Genome-Wide Association Study of Phenylalanine Derived Glucosinolates in Brassica rapa
Abstract
:1. Introduction
2. Results
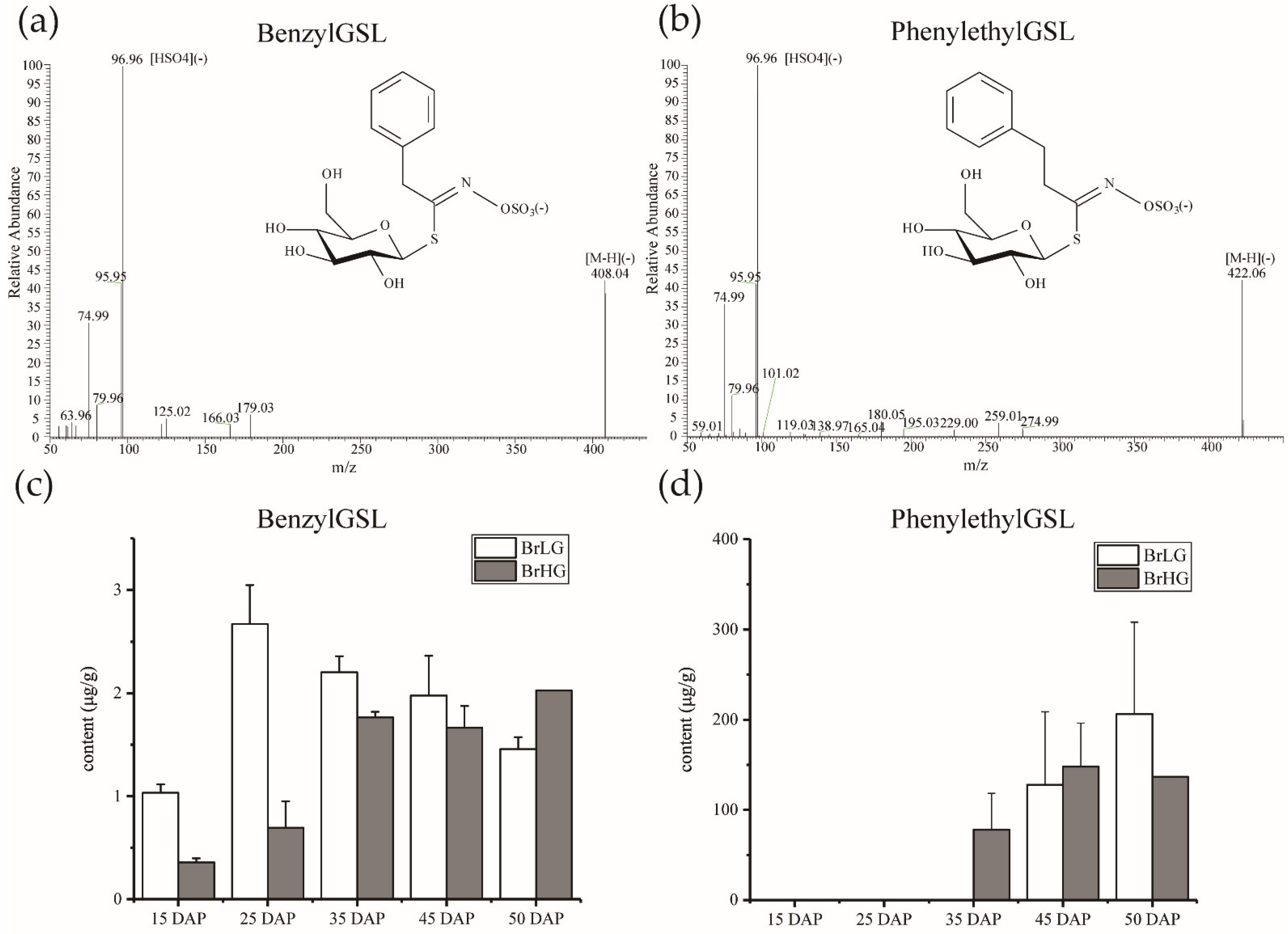
2.1. The Dynamic Accumulation of Phe-Derived Glucosinolates in B. rapa Seeds
2.2. QTNs (Quantitative Trait Nucleotides) Detected for Two Phe-Derived Glucosinolates
2.3. Screening of Candidate Genes for Two Phe-Derived Glucosinolates
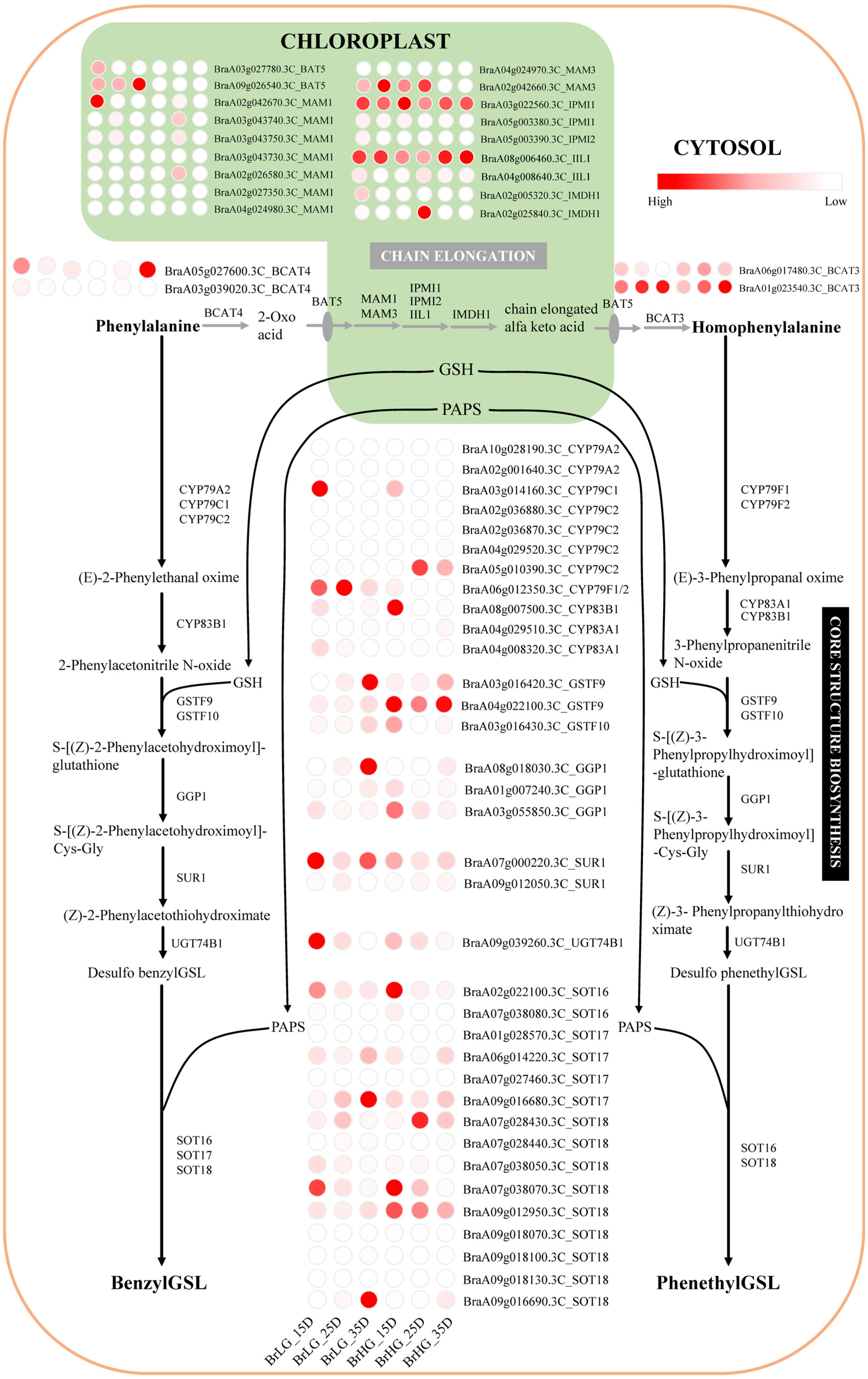
2.4. Regulation of Phe-Derived Glucosinolates Biosynthesis Pathway in B. rapa Seeds
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Sampling
4.2. Glucosinolates Extraction
4.3. UPLC-HESI-MS/MS Analysis
4.4. Data Processing and Glucosinolate Identification
4.5. Transcriptome Analysis
4.6. mGWAS Analysis of Phe-Derived Glucosinolates
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wittstock, U.; Halkier, B.A. Glucosinolate research in the Arabidopsis era. Trends Plant Sci. 2002, 7, 263–270. [Google Scholar] [CrossRef]
- Prieto, M.A.; Lopez, C.J.; Simal-Gandara, J. Glucosinolates: Molecular structure, breakdown, genetic, bioavailability, properties and healthy and adverse effects. Adv. Food Nutr. Res. 2019, 90, 305–350. [Google Scholar] [PubMed]
- Priyamedha, B.K.; Thomas, L.; Bala, M.; Singh, V.V.; Singh, D. Status and perspective of canola quality rapeseed-mustard cultivation in India: A review. J. Oilseed Brassica 2015, 6, 142–151. [Google Scholar]
- Lee, J.W.; Wang, S.; Huang, Y.; Seefeldt, T.; Donkor, A.; Logue, B.A.; Woyengo, T.A. Toxicity of canola-derived glucosinolates in pigs fed resistant starch-based diets. J. Anim. Sci. 2020, 98, skaa111. [Google Scholar] [CrossRef]
- Halkier, B.A.; Gershenzon, J. Biology and biochemistry of glucosinolates. Annu. Rev. Plant Biol. 2006, 57, 303–333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sønderby, I.E.; Geu-Flores, F.; Halkier, B.A. Biosynthesis of glucosinolates—Gene discovery and beyond. Trends Plant Sci. 2010, 15, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Lou, P.; Zhao, J.; He, H.; Hanhart, C.; Del Carpio, D.P.; Verkerk, R.; Custers, J.; Koornneef, M.; Bonnema, G. Quantitative trait loci for glucosinolate accumulation in Brassica rapa leaves. N. Phytol. 2008, 179, 1017–1032. [Google Scholar] [CrossRef]
