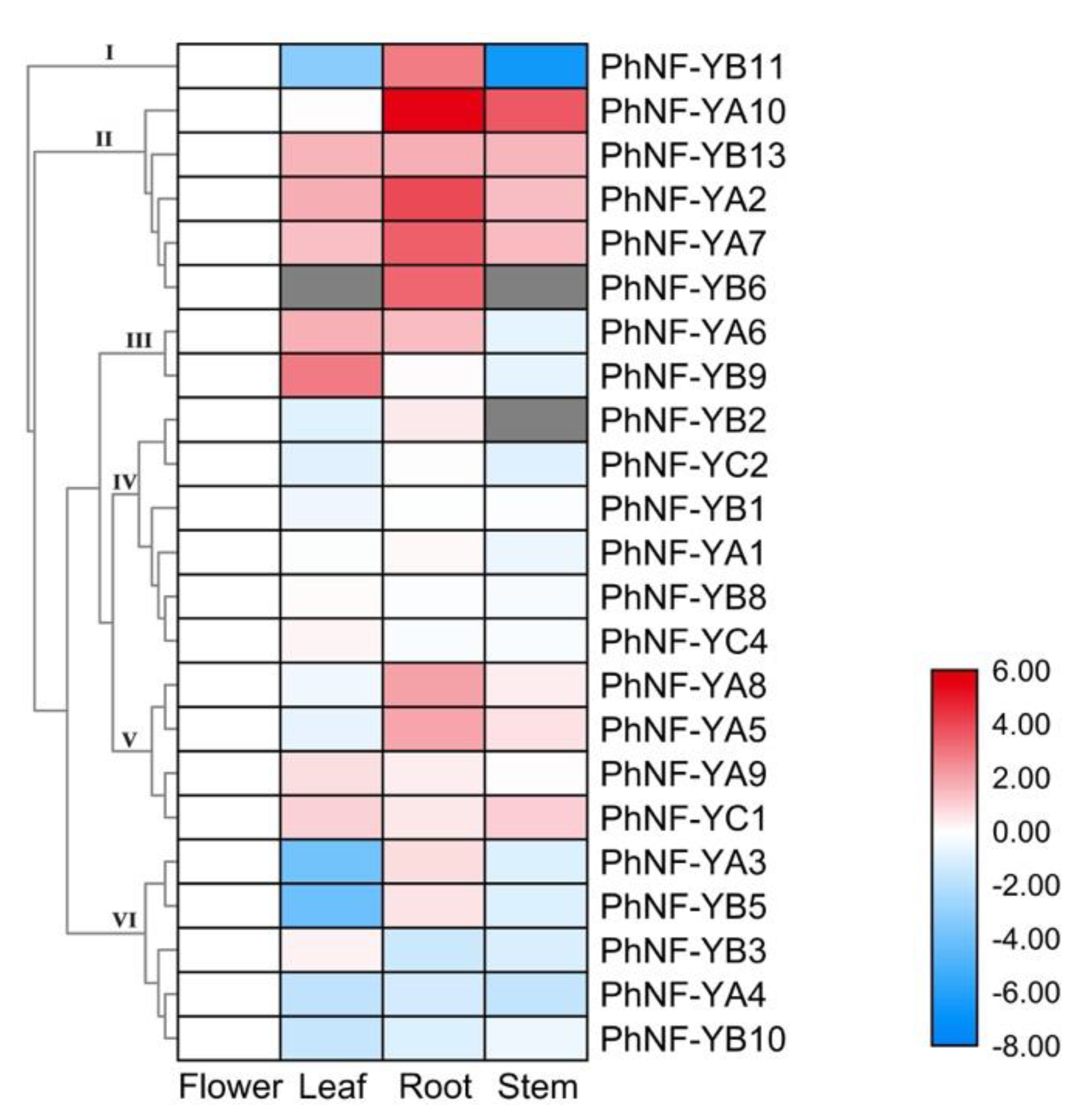
2.5. Expression Profiles of the PhNF-Ys in Different Tissues
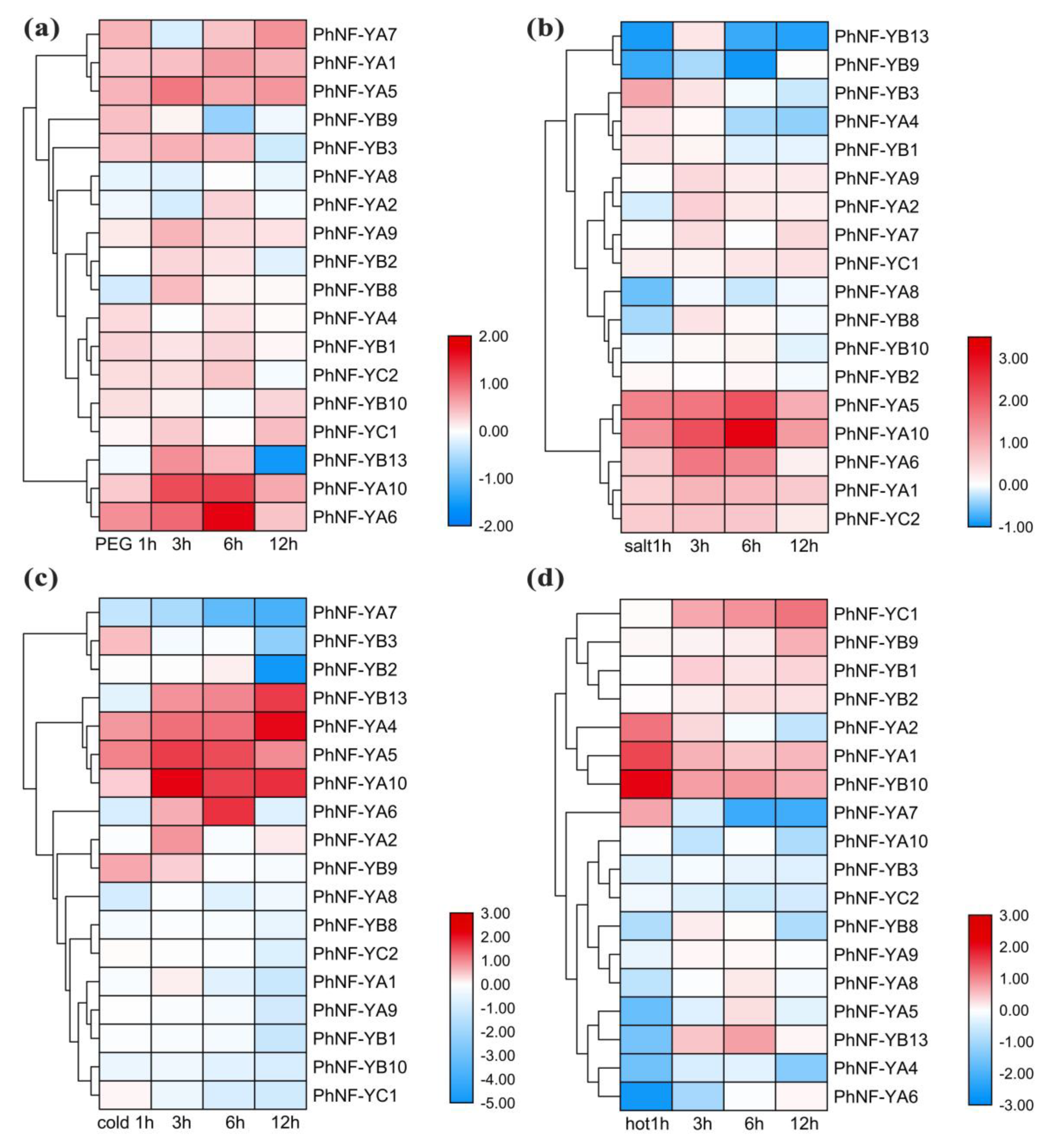
2.6. Expression Profiles of the PhNF-Ys under Abiotic Stress
3. Discussion
4. Materials and Methods
4.1. Identification, Alignments, and Phylogenetic Analysis of NF-Y Members
4.2. Conserved Motifs and Gene Structure Analyses
4.3. miRNA Target Site Prediction
4.4. Plant Material, Tissue, and Stress Treatment
4.5. Transcriptome Data Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Salehi-Lisar, S.Y.; Bakhshayeshan-Agdam, H. Drought Stress in Plants: Causes, Consequences, and Tolerance; Springer: Cham, Germany, 2016. [Google Scholar]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant. Sci. 2005, 10, 88–94. [Google Scholar] [CrossRef] [PubMed]
- Maity, S.N.; De Crombrugghe, B. Role of the CCAAT-binding protein CBF/NF-Y in transcription. Trends Biochem. Sci. 1998, 23, 174–178. [Google Scholar] [CrossRef]
- Li, M.; Li, G.; Liu, W.; Dong, X.; Zhang, A. Genome-wide analysis of the NF-Y gene family in peach (Prunus persica L.). BMC Genom. 2019, 20, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Kahle, J.; Baake, M.; Doenecke, D.; Albig, W. Subunits of the Heterotrimeric Transcription Factor NF-Y Are Imported into the Nucleus by Distinct Pathways Involving Importin and Importin 13. Mol. Cell. Biol. 2005, 25, 5339–5354. [Google Scholar] [CrossRef] [PubMed]
- Gusmaroli, G.; Tonelli, C.; Mantovani, R. Regulation of novel members of the Arabidopsis thaliana CCAAT-binding nuclear factor Y subunits. Gene 2002, 283, 41–48. [Google Scholar] [CrossRef]
- Miyoshi, K.; Ito, Y.; Serizawa, A.; Kurata, N. OsHAP3 genes regulate chloroplast biogenesis in rice. Plant J. 2003, 36, 532–540. [Google Scholar] [CrossRef]
- Malviya, N.; Jaiswal, P.; Yadav, D. Genome- wide characterization of Nuclear Factor Y (NF-Y) gene family of sorghum [Sorghum bicolor (L.) Moench]: A bioinformatics approach. Physiol. Mol. Biol. Plants 2016, 22, 33–49. [Google Scholar] [CrossRef]
- West, M.A.L.; Yee, K.M.; Danao, J.; Zimmerman, J.L.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. LEAFY COTYLEDON1 Is an Essential Regulator of Late Embryogenesis and Cotyledon Identity in Arabidopsis. Plant Cell 1994, 6, 1731–1745. [Google Scholar] [CrossRef]
