The Ubiquitin System and Jasmonate Signaling
Abstract
:1. Introduction to Jasmonates
2. Introduction to the Ub System
2.1. The Ub Conjugation Pathway: E1-E2-E3
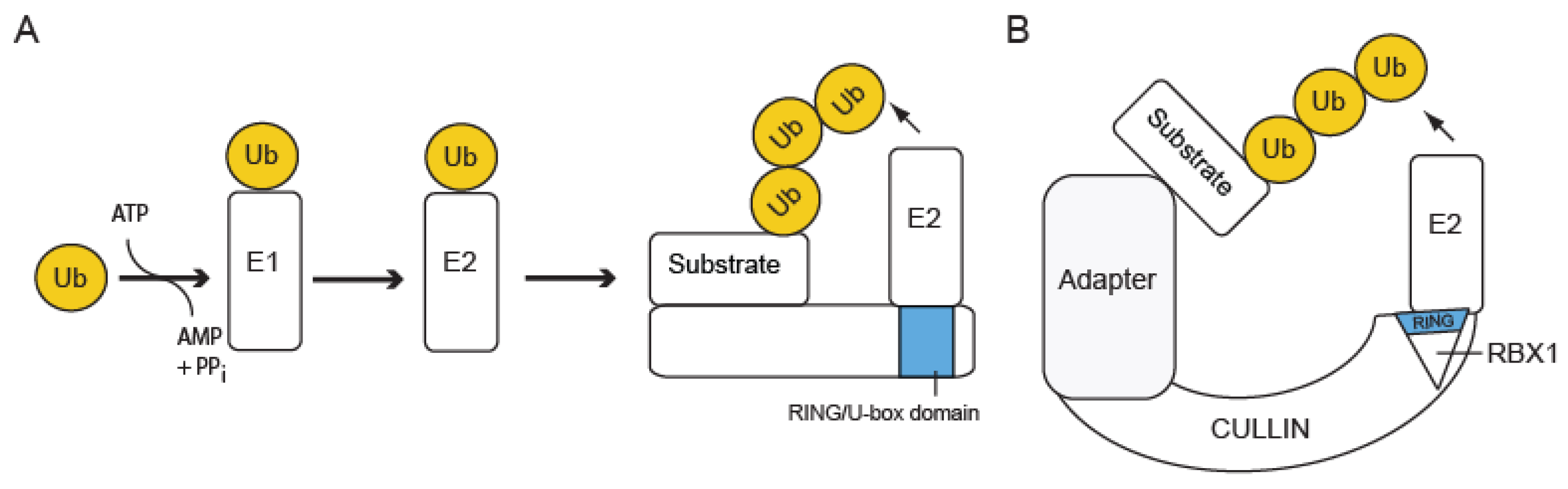
2.2. Classes of Ub Ligases (E3s)
2.3. Versatility in the Ubiquitin System
2.4. The 26S Proteasome
2.5. Ubiquitin-Like Modifiers
3. Mechanism and Importance of Ubiquitination of JAZ Proteins in JA Signaling
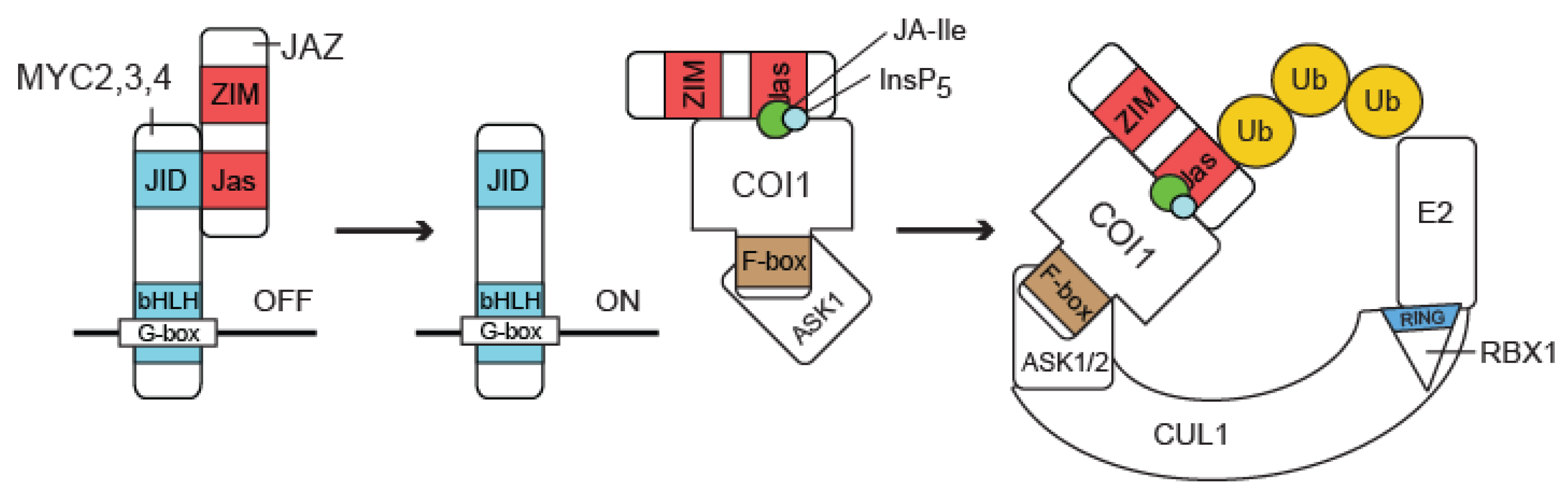
3.1. COI1 Links the Ubiquitin System to JA Signaling
3.2. JAZ Proteins Repress JA Signaling

3.3. MYC2 Regulates the Transcription of JA-Responsive Genes
4. Ubiquitination in JA Signaling beyond the Core Module
| Protein or Protein Complex | Ub-System Class | Candidate Target | Target Type | Evidence | Process | Reference |
|---|---|---|---|---|---|---|
| SCFCOI1 | Ub E3: CRL | JAZ | repressor | Binding, stability, genetic | JA signaling | [55,56] |
| DAF | Ub E3: RING | DAD1 | enzyme | Hypothetical | JA biosynthesis | [17,94] |
| KEG | Ub E3: | ABI5 | TF | Binding, stability, in vitro assay, genetic | ABA signaling, JA signaling | [67,95,96,97] |
| RING | ABF1 | TF | ||||
| ABF3 | TF | |||||
| RGLG3 RGLG4 | Ub E3: RING | - | - | - | JA signaling, SA-JA crosstalk | [98,99] |
| PUB10 | Ub E3: U-box | MYC2 | TF | In vitro assay | JA signaling | [100] |
| PUB10/11 | Ub E3: U-box | MYC3 MYC4 | TF | Binding, stability | JA signaling | [100] |
| BOI | Ub E3: RING | BOS1/MYB108 | TF | Binding, stability, in vitro assay | Cell death JA signaling | [101,102,103] |
| MtMKB1 | Ub E3: RING | MtHMGR | enzyme | Binding, stability | ERAD JA signaling | [104] |
| OsHOS1 | Ub E3: RING | OsEREBP1 OsEREBP2 | TFs | JA signaling, mechanosensing | [105] | |
| At/OsHOS1 | Ub E3: RING | At/OsICE1 | TF | JA signaling, abiotic stress responses | [106,107] | |
| - | - | JAV1 | - | Stability | JA signaling | [108] |
| - | - | ORA59 | TF | Stability | SA-JA crosstalk | [109] |
| TIC | - | MYC2 | TF | Binding, stability | JA signaling, circadian rhythm | [110] |
| UBP12 UBP13 | DUBs | TIFY8 | Transcrip-tional regulator | Binding | biotic stress responses, photoperiodicity | [111,112,113] |
| AXR1/AXL /ECR1 | Nedd8 E1 | CUL | CRL component | Genetic, stability | CRL activity regulation, JA signaling | [34,35,114] |
| RCE1/2 | Nedd8 E2 | CUL | CRL component | Genetic | CRL activity regulation, JA signaling | [36,115] |
| CSN | deneddylase | CUL | CRL component | Genetic | CRL activity regulation, JA signaling | [116] |
| SIZ1 | SUMO E3 | multiple | multiple | Genetic | Pleiotropic, JA signaling | [117] |
4.1. The Ub E3 Ligase DAF Is Involved in JA Biosynthesis
4.2. The Ub E3 Ligase KEG Interacts with JAZ12
4.3. The Ub E3 Ligases RGLG3/4 Positively Regulate JA Signaling
4.4. The Ub E3 Ligase PUB10 Regulates MYC2 Protein Levels
4.5. The Ub E3 Ligase BOI and Its Target BOS1 Link Cell Death to JA Signaling
4.6. The Ub E3 Ligase MtMKB1 Links JA Signaling to Protein Quality Control
4.7. The Ub E3 Ligase HOS1 Links JA Signaling to Multiple Developmental and Abiotic Stress Responses
4.8. JAV1 Protein Stability Is Regulated by COI1-Dependent JA Perception
4.9. Protein Stability of the Transcription Factor ORA59 Affects JA-SA Crosstalk
4.10. Light Signaling Alters JAZ Protein Stability through Gibberellic Acid (GA) Signaling
4.11. TIC Synchronizes JA Signaling to the Circadian Clock
4.12. A Possible Role for Deubiquitination in JA Signaling
4.13. Contribution of UBL-Mediated Modifications to JA Signaling
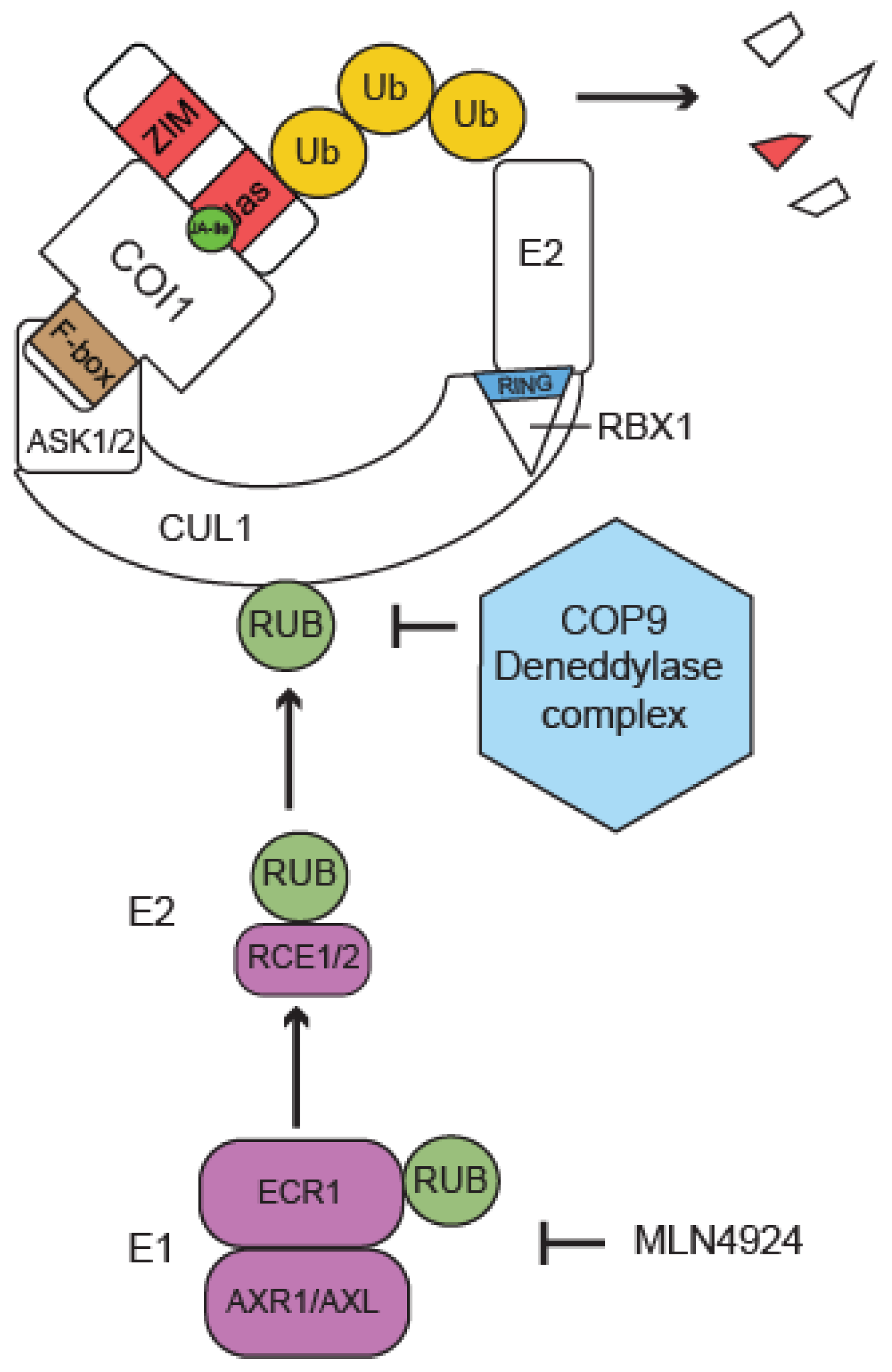
5. Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Browse, J. Jasmonate passes muster: A receptor and targets for the defense hormone. Annu. Rev. Plant Biol. 2009, 60, 183–205. [Google Scholar] [CrossRef] [PubMed]
- Wasternack, C.; Hause, B. Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 2013, 111, 1021–1058. [Google Scholar] [CrossRef] [PubMed]
- Wasternack, C. Action of jasmonates in plant stress responses and development—Applied aspects. Biotechnol. Adv. 2014, 32, 31–39. [Google Scholar] [CrossRef] [PubMed]
- De Geyter, N.; Gholami, A.; Goormachtig, S.; Goossens, A. Transcriptional machineries in jasmonate-elicited plant secondary metabolism. Trends Plant Sci. 2012, 17, 349–359. [Google Scholar] [CrossRef] [PubMed]
- Glazebrook, J. Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu. Rev. Phytopathol. 2005, 43, 205–227. [Google Scholar] [CrossRef] [PubMed]
- Ballaré, C.L. Light regulation of plant defense. Annu. Rev. Plant Biol. 2014, 65, 335–363. [Google Scholar] [CrossRef] [PubMed]
