Estimating Genetic Conformism of Korean Mulberry Cultivars Using Random Amplified Polymorphic DNA and Inter-Simple Sequence Repeat Profiling
Abstract
1. Introduction
2. Material and Methods
2.1. Plant Material Collection
2.2. Genomic DNA Extraction
2.3. RAPD Assay
2.4. ISSR Assay
2.5. Marker Scoring and Statistical Analysis
3. Results
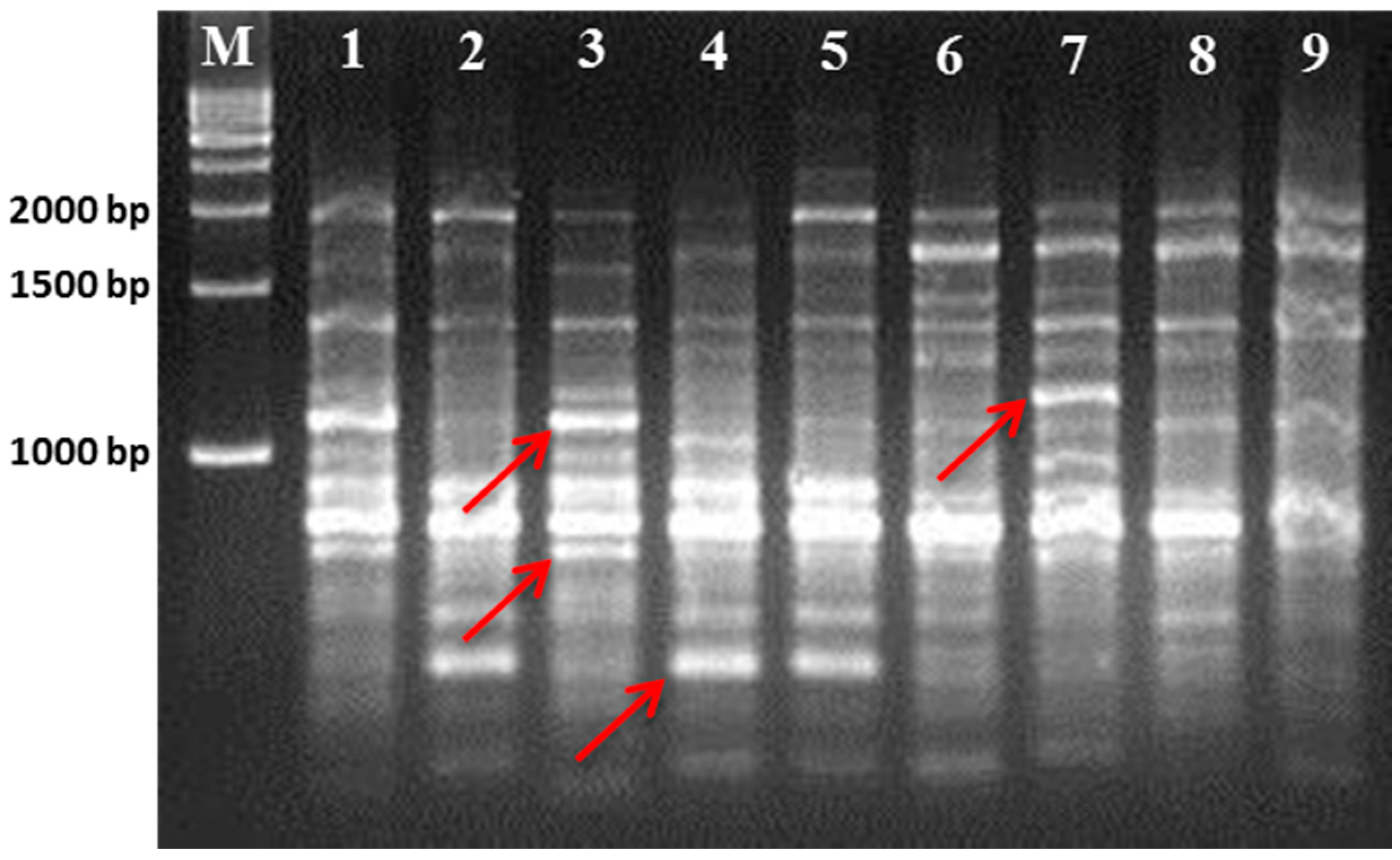
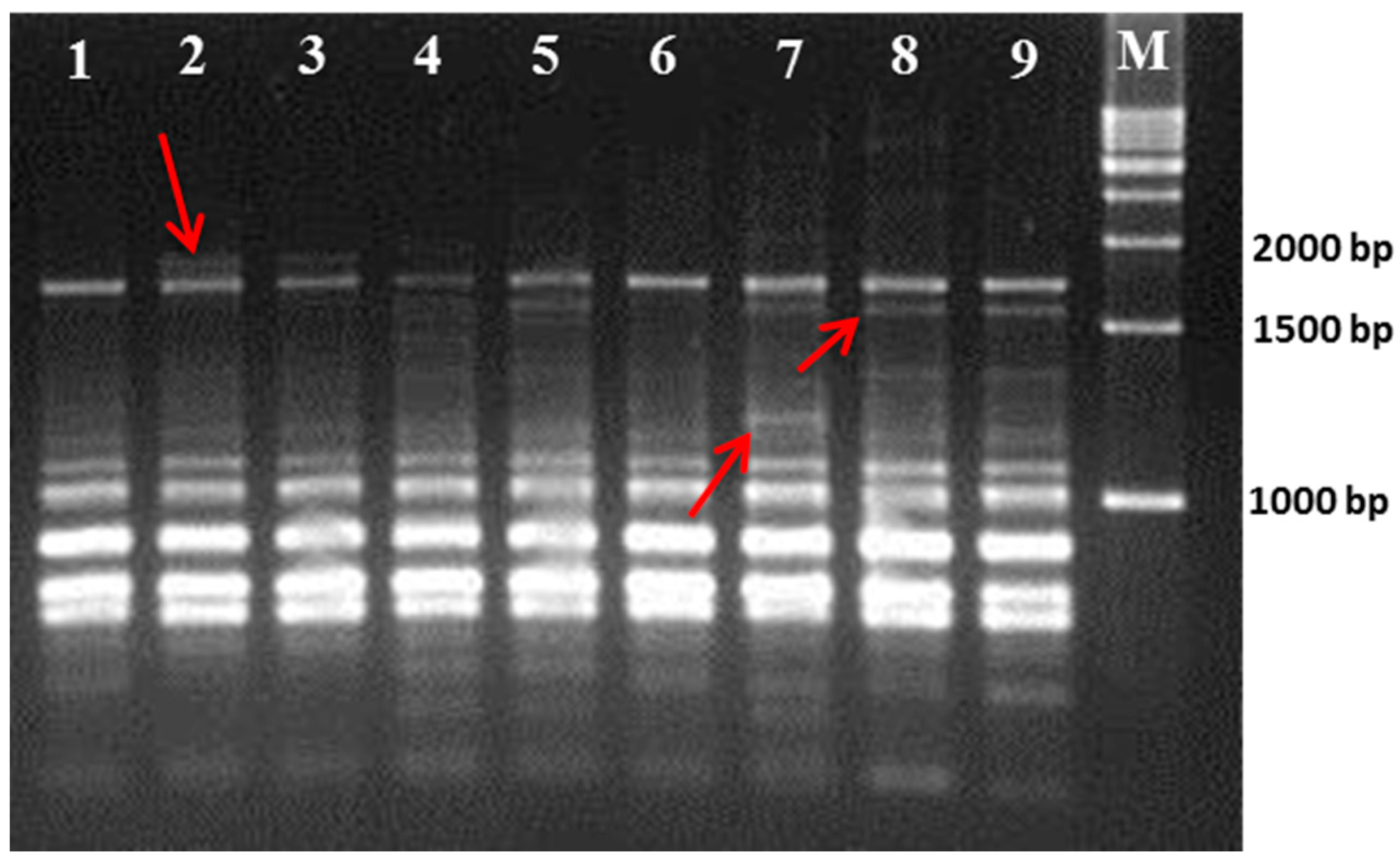
3.1. Polymorphism Displayed by the RAPD and ISSR Markers
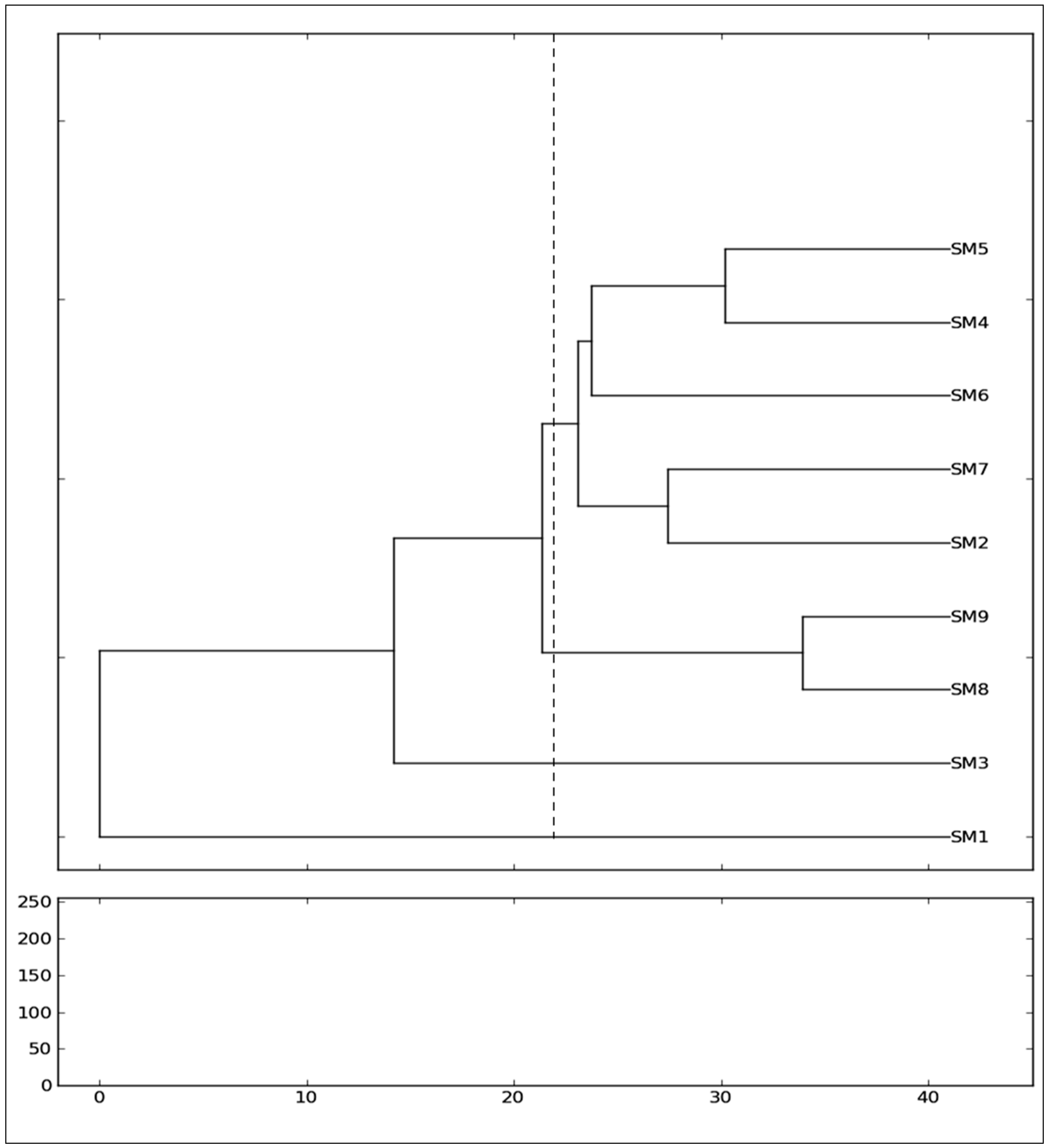
3.2. Genetic Variance and Clustering Analysis between the Genotypes
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of interest
References
- Watt, G. A Dictionary of the Economic Products of India; Cambridge University Press: Cambridge, UK, 2014. [Google Scholar]
- Vavilov, N.I. Studies on the Origin of Cultivated Plants; Leningrad Press: St. Petersburg, Russia, 1926. [Google Scholar]
- Vijayan, K.; Raju, P.J.; Tikader, A.; Saratchnadra, B. Biotechnology of mulberry (Morus L.)—A review. Emir. J. Food Agric. 2014, 26, 472. [Google Scholar] [CrossRef]
- Kalpana, D.; Choi, S.H.; Choi, T.K.; Senthil, K.; Lee, Y.S. Assessment of genetic diversity among varieties of mulberry using RAPD and ISSR fingerprinting. Sci. Hortic. 2012, 134, 79–87. [Google Scholar] [CrossRef]
- Rao, D.M.R.; Jhansilakshmi, K.; Saraswathi, P.; Rao, A.A.; Ramesh, S.; Borpuzari, M.; Manjula, A. Scope of pre-breeding in mulberry crop improvement—A review. Sci. Wkly. 2013, 1, 1–18. [Google Scholar]
- Vijayan, K.; Tikader, A.; Kar, P.; Srivastava, P.; Awasthi, A.; Thangavelu, K.; Saratchandra, B. Assessment of genetic relationships between wild and cultivated mulberry (Morus) species using PCR based markers. Genet. Resour. Crop Evol. 2006, 53, 873–882. [Google Scholar] [CrossRef]
