High Carotenoid Mutants of Chlorella vulgaris Show Enhanced Biomass Yield under High Irradiance
Abstract
1. Introduction
2. Results
2.1. Isolation of NFR-3 and NFR-13, High Car Content Mutants of Chlorella vulgaris
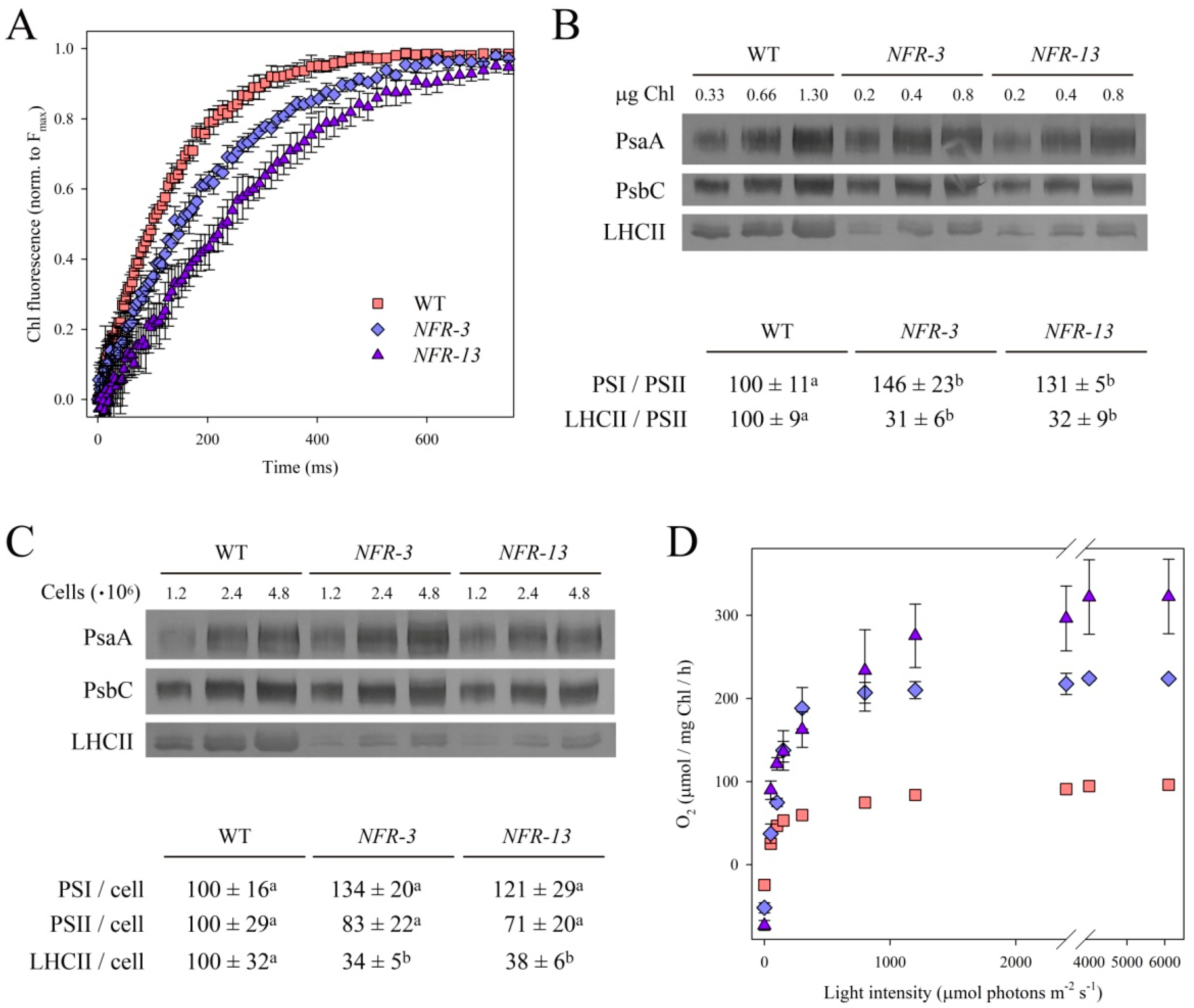
2.2. Stoichiometry of Pigment–Protein Complexes and Photosynthetic Efficiency
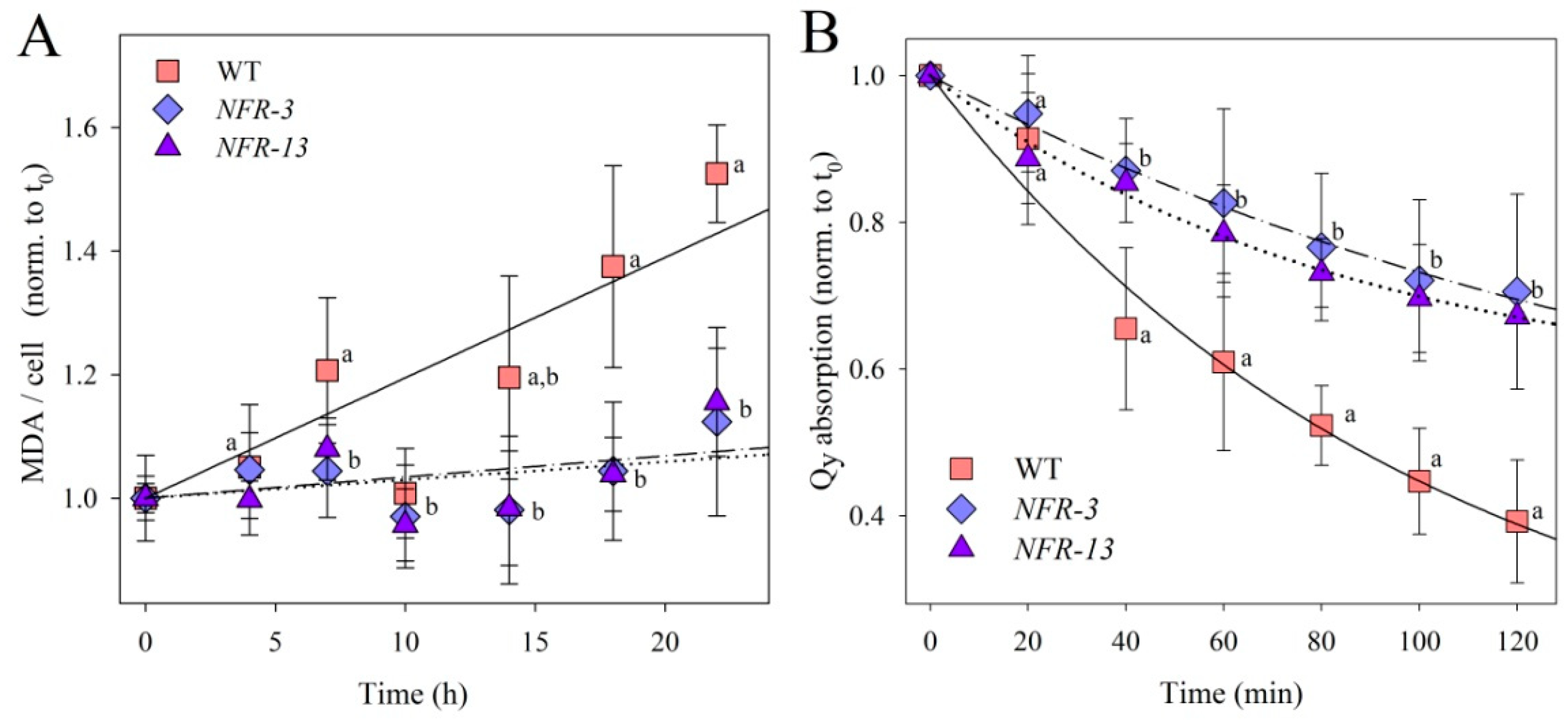
2.3. Phototolerance of WT and NFR Strains during EL Exposure
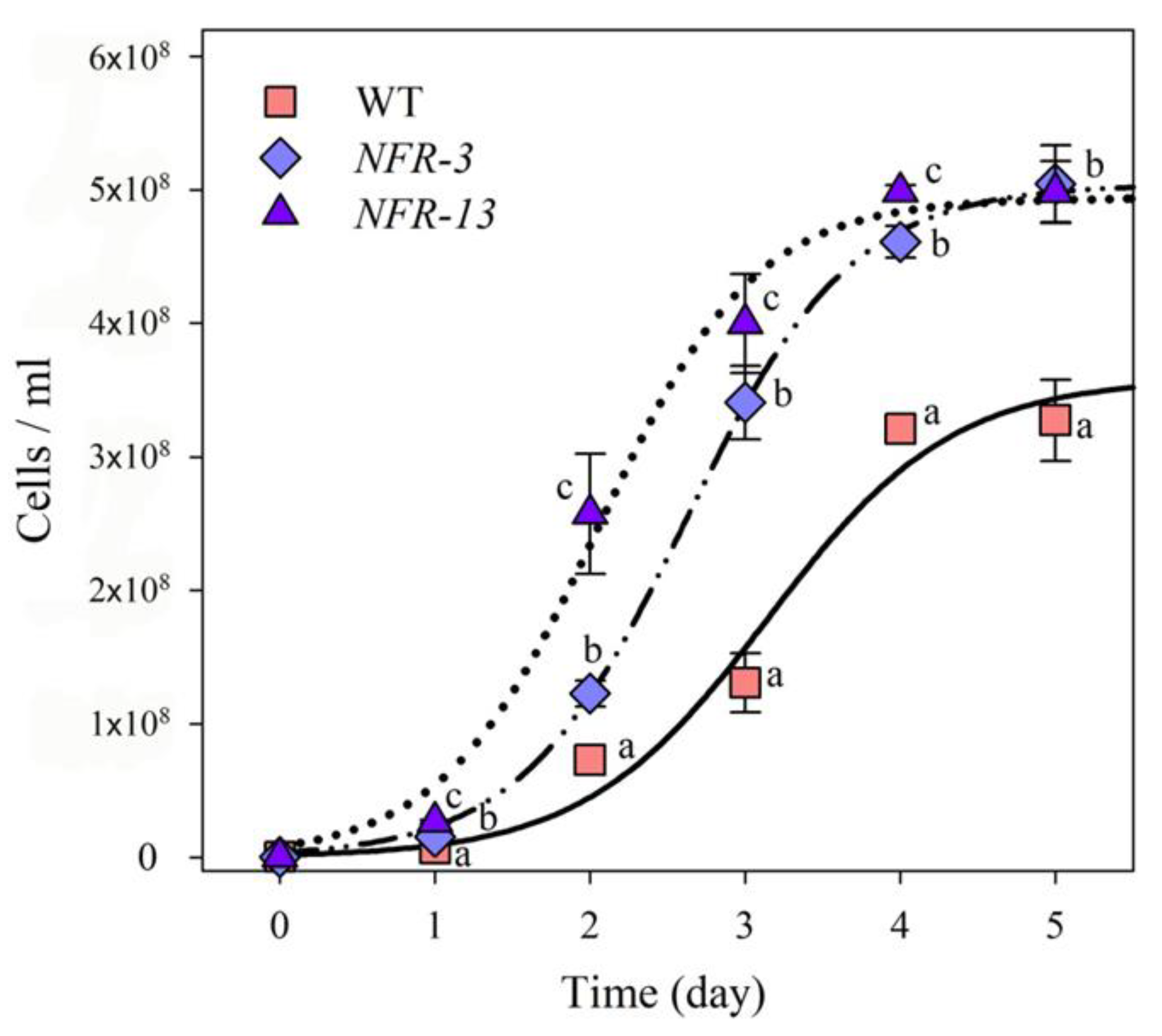
2.4. Biomass Productivity of WT and NFR Strains in Laboratory-Scale PBR
2.5. Lipid Productivity under Nitrogen Starvation
2.6. Genotypic Characterization of NFR Strains by Whole-Genome Sequencing
3. Discussion
3.1. Optimization of Light-Use Efficiency Enhances Biomass Yield
3.2. Influence of ROS-Resistance Traits on Stress-Induced Lipid Production
