Diversity Assessment and DNA-Based Fingerprinting of Sicilian Hazelnut (Corylus avellana L.) Germplasm
Abstract
:1. Introduction
2. Results
2.1. Evaluation of DNA Extraction Methods
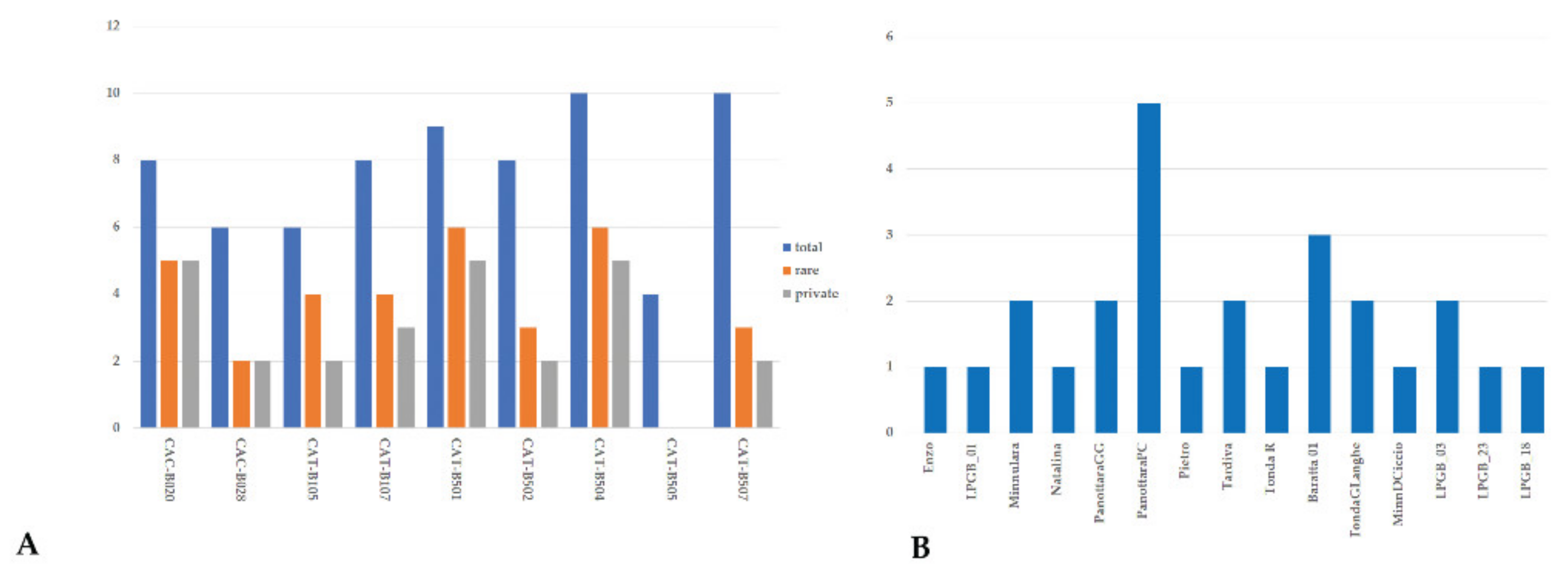
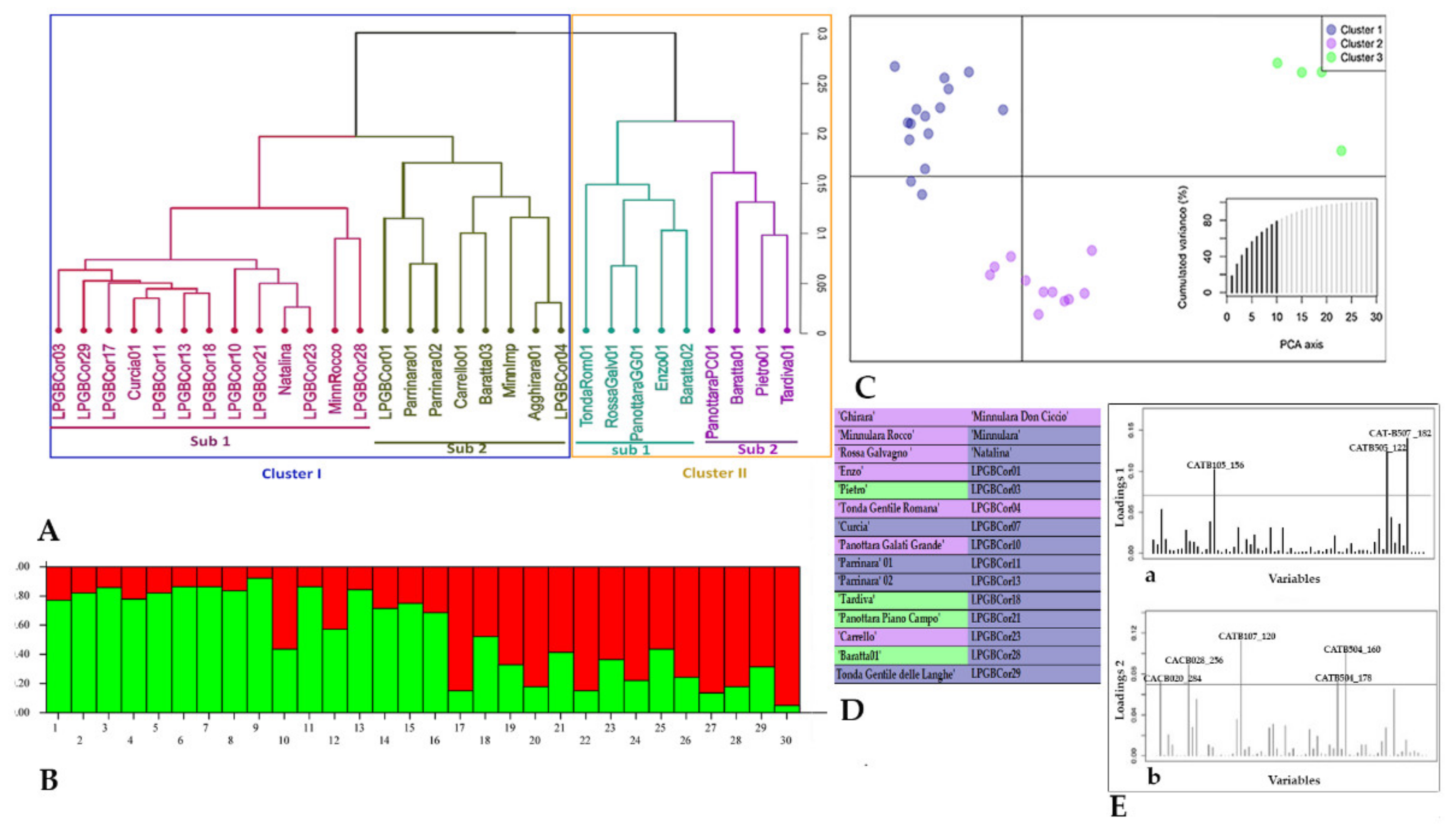
2.2. Genetic Diversity
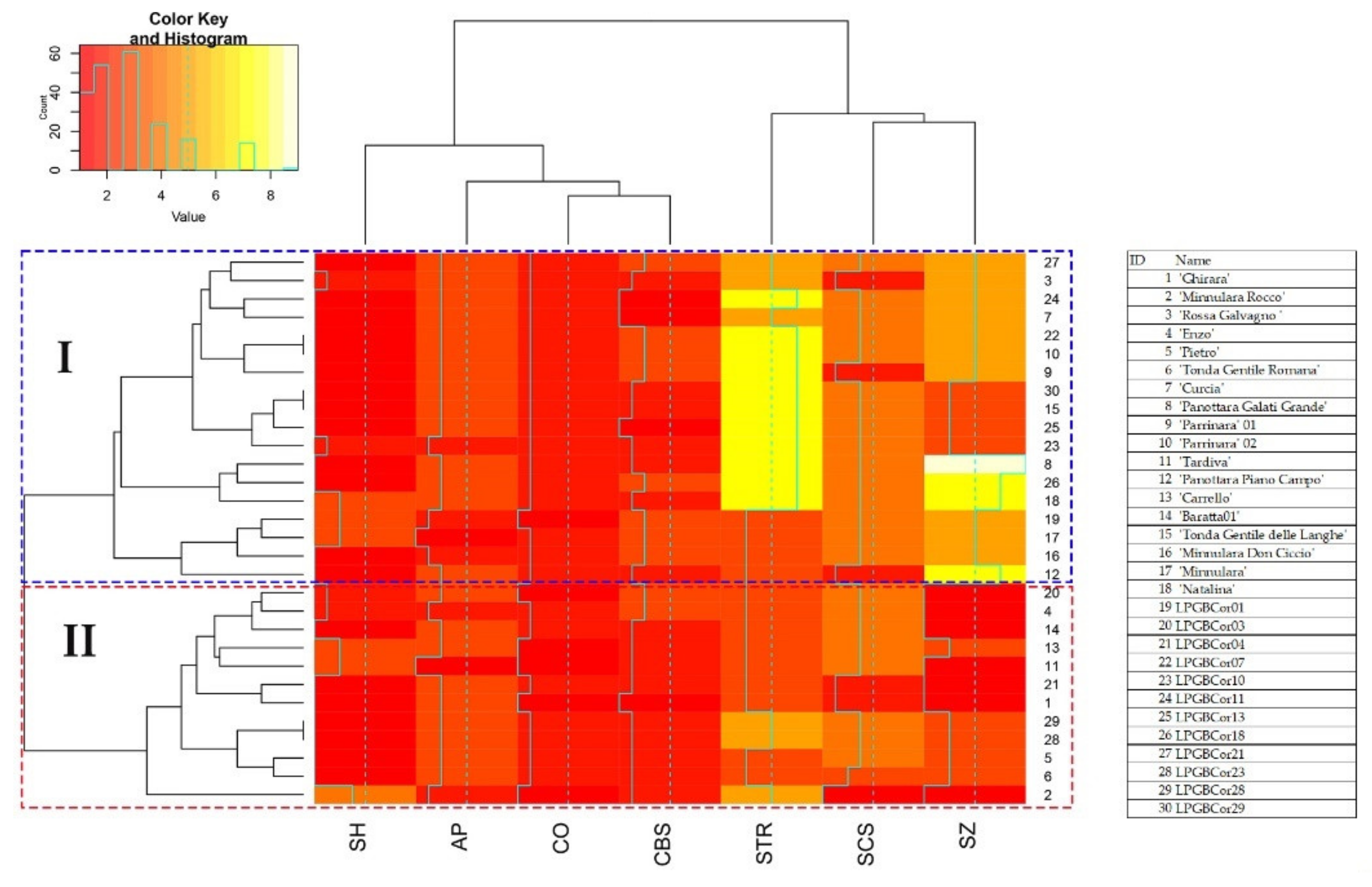
2.3. Nut Morphological Analysis
3. Discussion
4. Materials and Methods
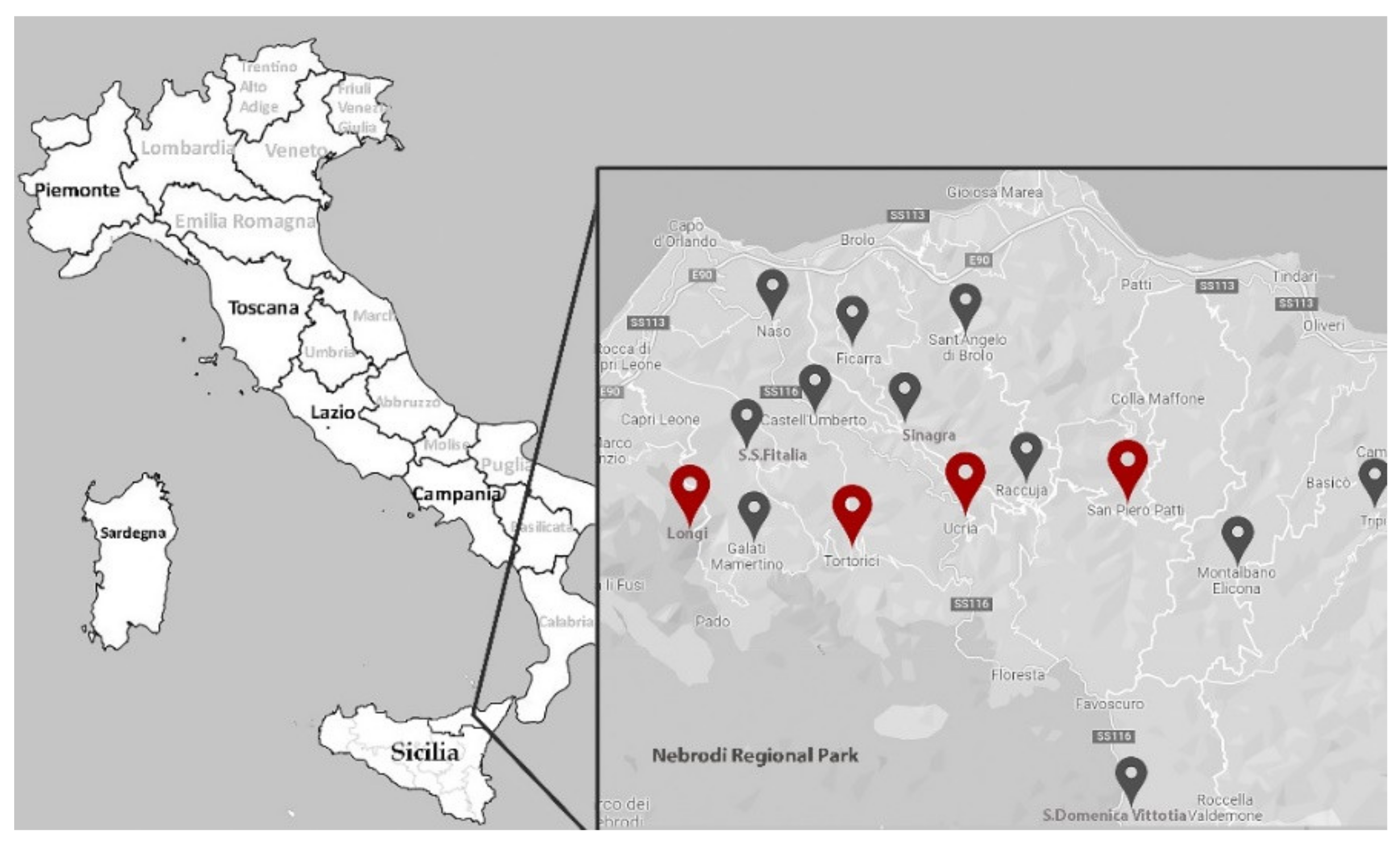
4.1. Plant Material and Nut Morphological Traits
4.2. DNA Extraction
4.3. SSR Fingerprinting
4.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dioscorides, P. The Greek Herbal of Dioscorides; Gunther, R.T., Ed.; Oxford Clarenton Press: Oxford, UK, 1934. [Google Scholar]
- Lagerstedt, H.B. Filberts. In Advances in Fruit Breeding; Janick, J., Moore, J.N., Eds.; Purdue University Press: West Lafayette, IN, USA; New York, NY, USA, 1975; pp. 456–488. ISBN 0911198369 9780911198362. [Google Scholar]
- Thompson, M.M.; Lagerstedt, H.B.; Mehlenbacher, S.A. Hazelnuts. In Fruit Breeding; Janick, J., Moore, J.N., Eds.; Wiley: New York, NY, USA, 1996; Volume 3, pp. 125–184. [Google Scholar]
- Molnar, T.J. Corylus. In Wild Crop Relatives: Genomic and Breeding Resources, Forest Trees; Kole, C., Ed.; Springer: Berlin, Germany, 2011; pp. 15–48. [Google Scholar] [CrossRef]
- Thompson, M.M. Genetics of incompatibility in Corylus avellana L. Theor. Appl. Genet. 1979, 54, 113–116. [Google Scholar] [CrossRef]
- Mehlenbacher, S. Revised dominance hierarchy for S-alleles in Corylus avellana L. Theor. Appl. Genet. 1997, 94, 360–366. [Google Scholar] [CrossRef]
- Boccacci, P.; Botta, R. Investigating the origin of hazelnut (Corylus avellana L.) cultivars using chloroplast microsatellites. Genet. Resour. Crop Evol. 2009, 56, 851–859. [Google Scholar] [CrossRef]
- Robinson, G.; Town, M.; Ballin, T.; Clarke, A.; Dunne, J.; Evershed, P.R.; Gardner, L.F.; Gibson, A.; Russ, H. Furness’s first farmers: Evidence of early neolithic settlement and dairying in Cumbria. Proc. Prehist. Soc. 2020, 86, 165–198. [Google Scholar] [CrossRef]
- Tipping, R. The form and the fate of Scotland’s woodlands. Proc. Soc. Antiqu. Scot. 1994, 124, 1–54. Available online: http://soas.is.ed.ac.uk/index.php/psas/article/view/9485 (accessed on 15 June 2021).
- Holst, D. Hazelnut economy of early Holocene hunter–gatherers: A case study from Mesolithic Duvensee, northern Germany. J. Archaeol. Sci. 2010, 37, 2871–2880. [Google Scholar] [CrossRef]
- Rottoli, M.; Castiglioni, E. Plant offerings from Roman cremations in northern Italy: A review. Veget. Hist. Archaeobot. 2011, 20, 495–506. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations. Available online: http://www.fao.org/faostat/en/#home (accessed on 12 January 2022).
