Huanglongbing Pandemic: Current Challenges and Emerging Management Strategies
Abstract
1. Introduction
2. Causative Agent, Genomics, and Pathogenesis Mechanism
| Sr. No | Candidatus Liberibacter spp. | Strain | Host | Sample Origin | Genome Size (Mb) | Number CDS Present in the Genome | Reference |
|---|---|---|---|---|---|---|---|
| 1 | Candidatus Liberibacter asiaticus | A4 (CP010804.1) | Citrus reticulata | China: Guangdong | 1.23025 | 1067 | [59] |
| 2 | Candidatus Liberibacter asiaticus | Gxpsy (CP004005.1) | Diaphorinacitri | China: Guangxi | 1.26824 | 1094 | [57] |
| 3 | Candidatus Liberibacter asiaticus | JRPAMB1 (CP040636.1) | Diaphorinacitri | USA: Florida | 1.23716 | 1072 | [62] |
| 4 | Candidatus Liberibacter asiaticus | TaiYZ2 (CP041385.1) | Citrus maxima | Thailand: Songkhla | 1.23062 | 1067 | [77] |
| 5 | Candidatus Liberibacter asiaticus | psy62 (CP001677.5) | - | USA: Florida | 1.22732 | 1049 | [56] |
| 6 | Candidatus Liberibacter asiaticus | JXGC (CP019958.1) | Citrus | China: Jiangxi | 1.22516 | 1033 | [60] |
| 7 | Candidatus Liberibacter asiaticus | Ishi-1 (AP014595.1) | Diaphorinacitri | Japan: Ishigaki | 1.19085 | 1001 | [58] |
| 8 | Candidatus Liberibacter asiaticus | AHCA1 (CP029348.1) | Diaphorinacitri | USA: California | 1.23375 | 1056 | [61] |
| 9 | Candidatus Liberibacter asiaticus | FL17 (JWHA00000000.1) | Citrus | USA: Florida | 1.22725 | 1019 | [78] |
| 10 | Candidatus Liberibacter asiaticus | YNJS7C (QXDO00000000) | Citrus | China: Yunnan | 1.25898 | 1102 | [79] |
| 11 | Candidatus Liberibacter asiaticus | YCPsy LIIM00000000) | Diaphorinacitri | China: Guangdong | 1.233647 | 1037 | [80] |
| 12 | Candidatus Liberibacter asiaticus | LBR19TX2 (VTMA00000000 | - | USA: Texas | 1.20275 | 1008 | [67] |
| 13 | Candidatus Liberibacter asiaticus | LBR23TX5 (VTMB00000000) | - | USA: Texas | 1.20347 | 1009 | [67] |
| 14 | Candidatus Liberibacter asiaticus | AHCA17 (VNFL00000000) | Citrus maxima | USA: California | 1.20862 | 1036 | [81] |
| 15 | Candidatus Liberibacter asiaticus | YNXP-1 (VIGA00000000) | Cuscuta | China: Yunnan | 1.20707 | 1031 | - |
| 16 | Candidatus Liberibacter asiaticus | SGCA16 (VTLZ00000000) | - | USA: San Gabriel | 1.20994 | 1015 | [67] |
| 17 | Candidatus Liberibacter asiaticus | JXGZ-1 (VIQL00000000) | Cuscuta | China: JiangXi | 1.21799 | 1040 | - |
| 18 | Candidatus Liberibacter asiaticus | DUR1TX1 VTLT00000000.1) | - | USA: Texas | 1.20629 | 1011 | [67] |
| 19 | Candidatus Liberibacter asiaticus | Mex8 (VTLU00000000.1) | - | Mexico: Mexicali | 1.24313 | 1042 | [67] |
| 20 | Candidatus Liberibacter asiaticus | SGCA5 (LMTO00000000.1) | Orange citrus | USA: San Gabriel | 1.20138 | 1001 | [80] |
| 21 | Candidatus Liberibacter asiaticus | CHUC (VTLV00000000) | - | China | 1.20845 | 1032 | [67] |
| 22 | Candidatus Liberibacter asiaticus | TX2351 (MTIM00000000) | Asian citrus psyllid | USA: Texas | 1.252 | 1129 | [82] |
| 23 | Candidatus Liberibacter asiaticus | GFR3TX3 (VTLR00000000) | - | USA: Texas | 1.20932 | 1013 | [67] |
| 24 | Candidatus Liberibacter asiaticus | HHCA16 (VTLY00000000) | - | USA: Hacienda Heights | 1.20705 | 1012 | [67] |
| 25 | Candidatus Liberibacter asiaticus | MFL16 (VTLX00000000) | - | USA: Florida | 1.19922 | 1012 | [67] |
| 26 | Candidatus Liberibacter asiaticus | DUR2TX1 (VTLS00000000) | - | USA: Texas | 1.21232 | 1009 | [67] |
| 27 | Candidatus Liberibacter asiaticus | CRCFL16 (VTLW00000000) | - | USA: Florida | 1.20828 | 1028 | [67] |
| 28 | Candidatus Liberibacter asiaticus | HHCA (JMIL00000000.2) | Citrus sp. | USA: Hacienda Heights | 1.15062 | 867 | [59] |
| 29 | Candidatus Liberibacter asiaticus | TX1712 (QEWL00000000) | Citrus sinensis | USA: Texas | 1.20333 | 0 | [83] |
| 30 | Candidatus Liberibacter asiaticus | SGpsy (QFZJ00000000.1) | Diaphorinacitri | USA: San Gabriel | 0.769888 | 0 | [61] |
| 31 | Candidatus Liberibacter asiaticus | SGCA1 (QFZT00000000.1) | USA: San Gabriel | 0.233414 | 557 | [61] | |
| 32 | Candidatus Liberibacter asiaticus | YCPsy (LIIM00000000) | Diaphorinacitri | Guangdong, China | 1.233647 | - | [80] |
| 33 | Candidatus Liberibacter asiaticus | PA19 (WOXD01000000) | Kinnow mandarin | Pakistan | 1.224156 | 1059 | [84] |
| 34 | Candidatus Liberibacter asiaticus | PA20 (WOUN01000000) | Kinnow mandarin | Pakistan | 1.226225 | 1062 | [84] |
| 35 | Candidatus Liberibacter asiaticus | CoFLP1 (CP054558.1) | Diaphorinacitri | Colombia: Municipio Dibulla | 1.231639 | 1048 | [64] |
| 36 | Candidatus Liberibacter asiaticus | 9PA (JABDRZ000000000.1) | Citrus sinensis | Brazil (South America) | 1.231881 | - | [85] |
| 37 | Candidatus Liberibacter asiaticus | MFL16 (VTLX00000000) | Citrus | USA: Florida | 1,199,225 bp | - | [67] |
| 38 | Candidatus Liberibacter asiaticus | CRCFL16 (VTLW00000000) | Citrus | USA: Florida | 1,208,280 bp | - | [67] |
| 39 | Candidatus Liberibacter asiaticus | ReuSP1 (CP061535.1) | Diaphorinacitri | France: La Reunion | 1.230064 | 1043 | [65] |
| 40 | Candidatus Liberibacter asiaticus | Tabriz.3(JAKQYA000000000.1) | Elaeagnus angustifolia | Iran: East Azerbaijan, Tabriz | 1.22409 | 589 | - |
| 41 | Candidatus Liberibacter asiaticus | YNHK-2 (WUUB01000000.1) | Citrus | China: Yunnan | 1.08957 | - | [86] |
| 42 | Candidatus Liberibacter asiaticus | A-SBCA19 (JADBIB010000000.1) | Diaphorinacitri | USA: California, San Bernardino County | 1.18688 | 1067 | [87] |
| 43 | Candidatus Liberibacter americanus | Sao Paulo (NC_022793) | Citrus sinensis | Brazil | 1.1952 | 945 | [73] |
| 44 | Candidatus Liberibacter americanus | PW_SP | Catharanthus roseus | Brazil: Sao Paulo | 1.17607 | 924 | - |
| 45 | Candidatus Liberibacter africanus | PTSAPSY | Psyllid | South Africa: Pretoria | 1.19223 | 1036 | - |
Pathogen Virulence Factors
3. HLB Diagnosis
3.1. Electron Microscopy
3.2. Molecular-Based Assays
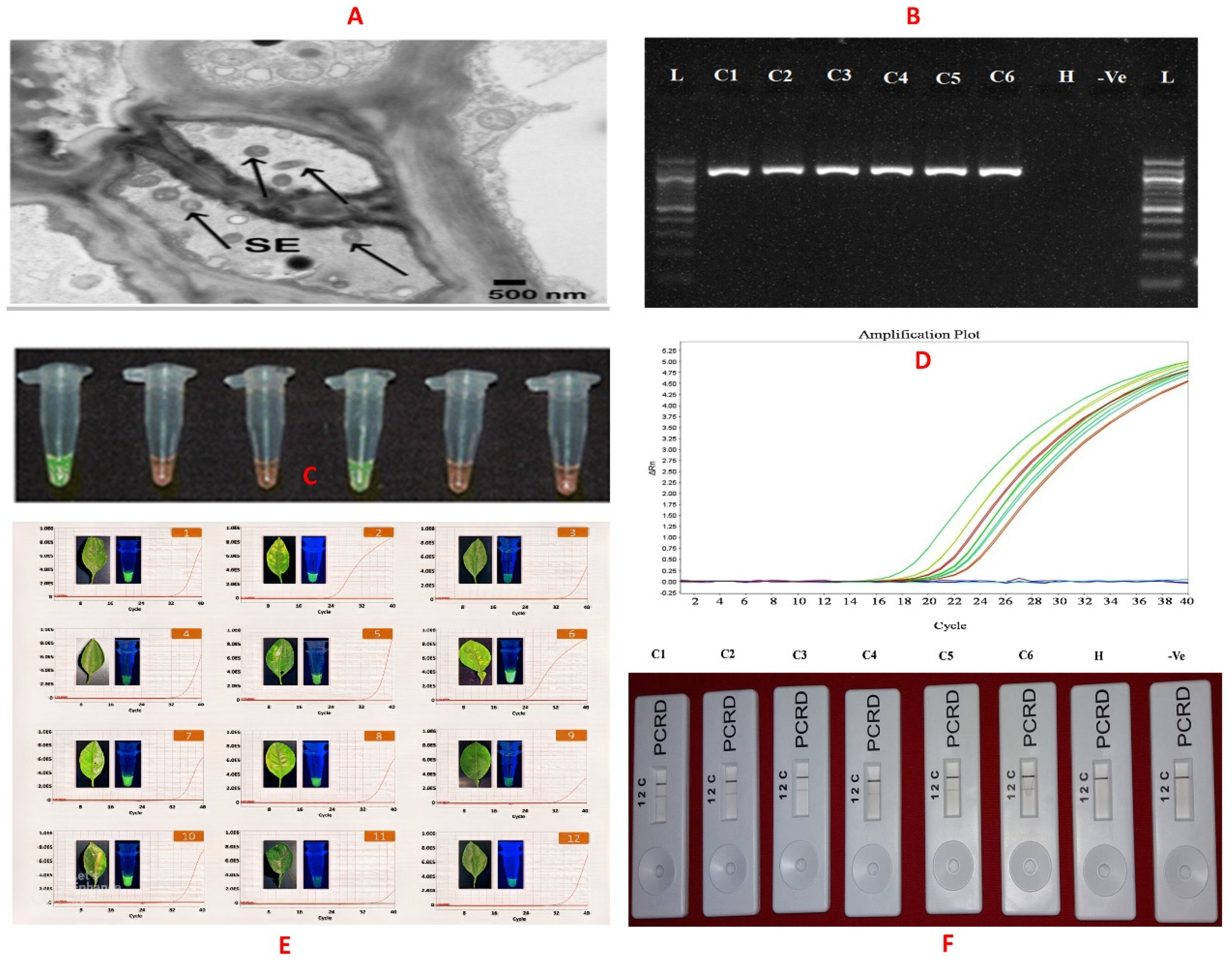
3.3. Early Diagnosis
4. Triangular Management Strategies
4.1. Disease Control Strategies at Pathogen Level
4.1.1. Protein-Based Approach/Targeting Important Proteins of CLas
- Transport protein
Znu System
Amino Acid Uptake System
Sec-Translocase/Translocon (SecY/SecE/SecG)
- Transcription regulator
- Hydrolase family enzyme
Phosphatase
Serine Protease (Protease IV Transmembrane Protein)
- Antioxidants Protein
- Nucleotide biosynthesis
Bifunctional Enzyme 5-Aminoimidazole-4-Carboxamide Ribonucleotide Formyl Transferase/Inosine Monophosphate Cyclohydrolase (ATIC)
Inosine-5′-Monophosphate Dehydrogenase
- Fatty acid biosynthesis
Enoyl-Acyl Carrier Protein Reductase I (FabI)
β-Hydroxyacyl-acyl Carrier Protein Dehydratase (FabZ)
- Amino Acid Biosynthesis
4.1.2. HLB management with Antimicrobial Chemicals
Engineered Nanomaterials (ENMs) for HLB Management
4.1.3. Thermotherapy and Cryotherapy
5. Management Strategies at Host Level
5.1. Activation of Plant Immune System
5.2. Transgenic Approach
5.2.1. Emerging Potential Genes as Transgene
2S Albumin
Miraculin-Like Proteins (MLPs)
Putranjiva Roxburghii Trypsin Inhibitor (PRTI)
5.3. Systemic Acquired Resistance (SAR)
6. Management of Asian Citrus Psyllid (ACP)
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wu, G.A.; Prochnik, S.; Jenkins, J.; Salse, J.; Hellsten, U.; Murat, F.; Perrier, X.; Ruiz, M.; Scalabrin, S.; Terol, J.; et al. Sequencing of diverse mandarin, pummelo and orange genomes reveals complex history of admixture during citrus domestication. Nat. Biotechnol. 2014, 32, 656–662. [Google Scholar] [CrossRef] [PubMed]
- Bové, J.M. Huanglongbing: A destructive, newly-emerging, century-old disease of citrus. J. Plant Pathol. 2006, 88, 7–37. [Google Scholar]
- Gottwald, T.R. Current epidemiological understanding of citrus huanglongbing. Annu. Rev. Phytopathol. 2010, 48, 119–139. [Google Scholar] [CrossRef] [PubMed]
- Reinking, O.A. Diseases of economic plants in Southern China. Philipp. Agric. 1919, 8, 109–135. [Google Scholar]
- Husain, M.A.; Nath, D. The citrus psylla (Diaphorinacitri, Kuw.) [Psyllidae: Homoptera]. Mem. Dept. Agric. India Entomol. Ser. 1927, 10, 5–27. [Google Scholar]
- Capoor, S.P. Decline of citrus in India. Bull. Natl. Inst. Sci. 1963, 24, 48–64. [Google Scholar]
- Beattie, G.A.C.; Holford, P.; Mabberley, D.J.; Haigh, A.M.; Bayer, R. Aspects and insights of Australia-Asia collaborative research on huanglongbing. In Proceedings of the International Workshop for the Prevention of Citrus Greening Disease in Severely Infected Areas, Ishigaki, Japan, 6–7 December 2006; Japanese Ministry of Agriculture, Forestry and Fisheries: Tokyo, Japan, 2006; pp. 47–64. [Google Scholar]
- Merwe, A.J.; Anderssen, F.G. Chromium and manganese toxicity. Is it important in Transvaal citrus greening. Farming S. Afr. 1937, 12, 439–440. [Google Scholar]
- Coletta-Filho, H.D.; Targon, M.; Takita, M.A.; De Negri, J.D.; Pompeu, J.; Machado, M.A.; do Amaral, A.M.; Muller, G.W. First report of the causal agent of Huanglongbing (Candidatus Liberibacter asiaticus) in Brazil. Plant Dis. 2004, 88, 1382. [Google Scholar] [CrossRef]
- Halbert, S.E. The discovery of huanglongbing in Florida. In Proceedings of the Second International Citrus Canker and Huanglongbing Research Workshop, Orlando, FL, USA, 7–11 November 2005; p. H-3. [Google Scholar]
- Molki, B.; Call, D.R.; Ha, P.T.; Omsland, A.; Gang, D.R.; Lindemann, S.R.; Killiny, N.; Beyenal, H. Growth of ‘Candidatus Liberibacter asiaticus’ in a host-free microbial culture is associated with microbial community composition. Enzym. Microb. Technol. 2020, 142, 109691. [Google Scholar] [CrossRef]
- da Graca, J.V.; Kunta, M.; Setamou, M.; Rascoe, J.; Li, W.; Nakhla, M.K.; Salas, B.; Bartels, D.W. Huanglongbing in Texas: Report on the first detections in commercial citrus. J. Citrus Pathol. 2015, 2, 27939. [Google Scholar] [CrossRef]
- Kumagai, L.B.; LeVesque, C.S.; Blomquist, C.L.; Madishetty, K.; Guo, Y.; Woods, P.W.; Rooney-Latham, S.; Rascoe, J.; Gallindo, T.; Schnabel, D.; et al. First report of Candidatus Liberibacter asiaticus associated with citrus huanglongbing in California. Plant Dis. 2013, 97, 283. [Google Scholar] [CrossRef]
- Halbert, S.E.; Manjunath, K.L.; Ramadugu, C.; Brodie, M.W.; Webb, S.E.; Lee, R.F. Trailers transporting organges to processing plants move asian citrus psyllids. Florida Entomol. 2010, 93, 33–38. [Google Scholar] [CrossRef]
- Luis, M.; Collazo, C.; Llauger, R.; Blanco, E.; Pena, I.; López, D.; González, C.; Casín, J.C.; Batista, L.; Kitajima, E.; et al. Occurrence of citrus Huanglongbing in Cuba and association of the disease with Candidatus Liberibacter asciaticus. J. Plant Pathol. 2009, 91, 709–712. [Google Scholar] [CrossRef]
- Oberheim, A.P.; Brown, S.E.; McLaughlin, W.A. The identification and distribution of citrus greening disease in Jamaica. In Proceedings of the 2nd International Research Conference on Huanglongbing, Orlando, FL, USA, 10–14 January 2011; p. 114. [Google Scholar]
- Manjunath, K.L.; Ramadugu, C.; Majil, V.M.; Williams, S.; Irey, M.; Lee, R.F. First report of the citrus huanglongbing associated bacterium ‘Candidatus Liberibacter asiaticus’ from sweet orange, mexican lime, and asian citrus psyllid in Belize. Plant Dis. 2010, 94, 781. [Google Scholar] [CrossRef] [PubMed]
- Trujillo-Arriaga, J.; Sanchez, A.H.; Robles, G.P.; de la Rosa, A.A.; Delgadillo, V.I.; Márquez, S.M. Antecedantes y situacion actual de Huanglongbing de loscıtricosen Mexico. In Proceedings of the 1er Simposio Nacional Sobre Investigacion para el Manejo del Psılido Asiatico de losCıtricos y el HLB en Mexico, Monterrey, Mexico, 8–9 December 2010; pp. 1–7. [Google Scholar]
- Faghihi, M.M.; Salehi, M.; Bagheri, A.; Izadpanah, K. First report of citrus huanglongbing disease on orange in Iran. Plant Pathol. 2009, 58, 793. [Google Scholar] [CrossRef]
- CABI. Citrus Huanglongbing (Greening) Disease (Citrus Greening). In Invasive Species Compendium; CABI International: Wallingford, UK, 2020. [Google Scholar]
- Catling, H.D.; Garnier, M.; Bove, J.M. Presence of citrus greening disease in Bangladesh and a new method for rapid diagnosis. FAO Plant Prot. Bull. 1978, 26, 16–18. [Google Scholar]
- EPPO (European and Mediterranean Plant Protection Organization). ‘Candidatus Liberibacter asiaticus’. EPPO Datasheets on Pests Recommended for Regulation. 2022. Available online: https://gd.eppo.int (accessed on 25 November 2022).