- Rey, T.; Laporte, P.; Bonhomme, M.; Jardinaud, M.F.; Huguet, S.; Balzergue, S.; Dumas, B.; Niebel, A.; Jacquet, C. MtNF-YA1, a central transcriptional regulator of symbiotic nodule development, is also a determinant of medicago truncatula susceptibility toward a root pathogen. Front. Plant Sci. 2016, 7, 1–14. [Google Scholar] [CrossRef]
- Battaglia, M.; Rípodas, C.; Clúa, J.; Baudin, M.; Aguilar, O.M.; Niebel, A.; Zanetti, M.E.; Blanco, F.A. A Nuclear Factor Y Interacting Protein of the GRAS Family Is Required for Nodule Organogenesis, Infection Thread Progression, and Lateral Root Growth. Plant Physiol. 2014, 164, 1430–1442. [Google Scholar] [CrossRef]
- Parcy, F.; Valon, C.; Kohara, A.; Miséra, S.; Giraudat, J. The ABSCISIC ACID-INSENSITIVE3, FUSCA3, and LEAFY COTYLEDON1 loci act in concert to control multiple aspects of Arabidopsis seed development. Plant Cell 1997, 9, 1265–1277. [Google Scholar]
- Hou, X.; Zhou, J.; Liu, C.; Liu, L.; Shen, L.; Yu, H. Nuclear factor Y-mediated H3K27me3 demethylation of the SOC1 locus orchestrates flowering responses of Arabidopsis. Nat. Commun. 2014, 5, 1–14. [Google Scholar] [CrossRef]
- Alam, M.M.; Tanaka, T.; Nakamura, H.; Ichikawa, H.; Kobayashi, K.; Yaeno, T.; Yamaoka, N.; Shimomoto, K.; Takayama, K.; Nishina, H.; et al. Overexpression of a rice heme activator protein gene (OsHAP2E) confers resistance to pathogens, salinity and drought, and increases photosynthesis and tiller number. Plant Biotechnol. J. 2015, 13, 85–96. [Google Scholar] [CrossRef]
- Li, W.X.; Oono, Y.; Zhu, J.; He, X.J.; Wu, J.M.; Iida, K.; Lu, X.Y.; Cui, X.; Jin, H.; Zhu, J.K. The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 2008, 20, 2238–2251. [Google Scholar] [CrossRef]
- Ma, X.; Zhu, X.; Li, C.; Song, Y.; Zhang, W.; Xia, G.; Wang, M. Overexpression of wheat NF-YA10 gene regulates the salinity stress response in Arabidopsis thaliana. Plant Physiol. Biochem. 2015, 86, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D.E.; Repetti, P.P.; Adams, T.R.; Creelman, R.A.; Wu, J.; Warner, D.C.; Anstrom, D.C.; Bensen, R.J.; Castiglioni, P.P.; Donnarummo, M.G.; et al. Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc. Natl. Acad. Sci. USA 2007, 104, 16450–16455. [Google Scholar] [CrossRef] [PubMed]