- Harun, S.; Abdullah-Zawawi, M.R.; Goh, H.H.; Mohamed-Hussein, Z.A. A Comprehensive Gene Inventory for Glucosinolate Biosynthetic Pathway in Arabidopsis thaliana. J. Agric. Food Chem. 2020, 68, 7281–7297. [Google Scholar] [CrossRef]
- Chhajed, S.; Mostafa, I.; He, Y.; Abou-Hashem, M.; El-Domiaty, M.; Chen, S. Glucosinolate Biosynthesis and the Glucosinolate–Myrosinase System in Plant Defense. Agronomy 2020, 10, 1786. [Google Scholar] [CrossRef]
- Geu-Flores, F.; Nielsen, M.T.; Nafisi, M.; Moldrup, M.E.; Olsen, C.E.; Motawia, M.S.; Halkier, B.A. Glucosinolate engineering identifies a gamma-glutamyl peptidase. Nat. Chem. Biol. 2009, 5, 575–577. [Google Scholar] [CrossRef]
- Wang, C.; Crocoll, C.; Agerbirk, N.; Halkier, B.A. Engineering and optimization of the 2-phenylethylglucosinolate production in Nicotiana benthamiana by combining biosynthetic genes from Barbarea vulgaris and Arabidopsis thaliana. Plant J. 2021, 106, 978–992. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Dissing, M.M.; Agerbirk, N.; Crocoll, C.; Halkier, B.A. Characterization of Arabidopsis CYP79C1 and CYP79C2 by Glucosinolate Pathway Engineering in Nicotiana benthamiana Shows Substrate Specificity Toward a Range of Aliphatic and Aromatic Amino Acids. Front. Plant Sci. 2020, 11, 57. [Google Scholar] [CrossRef] [PubMed]
- Wittstock, U.; Halkier, B.A. Cytochrome P450 CYP79A2 from Arabidopsis thaliana L. Catalyzes the conversion of L-phenylalanine to phenylacetaldoxime in the biosynthesis of benzylglucosinolate. J. Biol. Chem. 2000, 275, 14659–14666. [Google Scholar] [CrossRef] [Green Version]
- Augustine, R.; Mukhopadhyay, A.; Bisht, N.C. Targeted silencing of BjMYB28 transcription factor gene directs development of low glucosinolate lines in oilseed Brassica juncea. Plant Biotechnol. J. 2013, 11, 855–866. [Google Scholar] [CrossRef] [PubMed]
- Traka, M.H.; Saha, S.; Huseby, S.; Kopriva, S.; Walley, P.G.; Barker, G.C.; Moore, J.; Mero, G.; van den Bosch, F.; Constant, H.; et al. Genetic regulation of glucoraphanin accumulation in Beneforte broccoli. N. Phytol. 2013, 198, 1085–1095. [Google Scholar] [CrossRef] [Green Version]
- Zuluaga, D.L.; Graham, N.S.; Klinder, A.; van Ommen Kloeke, A.E.E.; Marcotrigiano, A.R.; Wagstaff, C.; Verkerk, R.; Sonnante, G.; Aarts, M.G.M. Overexpression of the MYB29 transcription factor affects aliphatic glucosinolate synthesis in Brassica oleracea. Plant Mol. Biol. 2019, 101, 65–79. [Google Scholar] [CrossRef] [Green Version]