- ISTAT Coltivazioni: Coltivazioni Legnose Fruttifere. Available online: www.istat.it (accessed on 12 January 2022).
- Alfonso, F. Monografia Sul Nocciuolo. In Tipografia dello Statuto; Consorzio Agrario delle Provincie Siciliane: Palermo, Italy, 1886; pp. 1–496. [Google Scholar]
- Granata, M.U.; Bracco, F.; Catoni, R. Carbon dioxide sequestration capability of hazelnut orchards: Daily and seasonal trends. Energ. Ecol. Environ. 2020, 5, 153–160. [Google Scholar] [CrossRef]
- Biasi, R.; Farina, R.; Brunori, E. Family farming plays an essential role in preserving soil functionality: A study on active managed and abandoned traditional tree crop-based systems. Sustainability 2021, 13, 3967. [Google Scholar] [CrossRef]
- Nera, E.; Paas, W.; Reidsma, P.; Paolini, G.; Antonioli, F.; Severini, S. Assessing the resilience and sustainability of a hazelnut farming system in central Italy with a participatory approach. Sustainability 2020, 12, 343. [Google Scholar] [CrossRef] [Green Version]
- Oliveira, I.; Silva, A.P.; Santos, T.; Igrejas, G.; Gonçalves, B. Hazelnut fruit and kernel traits: Influence of training systems and harvest year. Eur. J. Hortic. Sci. 2019, 84, 57–66. [Google Scholar] [CrossRef]
- Bostan, Z.S.; Islam, A.; Sen, S. Investigation on nut development in hazelnuts and determination of nut characteristic and variation within cultivars in some hazelnut cultivars. Acta Hort. 1997, 445, 101–107. [Google Scholar] [CrossRef]
- Klingenberg, C.P. Phenotypic plasticity, developmental instability, and robustness: The concepts and how they are connected. Front. Ecol. Evol. 2019, 7, 56–71. [Google Scholar] [CrossRef] [Green Version]
- Yao, Q.; Mehlenbacher, S.A. Heritability, variance components and correlation of morphological and phenological traits in hazelnut. Plant Breed. 2000, 119, 369–381. [Google Scholar] [CrossRef]
- Korir, N.K.; Han, J.; Shangguan, L.; Wang, C.; Kayesh, E.; Zhang, Y.; Fang, J. Plant variety and cultivar identification: Advances and prospects. Crit. Rev. Biotechnol. 2012, 33, 111–125. [Google Scholar] [CrossRef] [PubMed]
- Bacchetta, L.; Aramini, M.; Bernardini, C.; Rugini, E. In vitro propagation of traditional Italian hazelnut cultivars as a tool for the valorization and conservation of local genetic resources. HortScience 2008, 43, 562–566. [Google Scholar] [CrossRef]
- Dangl, G.S.; Yang, J.; Golino, D.A.; Gratziel, T. A practical method for almond cultivar identification and parental analysis using simple sequence repeat markers. Euphytica 2009, 168, 41–48. [Google Scholar] [CrossRef] [Green Version]