- Baomei, W.; Zhaoxia, L.; Qijun, R.; Peng, L.; Zhenghua, P.; Juren, Z. ZmNF-YB16 Overexpression Improves Drought Resistance and Yield by Enhancing Photosynthesis and the Antioxidant Capacity of Maize Plants. Front. Plant Sci. 2018, 9, 709. [Google Scholar]
- Shi, H.; Ye, T.; Zhong, B.; Liu, X.; Chan, Z. AtHAP5A modulates freezing stress resistance in Arabidopsis through binding to CCAAT motif of AtXTH21. New Phytol. 2014, 203, 554–567. [Google Scholar] [CrossRef]
- Liu, X.; Hu, P.; Huang, M.; Tang, Y.; Li, Y.; Li, L.; Hou, X. The NF-YC-RGL2 module integrates GA and ABA signalling to regulate seed germination in Arabidopsis. Nat. Commun. 2016, 7, 1–14. [Google Scholar] [CrossRef]
- Li, Y.J.; Fang, Y.; Fu, Y.R.; Huang, J.G.; Wu, C.A.; Zheng, C.C. NFYA1 Is Involved in Regulation of Postgermination Growth Arrest Under Salt Stress in Arabidopsis. PLoS ONE 2013, 8, e61289. [Google Scholar] [CrossRef]
- Huang, S.; Hu, L.Q.; Xu, D.B.; Li, W.W.; Xu, Z.S.; Li, L.C.; Zhou, Y.B.; Diao, X.M.; Jia, G.Q.; Ma, Y.Z. Transcription Factor Si NF-YA5 from Foxtail Millet (Setaria italica) Conferred Tolerance to High-salt Stress through ABA-independent Pathway in Transgenic Arabidopsis. Acta Agron. Sin. 2016, 42, 1787–1797. [Google Scholar] [CrossRef]
- Ni, Z.; Hu, Z.; Jiang, Q.; Zhang, H. GmNFYA3, a target gene of miR169, is a positive regulator of plant tolerance to drought stress. Plant Mol. Biol. 2013, 82, 113–129. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.K.; Kim, H.I.; Jang, G.; Chung, P.J.; Jeong, J.S.; Kim, Y.S.; Bang, S.W.; Jung, H.; Choi, Y.D.; Kim, J.K. The NF-YA transcription factor OsNF-YA7 confers drought stress tolerance of rice in an abscisic acid independent manner. Plant Sci. 2015, 241, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Luan, M.; Xu, M.; Lu, Y.; Zhang, Q.; Zhang, L.; Zhang, C.; Fan, Y.; Lang, Z.; Wang, L. Family-Wide Survey of miR169s and NF-YAs and Their Expression Profiles Response to Abiotic Stress in Maize Roots. PLoS ONE 2014, 9, e91369. [Google Scholar] [CrossRef]