3.3. NFR as Promising Car Producer Strains for Commercial Applications
3.4. Whole-Genome Sequencing to Identify Gene(s) Responsible for the Enhanced Light-Use Efficiency in C. vulgaris
- Glycerol-3-phosphate acyltransferase 3. A transversion (G to C) was observed at nucleotide 534 on the 6th exon. This results in missense amino acid substitution of a glutamic acid into an aspartic acid, a mutation shared by both NFR mutants, affecting a subunit involved in neutral lipid metabolism. Glutamic acid and aspartic acid are both acidic amino acids with side chains containing a carboxylic acid group which carries a full negative charge. In terms of interaction modes, both amino acids include ionic, van der Waals and H-bonds. However, glutamic acid differs from aspartic acid only in that its side chain is longer by one methylene group. This results in higher side chain flexibility in glutamic acid compared to aspartic acid whose side chain flexibility is moderate. Transmembrane topology prediction indicates that the site of mutation is at the boundary between cytoplasmic domain (residues 169–179) and the consecutive transmembrane region (residues 180–199). Multiple sequence alignments of top BLAST hits indicate that the last two residues of the cytoplasmic domain (E178 and R179) are conserved among green cut species (Figure S7). Glycerol-3-phosphate acyltransferase 3 is a key component of the diglyceride-3-phosphate synthesis pathway. Consistently, its activity has been reported to be directly correlated to the amount of glycerophospholipids in E. coli cells [82] and used as a target for membrane engineering to enhance production and accumulation of ß-carotene [83].
