Characterization of the TCP Gene Family in Chrysanthemum nankingense and the Role of CnTCP4 in Cold Tolerance
Abstract
:1. Introduction
2. Results
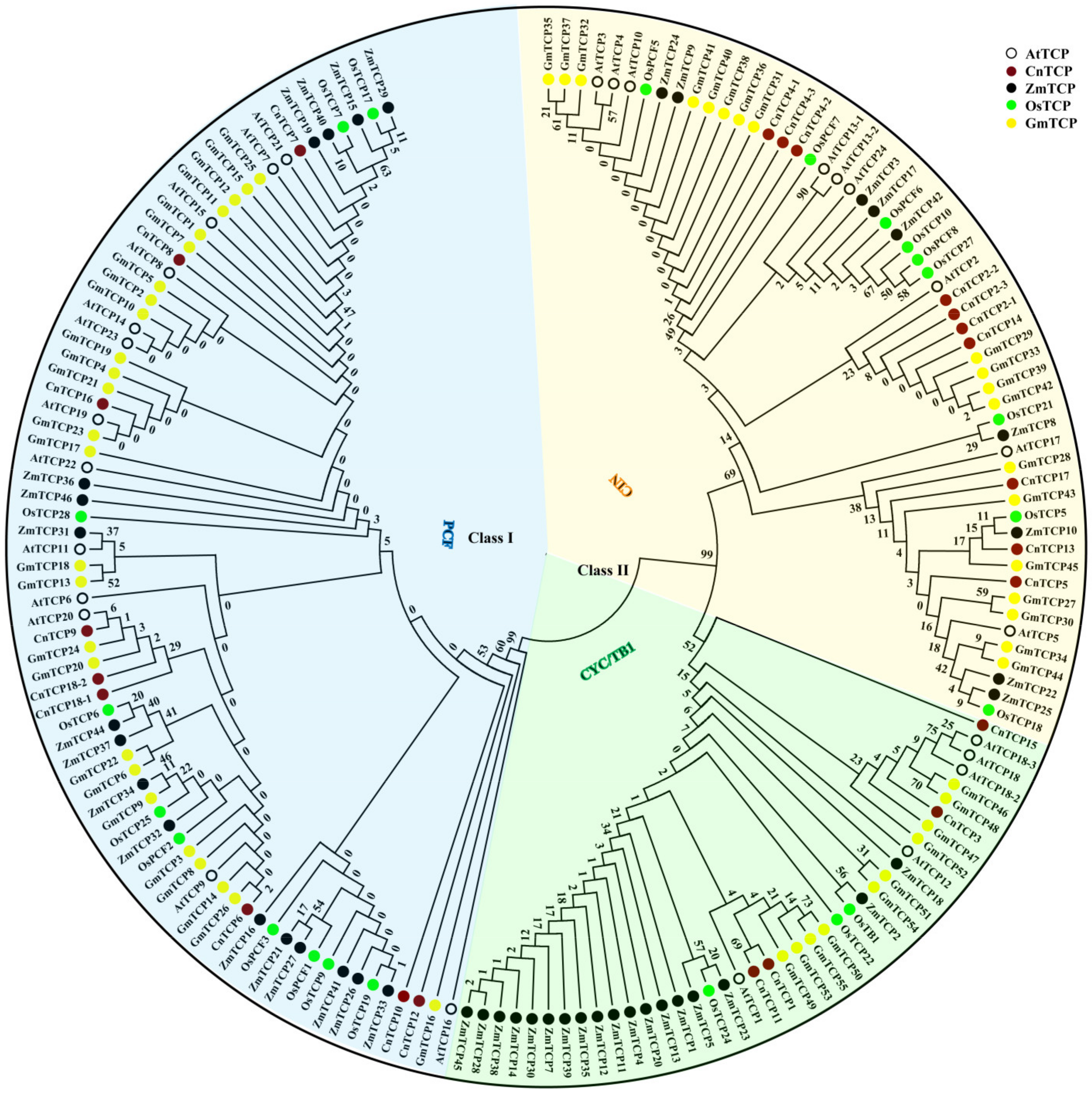
2.1. Classification and Phylogenetic Analysis of TCP Proteins
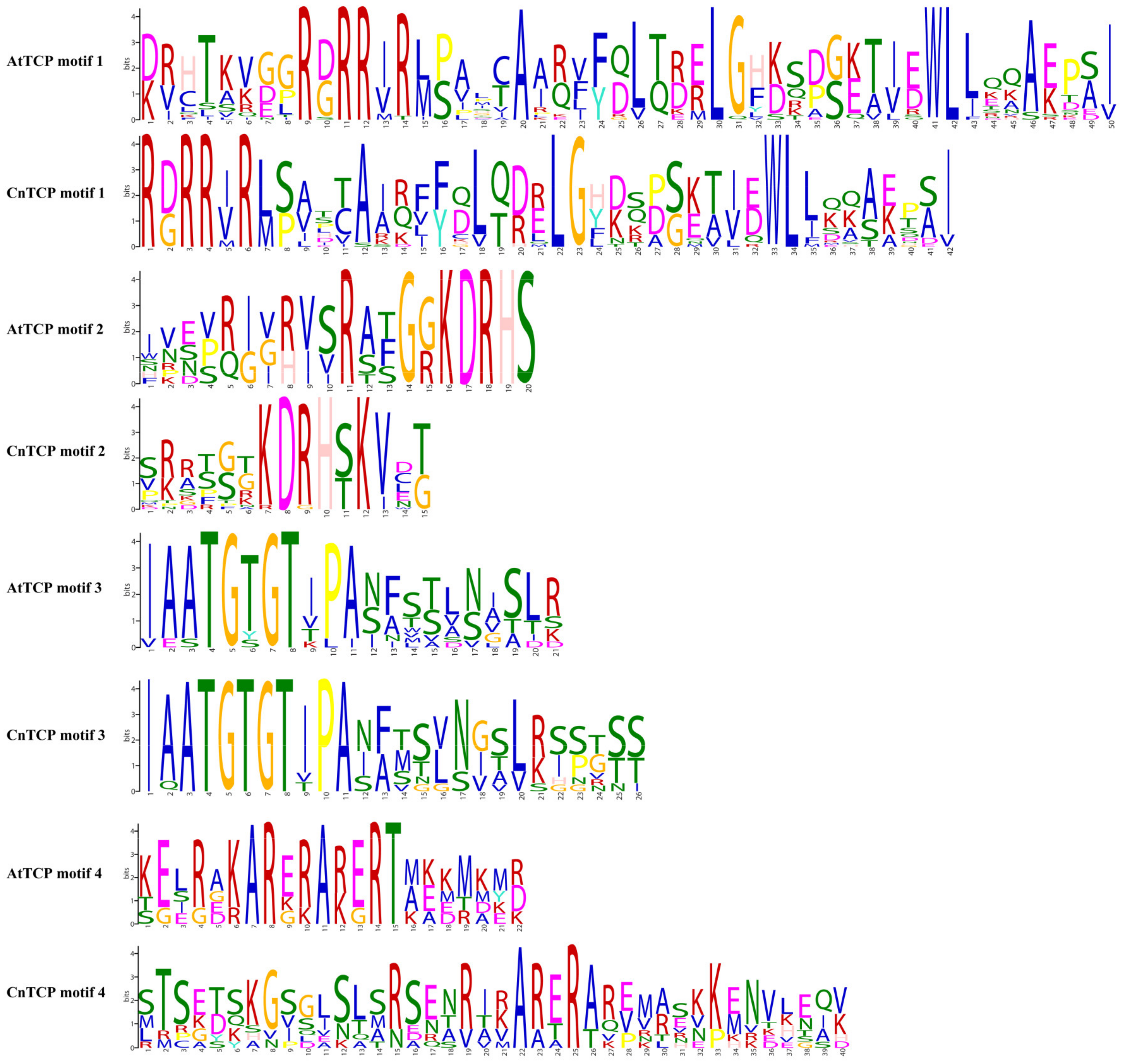
2.2. Structural Analysis and Protein–Protein Interactions of CnTCP Genes
2.3. Expression Profiling of CnTCP Genes in Response to Cold Stress
2.4. CnTCP4 Transcription Factors Are Involved in Cold Stress Response
3. Discussion
4. Materials and Methods
4.1. Phylogenetic Analysis of the CnTCP Family
4.2. Structural Analysis of the TCP Genes
4.3. Protein–Protein Interaction Network Analysis
4.4. Putative Cis-Elements in the Promoter Regions
4.5. Plant Growth Conditions and Cold Treatment
4.6. Verification of CnTCP Expression
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cubas, P.; Lauter, N.; Doebley, J.; Coen, E. The TCP Domain: A Motif Found in Proteins Regulating Plant Growth and Development. Plant J. 1999, 18, 215–222. [Google Scholar] [CrossRef] [Green Version]
- Kosugi, S.; Ohashi, Y. PCF1 and PCF2 Specifically Bind to cis Elements in the Rice Proliferating Cell Nuclear Antigen Gene. Plant Cell 1997, 9, 1607–1619. [Google Scholar] [CrossRef] [Green Version]
- Doebley, J.; Stec, A.; Hubbard, L. The Evolution of Apical Dominance in Maize. Nature 1997, 386, 485–488. [Google Scholar] [CrossRef]
- Luo, D.; Carpenter, R.; Vincent, C.; Copsey, L.; Coen, E. Origin of Floral Asymmetry in Antirrhinum. Nature 1996, 383, 794–799. [Google Scholar] [CrossRef]
- Martín-Trillo, M.; Cubas, P. TCP Genes: A Family Snapshot Ten Years Later. Trends Plant Sci. 2010, 15, 31–39. [Google Scholar] [CrossRef]
- Manassero, N.G.U.; Viola, I.L.; Welchen, E.; Gonzalez, D.H. TCP Transcription Factors: Architectures of Plant Form. Biomol. Concepts 2013, 4, 111–127. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.; Ma, H.; Wang, J.; Zhang, D. Genome-Wide Comparative Analysis and Expression Pattern of TCP Gene Families in Arabidopsis thaliana and Oryza sativa. J. Integr. Plant Biol. 2007, 49, 885–897. [Google Scholar] [CrossRef]
- Ling, L.; Zhang, W.; An, Y.; Du, B.; Wang, D.; Guo, C. Genome-Wide Analysis of the TCP Transcription Factor Genes in Five Legume Genomes and Their Response to Salt and Drought Stresses. Funct. Integr. Genom. 2020, 20, 537–550. [Google Scholar] [CrossRef]
- Ding, S.; Cai, Z.; Du, H.; Wang, H. Genome-Wide Analysis of TCP Family Genes in Zea mays L. Identified a Role for ZmTCP42 in Drought Tolerance. Int. J. Mol. Sci. 2019, 20, 2762. [Google Scholar] [CrossRef] [Green Version]
- Navaud, O.; Dabos, P.; Carnus, E.; Tremousaygue, D.; Hervé, C. TCP Transcription Factors Predate the Emergence of Land Plants. J. Mol. Evol. 2007, 65, 23–33. [Google Scholar] [CrossRef]
- Floyd, S.K.; Bowman, J.L. The Ancestral Developmental Tool Kit of Land Plants. Int. J. Plant Sci. 2007, 168, 1–35. [Google Scholar] [CrossRef]
- Ma, X.; Ma, J.; Fan, D.; Li, C.; Jiang, Y.; Luo, K. Genome-Wide Identification of TCP Family Transcription Factors from Populus euphratica and Their Involvement in Leaf Shape Regulation. Sci. Rep. 2016, 6, 32795. [Google Scholar] [CrossRef]
- Leng, X.; Wei, H.; Xu, X.; Ghuge, S.A.; Jia, D.; Liu, G.; Wang, Y.; Yuan, Y. Genome-Wide Identification and Transcript Analysis of TCP Transcription Factors in Grapevine. BMC Genom. 2019, 20, 786. [Google Scholar] [CrossRef] [Green Version]