- Luan, M.; Xu, M.; Lu, Y.; Zhang, L.; Fan, Y.; Wang, L. Expression of zma-miR169 miRNAs and their target ZmNF-YA genes in response to abiotic stress in maize leaves. Gene 2015, 555, 178–185. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Tang, S.; An, Y.; Zheng, D.C.; Xia, X.L.; Yin, W.L. Overexpression of the poplar NF-YB7 transcription factor confers drought tolerance and improves water-use efficiency in Arabidopsis. J. Exp. Bot. 2013, 64, 4589–4601. [Google Scholar] [CrossRef]
- Feng, Z.J.; He, G.H.; Zheng, W.J.; Lu, P.P.; Chen, M.; Gong, Y.M.; Ma, Y.Z.; Xu, Z.S. Foxtail millet NF-Y families: Genome-wide survey and evolution analyses identified two functional genes important in abiotic stresses. Front. Plant Sci. 2015, 6, 1–19. [Google Scholar] [CrossRef]
- Chen, M.; Zhao, Y.; Zhuo, C.; Lu, S.; Guo, Z. Overexpression of a NF-YC transcription factor from bermudagrass confers tolerance to drought and salinity in transgenic rice. Plant Biotechnol. J. 2014, 13, 482–491. [Google Scholar] [CrossRef]
- Shi, H.; Chan, Z. AtHAP5A modulates freezing stress resistance in arabidopsis independent of the CBF pathway. Plant Signal. Behav. 2014, 203, 554–567. [Google Scholar]
- Siefers, N.; Dang, K.K.; Kumimoto, R.W.; Bynum IV, W.E.; Tayrose, G.; Holt, B.F. Tissue-specific expression patterns of Arabidopsis NF-Y transcription factors suggest potential for extensive combinatorial complexity. Plant Physiol. 2009, 149, 625–641. [Google Scholar] [CrossRef]
- Stephenson, T.J.; McIntyre, C.L.; Collet, C.; Xue, G.-P. Genome-wide identification and expression analysis of the NF-Y family of transcription factors in Triticum aestivum. Plant Mol. Biol. 2007, 65, 77–92. [Google Scholar] [CrossRef] [PubMed]
- Thirumurugan, T.; Ito, Y.; Kubo, T.; Serizawa, A.; Kurata, N. Identification, characterization and interaction of HAP family genes in rice. Mol. Genet. Genom. 2008, 279, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Quach, T.N.; Nguyen, H.T.M.; Valliyodan, B.; Joshi, T.; Xu, D.; Nguyen, H.T. Genome-wide expression analysis of soybean NF-Y genes reveals potential function in development and drought response. Mol. Genet. Genom. 2015, 290, 1095–1115. [Google Scholar] [CrossRef] [PubMed]