- Bassil, N.V.; Botta, R.; Mehlenbacher, S.A. Microsatellite markers in hazelnut: Isolation, characterization and cross-species amplification. J. Am. Soc. Hortic. Sci. 2005, 130, 543–549. [Google Scholar] [CrossRef] [Green Version]
- Mehlenbacher, S.A.; Brown, R.N.; Nouhra, E.N.; Gökirmak, T.; Bassil, N.V.; Kubisiak, T.L. A genetic linkage map for hazelnut (Corylus avellana L.) based on RAPD and SSR markers. Genome 2006, 49, 122–133. [Google Scholar] [CrossRef] [Green Version]
- Boccacci, P.; Akkak, A.; Bassil, N.V.; Mehlenbacher, S.A.; Botta, R. Characterization and evaluation of microsatellite loci in European hazelnut (C. avellana) and their transferability to other Corylus species. Mol. Ecol. Notes 2005, 5, 934–937. [Google Scholar] [CrossRef]
- Boccacci, P.; Aramini, M.; Valentini, N.; Bacchetta, L.; Rovira, M.; Drogoudi, P.; Silva, A.P.; Solar, A.; Calizzano, F.; Erdoğan, V.; et al. Molecular and morphological diversity of on-farm hazelnut (Corylus avellana L.) landraces from southern Europe and their role in the origin and diffusion of cultivated germplasm. Tree Genet. Genomes 2013, 9, 1465–1480. [Google Scholar] [CrossRef]
- Bacchetta, L.; Rovira, M.; Tronci, C.; Aramini, M.; Drogoudi, P.; Silva, A.P.; Solar, A.; Avanzato, D.; Botta, R.; Valentini, N.; et al. Multidisciplinary approach to enhance the conservation and use of hazelnut Corylus avellana L. genetic resources. Genet. Resour. Crop Evol. 2015, 62, 649–663. [Google Scholar] [CrossRef]
- Demchik, M.; Kern, A.; Braun, L.; Fischbach, J.; Turnquist, K. Genetic diversity of American hazelnut in the Upper Midwest, USA. Agrofor. Syst. 2018, 92, 1507–1516. [Google Scholar] [CrossRef]
- Köse, M.A.; Gürcan, K. Genetic diversity and genetic comparison of hazelnuts (Corylus avellana L.) of Kayseri province of Turkey to major accessions of Azerbaijan, Georgia, Italy, Spain, and Turkey. Acta Hort. 2018, 1226, 93–99. [Google Scholar] [CrossRef]
- Tanhuanpää, P.; Heinonen, M.; Bitz, L.; Rokka, V.M. Genetic diversity and structure in the northern populations of European hazelnut (Corylus avellana L.). Genome 2019, 62, 537–548. [Google Scholar] [CrossRef]
- Kavas, M.; Yıldırım, K.; Seçgin, Z.; Gökdemir, G. Discovery of simple sequence repeat (SSR) markers in hazelnut (Corylus avellana L.) by transcriptome sequencing and SSR-based characterization of hazelnut cultivars. Scand. J. For. Res. 2020, 35, 227–237. [Google Scholar] [CrossRef]
- Boccacci, P.; Akkak, A.M.; Botta, R. DNA typing and genetic relations among European hazelnut (Corylus avellana L.) cultivars using microsatellite markers. Genome 2006, 49, 598–611. [Google Scholar] [CrossRef]