- E3 Ubiquitin-protein Ligase SP1-related (PTHR47568:SF2) encoding sequence contains a splice donor variant and is shared by both mutants, NFR-3 and NFR-13. A transition (G to A) occurs at the first nucleotide of the second intron affecting canonical GT-AG splice site pairs. Group of E3 ubiquitin–protein ligases, including SP1 from Arabidopsis (UniProtKB: Q8L7N4), are involved in regulation of plastid’s proteome via ubiquitination and subsequent degradation of translocon at the outer envelope membrane of chloroplasts (TOC) complexes [84]. In addition, it promotes stress tolerance by depleting the chloroplast protein import apparatus, which limits photosystem assembly and the potential for ROS formation [85]. Formighieri et al. [86] demonstrated that absence of ARSA1, a protein localized in the C. reinhardtii cytosol, led to a strongly decreased Chl content per cell; ARSA1 was shown to be required for optimal biogenesis of photosystems due to its involvement in the accumulation of TOC34, a component of the outer chloroplast membrane translocon complex. Interestingly, in both species, a mutation targeting a component interacting with TOC is associated with a pale green phenotype.
- Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A contains variation observed in NFR-13 mutant only. Transition mutation (C to T) results in substitution of an arginine by cysteine. 2A protein phosphatases are evolutionarily conserved and carry out multiple functions such as growth- and stress-related signaling, cell cycle regulation, vesicle trafficking, as well as regulation of the activities of a number of enzymes involved in key metabolic pathways [87]. The mutant 2A protein phosphatase in NFR-13 contains fatty acid desaturase domain (IPR005804) which is involved in the lipid metabolic process (GO:0006629).
- Both NFR-3 and NFR-13 share an intronic variation on Allophanate hydrolase (g7748). In Chlamydomonas reinhardtii, allophanate hydrolase is involved in urea hydrolysis to produce ammonium depending on the source of nitrogen available [88].
- Another exonic mutation found only in NFR-3 targets biosynthetic arginine decarboxylase (g9015) involved in the polyamine biosynthesis pathway. In plants, polyamine accumulation was shown to correlate with Car content [89].
- A missense amino acid substitution (Arg233Cys) was identified on S49 family peptidase (g6424) in NFR-3. The predicted protein contains peptidase_S49 domain (IPR002142) from residues 20–236. InterPro entry (IPR002142) describes proteolytic enzymes that exploit serine in their catalytic activity. The sequences of these ubiquitous enzymes are variously annotated in different taxonomic groups. In plants, these proteolytic sequences are annotated as Signal peptide peptidase A (SppA; protease IV; MEROPS identifier S49.001) which are involved in cleavage of signal peptides.
- NFR-13 mutant contains a point mutation resulting in Ala80Thr on Acylamino-acid-releasing enzyme-like (g7937). Acylamino-acid-releasing enzymes have been identified and characterized in plants. They are found to be mainly localized in the stroma of chloroplasts, and are possibly involved in degradation of glycated/oxidized proteins, such as glycated RuBisCO (ribulose-1,5-bisphosphate carboxylase/oxygenase), and thus contribute to sustain the chloroplast antioxidative system [90].