- Pauwels, L.; Inzé, D.; Goossens, A. Jasmonate-inducible gene: What does it mean? Trends Plant Sci. 2009, 14, 87–91. [Google Scholar] [CrossRef] [PubMed]
- Browse, J.; Howe, G.A. New weapons and a rapid response against insect attack. Plant Physiol. 2008, 146, 832–838. [Google Scholar] [CrossRef] [PubMed]
- Wasternack, C. Jasmonates: An update on biosynthesis, signal transduction and action in plant stress response, growth and development. Ann. Bot. 2007, 100, 681–697. [Google Scholar] [CrossRef] [PubMed]
- Staswick, P.E.; Tiryaki, I. The oxylipin signal jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell 2004, 16, 2117–2127. [Google Scholar] [CrossRef] [PubMed]
- Westfall, C.S.; Zubieta, C.; Herrmann, J.; Kapp, U.; Nanao, M.H.; Jez, J.M. Structural basis for prereceptor modulation of plant hormones by GH3 proteins. Science 2012, 336, 1708–1711. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, S.; Chini, A.; Hamberg, M.; Adie, B.; Porzel, A.; Kramell, R.; Miersch, O.; Wasternack, C.; Solano, R. (+)-7-iso-Jasmonoyl-L-isoleucine is the endogenous bioactive jasmonate. Nat. Chem. Biol. 2009, 5, 344–350. [Google Scholar] [CrossRef] [PubMed]
- Katsir, L.; Schilmiller, A.L.; Staswick, P.E.; He, S.Y.; Howe, G.A. COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. Proc. Natl. Acad. Sci. USA 2008, 105, 7100–7105. [Google Scholar] [CrossRef] [PubMed]
- Callis, J.; Pollmann, L.; Shanklin, J.; Wettern, M.; Vierstra, R.D. Sequence of a cDNA from Chlamydomonas reinhardii encoding a ubiquitin 52 amino acid extension protein. Nucleic Acids Res. 1989, 17, 8377. [Google Scholar] [CrossRef] [PubMed]
- Ciechanover, A.; Elias, S.; Heller, H.; Ferber, S.; Hershko, A. Characterization of the heat-stable polypeptide of the ATP-dependent proteolytic system from reticulocytes. J. Biol. Chem. 1980, 255, 7525–7528. [Google Scholar] [PubMed]
- Smalle, J.; Vierstra, R.D. The ubiquitin 26S proteasome proteolytic pathway. Annu. Rev. Plant Biol. 2004, 55, 555–590. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.-J.; Shih, C.-F.; Yang, J.-Y.; Tan, C.M.; Hsu, W.-H.; Huang, Y.-P.; Liao, P.-C.; Yang, C.-H. A RING-type E3 ligase controls anther dehiscence by activating the jasmonate biosynthetic pathway gene DEFECTIVE IN ANTHER DEHISCENCE1 in Arabidopsis. Plant J. 2013, 74, 310–327. [Google Scholar] [CrossRef] [PubMed]
- Kraft, E.; Stone, S.L.; Ma, L.; Su, N.; Gao, Y.; Lau, O.-S.; Deng, X.-W.; Callis, J. Genome analysis and functional characterization of the E2 and RING-type E3 ligase ubiquitination enzymes of Arabidopsis. Plant Physiol. 2005, 139, 1597–1611. [Google Scholar] [CrossRef] [PubMed]
- Hua, Z.; Vierstra, R.D. The cullin-RING ubiquitin-protein ligases. Annu. Rev. Plant Biol. 2011, 62, 299–334. [Google Scholar] [CrossRef] [PubMed]
- Callis, J. The ubiquitination machinery of the ubiquitin system. Arabidopsis Book 2014. [Google Scholar] [CrossRef] [PubMed]
- Stone, S.L.; Hauksdóttir, H.; Troy, A.; Herschleb, J.; Kraft, E.; Callis, J. Functional analysis of the RING-type ubiquitin ligase family of Arabidopsis. Plant Physiol. 2005, 137, 13–30. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Shen, W.-H. Dynamic regulation and function of histone monoubiquitination in plants. Front. Plant Sci. 2014, 5, 83. [Google Scholar] [CrossRef] [PubMed]
- Dikic, I.; Wakatsuki, S.; Walters, K.J. Ubiquitin-binding domains—From structures to functions. Nat. Rev. Mol. Cell Biol. 2009, 10, 659–671. [Google Scholar] [CrossRef] [PubMed]
- Meierhofer, D.; Wang, X.; Huang, L.; Kaiser, P. Quantitative analysis of global ubiquitination in HeLa cells by mass spectrometry. J. Proteome Res. 2008, 7, 4566–4576. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Duong, D.M.; Seyfried, N.T.; Cheng, D.; Xie, Y.; Robert, J.; Rush, J.; Hochstrasser, M.; Finley, D.; Peng, J. Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation. Cell 2009, 137, 133–145. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.-Y.; Scalf, M.; Smith, L.M.; Vierstra, R.D. Advanced proteomic analyses yield a deep catalog of ubiquitylation targets in Arabidopsis. Plant Cell 2013, 25, 1523–1540. [Google Scholar] [CrossRef] [PubMed]
- Fu, H.; Sadis, S.; Rubin, D.M.; Glickman, M.; van Nocker, S.; Finley, D.; Vierstra, R.D. Multiubiquitin chain binding and protein degradation are mediated by distinct domains within the 26 S proteasome subunit Mcb1. J. Biol. Chem. 1998, 273, 1970–1981. [Google Scholar] [CrossRef] [PubMed]
