Systematic Characterization of GATA Transcription Factors in Liriodendron chinense and Functional Validation in Abiotic Stresses
Abstract
:1. Introduction
2. Materials and Methods
2.1. Identification and Physicochemical Properties Analysis of GATA Family Members in Liriodendron chinense
2.2. Phylogenetic Tree, Conservation Domain, Cis-Element, and Protein-to-Protein Analyses
2.3. Plant Material Treatment and Transcriptome Data
2.3.1. RNA Quantification and Qualification
2.3.2. Library Preparation for Transcriptome Sequencing
2.4. qRT-PCR Expression Analysis
3. Results
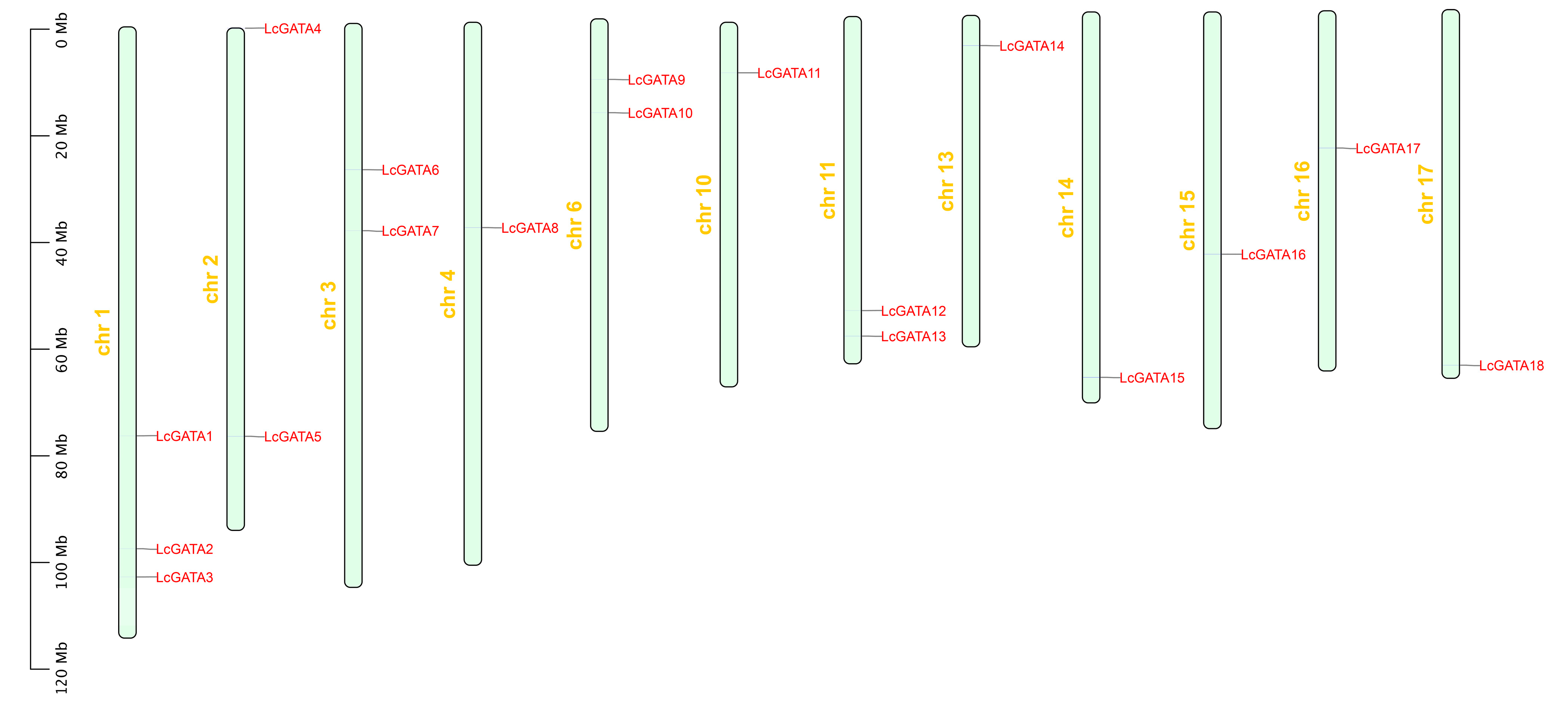
3.1. Characterization of GATA Factors in L. chinense and Their Physiological Properties
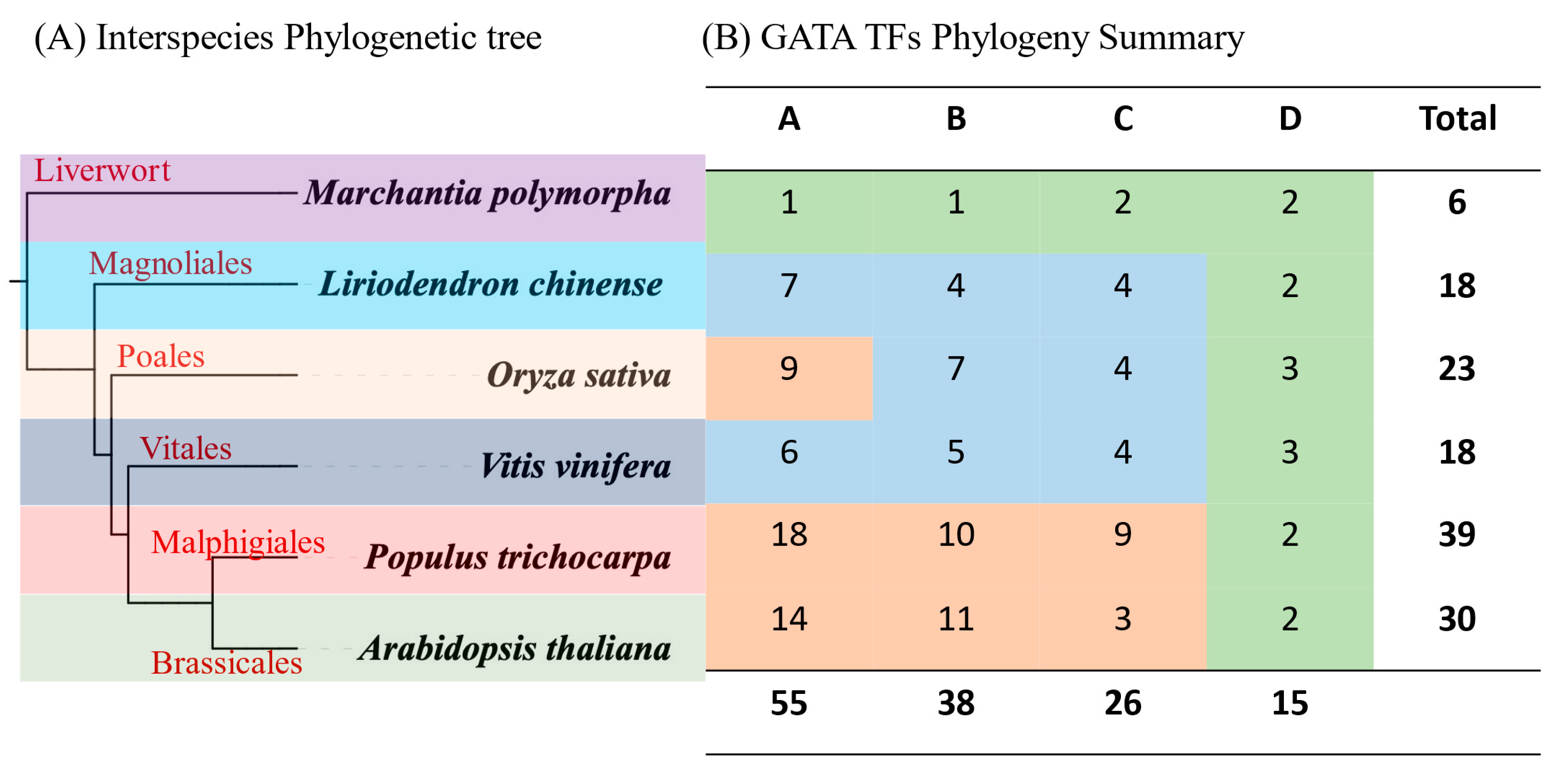
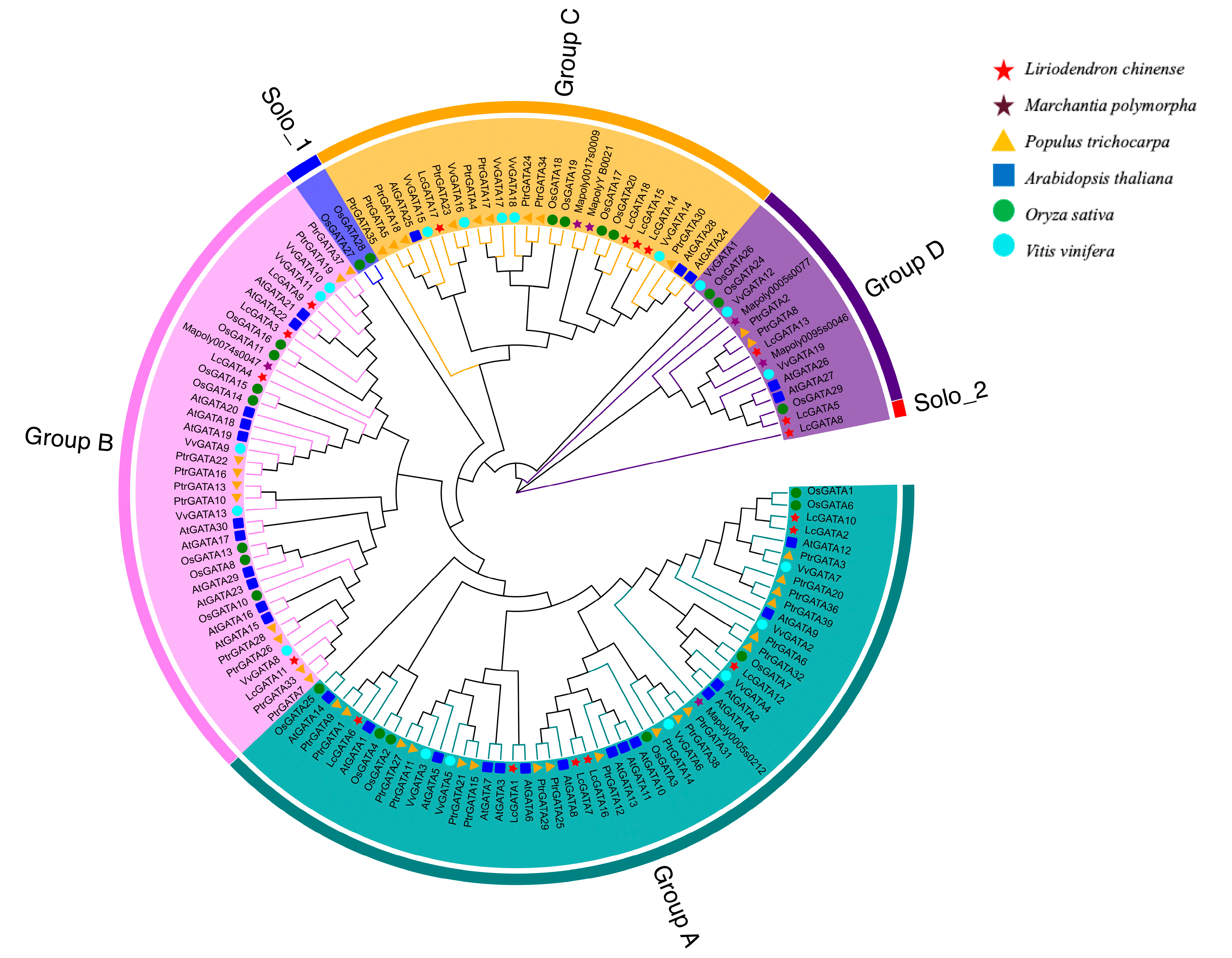
3.2. Phylogenetic Analysis of GATA Factors
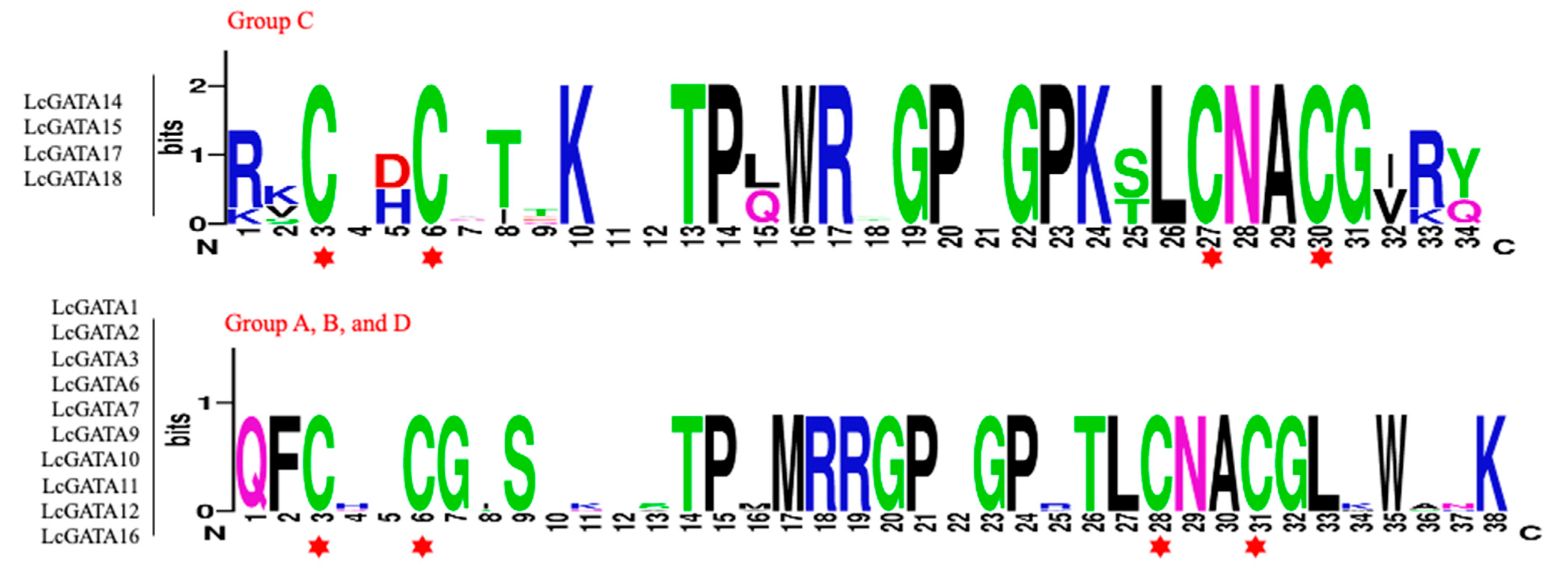
3.3. The Conserved Domain, Motif, and Gene Structure Analysis
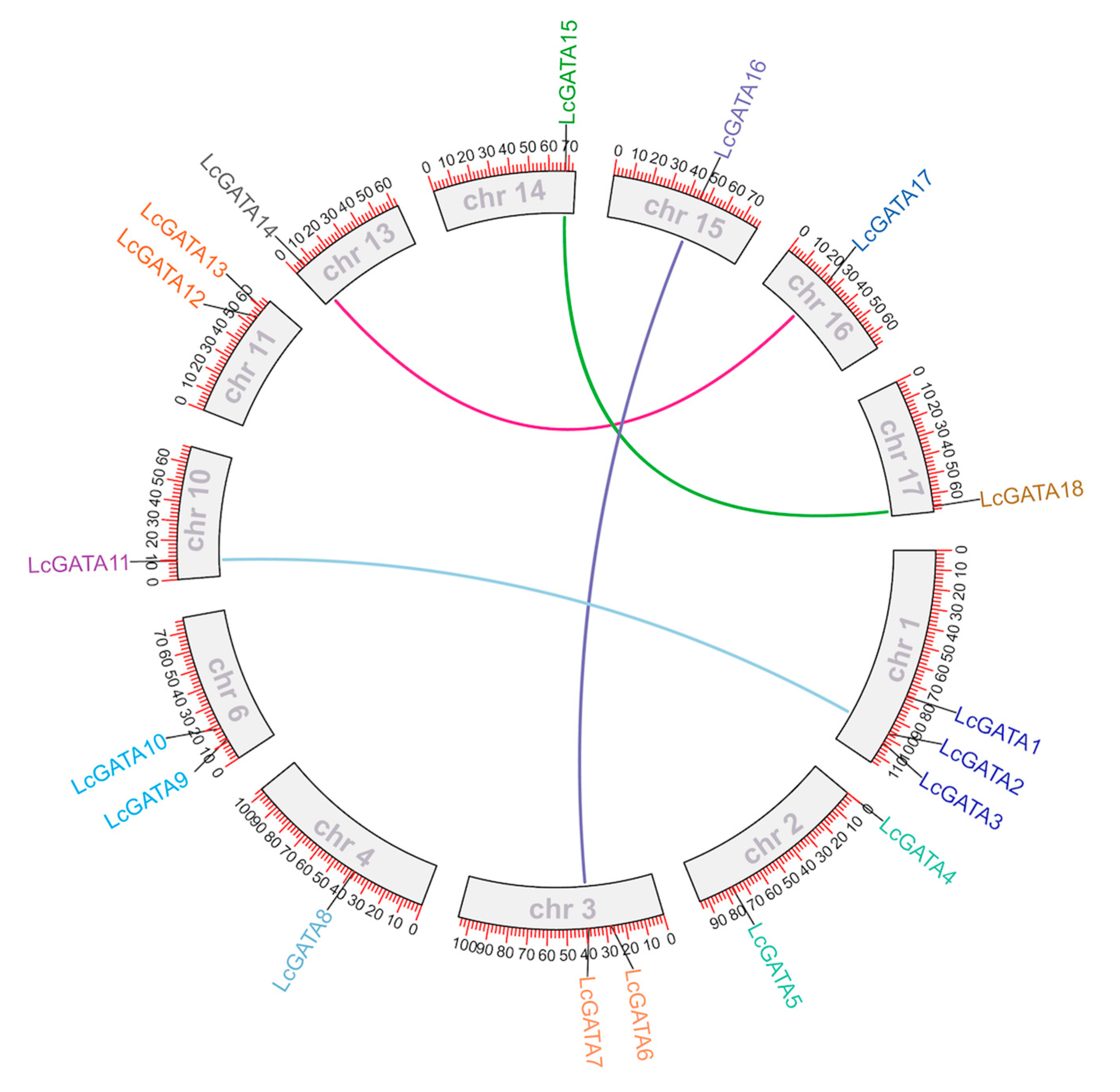
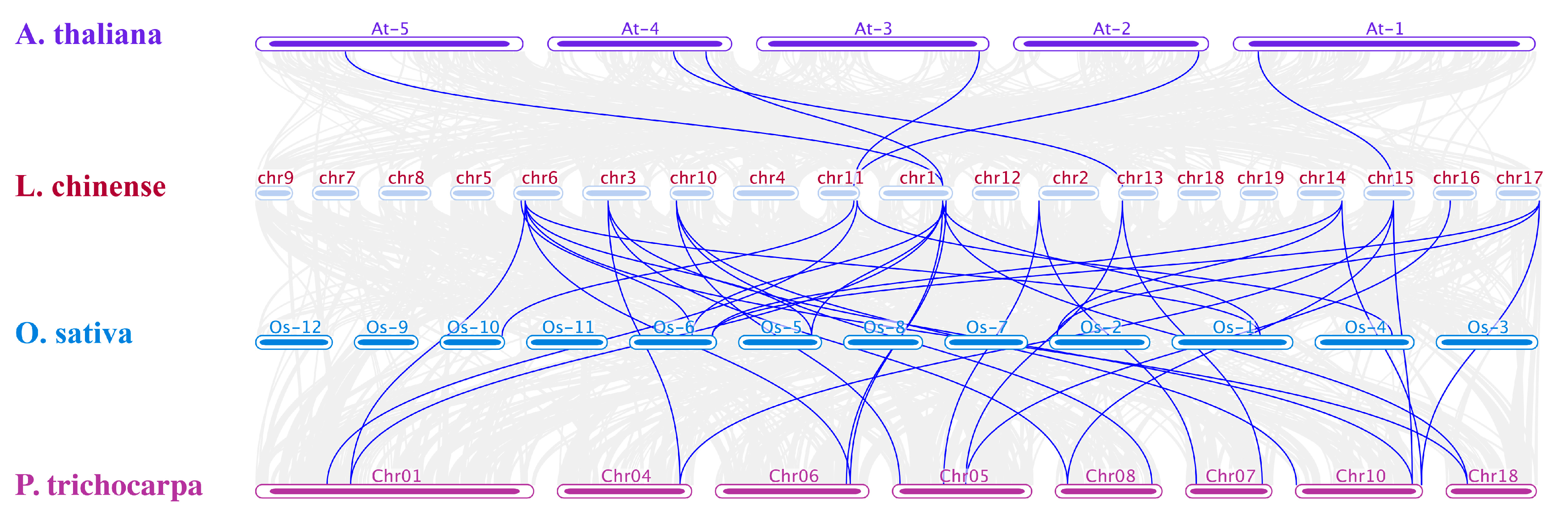
3.4. Collinearity, Selection Pressure, and Duplication Events
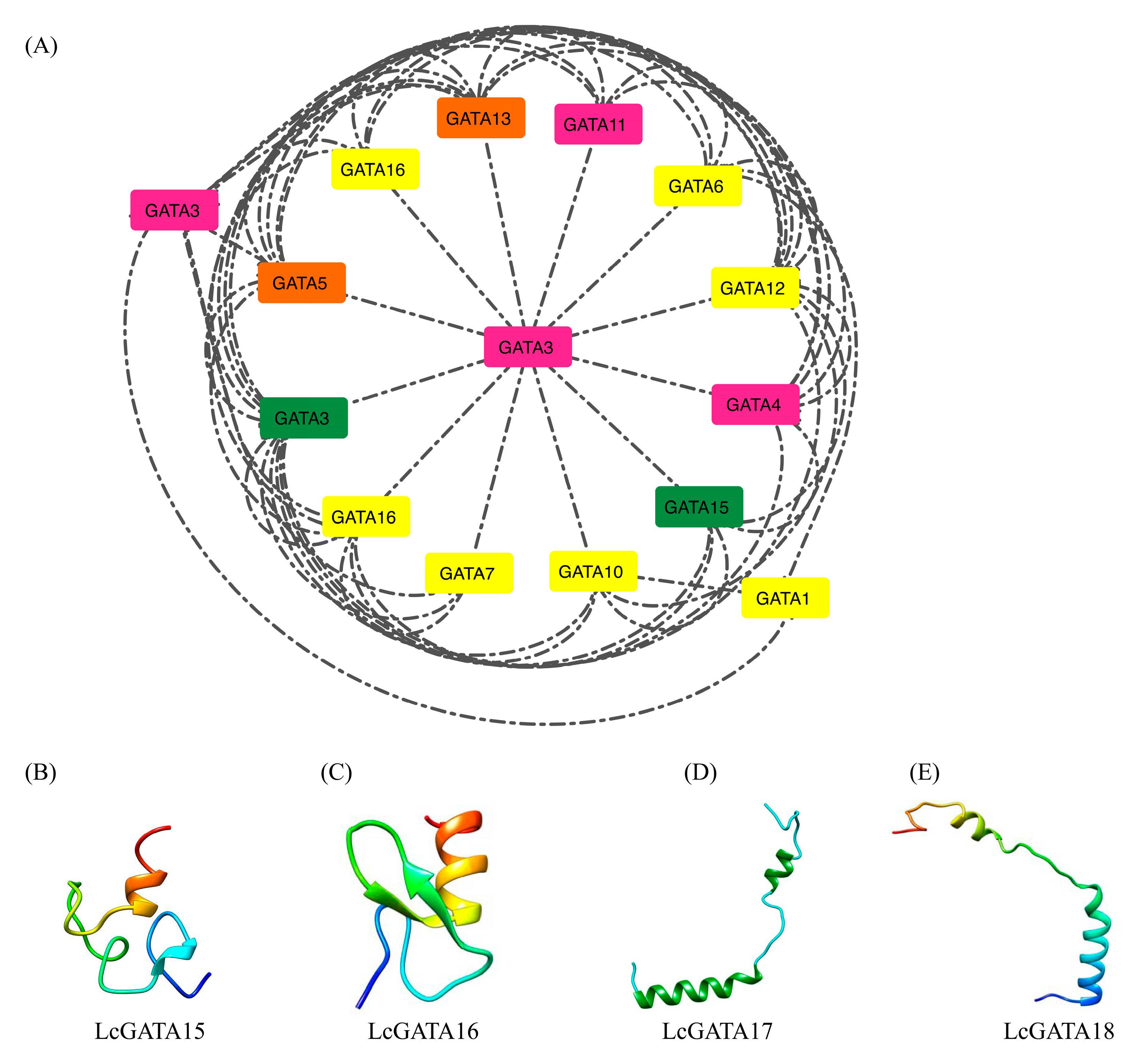
3.5. Protein-Protein Interaction and Secondary Structure of LcGATAs
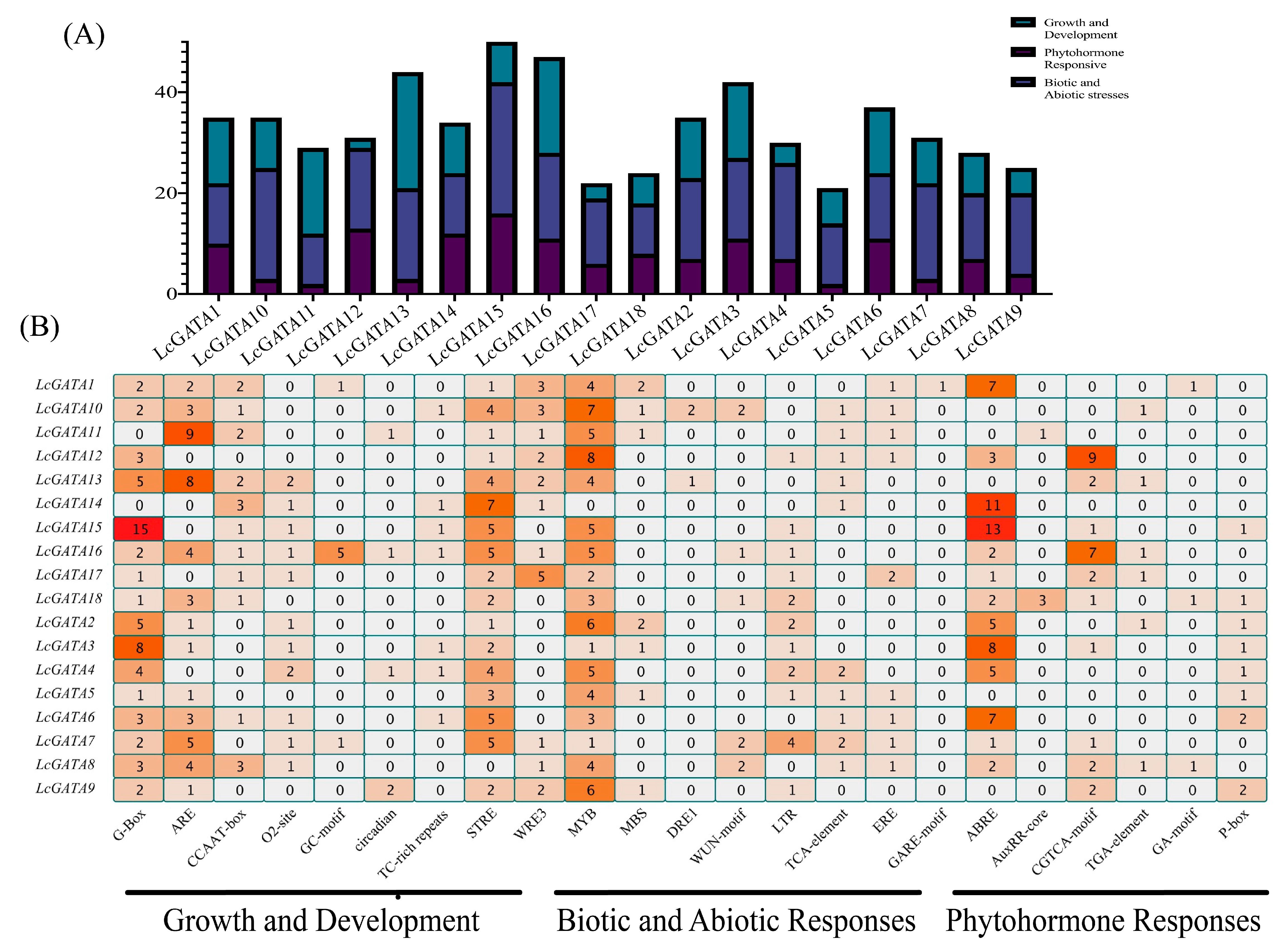
3.6. Analysis of Cis-Acting Elements in Promoters of LcGATAs
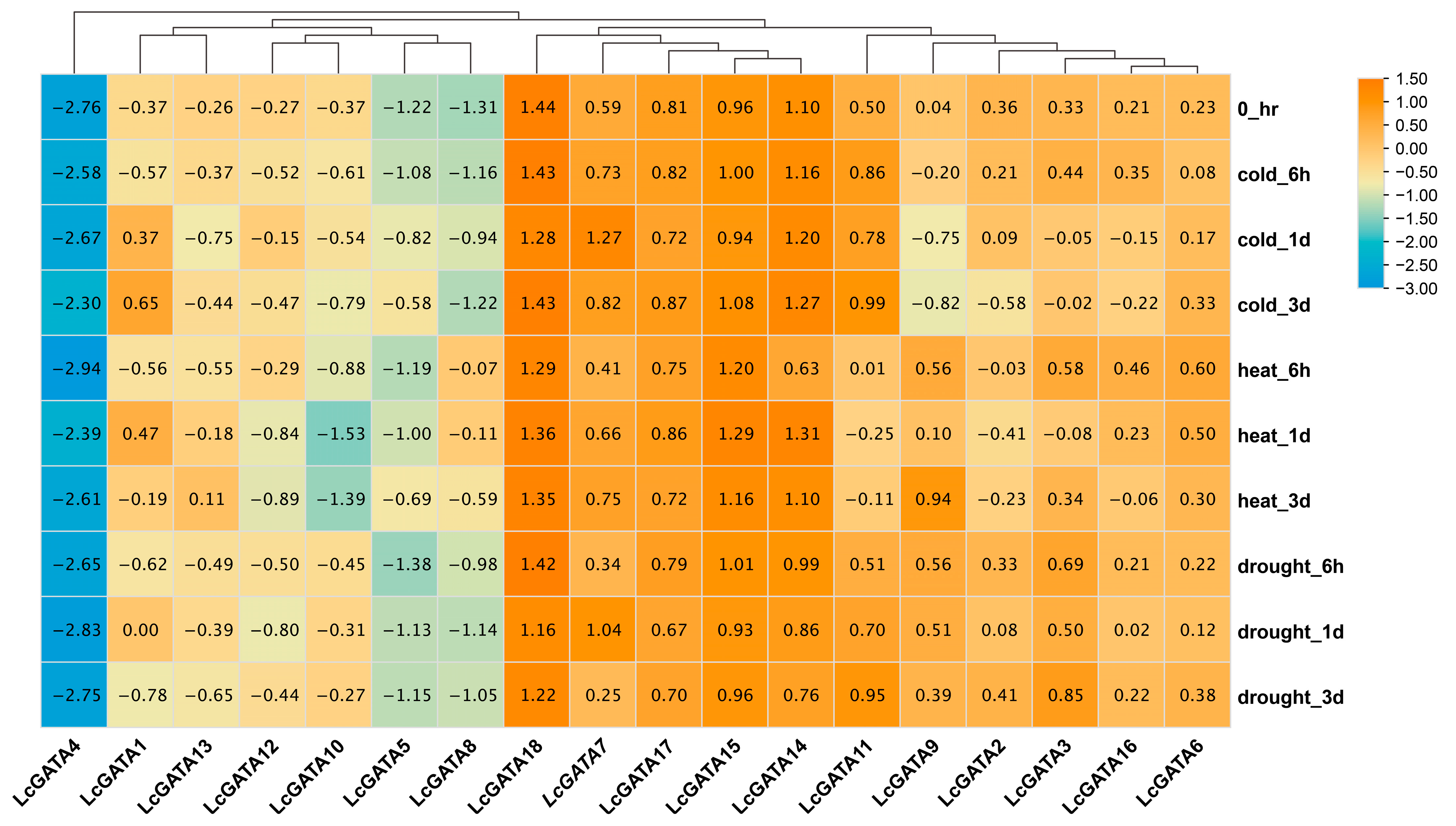
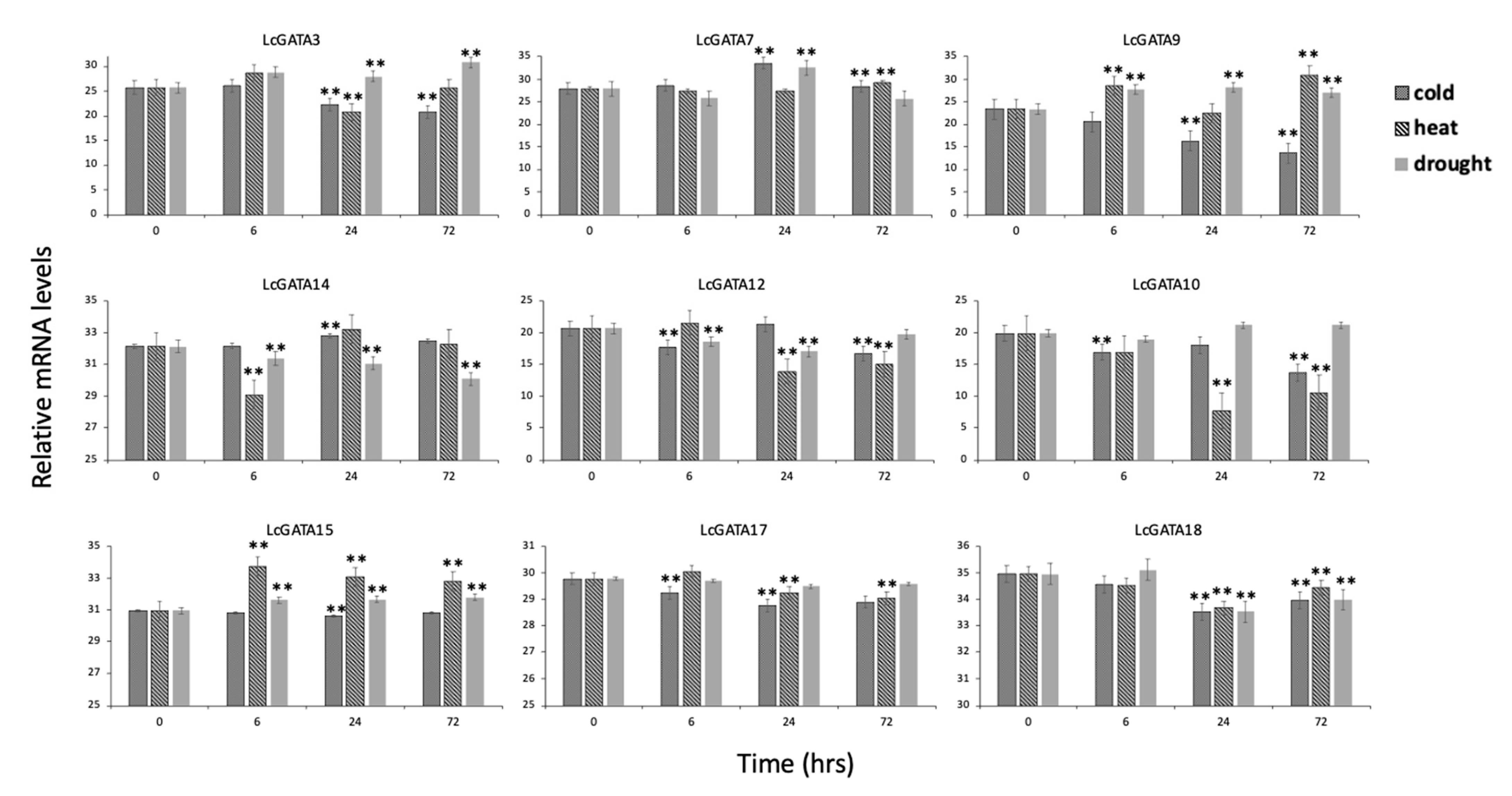
3.7. Transcriptomic and qRT-PCR Gene Expression Analysis of LcGATAs under Abiotic Stresses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cao, Y.; Feng, J.; Hwarari, D.; Ahmad, B.; Wu, H.; Chen, J.; Yang, L. Alterations in Population Distribution of Liriodendron chinense (Hemsl.) Sarg. and Liriodendron tulipifera Linn. Caused by Climate Change. Forests 2022, 13, 488. [Google Scholar] [CrossRef]
- Parks, C.R.; Wendel, J.F. Molecular divergence between Asian and North American species of Liriodendron (Magnoliaceae) with implications for interpretation of fossil floras. Am. J. Bot. 1990, 77, 1243–1256. [Google Scholar] [CrossRef]
- Chen, J.; Hao, Z.; Guang, X.; Zhao, C.; Wang, P.; Xue, L. Liriodendron genome sheds light on angiosperm phylogeny and species-pair differentiation. Nat. Plants 2019, 5, 18–25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, Y.; Zhou, Y.; Han, X.; Shen, C.; Wang, S.; Liu, C. The GATA transcription factor GNC plays an important role in photosynthesis and growth in poplar. J. Exp. Bot. 2020, 71, 1969–1984. [Google Scholar] [CrossRef]