- Tatematsu, K.; Nakabayashi, K.; Kamiya, Y.; Nambara, E. Transcription Factor AtTCP14 Regulates Embryonic Growth Potential During Seed Germination in Arabidopsis thaliana. Plant J. 2008, 53, 42–52. [Google Scholar] [CrossRef]
- Li, Z.Y.; Li, B.; Dong, A.W. The Arabidopsis Transcription Factor AtTCP15 Regulates Endoreduplication by Modulating Expression of Key Cell-Cycle Genes. Mol. Plant 2012, 5, 270–280. [Google Scholar] [CrossRef]
- Li, C.; Potuschak, T.; Colón-Carmona, A.; Gutiérrez, R.A.; Doerner, P. Arabidopsis TCP20 Links Regulation of Growth and Cell Division Control Pathways. Proc. Natl. Acad. Sci. USA 2005, 102, 12978–12983. [Google Scholar] [CrossRef] [Green Version]
- Takeda, T.; Amano, K.; Ohto, M.A.; Nakamura, K.; Sato, S.; Kato, T.; Tabata, S.; Ueguchi, C. RNA Interference of the Arabidopsis Putative Transcription Factor TCP16 Gene Results in Abortion of Early Pollen Development. Plant Mol. Biol. 2006, 61, 165–177. [Google Scholar] [CrossRef]
- Rueda-Romero, P.; Barrero-Sicilia, C.; Gómez-Cadenas, A.; Carbonero, P.; Oñate-Sánchez, L. Arabidopsis thaliana DOF6 Negatively Affects Germination in non-After-Ripened Seeds and Interacts with TCP14. J. Exp. Bot. 2012, 63, 1937–1949. [Google Scholar] [CrossRef] [Green Version]
- Schommer, C.; Palatnik, J.F.; Aggarwal, P.; Chételat, A.; Cubas, P.; Farmer, E.E.; Nath, U.; Weigel, D. Control of Jasmonate Biosynthesis and Senescence by miR319 Targets. PLoS Biol. 2008, 6, e230. [Google Scholar] [CrossRef] [Green Version]
- Giraud, E.; Ng, S.; Carrie, C.; Duncan, O.; Low, J.; Lee, C.P.; Van Aken, O.; Millar, A.H.; Murcha, M.; Whelan, J. TCP Transcription Factors Link the Regulation of Genes Encoding Mitochondrial Proteins with the Circadian Clock in Arabidopsis thaliana. Plant Cell 2010, 22, 3921–3934. [Google Scholar] [CrossRef] [Green Version]
- Pruneda-Paz, J.L.; Kay, S.A. An Expanding Universe of Circadian Networks in Higher Plants. Trends Plant Sci. 2010, 15, 259–265. [Google Scholar] [CrossRef] [Green Version]
- Qi, X.; Qu, Y.; Jiang, J.; Guan, Y.; Song, A.; Cao, P.; Guan, Z.; Fang, W.; Chen, S.; Chen, F.; et al. Heterologous Expression of Chrysanthemum nankingense TCP13 Suppresses Leaf Development in Arabidopsis thaliana. Plant Growth Regul. 2021, 95, 331–341. [Google Scholar] [CrossRef]
- Danisman, S.; van der Wal, F.; Dhondt, S.; Waites, R.; de Folter, S.; Bimbo, A.; van Dijk, A.D.; Muino, J.M.; Cutri, L.; Dornelas, M.C.; et al. Arabidopsis Class I and class II TCP Transcription Factors Regulate Jasmonic Acid Metabolism and Leaf Development Antagonistically. Plant Physiol. 2012, 159, 1511–1523. [Google Scholar] [CrossRef] [Green Version]
- Aguilar-Martínez, J.A.; Sinha, N. Analysis of the Role of Arabidopsis class I TCP Genes AtTCP7, AtTCP8, AtTCP22, and AtTCP23 in Leaf Development. Front. Plant Sci. 2013, 4, 406. [Google Scholar] [CrossRef] [Green Version]
- Ori, N.; Cohen, A.R.; Etzioni, A.; Brand, A.; Yanai, O.; Shleizer, S.; Menda, N.; Amsellem, Z.; Efroni, I.; Pekker, I.; et al. Regulation of LANCEOLATE by miR319 Is Required for Compound-Leaf Development in Tomato. Nat. Genet. 2007, 39, 787–791. [Google Scholar] [CrossRef]
- Camoirano, A.; Arce, A.L.; Ariel, F.D.; Alem, A.L.; Gonzalez, D.H.; Viola, I.L. Class I TCP Transcription Factors Regulate Trichome Branching and Cuticle Development in Arabidopsis. J. Exp. Bot. 2020, 71, 5438–5453. [Google Scholar] [CrossRef]