4. Conclusions
5. Materials and Methods
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Benedetti, M.; Vecchi, V.; Barera, S.; Dall’Osto, L. Biomass from microalgae: The potential of domestication towards sustainable biofactories. Microb. Cell Fact. 2018, 17, 173. [Google Scholar] [CrossRef] [PubMed]
- Nelson, N.; Ben-Shem, A. The complex architecture of oxygenic photosynthesis. Nat. Rev. Mol. Cell Biol. 2004, 5, 239–250. [Google Scholar] [CrossRef]
- Weyer, K.M.; Bush, D.R.; Darzins, A.; Willson, B.D. Theoretical maximum algal oil production. Bioenergy Res. 2010, 3, 204–213. [Google Scholar] [CrossRef]
- Chisti, Y. Biodiesel from microalgae. Biotechnol. Adv. 2007, 25, 294–306. [Google Scholar] [CrossRef]
- Rodolfi, L.; Chini Zittelli, G.; Bassi, N.; Padovani, G.; Biondi, N.; Bonini, G.; Tredici, M.R. Microalgae for oil: Strain selection, induction of lipid synthesis and outdoor mass cultivation in a low-cost photobioreactor. Biotechnol. Bioeng. 2009, 102, 100–112. [Google Scholar] [CrossRef] [PubMed]
- Woittiez, L.S.; van Wijk, M.T.; Slingerland, M.; van Noordwijk, M.; Giller, K.E. Yield gaps in oil palm: A quantitative review of contributing factors. Eur. J. Agron. 2017, 83, 57–77. [Google Scholar] [CrossRef]
- Leu, S.; Boussiba, S. Advances in the Production of High-Value Products by Microalgae. Ind. Biotechnol. 2014, 10, 169–183. [Google Scholar] [CrossRef]
- Chew, K.W.; Yap, J.Y.; Show, P.L.; Suan, N.H.; Juan, J.C.; Ling, T.C.; Lee, D.-J.; Chang, J.-S. Microalgae biorefinery: High value products perspectives. Bioresour. Technol. 2017, 229, 53–62. [Google Scholar] [CrossRef]
- Scott, S.A.; Davey, M.P.; Dennis, J.S.; Horst, I.; Howe, C.J.; Lea-Smith, D.J.; Smith, A.G. Biodiesel from algae: Challenges and prospects. Curr. Opin. Biotechnol. 2010, 21, 277–286. [Google Scholar] [CrossRef] [PubMed]
- Christenson, L.; Sims, R. Production and harvesting of microalgae for wastewater treatment, biofuels, and bioproducts. Biotechnol. Adv. 2011, 29, 686–702. [Google Scholar] [CrossRef] [PubMed]
- Pittman, J.K.; Dean, A.P.; Osundeko, O. The potenrial of sustainable algal biofuel production using wastewater resources. Bioresour. Technol. 2011, 102, 17–25. [Google Scholar] [CrossRef]
- Munkel, R.; Schmid-Staiger, U.; Werner, A.; Hirth, T. Optimization of outdoor cultivation in flat panel airlift reactors for lipid production by Chlorella vulgaris. Biotechnol. Bioeng. 2013, 110, 2882–2893. [Google Scholar] [CrossRef] [PubMed]
- Borowitzka, M.A. High-value products from microalgae—Their development and commercialisation. J. Appl. Phycol. 2013, 25, 743–756. [Google Scholar] [CrossRef]
- Chen, J.H.; Chen, C.Y.; Hasunuma, T.; Kondo, A.; Chang, C.H.; Ng, I.S.; Chang, J.S. Enhancing lutein production with mixotrophic cultivation of Chlorella sorokiniana MB-1-M12 using different bioprocess operation strategies. Bioresour. Technol. 2019, 278, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Novoveská, L.; Ross, M.E.; Stanley, M.S.; Pradelles, R.; Wasiolek, V.; Sassi, J.F. Microalgal carotenoids: A review of production, current markets, regulations, and future direction. Mar. Drugs 2019, 17, 640. [Google Scholar] [CrossRef]
- Nwachukwu, I.D.; Udenigwe, C.C.; Aluko, R.E. Lutein and zeaxanthin: Production technology, bioavailability, mechanisms of action, visual function, and health claim status. Trends Food Sci. Technol. 2016, 49, 74–84. [Google Scholar] [CrossRef]
- Chia, S.R.; Chew, K.W.; Show, P.L.; Yap, Y.J.; Ong, H.C.; Ling, T.C.; Chang, J.S. Analysis of Economic and Environmental Aspects of Microalgae Biorefinery for Biofuels Production: A Review. Biotechnol. J. 2018, 13. [Google Scholar] [CrossRef] [PubMed]