- Romano, O.; Miccio, A. GATA factor transcriptional activity: Insights from genome-wide binding profiles. IUBMB Life 2020, 72, 10–26. [Google Scholar] [CrossRef]
- Lowry, J.A.; Atchley, W.R. Molecular evolution of the GATA family of transcription factors: Conservation within the DNA-binding domain. J. Mol. Evol. 2000, 50, 103–115. [Google Scholar] [CrossRef] [PubMed]
- Cantor, A.B.; Orkin, S.H. Coregulation of GATA factors by the Friend of GATA (FOG) family of multitype zinc finger proteins. Semin. Cell Dev. Biol. 2005, 16, 117–128. [Google Scholar] [CrossRef]
- Scazzocchio, C. The fungal GATA factors. Curr. Opin. Microbiol. 2000, 3, 126–131. [Google Scholar] [CrossRef]
- Reyes, J.C.; Muro-Pastor, M.I.; Florencio, F.J. The GATA family of transcription factors in Arabidopsis and rice. Plant Physiol. 2004, 134, 1718–1732. [Google Scholar] [CrossRef] [Green Version]
- Bi, Y.M.; Zhang, Y.; Signorelli, T.; Zhao, R.; Zhu, T.; Rothstein, S. Genetic analysis of Arabidopsis GATA transcription factor gene family reveals a nitrate-inducible member important for chlorophyll synthesis and glucose sensitivity. Plant J. 2005, 44, 680–692. [Google Scholar] [CrossRef]
- Pomraning, K.R.; Bredeweg, E.L.; Baker, S.E. Regulation of Nitrogen Metabolism by GATA Zinc Finger Transcription Factors in Yarrowia lipolytica. mSphere 2017, 2, e00038-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.; Wang, G.; Deng, W.; Li, T. Distribution, evolution, and expression of GATA-TFs provide new insights into their functions in light response and fruiting body development of Tolypocladium guangdongense. PeerJ 2020, 8, e9784. [Google Scholar] [CrossRef]
- Niehaus, E.M.; Schumacher, J.; Burkhardt, I.; Rabe, P.; Spitzer, E.; Münsterkötter, M. The GATA-Type Transcription Factor Csm1 Regulates Conidiation and Secondary Metabolism in Fusarium fujikuroi. Front. Microbiol. 2017, 8, 1175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Münchow, S.; Sauter, C.; Jansen, R.P. Association of the class V myosin Myo4p with a localized messenger RNA in budding yeast depends on She proteins. J. Cell Sci. 1999, 112, 1511–1518. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.-M.; Lin, W.-H.; Zhu, S.; Zhu, J.-Y.; Sun, Y.; Fan, X.-Y. Integration of Light- and Brassinosteroid-Signaling Pathways by a GATA Transcription Factor in Arabidopsis. Dev. Cell 2010, 19, 872–883. [Google Scholar] [CrossRef] [Green Version]