- Schlötterer, C. The evolution of molecular markers—Just a matter of fashion? Nat. Rev. Genet. 2004, 5, 63. [Google Scholar] [PubMed]
- Koidzumi, G. Taxonomical discussion on Morus plants. Bull. Imp. Sericult. Exp. Stat. 1917, 3, 1–62. [Google Scholar]
- Zhou, Z.; Gilbert, M.G. Moraceae. Flora China 2003, 5, 21–73. [Google Scholar]
- Awasthi, A.K.; Nagaraja, G.; Naik, G.; Kanginakudru, S.; Thangavelu, K.; Nagaraju, J. Genetic diversity and relationships in mulberry (genus Morus) as revealed by RAPD and ISSR marker assays. BMC Genet. 2004, 5, 1. [Google Scholar] [CrossRef] [PubMed]
- Vijayan, K.; Kar, P.; Tikader, A.; Srivastava, P.; Awasthi, A.; Thangavelu, K.; Saratchandra, B. Molecular evaluation of genetic variability in wild populations of mulberry (Morus serrata Roxb.). Plant Breed. 2004, 123, 568–572. [Google Scholar] [CrossRef]
- Roldàn-Ruiz, I.; Calsyn, E.; Gilliland, T.; Coll, R.; Van Eijk, M.; De Loose, M. Estimating genetic conformity between related ryegrass (Lolium) varieties. 2. AFLP characterization. Mol. Breed. 2000, 6, 593–602. [Google Scholar] [CrossRef]
- Nei, M.; Li, W.-H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA 1979, 76, 5269–5273. [Google Scholar] [CrossRef] [PubMed]
- Marialouis, X.A.; Santhanam, A. Antibiotic resistance, RAPD-PCR typing of multiple drug resistant strains of Escherichia coli from urinary tract infection (UTI). J. Clin. Diagn. Res. JCDR 2016, 10, DC05. [Google Scholar] [CrossRef] [PubMed]