- Tredici, M.R. Photobiology of microalgae mass cultures: Understanding the tools for the next green revolution. Biofuels 2010, 1, 143–162. [Google Scholar] [CrossRef]
- Heaton, E.A.; Dohleman, F.G.; Long, S.P. Meeting US biofuel goals with less land: The potential of Miscanthus. Glob. Chang. Biol. 2008, 14, 2000–2014. [Google Scholar] [CrossRef]
- Stephenson, P.G.; Moore, C.M.; Terry, M.J.; Zubkov, M.V.; Bibby, T.S. Improving photosynthesis for algal biofuels: Toward a green revolution. Trends Biotechnol. 2011, 29, 615–623. [Google Scholar] [CrossRef]
- Ooms, M.D.; Dinh, C.T.; Sargent, E.H.; Sinton, D. Photon management for augmented photosynthesis. Nat. Commun. 2016, 7, 12699. [Google Scholar] [CrossRef]
- Ruban, A.V.; Johnson, M.P.; Duffy, C.D.P. The photoprotective molecular switch in the photosystem II antenna. Biochim. Biophys. Acta Bioenerg. 2012, 1817, 167–181. [Google Scholar] [CrossRef] [PubMed]
- Dall’Osto, L.; Bressan, M.; Bassi, R. Biogenesis of light harvesting proteins. Biochim. Biophys. Acta Bioenerg. 2015, 1847, 861–871. [Google Scholar] [CrossRef] [PubMed]
- Fuciman, M.; Enriquez, M.M.; Polívka, T.; Dallosto, L.; Bassi, R.; Frank, H.A. Role of xanthophylls in light harvesting in green plants: A Spectroscopic investigation of mutant LHCII and Lhcb pigment-protein complexes. J. Phys. Chem. B 2012, 116, 3834–3849. [Google Scholar] [CrossRef] [PubMed]
- Formighieri, C.; Franck, F.; Bassi, R. Regulation of the pigment optical density of an algal cell: Filling the gap between photosynthetic productivity in the laboratory and in mass culture. J. Biotechnol. 2012, 162, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Krieger-Liszkay, A.; Fufezan, C.; Trebst, A. Singlet oxygen production in photosystem II and related protection mechanism. Photosynth. Res. 2008, 98, 551–564. [Google Scholar] [CrossRef]
- Aro, E.M.; Virgin, I.; Andersson, B. Photoinhibition of Photosystem II. Inactivation, protein damage and turnover. BBA Bioenerg. 1993, 1143, 113–134. [Google Scholar] [CrossRef]
- Vecchi, V.; Barera, S.; Bassi, R.; Dall’osto, L. Potential and challenges of improving photosynthesis in algae. Plants 2020, 9, 67. [Google Scholar] [CrossRef]
- Melis, A. Solar energy conversion efficiencies in photosynthesis: Minimizing the chlorophyll antennae to maximize efficiency. Plant Sci. 2009, 177, 272–280. [Google Scholar] [CrossRef]
- Kulheim, C.; Agren, J.; Jansson, S. Rapid Regulation of Light Harvesting and Plant Fitness in the Field. Science 2002, 297, 91–93. [Google Scholar] [CrossRef]
- Zhu, X.G.; Ort, D.R.; Whitmarsh, J.; Long, S.P. The slow reversibility of photosystem II thermal energy dissipation on transfer from high to low light may cause large losses in carbon gain by crop canopies: A theoretical analysis. J. Exp. Bot. 2004, 55, 1167–1175. [Google Scholar] [CrossRef] [PubMed]
- Frank, H.A.; Cogdell, R.J. Carotenoids in photosynthesis. Photochem. Photobiol. 1996, 63, 257–264. [Google Scholar] [CrossRef]
- Dall’Osto, L.; Piques, M.; Ronzani, M.; Molesini, B.; Alboresi, A.; Cazzaniga, S.; Bassi, R. The Arabidopsis nox mutant lacking carotene hydroxylase activity reveals a critical role for xanthophylls in photosystem I biogenesis. Plant Cell 2013, 25, 591–608. [Google Scholar] [CrossRef]
- El Agamey, A.; Lowe, G.M.; McGarvey, D.J.; Mortensen, A.; Phillip, D.M.; Truscott, T.G.; Young, A.J. Carotenoid radical chemistry and antioxidant/pro-oxidant properties. Arch. Biochem. Biophys. 2004, 430, 37–48. [Google Scholar] [CrossRef]