- Bastakis, E.; Hedtke, B.; Klermundm, C.; Grimm, B.; Schwechheimer, C. LLM-Domain B-GATA Transcription Factors Play Multifaceted Roles in Controlling Greening in Arabidopsis. Plant Cell 2018, 30, 582–599. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, H.; Wu, T.; Li, Z.; Huang, K.; Kim, N.-E.; Ma, Z. OsGATA16, a GATA transcription factor, confers cold tolerance by repressing OsWRKY45–1 at the seedling stage in rice. Rice 2021, 14, 42. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.; Nutan, K.K.; Singla-Pareek, S.L.; Pareek, A. Abiotic Stresses Cause Differential Regulation of Alternative Splice Forms of GATA Transcription Factor in Rice. Front. Plant Sci. 2017, 13, 1944. [Google Scholar] [CrossRef] [Green Version]
- Nutan, K.K.; Singla-Pareek, S.L.; Pareek, A. The Saltol QTL-localized transcription factor OsGATA8 plays an important role in stress tolerance and seed development in Arabidopsis and rice. J. Exp. Bot. 2020, 71, 684–698. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Cao, X.; Zhang, D.; Li, Y.; Wang, Q.; Ma, F.; Xu, X.; Zhan, X.; Hu, T. SlGATA17, A tomato GATA protein, interacts with SlHY5 to modulate salinity tolerance and germination. Environ. Exp. Bot. 2023, 206, 105191. [Google Scholar] [CrossRef]
- Xu, Z. The Role of Anthocyanins and the GATA Transcription Factors GNC and CGA1 in the Plant Response to Stress. Ph.D. Thesis, University of Guelph, Guelph, ON, Canada, 2016. [Google Scholar]
- Richter, R.; Bastakis, E.; Schwechheimer, C. Cross-Repressive Interactions between SOC1 and the GATAs GNC and GNL/CGA1 in the Control of Greening, Cold Tolerance, and Flowering Time in Arabidopsis. Plant Physiol. 2013, 162, 1992–2004. [Google Scholar] [CrossRef] [Green Version]
- Guo, J.; Bai, X.; Dai, K.; Yuan, X.; Guo, P.; Zhou, M.; Shi, W.; Hao, C. Identification of GATA transcription factors in Brachypodium distachyon and functional characterization of BdGATA13 in drought tolerance and response to gibberellins. Front. Plant Sci. 2021, 12, 763665. [Google Scholar] [CrossRef]
- Xu, X.; Zhang, H.; Xie, T.; Xu, Y.; Zhao, L.; Tian, W. Effects of climate change on the potentially suitable climatic geographical range of Liriodendron chinense. Forests 2017, 8, 399. [Google Scholar] [CrossRef] [Green Version]
- Falk, T.; Herndon, N.; Grau, E.; Buehler, S.; Richter, P.; Zaman, S.; Baker, E.M.; Ramnath, R.; Ficklin, S.; Staton, M.; et al. Growing and cultivating the forest genomics database, TreeGenes. Database 2018, 2018, bay084. [Google Scholar] [CrossRef] [PubMed]