- Gürcan, K.; Mehlenbacher, S.A.; Erdoğan, V. Genetic diversity in hazelnut (Corylus avellana L.) cultivars from black sea countries assessed using SSR markers. Plant Breed. 2010, 129, 422–434. [Google Scholar] [CrossRef]
- Botta, R.; Molnar, T.J.; Erdogan, V.; Valentini, N.; Torello Marinoni, D.; Mehlenbacher, S.A. Hazelnut (Corylus spp.) Breeding. In Advances in Plant Breeding Strategies: Nut and Beverage Crops; Al-Khayri, J., Jain, S., Johnson, D., Eds.; Springer: Cham, Switzerland, 2019; pp. 157–219. [Google Scholar] [CrossRef]
- Gökirmak, T.; Mehlenbacher, S.A.; Bassil, N.V. Characterization of European hazelnut (Corylus avellana) cultivars using SSR markers. Genet. Resour. Crop Evol. 2009, 56, 147–172. [Google Scholar] [CrossRef]
- Boccacci, P.; Botta, R. Microsatellite variability and genetic structure in hazelnut (Corylus avellana L.) cultivars from different growing regions. Sci. Hortic. 2010, 124, 128–133. [Google Scholar] [CrossRef]
- Sathuvalli, V.R.; Mehlenbacher, S.A. Characterization of American hazelnut (Corylus americana) accessions and Corylus americana × Corylus avellana hybrids using microsatellite markers. Genet. Resour. Crop Evol. 2012, 59, 1055–1075. [Google Scholar] [CrossRef]
- Bassil, N.; Boccacci, P.; Botta, R.; Postman, J.; Mehlenbacher, S. Nuclear and chloroplast microsatellite markers to assess genetic diversity and evolution in hazelnut species, hybrids and cultivars. Genet. Resour. Crop Evol. 2013, 60, 543–568. [Google Scholar] [CrossRef] [Green Version]
- Gürcan, K.; Mehlenbacher, S.A.; Köse, M.A.; Balik, H.I. Population structure analysis of European hazelnut (Corylus avellana). Acta Hort. 2018, 1226, 87–92. [Google Scholar] [CrossRef]
- Martins, S.; Simões, F.; Mendonça, D.; Matos, S.; Silva, A.P.; Carnide, V. Western European Wild and Landraces Hazelnuts Evaluated by SSR Markers. Plant Mol. Biol. Rep. 2015, 33, 1712–1720. [Google Scholar] [CrossRef]
- Öztürk, S.C.; Balık, H.İ.; Balık, S.K.; Kızılcı, G.; Duyar, Ö.; Doğanlar, S.; Frary, A. Molecular genetic diversity of the Turkish national hazelnut collection and selection of a core set. Tree Genet. Genomes 2017, 13, 113. [Google Scholar] [CrossRef]
- Ozturk, S.C.; Ozturk, S.E.; Celik, I.; Celik, I.; Stampar, F.; Veberic, R.; Doganlar, S.; Solar, A.; Frary, A. Molecular genetic diversity and association mapping of nut and kernel traits in Slovenian hazelnut (Corylus avellana) germplasm. Tree Genet. Genomes 2017, 13, 16. [Google Scholar] [CrossRef]
- Honig, J.A.; Muehlbauer, M.F.; Capik, J.M.; Kubik, C.; Vaiciunas, J.N.; Mehlenbacher, S.A.; Molnar, T.J. Identification and mapping of eastern filbert blight resistance quantitative trait loci in european hazelnut using double digestion restriction site associated DNA sequencing. J. Amer. Soc. Hort. Sci. 2019, 144, 295–304. [Google Scholar] [CrossRef] [Green Version]