- Frerigmann, H.; Gigolashvili, T. MYB34, MYB51, and MYB122 distinctly regulate indolic glucosinolate biosynthesis in Arabidopsis thaliana. Mol. Plant 2014, 7, 814–828. [Google Scholar] [CrossRef] [Green Version]
- Zeng, X.; Yuan, H.; Dong, X.; Peng, M.; Jing, X.; Xu, Q.; Tang, T.; Wang, Y.; Zha, S.; Gao, M.; et al. Genome-wide Dissection of Co-selected UV-B Responsive Pathways in the UV-B Adaptation of Qingke. Mol. Plant 2020, 13, 112–127. [Google Scholar] [CrossRef] [Green Version]
- Fang, K.; Xia, Z.; Li, H.; Jiang, X.; Qin, D.; Wang, Q.; Wang, Q.; Pan, C.; Li, B.; Wu, H. Genome-wide association analysis identified molecular markers associated with important tea flavor-related metabolites. Hortic. Res. 2021, 8, 42. [Google Scholar] [CrossRef]
- Andini, S.; Araya-Cloutier, C.; Sanders, M.; Vincken, J.P. Simultaneous Analysis of Glucosinolates and Isothiocyanates by Reversed-Phase Ultra-High-Performance Liquid Chromatography-Electron Spray Ionization-Tandem Mass Spectrometry. J. Agric. Food Chem. 2020, 68, 3121–3131. [Google Scholar] [CrossRef] [Green Version]
- Li, Z.; Zheng, S.; Liu, Y.; Fang, Z.; Yang, L.; Zhuang, M.; Zhang, Y.; Lv, H.; Wang, Y.; Xu, D. Characterization of glucosinolates in 80 broccoli genotypes and different organs using UHPLC-Triple-TOF-MS method. Food Chem. 2021, 334, 127519. [Google Scholar] [CrossRef] [PubMed]
- Agerbirk, N.; Olsen, C.E. Glucosinolate structures in evolution. Phytochemistry 2012, 77, 16–45. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Huang, H.; Yi, X.; Zhang, Y.; Yang, Q.; Zhang, C.; Fan, C.; Zhou, Y. Dissection of genetic architecture for glucosinolate accumulations in leaves and seeds of Brassica napus by genome-wide association study. Plant Biotechnol. J. 2020, 18, 1472–1484. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, B.; Wu, Z.; Li, Z.; Zhang, Q.; Hu, J.; Xiao, Y.; Cai, D.; Wu, J.; King, G.J.; Li, H.; et al. Dissection of the genetic architecture of three seed-quality traits and consequences for breeding in Brassica napus. Plant Biotechnol. J. 2018, 16, 1336–1348. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Liu, T.; Duan, M.; Song, J.; Li, X. De novo Transcriptome Analysis of Sinapis alba in Revealing the Glucosinolate and Phytochelatin Pathways. Front. Plant Sci. 2016, 7, 259. [Google Scholar] [CrossRef] [Green Version]