- Aguilar-Martínez, J.A.; Poza-Carrión, C.; Cubas, P. Arabidopsis BRANCHED1 Acts as an Integrator of Branching Signals Within Axillary Buds. Plant Cell. 2007, 19, 458–472. [Google Scholar] [CrossRef]
- Takeda, T.; Suwa, Y.; Suzuki, M.; Kitano, H.; Ueguchi-Tanaka, M.; Ashikari, M.; Matsuoka, M.; Ueguchi, C. The OsTB1 Gene Negatively Regulates Lateral Branching in Rice. Plant J. 2003, 33, 513–520. [Google Scholar] [CrossRef]
- Narumi, T.; Aida, R.; Koyama, T.; Yamaguchi, H.; Sasaki, K.; Shikata, M.; Nakayama, M.; Ohme-Takagi, M.; Ohtsubo, N. Arabidopsis Chimeric TCP3 Repressor Produces Novel Floral Traits in Torenia fournieri and Chrysanthemum morifolium. Plant Biotechnol. 2011, 28, 131–140. [Google Scholar] [CrossRef] [Green Version]
- Li, D.B.; Zhang, H.; Mou, M.; Chen, Y.; Xiang, S.; Chen, L.; Yu, D. Arabidopsis class II TCP Transcription Factors Integrate with the FT–FD Module to Control Flowering. Plant Physiol. 2019, 181, 97–111. [Google Scholar] [CrossRef]
- Kubota, A.; Ito, S.; Shim, J.S.; Johnson, R.S.; Song, Y.H.; Breton, G.; Goralogia, G.S.; Kwon, M.S.; Laboy Cintrón, D.; Koyama, T.; et al. TCP4-Dependent Induction of CONSTANS Transcription Requires GIGANTEA in Photoperiodic Flowering in Arabidopsis. PLoS Genet. 2017, 13, e1006856. [Google Scholar] [CrossRef] [PubMed]
- Lucero, L.E.; Manavella, P.A.; Gras, D.E.; Ariel, F.D.; Gonzalez, D.H. Class Ⅰ and class Ⅱ TCP Transcription Factors Modulate SOC1-Dependent Flowering at Multiple Levels. Mol. Plant 2017, 10, 1571–1574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, J.; Cheng, X.; Liu, P.; Li, D.; Chen, T.; Gu, X.; Sun, J. MicroRNA319-Regulated TCPs Interact with FBHs and PFT1 to Activate CO Transcription and Control Flowering Time in Arabidopsis. PLoS Genet. 2017, 13, e1006833. [Google Scholar] [CrossRef] [Green Version]
- Davière, J.M.; Wild, M.; Regnault, T.; Baumberger, N.; Eisler, H.; Genschik, P.; Achard, P. Class I TCP-DELLA Interactions in Inflorescence Shoot Apex Determine Plant Height. Curr. Biol. 2014, 24, 1923–1928. [Google Scholar] [CrossRef] [Green Version]
- Sarvepalli, K.; Nath, U. Hyper-Activation of the TCP4 Transcription Factor in Arabidopsis thaliana Accelerates Multiple Aspects of Plant Maturation. Plant J. 2011, 67, 595–607. [Google Scholar] [CrossRef] [Green Version]
- Sarvepalli, K.; Nath, U. Interaction of TCP4-Mediated Growth Module with Phytohormones. Plant Signal. Behav. 2011, 6, 1440–1443. [Google Scholar] [CrossRef] [Green Version]
- Hao, J.; Tu, L.; Hu, H.; Tan, J.; Deng, F.; Tang, W.; Nie, Y.; Zhang, X. GbTCP, a Cotton TCP Transcription Factor, Confers Fibre Elongation and Root Hair Development by a Complex Regulating System. J. Exp. Bot. 2012, 63, 6267–6281. [Google Scholar] [CrossRef] [Green Version]
- Koyama, T.; Furutani, M.; Tasaka, M.; Ohme-Takagi, M. TCP Transcription Factors Control the Morphology of Shoot Lateral Organs via Negative Regulation of the Expression of Boundary-Specific Genes in Arabidopsis. Plant Cell 2007, 19, 473–484. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.T.; Sun, X.L.; Hoshino, Y.; Yu, Y.; Jia, B.; Sun, Z.W.; Sun, M.Z.; Duan, X.B.; Zhu, Y.M. MicroRNA319 Positively Regulates Cold Tolerance by Targeting OsPCF6 and OsTCP21 in Rice (Oryza sativa L.). PLoS ONE 2014, 9, e91357. [Google Scholar] [CrossRef] [Green Version]
- Kosugi, S.; Ohashi, Y. DNA Binding and Dimerization Specificity and Potential Targets for the TCP Protein Family. Plant J. 2002, 30, 337–348. [Google Scholar] [CrossRef]