- Torello Marinoni, D.; Valentini, N.; Portis, E.; Acquadro, A.; Beltramo, C. High density SNP mapping and QTL analysis for time of leaf budburst in Corylus avellana L. PLoS ONE 2018, 13, e0195408. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Helmstetter, A.J.; Oztolan-Erol, N.; Lucas, S.J.; Buggs, R.J.A. Genetic diversity and domestication of hazelnut (Corylus avellana) in Turkey. Plants People Planet 2020, 2, 326–329. [Google Scholar] [CrossRef] [Green Version]
- Lombardoni, J.J.; Honig, J.A.; Vaiciunas, J.; Kubik, C.; Capik, J.; Mehlenbacher, S.; Molnar, T.J. Evaluation of European hazelnut (Corylus avellana) genetic diversity using a genotyping-by-sequencing approach. Acta Hort. 2020, 1280, 11–16. [Google Scholar] [CrossRef]
- Oztolan-Erol, N.; Helmstetter, A.J.; İnan, A.; Buggs, R.J.A.; Lucas, S.J. Unraveling genetic diversity amongst European hazelnut (Corylus avellana L.) varieties in Turkey. Front. Plant Sci. 2021, 12, 1250. [Google Scholar] [CrossRef]
- Scaglione, D.; Fornasiero, A.; Cattonaro, F.; Pitoni, A.; Torelli, D.; Vittori, D.; De Pace, C. High-density marker genotyping in hazelnut (Corylus avellana). I. Genotyping-by-sequencing approach for efficient SNP discovery and DNA typing. Acta Hort. 2015, 1100, 63–67. [Google Scholar] [CrossRef]
- Bacchetta, L.; Aramini, M.; Zini, A.; Di Gianmatteo, V.; Spera, D.; Drogoudi, P.; Rovira, M.; Silva, P.; Solar, A.; Botta, R. Fatty acids and alpha-tocopherol composition in hazelnut (Corylus avellana L.): A chemometric approach to emphasize the quality of European germplasm. Euphytica 2013, 191, 57–73. [Google Scholar] [CrossRef]
- Campa Negrillo, A.; Rodríguez Madrera, R.; Suárez Valles, B.; Ferreira, J.J. Variation of morphological, agronomic and chemical composition traits of local hazelnuts collected in northern Spain. Front. Plant Sci. 2021, 12, 659510. [Google Scholar] [CrossRef]
- Cittadini, M.C.; Martín, D.; Gallo, S.; Fluente, G.; Bodara, R.M.; Martinez, M.L.; Maestri, D. Evaluation of hazelnut and walnut oil chemical traits from conventional cultivars and native genetic resources in a non-traditional crop environment from Argentina. Eur. Food Res. Technol. 2020, 246, 833–843. [Google Scholar] [CrossRef]
- Martins, S.; Simões, F.; Matos, J.; Silva, A.P.; Carnide, V. Genetic relationship among wild, landraces and cultivars of hazelnut (Corylus avellana) from Portugal revealed through ISSR and AFLP markers. Plant Syst. Evol. 2014, 300, 1035–1046. [Google Scholar] [CrossRef]