- Garnier, M.; Bové, J.M. Huanglongbing in Cambodia, Laos and Myanmar. In Proceedings of the 14th Conference of the International Organization of Citrus Virologists, Riverside, CA, USA, 2000; da Graça, J.V., Lee, R.F., Yokomi, R.K., Eds.; IOCV: Concordia, Argentina, 2000; Volume 14, pp. 378–380. [Google Scholar] [CrossRef]
- Bové, J.M.; Garnier, M.; Ahlawat, Y.S.; Chakraborty, N.K.; Varma, A. Detection of the Asian strains of the greening BLO by DNA-DNA hybridization in Indian orchard trees and Malaysian Diaphorina Citri Psyllids. Int. Organ. Citrus Virol. Conf. Proc. 1993, 12, 1957–2010. [Google Scholar] [CrossRef]
- Aubert, B.; Garnier, M.; Guillaumin, D.; Herbagyandono, B.; Setiobudi, L.; Nurhadi, F. Greening, a serious threat for the citrus productions of the Indonesian archipelago. Future prospects of integrated control. Fruits 1985, 40, 549–563. [Google Scholar]
- Salehi, M.; Faghihi, M.M.; Khanchezar, A.; Bagheree, A.; Izadpanah, K. Distributioin of citrus Huanglongbing disease and its vector in southern Iran. Iranian J. Plant Pathol. 2012, 48, 195–208. [Google Scholar]
- Miyakawa, T.; Tsuno, K. Occurrence of citrus greening disease in the southern islands of Japan. Jpn. J. Phytopathol. 1989, 55, 667–670. [Google Scholar] [CrossRef]
- Regmi, C.; Garnier, M.; Bové, J.M. Detection of the Asian Huanglongbing (Greening) Liberobacter in Nepal by DNA-DNA hybridization. Int. Organ. Citrus Virol. Conf. Proc. 1996, 13, 267–270. [Google Scholar]
- Garnier, M.; Bové, J.M. Distribution of the huanglongbing (greening) Liberobacter species in fifteen African and Asian countries. Int. Organ. Citrus Virol. Conf. Proc. 1996, 13, 1957–2010. [Google Scholar] [CrossRef]
- Bové, J.M.; Garnier, M. Citrus greening and psylla vectors of the disease in the Arabian Peninsula. Int. Organ. Citrus Virol. Conf. Proc. 1984, 9, 1957–2010. [Google Scholar] [CrossRef]
- Promintara, M. Simultaneous infection of mandarin (Citrus reticulata Bl.) with tristeza virus and greening organism in Thailand. In Proceedings of the 4th International Asia Pacific Conference on Citrus Rehabilitation, Chiang Mai, Thailand, 4–10 February 1990; FAO-UNDP RAS/86/022 Regional Project. pp. 96–99. [Google Scholar]
- Bové, J.M.; Navarro, L.; Bonnet, P.; Zreik, L.; Garnier, M. Reaction of citrus cultivars to graft-inoculation of Phytoplasma aurantifolia-infected lime shoots. Int. Organ. Citrus Virol. Conf. Proc. (1957–2010) 1996, 13, 13. [Google Scholar] [CrossRef]
- MAG. Declaraemergenciafitosanitarianacionalporpresencia de HLB, Enfermedad de Loscítricos, El Salvador, Ministerio de Agricultura y Ganadería. 2020. Available online: http://www.mag.gob.sv/mag-declara-emergencia-fitosanitaria-nacional-por-presencia-de-hlb-enfermedad-de-los-citricos/ (accessed on 25 November 2022).
- Cellier, G.; Moreau, A.; Cassam, N.; Hostachy, B.; Ryckewaert, P.; Aurela, L.; Picard, R.; Lombion, K.; Rioualec, A.L. First report of ‘Candidatus Liberibacter asiaticus’ associated with huanglongbing on Citrus latifolia in Martinique and Guadeloupe, French West Indies. Plant Dis. 2014, 98, 683. [Google Scholar] [CrossRef]
- Leite, R., Jr.; Cordeiro, A.B.; Meneguim, L. Primera deteccion de la enfermedadHuanglongbing (HLB) enasociacion con la bacteria Candidatus Liberibacter asiaticus en Paraguay. In Proceedings of the VII Congresso Argentino de Citricultura, Puerto Iguazu, Missiones, Argentina, 15–17 May 2013. [Google Scholar]
- Marys, E.; Rodríguez-Román, E.; Mejías, R.; Mejías, A.; Mago, M.; Hernández, Y. First report on molecular evidence of Candidatus Liberibacter asiaticus associated with citrus Huanglongbing in Venezuela. J. Plant Pathol. 2020, 102, 1333. [Google Scholar] [CrossRef]
- Aubert, B.; Garnier, M.; Cassin, J.E.; Bertin, Y. Citrus greening disease survey in East and West African countries South of Sahara. Int. Organ. Citrus Virol. Conf. Proc. (1957–2010) 1988, 10, 231–237. [Google Scholar] [CrossRef]
- Garnier, M.; Jagoueix-Eveillard, S.; Cronje, P.R.; Le Roux, H.F.; Bove, J.M. Genomic characterization of a liberibacter present in an ornamental rutaceous tree, Calodendrumcapense, in the Western Cape Province of South Africa. Proposal of ‘Candidatus Liberibacter africanus subsp. capensis’. Int. J. Syst. Evol. Microbiol. 2000, 50, 2119–2125. [Google Scholar] [CrossRef] [PubMed]
- Korsten, L.; Jagoueix, S.; Bové, J.M.; Garnier, M. Huanglongbing (greening) detection in South Africa. Int. Organ. Citrus Virol. Conf. Proc. (1957–2010) 1996, 13. [Google Scholar] [CrossRef]
- Kalyebi, A.; Aisu, G.; Ramathani, I.; Ogwang, J.; McOwen, N.; Russell, P. Detection and identification of etiological agents (Liberibacter spp.) associated with citrus greening disease in Uganda. Uganda J. Agric. Sci. 2015, 16, 43–54. [Google Scholar] [CrossRef][Green Version]
- Ghosh, D.; Bhose, S.; Mukherjee, K.; Baranwal, V.K. Sequence and evolutionary analysis of ribosomal DNA from Huanglongbing (HLB) isolates of western India. Phytoparasitica 2013, 41, 295–305. [Google Scholar] [CrossRef]
- Nageswara-Rao, M.; Miyata, S.I.; Ghosh, D.; Irey, M.; Garnsey, S.M.; Gowda, S. Development of rapid, sensitive and non-radioactive tissue-blot diagnostic method for the detection of citrus greening. Mol. Cell. Probes 2013, 27, 176–183. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.Z.; Wang, X.F.; Zhou, Y.; Li, Z.A.; Tang, K.Z.; Zhou, C. Diversity of the OMP gene in ‘Candidatus Liberibacter asiaticus’ in China. J. Plant Pathol. 2011, 93, 211–214. [Google Scholar]
- Kokane, S.B.; Bhose, S.; Kokane, A.; Gubyad, M.; Ghosh, D.K. Molecular detection, identification, and sequence analysis of Candidatus Liberibacter asiaticus associated with Huanglongbing disease of citrus in North India. 3Biotech 2020, 10, 341. [Google Scholar] [CrossRef]
- Kokane, S.B.; Misra, P.; Kokane, A.D.; Gubyad, M.G.; Warghane, A.J.; Surwase, D.; Reddy, M.K.; Ghosh, D.K. Development of a real-time RT-PCR method for the detection of Citrus tristeza virus (CTV) and its implication in studying virus distribution in planta. 3Biotech 2021, 11, 431. [Google Scholar] [CrossRef]
- Ghosh, D.K.; Kokane, A.; Kokane, S.B.; Tenzin, J.; Gubyad, M.G.; Wangdi, P.; Murkute, A.A.; Sharma, A.K.; Gowda, S. Detection and molecular characterization of Candidatus Liberibacter asiaticus and Citrus tristeza virus associated with citrus decline in Bhutan. Phytopathology 2021, 111, 870–881. [Google Scholar] [CrossRef]
- Brlansky, R.H. Blight. In Compendium of Citrus Disease, 2nd ed.; Timmer, L.W., Garnsey, S.M., Graham, J.H., Eds.; APS Press: St. Paul, MN, USA, 2000; pp. 65–66. [Google Scholar]
- Tsai, J.H.; Liu, Y.H. Biology of Diaphorinacitri (Homoptera: Psyllidae) on four host plants. J. Econ. Entomol. 2000, 93, 1721–1725. [Google Scholar] [CrossRef]
- Jagoueix, S.; Bové, J.M.; Garnier, M. The phloem-limited bacterium of greening disease of citrus is a member of the alpha subdivision of the proteobacteria. Int. J. Syst. Evol. Microbiol. 1994, 44, 379–386. [Google Scholar] [CrossRef]
- Kokane, S.B.; Kokane, A.D.; Misra, P.; Warghane, A.; Kumar, P.; Gubyad, M.G.; Sharma, A.K.; Biswas, K.K.; Ghosh, D.K. In-silico characterization and RNA-binding protein based polyclonal antibodies production for detection of Citrus tristeza virus. Mol. Cell. Probes. 2020, 54, 101654. [Google Scholar] [CrossRef]
- Ghosh, D.K.; Motghare, M.; Kokane, A.; Kokane, S.; Warghane, A.; Bhose, S.; Surwase, D.; Ladaniya, M.S. First report of a ‘Candidatus phytoplasma cynodontis’-related strain (group 16SrXIV) associated with Huanglongbing disease on Citrus grandis in India. Aus. Plant Dis. Notes 2019, 14, 9. [Google Scholar] [CrossRef]
- Wang, N. A perspective of citrus Huanglongbing in the context of the mediterranean Basin. J. Plant Pathol. 2020, 102, 635–640. [Google Scholar] [CrossRef]
- Pelz-Stelinski, K.S.; Brlansky, R.H.; Ebert, T.A.; Rogers, M.E. Transmission parameters for ‘Candidatus Liberibacter asiaticus’ by Asian citrus psyllid (Hemiptera: Psyllidae). J. Econ. Entomol. 2010, 103, 1531–1541. [Google Scholar] [CrossRef] [PubMed]
- El-Shesheny, I.; Hijaz, F.; El-Hawary, I.; Mesbah, I.; Killiny, N. Impact of different temperatures on survival and energy metabolism in the Asian citrus psyllid, Diaphorina citri Kuwayama. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2016, 192, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Mann, R.S.; Pelz-Stelinski, K.; Hermann, S.L.; Tiwari, S.; Stelinski, L.L. Sexual transmission of a plant pathogenic bacterium, ‘Candidatus Liberibacter asiaticus’, between conspecific insect vectors during mating. PLoS ONE 2011, 6, e29197. [Google Scholar] [CrossRef]
- Duan, Y.; Zhou, L.; Hall, D.G.; Li, W.; Doddapaneni, H.; Lin, H.; Liu, L.; Vahling, C.M.; Gabriel, D.W.; Kelly, P.; et al. Complete genome sequence of citrus huanglongbing bacterium, ‘Candidatus Liberibacter asiaticus’ obtained through metagenomics. Mol. Plant-Microbe Interact. 2009, 22, 1011–1020. [Google Scholar] [CrossRef]
- Lin, H.; Han, C.S.; Liu, B.; Lou, B.; Bai, X.; Deng, C.; Civerolo, E.L.; Gupta, G. Complete genome sequence of a chinese strain of “Candidatus Liberibacter asiaticus. ” Genome Announc. 2013, 1, e00184-13. [Google Scholar] [CrossRef]
- Katoh, H.; Miyata, S.; Inoue, H.; Iwanami, T. Unique features of a Japanese ‘Candidatus Liberibacter asiaticus’ strain revealed by whole genome sequencing. PLoS ONE 2014, 9, e106109. [Google Scholar] [CrossRef]
- Zheng, Z.; Deng, X.; Chen, J. Whole-genome sequence of ‘Candidatus Liberibacter asiaticus’ from Guangdong, China. Genome Announc. 2014, 2, e00273-14. [Google Scholar] [CrossRef]
- Zheng, Z.; Bao, M.; Wu, F.; Van Horn, C.; Chen, J.; Deng, X. A Type 3 Prophage of ‘Candidatus Liberibacter asiaticus’ carrying a restriction-modification system. Phytopathology 2018, 108, 454–461. [Google Scholar] [CrossRef]
- Dai, Z.; Wu, F.; Zheng, Z.; Yokomi, R.; Kumagai, L.; Cai, W.; Rascoe, J.; Polek, M.; Chen, J.; Deng, X. Prophage diversity of ‘Candidatus Liberibacter asiaticus’ strains in california. Phytopathology 2019, 109, 551–559. [Google Scholar] [CrossRef]
- Petrone, J.R.; Muñoz-Beristain, A.; Glusberger, P.R.; Russell, J.T.; Triplett, E.W. Unamplified, Long-Read Metagenomic Sequencing Approach to Close Endosymbiont Genomes of Low-Biomass Insect Populations. Microorganisms 2022, 10, 513. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Wu, F.; Duan, Y.; Singerman, A.; Guan, Z. Citrus Greening: Management strategies and their economic impact. Hort. Sci. 2020, 55, 604. [Google Scholar] [CrossRef]
- Wang, Y.; Kondo, T.; He, Y.; Zhou, Z.; Lu, J. Genome Sequence Resource of ‘Candidatus Liberibacter asiaticus’ from Diaphorinacitri Kuwayama (Hemiptera: Liviidae) in Colombia. Plant Dis. 2021, 105, 193–195. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Delatte, H.; Reynaud, B.; Beattie, G.A.; Holford, P.; Cen, Y.; Wang, Y. Genome sequence resource of ‘Candidatus Liberibacter asiaticus’ from Diaphorinacitri Kuwayama (Hemiptera: Liviidae) from La Réunion. Plant Dis. 2021, 105, 1171–1173. [Google Scholar] [CrossRef]
- Toruno, T.Y.; Stergiopoulos, I.; Coaker, G. Plant-pathogen effectors: Cellular probes interfering with plant defenses in spatial and temporal manners. Annu. Rev. Phytopathol. 2016, 54, 419–441. [Google Scholar] [CrossRef]
- Thapa, S.P.; De Francesco, A.; Trinh, J.; Gurung, F.B.; Pang, Z.; Vidalakis, G.; Wang, N.; Ancona, V.; Ma, W.; Coaker, G. Genome-wide analyses of Liberibacter species provides insights into evolution, phylogenetic relationships, and virulence factors. Mol. Plant Pathol. 2020, 21, 716–731. [Google Scholar] [CrossRef]
- Kruse, A.; Fleites, L.A.; Heck, M. Lessons from one fastidious bacterium to another: What can we learn about liberibacter species from Xylella fastidiosa. Insects 2019, 10, 300. [Google Scholar] [CrossRef]
- Du, P.; Zhang, C.; Zou, X.; Zhu, Z.; Yan, H.; Wuriyanghan, H.; Li, W. “Candidatus Liberibacter asiaticus” secretes nonclassically secreted proteins that suppress host hypersensitive cell death and induce expression of plant pathogenesis-related proteins. Appl. Environ. Microbiol. 2021, 87, e00019-21. [Google Scholar] [CrossRef]
- Pitino, M.; Armstrong, C.M.; Cano, L.M.; Duan, Y. Transient expression of Candidatus Liberibacter asiaticus effector induces cell death in Nicotiana benthamiana. Front. Plant Sci. 2016, 7, 982. [Google Scholar] [CrossRef]
- Pitino, M.; Allen, V.; Duan, Y. LasΔ5315 effector induces extreme starch accumulation and chlorosis as Candidatus Liberibacter asiaticus infection in Nicotiana benthamiana. Front. Plant Sci. 2018, 9, 113. [Google Scholar] [CrossRef]
- Singh, A.; Kumar, N.; Tomar, P.P.; Bhose, S.; Ghosh, D.K.; Roy, P.; Sharma, A.K. Characterization of a bacterioferritin comigratory protein family 1-Cys peroxiredoxin from Candidatus Liberibacter asiaticus. Protoplasma 2017, 254, 1675–1691. [Google Scholar] [CrossRef] [PubMed]
- Wulff, N.A.; Zhang, S.; Setubal, J.C.; Almeida, N.F.; Martins, E.C.; Harakava, R.; Kumar, D.; Rangel, L.T.; Foissac, X.; Bové, J.M.; et al. The complete genome sequence of ‘Candidatus Liberibacter americanus’, associated with citrus huanglongbing. Mol. Plant Microbe Interact. 2014, 27, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Killiny, N. Quorum sensing controls Candidatus Liberibacter asiaticus interactions with host plant and insect vector. APS Annu. Meet. 2015, 105, 72. [Google Scholar]
- Shahzad, F.M.; Kadyampakeni, D.; Vashisth, T. Effect of Growing Media pH on Performance of Huanglongbing-Affected Young Citrus Trees. Agronomy 2021, 11, 439. [Google Scholar] [CrossRef]
- Ramsey, J.S.; Chavez, J.D.; Johnson, R.; Hosseinzadeh, S.; Mahoney, J.E.; Mohr, J.P.; Robison, F.; Zhong, X.; Hall, D.G.; MacCoss, M.; et al. Protein interaction networks at the host-microbe interface in Diaphorinacitri, the insect vector of the citrus greening pathogen. Royal Soc. Open Sci. 2017, 4, 160545. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Thaochan, N.; Huang, J.; Chen, J.; Deng, X.; Zheng, Z. Genome sequence resource of ‘Candidatus Liberibacter asiaticus’ from Thailand. Plant Dis. 2020, 104, 624–626. [Google Scholar] [CrossRef]
- Zheng, Z.; Sun, X.; Deng, X.; Chen, J. Whole-genome sequence of “Candidatus Liberibacter asiaticus” from a huanglongbing-affected citrus tree in central Florida. Genome Announc. 2015, 3, e00169-15. [Google Scholar] [CrossRef]
- Chen, Y.; Li, T.; Zheng, Z.; Xu, M.; Deng, X. Draft whole-genome sequence of a “Candidatus Liberibacter asiaticus” strain from Yunnan, China. Microbiol. Res. Announc. 2019, 8, e01413-18. [Google Scholar] [CrossRef]
- Wu, F.; Zheng, Z.; Deng, X.; Cen, Y.; Liang, G.; Chen, J. Draft genome sequence of “Candidatus Liberibacter asiaticus” from Diaphorinacitri in Guangdong, China. Genome Announc. 2015, 3, e01316-15. [Google Scholar] [CrossRef]
- Cai, W.; Nunziata, S.; Costanzo, S.; Kumagai, L.; Rascoe, J.; Stulberg, M.J. Genome resource for the huanglongbing causal agent ‘Candidatus Liberibacter asiaticus’ strain AHCA17 from citrus root tissue in California, USA. Plant Dis. 2020, 104, 627–629. [Google Scholar] [CrossRef]
- Kunta, M.; Zheng, Z.; Wu, F.; da Graca, J.V.; Park, J.W.; Deng, X.; Chen, J. Draft whole-genome sequence of “Candidatus Liberibacter asiaticus” strain TX2351 isolated from Asian citrus psyllids in Texas, USA. Genome Announc. 2017, 5, e00170-17. [Google Scholar] [CrossRef] [PubMed]
- Cai, W.; Yan, Z.; Rascoe, J.; Stulberg, M.J. Draft whole-genome sequence of “Candidatus Liberibacter asiaticus” strain TX1712 from citrus in texas. Genome Announc. 2018, 6, e00554-18. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Atta, S.; Cui, X.; Zeng, C.; Chen, J.; Zhou, C.; Wang, X. Genome sequence resources of two ‘Candidatus Liberibacter asiaticus’ strains from Pakistan. Plant Dis. 2020, 104, 2048–2050. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.A.; Huang, J.; Wulff, N.A.; Zheng, Z.; Krugner, R.; Chen, J. Genome sequence resource of ‘Candidatus Liberibacter asiaticus’ strain 9PA from Brazil. Plant Dis. 2021, 105, 199–201. [Google Scholar] [CrossRef]
- Cui, X.J.; Zeng, C.H.; Liu, K.H.; Teng, C.L.; Zhou, C.Y.; Wang, X.F. Decreasing detection frequency of MITE (MCLas-A) in the population of ‘Candidatus Liberibacter asiaticus’ recently collected in southern China. J. Integr. Agric. 2020, 19, 2597–2601. [Google Scholar] [CrossRef]
- Huang, J.; Dai, Z.; Zheng, Z.; Silvia, P. Metagenomic Analyses of ACP and Citrus Samples Infected with “Candidatus Liberibacter asiaticus” in California. In American Phytopathological Society Abstracts; USDA: Washington, DC, USA, 2020. Available online: https://www.ars.usda.gov/research/publications/publication/?seqNo115=373141 (accessed on 25 November 2022).