- Hwarari, D.; Guan, Y.; Li, R.; Movahedi, A.; Chen, J.; Yang, L. Comprehensive Bioinformatics and Expression Analysis of TCP Transcription Factors in Liriodendron chinense Reveals Putative Abiotic Stress Regulatory Roles. Forests 2022, 13, 1401. [Google Scholar] [CrossRef]
- Poole, R.L. The TAIR database. In Plant Bioinformatics: Methods and Protocols; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2007; pp. 179–212. [Google Scholar]
- Jin, J.; Tian, F.; Yang, D.C.; Meng, Y.Q.; Kong, L.; Luo, J. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2017, 45, D1040–D1045. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol Evol. 2018, 35, 1547. [Google Scholar] [CrossRef] [PubMed]
- Thumuluri, V.; Almagro Armenteros, J.J.; Johansen, A.R.; Nielsen, H.; Winther, O. DeepLoc 2.0: Multi-label subcellular localization prediction using protein language models. Nucl Acids Res. 2022, 50, W228–W234. [Google Scholar] [CrossRef]
- Ke, Y.; Xu, M.; Hwarari, D.; Ahmad, B.; Li, R.; Guan, Y. OSCA Genes in Liriodendron chinense: Characterization, Evolution, and Response to Abiotic Stress. Forests 2022, 13, 1835. [Google Scholar] [CrossRef]
- Yang, Y.; Xu, M.; Luo, Q.; Wang, J.; Li, H. De novo transcriptome analysis of Liriodendron chinense petals and leaves by Illumina sequencing. Gene 2014, 534, 155–162. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucl. Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef] [Green Version]
- Ragueneau, E.; Shrivastava, A.; Morris, J.H.; Del-Toro, N.; Hermjakob, H.; Porras, P. IntAct App: A Cytoscape application for molecular interaction network visualization and analysis. Bioinformatics 2021, 37, 3684–3685. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Radani, Y.; Ahmad, B.; Movahedi, A.; Yang, L. Identification and characteristics of SnRK genes and cold stress-induced expression profiles in Liriodendron chinense. BMC Genom. 2022, 23, 708. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Hwarari, D.; Li, Y.; Ahmad, B.; Min, T.; Zhang, W. The bZIP transcription factors in Liriodendron chinense: Genome-wide recognition, characteristics, and cold stress response. Front. Plant Sci. 2022, 13, 1035627. [Google Scholar] [CrossRef] [PubMed]
- Mitteer, D.R.; Greer, B.D.; Randall, K.R.; Briggs, A.M. Further Evaluation of Teaching Behavior Technicians to Input Data and Graph Using GraphPad Prism. Behav. Anal. 2020, 20, 81–93. [Google Scholar] [CrossRef] [PubMed]