- Bacchetta, L.; Avanzato, D.; Giovanni, B.; Botta, R.; Boccacci, P.; Drogoudi, P.; Metzidakis, I.; Rovira, M.; Sarraquigne, J.P.; Silva, A.P.; et al. The reorganisation of European hazelnut genetic resources in the SAFENUT (AGRI GEN RES) project. Acta Hort. 2014, 1052, 67–74. [Google Scholar] [CrossRef]
- Ordidge, M.; Litthauer, S.; Venison, E.; Blouin-Delmas, M.; Fernandez-Fernandez, F.; Höfer, M.; Kägi, C.; Kellerhals, M.; Marchese, A.; Mariette, S.; et al. Towards a Joint International Database: Alignment of SSR marker data for European collections of cherry germplasm. Plants 2021, 10, 1243. [Google Scholar] [CrossRef]
- Nybom, H.; Giovannini, D.; Ordidge, M.; Hjeltnes, S.H.; Grahić, J.; Gaši, F. ECPGR recommended SSR loci for analyses of European plum (Prunus domestica) collections. Genet. Resour. 2020, 1, 40–48. [Google Scholar] [CrossRef]
- FAO; CIHEAM. Biodiversity. In Descriptors for Hazelnut (Corylus avellana L.); Biodiversity International: Rome, Italy; Food and Agriculture Organization of the United Nations: Rome, Italy; International Centre for Advanced Mediterranean Agronomic Studies: Zaragoza, Spain, 2008. [Google Scholar]
- Amiteye, S. Basic concepts and methodologies of DNA marker systems in plant molecular breeding. Heliyon 2021, 7, e080932. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15, ISBN: 978-92-9043-762-8. [Google Scholar]
- Martínez-González, C.R.; Ramírez-Mendoza, R.; Jiménez-Ramírez, J.; Gallegos-Vázquez, C.; Luna-Vega, I. Improved method for genomic DNA extraction for Opuntia Mill. (Cactaceae). Plant Methods 2017, 13, 82–92. [Google Scholar] [CrossRef] [Green Version]
- Nybom, H.; Weising, K. DNA-Based Identification of Clonally Propagated Cultivars. In Plant Breeding Reviews; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2011; pp. 221–295. [Google Scholar] [CrossRef]
- Yang, Y.; Tian, H.; Wang, R.; Wang, L.; Yi, H.; Liu, Y.; Xu, L.; Fan, Y.; Zhao, J.; Wang, F. Variety Discrimination Power: An Appraisal Index for Loci Combination Screening Applied to Plant Variety Discrimination. Front. Plant Sci. 2021, 12, 566796. [Google Scholar] [CrossRef]
- Alberghina, O. Indagine sulla corilicoltura siciliana. Frutticoltura 1982, 2, 27–30. [Google Scholar]
- Shete, S.; Tiwari, H.; Elston, R.C. On estimating the heterozygosity and polymorphism information content value. Theor. Popul. Biol. 2000, 57, 265–271. [Google Scholar] [CrossRef]
- Alberghina, O.; Damigella, P. Indagine sulla Composizione Varietale di Alcuni Centri Corilicoli del Messinese. In Atti del Convegno Nazionale su: II Miglioramento della Coltura del Mandorlo e del Nocciolo; Messina e Siracusa, Italy, 1979; pp. 145–194. Available online: https://www.unipa.it/biblioteche/ (accessed on 25 February 2021).