- Leitner, J.; Petrášek, J.; Tomanov, K.; Retzer, K.; Pařezová, M.; Korbei, B.; Bachmair, A.; Zažímalová, E.; Luschnig, C. Lysine63-linked ubiquitylation of PIN2 auxin carrier protein governs hormonally controlled adaptation of Arabidopsis root growth. Proc. Natl. Acad. Sci. USA 2012, 109, 8322–8627. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Schmidt, W. A lysine-63-linked ubiquitin chain-forming conjugase, UBC13, promotes the developmental responses to iron deficiency in Arabidopsis roots. Plant J. 2010, 62, 330–343. [Google Scholar] [CrossRef] [PubMed]
- Heinemeyer, W.; Fischer, M.; Krimmer, T.; Stachon, U.; Wolf, D.H. The active sites of the eukaryotic 20 S proteasome and their involvement in subunit precursor processing. J. Biol. Chem. 1997, 272, 25200–25209. [Google Scholar] [CrossRef] [PubMed]
- Vierstra, R.D. The ubiquitin-26S proteasome system at the nexus of plant biology. Nat. Rev. Mol. Cell Biol. 2009, 10, 385–397. [Google Scholar] [CrossRef] [PubMed]
- Vierstra, R.D. The expanding universe of ubiquitin and ubiquitin-like modifiers. Plant Physiol. 2012, 160, 2–14. [Google Scholar] [CrossRef] [PubMed]
- Bostick, M.; Lochhead, S.R.; Honda, A.; Palmer, S.; Callis, J. Related to ubiquitin 1 and 2 are redundant and essential and regulate vegetative growth, auxin signaling, and ethylene production in Arabidopsis. Plant Cell 2004, 16, 2418–2432. [Google Scholar] [CrossRef] [PubMed]
- Del Pozo, J.C.; Dharmasiri, S.; Hellmann, H.; Walker, L.; Gray, W.M.; Estelle, M. AXR1-ECR1-dependent conjugation of RUB1 to the Arabidopsis Cullin AtCUL1 is required for auxin response. Plant Cell 2002, 14, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Dharmasiri, N.; Dharmasiri, S.; Weijers, D.; Karunarathna, N.; Jurgens, G.; Estelle, M. AXL and AXR1 have redundant functions in RUB conjugation and growth and development in Arabidopsis. Plant J. 2007, 52, 114–123. [Google Scholar] [CrossRef] [PubMed]
- Del Pozo, J.C.; Estelle, M. The Arabidopsis cullin AtCUL1 is modified by the ubiquitin-related protein RUB1. Proc. Natl. Acad. Sci. USA 1999, 96, 15342–15347. [Google Scholar] [CrossRef] [PubMed]
- Kamura, T.; Conrad, M.N.; Yan, Q.; Conaway, R.C.; Conaway, J.W. The Rbx1 subunit of SCF and VHL E3 ubiquitin ligase activates Rub1 modification of cullins Cdc53 and Cul2. Genes Dev. 1999, 13, 2928–2933. [Google Scholar] [CrossRef] [PubMed]
- Morimoto, M.; Nishida, T.; Nagayama, Y.; Yasuda, H. Nedd8-modification of Cul1 is promoted by Roc1 as a Nedd8-E3 ligase and regulates its stability. Biochem. Biophys. Res. Commun. 2003, 301, 392–398. [Google Scholar] [CrossRef]
- Enchev, R.I.; Schulman, B.A.; Peter, M. Protein neddylation: beyond cullin-RING ligases. Nat. Rev. Mol. Cell Biol. 2015, 16, 30–44. [Google Scholar] [CrossRef] [PubMed]
- Mergner, J.; Schwechheimer, C. The NEDD8 modification pathway in plants. Front. Plant Sci. 2014, 5, 103. [Google Scholar] [CrossRef] [PubMed]
- Mergner, J.; Heinzlmeir, S.; Kuster, B.; Schwechheimer, C. DENEDDYLASE1 deconjugates NEDD8 from non-cullin protein substrates in Arabidopsis thaliana. Plant Cell 2015, 27, 741–753. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Schwechheimer, C.; Isono, E. The COP9 signalosome and its role in plant development. Eur. J. Cell Biol. 2010, 89, 157–162. [Google Scholar] [CrossRef] [PubMed]
- Saracco, S.A.; Miller, M.J.; Kurepa, J.; Vierstra, R.D. Genetic analysis of SUMOylation in Arabidopsis: conjugation of SUMO1 and SUMO2 to nuclear proteins is essential. Plant Physiol. 2007, 145, 119–134. [Google Scholar] [CrossRef] [PubMed]
- Novatchkova, M.; Tomanov, K.; Hofmann, K.; Stuible, H.-P.; Bachmair, A. Update on sumoylation: defining core components of the plant SUMO conjugation system by phylogenetic comparison. New Phytol. 2012, 195, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Melchior, F. SUMO—nonclassical ubiquitin. Annu. Rev. Cell Dev. Biol. 2000, 16, 591–626. [Google Scholar] [CrossRef] [PubMed]
- Kurepa, J.; Walker, J.M.; Smalle, J.; Gosink, M.M.; Davis, S.J.; Durham, T.L.; Sung, D.-Y.; Vierstra, R.D. The small ubiquitin-like modifier (SUMO) protein modification system in Arabidopsis. Accumulation of SUMO1 and -2 conjugates is increased by stress. J. Biol. Chem. 2003, 278, 6862–6872. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.J.; Scalf, M.; Rytz, T.C.; Hubler, S.L.; Smith, L.M.; Vierstra, R.D. Quantitative proteomics reveals factors regulating RNA biology as dynamic targets of stress-induced SUMOylation in Arabidopsis. Mol. Cell. Proteomics 2013, 12, 449–463. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, M.; Chiba, T.; Tatsumi, K.; Iemura, S.; Tanida, I.; Okazaki, N.; Ueno, T.; Kominami, E.; Natsume, T.; Tanaka, K. A novel protein-conjugating system for Ufm1, a ubiquitin-fold modifier. EMBO J. 2004, 23, 1977–1986. [Google Scholar] [CrossRef] [PubMed]