- Viola, I.L.; Uberti Manassero, N.; Ripoll, R.; Gonzalez, D. The Arabidopsis class I TCP Transcription Factor AtTCP11 Is a Developmental Regulator with Distinct DNA-Binding Properties due to the Presence of a Threonine Residue at position 15 of the TCP Domain. Biochem. J. 2011, 435, 143–155. [Google Scholar] [CrossRef] [Green Version]
- Parapunova, V.; Busscher, M.; Busscher-Lange, J.; Lammers, M.; Karlova, R.; Bovy, A.G.; Angenent, G.C.; de Maagd, R.A. Identification, Cloning and Characterization of the Tomato TCP Transcription Factor Family. BMC Plant Biol. 2014, 14, 157. [Google Scholar] [CrossRef] [Green Version]
- Viola, I.L.; Reinheimer, R.; Ripoll, R.; Manassero, N.G.U.; Gonzalez, D.H. Determinants of the DNA Binding Specificity of Class I and class II TCP Transcription Factors. J. Biol. Chem. 2012, 287, 347–356. [Google Scholar] [CrossRef] [Green Version]
- Tähtiharju, S.; Rijpkema, A.S.; Vetterli, A.; Albert, V.A.; Teeri, T.H.; Elomaa, P. Evolution and Diversification of the CYC/TB1 Gene Family in Asteraceae—A Comparative Study in Gerbera (Mutisieae) and Sunflower (Heliantheae). Mol. Biol. Evol. 2012, 29, 1155–1166. [Google Scholar] [CrossRef] [Green Version]
- Danisman, S.; van Dijk, A.D.; Bimbo, A.; van der Wal, F.; Hennig, L.; de Folter, S.; Angenent, G.C.; Immink, R.G. Analysis of Functional Redundancies Within the Arabidopsis TCP Transcription Factor Family. J. Exp. Bot. 2013, 64, 5673–5685. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Guan, Y.; Ding, L.; Li, P.; Zhao, W.; Jiang, J.; Chen, S.; Chen, F. The CmTCP20 Gene Regulates Petal Elongation Growth in Chrysanthemum morifolium. Plant Sci. 2019, 280, 248–257. [Google Scholar] [CrossRef]
- Gull, A.; Lone, A.A.; Wani, N. Biotic and Abiotic Stresses in Plants. In Abiotic Biotic Stress Plants; InTech Open: London, UK, 2019. [Google Scholar]
- Chinnusamy, V.; Zhu, J.; Zhu, J.K. Cold Stress Regulation of Gene Expression in Plants. Trends Plant Sci. 2007, 12, 444–451. [Google Scholar] [CrossRef]
- Lissarre, M.; Ohta, M.; Sato, A.; Miura, K. Cold-Responsive Gene Regulation During Cold Acclimation in Plants. Plant Signal. Behav. 2010, 5, 948–952. [Google Scholar] [CrossRef] [Green Version]
- Zhao, C.; Lang, Z.; Zhu, J.K. Cold Responsive Gene Transcription Becomes More Complex. Trends Plant Sci. 2015, 20, 466–468. [Google Scholar] [CrossRef] [Green Version]
- Shi, Y.; Ding, Y.; Yang, S. Molecular Regulation of CBF Signaling in Cold Acclimation. Trends Plant Sci. 2018, 23, 623–637. [Google Scholar] [CrossRef]
- Dong, C.H.; Agarwal, M.; Zhang, Y.; Xie, Q.; Zhu, J.K. The Negative Regulator of Plant Cold Responses, HOS1, is a RING E3 Ligase That Mediates the Ubiquitination and Degradation of ICE1. Proc. Natl. Acad. Sci. USA 2006, 103, 8281–8286. [Google Scholar] [CrossRef] [Green Version]
- Bergonzi, S.; Albani, M.C.; Ver Loren van Themaat, E.; Nordström, K.J.; Wang, R.; Schneeberger, K.; Moerland, P.D.; Coupland, G. Mechanisms of Age-Dependent Response to Winter Temperature in Perennial Flowering of Arabis alpina. Science 2013, 340, 1094–1097. [Google Scholar] [CrossRef]
- Yang, C.; Li, D.; Mao, D.; Liu, X.; Ji, C.; Li, X.; Zhao, X.; Cheng, Z.; Chen, C.; Zhu, L. Overexpression of microRNA319 Impacts Leaf Morphogenesis and Leads to Enhanced Cold Tolerance in Rice (Oryza sativa L.). Plant Cell Environ. 2013, 36, 2207–2218. [Google Scholar] [CrossRef]
- Thiebaut, F.; Rojas, C.A.; Almeida, K.L.; Grativol, C.; Domiciano, G.C.; Lamb, C.R.; Engler, J.A.; Hemerly, A.S.; Ferreira, P.C. Regulation of miR319 During Cold Stress in Sugarcane. Plant Cell Environ. 2012, 35, 502–512. [Google Scholar] [CrossRef]