- Naur, P.; Petersen, B.L.; Mikkelsen, M.D.; Bak, S.; Rasmussen, H.; Olsen, C.E.; Halkier, B.A. CYP83A1 and CYP83B1, two nonredundant cytochrome P450 enzymes metabolizing oximes in the biosynthesis of glucosinolates in Arabidopsis. Plant Physiol. 2003, 133, 63–72. [Google Scholar] [CrossRef] [Green Version]
- Chen, S.; Glawischnig, E.; Jørgensen, K.; Naur, P.; Jørgensen, B.; Olsen, C.-E.; Hansen1, C.H.; Rasmussen, H.; Pickett, J.A.; Halkier, B.A. CYP79F1 and CYP79F2 have distinct functions in the biosynthesis of aliphatic glucosinolates in Arabidopsis. Plant J. 2003, 33, 923–937. [Google Scholar] [CrossRef] [Green Version]
- Kittipol, V.; He, Z.; Wang, L.; Doheny-Adams, T.; Langer, S.; Bancroft, I. Genetic architecture of glucosinolate variation in Brassica napus. J. Plant Physiol. 2019, 240, 152988. [Google Scholar] [CrossRef]
- Wentzell, A.M.; Rowe, H.C.; Hansen, B.G.; Ticconi, C.; Halkier, B.A.; Kliebenstein, D.J. Linking metabolic QTLs with network and cis-eQTLs controlling biosynthetic pathways. PLoS Genet. 2007, 3, e162. [Google Scholar] [CrossRef] [Green Version]
- Mikkelsen, M.D.; Naur, P.; Halkier, B.A. Arabidopsis mutants in the C-S lyase of glucosinolate biosynthesis establish a critical role for indole-3-acetaldoxime in auxin homeostasis. Plant J. 2004, 37, 770–777. [Google Scholar] [CrossRef]
- Grubb, C.D.; Zipp, B.J.; Ludwig-Muller, J.; Masuno, M.N.; Molinski, T.F.; Abel, S. Arabidopsis glucosyltransferase UGT74B1 functions in glucosinolate biosynthesis and auxin homeostasis. Plant J. 2004, 40, 893–908. [Google Scholar] [CrossRef] [PubMed]
- Piotrowski, M.; Schemenewitz, A.; Lopukhina, A.; Muller, A.; Janowitz, T.; Weiler, E.W.; Oecking, C. Desulfoglucosinolate sulfotransferases from Arabidopsis thaliana catalyze the final step in the biosynthesis of the glucosinolate core structure. J. Biol. Chem. 2004, 279, 50717–50725. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Klein, M.; Reichelt, M.; Gershenzon, J.; Papenbrock, J. The three desulfoglucosinolate sulfotransferase proteins in Arabidopsis have different substrate specificities and are differentially expressed. FEBS J. 2006, 273, 122–136. [Google Scholar] [CrossRef] [PubMed]