- Jain, M.; Fleites, L.; Gabriel, D.W. Prophage encoded peroxidase in “Candidatus Liberibacter asiaticus” is a secreted effector that suppresses plant defenses. Mol. Plant-Microbe Interac. 2015, 28, 1330–1337. [Google Scholar] [CrossRef]
- Dominguez-Mirazo, M.; Jin, R.; Weitz, J.S. Functional and comparative genomic analysis of integrated prophage-like sequences in “Candidatus Liberibacter asiaticus”. mSphere 2019, 4, e00409-19. [Google Scholar] [CrossRef]
- Zhang, S.; Flores-Cruz, Z.; Zhou, L.; Kang, B.H.; Fleites, L.A.; Gooch, M.D.; Wulff, N.A.; Davis, M.J.; Duan, Y.P.; Gabriel, D.W. ‘Ca. Liberibacter asiaticus’ carries an excision plasmid prophage and a chromosomally integrated prophage that becomes lytic in plant infections. Mol. Plant-Microbe Interact. 2011, 24, 458–468. [Google Scholar] [CrossRef]
- Fu, S.M.; Hartung, J.; Zhou, C.Y.; Su, H.N.; Tan, J.; Li, Z.A. Ultrastructural changes and putative phage particles observed in sweet orange leaves infected with ‘Candidatus Liberibacter asiaticus’. Plant Dis. 2015, 99, 320–324. [Google Scholar] [CrossRef]
- Zheng, Z.; Bao, M.; Wu, F.; Chen, J.; Deng, X. Predominance of single prophage carrying a CRISPR/cas system in “Candidatus Liberibacter asiaticus” strains in Southern China. PLoS ONE 2016, 11, e0146422. [Google Scholar] [CrossRef]
- Castro, M.S.; Fontes, W. Plant defense and antimicrobial peptides. Protein Pept. Lett. 2005, 12, 11–16. [Google Scholar] [CrossRef]
- Casteel, C.L.; Hansen, A.K.; Walling, L.L.; Paine, T.D. Manipulation of plant defense responses by the Tomato psyllid (Bactericercacockerelli) and its associated endosymbiont Candidatus Liberibacterpsyllaurous. PLoS ONE 2012, 7, e35191. [Google Scholar] [CrossRef]
- Li, J.; Pang, Z.; Trivedi, P.; Zhou, X.; Ying, X.; Jia, H.; Wang, N. “Candidatus Liberibacter asiaticus” encodes a functional salicylic acid (SA) hydroxylase that degrades SA to suppress plant defenses. Mol. Plant-Microbe Interac. 2017, 30, 620–630. [Google Scholar] [CrossRef]
- Clark, K.; Franco, J.Y.; Schwizer, S.; Pang, Z.Q.; Hawara, E.; Thomas, W.H.; Pagliaccia, D.; Zeng, L.; Gurung, F.B.; Wang, P.; et al. An effector from the Huanglongbing-associated pathogen targets citrus proteases. Nat. Commun. 2018, 9, 1718. [Google Scholar] [CrossRef] [PubMed]
- Ying, X.; Wan, M.; Hu, L.; Zhang, J.; Li, H.; Dianqiu, L. Identification of the virulence factors of Candidatus Liberibacter asiaticus via heterologous expression in Nicotiana benthamiana using Tobacco mosaic virus. Int. J. Mol. Sci. 2019, 20, 5575. [Google Scholar] [CrossRef]
- Ghosh, D.K.; Kokane, A.D.; Kokane, S.B.; Mukherjee, K.; Tenzin, J.; Surwase, D.; Deshmukh, D.; Gubyad, M.; Biswas, K.K. A comprehensive analysis of Citrus tristeza variants of Bhutan and across the World. Front. Microbiol. 2022, 13, 797463. [Google Scholar] [CrossRef] [PubMed]
- Lafléche, D.; Bové, J.M. Structures de type mycoplasme dans les feuillesdd’orangers attaints de l maladie du greening. CR. Acad. Sci. Ser. D 1970, 270, 1915–1917. [Google Scholar]
- Cevallos-Cevallos, J.M.; Reyes-De-Corcuera, J.I.; Etxeberria, E.; Danyluk, M.D.; Rodrick, G.E. Metabolomic analysis in food science: A review. Trends Food Sci. Technol. 2009, 20, 557–566. [Google Scholar] [CrossRef]
- Motghare, M.; Kokane, A.; Warghane, A.; Kokane, S.B.; Sharma, A.K.; Ghosh, D.K. Quantitative distribution of Citrus yellow mosaic badnavirus in sweet orange (Citrus sinensis) and its implication in developing disease diagnostics. J. Virol. Meth. 2018, 259, 25–31. [Google Scholar] [CrossRef]
- Warghane, A.; Kokane, D.; Kokane, S.B.; Motghare, M.; Surwase, D.; Biswas, K.; Ghosh, D.K. Molecular detection and coat protein gene based characterization of Citrus tristeza virus prevalent in Sikkim state of India. Indian Phytopathol. 2020, 73, 135–143. [Google Scholar] [CrossRef]
- Kokane, A.D.; Lawrence, K.; Kokane, S.B.; Gubyad, M.G.; Misra, P.; Reddy, M.K.; Ghosh, D.K. Development of a SYBR green-based RT-qPCR assay for the detection of Indian citrus ringspot virus. 3Biotech 2021, 11, 359. [Google Scholar] [CrossRef]
- Li, W.; Hartung, J.S.; Levy, L. Quantitative real-time PCR for detection and identification of Candidatus Liberibacter species associated with citrus huanglongbing. J. Microbiol. Methods 2006, 66, 104–115. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Levy, L.; Hartung, J.S. Quantitative distribution of ‘Candidatus Liberibacter asiaticus’ in citrus plants with citrus Huanglongbing. Phytopathology 2009, 99, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, D.K.; Kokane, S.; Kumar, P.; Ozcan, A.; Warghane, A.; Motghare, M.; Santra, S.; Sharma, A.K. Antimicrobial nano-zinc oxide-2S albumin protein formulation significantly inhibits growth of “Candidatus Liberibacter asiaticus” in planta. PLoS ONE 2018, 13, e0204702. [Google Scholar] [CrossRef] [PubMed]
- Gopal, K.; Sudarsan, S.; Gopi, V.; Naidu, L.N.; Ramaiah, M.; Sreenivasulu, Y.; Wesley, E. Detection of huanglongbing (Citrus greening) disease by nucleic acid spot hybridization. Z. Für Nat. C 2009, 64, 711–716. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Albrecht, U.; Bowman, K.D. Gene expression in Citrus sinensis (L.) Osbeck following infection with the bacterial pathogen Candidatus Liberibacter asiaticus causing Huangloangbing in Florida. Plant Sci. 2008, 175, 291–306. [Google Scholar] [CrossRef]
- Ghosh, D.K.; Kokane, S.B.; Gowda, S. Development of a reverse transcription recombinase polymerase based isothermal amplification coupled with lateral flow immune chromatographic assay (CTV-RT-RPA-LFICA) for rapid detection of Citrus tristeza virus. Sci. Rep. 2020, 10, 20593. [Google Scholar] [CrossRef]
- Kokane, A.; Kokane, S.B.; Warghane, A.J.; Gubyad, M.G.; Sharma, A.K.; Reddy, M.K.; Ghosh, D.K. A rapid and sensitive reverse transcription-loop-mediated isothermal amplification (RT-LAMP) assay for the detection of Indian citrus ringspot virus. Plant Dis. 2021, 105, 1346–1355. [Google Scholar] [CrossRef]
- Rigano, L.A.; Malamud, F.; Orce, I.G.; Filippone, M.P.; Marano, M.R.; do Amaral, A.M.; Castagnaro, A.P.; Vojnov, A.A. Rapid and sensitive detection of Candidatus Liberibacter asiaticus by loop mediated isothermal amplification combined with a lateral flow dipstick. BMC Microbiol. 2014, 14, 86. [Google Scholar] [CrossRef]
- Ghosh, D.K.; Bhose, S.; Warghane, A.; Motghare, M.; Sharma, A.K.; Dhar, A.K.; Gowda, S. Loop-mediated isothermal amplification (LAMP) based method for rapid and sensitive detection of ‘Candidatus Liberibacter asiaticus’ in citrus and the psyllid vector, Diaphorinacitri Kuwayama. J. Plant Biochem. Biotechol. 2016, 25, 219–223. [Google Scholar] [CrossRef]
- Qian, W.; Lu, Y.; Meng, Y.; Ye, Z.; Wang, L.; Wang, R.; Zheng, Q.; Wu, H.; Wu, J. Field detection of citrus Huanglongbing associated with ‘Candidatus Liberibacter asiaticus’ by recombinase polymerase amplification within 15 min. J. Agric. Food Chem. 2018, 66, 5473–5480. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, D.K.; Kokane, S.B.; Kokane, A.D.; Warghane, A.J.; Motghare, M.R.; Bhose, S.; Sharma, A.K.; Reddy, M.K. Development of a recombinase polymerase based isothermal amplification combined with lateral flow assay (HLB-RPA-LFA) for rapid detection of “Candidatus Liberibacter asiaticus”. PLoS ONE 2018, 13, e0208530. [Google Scholar] [CrossRef] [PubMed]
- Folimonova, S.Y.; Achor, D.S. Early events of Citrus greening (huanglongbing) disease development at the ultrastructural level. Phytopathology 2010, 100, 949–958. [Google Scholar] [CrossRef] [PubMed]
- Sankaran, S.; Mishra, A.; Ehsani, R.; Davis, C. A review of advanced techniques for detecting plant diseases. Comput. Electron. Agric. 2010, 72, 1–13. [Google Scholar] [CrossRef]
- Lipchock, S.V.; Reed, D.R.; Mennella, J.A. The gustatory and olfactory systems during infancy: Implications for development of feeding behaviors in the high-risk neonate. Clin. Perinatol. 2011, 38, 627–641. [Google Scholar] [CrossRef]
- Gottwald, T.; Poole, G.; McCollum, T.; Hall, D.; Hartung, J.; Bai, J.; Luo, W.; Posny, D.; Duan, Y.P.; Taylor, E.; et al. Canine olfactory detection of a vectored phytobacterial pathogen, Liberibacter asiaticus, and integration with disease control. Proc. Natl. Acad. Sci. USA 2020, 117, 3492–3501. [Google Scholar] [CrossRef]
- Fujiwara, K.; Iwanami, T.; Fujikawa, T. Alterations of Candidatus Liberibacter asiaticus-associated microbiota decrease survival of Ca. L. asiaticus in vitro assays. Front. Microbiol. 2018, 9, 3089. [Google Scholar] [CrossRef]
- Zhang, M.; Guo, Y.; Powell, C.A.; Doud, M.S.; Yang, C.; Duan, Y. Effective antibiotics against “Candidatus Liberibacter asiaticus” in HLB-affected citrus plants identified via the graft-based evaluation. PLoS ONE 2014, 9, e111032. [Google Scholar] [CrossRef]
- Li, W.; Cong, Q.; Pei, J.; Kinch, L.N.; Grishin, N.V. The ABC transporters in Candidatus Liberibacter asiaticus. Proteins 2012, 80, 2614–2628. [Google Scholar] [CrossRef]
- Vahling-Armstrong, C.M.; Zhou, H.; Benyon, L.; Morgan, J.K.; Duan, Y.P. Two plant bacteria, S. meliloti and Ca. Liberibacter asiaticus, share functional znuABC homologues that encode for a high affinity zinc uptake system. PLoS ONE 2012, 7, e37340. [Google Scholar] [CrossRef]
- Kokane, S.B.; Pathak, R.K.; Singh, M.; Kumar, A. The role of tripartite interaction of calcium sensors and transporters in the accumulation of calcium in finger millet grain. Biol. Plant. 2018, 62, 325–334. [Google Scholar] [CrossRef]
- Jones, P.M.; George, A.M. The ABC transporter structure and mechanism: Perspectives on recent research. CMLS Cell. Mol. Life Sci. 2004, 61, 682–699. [Google Scholar] [CrossRef] [PubMed]
- Locher, K.P. Structure and mechanism of ATP-binding cassette transporters. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2009, 364, 239–245. [Google Scholar] [CrossRef] [PubMed]
- Quiocho, F.A.; Phillips, G.N., Jr.; Parsons, R.G.; Hogg, R.W. Crystallographic data of an L-arabinose-binding protein from Escherichia coli. J. Mol. Biol. 1974, 86, 491–493. [Google Scholar] [CrossRef]
- Sun, Y.J.; Rose, J.; Wang, B.C.; Hsiao, C.D. The structure of glutamine-binding protein complexed with glutamine at 1.94 Å resolution: Comparisons with other amino acid binding proteins. J. Mol. Biol. 1998, 278, 219–229. [Google Scholar] [CrossRef]
- Tang, C.; Schwieters, C.D.; Clore, G.M. Open-to-closed transition in apo maltose-binding protein observed by paramagnetic NMR. Nature 2007, 449, 1078–1082. [Google Scholar] [CrossRef]
- Millet, O.; Hudson, R.P.; Kay, L.E. The energetic cost of domain reorientation in maltose-binding protein as studied by NMR and fluorescence spectroscopy. Proc. Natl. Acad. Sci. USA 2003, 100, 12700–12705. [Google Scholar] [CrossRef]
- Fukamizo, T.; Kitaoku, Y.; Suginta, W. Periplasmic solute-binding proteins: Structure classification and chitooligosaccharide recognition. Int. J. Biol. Macromol. 2019, 128, 985–993. [Google Scholar] [CrossRef]
- de Vries, S.P.; Bootsma, H.J.; Hays, J.P.; Hermans, P.W. Molecular aspects of Moraxella catarrhalis pathogenesis. Microbiol. Mol. Biol. Rev. 2009, 73, 389–406. [Google Scholar] [CrossRef] [PubMed]
- Xayarath, B.; Marquis, H.; Port, G.C.; Freitag, N.E. Listeria monocytogenes CtaP is a multifunctional cysteine transport-associated protein required for bacterial pathogenesis. Mol. Microbiol. 2009, 74, 956–973. [Google Scholar] [CrossRef] [PubMed]
- Tuinema, B.R.; Reid-Yu, S.A.; Coombes, B.K. Salmonella evades D-amino acid oxidase to promote infection in neutrophils. MBio 2014, 5, e01886-14. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.J.; Song, S.; Mason, K.; Pinkett, H.W. Selective substrate uptake: The role of ATP-binding cassette (ABC) importers in pathogenesis. Biochim. Biophy. Acta -Biomemb. 2018, 1860, 868–877. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Sreedharan, A.; Wei, S.; Wang, J.; Pelz-Stelinski, K.; Folimonova, S.; Wang, N. Global gene expression changes in ‘Candidatus Liberibacter asiaticus’ during the transmission in distinct hosts between plant and insect. Mol. Plant Pathol. 2013, 14, 391–404. [Google Scholar] [CrossRef] [PubMed]
- Gabbianelli, R.; Scotti, R.; Ammendola, S.; Petrarca, P.; Nicolini, L.; Battistoni, A. Role of ZnuABC and ZinT in Escherichia coli O157: H7 zinc acquisition and interaction with epithelial cells. BMC Microb. 2011, 11, 1–11. [Google Scholar] [CrossRef]
- Vahedi-Faridi, A.; Eckey, V.; Scheffel, F.; Alings, C.; Landmesser, H.; Schneider, E.; Saenger, W. Crystal structures and mutational analysis of the arginine-, lysine-, histidine-binding protein ArtJ from Geobacillus stearothermophilus. Implications for interactions of ArtJ with its cognate ATP-binding cassette transporter, Art (MP)2. J. Mol. Biol. 2008, 375, 448–459. [Google Scholar] [CrossRef]
- Bhattacharyya, N.; Nkumama, I.N.; Newland-Smith, Z.; Lin, L.Y.; Yin, W.; Cullen, R.E.; Griffiths, J.S.; Jarvis, A.R.; Price, M.J.; Chong, P.Y.; et al. An aspartate-specific solute-binding protein regulates protein kinase G activity to control glutamate metabolism in mycobacteria. MBio 2018, 9, e00931-18. [Google Scholar] [CrossRef]
- Hu, Y.; Fan, C.P.; Fu, G.; Zhu, D.; Jin, Q.; Wang, D.-C. Crystal structure of a glutamate/aspartate binding protein complexed with a glutamate molecule: Structural basis of ligand specificity at atomic resolution. J. Mol. Biol. 2008, 382, 99–111. [Google Scholar] [CrossRef]
- Fulyani, F.; Schuurman-Wolters, G.K.; Žagar, A.V.; Guskov, A.; Slotboom, D.J.; Poolman, B. Functional diversity of tandem substrate-binding domains in ABC transporters from pathogenic bacteria. Structure 2013, 21, 1879–1888. [Google Scholar] [CrossRef]
- Müller, A.; Thomas, G.H.; Horler, R.; Brannigan, J.A.; Blagova, E.; Levdikov, V.M.; Fogg, M.J.; Wilson, K.S.; Wilkinson, A.J. An ATP-binding cassette-type cysteine transporter in Campylobacter jejuni inferred from the structure of an extracytoplasmic solute receptor protein. Mol. Microbiol. 2005, 57, 143–155. [Google Scholar] [CrossRef]
- Yao, N.; Trakhanov, S.; Quiocho, F.A. Refined 1.89-ANG. A structure of the histidine-binding protein complexed with histidine and its relationship with many other active transport/chemosensory proteins. Biochemistry 1994, 33, 4769–4779. [Google Scholar] [CrossRef]
- Oh, B.H.; Kang, C.H.; De Bondt, H.; Kim, S.H.; Nikaido, K.; Joshi, A.K.; Ames, G.F. The bacterial periplasmic histidine-binding protein. structure/function analysis of the ligand-binding site and comparison with related proteins. J. Biol. Chem. 1994, 269, 4135–4143. [Google Scholar] [CrossRef] [PubMed]
- Bulut, H.; Moniot, S.; Licht, A.; Scheffel, F.; Gathmann, S.; Saenger, W.; Schneider, E. Crystal structures of two solute receptors for L-cystine, respectively, of the human pathogen Neisseria gonorrhoeae. J. Mol. Bio. 2012, 415, 560–572. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Kesari, P.; Kokane, S.; Ghosh, D.K.; Kumar, P.; Sharma, A.K. Crystal structures of a putative periplasmic cystine-binding protein from Candidatus Liberibacter asiaticus: Insights into an adapted mechanism of ligand binding. FEBS J. 2019, 286, 3450–3472. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Dalal, V.; Kokane, A.; Singh, S.; Lonare, S.; Kaur, H.; Ghosh, D.K.; Kumar, P.; Sharma, A.K. Mutation studies and structure-based identification of potential inhibitor molecules against periplasmic amino acid binding protein of Candidatus Liberibacter asiaticus (CLasTcyA). Int. J. Biol. Macromol. 2020, 147, 1228–1238. [Google Scholar] [CrossRef]
- Sharma, N.; Selvakumar, P.; Bhose, S.; Ghosh, D.K.; Kumar, P.; Sharma, A.K. Crystal structure of a periplasmic solute binding protein in metal-free, intermediate and metal-bound states from Candidatus Liberibacter asiaticus. J. Struct. Biol. 2015, 189, 184–194. [Google Scholar] [CrossRef]
- Sharma, N.; Selvakumar, P.; Saini, G.; Warghane, A.; Ghosh, D.K.; Sharma, A.K. Crystal structure analysis in Zn2+ -bound state and biophysical characterization of CLas-ZnuA2. Biochim. Biophys. Acta- Prot. Proteom. 2016, 1864, 1649–1657. [Google Scholar] [CrossRef] [PubMed]
- Saini, G.; Sharma, N.; Dalal, V.; Warghane, A.; Ghosh, D.K.; Kumar, P.; Sharma, A.K. The analysis of subtle internal communications through mutation studies in periplasmic metal uptake protein CLas-ZnuA2. J. Struct. Biol. 2018, 204, 228–239. [Google Scholar] [CrossRef]
- Magnusson, U.; Salopek-Sondi, B.; Luck, L.A.; Mowbray, S.L. X-ray structures of the leucine-binding protein illustrate conformational changes and the basis of ligand specificity. J. Biol. Chem. 2004, 279, 8747–8752. [Google Scholar] [CrossRef]
- Osborne, A.R.; Rapoport, T.A.; van den Berg, B. Protein translocation by the Sec61/SecY channel. Annu. Rev. Cell Dev. Biol. 2005, 21, 43209–43217. [Google Scholar] [CrossRef]
- Vrontou, E.; Economou, A. Structure and function of SecA, the preprotein translocase nanomotor. Biochim. Biophys. Acta 2004, 1694, 67–80. [Google Scholar] [CrossRef]
- Driessen, A.J.; Nouwen, N. Protein translocation across the bacterial cytoplasmic membrane. Annu. Rev. Biochem. 2008, 77, 643–667. [Google Scholar] [CrossRef] [PubMed]
- Natale, P.; Brüser, T.; Driessen, A.J. Sec-and Tat-mediated protein secretion across the bacterial cytoplasmic membrane-distinct translocases and mechanisms. Biochim. Biophys. Acta Biomembranes 2008, 1778, 1735–1756. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.G.; Rollo, E.E.; Grodberg, J.; Oliver, D.B. Nucleotide sequence of the secA gene and secA (Ts) mutations preventing protein export in Escherichia coli. J. Bacterial. 1988, 170, 3404–3414. [Google Scholar] [CrossRef] [PubMed]
- Papanikolau, Y.; Papadovasilaki, M.; Ravelli, R.B.; McCarthy, A.A.; Cusack, S.; Economou, A.; Petratos, K. Structure of dimeric SecA, the Escherichia coli preprotein translocase motor. J. Mol. Biol. 2007, 366, 1545–1557. [Google Scholar] [CrossRef] [PubMed]