- Crooks, G.E.; Hon, G.; Chandonia, J.M.; Brenner, S.E. WebLogo: A sequence logo generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, H.; Hwarari, D.; Ma, H.; Xu, H.; Yang, L.; Luo, Y. Genomic survey of TCP transcription factors in plants: Phylogenomics, evolution and their biology. Front. Genet. 2022, 13, 1060546. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Xi, H.; Park, S.; Yun, Y.; Park, J. Genome-wide comparative analyses of GATA transcription factors among seven Populus genomes. Sci. Rep. 2021, 11, 16578. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Ren, C.; Zou, L.; Wang, Y.; Li, S.; Liang, Z. Characterization of the GATA gene family in Vitis vinifera: Genome-wide analysis, expression profiles, and involvement in light and phytohormone response. Genome 2018, 61, 713–723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Emms, D.M.; Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 2019, 20, 238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilbrandt, J.; Misof, B.; Panfilio, K.A.; Niehuis, O. Repertoire-wide gene structure analyses: A case study comparing automatically predicted and manually annotated gene models. BMC Genom. 2019, 20, 753. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peng, W.; Li, W.; Song, N.; Tang, Z.; Liu, J.; Wang, Y.; Pan, S.; Dai, L.; Wang, B. Genome-Wide Characterization, Evolution, and Expression Profile Analysis of GATA Transcription Factors in Brachypodium distachyon. Int J. Mol. Sci. 2021, 22, 2026. [Google Scholar] [CrossRef]
- Li, J.; Zhang, Z.; Vang, S.; Jun, Y.; Gane, K.-S.W.; Jun, W. Correlation Between Ka/Ks and Ks is Related to Substitution Model and Evolutionary Lineage. J. Mol. Evol. 2009, 68, 414–423. [Google Scholar] [CrossRef] [PubMed]
- Rao, V.S.; Srinivas, K.; Sujini, G.N.; Kumar, G.N. Protein-protein interaction detection: Methods and analysis. Int. J. Proteom. 2014, 2014, 147648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucl. Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Guan, Y.; Liu, S.; Wu, W.; Hong, K.; Li, R.; Zhu, L. Genome-wide identification and cold stress-induced expression analysis of the CBF gene family in Liriodendron chinense. J. Res. 2021, 32, 2531–2543. [Google Scholar] [CrossRef]