- Knill, T.; Schuster, J.; Reichelt, M.; Gershenzon, J.; Binder, S. Arabidopsis branched-chain aminotransferase 3 functions in both amino acid and glucosinolate biosynthesis. Plant Physiol. 2008, 146, 1028–1039. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gigolashvili, T.; Yatusevich, R.; Rollwitz, I.; Humphry, M.; Gershenzon, J.; Flugge, U.I. The plastidic bile acid transporter 5 is required for the biosynthesis of methionine-derived glucosinolates in Arabidopsis thaliana. Plant Cell 2009, 21, 1813–1829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knill, T.; Reichelt, M.; Paetz, C.; Gershenzon, J.; Binder, S. Arabidopsis thaliana encodes a bacterial-type heterodimeric isopropylmalate isomerase involved in both Leu biosynthesis and the Met chain elongation pathway of glucosinolate formation. Plant Mol. Biol. 2009, 71, 227–239. [Google Scholar] [CrossRef] [Green Version]
- Sawada, Y.; Kuwahara, A.; Nagano, M.; Narisawa, T.; Sakata, A.; Saito, K.; Hirai, M.Y. Omics-based approaches to methionine side chain elongation in Arabidopsis: Characterization of the genes encoding methylthioalkylmalate isomerase and methylthioalkylmalate dehydrogenase. Plant Cell Physiol. 2009, 50, 1181–1190. [Google Scholar] [CrossRef] [Green Version]
- He, Y.; Mawhinney, T.P.; Preuss, M.L.; Schroeder, A.C.; Chen, B.; Abraham, L.; Jez, J.M.; Chen, S. A redox-active isopropylmalate dehydrogenase functions in the biosynthesis of glucosinolates and leucine in Arabidopsis. Plant J. 2009, 60, 679–690. [Google Scholar] [CrossRef]
- Liu, T.; Zhang, X.; Yang, H.; Agerbirk, N.; Qiu, Y.; Wang, H.; Shen, D.; Song, J.; Li, X. Aromatic Glucosinolate Biosynthesis Pathway in Barbarea vulgaris and its Response to Plutella xylostella Infestation. Front. Plant Sci. 2016, 7, 83. [Google Scholar] [CrossRef] [Green Version]
- Cai, C.; Yuan, W.; Miao, H.; Deng, M.; Wang, M.; Lin, J.; Zeng, W.; Wang, Q. Functional Characterization of BoaMYB51s as Central Regulators of Indole Glucosinolate Biosynthesis in Brassica oleracea var. alboglabra Bailey. Front. Plant Sci. 2018, 9, 1599. [Google Scholar] [CrossRef]
- Yu, Q.; Hao, G.; Zhou, J.; Wang, J.; Evivie, E.R.; Li, J. Identification and expression pattern analysis of BoMYB51 involved in indolic glucosinolate biosynthesis from broccoli (Brassica oleracea var. italica). Biochem. Biophys. Res. Commun. 2018, 501, 598–604. [Google Scholar] [CrossRef] [PubMed]