- Tatsumi, K.; Sou, Y.S.; Tada, N.; Nakamura, E.; Iemura, S.I.; Natsume, T.; Kang, S.H.; Chung, C.H.; Kasahara, M.; Kominami, E.; et al. A novel type of E3 ligase for the Ufm1 conjugation system. J. Biol. Chem. 2010, 285, 5417–5427. [Google Scholar] [CrossRef] [PubMed]
- Downes, B.P.; Saracco, S.A.; Lee, S.S.; Crowell, D.N.; Vierstra, R.D. MUBs, a family of ubiquitin-fold proteins that are plasma membrane-anchored by prenylation. J. Biol. Chem. 2006, 281, 27145–27157. [Google Scholar] [CrossRef] [PubMed]
- Ohsumi, Y. Molecular dissection of autophagy: two ubiquitin-like systems. Nat. Rev. Mol. Cell Biol. 2001, 2, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Vierstra, R.D. Autophagy: A multifaceted intracellular system for bulk and selective recycling. Trends Plant Sci. 2012, 17, 526–537. [Google Scholar] [CrossRef] [PubMed]
- Marshall, R.S.; Li, F.; Gemperline, D.C.; Book, A.J.; Vierstra, R.D. Autophagic degradation of the 26S proteasome is mediated by the dual ATG8/ubiquitin receptor RPN10 in Arabidopsis. Mol. Cell 2015, 58, 1053–1066. [Google Scholar] [CrossRef] [PubMed]
- Lorenzo, O.; Chico, J.M.; Sánchez-Serrano, J.J.; Solano, R. JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 2004, 16, 1938–1950. [Google Scholar] [CrossRef] [PubMed]
- Chini, A.; Fonseca, S.; Fernández, G.; Adie, B.; Chico, J.M.; Lorenzo, O.; García-Casado, G.; López-Vidriero, I.; Lozano, F.M.; Ponce, M.R.; et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 2007, 448, 666–671. [Google Scholar] [CrossRef] [PubMed]
- Thines, B.; Katsir, L.; Melotto, M.; Niu, Y.; Mandaokar, A.; Liu, G.; Nomura, K.; He, S.Y.; Howe, G.A.; Browse, J. JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signalling. Nature 2007, 448, 661–665. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Zhang, C.; Gu, M.; Bai, Z.; Zhang, W.; Qi, T.; Cheng, Z.; Peng, W.; Luo, H.; Nan, F.; et al. The Arabidopsis CORONATINE INSENSITIVE1 protein is a jasmonate receptor. Plant Cell 2009, 21, 2220–2236. [Google Scholar] [CrossRef] [PubMed]
- Pauwels, L.; Barbero, G.F.; Geerinck, J.; Tilleman, S.; Grunewald, W.; Cuéllar Pérez, A.; Chico, J.M.; vanden Bossche, R.; Sewell, J.; Gil, E.; et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 2010, 464, 788–791. [Google Scholar] [CrossRef] [PubMed]
- Chico, J.M.; Chini, A.; Fonseca, S.; Solano, R. JAZ repressors set the rhythm in jasmonate signaling. Curr. Opin. Plant Biol. 2008, 11, 486–494. [Google Scholar] [CrossRef] [PubMed]
- Pauwels, L.; Goossens, A. The JAZ proteins: a crucial interface in the jasmonate signaling cascade. Plant Cell 2011, 23, 3089–3100. [Google Scholar] [CrossRef] [PubMed]
- Feys, B.J.F.; Benedetti, C.E.; Penfold, C.N.; Turner, J.G. Arabidopsis mutants selected for resistance to the phytotoxin coronatine are male sterile, insensitive to methyl jasmonate, and resistant to a bacterial pathogen. Plant Cell 1994, 6, 751–759. [Google Scholar] [CrossRef] [PubMed]
- Vijayan, P.; Shockey, J.; Lévesque, C.A.; Cook, R.J.; Browse, J. A role for jasmonate in pathogen defense of Arabidopsis. Proc. Natl. Acad. Sci. USA 1998, 95, 7209–7214. [Google Scholar] [CrossRef] [PubMed]
- Howe, G.A.; Jander, G. Plant immunity to insect herbivores. Annu. Rev. Plant Biol. 2008, 59, 41–66. [Google Scholar] [CrossRef] [PubMed]
- Xie, D.-X.; Feys, B.F.; James, S.; Nieto-Rostro, M.; Turner, J.G. COI1: an Arabidopsis gene required for jasmonate-regulated defense and fertility. Science 1998, 280, 1091–1094. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Liu, F.; Lechner, E.; Genschik, P.; Crosby, W.L.; Ma, H.; Peng, W.; Huang, D.; Xie, D. The SCFCOl1 ubiquitin-ligase complexes are required for jasmonate response in Arabidopsis. Plant Cell 2002, 14, 1919–1935. [Google Scholar] [CrossRef] [PubMed]
- Dai, L.; Xu, L.; Huang, D.; Li, X.; Luo, K.; Guan, C. ASK1 physically interacts with COI1 and is required for male fertility in Arabidopsis. Sci. China C Life Sci. 2002, 45, 631–636. [Google Scholar] [CrossRef] [PubMed]
- Pauwels, L.; Ritter, A.; Goossens, J.; Nagels Durand, A.; Liu, H.; Gu, Y.; Geerinck, J.; Boter, M.; vanden Bossche, R.; de Clercq, R.; et al. The RING E3 ligase KEEP ON GOING modulates JASMONATE ZIM-DOMAIN12 stability. Plant Physiol. 2015, 169, 1405–1417. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Ma, L.; Wang, X.; Xie, D.; Dinesh-Kumar, S.P.; Wei, N.; Deng, X.W. The COP9 signalosome interacts physically with SCFCOI1 and modulates jasmonate responses. Plant Cell 2003, 15, 1083–1094. [Google Scholar] [CrossRef] [PubMed]