- Akula, N.; Zheng, H.; Han, F.Q.; Wang, N. Discovery of novel SecA inhibitors of Candidatus Liberibacter asiaticus by structure-based design. Bioorg. Med. Chem. Lett. 2011, 21, 4183–4188. [Google Scholar] [CrossRef]
- Coyle, J.F.; Lorca, G.L.; Gonzalez, C.F. Understanding the physiology of Liberibacter asiaticus: An overview of the demonstrated molecular mechanisms. J. Mol. Microbiol. Biotechnol. 2018, 28, 116–127. [Google Scholar] [CrossRef]
- Gardner, C.L.; Pagliai, F.A.; Pan, L.; Bojilova, L.; Torino, M.I.; Lorca, G.L.; Gonzalez, C.F. Drug repurposing: Tolfenamic acid inactivates PrbP, a transcriptional accessory protein in Liberibacter asiaticus. Front. Microbiol. 2016, 7, 1630. [Google Scholar] [CrossRef]
- Pagliai, F.A.; Coyle, J.F.; Kapoor, S.; Gonzalez, C.F.; Lorca, G.L. LdtR is a master regulator of gene expression in Liberibacter asiaticus. Microb. Biotechnol. 2017, 10, 896–909. [Google Scholar] [CrossRef]
- Sourjik, V.; Muschler, P.; Scharf, B.; Schmitt, R. VisN and VisR are global regulators of chemotaxis, flagellar, and motility genes in Sinorhizobium (Rhizobium) meliloti. J. Bacteriol. 2000, 182, 782–788. [Google Scholar] [CrossRef]
- Charoenpanich, P.; Meyer, S.; Becker, A.; McIntosh, M. Temporal expression program of quorum sensing-based transcription regulation in Sinorhizobiummeliloti. J. Bacteriol. 2013, 195, 3224–3236. [Google Scholar] [CrossRef]
- Tang, G.; Wang, Y.; Luo, L. Transcriptional regulator LsrB of Sinorhizobiummeliloti positively regulates the expression of genes involved in lipopolysaccharide biosynthesis. Appl. Environ. Microbiol. 2014, 80, 5265–5273. [Google Scholar] [CrossRef]
- Pini, F.; De Nisco, N.J.; Ferri, L.; Penterman, J.; Fioravanti, A.; Brilli, M.; Mengoni, A.; Bazzicalupo, M.; Viollier, P.H.; Walker, G.C.; et al. Cell cycle control by the master regulator CtrA in Sinorhizobiummeliloti. PLoS Genet. 2015, 11, e1005232. [Google Scholar] [CrossRef]
- Chao, T.C.; Buhrmester, J.; Hansmeier, N.; Pühler, A.; Weidner, S. Role of the regulatory gene rirA in the transcriptional response of Sinorhizobiummeliloti to iron limitation. Appl. Environ. Microbiol. 2005, 71, 5969–5982. [Google Scholar] [CrossRef]
- Pereira, S.F.; Goss, L.; Dworkin, J. Eukaryote-like serine/threonine kinases and phosphatases in bacteria. Microbiol. Mol. Biol. Rev. 2011, 75, 192–212. [Google Scholar] [CrossRef]
- Pitzschke, A.; Schikora, A.; Hirt, H. MAPK cascade signaling networks in plant defence. Curr. Opin. Plant Biol. 2009, 12, 421–426. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsova, E.; Nocek, B.; Brown, G.; Makarova, K.S.; Flick, R.; Wolf, Y.I.; Khusnutdinova, A.; Evdokimova, E.; Jin, K.; Tan, K.; et al. Functional diversity of haloacid dehalogenase superfamily phosphatases from Saccharomyces cerevisiae biochemical, structural, and evolutionary insights. J. Biol. Chem. 2015, 290, 18678–18698. [Google Scholar] [CrossRef]
- Nam, S.E.; Kim, A.C.; Paetzel, M. Crystal structure of Bacillus subtilis signal peptide peptidase A. J. Mol. Biol. 2012, 419, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Shim, E.; Cravatt, B.; Jacobsen, R.; Schoeniger, J. Escherichia Coli signal peptide peptidase A is a serine-lysine protease with a lysine recruited to the nonconserved amino-terminal domain in the S49 protease family. Biochemistry 2008, 47, 6361–6369. [Google Scholar] [CrossRef]
- Damsteegt, V.D.; Postnikova, E.N.; Stone, A.L.; Kuhlmann, M.; Wilson, C. Murraya paniculata and related species as potential hosts and inoculum reservoirs of ‘Candidatus Liberibacter asiaticus’, causal agent of Huanglongbing. Plant Dis. 2010, 94, 528–533. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Fan, J.; Chen, C.; Yu, Q.; Brlansky, R.H.; Li, Z.G. Comparative iTRAQ proteome and transcriptome analyses of sweet orange infected by “Candidatus Liberibacter asiaticus”. Physiol. Plant. 2011, 143, 235–245. [Google Scholar] [CrossRef] [PubMed]
- Shee, C.; Islam, A.; Ahmad, F.; Sharma, A.K. Structure-function studies of Murrayakoenigii trypsin inhibitor revealed a stable core beta sheet structure surrounded by α-helices with a possible role for α-helix in inhibitory function. Int. J. Biol. Macromol. 2007, 41, 410–414. [Google Scholar] [CrossRef] [PubMed]
- Tsukuda, S.; Gomi, K.; Yamamoto, H.; Akimitsu, K. Characterization of cDNAs encoding two distinct miraculin-like proteins and stress-related modulation of the corresponding mRNAs in Citrus jambhiri Lush. Plant Mol. Biol. 2006, 60, 125–136. [Google Scholar] [CrossRef]
- Verma, P.; Kar, B.; Varshney, R.; Roy, P.; Sharma, A.K. Characterization of AICAR transformylase /IMP cyclohydrolase (ATIC) from Staphylococcus lugdunensis. FEBS J. 2017, 284, 4233–4261. [Google Scholar] [CrossRef] [PubMed]
- Wood, Z.A.; Schroder, E.; Harris, R.J.; Poole, L.B. Structure, mechanism and regulation of peroxiredoxins. Trends Biochem Sci. 2003, 28, 32–40. [Google Scholar] [CrossRef]
- Kalckar, H.M.; Shafran, M. Differential spectrophotometry of purine compounds by means of specific enzymes: I. Determination of hydroxypurine compounds. J. Biol. Chem. 1947, 167, 429–443. [Google Scholar] [CrossRef] [PubMed]
- Beardsley, G.P.; Rayl, E.A.; Gunn, K.; Moroson, B.A.; Seow, H.; Anderson, K.S.; Vergis, J.; Fleming, K.; Worland, S.; Condon, B.; et al. Structure and functional relationships in human pur H. In Purine and Pyrimidine Metabolism in Man IX. Advances in Experimental Medicine and Biology; Springer: Boston, MA, USA, 1998; Volume 43, pp. 221–226. [Google Scholar] [CrossRef]
- Verma, P.; Patel, G.K.; Kar, B.; Sharma, A.K. A case of neofunctionalization of a Putranjiva roxburghii PNP proteins to trypsin inhibitor by disruption of PNP-UDP domain through an in insert containing inhibitory site. Plant Sci. 2017, 260, 19–30. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Xu, L.; Wolan, D.W.; Wilson, I.A.; Olson, A.J. Virtual screening of human 5-aminoimidazole-4-carboxamide ribonucleotide transformylase against the NCI diversity set by use of AutoDock to identify novel nonfolate inhibitors. J. Med. Chem. 2004, 47, 6681–6690. [Google Scholar] [CrossRef]
- Maier, T.; Jenni, S.; Ban, N. Architecture of mammalian fatty acid synthase at 4.5 Å resolution. Science 2006, 311, 1258–1262. [Google Scholar] [CrossRef]
- Tasdemir, D. Type II fatty acid biosynthesis, a new approach in antimalarial natural product discovery. Phytochem. Rev. 2006, 5, 99–108. [Google Scholar] [CrossRef]
- Rafi, S.; Novichenok, P.; Kolappan, S.; Stratton, C.F.; Rawat, R.; Kisker, C.; Simmerling, C.; Tonge, P. Structure of acyl carrier protein bound to FabI, the FASII enoyl reductase from Escherichia coli. J. Biol. Chem. 2006, 281, 39285–39293. [Google Scholar] [CrossRef]
- Jiang, L.; Gao, Z.; Li, Y.; Wang, S.; Dong, Y. Crystal structures and kinetic properties of enoyl-acyl carrier protein reductase I from Candidatus Liberibacter asiaticus. Protein Sci. 2014, 23, 366–377. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; GAO, Z.; LI, Y.; Jiang, L.; Dong, Y.H. β- Hydroxyacyl-acyl carrier protein dehydratase (FabZ) from Candidatus Liberibacterasiaticum: Protein characterization and structural modeling. Austin J. Microbiol. 2015, 1, 1007. [Google Scholar]
- Gilkes, J.M.; Frampton, R.A.; Board, A.J.; Sheen, C.R.; Smith, G.R.; Dobson, R.C.J.D. Dihydrodipicolinate synthase bound with allosteric inhibitor (S)-lysine from Candidatus Liberibacter solanacearum. Nat. Struct. Biol. 2021, 10, 980. [Google Scholar] [CrossRef]
- Nan, J.; Zhang, S.; Zhan, P.; Jiang, L. Evaluation of bronopol and disulfiram as potential Candidatus Liberibacter asiaticus inosine 5′-monophosphate dehydrogenase inhibitors by using molecular docking and enzyme kinetic. Molecules 2020, 25, 2313. [Google Scholar] [CrossRef]
- Andrade, M.; Li, J.; Wang, N. Candidatus Liberibacter asiaticus: Virulence traits and control strategies. Trop. Plant Pathol. 2020, 45, 285–297. [Google Scholar] [CrossRef]
- Nariani, T.K.; Ghosh, S.K.; Kumar, D.; Raychaudhuri, S.P.; Viswanath, S.M. Detection and possibilities of therapeutic control of the greening disease of citrus caused by mycoplasma. Proc. Indian Natl. Sci. Acad. 1975, 41, 334–339. [Google Scholar]
- Shokrollah, H.; Lee, T.; Sijam, K.; Nor, S.; Abdullah, A. Potential use of selected citrus rootstocks and interstocks against HLB disease in Malaysia. Crop Prot. 2011, 30, 521–525. [Google Scholar] [CrossRef]
- Zhang, X.; Francis, M.I.; Dawson, W.O.; Graham, J.H.; Orbović, V.; Triplett, E.W.; Mou, Z. Over-expression of the Arabidopsis NPR1 gene in citrus increases resistance to citrus canker. European J. Plant Pathol. 2010, 128, 91–100. [Google Scholar] [CrossRef]
- Zhang, M.; Powell, C.A.; Guo, Y.; Doud, M.S.; Duan, Y. A graft-based chemo-therapy method for screening effective molecules and rescuing huanglongbing- affected citrus plants. Phytopathology 2012, 102, 567e574. [Google Scholar] [CrossRef]
- Blaustein, R.A.; Graciela, L.L.; Max, T. Challenges for managing Candidatus Liberibacter spp. (Huanglongbing Disease Pathogen): Current control measures and future directions. Phytopathology 2018, 108, 424–435. [Google Scholar] [CrossRef]
- Li, J.; Li, L.; Pang, Z.; Kolbasov, V.G.; Ehsani, R.; Carter, E.W.; Wang, N. Developing citrus huanglongbing (HLB) management strategies based on the severity of symptoms in HLB-endemic citrus-producing regions. Phytopathology 2019, 109, 582–592. [Google Scholar] [CrossRef] [PubMed]
- Killiny, N.; Hijaz, F.; Gonzalez-Blanco, P.; Jones, S.E.; Pierre, M.O.; Vincent, C.I. Effect of adjuvants on oxytetracycline uptake upon foliar application in citrus. Antibiotics 2020, 9, 677. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Jiang, J.; Wang, N. Control of citrus huanglongbing (HLB) via trunk injection of plant activators and antibiotics. Phytopathology 2017, 108, 186–195. [Google Scholar] [CrossRef] [PubMed]
- Al-Rimawi, F.; Hijaz, F.; Nehela, Y.; Batuman, O.; Killiny, N. Uptake, translocation, and stability of oxytetracycline and streptomycin in citrus plants. Antibiotics 2019, 8, 196. [Google Scholar] [CrossRef]
- Canales, E.; Coll, Y.; Hernández, I.; Portieles, R.; García, M.R.; Lopez, Y.; Aranguren, M.; Alonso, E.; Delgado, R.; Luis, M.; et al. “Candidatus Liberibacter asiaticus,” causal agent of citrus huanglongbing, is reduced by treatment with brassinosteroids. PLoS ONE 2016, 11, e0146223. [Google Scholar] [CrossRef]
- War, A.R.; Paulraj, M.G.; War, M.Y.; Ignacimuthu, S. Role of salicylic acid in induction of plant defense system in chickpea (Cicer arietinum L.). Plant Signal. Behav. 2011, 6, 178–179. [Google Scholar] [CrossRef]
- Tamaoki, D.; Seo, S.; Yamada, S.; Kano, A.; Miyamoto, A.; Shishido, H.; Miyoshi, S.; Taniguchi, S.; Akimitsu, K.; Gomi, K. Jasmonic acid and salicylic acid activate a common defense system in rice. Plant Signal. Behav. 2013, 8, e24260. [Google Scholar] [CrossRef]
- Li, J.; Trivedi, P.; Wang, N. Field evaluation of plant defense inducers for the control of citrus huanglongbing. Phytopathology 2016, 106, 37–46. [Google Scholar] [CrossRef]
- Alves, M.N.; Lopes, S.A.; Raiol-Junior, L.L.; Wulff, N.A.; Girardi, E.A.; Ollitrault, P.; Pena, L. Resistance to ‘Candidatus Liberibacter asiaticus,’ the huanglongbing associated bacterium, in sexually and/or graft-compatible citrus relatives. Front. Plant Sci. 2021, 11, 2166. [Google Scholar] [CrossRef]
- Huang, C.Y.; Araujo, K.; Sánchez, J.N.; Kund, G.; Trumble, J.; Roper, C.; Godfrey, K.E.; Jin, H. A stable antimicrobial peptide with dual functions of treating and preventing citrus Huanglongbing. Proc. Natl. Acad. Sci. USA 2021, 118, e2019628118. [Google Scholar] [CrossRef]
- Huang, C.Y.; Niu, D.; Kund, G.; Jones, M.; Albrecht, U.; Nguyen, L.; Bui, C.; Ramadugu, C.; Bowman, K.D.; Trumble, J.; et al. Identification of citrus immune regulators involved in defence against Huanglongbing using a new functional screening system. Plant Biotechnol. J. 2021, 19, 757–766. [Google Scholar] [CrossRef] [PubMed]
- Sekhon, B.S. Nanotechnology in agri-food production: An overview. Nanotechnol. Sci. Appl. 2014, 7, 31–53. [Google Scholar] [CrossRef] [PubMed]
- Gilbertson, L.M.; Zimmerman, J.B.; Plata, D.L.; Hutchison, J.E.; Anastas, P.T. Designing nanomaterials to maximize performance and minimize undesirable implications guided by the Principles of Green Chemistry. Chem. Soc. Rev. 2015, 44, 5758–5777. [Google Scholar] [CrossRef] [PubMed]
- Kah, M.; Navarro, D.; Kookana, R.S.; Kirby, J.K.; Santra, S.; Ozcan, A.; Kabiri, S. Impact of (nano) formulations on the distribution and wash-off of copper pesticides and fertilisers applied on citrus leaves. Environ. Chem. 2019, 16, 401–410. [Google Scholar] [CrossRef]
- Wu, H.; Nißler, R.; Morris, V.; Herrmann, N.; Hu, P.; Jeon, S.; Kruss, S.; Giraldo, J.P. Monitoring plant health with near-infrared fluorescent H2O2 nanosensors. Nano Lett. 2020, 20, 2432–2442. [Google Scholar] [CrossRef]
- Adisa, I.O.; Pullagurala, V.L.R.; Peralta-Videa, J.R.; Dimkpa, C.O.; Elmer, W.H.; Gardea-Torresdey, J.L.; White, J.C. Recent advances in nano-enabled fertilizers and pesticides: A critical review of mechanisms of action. Environ. Sci. Nano 2019, 6, 2002–2030. [Google Scholar] [CrossRef]
- Dimkpa, C.O.; Campos, M.G.N.; Fugice, J.; Glass, K.; Ozcan, A.; Huang, Z.; Singh, U.; Santra, S. Synthesis and characterization of novel dual-capped Zn–urea nanofertilizers and application in nutrient delivery in wheat. Environ Sci. Adv. 2022, 1, 47–58. [Google Scholar] [CrossRef]
- Lowry, G.V.; Avellan, A.; Gilbertson, L.M. Opportunities and challenges for nanotechnology in the agri-tech revolution. Nat. Nanotechnol. 2019, 14, 517–522. [Google Scholar] [CrossRef]
- Gilbertson, L.M.; Pourzahedi, L.; Laughton, S.; Gao, X.; Zimmerman, J.B.; Theis, T.L.; Westerhoff, P.; Lowry, G.V. Guiding the design space for nanotechnology to advance sustainable crop production. Nat. Nanotechnol. 2020, 15, 801–810. [Google Scholar] [CrossRef]
- Zhang, Y.; Yan, J.; Avellan, A.; Gao, X.; Matyjaszewski, K.; Tilton, R.D. Temperature- and pH-responsive star polymers as nanocarriers with potential for in vivo agrochemical delivery. ACS Nano 2020, 14, 10954–10965. [Google Scholar] [CrossRef]
- Strayer-Scherer, A.L.; Liao, Y.Y.; Young, M.; Ritchie, L.; Vallad, G.; Santra, S.; Clark, F.D.; jones, J.B.; Paret, M.L. Advanced copper composites against copper-tolerant Xanthomonas perforans and Tomato bacterial spot. Phytopathology 2018, 108, 196–205. [Google Scholar] [CrossRef] [PubMed]
- Dietz, K.J.; Herth, S. Plant nanotoxicology. Trends Plant Sci. 2011, 16, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Ashworth, V.; Kim, C.; Adeleye, A.S.; Rolshausen, P.; Roper, C.; White, J.; Jassby, D. Delivery, uptake, fate, and transport of engineered nanoparticles in plants: A critical review and data analysis. Environ. Sci. Nano 2019, 6, 2311–2331. [Google Scholar] [CrossRef]
- Etxeberria, E.; Gonzalez, P.; Bhattacharya, P.; Sharma, P.; Ke, P.C. Determining the size exclusion for nanoparticles in Citrus leaves. Hort Sci. 2016, 51, 732–737. [Google Scholar] [CrossRef]
- Avellan, A.; Yun, J.; Zhang, Y.; Spielman-Sun, E.; Unrine, J.M.; Thieme, J.; Lombi, E.; Bland, G.; Lowry, G.V. Nanoparticle size and coating chemistry control foliar uptake pathways, translocation and leaf-to-Rhizosphere transport in Wheat. ACS Nano 2019, 13, 5291–5305. [Google Scholar] [CrossRef] [PubMed]
- Raliya, R.; Franke, C.; Chavalmane, S.; Nair, R.; Reed, N.; Biswas, P. Quantitative understanding of nanoparticle uptake in watermelon plants. Front. Plant Sci. 2016, 7, 1288. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Ashworth, V.E.T.M.; Geitner, N.K.; Wiesner, M.R.; Ginnan, N.; Rolshausen, P.; Roper, C.; Jassby, D. Delivery, fate, and mobility of silver nanoparticles in citrus trees. ACS Nano 2020, 14, 2966–2981. [Google Scholar] [CrossRef] [PubMed]
- Elmer, W.; Ma, C.; White, J. Nanoparticles for plant disease management. Curr. Opin. Environ. Sci. Health. 2018, 6, 66–70. [Google Scholar] [CrossRef]
- Strayer-Scherer, A.; Ocsoy, I.; Tan, W.; Jones, J.; Paret, M.L. Low concentrations of a silver-based nanocomposite to manage bacterial spot of tomato in the greenhouse. Plant Dis. 2015, 100, 1460–1465. [Google Scholar] [CrossRef]
- Servin, A.; Elmer, W.; Mukherjee, A.; De La Torre-Roche, R.; Hamdi, H.; White, J.C.; Bindraban, P.; Dimpka, C. A review of the use of engineered nanomaterials to suppress plant disease and enhance crop yield. J. Nanopart. Res. 2015, 17, 92. [Google Scholar] [CrossRef]
- Stephano-Hornedo, J.L.; Torres-Gutiérrez, O.; Toledano-Magaña, Y.; Gradilla-Martínez, I.; Pestryakov, A.; Sanchez-Gonzalez, A.; Sanchez-gonzalez, A.; Garacia-Ramos, J.C.; Bogdanchikova, N. Argovit™ silver nanoparticles to fight Huanglongbing disease in Mexican limes (Citrus aurantifolia Swingle). RSC Adv. 2020, 10, 6146–6155. [Google Scholar] [CrossRef] [PubMed]
- Nair, A.; Lian, W.Y.; Gong, Z.; Valiyaveettil, S. Toxicity of silver nanoparticles in Zebrafish models. Nanotechnology 2008, 19, 255102. [Google Scholar]
- Navarro, E.; Piccapietra, F.; Wagner, B.; Marconi, F.; Kaegi, R.; Odzak, N.; Sigg, L.; Behra, R. Toxicity of silver nanoparticles to Chlamydomonas reinhardtii. Environ. Sci. Technol. 2008, 42, 8959–8964. [Google Scholar] [CrossRef] [PubMed]
- Beer, C.; Foldbjerg, R.; Hayashi, Y.; Sutherland, D.; Autrup, H. Toxicity of silver nanoparticles-Nanoparticle or silver ion? Toxicol. Lett. 2011, 208, 286–292. [Google Scholar] [CrossRef] [PubMed]
- Wells, B.; Fishel, F.M. Agricultural Pesticide Use in Florida: A Summary, 2007–2009. EDIS. 2011. Available online: http://edis.ifas.ufl.edu (accessed on 25 November 2022).