- Schwechheimer, C.; Schröder, P.M.; Blaby-Haas, C.E. Plant GATA Factors: Their Biology, Phylogeny, and Phylogenomics. Ann. Rev. Plant Biol. 2022, 73, 123–148. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Yang, Y.; Lou, S.; Wei, W.; Zhao, Z.; Ren, Y. Genome-wide characterization and gene expression analyses of GATA transcription factors in Moso bamboo (Phyllostachys edulis). Int. J. Mol. Sci. 2019, 21, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, X.; Yu, Q.; Zeng, J.; He, X.; Liu, W. Genome-wide identification and characterization of GATA family genes in wheat. BMC Plant Biol. 2022, 22, 372. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zou, X.; Huang, Z.; Fan, S.; Qun, G.; Liu, A. Genome-wide identification and analysis of the evolution and expression patterns of the GATA transcription factors in three species of Gossypium genus. Gene 2019, 680, 72–83. [Google Scholar] [CrossRef] [PubMed]
- Nishii, A.; Takemura, M.; Fujita, H.; Shikata, M.; Yokota, A.; Kohchi, T. Characterization of a novel gene encoding a putative single zinc-finger protein, ZIM, expressed during the reproductive phase in Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 2000, 64, 1402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wei, X.; Li, Y.; Zhu, X.; Liu, X.; Ye, X.; Zhou, M.; Zhang, Z. The GATA transcription factor TaGATA1 recruits demethylase TaELF6-A1 and enhances seed dormancy in wheat by directly regulating TaABI5. J. Integr. Plant Biol. 2022, 65, 1262–1276. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, H.; Zhao, C.; Yang, C.; Xu, Q.; Yuan, H.; Yang, H.; Zeng, X. Identification of a novel transcription factor under long-term drought resistance in highland barley: A DNA affinity purification sequencing-based transcriptomic analysis. Chem. Biol. Technol. Agric. 2023, 10, 1. [Google Scholar] [CrossRef]
- Kohchi, T.; Yamato, K.T.; Ishizaki, K.; Yamaoka, S.; Nishihama, R. Development and Molecular Genetics of Marchantia polymorpha. Annu. Rev. Plant Biol. 2021, 72, 677–702. [Google Scholar] [CrossRef]
- Zhang, X.; Zhou, Y.; Ding, L.; Wu, Z.; Liu, R.; Meyerowitz, E.M. Transcription repressor HANABA TARANU controls flower development by integrating the actions of multiple hormones, floral organ specification genes, and GATA3 family genes in Arabidopsis. Plant Cell 2013, 25, 83–101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clark, J.W.; Donoghue, P.C. Whole-genome duplication and plant macroevolution. Trends Plant Sci. 2018, 23, 933–945. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, W.; Guo, Y.; Chen, Y.; Wu, D.; Jiang, L. Genome-wide identification, phylogenetic and expression pattern analysis of GATA family genes in Brassica napus. BMC Plant Biol. 2020, 20, 543. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Liu, F.; Wang, L.; Huang, S.; Yu, J. Nonsynonymous substitution rate (Ka) is a relatively consistent parameter for defining fast-evolving and slow-evolving protein-coding genes. Biol. Direct 2011, 6, 13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, W.; Zhu, S.; Xu, L.; Zhu, L.; Wang, D.; Liu, Y.; Liu, S.; Hao, Z.; Lu, Y.; Yang, L.; et al. Genome-wide identification of the Liriodendron chinense WRKY gene family and its diverse roles in response to multiple abiotic stress. BMC Plant Biol. 2022, 22, 25. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Lv, G.; Ge, J.; He, B.; Zhang, Z.; Hu, Z. Genome-wide identification of the GATA transcription factor family and their expression patterns under temperature and salt stress in Aspergillus oryzae. AMB Express 2021, 11, 56. [Google Scholar] [CrossRef]
- Wang, J.; Sheng, J.; Zhu, J.; Hu, Z.; Diao, Y. Comparative transcriptome analysis and identification of candidate adaptive evolution genes of Miscanthus lutarioriparius and Miscanthus sacchariflorus. Physiol. Mol. Biol. Plants 2021, 27, 1499–1512. [Google Scholar] [CrossRef] [PubMed]
- Díaz, M.L.; Soresi, D.S.; Basualdo, J.; Cuppari, S.J.; Carrera, A. Transcriptomic response of durum wheat to cold stress at reproductive stage. Mol. Biol. Rep. 2019, 46, 2427–2445. [Google Scholar] [CrossRef] [PubMed]
- Kashyap, P.; Deswal, R. Two ICE isoforms showing differential transcriptional regulation by cold and hormones participate in Brassica juncea cold stress signaling. Gene 2019, 695, 32–41. [Google Scholar] [CrossRef] [PubMed]



| Gene ID | Gene Name | Chromosome | Chromosome Location | aa | Mw(KDa) | pI | Localization |
|---|---|---|---|---|---|---|---|
| Lchi11034 | LcGATA1 | Chr 1 | 76673911; 76676124 | 327 | 36.0 | 5.53 | Nucleus |
| Lchi00930 | LcGATA2 | Chr 1 | 97844054; 97846142 | 321 | 35.5 | 6.76 | Nucleus |
| Lchi00761 | LcGATA3 | Chr 1 | 103142854; 103144266 | 297 | 32.7 | 9.34 | Nucleus |
| Lchi33577 | LcGATA4 | Chr 2 | 27371; 35262 | 135 | 15.1 | 5.25 | Nucleus |
| Lchi01856 | LcGATA5 | Chr 2 | 76545610; 76556256 | 393 | 43.5 | 8.4 | Nucleus |
| Lchi24152 | LcGATA6 | Chr 3 | 27419892; 27425752 | 222 | 25.2 | 6.15 | Nucleus |
| Lchi05395 | LcGATA7 | Chr 3 | 38862093; 38864782 | 330 | 35.4 | 5.67 | Nucleus |