- Tiryaki, I.; Staswick, P.E. An Arabidopsis mutant defective in jasmonate response is allelic to the auxin-signaling mutant axr1. Plant Physiol. 2002, 130, 887–894. [Google Scholar] [CrossRef] [PubMed]
- Sheard, L.B.; Tan, X.; Mao, H.; Withers, J.; Ben-Nissan, G.; Hinds, T.R.; Kobayashi, Y.; Hsu, F.-F.; Sharon, M.; Browse, J.; et al. Jasmonate perception by inositol-phosphate-potentiated COI1–JAZ co-receptor. Nature 2010, 468, 400–405. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.S.; Howe, G.A. A critical role for the TIFY motif in repression of jasmonate signaling by a stabilized splice variant of the JASMONATE ZIM-domain protein JAZ10 in Arabidopsis. Plant Cell 2009, 21, 131–145. [Google Scholar] [CrossRef] [PubMed]
- Shyu, C.; Figueroa, P.; DePew, C.L.; Cooke, T.F.; Sheard, L.B.; Moreno, J.E.; Katsir, L.; Zheng, N.; Browse, J.; Howe, G.A. JAZ8 lacks a canonical degron and has an EAR motif that mediates transcriptional repression of jasmonate responses in Arabidopsis. Plant Cell 2012, 24, 536–550. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Li, H.; Li, S.; Yao, R.; Deng, H.; Xie, Q.; Xie, D. The Arabidopsis F-box protein CORONATINE INSENSITIVE1 is stabilized by SCFCOI1 and degraded via the 26S proteasome pathway. Plant Cell 2013, 25, 486–498. [Google Scholar] [CrossRef] [PubMed]
- Kazan, K.; Manners, J.M. Jasmonate signaling: toward an integrated view. Plant Physiol. 2008, 146, 1459–1468. [Google Scholar] [CrossRef] [PubMed]
- Thireault, C.; Shyu, C.; Yoshida, Y.; St.Aubin, B.; Campos, M.L.; Howe, G.A. Repression of jasmonate signaling by a non-TIFY JAZ protein in Arabidopsis. Plant J. 2015, 82, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Ye, H.; Du, H.; Tang, N.; Li, X.; Xiong, L. Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Mol. Biol. 2009, 71, 291–305. [Google Scholar] [CrossRef] [PubMed]
- Melotto, M.; Mecey, C.; Niu, Y.; Chung, H.S.; Katsir, L.; Yao, J.; Zeng, W.; Thines, B.; Staswick, P.; Browse, J.; et al. A critical role of two positively charged amino acids in the Jas motif of Arabidopsis JAZ proteins in mediating coronatine- and jasmonoyl isoleucine-dependent interactions with the COI1 F-box protein. Plant J. 2008, 55, 979–988. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Calvo, P.; Chini, A.; Fernández-Barbero, G.; Chico, J.-M.; Gimenez-Ibanez, S.; Geerinck, J.; Eeckhout, D.; Schweizer, F.; Godoy, M.; Franco-Zorrilla, J.M.; et al. The Arabidopsis bHLH transcription factors MYC3 and MYC4 are targets of JAZ repressors and act additively with MYC2 in the activation of jasmonate responses. Plant Cell 2011, 23, 701–715. [Google Scholar] [CrossRef] [PubMed]
- Vanholme, B.; Grunewald, W.; Bateman, A.; Kohchi, T.; Gheysen, G. The tify family previously known as ZIM. Trends Plant Sci. 2007, 12, 239–244. [Google Scholar] [CrossRef] [PubMed]
- Chini, A.; Fonseca, S.; Chico, J.M.; Fernández-Calvo, P.; Solano, R. The ZIM domain mediates homo- and heteromeric interactions between Arabidopsis JAZ proteins. Plant J. 2009, 59, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Saracco, S.A.; Hansson, M.; Scalf, M.; Walker, J.M.; Smith, L.M.; Vierstra, R.D. Tandem affinity purification and mass spectrometric analysis of ubiquitylated proteins in Arabidopsis. Plant J. 2009, 59, 344–358. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.; Bell, E.; Mullet, J.E. Two methyl jasmonate-insensitive mutants show altered expression of AtVsp in response to methyl jasmonate and wounding. Plant Physiol. 1996, 111, 525–531. [Google Scholar] [PubMed]
- Kazan, K.; Manners, J.M. MYC2: the master in action. Mol. Plant 2013, 6, 686–703. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Chen, J.; Xie, Z.; Gao, J.; Ren, G.; Gao, S.; Zhou, X.; Kuai, B. Jasmonic acid promotes degreening via MYC2/3/4- and ANAC019/055/072-mediated regulation of major chlorophyll catabolic genes. Plant J. 2015, 84, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Toledo-Ortiz, G.; Huq, E.; Quail, P.H. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 2003, 15, 1749–1770. [Google Scholar] [CrossRef] [PubMed]
- Dombrecht, B.; Xue, G.P.; Sprague, S.J.; Kirkegaard, J.A.; Ross, J.J.; Reid, J.B.; Fitt, G.P.; Sewelam, N.; Schenk, P.M.; Manners, J.M.; et al. MYC2 differentially modulates diverse jasmonate-dependent functions in Arabidopsis. Plant Cell 2007, 19, 2225–2245. [Google Scholar] [CrossRef] [PubMed]
- Godoy, M.; Franco-Zorrilla, J.M.; Pérez-Pérez, J.; Oliveros, J.C.; Lorenzo, Ó.; Solano, R. Improved protein-binding microarrays for the identification of DNA-binding specificities of transcription factors. Plant J. 2011, 66, 700–711. [Google Scholar] [CrossRef] [PubMed]