- Lamichhane, J.R.; Osdaghi, E.; Behlau, F.; Köhl, J.; Jones, J.B.; Aubertot, J.N. Thirteen decades of antimicrobial copper compounds applied in agriculture. A review. Agron. Sustain. Dev. 2018, 38, 28. [Google Scholar] [CrossRef]
- Reed, R.; Ladner, D.; Higgins, C.; Westerhoff, P.; Ranville, J. Solubility of nano-zinc oxide in environmentally and biologically important matrices. Environ. Toxicol. Chem. 2012, 31, 93–99. [Google Scholar] [CrossRef]
- Tian, S.; Lu, L.; Labavitch, J.M.; Webb, S.M.; Yang, X.; Brown, P.H.; He, Z. Spatial imaging of Zn and other elements in Huanglongbing-affected grapefruit by synchrotron-based micro X-ray fluorescence investigation. J. Exp. Bot. 2014, 65, 953–964. [Google Scholar] [CrossRef]
- Raman, N.; Dhaveethu, R.J.; Sakthivel, A. Synthesis, spectral characterization of schiff base transition metal complexes: DNA cleavage and antimicrobial activity studies. J. Chem. Sci. 2007, 119, 303–310. [Google Scholar] [CrossRef]
- Liu, S.H.; Rawal, T.B.; Soliman, M.; Lee, B.; Maxwell, T.; Rajasekaran, P.; Mendis, H.C.; Labbé, N.; Santra, S.; Tetard, L.; et al. Antimicrobial Zn-Based “TSOL” for citrus greening management: Insights from spectroscopy and molecular simulation. J. Agric. Food Chem. 2019, 67, 6970–6977. [Google Scholar] [CrossRef]
- Mendis, H.C.; Ozcan, A.; Santra, S.; De La Fuente, L. A novel Zn chelate (TSOL) that moves systemically in citrus plants inhibits growth and biofilm formation of bacterial pathogens. PLoS ONE 2019, 14, e0218900. [Google Scholar] [CrossRef]
- Smith, S.L.; Campos, M.G.N.; Ozcan, A.; Mendis, H.C.; Young, M.; Myers, E.M.; Atilola, M.; Doomra, M.; Thwin, Z.; Johnson, E.G.; et al. Multifunctional surface, subsurface, and systemic therapeutic (MS3T) formulation for the control of citrus canker. J. Agril. Food Chem. 2021, 69, 10807–10818. [Google Scholar] [CrossRef]
- Zhang, M.Q.; Guo, Y.; Powell, C.A.; Doud, M.S.; Yang, C.Y.; Zhou, H.; Duan, Y.P. Zinc treatment increases the titre of ‘Candidatus Liberibacter asiaticus’ in huanglongbing-affected citrus plants while affecting the bacterial microbiomes. J. Appl. Microbiol. 2016, 120, 1616–1628. [Google Scholar] [CrossRef] [PubMed]
- Dimkpa, C.O.; White, J.C.; Elmer, W.H.; Gardea-Torresdey, J. Nanoparticle and ionic Zn promote nutrient loading of sorghum grain under low NPK fertilization. J. Agril. Food Chem. 2017, 65, 8552–8559. [Google Scholar] [CrossRef] [PubMed]
- Faizan, M.; Faraz, A.; Yusuf, M.; Khan, S.T.; Hayat, S. Zinc oxide nanoparticle-mediated changes in photosynthetic efficiency and antioxidant system of tomato plants. Photosynthetica 2018, 56, 678–686. [Google Scholar] [CrossRef]
- Xu, J.; Luo, X.; Wang, Y.; Feng, Y. Evaluation of zinc oxide nanoparticles on lettuce (Lactuca sativa L.) growth and soil bacterial community. Envirol. Sci. Pollut. Res. 2018, 25, 6026–6035. [Google Scholar] [CrossRef] [PubMed]
- López-Moreno, M.L.; de la Rosa, G.; Cruz-Jiménez, G.; Castellano, L.; Peralta-Videa, J.R.; Gardea-Torresdey, J.L. Effect of ZnO nanoparticles on corn seedlings at different temperatures; X-ray absorption spectroscopy and ICP/OES studies. Microchem. J. 2017, 134, 54–61. [Google Scholar] [CrossRef]
- Deshpande, P.; Dapkekar, A.; Oak, M.D.; Paknikar, K.M.; Rajwade, J.M. Zinc complexed chitosan/TPP nanoparticles: A promising micronutrient nanocarrier suited for foliar application. Carbohydr. Polym. 2017, 165, 394–401. [Google Scholar] [CrossRef]
- De Lucas-Gil, E.; Del Campo, A.; Pascual, L.; Monte-Serrano, M.; Menéndez, J.; Fernández, J.; Rubio-Marcos, F. The fight against multidrug-resistant organisms: The role of ZnO crystalline defects. Mater. Sci. Eng. C 2019, 99, 575–581. [Google Scholar] [CrossRef]
- Sirelkhatim, A.; Mahmud, S.; Seeni, A.; Kaus, N.H.M.; Ann, L.C.; Bakhori, S.K.M.; Hasan, H.; Mohamad, D. Review on zinc oxide nanoparticles: Antibacterial activity and toxicity mechanism. Nano-Micro Lett. 2015, 7, 219–242. [Google Scholar] [CrossRef]
- Nair, M.G.; Nirmala, M.; Rekha, K.; Anukaliani, A. Structural, optical, photo catalytic and antibacterial activity of ZnO and Co doped ZnO nanoparticles. Mater. Lett. 2011, 65, 1797–1800. [Google Scholar] [CrossRef]
- Lakshmi Prasanna, V.; Vijayaraghavan, R. Insight into the mechanism of antibacterial activity of ZnO: Surface defects mediated reactive oxygen species even in the dark. Langmuir 2015, 31, 9155–9162. [Google Scholar] [CrossRef] [PubMed]
- Santra, S.; Berroth, M. Compositions Including a Vacancy-Engineered (VE)-Zinc Oxide Nanocomposite, Methods of Making a Composition, Method of Using a Composition. U.S. Patent 9,215,877, 22 December 2015. [Google Scholar]
- Rawal, T.B.; Ozcan, A.; Liu, S.-H.; Pingali, S.V.; Akbilgic, O.; Tetard, L.; O’Neill, H.; Santra, S.; Petridis, L. Interaction of zinc oxide nanoparticles with water: Implications for catalytic activity. ACS Appl. Nano Mater. 2019, 2, 4257–4266. [Google Scholar] [CrossRef]
- Naranjo, E.; Merfa, M.V.; Santra, S.; Ozcan, A.; Johnson, E.; Cobine, P.A.; De La Fuente, L. Zinkicide Is a ZnO-Based nanoformulation with bactericidal activity against Liberibactercrescens in batch cultures and in microfluidic chambers simulating plant vascular systems. Appl. Envirol. Microbiol. 2020, 86, e00788-720. [Google Scholar] [CrossRef]
- Soliman, M.; Lee, B.; Ozcan, A.; Rawal, T.B.; Young, M.; Mendis, H.C.; Rajasekaran, P.; Washington, T.; Pingali, S.V.; O’Neil, H.; et al. Engineered zinc oxide-based nanotherapeutics boost systemic antibacterial efficacy against phloem-restricted diseases. Environ. Sci. Nano 2022, 9, 2869–2886. [Google Scholar] [CrossRef]
- Graham, J.H.; Johnson, E.G.; Myers, M.E.; Young, M.; Rajasekaran, P.; Das, S.; Santra, S. Potential of nano-formulated zinc oxide for control of citrus canker on grapefruit trees. Plant Dis. 2016, 100, 2442–2447. [Google Scholar] [CrossRef]
- Santra, S.; Ozcan, A.; Ghosh, D.K.; Kokane, S.; Kumar, P.; Sharma, A.K. Compositions and Methods for Systemic Delivery of Cargos in Vascular Plants. U.S. Patent 11,229,608, 25 January 2022. [Google Scholar]
- Kundu, A.; Nogueira Campos, M.G.; Santra, S.; Rajaraman, S. Precision vascular delivery of agrochemicals with micromilled microneedles (µMMNs). Sci. Rep. 2019, 9, 14008. [Google Scholar] [CrossRef]
- Xin, X.; He, Z.; Hill, M.R.; Niedz, R.P.; Jiang, X.; Sumerlin, B.S. Efficiency of biodegradable and pH-responsive polysuccinimide nanoparticles (PSI-NPs) as smart nanodelivery systems in grapefruit: In vitro cellular investigation. Macromol. Biosci. 2018, 18, 1800159. [Google Scholar] [CrossRef]
- Kah, M.; Tufenkji, N.; White, J.C. Nano-enabled strategies to enhance crop nutrition and protection. Nat. Nanotechnol. 2019, 14, 532–540. [Google Scholar] [CrossRef]
- Maxwell, T.J.; Rajasekaran, P.; Young, M.; Schaff, M.; Heetai, R.; Santra, S. Non-phytotoxic zinc based nanoparticle adjuvant for improving rainfastness and sustained release of streptomycin. Environ. Nanotechnol. Monit. Manag. 2020, 14, 100355. [Google Scholar] [CrossRef]
- Miernicki, M.; Hofmann, T.; Eisenberger, I.; von der Kammer, F.; Praetorius, A. Legal and practical challenges in classifying nanomaterials according to regulatory definitions. Nat. Nanotechnol. 2019, 14, 208–216. [Google Scholar] [CrossRef]
- Rodrigues, S.; Demokritou, P.; Dokoozlian, N.; Hendren, C.O.; Karn, B.; Mauter, M.S.; Sadik, O.A.; Safarpour, M.; Unrine, J.M.; Viers, J.; et al. Nanotechnology for sustainable food production: Promising opportunities and scientific challenges. Environ. Sci. Nano 2017, 4, 767–781. [Google Scholar] [CrossRef]
- Hoffman, M.T.; Doud, M.S.; Williams, L.; Zhang, M.Q.; Ding, F.; Stover, E.; Hall, D.; Zhang, S.; Jones, L.; Gooch, M.; et al. Heat treatment eliminates “Candidatus Liberibacter asiaticus” from Infected citrus trees under controlled conditions. Phytopathology 2013, 103, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Kah, M.; Kookana, R. Emerging investigator series: Nanotechnology to develop novel agrochemicals: Critical issues to consider in the global agricultural context. Environ. Sci. Nano 2020, 7, 1867–1873. [Google Scholar] [CrossRef]
- Fan, G.C.; Xia, Y.L.; Lin, X.J.; Hu, H.Q.; Wang, X.D.; Ruan, C.Q.; Lu, L.; Liu, B. Evaluation of thermotherapy against Huanglongbing (citrus greening) in the greenhouse. J. Integr. Agric. 2016, 15, 111–119. [Google Scholar] [CrossRef]
- Yang, C.; Powell, C.A.; Duan, Y.; Shatters, R.; Fang, J.; Zhang, M. Deciphering the Bacterial Microbiome in Huanglongbing-Affected Citrus Treated with Thermotherapy and Sulfonamide Antibiotics. PLoS ONE 2016, 11, e015547. [Google Scholar] [CrossRef]
- Doud, M.M.; Wang, Y.; Hoffman, M.T.; Latza, C.L.; Luo, W.; Armstrong, C.M.; Gottwald, T.R.; Dai, L.; Luo, F.; Duan, Y. Solar thermotherapy reduces the titer of Candidatus Liberibacter asiaticus and enhances canopy growth by altering gene expression profiles in HLB-affected citrus plants. Hort. Res. 2017, 4, 17054. [Google Scholar] [CrossRef]
- Ghatrehsamani, S.; Abdulridha, J.; Balafoutis, A.; Zhang, X.; Ehsani, R.; Ampatzidis, Y. Development and evaluation of a mobile thermotherapy technology for in-field treatment of Huanglongbing (HLB) affected trees. Biosyst. Eng. 2019, 182, 1–15. [Google Scholar] [CrossRef]
- Ghatrehsamani, S.; Czarnecka, E.; Verner, F.L.; Gurley, W.B.; Ehsani, R.; Ampatzidis, Y. Evaluation of mobile heat treatment system for treating in-field HLB-affected trees by analyzing survival rate of surrogate bacteria. Agronomy 2019, 9, 540. [Google Scholar] [CrossRef]
- Wang, Q.; Valkonen, J.P. Cryotherapy of shoot tips: Novel pathogen eradication method. Trends Plant Sci. 2009, 14, 119–122. [Google Scholar] [CrossRef]
- Wang, Q.C.; Panis, B.; Engelmann, F.; Lambardi, M.; Valkonen, J.P.T. Cryotherapy of shoot tips: A technique for pathogen eradication to produce healthy planting materials and prepare healthy plant genetic resources for cryopreservation. Ann. Appl. Biol. 2009, 154, 351–363. [Google Scholar] [CrossRef]
- Ding, F.; Jin, S.; Hong, N.; Zhong, Y.; Cao, Q.; Yi, G.; Wang, G. Vitrification-cryopreservation, an efficient method for eliminating Candidatus Liberibacter asiaticus, the citrus Huanglongbing pathogen, from in vitro adult shoot tips. Plant Cell Rep. 2008, 27, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, F.; Tsuda, K. Understanding the plant immune system. Mol. Plant-Microbe Interact. 2010, 23, 1531–1536. [Google Scholar] [CrossRef]
- Hou, S.; Liu, Z.; Shen, H.; Wu, D. Damage-associated molecular pattern-triggered immunity in plants. Front. Plant Sci. 2019, 10, 646. [Google Scholar] [CrossRef]
- da Graça, J.V.; Douhan, G.W.; Halbert, S.E.; Keremane, M.L.; Lee, R.F.; Vidalakis, G.; Zhao, H. Huanglongbing: An overview of a complex pathosystem ravaging the world’s citrus. J. Integr. Plant. Biol. 2016, 58, 373–387. [Google Scholar] [CrossRef] [PubMed]
- Orozco-Cardenas, M.; Ryan, C.A. Hydrogen peroxide is generated systemically in plant leaves by wounding and systemin via the octadecanoid pathway. Proc. Natl. Acad. Sci. USA 1999, 96, 6553–6557. [Google Scholar] [CrossRef] [PubMed]
- Martinelli, F.; Uratsu, S.L.; Albrecht, U.; Reagan, R.L.; Phu, M.L.; Britton, M.; Buffalo, V.; Fass, J.; Leicht, E.; Zhao, W.; et al. Transcriptome profiling of citrus fruit response to huanglongbing disease. PLoS ONE 2012, 7, e38039. [Google Scholar] [CrossRef]
- Halbert, S.E.; Manjunath, K.L. Asian citrus psyllids (Sternorrhyncha: Psyllidae) and greening disease of citrus: A literature review and assessment of risk in Florida. Florida Entomol. 2004, 87, 330–353. [Google Scholar] [CrossRef]
- Albrecht, U.; Bowman, K.D. Plant Science transcriptional response of susceptible and tolerant citrus to infection with Candidatus Liberibacter asiaticus. Plant Sci. 2012, 185, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Zou, X.; Jiang, X.; Xu, L.; Lei, T.; Peng, A.; He, Y.; Yao, L.; Chen, S. Transgenic citrus expressing synthesized cecropin B genes in the phloem exhibits decreased susceptibility to Huanglongbing. Plant. Mol. Biol. 2017, 93, 341–353. [Google Scholar] [CrossRef]
- Grosser, J.W.; Dutt, M.; Omar, A.; Orbovic, V.; Barthe, G. Progress towards the development of transgenic disease resistance in citrus. II Int. Sympo. Citrus Biotechnol. 2009, 892, 101–107. [Google Scholar] [CrossRef]
- Gurr, S.J.; Rushton, P.J. Engineering plants with increased disease resistance: What are we going to express. Trends Biotechnol. 2005, 23, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Mirkov, T.E. Genetic Transformation of Citrus with Spinach Defensin for Broad Spectrum Resistance to Bacteria and Fungi. In Presentation, “Breeding for HLB Resistance”, Citrus Show Fla. 2010. Available online: http://stlucie.ifas.ufl.edu/pdfs/citrus/201 (accessed on 25 November 2022).