- Dall’Osto, L.; Holt, N.E.; Kaligotla, S.; Fuciman, M.; Cazzaniga, S.; Carbonera, D.; Frank, H.A.; Alric, J.; Bassi, R. Zeaxanthin protects plant photosynthesis by modulating chlorophyll triplet yield in specific light-harvesting antenna subunits. J. Biol. Chem. 2012, 287, 41820–41834. [Google Scholar] [CrossRef] [PubMed]
- Dall’Osto, L.; Cazzaniga, S.; Guardini, Z.; Barera, S.; Benedetti, M.; Mannino, G.; Maffei, M.E.; Bassi, R. Combined resistance to oxidative stress and reduced antenna size enhance light-to-biomass conversion efficiency in Chlorella vulgaris cultures. Biotechnol. Biofuels 2019, 12, 221. [Google Scholar] [CrossRef]
- Day, J.G.; Slocombe, S.P.; Stanley, M.S. Overcoming biological constraints to enable the exploitation of microalgae for biofuels. Bioresour. Technol. 2012, 109, 245–251. [Google Scholar] [CrossRef]
- Dall’Osto, L.; Bassi, R.; Ruban, A. V Photoprotective Mechanisms: Carotenoids. In Advances in Plant Biology 5; Theg, S.M., Wollman, F.-A., Eds.; Plastid Biology; Springer Science+Business Media: New York, NY, USA, 2014; pp. 393–435. [Google Scholar]
- Cordero, B.F.; Obraztsova, I.; Couso, I.; Leon, R.; Vargas, M.A.; Rodriguez, H. Enhancement of Lutein Production in Chlorella sorokiniana (Chorophyta) by Improvement of Culture Conditions and Random Mutagenesis. Mar. Drugs 2011, 9, 1607–1624. [Google Scholar] [CrossRef]
- Chamovitz, D.; Pecker, I.; Hirschberg, J. The molecular basis of resistance to the herbicide norflurazon. Plant Mol. Biol. 1991, 16, 967–974. [Google Scholar] [CrossRef]
- Del Campo, J.A.; García-González, M.; Guerrero, M.G. Outdoor cultivation of microalgae for carotenoid production: Current state and perspectives. Appl. Microbiol. Biotechnol. 2007, 74, 1163–1174. [Google Scholar] [CrossRef]
- Malkin, S.; Armond, P.A.; Mooney, H.A.; Fork, D.C. Photosystem II photosynthetic unit sizes from fluorescence induction in leaves. Correlation to photosynthetic capacity. Plant Physiol. 1981, 67, 570–579. [Google Scholar] [CrossRef]
- De Marchin, T.; Ghysels, B.; Nicolay, S.; Franck, F. Analysis of PSII antenna size heterogeneity of Chlamydomonas reinhardtii during state transitions. Biochim. Biophys. Acta Bioenerg. 2014, 1837, 121–130. [Google Scholar] [CrossRef]
- Prasad, A.; Ferretti, U.; Sedlaová, M.; Pospíšil, P. Singlet oxygen production in Chlamydomonas reinhardtii under heat stress. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Del Campo, J.A.; Rodríguez, H.; Moreno, J.; Vargas, M.Á.; Rivas, J.; Guerrero, M.G. Accumulation of astaxanthin and lutein in Chlorella zofingiensis (Chlorophyta). Appl. Microbiol. Biotechnol. 2004, 64, 848–854. [Google Scholar] [CrossRef]
- Kirst, H.; Garcia-Cerdan, J.G.; Zurbriggen, A.; Ruehle, T.; Melis, A. Truncated Photosystem Chlorophyll Antenna Size in the Green Microalga Chlamydomonas reinhardtii upon Deletion of the TLA3-CpSRP43 Gene. Plant Physiol. 2012, 160, 2251–2260. [Google Scholar] [CrossRef]
- Cazzaniga, S.; Dall’Osto, L.; Szaub, J.; Scibilia, L.; Ballottari, M.; Purton, S.; Bassi, R. Domestication of the green alga Chlorella sorokiniana: Reduction of antenna size improves light-use efficiency in a photobioreactor. Biotechnol. Biofuels 2014, 7, 157. [Google Scholar] [CrossRef]
- Perin, G.; Bellan, A.; Segalla, A.; Meneghesso, A.; Alboresi, A.; Morosinotto, T. Generation of random mutants to improve light-use efficiency of Nannochloropsis gaditana cultures for biofuel production. Biotechnol. Biofuels 2015, 8, 161. [Google Scholar] [CrossRef]
- Niyogi, K.K. Safety valves for photosynthesis. Curr. Opin. Plant Biol. 2000, 3, 455–460. [Google Scholar] [CrossRef]