- Liang, M.; Yin, X.; Lin, Z.; Zheng, Q.; Liu, G.; Zhao, G. Identification and characterization of NF-Y transcription factor families in Canola (Brassica napus L.). Planta 2014, 239, 107–126. [Google Scholar] [CrossRef]
- Cao, S.; Kumimoto, R.W.; Siriwardana, C.L.; Risinger, J.R.; Holt, B.F. Identification and characterization of NF-Y transcription factor families in the monocot model plant Brachypodium distachyon. PLoS ONE 2011, 6, e21805. [Google Scholar] [CrossRef]
- Ren, C.; Zhang, Z.; Wang, Y.; Li, S.; Liang, Z. Genome-wide identification and characterization of the NF-Y gene family in grape (vitis vinifera L.). BMC Genom. 2016, 17, 1–16. [Google Scholar] [CrossRef]
- Yang, J.; Zhu, J.; Yang, Y. Genome-Wide Identification and Expression Analysis of NF-Y Transcription Factor Families in Watermelon (Citrullus lanatus). J. Plant Growth Regul. 2017, 36, 590–607. [Google Scholar] [CrossRef]
- Pereira, S.L.S.; Martins, C.P.S.; Sousa, A.O.; Camillo, L.R.; Araújo, C.P.; Alcantara, G.M.; Camargo, D.S.; Cidade, L.C.; De Almeida, A.A.F.; Costa, M.G.C. Genome-wide characterization and expression analysis of citrus NUCLEAR FACTOR-Y (NF-Y) transcription factors identified a novel NF-YA gene involved in drought-stress response and tolerance. PLoS ONE 2018, 13, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Ying, F.U.; Zhou, M. Genome-wide Identification and Expression Analysis of the NF-Y Family Genes in Phyllostachys edulis. J. Nucl. Agric. Sci. 2018, 32, 1513–1527. [Google Scholar]
- Xing, Y.; Fikes, J.D.; Guarente, L. Mutations in yeast HAP2/HAP3 define a hybrid CCAAT box binding domain. EMBO J. 1993, 12, 4647–4655. [Google Scholar] [CrossRef] [PubMed]
- Thön, M.; Abdallah, Q.A.; Hortschansky, P.; Scharf, D.H.; Eisendle, M.; Haas, H.; Brakhage, A.A. The CCAAT-binding complex coordinates the oxidative stress response in eukaryotes. Nucleic Acids Res. 2009, 38, 1098–1113. [Google Scholar] [CrossRef] [PubMed]
- Hackenberg, D.; Wu, Y.; Voigt, A.; Adams, R.; Schramm, P.; Grimm, B. Studies on differential nuclear translocation mechanism and assembly of the three subunits of the arabidopsis thaliana transcription factor NF-Y. Mol. Plant 2012, 5, 876–888. [Google Scholar] [CrossRef] [PubMed]