- Çevik, V.; Kidd, B.N.; Zhang, P.; Hill, C.; Kiddle, S.; Denby, K.J.; Holub, E.B.; Cahill, D.M.; Manners, J.M.; Schenk, P.M.; et al. MEDIATOR25 acts as an integrative hub for the regulation of jasmonate-responsive gene expression in Arabidopsis. Plant Physiol. 2012, 160, 541–555. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Jiang, H.; Li, L.; Zhai, Q.; Qi, L.; Zhou, W.; Liu, X.; Li, H.; Zheng, W.; Sun, J.; et al. The Arabidopsis mediator subunit MED25 differentially regulates jasmonate and abscisic acid signaling through interacting with the MYC2 and ABI5 transcription factors. Plant Cell 2012, 24, 2898–2916. [Google Scholar] [CrossRef] [PubMed]
- Goossens, J.; Swinnen, G.; vanden Bossche, R.; Pauwels, L.; Goossens, A. Change of a conserved amino acid in the MYC2 and MYC3 transcription factors leads to release of JAZ repression and increased activity. New Phytol. 2015, 206, 1229–1237. [Google Scholar] [CrossRef] [PubMed]
- Gasperini, D.; Chételat, A.; Acosta, I.F.; Goossens, J.; Pauwels, L.; Goossens, A.; Dreos, R.; Alfonso, E.; Farmer, E.E. Multilayered organization of jasmonate signalling in the regulation of root growth. PLoS Genet. 2015, 11, e1005300. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Yao, J.; Ke, J.; Zhang, L.; Lam, V.Q.; Xin, X.-F.; Zhou, X.E.; Chen, J.; Brunzelle, J.; Griffin, P.R.; et al. Structural basis of JAZ repression of MYC transcription factors in jasmonate signalling. Nature 2015, 525, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Kazan, K.; Manners, J.M. JAZ repressors and the orchestration of phytohormone crosstalk. Trends Plant Sci. 2012, 17, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Ishiguro, S.; Kawai-Oda, A.; Ueda, J.; Nishida, I.; Okada, K. The DEFECTIVE IN ANTHER DEHISCENCE1 gene encodes a novel phospholipase A1 catalyzing the initial step of jasmonic acid biosynthesis, which synchronizes pollen maturation, anther dehiscence, and flower opening in Arabidopsis. Plant Cell 2001, 13, 2191–2209. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-T.; Liu, H.; Stone, S.; Callis, J. ABA and the ubiquitin E3 ligase KEEP ON GOING affect proteolysis of the Arabidopsis thaliana transcription factors ABF1 and ABF3. Plant J. 2013, 75, 965–976. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Stone, S.L. Cytoplasmic degradation of the Arabidopsis transcription factor ABSCISIC ACID INSENSITIVE 5 is mediated by the RING-type E3 ligase KEEP ON GOING. J. Biol. Chem. 2013, 288, 20267–20279. [Google Scholar] [CrossRef] [PubMed]
- Stone, S.L.; Williams, L.A.; Farmer, L.M.; Vierstra, R.D.; Callis, J. KEEP ON GOING, a RING E3 ligase essential for Arabidopsis growth and development, is involved in abscisic acid signaling. Plant Cell 2006, 18, 3415–3428. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wu, Q.; Cui, S.; Ren, J.; Qian, W.; Yang, Y.; He, S.; Chu, J.; Sun, X.; Yan, C.; et al. Hijacking of the jasmonate pathway by the mycotoxin fumonisin B1 (FB1) to initiate programmed cell death in Arabidopsis is modulated by RGLG3 and RGLG4. J. Exp. Bot. 2015, 66, 2709–2721. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wu, Q.; Ren, J.; Qian, W.; He, S.; Huang, K.; Yu, X.; Gao, Y.; Huang, P.; An, C. Two novel RING-type ubiquitin ligases, RGLG3 and RGLG4, are essential for jasmonate-mediated responses in Arabidopsis. Plant Physiol. 2012, 160, 808–822. [Google Scholar] [CrossRef] [PubMed]
- Jung, C.; Zhao, P.; Seo, J.S.; Mitsuda, N.; Deng, S.; Chua, N.H. PLANT U-BOX PROTEIN10 regulates MYC2 stability in Arabidopsis. Plant Cell 2015, 27, 2016–2031. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Laluk, K.; Lai, Z.; Veronese, P.; Song, F.; Mengiste, T. The Arabidopsis Botrytis Susceptible1 Interactor defines a subclass of RING E3 ligases that regulate pathogen and stress responses. Plant Physiol. 2010, 154, 1766–1782. [Google Scholar] [CrossRef] [PubMed]
- Mandaokar, A.; Browse, J. MYB108 acts together with MYB24 to regulate jasmonate-mediated stamen maturation in Arabidopsis. Plant Physiol. 2009, 149, 851–862. [Google Scholar] [CrossRef] [PubMed]
- Mengiste, T.; Chen, X.; Salmeron, J.; Dietrich, R. The BOTRYTIS SUSCEPTIBLE1 gene encodes an R2R3MYB transcription factor protein that is required for biotic and abiotic stress responses in Arabidopsis. Plant Cell 2003, 15, 2551–2565. [Google Scholar] [CrossRef] [PubMed]
- Pollier, J.; Moses, T.; González-Guzmán, M.; de Geyter, N.; Lippens, S.; vanden Bossche, R.; Marhavý, P.; Kremer, A.; Morreel, K.; Guérin, C.J.; et al. The protein quality control system manages plant defence compound synthesis. Nature 2013, 504, 148–152. [Google Scholar] [CrossRef] [PubMed]