- Pavel, A.B.; Vasile, C.I. PyElph-a software tool for gel images analysis and phylogenetics. BMC BIoinform. 2012, 13, 9. [Google Scholar] [CrossRef] [PubMed]
- Saber, M.; AMA, A.; Rahman, T.G.A.; Alsaidi, K. Genetic diversity of fusarium species causing potato dry rot disease using random amplified polymorphic DNA (rapd-pcr). Environ. Sci. 2017, 12, 237–247. [Google Scholar]
- Srivastava, P.P.; Vijayan, K.; Awasthi, A.K.; Saratchandra, B. Genetic Analysis of Morus alba through RAPD and ISSR Markers; CSIR: Cape Town, South Africa, 2004. [Google Scholar]
- He, N.; Zhang, C.; Qi, X.; Zhao, S.; Tao, Y.; Yang, G.; Lee, T.-H.; Wang, X.; Cai, Q.; Li, D. Draft genome sequence of the mulberry tree Morus notabilis. Nat. Commun. 2013, 4, 2445. [Google Scholar] [CrossRef] [PubMed]
- Yile, P.; Zhifang, Z.; Xuexia, M. Phylogeny of the genus Morus (Urticales: Moraceae) inferred from ITS and trnL-F sequences. Afr. J. Biotechnol. 2005, 4, 563. [Google Scholar]
- Biasiolo, M.; Da Canal, M.T.; Tornadore, N. Micromorphological Characterization of Ten Mulberry Cultivars (Morus spp.). Econ. Bot. 2004, 58, 639–646. [Google Scholar] [CrossRef]
- Gupta, S.; Srivastava, M.; Mishra, G.; Naik, P.; Chauhan, R.; Tiwari, S.; Kumar, M.; Singh, R. Analogy of ISSR and RAPD markers for comparative analysis of genetic diversity among different Jatropha curcas genotypes. Afr. J. Biotechnol. 2008, 7, 4230–4243. [Google Scholar]
- Vijayan, K.; Chatterjee, S. ISSR profiling of Indian cultivars of mulberry (Morus spp.) and its relevance to breeding programs. Euphytica 2003, 131, 53–63. [Google Scholar] [CrossRef]
- Manimekalai, R.; Nagarajan, P. Interrelationships among coconut (Cocos nucifera L.) accessions using RAPD technique. Genet. Resour. Crop Evol. 2006, 53, 1137–1144. [Google Scholar] [CrossRef]