- Capovilla, G.; Pajoro, A.; Immink, R.G.; Schmid, M. Role of Alternative Pre-mRNA Splicing in Temperature Signaling. Curr. Opin. Plant Biol. 2015, 27, 97–103. [Google Scholar] [CrossRef]
- Ren, L.; Sun, J.; Chen, S.; Gao, J.; Dong, B.; Liu, Y.; Xia, X.; Wang, Y.; Liao, Y.; Teng, N.; et al. A Transcriptomic Analysis of Chrysanthemum nankingense Provides Insights into the Basis of Low Temperature Tolerance. BMC Genom. 2014, 15, 844. [Google Scholar] [CrossRef] [Green Version]
- Dong, B.; Wang, H.; Liu, T.; Cheng, P.; Chen, Y.; Chen, S.; Guan, Z.; Fang, W.; Jiang, J.; Chen, F. Whole Genome Duplication Enhances the Photosynthetic Capacity of Chrysanthemum nankingense. Mol. Genet. Genom. 2017, 292, 1247–1256. [Google Scholar] [CrossRef]
- Wang, H.; Jiang, J.; Chen, S.; Fang, W.; Guan, Z.; Liao, Y.; Chen, F. Rapid Genomic and Transcriptomic Alterations Induced by Wide Hybridization: Chrysanthemum nankingense × Tanacetum vulgare and C. crassum × Crossostephium chinense (Asteraceae). BMC Genom. 2013, 14, 902. [Google Scholar] [CrossRef] [Green Version]
- Qi, X.; Qu, Y.; Gao, R.; Jiang, J.; Fang, W.; Guan, Z.; Zhang, F.; Zhao, S.; Chen, S.; Chen, F.; et al. The Heterologous Expression of a Chrysanthemum nankingense TCP Transcription Factor Blocks Cell Division in Yeast and Arabidopsis thaliana. Int. J. Mol. Sci. 2019, 20, 4848. [Google Scholar] [CrossRef] [Green Version]
- Song, C.; Liu, Y.; Song, A.; Dong, G.; Zhao, H.; Sun, W.; Ramakrishnan, S.; Wang, Y.; Wang, S.; Li, T.; et al. The Chrysanthemum nankingense Genome Provides Insights into the Evolution and Diversification of Chrysanthemum Flowers and Medicinal Traits. Mol. Plant 2018, 11, 1482–1491. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Jiang, J.; Chen, S.; Qi, X.; Peng, H.; Li, P.; Song, A.; Guan, Z.; Fang, W.; Liao, Y.; et al. Next-Generation Sequencing of the Chrysanthemum nankingense (Asteraceae) Transcriptome Permits Large-Scale Unigene Assembly and SSR Marker Discovery. PLoS ONE 2013, 8, e62293. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Badouin, H.; Gouzy, J.; Grassa, C.J.; Murat, F.; Staton, S.E.; Cottret, L.; Lelandais-Brière, C.; Owens, G.L.; Carrère, S.; Mayjonade, B.; et al. The Sunflower Genome Provides Insights into Oil Metabolism, Flowering and Asterid Evolution. Nature 2017, 546, 148–152. [Google Scholar] [CrossRef] [PubMed]
- Masuda, H.P.; Cabral, L.M.; De Veylder, L.; Tanurdzic, M.; de Almeida Engler, J.; Geelen, D.; Inzé, D.; Martienssen, R.A.; Ferreira, P.C.; Hemerly, A.S. ABAP1 Is a Novel Plant Armadillo BTB Protein Involved in DNA Replication and Transcription. EMBO J. 2008, 27, 2746–2756. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, L.; Mao, Y.; Ji, Q.; Dersh, D.; Yewdell, J.W.; Qian, S.B. Decoding mRNA Translatability and Stability from the 5′ UTR. Nat. Struct. Mol. Biol. 2020, 27, 814–821. [Google Scholar] [CrossRef]
- Chung, B.Y.W.; Simons, C.; Firth, A.E.; Brown, C.M.; Hellens, R.P. Effect of 5′UTR Introns on Gene Expression in Arabidopsis thaliana. BMC Genom. 2006, 7, 120. [Google Scholar] [CrossRef] [Green Version]
- Hogg, J.R.; Goff, S.P. Upf1 Senses 3′UTR Length to Potentiate mRNA Decay. Cell 2010, 143, 379–389. [Google Scholar] [CrossRef] [Green Version]
- Shaul, O. How Introns Enhance Gene Expression. Int. J. Biochem. Cell Biol. 2017, 91, 145–155. [Google Scholar] [CrossRef]