- Lourenco, T.F.; Serra, T.S.; Cordeiro, A.M.; Swanson, S.J.; Gilroy, S.; Saibo, N.J.M.; Oliveira, M.M. The rice E3-ubiquitin ligase HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENE1 modulates the expression of ROOT MEANDER CURLING, a gene involved in root mechanosensing, through the interaction with two ETHYLENE-RESPONSE FACTOR transcription factors. Plant Physiol. 2015, 169, 2275–2287. [Google Scholar] [PubMed]
- Dong, C.H.; Agarwal, M.; Zhang, Y.; Xie, Q.; Zhu, J.-K. The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degradation of ICE1. Proc. Natl. Acad. Sci. USA 2006, 103, 8281–8286. [Google Scholar] [CrossRef] [PubMed]
- Lourenço, T.; Sapeta, H.; Figueiredo, D.D.; Rodrigues, M.; Cordeiro, A.; Abreu, I.A.; Saibo, N.J.M.; Oliveira, M.M. Isolation and characterization of rice (Oryza sativa L.) E3-ubiquitin ligase OsHOS1 gene in the modulation of cold stress response. Plant Mol. Biol. 2013, 83, 351–363. [Google Scholar] [CrossRef] [PubMed]
- Hu, P.; Zhou, W.; Cheng, Z.; Fan, M.; Wang, L.; Xie, D. JAV1 controls jasmonate-regulated plant defense. Mol. Cell 2013, 50, 504–515. [Google Scholar] [CrossRef] [PubMed]
- Van der Does, D.; Leon-Reyes, A.; Koornneef, A.; Van Verk, M.C.; Rodenburg, N.; Pauwels, L.; Goossens, A.; Körbes, A.P.; Memelink, J.; Ritsema, T.; et al. Salicylic acid suppresses jasmonic acid signaling downstream of SCFCOI1-JAZ by targeting GCC promoter motifs via transcription factor ORA59. Plant Cell 2013, 25, 744–761. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.; Heidrich, K.; Sanchez-Villarreal, A.; Parker, J.E.; Davis, S.J. TIME FOR COFFEE represses accumulation of the MYC2 transcription factor to provide time-of-day regulation of jasmonate signaling in Arabidopsis. Plant Cell 2012, 24, 2470–2482. [Google Scholar] [CrossRef] [PubMed]
- Cuéllar Pérez, A.; Nagels Durand, A.; vanden Bossche, R.; de Clercq, R.; Persiau, G.; van Wees, S.C.M.; Pieterse, C.M.J.; Gevaert, K.; de Jaeger, G.; Goossens, A.; et al. The non-JAZ TIFY protein TIFY8 from Arabidopsis thaliana is a transcriptional repressor. PLoS ONE 2014, 9, e84891. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Lu, F.; Li, Y.; Xue, Y.; Kang, Y.; Zhang, S.; Qiu, Q.; Cui, X.; Zheng, S.; Liu, B.; et al. Ubiquitin-specific proteases UBP12 and UBP13 act in circadian clock and photoperiodic flowering regulation in Arabidopsis. Plant Physiol. 2013, 162, 897–906. [Google Scholar] [CrossRef] [PubMed]
- Ewan, R.; Pangestuti, R.; Thornber, S.; Craig, A.; Carr, C.; O'Donnell, L.; Zhang, C.; Sadanandom, A. Deubiquitinating enzymes AtUBP12 and AtUBP13 and their tobacco homologue NtUBP12 are negative regulators of plant immunity. New Phytol. 2011, 191, 92–106. [Google Scholar] [CrossRef] [PubMed]
- Hakenjos, J.P.; Richter, R.; Dohmann, E.M.N.; Katsiarimpa, A.; Isono, E.; Schwechheimer, C. MLN4924 Is an Efficient Inhibitor of NEDD8 Conjugation in Plants. Plant Physiol. 2011, 156, 527–536. [Google Scholar] [CrossRef] [PubMed]
- Larsen, P.B.; Cancel, J.D. A recessive mutation in the RUB1-conjugating enzyme, RCE1, reveals a requirement for RUB modification for control of ethylene biosynthesis and proper induction of basic chitinase and PDF1.2 in Arabidopsis. Plant J. 2004, 38, 626–638. [Google Scholar] [CrossRef] [PubMed]
- Lozano-Durán, R.; Rosas-Díaz, T.; Gusmaroli, G.; Luna, A.P.; Taconnat, L.; Deng, X.W.; Bejarano, E.R. Geminiviruses subvert ubiquitination by altering CSN-mediated derubylation of SCF E3 ligase complexes and inhibit jasmonate signaling in Arabidopsis thaliana. Plant Cell 2011, 23, 1014–1032. [Google Scholar] [CrossRef] [PubMed]
- Catala, R.; Ouyang, J.; Abreu, I.A.; Hu, Y.; Seo, H.; Zhang, X.; Chua, N.-H. The Arabidopsis E3 SUMO ligase SIZ1 regulates plant growth and drought responses. Plant Cell 2007, 19, 2952–2966. [Google Scholar] [CrossRef] [PubMed]
- Serrano, M.; Parra, S.; Alcaraz, L.D.; Guzmán, P. The ATL gene family from Arabidopsis thaliana and Oryza sativa comprises a large number of putative ubiquitin ligases of the RING-H2 type. J. Mol. Evol. 2006, 62, 434–445. [Google Scholar] [CrossRef] [PubMed]
- Andersen, P.; Kragelund, B.B.; Olsen, A.N.; Larsen, F.H.; Chua, N.-H.; Poulsen, F.M.; Skriver, K. Structure and biochemical function of a prototypical Arabidopsis U-box domain. J. Biol. Chem. 2004, 279, 40053–40061. [Google Scholar] [CrossRef] [PubMed]
- Son, O.; Cho, S.K.; Kim, E.Y.; Kim, W.T. Characterization of three Arabidopsis homologs of human RING membrane anchor E3 ubiquitin ligase. Plant Cell Rep. 2009, 28, 561–569. [Google Scholar] [CrossRef] [PubMed]