- Medhi, K.; Sarmah, D.; Deka, M.; Bhau, B. High gene flow and genetic diversity in three economically important Zanthoxylum Spp. of Upper Brahmaputra Valley Zone of NE India using molecular markers. Meta Gene 2014, 2, 706–721. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Wang, Y.; Chen, T.; Jia, G.; Wang, X.; Qi, J.; Pang, Y.; Wang, S.; Li, Z.; Huang, Y. Genetic structure of mulberry from different ecotypes revealed by ISSRs in China: An implications for conservation of local mulberry varieties. Sci. Hortic. 2007, 115, 47–55. [Google Scholar] [CrossRef]
- Slatkin, M. Gene flow and the geographic structure of natural populations. Science 1987, 236, 787–792. [Google Scholar] [CrossRef] [PubMed]
- Tilwari, A.; Chauhan, D.; Sharma, R.; Singh, R. Assessment of Genetic Variations among Medicinal Plant Cassia tora from Different Geographic Regions of Central India Using RAPD Markers. Med. Aromat. Plants (Los Angel.) 2016. [Google Scholar] [CrossRef]
| Sample No | Species | Morphological Characters | Leaf Size (cm) | Cultivar | |
|---|---|---|---|---|---|
| Width | Length | ||||
| SM1 | M. lhou | Leaf bluntly tipped, cros-venulate, margin serrate, rough surface, unlobed. | 9.63 ± 3.12 | 12.04± 1.20 | Dae-Saug |
| SM2 | M. alba | Leaf ovate with serrate, often lobed, small smooth base. | 10.33 ± 1.97 | 13.3 ± 0.57 | Chung IL |
| SM3 | M. alba | Leaf slightly cordate, margin serrate, lobed, comparatively smooth base, cros-venulate. | 12.85 ± 0.27 | 15.3 ± 1.87 | Hong-ol |
| SM4 | M. alba | Leaf ovate with often lobed and serrate, cros-venulate, rough surface. | 8.50 ± 2.42 | 10.3 ± 1.33 | Gae-ryang |