- Singh, K.B.; Foley, R.C.; Oñate-Sánchez, L. Transcription Factors in Plant Defense and Stress Responses. Curr. Opin. Plant Biol. 2002, 5, 430–436. [Google Scholar] [CrossRef]
- Oda, A.; Narumi, T.; Li, T.; Kando, T.; Higuchi, Y.; Sumitomo, K.; Fukai, S.; Hisamatsu, T. CsFTL3, A Chrysanthemum FLOWERING LOCUS T-like Gene, is A Key Regulator of Photoperiodic Flowering in Chrysanthemums. J. Exp. Bot. 2012, 63, 1461–1477. [Google Scholar] [CrossRef] [Green Version]
- Chen, M.-D.; Wang, B.; Li, Y.-P.; Zeng, M.-J.; Liu, J.-T.; Ye, X.-R.; Zhu, H.-S.; Wen, Q.-F. Reference Gene Selection for qRT-PCR Analyses of Luffa (Luffa cylindrica) Plants under Abiotic Stress Conditions. Sci. Rep. 2021, 11, 3161. [Google Scholar] [CrossRef]
- Cai, J.; Li, P.; Luo, X.; Chang, T.; Li, J.; Zhao, Y.; Xu, Y. Selection of Appropriate Reference Genes for the Detection of Rhythmic Gene Expression via Quantitative Real-time PCR in Tibetan hulless Barley. PLoS ONE 2018, 13, e0190559. [Google Scholar] [CrossRef] [Green Version]
- Salone, V.; Rederstorff, M. Stem-Loop RT-PCR Based Quantification of Small Non-Coding RNAs. In Small Non-Coding RNAs; Rederstorff, M., Ed.; Methods in Molecular Biology Series; Springer: Cham, Switzerland, 2015; Volume 1296. [Google Scholar] [CrossRef]
- Galiveti, C.R.; Rozhdestvensky, T.S.; Brosius, J.; Lehrach, H.; Konthur, Z. Application of Housekeeping npcRNAs for Quantitative Expression Analysis of Human Transcriptome by Real-time PCR. RNA 2010, 16, 450–461. [Google Scholar] [CrossRef] [Green Version]
- Mestdagh, P.; Van Vlierberghe, P.; De Weer, A.; Muth, D.; Westermann, F.; Speleman, F.; Vandesompele, J. A Novel and Universal Method for microRNA RT-qPCR Data Normalization. Genome Biol. 2009, 10, R64. [Google Scholar] [CrossRef] [Green Version]
- Novillo, F.; Alonso, J.M.; Ecker, J.R.; Salinas, J. CBF2/DREB1C Is a Negative Regulator of CBF1/DREB1B and CBF3/DREB1A Expression and Plays a Central Role in Stress Tolerance in Arabidopsis. Proc. Natl. Acad. Sci. USA 2004, 101, 3985–3990. [Google Scholar] [CrossRef] [Green Version]
- Zhao, C.; Zhang, Z.; Xie, S.; Si, T.; Li, Y.; Zhu, J.K. Mutational Evidence for the Critical Role of CBF Transcription Factors in Cold Acclimation in Arabidopsis. Plant Physiol. 2016, 171, 2744–2759. [Google Scholar] [CrossRef] [Green Version]
- Wu, J.F.; Tsai, H.L.; Joanito, I.; Wu, Y.C.; Chang, C.W.; Li, Y.H.; Wang, Y.; Hong, J.C.; Chu, J.W.; Hsu, C.P.; et al. LWD–TCP Complex Activates the Morning Gene CCA1 in Arabidopsis. Nat. Commun. 2016, 7, 13181. [Google Scholar] [CrossRef] [Green Version]
- Moon, J.; Suh, S.S.; Lee, H.; Choi, K.R.; Hong, C.B.; Paek, N.C.; Kim, S.G.; Lee, I. The SOC1 MADS-Box Gene Integrates Vernalization and Gibberellin Signals for Flowering in Arabidopsis. Plant J. 2003, 35, 613–623. [Google Scholar] [CrossRef]
- Feng, Z.J.; Xu, S.C.; Liu, N.; Zhang, G.W.; Hu, Q.Z.; Gong, Y.M. Soybean TCP Transcription Factors: Evolution, Classification, Protein Interaction and Stress and Hormone Responsiveness. Plant Physiol. Biochem. 2018, 127, 129–142. [Google Scholar] [CrossRef]
- Ge, W.; Steber, C.M. Positive and Negative Regulation of Seed Germination by the Arabidopsis GA Hormone Receptors, GID1a, b, and c. Plant Direct. 2018, 2, e00083. [Google Scholar] [CrossRef]
- Cubas, P. Role of TCP Genes in the Evolution of Key Morphological Characters in Angiosperms. In Developmental Genetics and Plant Evolution; Taylor & Francis: Abingdon, UK, 2002; pp. 247–266. [Google Scholar]