- Damigella, P.; Alberghina, O. Indagine sulla Composizione Varietale di Alcuni Centri Corilicoli del Messinese: Le Cultivar Presenti nei Territori Comunali di Tortorici e Sinagra. In Atti Convegno Internazionale sul Nocciuolo; Avellino, Italy, 1983; pp. 363–377. Available online: https://www.unipa.it/biblioteche/ (accessed on 25 February 2021).
- Lucas, S.J.; Kahraman, K.; Avşar, B.; Buggs, R.J.; Bilge, I. A chromosome-scale genome assembly of European hazel (Corylus avellana L.) reveals targets for crop improvement. Plant J. 2021, 105, 1413–1430. [Google Scholar] [CrossRef]
- Li, C.; Qi, R.; Yuan, J.; Han, L.; Wang, S.; Li, W.; Han, W. In silico study to predict potential precursors of human dipeptidyl peptidase-IV inhibitors from hazelnut. J. Biomol. Struct. Dyn. 2021, 5, 1–12. [Google Scholar] [CrossRef]
- Rowley, E.R.; VanBuren, R.; Bryant, D.W.; Priest, H.D.; Mehlenbacher, S.A.; Mockler, T.C. A Draft Genome and High-Density Genetic Map of European Hazelnut (Corylus avellana L.). bioRxiv 2018, 469015. [Google Scholar] [CrossRef] [Green Version]
- Freixas-Coutin, J.A.; An, S.; Postman, J.; Bassil, N.V.; Yates, B.; Shukla, M.; Saxena, P.K. Development of a reliable Corylus sp. reference database through the implementation of a DNA fingerprinting test. Planta 2019, 249, 1863–1874. [Google Scholar] [CrossRef]
- Boccacci, P.; Aramini, M.; Ordidge, M.; van Hantum, T.J.L.; Torello Marinoni, D.; Valentini, N.; Sarraquigne, J.; Solar, A.; Rovira, M.; Bacchetta, L.; et al. Comparison of selection methods for the establishment of a core collection using SSR markers for hazelnut (Corylus avellana L.) accessions from European germplasm repositories. Tree Genet. Genomes 2021, 17, 48. [Google Scholar] [CrossRef]
- Garrone, W.; Vacchetti, M. Hazelnut quality in relation with the requirements of confectionary industry. Acta Hort. 1994, 351, 641–648. [Google Scholar] [CrossRef]
- Cristofori, V.; Bertazza, G.; Bignami, C. Changes in kernel chemical composition during nut development of three Italian hazelnut cultivars. Fruits 2015, 70, 311–322. [Google Scholar] [CrossRef]
- Silvestri, C.; Bacchetta, L.; Bellincontro, A.; Cristofori, V. Advances in cultivar choice, hazelnut orchard management, and nut storage to enhance product quality and safety: An overview. J. Sci. Food Agric. 2021, 7, 27–43. [Google Scholar] [CrossRef]
- International Union for the Protection of New Varieties of Plants (UPOV). In Proceedings of the Technical Working Party for Fruit Crops at Its Fifty-Second Session, Zhengzhou, China, 12–16 July 2021; Available online: https://www.upov.int (accessed on 21 December 2021).