| SM5 | M. alba | Leaf bluntly tipped with small point, comparatively big rough surface, cros-venulate, unlobed, margin serrate. | 16.20 ± 0.57 | 18.3 ± 1.22 | SangchonJo Sang Saeng |
| SM6 | M. alba | Elliptical leaf, unlobed, margin serrate, less rough surface, comparatively deep green, cros-venulate. | 11.85 ± 2.27 | 12.87 ± 1.1 | Soo hyang |
| SM7 | M. alba | Elliptical leaf, unlobed, margin serrate, less rough surface, cros-venulate. | 12.34 ± 1.46 | 15.3 ± 0.37 | Gwasang II |
| SM8 | M. alba | Leaf ovate with lobed and serrate, cros-venulate, rough surface. | 9.42 ± 0.79 | 13.5 ± 2.04 | Soosang |
| SM9 | M. alba | Leaf slightly ovate with unlobed and serrate, cros-venulate, rough surface. | 11.78 ± 1.70 | 13.4 ± 0.50 | Hong-Saug |
| Primer Name | Sequence (5ʹ–3ʹ) | Polymorphic Band | Polymorphism (%) | PIC |
|---|---|---|---|---|
| OPA-02 | TGCCGAGCTG | 4 | 63.7 | 0.211 |
| OPA-06 | GGTCCCTGAC | 10 | 90 | 0.335 |
| OPA-07 | GAAACGGGTG | 7 | 71.82 | 0.415 |
| OPA-14 | TCTGTGCTGG | 3 | 62.5 | 0.337 |
| OPA-15 | TTCCGAACCC | 4 | 67.76 | 0.352 |
| OPA-17 | GACCGCTTGT | 8 | 100 | 0.601 |
| OPA-18 | AGGTGACCGT | 3 | 44.7 | 0.173 |
| OPY-07 | AGAGCCGTCA | 9 | 100 | 0.792 |
| OPY-15 | AGTCGCCCTT | 7 | 91.3 | 0.577 |
| OPY-20 | AGCCGTGGAA | 7 | 100 | 0.775 |
| Primer Name | Sequence 5ʹ–3ʹ | Polymorphic Band | Polymorphism (%) | PIC |
|---|---|---|---|---|
| UBC-807 | AGAGAGAGAGAGAGAGT | 3 | 60 | 0.411 |
| UBC-808 | AGAGAGAGAGAGAGAGC | 5 | 50 | 0.335 |
| UBC-809 | AGAGAGAGAGAGAGAGG | 2 | 72.12 | 0.615 |
| UBC-810 | GAGAGAGAGAGAGAGAT | 5 | 62.5 | 0.567 |
| UBC-814 | CTCTCTCTCTCTCTCTA | 7 | 77.15 | 0.652 |