- Dutt, M.; Barthe, G.; Irey, M.; Grosser, J. Transgenic citrus expressing an Arabidopsis NPR1 gene exhibit enhanced resistance against Huanglongbing (HLB, Citrus greening). PLoS ONE 2015, 10, e0137134. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, L.; Yu, X.; Stover, E.; Luo, F.; Duan, Y. Transcriptome profiling of huanglongbing (HLB) tolerant and susceptible Citrus plants reveals the role of basal resistance in HLB tolerance. Front. Plant Sci. 2016, 7, 933. [Google Scholar] [CrossRef] [PubMed]
- Ramadugu, C.; Keremane, M.L.; Halbert, S.E.; Duan, Y.P.; Roose, M.L.; Stover, E.; Lee, F. Long-term field evaluation reveals huanglongbing resistance in Citrus relatives. Plant Dis. 2016, 100, 1858–1869. [Google Scholar] [CrossRef]
- Shi, Q.; Febres, V.J.; Jones, J.B.; Moore, G.A. A survey of FLS2 genes from multiple citrus species identifies candidates for enhancing disease resistance to Xanthomonas citri ssp. citri. Hortic. Res. 2016, 3, 16022. [Google Scholar] [CrossRef]
- Soares, J.M.; Tanwir, S.E.; Grosser, J.W.; Dutt, M. Development of genetically modified citrus plants for the control of citrus canker and huanglongbing. Trop. Plant Pathol. 2020, 45, 237–250. [Google Scholar] [CrossRef]
- Xie, K.; Yang, Y. RNA-guided genome editing in plants using a CRISPR-Cas system. Mol. Plant. 2013, 6, 1975–1983. [Google Scholar] [CrossRef]
- Nekrasov, V.; Staskawicz, B.; Weigel, D.; Jones, J.D.G.; Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 691–693. [Google Scholar] [CrossRef]
- Jia, H.; Zhang, Y.; Orbovi’c, V.; Xu, J.; White, F.F.; Jones, J.B.; Wang, N. Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnol. J. 2016, 15, 817–823. [Google Scholar] [CrossRef]
- Peng, A.; Chen, S.; Lei, T.; Xu, L.; He, Y.; Wu, L.; Yao, L.; Zou, X. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene Cs LOB 1 promoter in citrus. Plant Biotechnol. J. 2017, 15, 1509–1519. [Google Scholar] [CrossRef]
- Hammond, J.; Hsu, H.T.; Huang, Q.; Jordan, R.; Kamo, K.; Pooler, M. Transgenic approaches to disease resistance in ornamental crops. J. Crop Improv. 2006, 17, 155–210. [Google Scholar] [CrossRef]
- Müntz, K. Deposition of storage proteins. Plant Mol. Biol. 1998, 38, 77–99. [Google Scholar] [CrossRef] [PubMed]
- Ryan, C.A. Proteinase inhibitors in plant leaves: A biochemical model for pest-induced natural plant protection. Trends Biochem. Sci. 1978, 3, 148–150. [Google Scholar] [CrossRef]
- da Rocha Tavano, E.C.; Vieira, M.L.C.; de A. Alves-Mourao, F.; Harakava, R.; Mendes, B.M.J. Genetic transformation of Citrus sinensis ‘Hamlin’ with attacin A driven by a phloem tissue-specific promoter for resistance to Candidatusliberibacter spp. Acta Horti. 2015, 1065, 695–701. [Google Scholar] [CrossRef]
- Rawat, N.; Kumar, B.; Albrecht, U.; Du, D.; Huang, M.; Yu, Q.; Zhang, Y.; Duan, Y.; Bowman, K.D.; Gmitter, F.G.; et al. Genome resequencing and transcriptome profiling reveal structural diversity and expression patterns of constitutive disease resistance genes in Huanglongbing-tolerant Poncirus trifoliata and its hybrids. Hortic. Res. 2017, 4, 17064. [Google Scholar] [CrossRef] [PubMed]
- Tavano, E.C.; Erpen, L.; Aluisi, B.; Harakava, R.; Lopes, J.R.; Vieira, M.L.; De Stefano Piedade, S.M.; Mendes, B.M.; Filho, F. Sweet orange genetic transformation with the attacin A gene under the control of phloem-specific promoters and inoculation with Candidatus Liberibacter asiaticus. J. Hortic. Sci. Biotechnol. 2019, 94, 210–219. [Google Scholar] [CrossRef]
- Chéreau, D.; Videcoq, P.; Ruffieux, C.; Pichon, L.; Motte, j.; Belaid, S.; Ventureira, J.; Lopez, M. Combination of existing and alternative technologies to promote oilseeds and pulses proteins in food applications. OCL 2016, 23, D406. [Google Scholar] [CrossRef]
- Youle, R.J.; Huang, A.H. Occurrence of low molecular weight and high cysteine containing albumin storage proteins in oilseeds of diverse species. Am. J. Bot. 1981, 68, 44–48. [Google Scholar] [CrossRef]
- Neumann, G.M.; Condron, R.; Polya, G.M. Purification and sequencing of yellow mustard seed napin small and large chains that are phosphorylated by plant calcium-dependent protein kinase and are calmodulin antagonists. Plant Sci. 1996, 119, 49–66. [Google Scholar] [CrossRef]
- Svendsen, I.B.; Nicolova, D.; Goshev, I.; Genov, N. Isolation and characterization of a trypsin inhibitor from the seeds of kohlrabi (Brassica napus var. rapifera) belonging to the napin family of storage proteins. Carlsberg Res. Commun. 1989, 54, 231. [Google Scholar] [CrossRef]
- Svendsen, I.B.; Nicolova, D.; Goshev, I.; Genov, N. Primary structure, spectroscopic and inhibitory properties of a two-chain trypsin inhibitor from the seeds of charlock (Sinapis arvensis L), a member of the napin protein family. Int. J. Pept. Protein Res. 1994, 43, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Genov, N.; Goshev, I.; Nikolova, D.; Georgieva, D.N.; Filippi, B.; Svendsen, I. A novel thermostable inhibitor of trypsin and subtilisin from the seeds of Brassica nigra: Amino acid sequence, inhibitory and spectroscopic properties and thermostability. Biochim. Biophys. Acta-ProtStruc. Mol. Enzymol. 1997, 1341, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Ullah, A.; Mariutti, R.B.; Masood, R.; Caruso, I.P.; Costa, G.H.G.; de Freita, C.M.; Santos, C.R.; Zanphorlin, L.M.; Mutton, M.J.; Murrakami, M.T.; et al. Crystal structure of mature 2S albumin from Moringa oleifera seeds. Biochem. Biophys. Res. Commun. 2015, 468, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Kumar, P.; Kesari, P.; Neetu; Neetu Katiki, M.; Mishra, M.; Singh, P.K.; Gurjar, B.R.; Sharma, A.K.; Tomar, S.; et al. Purification and Characterization of 2S albumin from seeds of Wrightia tinctoria exhibiting Antibacterial and DNase Activity. Protein Pept. Lett. 2017, 24, 368–378. [Google Scholar] [CrossRef]
- Tomar, P.P.S.; Chaudhary, N.S.; Mishra, P.; Gahloth, D.; Patel, G.K.; Sharma, A.K. Purification, characterization and cloning of a 2S albumin with DNase, RNase and antifungal activities from Putranjiva roxburghii. App. Biochem. Biotechnol. 2014, 174, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Tomar, P.P.S.; Nikhil, K.; Singh, A.; Selvakumar, P.; Roy, P.; Sharma, A.K. Characterization of anticancer, DNase and antifungal activity of pumpkin 2S albumin. Biochem. Biophys. Res. Commun. 2014, 448, 349–354. [Google Scholar] [CrossRef]
- Theerasilp, S.; Kurihara, Y. Complete purification and characterization of the taste-modifying protein, miraculin, from miracle fruit. J. Biol. Chem. 1988, 263, 11536–11539. [Google Scholar] [CrossRef]
- Selvakumar, P.; Gahloth, D.; Tomar, P.P.S.; Sharma, N.; Sharma, A.K. Molecular evolution of miraculin-like proteins in soybean kunitz super-family. J. Mol. Evol. 2011, 73, 369–379. [Google Scholar] [CrossRef]
- Takai, A.; Satoh, M.; Matsuyama, T.; Ito, A.; Nakata, R.; Aoyama, T.; Inoue, H. Secretion of miraculin through the function of a signal peptide conserved in the Kunitz-type soybean trypsin inhibitor family. FEBS Lett. 2013, 587, 1767–1772. [Google Scholar] [CrossRef]
- Song, H.K.; Suh, S.W. Kunitz-type soybean trypsin inhibitor revisited: Refined structure of its complex with porcine trypsin reveals an insight into the interaction between a homologous inhibitor from Erythrina caffra and tissue-type plasminogen activator. J. Mol. Biol. 1998, 275, 347–363. [Google Scholar] [CrossRef]
- Gahloth, D.; Selvakumar, P.; Shee, C.; Kumar, P.; Sharma, A.K. Cloning, sequence analysis and crystal structure determination of a miraculin-like protein from Murrayakoenigii. Arc. Biochem. Biophys. 2010, 494, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Prasad, E.R.; Dutta-Gupta, A.; Padmasree, K. Inhibitors from pigeonpea active against lepidopteran gut proteinases. J. Econ. Entomol. 2009, 102, 2343–2349. [Google Scholar] [CrossRef] [PubMed]
- Gahloth, D.; Shukla, U.; Birah, A.; Gupta, G.P.; Ananda Kumar, P.; Dhaliwal, H.S.; Sharma, A.K. Bioinsecticidal activity of Murrayakoenigii miraculin-like protein against Helicoverpa armigera and Spodoptera litura. Arch. Insect Biochem. Physiol. 2011, 78, 132–144. [Google Scholar] [CrossRef] [PubMed]
- Verberne, M.C.; Verpoorte, R.; Bol, J.F.; Mercado-Blanco, J.; Linthorst, H.J. Over production of salicylic acid in plants by bacterial transgenes enhances pathogen resistance. Nat. Biotechnol. 2000, 18, 779–783. [Google Scholar] [CrossRef]
- Malamy, J.; Carr, J.P.; Klessig, D.F.; Raskin, I. Salicylic acid: A likely endogenous signal in the resistance response of tobacco to viral infection. Science 1990, 250, 1002–1004. [Google Scholar] [CrossRef]
- Silva, K.J.; Mahna, N.; Mou, Z.; Folta, K.M. NPR1 as a transgenic crop protection strategy in horticultural species. Hortic Res. 2018, 5, 15. [Google Scholar] [CrossRef]
- Dutt, M.; Barthe, G.; Irey, M.; Grosser, J. Correction: Transgenic citrus expressing an Arabidopsis NPR1 gene exhibit enhanced resistance against Huanglongbing (HLB, Citrus greening). PLoS ONE 2016, 11, e0147657. [Google Scholar] [CrossRef]
- Barbosa-Mendes, J.M.; Mourão Filho, F.D.; Bergamin Filho, A.; Harakava, R.; Beer, S.V.; Mendes, B.M. Genetic transformation of Citrus sinensis cv. Hamlin with hrpN gene from Erwinia amylovora and evaluation of the transgenic lines for resistance to citrus canker. Sci. Hortic. 2009, 122, 109–115. [Google Scholar] [CrossRef]
- Durner, J.; Klessig, D.F. Inhibition of ascorbate peroxidase by salicylic acid and 2, 6-dichloroisonicotinic acid, two inducers of plant defense responses. Proc. Natl. Acad. Sci. USA 1995, 92, 11312–11316. [Google Scholar] [CrossRef]
- Bi, Y.M.; Kenton, P.; Mur, L.; Darby, R.; Draper, J. Hydrogen peroxide does not function downstream of salicylic acid in the induction of PR protein expression. Plant J. 1995, 8, 235–245. [Google Scholar] [CrossRef]
- Inoue, H.; Ohnishi, J.; Ito, T.; Tomimura, K.; Miyata, S.; Iwanami, T. Enhanced proliferation and efficient transmission of Candidatus Liberibacter asiaticus by adult Diaphorinacitri after acquisition feeding in the nymphal stage. Ann. Appl. Biol. 2009, 155, 29–36. [Google Scholar] [CrossRef]
- Ammar, E.-D.; Ramos, J.E.; Hall, D.G.; Dawson, W.O.; Shatters, R.G., Jr. Acquisition, Replication and Inoculation of Candidatus Liberibacter asiaticus following Various Acquisition Periods on Huanglongbing-Infected Citrus by Nymphs and Adults of the Asian Citrus Psyllid. PLoS ONE 2016, 11, e0159594. [Google Scholar] [CrossRef] [PubMed]
- Pelz-Stelinski, K.S.; Killiny, N. Better together: Association with ‘Candidatus Liberibacter asiaticus’ increases the reproductive fitness of its insect vector, Diaphorinacitri (Hemiptera: Liviidae). Ann. Entomol. Soc. Am. 2016, 109, 371–376. [Google Scholar] [CrossRef] [PubMed]
- Rogers, M.E.; Stansley, P.R.; Stelinski, L.L. Florida Citrus Pest Management Guide: Asian Citrus Psyllid and Citrus Leafminer; Entomology and Nematology Department, Florida Cooperative Extension Service, Institute of Food and Agricultural Sciences, University of Florida: Gainesville, FL, USA, 2012; Available online: http://edis.ifas.ufl.edu/in686/ENY734 (accessed on 25 November 2022).
- Cocco, A.; Hoy, M.A. Toxicity of organosilicone adjuvants and selected pesticides to the Asian citrus psyllid (Hemiptera: Psyllidae) and its parasitoid Tamarixia radiata (Hymenoptera: Eulophidae). Fla. Entomol. 2008, 91, 610–620. [Google Scholar]
- Hall, D.G.; Nguyen, R. Toxicity of pesticides to Tamarixia radiata, a parasitoid of the Asian citrus psyllid. Bio Control 2010, 55, 601–611. [Google Scholar] [CrossRef]
- Avery, P.B.; Pick, D.A.; Aristizábal, L.F.; Kerrigan, J.; Powell, C.A.; Rogers, M.E.; Arthurs, S.P. Compatibility of Isariafumosorosea (Hypocreales: Cordycipitaceae) blastospores with agricultural chemicals used for management of the Asian citrus psyllid, Diaphorinacitri (Hemiptera: Liviidae). Insects 2013, 4, 694–711. [Google Scholar] [CrossRef]
- Casique-Valdez, R.; Reyes-Martínez, A.Y.; Sánchez-Peña, S.R.; Bidochka, M.J.; López-Arroyo, J.I. Pathogenicity of Hirsutellacitriformis (Ascomycota: Cordycipitaceae) to Diaphorinacitri (Hemiptera: Psyllidae) and Bactericeracockerelli (Hemiptera: Triozidae). Fla. Entomol. 2011, 94, 703–705. [Google Scholar] [CrossRef]
- Diniz, A.J.F.; Garcia, A.G.; Alves, G.R.; Reigada, C.; Vieira, J.M.; Parra, J.R.P. The enemy is outside: Releasing the parasitoid Tamarixia radiata (Hymenoptera: Eulophidae) in external sources of HLB inocula to control the Asian Citrus Psyllid Diaphorinacitri (Hemiptera: Liviidae). Neotrop. Entomol. 2020, 49, 250–257. [Google Scholar] [CrossRef]
- Conceschi, M.R.; D’Alessandro, C.P.; Moral, R.; Demetrio, C.G.B.; Júnior, I.D. Transmission potential of the entomopathogenic fungi Isariafumosorosea and Beauveria bassiana from sporulated cadavers of Diaphorinacitri and Toxopteracitricida to uninfected D. citri adults. Bio Control 2016, 61, 567–577. [Google Scholar]
- Dorta, S.O.; Balbinotte, J.; Monnerat, R.; Lopes, J.R.S.; Cunha, T.; Zanardi, O.Z.; Miranda, M.P. Selection of Bacillus thuringiensis strains in citrus and their pathogenicity to Diaphorinacitri (Hemiptera: Liviidae) nymphs. Insect Sci. 2020, 27, 519–530. [Google Scholar] [CrossRef]
- Boina, D.R.; Youn, Y.; Folimonova, S.; Stelinski, L.L. Effects of pymetrozine, an antifeedant of Hemiptera, on Asian citrus psyllid, Diaphorinacitri, feeding behavior, survival and transmission of Candidatus Liberibacter asiaticus. Pest Manag. Sci. 2011, 67, 146–155. [Google Scholar] [CrossRef] [PubMed]
- Grafton-Cardwell, E.E.; Stelinski, L.L.; Stansly, P.A. Biology and management of asian citrus psyllid, vector of the Huanglongbing Pathogens. Annu. Rev. Entomol. 2013, 58, 413–432. [Google Scholar] [CrossRef] [PubMed]
- Stuchi, E.S.; Giradi, E.A. Use of horticultural practices in citriculture to survive huanglongbing. Brazilian agricultural research corporation, embrapa cassava and fruits. Cruz Almas Embrapa Mandioca Frutic. 2010, 68. Available online: http://www.infoteca.cnptia.embrapa.br/infoteca/handle/doc/873095 (accessed on 25 November 2022).
- Moreira, A.S.; Stuchi, E.S.; Silva, P.R.B.; Bassanezi, R.B.; Girardi, E.A.; Laranjeira, F.F. Could tree density play a role in managing citrus huanglongbing epidemics? Trop. Plant Pathol. 2019, 44, 268–274. [Google Scholar] [CrossRef]
- de Andrade, E.C.; Girardi, E.A.; Stuchi, E.S.; Moreira, A.S.; Freitas-Astua, J.; Fancelli, M.; De Brito Silva, S.X.; Laranjeira, F.F. Citrus huanglongbing (HLB) and the Brazilian efforts to overcome the disease. Outlooks Pest Manag. 2021, 32, 189–194. [Google Scholar] [CrossRef]
- Ladaniya, M.S.; Marathe, R.A.; Das, A.K.; Rao, C.N.; Huchche, A.D.; Shirgure, P.S.; Murkute, A.A. High density planting studies in acid lime (Citrus aurantifolia Swingle). Sci. Hortic. 2020, 261, 108935. [Google Scholar] [CrossRef]
- Rodrigues, J.D.B.; Moreira, A.S.; Stuchi, E.S.; Bassanezi, R.B.; Laranjeira, F.F.; Girardi, E.A. Huanglongbing incidence, canopy volume, and sprouting dynamics of ‘Valencia’ sweet orange grafted onto 16 rootstocks. Trop. Plant Pathol. 2020, 45, 611–619. [Google Scholar] [CrossRef]
- de C Felisberto, P.A.; Girardi, E.A.; Peña, L.; Felisberto, G.; Beattie, G.A.; Lopes, S.A. Unsuitability of indigenous South American rutaceae as potential hosts of Diaphorinacitri. Pest Manag. Sci. 2019, 75, 1911–1920. [Google Scholar] [CrossRef]
- Tiwari, S.; Gondhalekar, A.D.; Mann, R.S.; Scharf, M.E.; Stelinski, L.L. Characterization of five CYP4 genes from Asian citrus psyllid and their expression levels in Candidatus Liberibacter asiaticus-infected and uninfected psyllids. Insect Mol. Biol. 2011, 20, 733–744. [Google Scholar] [CrossRef]
- Killiny, N.; Hajeri, S.; Tiwari, S.; Gowda, S.; Stelinski, L.L. Double-stranded RNA uptake through topical application, mediates silencing of five CYP4 genes and suppresses insecticide resistance in Diaphorinacitri. PLoS ONE 2014, 9, e110536. [Google Scholar] [CrossRef]
- El-Shesheny, I.; Hajeri, S.; El-Hawary, I.; Gowda, S.; Killiny, N. Silencing abnormal wing disc gene of the Asian Citrus Psyllid, Diaphorinacitri Disrupts Adult wing development and increases nymph mortality. PLoS ONE 2013, 8, e65392. [Google Scholar] [CrossRef] [PubMed]
- Ferrara, T.F.D.S.; Schneider, V.K.; Kishi, L.T.; Carmona, A.K.; Alves, M.F.M.; Belasque-Júnior, J.; Rosa, J.C.; Hunter, W.B.; Henrique-Silva, F.; Soares-Costa, A. Characterization of a recombinant cathepsin B-Like cysteine peptidase from Diaphorinacitri kuwayama (Hemiptera: Liviidae): A putative target for control of citrus huanglongbing. PLoS ONE 2015, 10, e0145132. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Gowda, S.; Killiny, N. Double-stranded RNA delivery through soaking mediates silencing of the muscle protein 20 and increases mortality to the Asian citrus psyllid, Diaphorinacitri. Pest Manag. Sci. 2017, 73, 1846–1853. [Google Scholar] [CrossRef] [PubMed]
- Kishk, A.; Anber, H.A.; AbdEl-Raof, T.K.; El-Sherbeni, A.H.; Hamed, S.; Gowda, S.; Killiny, N. RNA interference of carboxyesterases causes nymph mortality in the Asian citrus psyllid, Diaphorinacitri. Arch. Insect Biochem. Physiol. 2017, 94, e21377. [Google Scholar] [CrossRef] [PubMed]
- Galdeano, D.M.; Breton, M.C.; Lopes, J.R.S.; Falk, B.W.; Machado, M.A. Oral delivery of double-stranded RNAs induces mortality in nymphs and adults of the Asian citrus psyllid, Diaphorinacitri. PLoS ONE 2017, 12, e0171847. [Google Scholar] [CrossRef]
- Killiny, N.; Kishk, A. Delivery of dsRNA through topical feeding for RNA interference in the citrus sap piercing-sucking hemipteran, Diaphorinacitri. Arch. Insect Biochem Physiol. 2017, 95, e21394. [Google Scholar] [CrossRef]
- Hajeri, S.; Killiny, N.; El-Mohtar, C.; Dawson, W.O.; Gowda, S. Citrus tristeza virus-based RNAi in citrus plants induces gene silencing in Diaphorinacitri, a phloem-sap sucking insect vector of citrus greening disease (Huanglongbing). J. Biotechnol. 2014, 176, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Dawson, W.O.; Bar-Joseph, M.; Garnsey, S.M.; Moreno, P. Citrus tristeza virus: Making an ally from an enemy. Annu. Rev. Phytopathol. 2015, 53, 137–155. [Google Scholar] [CrossRef]
- Yu, X.; Killiny, N. RNA interference-mediated control of Asian citrus psyllid, the vector of the huanglongbing bacterial pathogen. Trop.Plant Pathol. 2020, 45, 298–305. [Google Scholar] [CrossRef]
- Folimonova, S.Y.; Sun, Y.D. Citrus Tristeza Virus: From Pathogen to Panacea. Annu. Rev. Virol. 2022, 9, 417–435. [Google Scholar] [CrossRef]
- Harris, A.F.; Nimmo, D.; McKemey, A.R.; Kelly, N.; Scaife, S.; Donnelly, C.A.; Beech, C.; Petrie, W.D.; Alphey, L. Field performance of engineered male mosquitoes. Nat. Biotechnol. 2011, 29, 1034–1037. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, D.O.; McKemey, A.R.; Garziera, L.; Lacroix, R.; Donnelly, C.A.; Alphey, L.; Malavasi, A.; Capurro, M.L. Suppression of a field population of Aedes aegypti in Brazil by sustained release of transgenic male mosquitoes. PLoS Negl. Trop. Dis. 2015, 9, e0003864. [Google Scholar] [CrossRef] [PubMed]
- Thomé, R.C.A.; Yang, H.M.; Esteva, L. Optimal control of Aedes aegypti mosquitoes by the sterile insect technique and insecticide. Math. Biosci. 2010, 223, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Bouyer, J.; Lefrançois, T. Boosting the sterile insect technique to control mosquitoes. Trends Parasitol. 2014, 30, 271–273. [Google Scholar] [CrossRef] [PubMed]
- Marco, F.G.; Yoshimoto, A.; Simmons, G.; Stouthamer, R. First Steps in the Development of the Sterile Insect Technique as a Control Tactic for the Asian Citrus Psyllid, Diaphorina Citri Kuwayama (Hemiptera: Liviidae). Entomology 2017, 368. Available online: https://esa.confex.com/esa/2017/meetingapp.cgi/Paper/125648 (accessed on 25 November 2022).