- He, Z.; Zhao, X.; Kong, F.; Zuo, Z.; Liu, X. TCP2 Positively Regulates HY5/HYH and Photomorphogenesis in Arabidopsis. J. Exp. Bot. 2016, 67, 775–785. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Shi, Y.; Li, M.; Fu, D.; Wu, S.; Li, J.; Gong, Z.; Liu, H.; Yang, S. The CRY2-COP1-HY5-BBX7/8 Module Regulates Blue Light-Dependent Cold Acclimation in Arabidopsis. Plant Cell 2021, 33, 3555–3573. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, H. Genome Evoultion and Epigenetic of Chrysanthemun and Its Related Species. Ph.D. Thesis, Nanjing Agricultural University, Nanjing, China, 2014. [Google Scholar]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Tian, F.; Yang, D.C.; Meng, Y.Q.; Jin, J.; Gao, G. PlantRegMap: Charting Functional Regulatory Maps in Plants. Nucleic Acids Res. 2020, 48, D1104–D1113. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a Database of Plant cis-Acting Regulatory Elements and a Portal to Tools for In Silico Analysis of Promoter Sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Ding, R.; Che, X.; Shen, Z.; Zhang, Y. Metabolome and Transcriptome Profiling Provide Insights into Green Apple Peel Reveals Light- and UV-B-Responsive Pathway in Anthocyanins Accumulation. BMC Plant Biol. 2021, 21, 351. [Google Scholar] [CrossRef]
- Zheng, Q.; Zuo, J.; Gu, S.; Gao, L.; Hu, W.; Wang, Q.; Jiang, A. Putrescine Treatment Reduces Yellowing During Senescence of Broccoli (Brassica oleracea L. var. italica). Postharvest Biol. Technol. 2019, 152, 29–35. [Google Scholar] [CrossRef]
- Li, W.; Huang, D.; Wang, B.; Hou, X.; Zhang, R.; Yan, M.; Liao, W. Changes of Starch and Sucrose Content and Related Gene Expression During the Growth and Development of Lanzhou Lily Bulb. PLoS ONE 2022, 17, e0262506. [Google Scholar] [CrossRef]





| CnTCPS | CnTCP4-1 | CnTCP4-3 | CnTCP4-2 | CnTCP14 | CnTCP2-3 | CnTCP2-2 | CnTCP2-1 | CnTCP13 | CnTCP5 | CnTCP17 | CnTCP3 | CnTCP15 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CnTCP10 | 10.70% | 11.40% | 11.00% | 11.60% | 11.80% | 12.10% | 12.10% | 10.30% | 12.90% | 16.30% | 12.30% | 11.20% |
| CnTCP12 | 11.20% | 11.90% | 11.20% | 12.40% | 11.30% | 11.40% | 11.30% | 10.60% | 14.20% | 15.50% | 12.00% | 11.20% |
| CnTCP16 | 14.10% | 14.20% | 14.10% | 13.10% | 15.50% | 15.50% | 15.50% | 12.90% | 13.80% | 16.10% | 15.50% | 15.20% |
| CnTCP9 | 15.80% | 15.70% | 15.70% | 11.80% | 12.60% | 12.70% | 12.70% | 13.20% | 18.80% | 17.60% | 15.10% | 14.40% |
| CnTCP7 | 15.30% | 14.80% | 14.80% | 14.70% | 14.80% | 15.00% | 14.80% | 12.00% | 14.20% | 13.80% | 15.50% | 16.80% |
| CnTCP8 | 14.00% | 13.40% | 13.70% | 9.40% | 11.00% | 11.10% | 11.00% | 12.20% | 12.90% | 12.80% | 17.40% | 14.10% |
| CnTCP18-1 | 11.80% | 11.90% | 11.80% | 13.40% | 13.60% | 13.60% | 13.80% | 14.20% | 15.50% | 18.10% | 16.40% | 16.10% |
| CnTCP18-2 | 12.30% | 12.40% | 12.30% | 14.00% | 13.70% | 13.70% | 14.00% | 14.70% | 16.00% | 17.80% | 16.50% | 16.20% |
| CnTCP6 | 16.40% | 16.10% | 16.40% | 15.70% | 14.90% | 14.70% | 14.60% | 15.80% | 17.60% | 17.40% | 14.80% | 15.80% |
| CnTCP1 | 19.80% | 20.60% | 20.60% | 19.10% | 17.60% | 17.60% | 17.50% | 17.50% | 19.70% | 22.50% | 31.10% | 29.20% |
| CnTCP11 | 20.70% | 21.50% | 20.80% | 15.70% | 17.40% | 17.40% | 17.30% | 17.30% | 14.90% | 19.10% | 27.00% | 25.90% |
| CnTCP15 | 16.00% | 16.50% | 16.00% | 17.60% | 18.80% | 18.80% | 18.70% | 16.00% | 16.20% | 19.90% | 46.20% | 100% |