| Lchi11226 | LcGATA8 | Chr 4 | 38490148; 38502630 | 366 | 41.0 | 4.63 | Cytoplasm/Nucleus |
| Lchi03424 | LcGATA9 | Chr 6 | 11359264; 11359944 | 199 | 22.4 | 9.94 | Nucleus |
| Lchi08934 | LcGATA10 | Chr 6 | 17595500; 17597071 | 343 | 37.2 | 5.83 | Nucleus |
| Lchi30686 | LcGATA11 | Chr 10 | 9467120; 9467702 | 120 | 13.4 | 9.71 | Nucleus |
| Lchi01461 | LcGATA12 | Chr 11 | 55131852; 55133154 | 268 | 29.1 | 6.30 | Nucleus |
| Lchi01268 | LcGATA13 | Chr 11 | 59911870; 59920955 | 583 | 64.3 | 6.93 | Nucleus |
| Lchi13044 | LcGATA14 | Chr 13 | 5648353; 5661943 | 624 | 67.7 | 6.1 | Nucleus |
| Lchi03334 | LcGATA15 | Chr 14 | 68497886; 68527212 | 504 | 55.9 | 9.81 | Nucleus |
| Lchi08164 | LcGATA16 | Chr 15 | 45414499; 45425579 | 289 | 31.7 | 10.05 | Nucleus |
| Lchi17957 | LcGATA17 | Chr 16 | 25722343; 25736319 | 339 | 36.7 | 5.83 | Nucleus |
| Lchi05152 | LcGATA18 | Chr 17 | 66662271; 66670843 | 299 | 33.0 | 8.33 | Nucleus |
| SEQ_1 | SEQ_2 | KA | KS | KA_KS |
|---|---|---|---|---|
| LCGATA10 | LcGATA2 | 0.17044698 | 0.85528245 | 0.19928735 |
| LCGATA16 | LcGATA7 | 0.25063103 | 1.27657256 | 0.19633121 |
| LCGATA18 | LcGATA15 | 0.3122753 | 1.03194528 | 0.30260839 |
| LCGATA13 | LcGATA5 | 0.45604965 | 2.18390847 | 0.20882269 |
| LCGATA17 | LcGATA14 | 0.35202336 | 1.50301022 | 0.23421222 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hwarari, D.; Radani, Y.; Guan, Y.; Chen, J.; Liming, Y. Systematic Characterization of GATA Transcription Factors in Liriodendron chinense and Functional Validation in Abiotic Stresses. Plants 2023, 12, 2349. https://doi.org/10.3390/plants12122349
Hwarari D, Radani Y, Guan Y, Chen J, Liming Y. Systematic Characterization of GATA Transcription Factors in Liriodendron chinense and Functional Validation in Abiotic Stresses. Plants. 2023; 12(12):2349. https://doi.org/10.3390/plants12122349
Chicago/Turabian StyleHwarari, Delight, Yasmina Radani, Yuanlin Guan, Jinhui Chen, and Yang Liming. 2023. "Systematic Characterization of GATA Transcription Factors in Liriodendron chinense and Functional Validation in Abiotic Stresses" Plants 12, no. 12: 2349. https://doi.org/10.3390/plants12122349
APA StyleHwarari, D., Radani, Y., Guan, Y., Chen, J., & Liming, Y. (2023). Systematic Characterization of GATA Transcription Factors in Liriodendron chinense and Functional Validation in Abiotic Stresses. Plants, 12(12), 2349. https://doi.org/10.3390/plants12122349