- Son, O.; Cho, S.K.; Kim, S.J.; Kim, W.T. In vitro and in vivo interaction of AtRma2 E3 ubiquitin ligase and auxin binding protein 1. Biochem. Biophys. Res. Commun. 2010, 393, 492–497. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, Y.; Zhang, D.; Dai, X.; Estelle, M.; Zhao, Y. Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development. Proc. Natl. Acad. Sci. USA 2015, 112, 2275–2280. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Jiang, L.; Wang, F.; Yu, D. Jasmonate regulates the INDUCER OF CBF EXPRESSION–C-REPEAT BINDING FACTOR/DRE BINDING FACTOR1 cascade and freezing tolerance in Arabidopsis. Plant Cell 2013, 25, 2907–2924. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Li, J.; Xu, Y.; Han, Y.; Bai, Y.; Zhou, G.; Lou, Y.; Xu, Z.; Chong, K. RNAi knockdown of Oryza sativa root meander curling gene led to altered root development and coiling which were mediated by jasmonic acid signalling in rice. Plant Cell Environ. 2007, 30, 690–699. [Google Scholar] [CrossRef] [PubMed]
- Leone, M.; Keller, M.M.; Cerrudo, I.; Ballaré, C.L. To grow or defend? Low red : far-red ratios reduce jasmonate sensitivity in Arabidopsis seedlings by promoting DELLA degradation and increasing JAZ10 stability. New Phytol. 2014, 204, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Pieterse, C.M.J.; Pierik, R.; van Wees, S.C.M. Different shades of JAZ during plant growth and defense. New Phytol. 2014, 204, 261–264. [Google Scholar] [CrossRef] [PubMed]
- Goodspeed, D.; Chehab, E.W.; Min-Venditti, A.; Braam, J.; Covington, M.F. Arabidopsis synchronizes jasmonate-mediated defense with insect circadian behavior. Proc. Natl. Acad. Sci. USA 2012, 109, 4674–4677. [Google Scholar] [CrossRef] [PubMed]
- Ingle, R.A.; Stoker, C.; Stone, W.; Adams, N.; Smith, R.; Grant, M.; Carré, I.; Roden, L.C.; Denby, K.J. Jasmonate signalling drives time-of-day differences in susceptibility of Arabidopsis to the fungal pathogen Botrytis cinerea. Plant J. 2015, 84, 937–948. [Google Scholar] [CrossRef] [PubMed]
- Stuttmann, J.; Lechner, E.; Guérois, R.; Parker, J.E.; Nussaume, L.; Genschik, P.; Noël, L.D. COP9 signalosome-and 26S proteasome-dependent regulation of SCFTIR1 accumulation in Arabidopsis. J. Biol. Chem. 2009, 284, 7920–7930. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Zhang, Y.; Moss, B.L.; Bargmann, B.O.R.; Wang, R.; Prigge, M.; Nemhauser, J.L.; Estelle, M. Untethering the TIR1 auxin receptor from the SCF complex increases its stability and inhibits auxin response. Nat. Plants 2015, 1, 14030. [Google Scholar] [CrossRef] [PubMed]
- Ramadan, A.; Nemoto, K.; Seki, M.; Shinozaki, K.; Takeda, H.; Takahashi, H.; Sawasaki, T. Wheat germ-based protein libraries for the functional characterisation of the Arabidopsis E2 ubiquitin conjugating enzymes and the RING-type E3 ubiquitin ligase enzymes. BMC Plant Biol. 2015, 15, 275. [Google Scholar] [CrossRef] [PubMed]
- Keren-Kaplan, T.; Attali, I.; Motamedchaboki, K.; Davis, B.A.; Tanner, N.; Reshef, Y.; Laudon, E.; Kolot, M.; Levin-Kravets, O.; Kleifeld, O.; et al. Synthetic biology approach to reconstituting the ubiquitylation cascade in bacteria. EMBO J. 2012, 31, 378–390. [Google Scholar] [CrossRef] [PubMed]
- Gilkerson, J.; Kelley, D.R.; Tam, R.; Estelle, M.; Callis, J. Lysine residues are not required for proteasome-mediated proteolysis of the auxin/indole acidic acid protein IAA1. Plant Physiol. 2015, 168, 708–720. [Google Scholar] [CrossRef] [PubMed]
- Stes, E.; Laga, M.; Walton, A.; Samyn, N.; Timmerman, E.; De Smet, I.; Goormachtig, S.; Gevaert, K. A COFRADIC protocol to study protein ubiquitination. J. Proteome Res. 2014, 13, 3107–3113. [Google Scholar] [CrossRef] [PubMed]
- Larrieu, A.; Champion, A.; Legrand, J.; Lavenus, J.; Mast, D.; Brunoud, G.; Oh, J.; Guyomarc'h, S.; Pizot, M.; Farmer, E.E.; et al. A fluorescent hormone biosensor reveals the dynamics of jasmonate signalling in plants. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nagels Durand, A.; Pauwels, L.; Goossens, A. The Ubiquitin System and Jasmonate Signaling. Plants 2016, 5, 6. https://doi.org/10.3390/plants5010006
Nagels Durand A, Pauwels L, Goossens A. The Ubiquitin System and Jasmonate Signaling. Plants. 2016; 5(1):6. https://doi.org/10.3390/plants5010006
Chicago/Turabian StyleNagels Durand, Astrid, Laurens Pauwels, and Alain Goossens. 2016. "The Ubiquitin System and Jasmonate Signaling" Plants 5, no. 1: 6. https://doi.org/10.3390/plants5010006
APA StyleNagels Durand, A., Pauwels, L., & Goossens, A. (2016). The Ubiquitin System and Jasmonate Signaling. Plants, 5(1), 6. https://doi.org/10.3390/plants5010006