- Gatineau, F.; Loc, H.T.; Tuyen, N.D.; Tuan, T.M.; Hien, N.T.D.; Truc, N.T.N. Effects of two insecticide practices on population dynamics of Diaphorinacitri and huanglongbing incidence in south Vietnam. In Proceedings of the Huanglongbing–Greening International Workshop, Ribeirão Preto, Brazil, 6 July 2006; p. 110. [Google Scholar]
- Gottwald, T.R.; Graham, J.H.; Irey, M.S.; McCollum, T.G.; Wood, B.W. Inconsequential effect of nutritional treatments on huanglongbing control, fruit quality, bacterial titer and disease progress. Crop Prot. 2012, 36, 73–82. [Google Scholar] [CrossRef]
- Zaka, S.M.; Zeng, X.N.; Holford, P.; Beattie, G.A.C. Repellent effect of guava leaf volatiles on settlement of adults of citrus psylla, Diaphorinacitri Kuwayama, on citrus. Insect Sci. 2010, 17, 39–45. [Google Scholar] [CrossRef]
- Alquézar, B.; Volpe, H.X.L.; Magnani, R.F.; de Miranda, M.P.; Santos, M.A.; Wulff, N.A.; Bento, J.M.; Parra, J.R.; Bouwmeester, H.; Pena, L. β-caryophyllene emitted from a transgenic Arabidopsis or chemical dispenser repels Diaphorinacitri, vector of Candidatus Liberibacters. Sci. Rep. 2017, 7, 5639. [Google Scholar] [CrossRef]
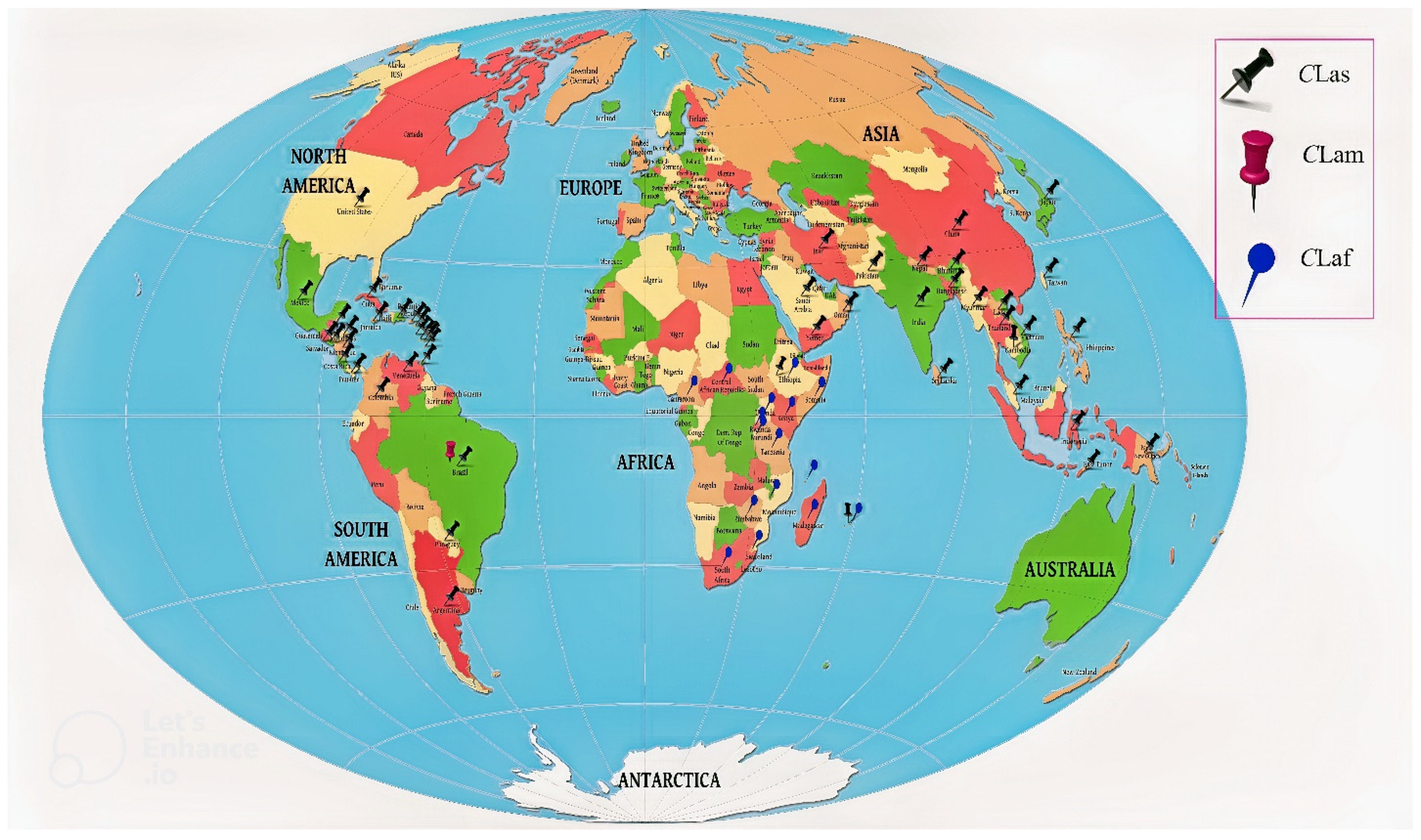



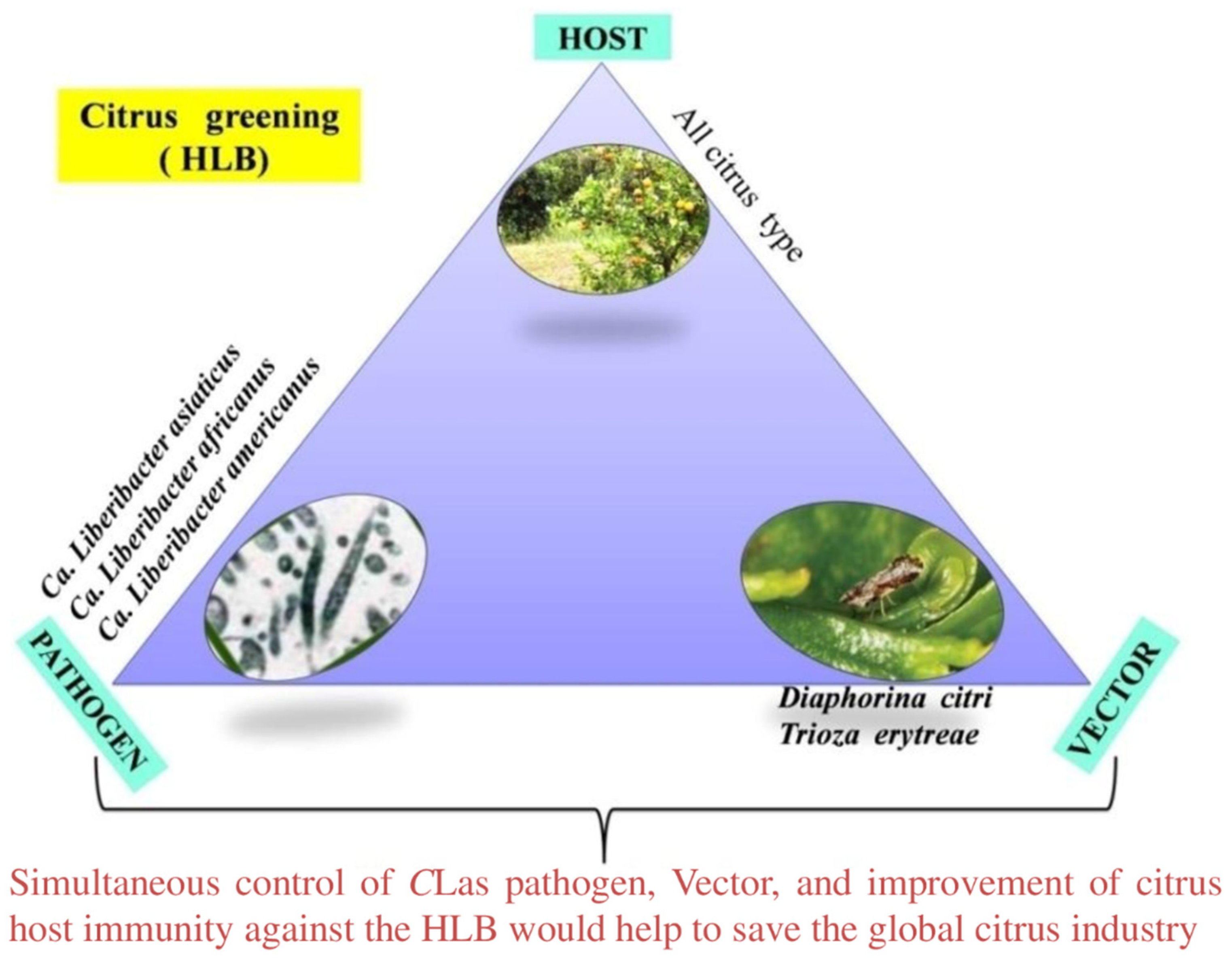
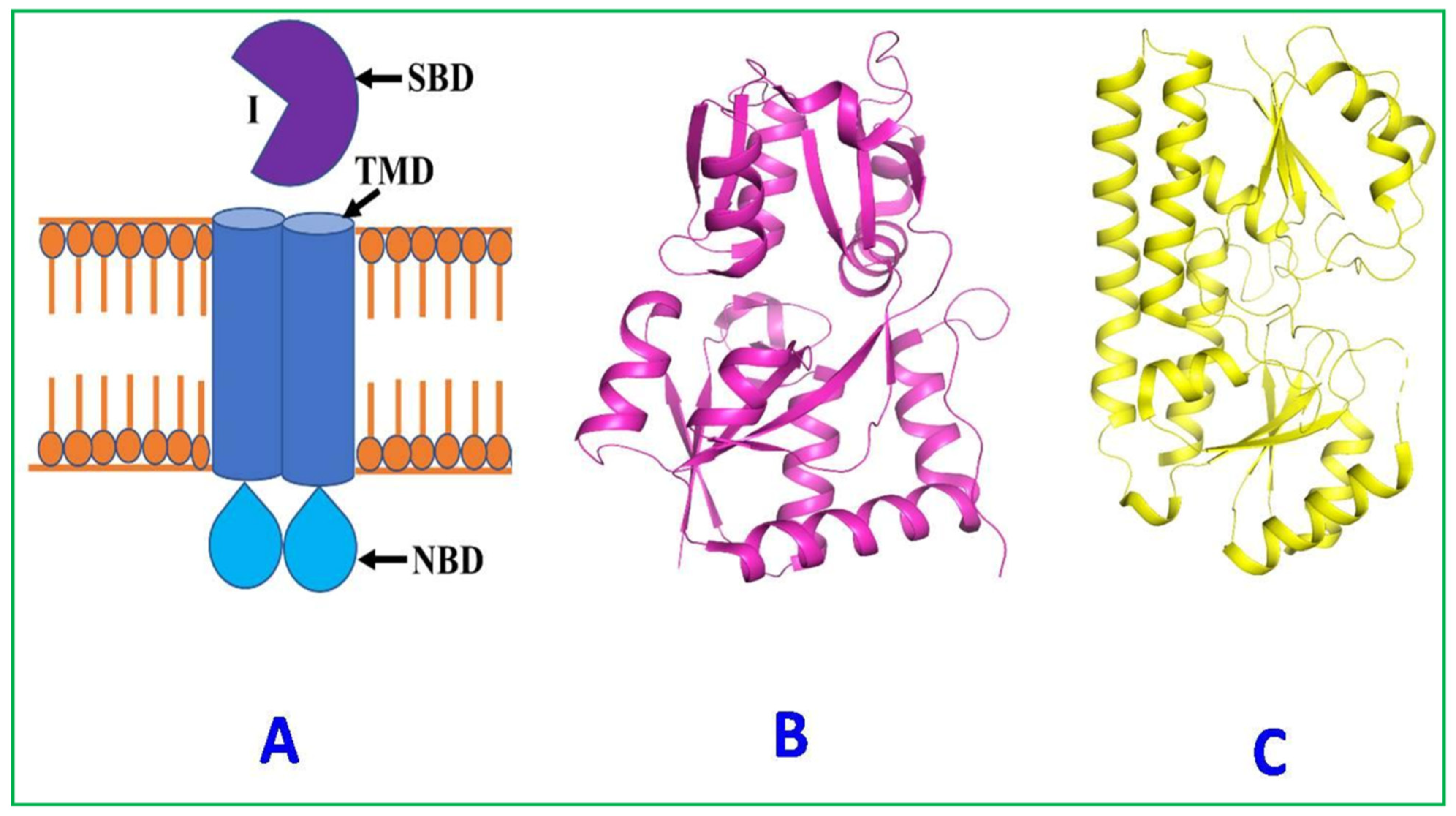
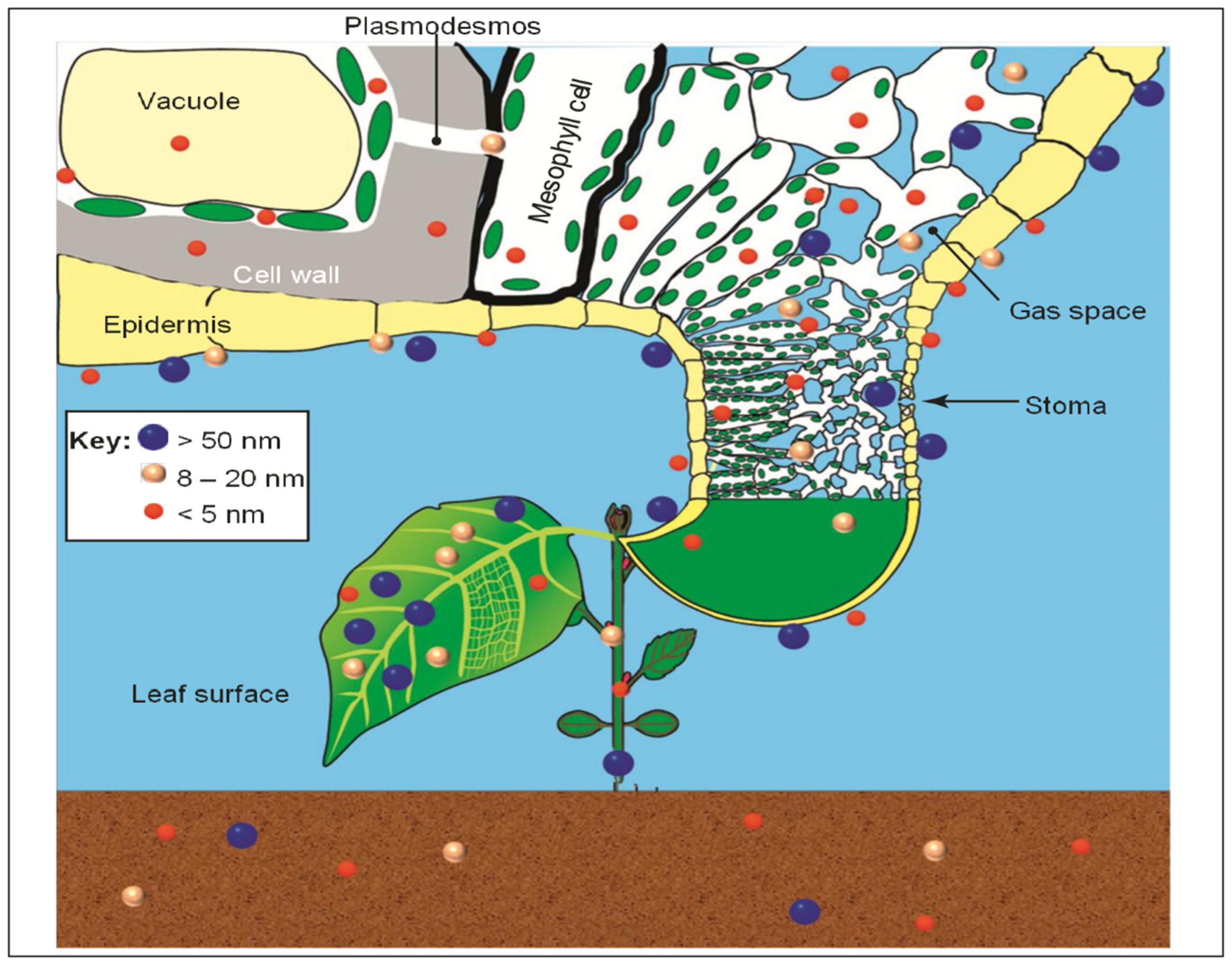
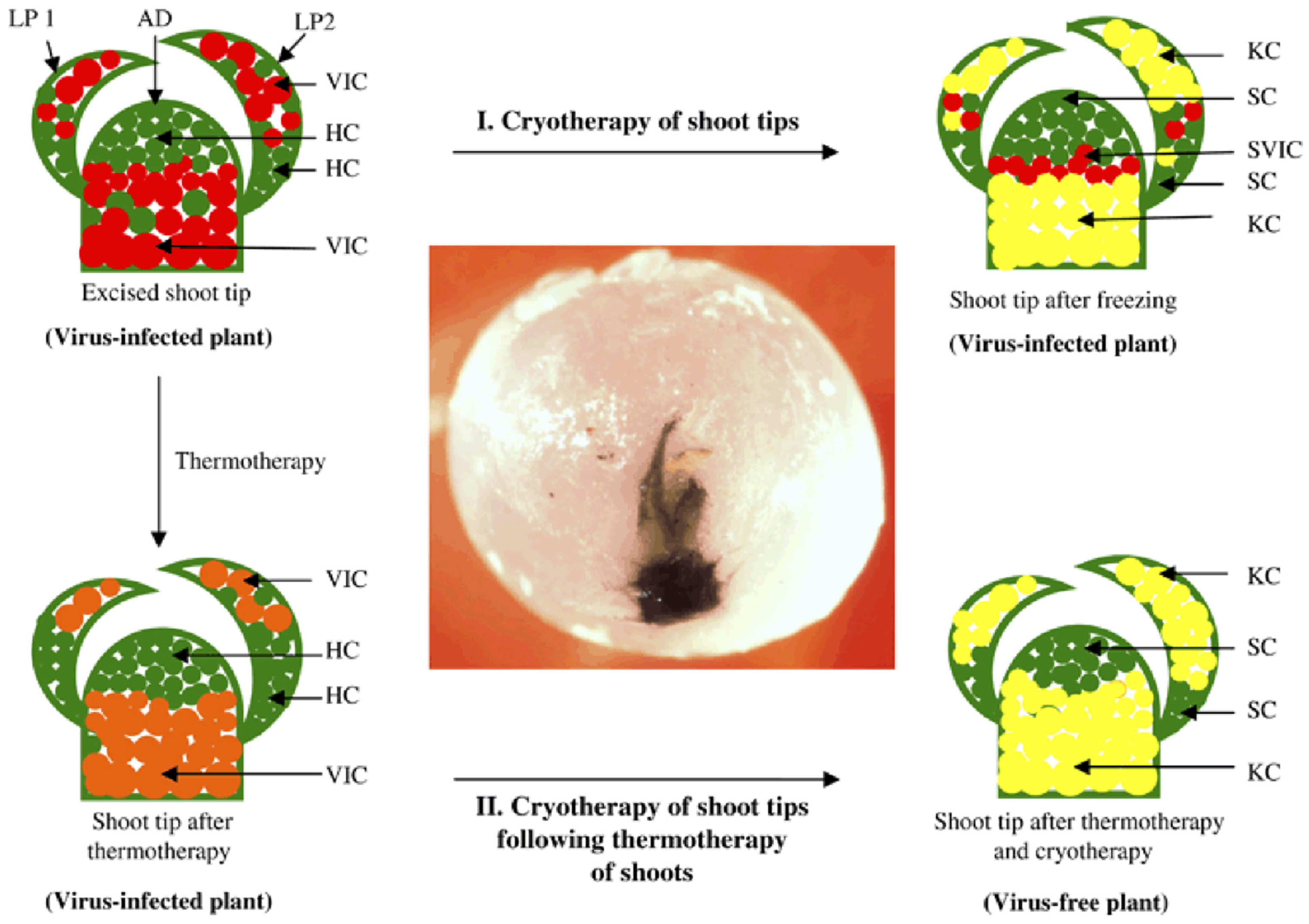

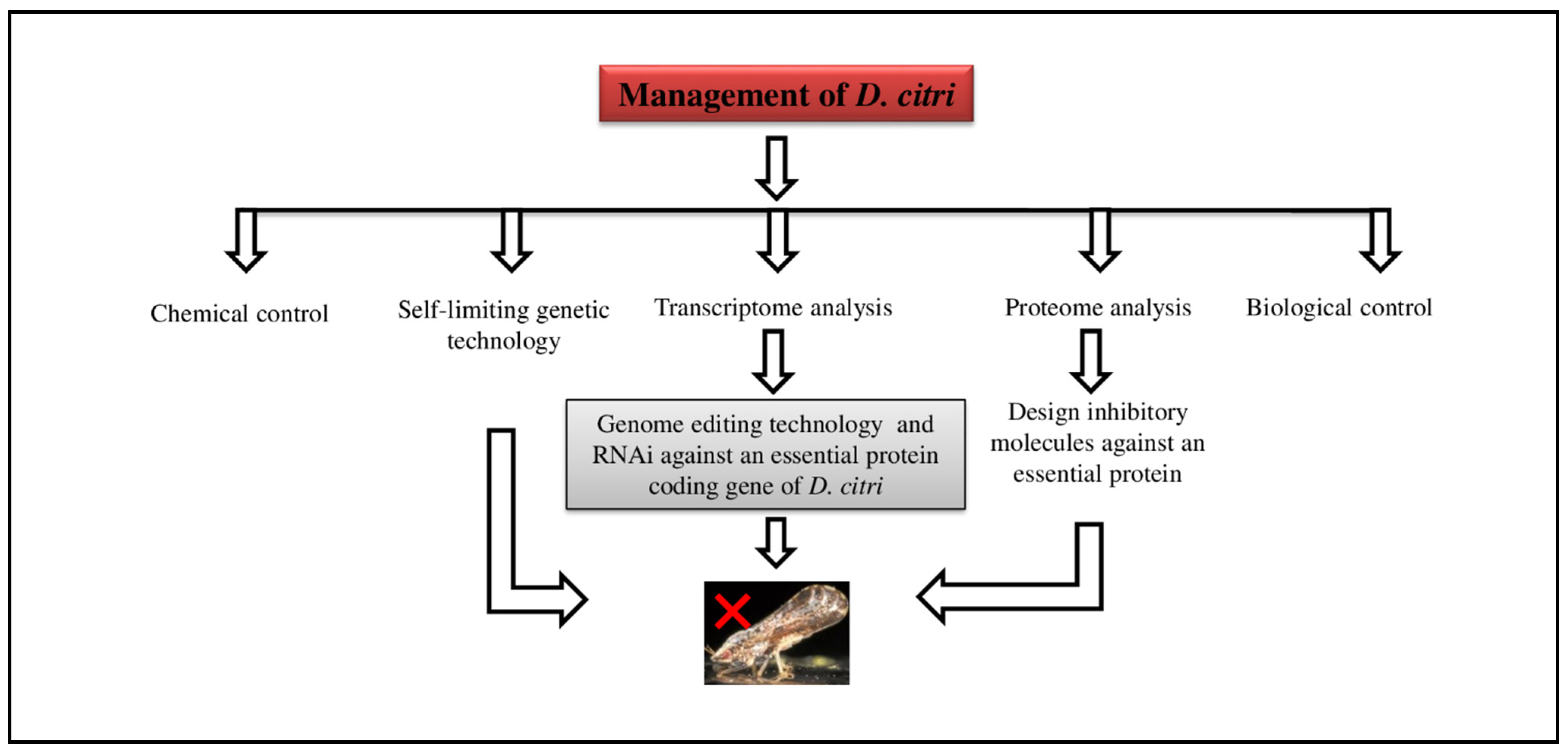


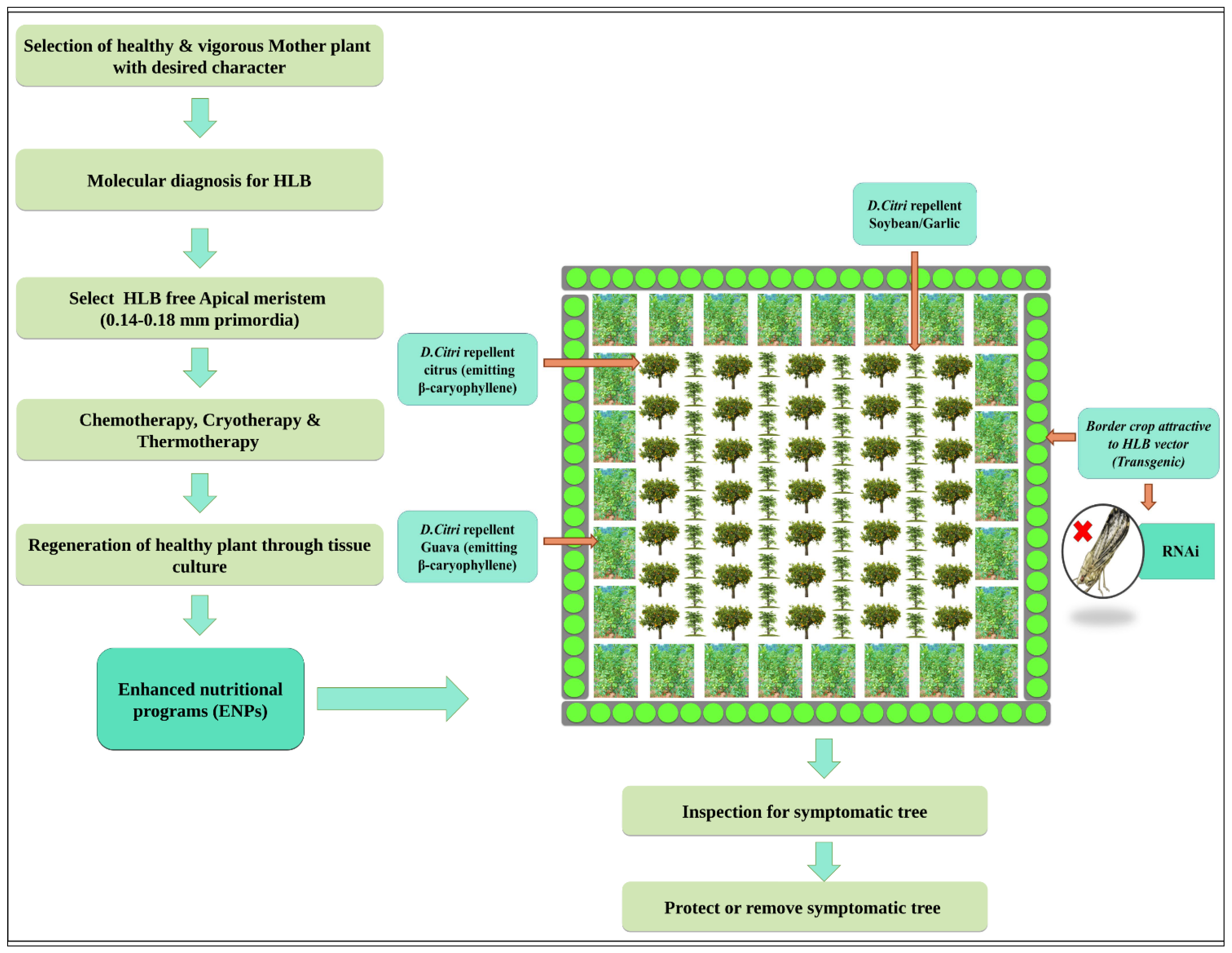
| Sr. No | Continent/Country/Region | Distribution of HLB | Causal Organism | Reference |
|---|---|---|---|---|
| Asia | ||||
| 1 | Bangladesh | Present | Ca. Liberibacter asiaticus | [21] |
| 2 | Bhutan | Present | Ca. Liberibacter asiaticus | [22] |
| 3 | Cambodia | Present | Ca. Liberibacter asiaticus | [23] |
| 4 | China | Present | Ca. Liberibacter asiaticus | [22] |
| 5 | India | Present, Widespread | Ca. Liberibacter asiaticus | [24] |
| 6 | Indonesia | Present | Ca. Liberibacter asiaticus | [25] |
| 7 | Iran | Present, Localized | Ca. Liberibacter asiaticus | [26] |
| 8 | Japan | Present | Ca. Liberibacter asiaticus | [27] |
| 9 | Laos | Present | Ca. Liberibacter asiaticus | [23] |
| 10 | Malaysia | Present, Localized | Ca. Liberibacter asiaticus | [24] |
| 11 | Myanmar | Present | Ca. Liberibacter asiaticus | [23] |
| 12 | Nepal | Present, Widespread | Ca. Liberibacter asiaticus | [28] |
| 13 | Oman | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 14 | Pakistan | Present | Ca. Liberibacter asiaticus | [29] |
| 15 | Philippines | Present, Widespread | Ca. Liberibacter asiaticus | [29] |
| 16 | Saudi Arabia | Present, Localized | Ca. Liberibacter asiaticus | [30] |
| 17 | Sri Lanka | Present | Ca. Liberibacter asiaticus | [22] |
| 18 | Taiwan | Present, Widespread | Ca. Liberibacter asiaticus | [22] |
| 19 | Thailand | Present | Ca. Liberibacter asiaticus | [31] |
| 20 | Vietnam | Present, Localized | Ca. Liberibacter asiaticus | [32] |
| 21 | Yemen | Present, Localized | Ca. Liberibacter asiaticus | [30] |
| North America | Ca. Liberibacter asiaticus | |||
| 22 | Barbados | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 23 | Belize | Present, Localized | Ca. Liberibacter asiaticus | [17] |
| 24 | Costa Rica | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 25 | Cuba | Present, Widespread | Ca. Liberibacter asiaticus | [22] |
| 26 | Dominica | Present, Few occurrences | Ca. Liberibacter asiaticus | [22] |
| 27 | Dominican Republic | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 28 | El Salvador | Present | Unknown | [33] |
| 29 | Guadeloupe | Present, Localized | Unknown | [34] |
| 30 | Guatemala | Present | Ca. Liberibacter asiaticus | [22] |
| 31 | Honduras | Present | Ca. Liberibacter asiaticus | [22] |
| 32 | Jamaica | Present, Widespread | Ca. Liberibacter asiaticus | [22] |
| 33 | Martinique | Present, Localized | Ca. Liberibacter asiaticus | [34] |
| 34 | Mexico | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 35 | Nicaragua | Present | Ca. Liberibacter asiaticus | [22] |
| 36 | Panama | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 37 | Puerto Rico | Present | Ca. Liberibacter asiaticus | [22] |
| 38 | Trinidad and Tobago | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 39 | U.S. Virgin Islands | Present, Few occurrences | Ca. Liberibacter asiaticus | [22] |
| 40 | United States | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| South America | ||||
| 41 | Argentina | Present, Localized | Ca. Liberibacter asiaticus | [22] |
| 42 | Brazil | Present, Localized | Ca. Liberibacter americanus and Ca. Liberibacter asiaticus | [9] |
| 43 | Colombia | Present, Few occurrences | Ca.Liberibacter asiaticus | [22] |
| 44 | Paraguay | Present, Localized | Ca. Liberibacter asiaticus | [35] |
| 45 | Venezuela | Present | Ca. Liberibacter asiaticus | [36] |
| Africa | ||||
| 46 | Burundi | Present | Ca. Liberibacter africanus | [37] |
| 47 | Cameroon | Present | Ca. Liberibacter africanus | [37] |
| 48 | Central African Republic | Present | Ca. Liberibacter africanus | [37] |
| 49 | Comoros | Present | Ca. Liberibacter africanus | [22] |
| 50 | Eswatini | Present | Ca. Liberibacter africanus | [38] |
| 51 | Ethiopia | Present | Ca. Liberibacter africanus and Ca. Liberibacter asiaticus | [37] |
| 52 | Kenya | Present | Ca. Liberibacter africanus | [29] |
| 53 | Madagascar | Present | Ca. Liberibacter africanus | [38] |
| 54 | Malawi | Present | Ca. Liberibacter africanus | [37] |
| 55 | Mauritius | Present | Ca. Liberibacter africanus and Ca. Liberibacter asiaticus | [22] |
| 56 | Rwanda | Present | Ca. Liberibacter africanus | [37] |
| 57 | Somalia | Present | Ca. Liberibacter africanus | [22] |
| 58 | South Africa | Present, Localized | Ca. Liberibacter africanus | [39] |
| 59 | Tanzania | Present, Localized | Ca. Liberibacter africanus | [38] |
| 60 | Uganda | Present | Ca. Liberibacter africanus | [40] |
| 61 | Zimbabwe | Present, Localized | Ca. Liberibacter africanus | [29] |