| CnTCP3 | 16.50% | 15.70% | 16.10% | 15.40% | 17.00% | 17.00% | 16.90% | 16.70% | 17.80% | 17.90% | 100% | |
| CnTCP17 | 20.10% | 20.30% | 20.40% | 25.30% | 26.20% | 26.10% | 26.00% | 37.20% | 52.10% | 100% | ||
| CnTCP5 | 19.50% | 20.10% | 19.50% | 23.90% | 23.70% | 23.70% | 23.60% | 34.70% | 100% | |||
| CnTCP13 | 19.90% | 21.00% | 21.00% | 26.60% | 24.60% | 24.60% | 24.80% | 100% | ||||
| CnTCP2-1 | 20.20% | 19.40% | 20.00% | 57.40% | 99.50% | 99.80% | 100% | |||||
| CnTCP2-2 | 20.30% | 19.50% | 20.10% | 57.10% | 99.80% | 100% | ||||||
| CnTCP2-3 | 20.40% | 19.60% | 20.10% | 57.10% | 100% | |||||||
| CnTCP14 | 22.50% | 21.90% | 22.40% | 100% | ||||||||
| CnTCP4-2 | 93.80% | 96.00% | 100% | |||||||||
| CnTCP4-3 | 90.10% | 100% | ||||||||||
| CnTCP4-1 | 100% | |||||||||||
| CnTCPS | CnTCP11 | CnTCP1 | CnTCP6 | CnTCP18-2 | CnTCP18-1 | CnTCP8 | CnTCP7 | CnTCP9 | CnTCP16 | CnTCP12 | CnTCP10 | |
| CnTCP10 | 12.40% | 11.90% | 26.80% | 28.00% | 28.30% | 34.20% | 28.70% | 36.00% | 50.00% | 68.10% | 100% | |
| CnTCP12 | 11.60% | 13.30% | 26.40% | 28.40% | 28.60% | 34.00% | 29.10% | 34.00% | 50.60% | 100% | ||
| CnTCP16 | 11.50% | 13.80% | 27.40% | 31.30% | 31.70% | 38.10% | 35.70% | 39.30% | 100% | |||
| CnTCP9 | 14.80% | 18.20% | 28.70% | 31.50% | 31.50% | 28.90% | 33.10% | 100% | ||||
| CnTCP7 | 16.20% | 16.20% | 28.80% | 30.20% | 29.20% | 38.50% | 100% | |||||
| CnTCP8 | 13.00% | 14.50% | 28.80% | 29.70% | 29.20% | 100% | ||||||
| CnTCP18-1 | 14.10% | 16.20% | 41.70% | 97.30% | 100% | |||||||
| CnTCP18-2 | 14.20% | 16.20% | 41.30% | 100% | ||||||||
| CnTCP6 | 16.30% | 13.90% | 100% | |||||||||
| CnTCP1 | 52.30% | 100% | ||||||||||
| CnTCP11 | 100% |
| Family | AtTCP | CnTCP | Family | AtTCP | CnTCP |
|---|---|---|---|---|---|
| AP2 | 24 | 18 | TCP | 22 | 17 |
| B3 | 24 | 18 | ARF | 24 | 16 |
| bHLH | 24 | 18 | BBR-BPC | 24 | 16 |
| bZIP | 24 | 18 | WRKY | 24 | 16 |
| C2H2 | 24 | 18 | ZF-HD | 22 | 16 |
| Dof | 24 | 18 | LBD | 21 | 16 |
| ERF | 24 | 18 | SBP | 21 | 16 |
| G2-like | 24 | 18 | C3H | 20 | 16 |
| GATA | 24 | 18 | WOX | 22 | 15 |
| HD-ZIP | 24 | 18 | CPP | 21 | 15 |
| MIKC_MADS | 24 | 18 | EIL | 18 | 15 |
| MYB | 24 | 18 | NF-YB | 16 | 15 |
| MYB_related | 24 | 18 | E2F/DP | 23 | 14 |
| NAC | 24 | 18 | Nin-like | 22 | 14 |
| Trihelix | 23 | 18 | RAV | 22 | 14 |
| GRAS | 23 | 17 | SRS | 21 | 13 |
| ARR-B | 20 | 7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tian, C.; Zhai, L.; Zhu, W.; Qi, X.; Yu, Z.; Wang, H.; Chen, F.; Wang, L.; Chen, S. Characterization of the TCP Gene Family in Chrysanthemum nankingense and the Role of CnTCP4 in Cold Tolerance. Plants 2022, 11, 936. https://doi.org/10.3390/plants11070936
Tian C, Zhai L, Zhu W, Qi X, Yu Z, Wang H, Chen F, Wang L, Chen S. Characterization of the TCP Gene Family in Chrysanthemum nankingense and the Role of CnTCP4 in Cold Tolerance. Plants. 2022; 11(7):936. https://doi.org/10.3390/plants11070936
Chicago/Turabian StyleTian, Chang, Lisheng Zhai, Wenjing Zhu, Xiangyu Qi, Zhongyu Yu, Haibin Wang, Fadi Chen, Likai Wang, and Sumei Chen. 2022. "Characterization of the TCP Gene Family in Chrysanthemum nankingense and the Role of CnTCP4 in Cold Tolerance" Plants 11, no. 7: 936. https://doi.org/10.3390/plants11070936
APA StyleTian, C., Zhai, L., Zhu, W., Qi, X., Yu, Z., Wang, H., Chen, F., Wang, L., & Chen, S. (2022). Characterization of the TCP Gene Family in Chrysanthemum nankingense and the Role of CnTCP4 in Cold Tolerance. Plants, 11(7), 936. https://doi.org/10.3390/plants11070936