| UBC-815 | CTCTCTCTCTCTCTCTG | 9 | 45.2 | 0.287 |
| UBC-817 | CACACACACACACACAA | 6 | 70 | 0.603 |
| UBC-820 | GTGTGTGTGTGTGTGTC | 3 | 100 | 0.753 |
| UBC-824 | TCTCTCTCTCTCTCTCG | 5 | 80.2 | 0.632 |
| UBC-825 | ACACACACACACACACT | 7 | 81.3 | 0.645 |
| Genotypes | SM1 | SM2 | SM3 | SM4 | SM5 | SM6 | SM7 | SM8 | SM9 |
|---|---|---|---|---|---|---|---|---|---|
| SM1 | 0 | - | - | - | - | - | - | - | - |
| SM2 | 0.733 | 0 | - | - | - | - | - | - | - |
| SM3 | 0.780 | 0.714 | 0 | - | - | - | - | - | - |
| SM4 | 0.620 | 0.647 | 0.685 | 0 | - | - | - | - | - |
| SM5 | 0.650 | 0.669 | 0.714 | 0.566 | 0 | - | - | - | - |
| SM6 | 0.714 | 0.658 | 0.833 | 0.700 | 0.727 | 0 | - | - | - |
| SM7 | 0.685 | 0.633 | 0.738 | 0.737 | 0.633 | 0.790 | 0 | - | - |
| SM8 | 0.766 | 0.620 | 0.662 | 0.592 | 0.620 | 0.678 | 0.727 | 0 | - |
| SM9 | 0.452 | 0.460 | 0.592 | 0.361 | 0.564 | 0.614 | 0.589 | 0.483 | 0 |
| Genotypes | SM1 | SM2 | SM3 | SM4 | SM5 | SM6 | SM7 | SM8 | SM9 |
|---|---|---|---|---|---|---|---|---|---|
| SM1 | 0 | - | - | - | - | - | - | - | - |
| SM2 | 0.153 | 0 | - | - | - | - | - | - | - |
| SM3 | 0.222 | 0.588 | 0 | - | - | - | - | - | - |
| SM4 | 0.526 | 0.444 | 0.521 | 0 | - | - | - | - | - |
| SM5 | 0.470 | 0.500 | 0.571 | 0.727 | 0 | - | - | - | - |
| SM6 | 0.235 | 0.625 | 0.381 | 0.545 | 0.601 | 0 | - | - | - |
| SM7 | 0.285 | 0.600 | 0.480 | 0.535 | 0.583 | 0.750 | 0 | - | - |
| SM8 | 0.526 | 0.556 | 0.527 | 0.665 | 0.636 | 0.545 | 0.615 | 0 | - |
| SM9 | 0.434 | 0.545 | 0.667 | 0.642 | 0.615 | 0.538 | 0.666 | 0.714 | 0 |
| Genotypes | SM1 | SM2 | SM3 | SM4 | SM5 | SM6 | SM7 | SM8 | SM9 |
|---|---|---|---|---|---|---|---|---|---|
| SM1 | 0 | - | - | - | - | - | - | - | - |