- Sinha, S.; Kim, I.S.; Sohn, K.Y.; de Crombrugghe, B.; Maity, S.N. Three classes of mutations in the A subunit of the CCAAT-binding factor CBF delineate functional domains involved in the three-step assembly of the CBF-DNA complex. Mol. Cell. Biol. 1996, 16, 328–337. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chen, X. microRNA biogenesis and function in plants. FEBS Lett. 2005, 579, 5923–5931. [Google Scholar] [CrossRef]
- Bombarely, A.; Moser, M.; Amrad, A.; Bao, M.; Bapaume, L.; Barry, C.S.; Bliek, M.; Boersma, M.R.; Borghi, L.; Bruggmann, R.; et al. Insight into the evolution of the Solanaceae from the parental genomes of Petunia hybrida. Nat. Plants 2016, 2, 1–9. [Google Scholar] [CrossRef]
- Xu, M.; Zhu, J.; Zhang, M.; Wang, L. Advances on plant miR169/NF-YA regulation modules. Yi Chuan Hereditas 2016, 38, 700–706. [Google Scholar]
- Zhao, B.; Ge, L.; Liang, R.; Wei, L.; Jin, Y. Members of miR-169 family are induced by high salinity and transiently inhibit the NF-YA transcription factor. BMC Mol. Biol. 2009, 10, 29. [Google Scholar] [CrossRef]
- Swain, S.; Myers, Z.A.; Siriwardana, C.L.; Holt, B.F. The multifaceted roles of NUCLEAR FACTOR-Y in Arabidopsis thaliana development and stress responses. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 636–644. [Google Scholar] [CrossRef]
- Zemzoumi, K.; Frontini, M.; Bellorini, M.; Mantovani, R.; Milano, Á. NF-Y Histone Fold a 1 Helices Help Impart CCAAT Specificity. J. Mol. Biol. 1999, 286, 327–337. [Google Scholar] [CrossRef]
- Zhang, T.; Zhang, D.; Liu, Y.; Luo, C.; Zhou, Y.; Zhang, L. Overexpression of a NF-YB3 transcription factor from Picea wilsonii confers tolerance to salinity and drought stress in transformed Arabidopsis thaliana. Plant Physiol. Biochem. 2015, 94, 153–164. [Google Scholar] [CrossRef]
- Mu, J.; Tan, H.; Hong, S.; Liang, Y.; Zuo, J. Arabidopsis transcription factor genes NF-YA1, 5, 6, and 9 play redundant roles in male gametogenesis, embryogenesis, and seed development. Mol. Plant 2013, 6, 188–201. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zou, Z.; Gong, P.; Zhang, J.; Ziaf, K.; Li, H.; Xiao, F.; Ye, Z. Over-expression of microRNA169 confers enhanced drought tolerance to tomato. Biotechnol. Lett. 2011, 33, 403–409. [Google Scholar] [CrossRef] [PubMed]