- Araki, R.; Hasumi, A.; Nishizawa, O.I.; Sasaki, K.; Kuwahara, A.; Sawada, Y.; Totoki, Y.; Toyoda, A.; Sakaki, Y.; Li, Y.; et al. Novel bioresources for studies of Brassica oleracea: Identification of a kale MYB transcription factor responsible for glucosinolate production. Plant Biotechnol. J. 2013, 11, 1017–1027. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Long, Y.; Shi, L.; Shi, J.; Barker, G.; Meng, J. Characterization of metabolite quantitative trait loci and metabolic networks that control glucosinolate concentration in the seeds and leaves of Brassica napus. N. Phytol. 2012, 193, 96–108. [Google Scholar] [CrossRef] [PubMed]
- Qu, C.; Yin, N.; Chen, S.; Wang, S.; Chen, X.; Zhao, H.; Shen, S.; Fu, F.; Zhou, B.; Xu, X.; et al. Comparative Analysis of the Metabolic Profiles of Yellow-versus Black-Seeded Rapeseed Using UPLC-HESI-MS/MS and Transcriptome Analysis. J. Agric. Food Chem. 2020, 68, 3033–3049. [Google Scholar] [CrossRef]
- Yin, N.W.; Wang, S.X.; Jia, L.D.; Zhu, M.C.; Yang, J.; Zhou, B.J.; Yin, J.M.; Lu, K.; Wang, R.; Li, J.N.; et al. Identification and Characterization of Major Constituents in Different-Colored Rapeseed Petals by UPLC-HESI-MS/MS. J. Agric. Food Chem. 2019, 67, 11053–11065. [Google Scholar] [CrossRef]
- Tsugawa, H.; Ikeda, K.; Takahashi, M.; Satoh, A.; Mori, Y.; Uchino, H.; Okahashi, N.; Yamada, Y.; Tada, I.; Bonini, P.; et al. MS-DIAL 4: Accelerating lipidomics using an MS/MS, CCS, and retention time atlas. bioRxiv 2020. [Google Scholar] [CrossRef]
- Millan, S.; Sampedro, M.C.; Gallejones, P.; Castellon, A.; Ibargoitia, M.L.; Goicolea, M.A.; Barrio, R.J. Identification and quantification of glucosinolates in rapeseed using liquid chromatography-ion trap mass spectrometry. Anal. Bioanal. Chem. 2009, 394, 1661–1669. [Google Scholar] [CrossRef]
- Hwang, I.M.; Park, B.; Dang, Y.M.; Kim, S.-Y.; Seo, H.Y. Simultaneous direct determination of 15 glucosinolates in eight Brassica species by UHPLC-Q-Orbitrap-MS. Food Chem. 2019, 282, 127–133. [Google Scholar] [CrossRef]
- Dong, M.; Tian, Z.; Ma, Y.; Yang, Z.; Ma, Z.; Wang, X.; Li, Y.; Jiang, H. Rapid screening and characterization of glucosinolates in 25 Brassicaceae tissues by UHPLC-Q-exactive orbitrap-MS. Food Chem. 2021, 365, 130493. [Google Scholar] [CrossRef]