| SM2 | 0.253 | 0 | - | - | - | - | - | - | - |
| SM3 | 0.230 | 0.310 | 0 | - | - | - | - | - | - |
| SM4 | 0.226 | 0.244 | 0.221 | 0 | - | - | - | - | - |
| SM5 | 0.183 | 0.270 | 0.271 | 0.747 | 0 | - | - | - | - |
| SM6 | 0.195 | 0.425 | 0.312 | 0.635 | 0.621 | 0 | - | - | - |
| SM7 | 0.315 | 0.678 | 0.327 | 0.535 | 0.583 | 0.650 | 0 | - | - |
| SM8 | 0.422 | 0.556 | 0.527 | 0.465 | 0.436 | 0.345 | 0.315 | 0 | - |
| SM9 | 0.434 | 0.545 | 0.567 | 0.442 | 0.415 | 0.338 | 0.366 | 0.814 | 0 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sheet, S.; Ghosh, K.; Acharya, S.; Kim, K.-P.; Lee, Y.S. Estimating Genetic Conformism of Korean Mulberry Cultivars Using Random Amplified Polymorphic DNA and Inter-Simple Sequence Repeat Profiling. Plants 2018, 7, 21. https://doi.org/10.3390/plants7010021
Sheet S, Ghosh K, Acharya S, Kim K-P, Lee YS. Estimating Genetic Conformism of Korean Mulberry Cultivars Using Random Amplified Polymorphic DNA and Inter-Simple Sequence Repeat Profiling. Plants. 2018; 7(1):21. https://doi.org/10.3390/plants7010021
Chicago/Turabian StyleSheet, Sunirmal, Kuntal Ghosh, Satabdi Acharya, Kwang-Pyo Kim, and Yang Soo Lee. 2018. "Estimating Genetic Conformism of Korean Mulberry Cultivars Using Random Amplified Polymorphic DNA and Inter-Simple Sequence Repeat Profiling" Plants 7, no. 1: 21. https://doi.org/10.3390/plants7010021
APA StyleSheet, S., Ghosh, K., Acharya, S., Kim, K.-P., & Lee, Y. S. (2018). Estimating Genetic Conformism of Korean Mulberry Cultivars Using Random Amplified Polymorphic DNA and Inter-Simple Sequence Repeat Profiling. Plants, 7(1), 21. https://doi.org/10.3390/plants7010021