- Zhuo, X.; Zheng, T.; Zhang, Z.; Zhang, Y.; Jiang, L.; Ahmad, S.; Sun, L.; Wang, J.; Cheng, T.; Zhang, Q. Genome-wide analysis of the NAC transcription factor gene family reveals differential expression patterns and cold-stress responses in the woody plant prunus mume. Genes 2018, 9, 494. [Google Scholar] [CrossRef] [PubMed]
- Wei, Q.; Ma, C.; Xu, Y.; Wang, T.; Chen, Y.; Lü, J.; Zhang, L.; Jiang, C.Z.; Hong, B.; Gao, J. Control of chrysanthemum flowering through integration with an aging pathway. Nat. Commun. 2017, 8, 1–11. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Song, A.; An, J.; Guan, Z.; Jiang, J.; Chen, F.; Lou, W.; Fang, W.; Liu, Z.; Chen, S. The constitutive expression of a two transgene construct enhances the abiotic stress tolerance of chrysanthemum. Plant Physiol. Biochem. 2014, 80, 114–120. [Google Scholar] [CrossRef]
- Sun, D.; Nandety, R.S.; Zhang, Y.; Reid, M.S.; Niu, L.; Jiang, C.Z. A petunia ethylene-responsive element binding factor, PhERF2, plays an important role in antiviral RNA silencing. J. Exp. Bot. 2016, 67, 3353–3365. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; He, Y.; Xia, R. TBtools, a Toolkit for Biologists integrating various biological data handling tools with a user-friendly interface. BioRxiv 2018. [Google Scholar] [CrossRef]


| Gene ID | Gene Name | mRNA Length (bp) | No. of Introns | Peptide Residues (aa) | MW (kDa) | pI |
|---|---|---|---|---|---|---|
| Peaxi162Scf00004g02521 | PhNF-YA1 | 933 | 4 | 310 | 33.91 | 7.34 |
| Peaxi162Scf00339g00512 | PhNF-YA2 | 957 | 4 | 312 | 34.05 | 9.85 |
| Peaxi162Scf00038g01819 | PhNF-YA3 | 1065 | 4 | 327 | 36.13 | 9.85 |
| Peaxi162Scf00267g00038 | PhNF-YA4 | 792 | 4 | 286 | 31.70 | 9.37 |
| Peaxi162Scf00953g00217 | PhNF-YA5 | 861 | 5 | 298 | 33.54 | 9.33 |
| Peaxi162Scf00003g04130 | PhNF-YA6 | 585 | 2 | 354 | 39.76 | 6.94 |
| Peaxi162Scf00498g00617 | PhNF-YA7 | 897 | 5 | 318 | 34.72 | 10.04 |
| Peaxi162Scf00809g00019 | PhNF-YA8 | 987 | 4 | 263 | 29.51 | 9.93 |
| Peaxi162Scf00032g00323 | PhNF-YA9 | 939 | 4 | 296 | 33.26 | 7.47 |
| Peaxi162Scf00211g00239 | PhNF-YA10 | 888 | 6 | 194 | 22.19 | 10.39 |