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [Green Version]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. Peer J. 2014, 2, e281. [Google Scholar] [CrossRef] [Green Version]
- Kalinowski, S.T.; Taper, M.L.; Marshall, T.C. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 2007, 16, 1099–1106. [Google Scholar] [CrossRef]
- Marshal, D.R.; Brown, A.H.D. Optimum Sampling Strategies in Genetic Conservation. In Crop Genetic Resources for Today and Tomorrow; Frankel, O.H., Hawkes, J.G., Eds.; Cambridge University Press: London, UK, 1975; pp. 53–58. [Google Scholar]
- Tessier, C.; David, J.; This, P.; Boursiquot, J.M.; Charrier, A. Optimization of the choice of molecular markers for varietal identification in Vitis vinifera L. Theor. App. Genet. 1999, 98, 171–177. [Google Scholar] [CrossRef]
- Bruvo, R.; Michiels, N.K.; D’Souza, T.G.; Schulenburg, H. A simple method for the calculation of microsatellite genotype distances irrespective of ploidy level. Mol. Ecol. 2004, 13, 2101–2106. [Google Scholar] [CrossRef]
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 2010, 11, 94–109. [Google Scholar] [CrossRef] [Green Version]
- Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [Green Version]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. Available online: http://www.genetics.org/content/155/2/945 (accessed on 21 December 2021). [CrossRef]
- Earl, D.A.; von Holdt, B.M. Structure Harvester: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [Green Version]
- Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 1967, 27, 209–220. Available online: https://cancerres.aacrjournals.org/content/27/2_Part_1/209 (accessed on 21 December 2021).


| Locus | Na | Ne | Ho | He | uHe | PIC | D |
|---|---|---|---|---|---|---|---|
| CAC-B020 | 8 | 3.051 | 0.833 | 0.672 | 0.684 | 0.634 | 0.600 |
| CAC-B028 | 6 | 3.523 | 0.933 | 0.716 | 0.728 | 0.678 | 0.612 |
| CAT-B105 | 6 | 1.532 | 0.367 | 0.347 | 0.353 | 0.332 | 0.328 |
| CAT-B107 | 8 | 3.488 | 0.800 | 0.713 | 0.725 | 0.675 | 0.608 |
| CAT-B501 | 9 | 3.147 | 0.900 | 0.682 | 0.694 | 0.649 | 0.516 |
| CAT-B502 | 8 | 3.383 | 0.767 | 0.704 | 0.716 | 0.638 | 0.267 |
| CAT-B504 | 10 | 4.174 | 0.862 | 0.760 | 0.774 | 0.744 | 0.659 |
| CAT-B505 | 4 | 3.186 | 1.00 | 0.686 | 0.698 | 0.622 | 0.426 |
| CAT-B507 | 10 | 3.922 | 0.900 | 0.745 | 0.758 | 0.717 | 0.650 |
| Average | 7.67 | 3.267 | 0.818 | 0.670 | 0.684 | 0.681 | 0.518 |
| Cluster | Inferred Cluster | Mean Fst | Expected Heterozygosity | Number of Accessions |
|---|---|---|---|---|
| I | 0.460 | 0.0016 | 0.8247 | 13 |
| II | 0.540 | 0.4482 | 0.5166 | 17 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fiore, M.C.; Marchese, A.; Mauceri, A.; Digangi, I.; Scialabba, A. Diversity Assessment and DNA-Based Fingerprinting of Sicilian Hazelnut (Corylus avellana L.) Germplasm. Plants 2022, 11, 631. https://doi.org/10.3390/plants11050631
Fiore MC, Marchese A, Mauceri A, Digangi I, Scialabba A. Diversity Assessment and DNA-Based Fingerprinting of Sicilian Hazelnut (Corylus avellana L.) Germplasm. Plants. 2022; 11(5):631. https://doi.org/10.3390/plants11050631
Chicago/Turabian StyleFiore, Maria Carola, Annalisa Marchese, Antonio Mauceri, Ignazio Digangi, and Anna Scialabba. 2022. "Diversity Assessment and DNA-Based Fingerprinting of Sicilian Hazelnut (Corylus avellana L.) Germplasm" Plants 11, no. 5: 631. https://doi.org/10.3390/plants11050631
APA StyleFiore, M. C., Marchese, A., Mauceri, A., Digangi, I., & Scialabba, A. (2022). Diversity Assessment and DNA-Based Fingerprinting of Sicilian Hazelnut (Corylus avellana L.) Germplasm. Plants, 11(5), 631. https://doi.org/10.3390/plants11050631