- Zhang, L.; Cai, X.; Wu, J.; Liu, M.; Grob, S.; Cheng, F.; Liang, J.; Cai, C.; Liu, Z.; Liu, B.; et al. Improved Brassica rapa reference genome by single-molecule sequencing and chromosome conformation capture technologies. Hortic. Res. 2018, 5, 50. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [Green Version]
- Liao, Y.; Smyth, G.K.; Shi, W. The Subread aligner: Fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 2013, 41, e108. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of Population Structure Using Multilocus Genotype Data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef]
- Zhang, Y.W.; Tamba, C.L.; Wen, Y.J.; Li, P.; Ren, W.L.; Ni, Y.L.; Gao, J.; Zhang, Y.M. mrMLM v4.0.2: An R Platform for Multi-locus Genome-wide Association Studies. Genom. Proteom. Bioinform. 2020, 18, 481–487. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Franceschini, A.; Kuhn, M.; Simonovic, M.; Roth, A.; Minguez, P.; Doerks, T.; Stark, M.; Muller, J.; Bork, P.; et al. The STRING database in 2011: Functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 2011, 39, D561–D568. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shang, G.; Zhao, H.; Tong, L.; Yin, N.; Hu, R.; Jiang, H.; Kamal, F.; Zhao, Z.; Xu, L.; Lu, K.; et al. Genome-Wide Association Study of Phenylalanine Derived Glucosinolates in Brassica rapa. Plants 2022, 11, 1274. https://doi.org/10.3390/plants11091274
Shang G, Zhao H, Tong L, Yin N, Hu R, Jiang H, Kamal F, Zhao Z, Xu L, Lu K, et al. Genome-Wide Association Study of Phenylalanine Derived Glucosinolates in Brassica rapa. Plants. 2022; 11(9):1274. https://doi.org/10.3390/plants11091274
Chicago/Turabian StyleShang, Guoxia, Huiyan Zhao, Linhui Tong, Nengwen Yin, Ran Hu, Haiyan Jiang, Farah Kamal, Zhi Zhao, Liang Xu, Kun Lu, and et al. 2022. "Genome-Wide Association Study of Phenylalanine Derived Glucosinolates in Brassica rapa" Plants 11, no. 9: 1274. https://doi.org/10.3390/plants11091274
APA StyleShang, G., Zhao, H., Tong, L., Yin, N., Hu, R., Jiang, H., Kamal, F., Zhao, Z., Xu, L., Lu, K., Li, J., Qu, C., & Du, D. (2022). Genome-Wide Association Study of Phenylalanine Derived Glucosinolates in Brassica rapa. Plants, 11(9), 1274. https://doi.org/10.3390/plants11091274