| Peaxi162Scf00195g00082 | PhNF-YB1 | 492 | 3 | 163 | 17.54 | 5.32 |
| Peaxi162Scf00146g01325 | PhNF-YB2 | 420 | 2 | 140 | 15.58 | 5.29 |
| Peaxi162Scf00047g02011 | PhNF-YB3 | 606 | 0 | 201 | 20.52 | 6.68 |
| Peaxi162Scf00037g00085 | PhNF-YB4 | 423 | 0 | 140 | 15.90 | 4.56 |
| Peaxi162Scf00534g00022 | PhNF-YB5 | 420 | 0 | 139 | 15.71 | 4.89 |
| Peaxi162Scf00827g00055 | PhNF-YB6 | 642 | 0 | 213 | 23.69 | 5.10 |
| Peaxi162Scf00525g00067 | PhNF-YB7 | 696 | 0 | 231 | 26.03 | 6.95 |
| Peaxi162Scf00208g00014 | PhNF-YB8 | 534 | 3 | 177 | 19.36 | 5.73 |
| Peaxi162Scf00139g00134 | PhNF-YB9 | 681 | 1 | 226 | 25.43 | 6.79 |
| Peaxi162Scf00005g00493 | PhNF-YB10 | 546 | 4 | 181 | 19.68 | 6.79 |
| Peaxi162Scf00067g00163 | PhNF-YB11 | 501 | 0 | 166 | 18.71 | 8.06 |
| Peaxi162Scf00399g00108 | PhNF-YB12 | 510 | 1 | 169 | 19.32 | 6.94 |
| Peaxi162Scf01006g00035 | PhNF-YB13 | 588 | 0 | 195 | 21.66 | 6.54 |
| Peaxi162Scf00089g00023 | PhNF-YC1 | 732 | 0 | 243 | 26.20 | 5.03 |
| Peaxi162Scf00134g02029 | PhNF-YC2 | 873 | 1 | 291 | 32.20 | 6.08 |
| Peaxi162Scf00128g00124 | PhNF-YC3 | 513 | 0 | 170 | 18.04 | 4.25 |
| Peaxi162Scf00102g01519 | PhNF-YC4 | 327 | 1 | 108 | 11.92 | 10.25 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wei, Q.; Wen, S.; Lan, C.; Yu, Y.; Chen, G. Genome-Wide Identification and Expression Profile Analysis of the NF-Y Transcription Factor Gene Family in Petunia hybrida. Plants 2020, 9, 336. https://doi.org/10.3390/plants9030336
Wei Q, Wen S, Lan C, Yu Y, Chen G. Genome-Wide Identification and Expression Profile Analysis of the NF-Y Transcription Factor Gene Family in Petunia hybrida. Plants. 2020; 9(3):336. https://doi.org/10.3390/plants9030336
Chicago/Turabian StyleWei, Qian, Shiyun Wen, Chuying Lan, Yixun Yu, and Guoju Chen. 2020. "Genome-Wide Identification and Expression Profile Analysis of the NF-Y Transcription Factor Gene Family in Petunia hybrida" Plants 9, no. 3: 336. https://doi.org/10.3390/plants9030336
APA StyleWei, Q., Wen, S., Lan, C., Yu, Y., & Chen, G. (2020). Genome-Wide Identification and Expression Profile Analysis of the NF-Y Transcription Factor Gene Family in Petunia hybrida. Plants, 9(3), 336. https://doi.org/10.3390/plants9030336