| Protein | Organism | PDB Id | Ligand | Dissociation Constant (KD) | References | |
|---|---|---|---|---|---|---|
| ArtJ | Geobacillus stearothermophilus | 2Q2A | Arginine | 0.039 µM | [137] | |
| 2Q2C | Histidine | 0.42 µM | ||||
| 2PVU | Lysine | 0.49 µM | ||||
| GlnH | Mycobacterium tuberculosis | 6H2T | Glutamate | 15.2 µM | [138] | |
| 6H1U | Aspartate | 4.8µM | ||||
| 6H20 | Aspargine | |||||
| DEBP | Shigella flexneri | 2VHA | Glutamate | 1.6 µM | [139] | |
| Aspartate | 5.2 µM | |||||
| GlnP | Lactococcuslactis | 4KQP | Glutamine | 1.49 µM | [140] | |
| CjaA | Campylobacter jejuni | 1XT8 | Cysteine | 100 µM | [141] | |
| GlnBP | Escherichia coli | 1WDN | Glutamine | 0.5µM | [127] | |
| HBP | Escherichia coli | 1HSL | Histidine | 0.064 µM | [142] | |
| HisJ | Salmonella typhimurium | 1HPB | Histidine | 0.030 µM | [143] | |
| LAO | Salmonella typhimurium | 1LST | Lysine | 0.014 µM | [143] | |
| Ngo0372 | Neisseria gonorrhoeae | 2YLN | Cystine | 0.021 µM | [144] | |
| Ngo2014 | Neisseria gonorrhoeae | 2YJP | Cysteine | 0.026 µM | [144] | |
| CLasTcyA | Candidatus Liberibacter asiaticus | 6A80 | Cystine | 1.26 µM | [145] | |
| ClasTcyA Mutant (V58W) | Candidatus Liberibacter asiaticus | - | Cystine | 0.22 µM | [146] | |
| CLas-ZnuA2 | Candidatus Liberibacterasiaticus | 4UDO | Mn2+ | 370 µM | [147,148] | |
| 5AFS | Zn2+ | 430 µM | ||||
| 6IXI | Cd2+ | [146] | ||||
| CLas-ZnuA2 Mutant | (S38A) | Candidatus Liberibacterasiaticus | 5Z2K | Mn2+ | 340 µM | [149] |
| (Y68F) | Candidatus Liberibacterasiaticus | 5ZHA | Mn2+ | 540 µM | ||
| LBP | Escherichia coli | 1USK | Leucine | 0.4 µM | [150] | |
| 1USI | Phenylalanine | 0.18 µM | ||||
| GenBank Accession No £. | Super Family | Species | Domain | Putative Function | CLas Relative Expression $ | |
|---|---|---|---|---|---|---|
| Query Coverage € | Identity € | |||||
| ACT56612 | MlaF | Acinetobacter baumanii | NBD | Putative ATP-binding component of ABC transporter: Involved in resistance to organic component | 7.60 | |
| 92% | 38.49% | |||||
| ACT56643 | PBP2_ BztA | Brucellaovis | SBD | Putative cationic amino acid ABC transporter | −1.29 | |
| 93% | 57.32% | |||||
| ACT56645 (aapM) | TM_ PBP2 | Caldanaerobactersubterraneus | Permease | Putative general L-amino acid transport system permease | 3.64 | |
| 54% | 30.77% | |||||
| ACT56815 (proX) | PBP2_ ChoX | Sinorhizobiummeliloti | SBD | Putative glycine betaine/proline ABC transporter | 1.84 | |
| 89% | 50.0% | |||||
| ACT56816 | FieF | Escherichia coli | Efflux protein | Predicted cation (Co/Zn/Cd) efflux transporters | 3.88 | |
| 91% | 26.83% | |||||
| ACT57010 (ZnuA2) | TroA-like | Yersinia pestis | SBD | Fi/Mn transport | 7.92 | |
| 92% | 49.26% | |||||
| ACT57013 | ZnuB | - | Permease | Putative Mn2+/Zn2+ transport system | 5.72 | |
| - | - | |||||
| ACT57585 | PBP2 | Neisseria gonorrhoeae | SBD | General L-amino acid Transport | 2.97 | |
| 83% | 32.33% | |||||
| ACT57586 | TM_ PBP2 | Caldanaerobactersubterraneus | Permease | ABC-type amino acid transport system | 7.96 | |
| 83% | 34.04% | |||||
| ACT57180 | PBP2 | Planctopiruslimnophila | SBD | Putative periplasmic phosphate-binding protein | 1.28 | |
| 82% | 29.01% | |||||
| ACT57178 | PstA | - | Permease | Putative phosphate transporter | 2.23 | |
| - | - | |||||
| ACT57369 | livM | - | Permease | Putative branch chain aminoacid (Leucine, Isoleucine, Valine) transporter | - | |
| - | - | |||||
| GenBank Accession No | Regulator Type | Protein Name | Putative Function | Mw (kDa) | CLas Relative Expression $ | References |
|---|---|---|---|---|---|---|
| ACT56890 | CarD | PrbP | Regulate some ribosomal gene expression (Activator) | 21.3 Kda | 1.24 | [159] |
| ACT56824 | MarR | LdtR | Activator and Repressor of gene expression | 19.6 Kda | 5.23 | [160] |
| ACT57167 | LuxR | VisN | Activator of chemotaxis, flagellar, and motility genes. | 28.5 Kda | 2.75 | [161] |
| ACT56755 | LysR | LsrB | Activator of oxidativestress lipopolysaccharide biosynthesis gene. | 34.3 Kda | 2.23 | [163] |
| ACT56897 | HTH-XRE | PhrR1 | Gene related to quorum sensing | 16.5 Kda | Not reported | [162] |
| ACT57366 | Response regulator | CtrA | Control cell cycle | 26.7 Kda | 1.4 | [164] |
| ACT57093 | IscR | RirA | Response to iron limitation | 16.1 Kda | 1.04 | [165] |
| Protein | GenBank Accession No | PDB ID | References |
|---|---|---|---|
| Periplasmic metal-binding protein | ACT57010 | 4CL2, 4UDN, 4UDO, 5AFS, 5Z2J, 5Z2K, 5Z35, 5ZHA, 6IXI | [146,147,148,149] |
| Putative amino-acid-binding periplasmic ABC transporter protein | ACT57585 | 6A80, 6AA1, 6AAL, 6A8S | [145] |
| Enoyl-Acyl Carrier Protein Reductase I (FabI) | KAE9510327 | 4NK4, 4NK5 | [183] |
| beta-Hydroxyacyl-acyl carrier protein dehydratase (FabZ) | WP_015452391 | 4ZW0 | [185] |
| Inosine 5′-monophosphate Dehydrogenase | ACT57362 | 6KCF | [187] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghosh, D.; Kokane, S.; Savita, B.K.; Kumar, P.; Sharma, A.K.; Ozcan, A.; Kokane, A.; Santra, S. Huanglongbing Pandemic: Current Challenges and Emerging Management Strategies. Plants 2023, 12, 160. https://doi.org/10.3390/plants12010160
Ghosh D, Kokane S, Savita BK, Kumar P, Sharma AK, Ozcan A, Kokane A, Santra S. Huanglongbing Pandemic: Current Challenges and Emerging Management Strategies. Plants. 2023; 12(1):160. https://doi.org/10.3390/plants12010160
Chicago/Turabian StyleGhosh, Dilip, Sunil Kokane, Brajesh Kumar Savita, Pranav Kumar, Ashwani Kumar Sharma, Ali Ozcan, Amol Kokane, and Swadeshmukul Santra. 2023. "Huanglongbing Pandemic: Current Challenges and Emerging Management Strategies" Plants 12, no. 1: 160. https://doi.org/10.3390/plants12010160
APA StyleGhosh, D., Kokane, S., Savita, B. K., Kumar, P., Sharma, A. K., Ozcan, A., Kokane, A., & Santra, S. (2023). Huanglongbing Pandemic: Current Challenges and Emerging Management Strategies. Plants, 12(1), 160. https://doi.org/10.3390/plants12010160