- Park, S.; Steen, C.J.; Lyska, D.; Fischer, A.L.; Endelman, B.; Iwai, M.; Niyogi, K.K.; Fleming, G.R. Chlorophyll–carotenoid excitation energy transfer and charge transfer in Nannochloropsis oceanica for the regulation of photosynthesis. Proc. Natl. Acad. Sci. USA 2019, 116, 3385–3390. [Google Scholar] [CrossRef]
- Solovchenko, A.; Khozin-Goldberg, I.; Recht, L.; Boussiba, S. Stress-induced changes in optical properties, pigment and fatty acid content of Nannochloropsis sp.: Implications for non-destructive assay of total fatty acids. Mar. Biotechnol. 2011, 13, 527–535. [Google Scholar] [CrossRef]
- Hu, Q.; Sommerfeld, M.; Jarvis, E.; Ghirardi, M.; Posewitz, M.; Seibert, M.; Darzins, A. Microalgal triacylglycerols as feedstocks for biofuel production: Perspectives and advances. Plant J. 2008, 54, 621–639. [Google Scholar] [CrossRef]
- Menon, K.R.; Balan, R.; Suraishkumar, G.K. Stress induced lipid production in Chlorella vulgaris: Relationship with specific intracellular reactive species levels. Biotechnol. Bioeng. 2013, 110, 1627–1636. [Google Scholar] [CrossRef]
- Gimpel, J.A.; Henríquez, V.; Mayfield, S.P. In Metabolic Engineering of Eukaryotic Microalgae: Potential and Challenges Come with Great Diversity. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef]
- Richmond, A.; Cheng-Wu, Z.; Zarmi, Y. Efficient use of strong light for high photosynthetic productivity: Interrelationships between the optical path, the optimal population density and cell-growth inhibition. In Proceedings of the Biomolecular Engineering; Elsevier: Amsterdam, The Netherlands, 2003; Volume 20, pp. 229–236. [Google Scholar]
- Polle, J.E.W.; Kanakagiri, S.-D.; Melis, A. tla1, a DNA insertional transformant of the green alga Chlamydomonas reinhardtii with a truncated light-harvesting chlorophyll antenna size. Planta 2003, 217, 49–59. [Google Scholar] [CrossRef]
- Yang, Y.; Gao, K. Effects of CO2 concentrations on the freshwater microalgae, Chlamydomonas reinhardtii, Chlorella pyrenoidosa and Scenedesmus obliquus (Chlorophyta). J. Appl. Phycol. 2003, 15, 379–389. [Google Scholar] [CrossRef]
- Domingues, N.; Matos, A.R.; da Silva, J.M.; Cartaxana, P. Response of the Diatom Phaeodactylum tricornutum to photooxidative stress resulting from high light exposure. PLoS ONE 2012, 7, e38162. [Google Scholar] [CrossRef]
- Carvalho, A.P.; Silva, S.O.; Baptista, J.M.; Malcata, F.X. Light requirements in microalgal photobioreactors: An overview of biophotonic aspects. Appl. Microbiol. Biotechnol. 2011, 89, 1275–1288. [Google Scholar] [CrossRef]
- Tikkanen, M.; Grieco, M.; Nurmi, M.; Rantala, M.; Suorsa, M.; Aro, E.M. Regulation of the photosynthetic apparatus under fluctuating growth light. Philos. Trans. R. Soc. B-Biol. Sci. 2012, 367, 3486–3493. [Google Scholar] [CrossRef]
- Li, Z.R.; Wakao, S.; Fischer, B.B.; Niyogi, K.K. Sensing and responding to excess Light. Ann. Rev. Plant Biol. 2009, 60, 239–260. [Google Scholar] [CrossRef]
- Havaux, M.; Dall’Osto, L.; Bassi, R. Zeaxanthin has enhanced antioxidant capacity with respect to all other xanthophylls in Arabidopsis leaves and functions independent of binding to PSII antennae. Plant Physiol. 2007, 145, 1506–1520. [Google Scholar] [CrossRef]
- Erickson, E.; Wakao, S.; Niyogi, K.K. Light stress and photoprotection in Chlamydomonas reinhardtii. Plant J. 2015, 82, 449–465. [Google Scholar] [CrossRef]
- Meagher, E.; Rangsrikitphoti, P.; Faridi, B.; Zamzam, G.; Durnford, D.G. Photoacclimation to high-light stress in Chlamydomonas reinhardtii during conditional senescence relies on generating pH-dependent, high-quenching centres. Plant Physiol. Biochem. 2021, 158, 136–145. [Google Scholar] [CrossRef]
- Kondo, T.; Pinnola, A.; Chen, W.J.; Dall’Osto, L.; Bassi, R.; Schlau-Cohen, G.S. Single-molecule spectroscopy of LHCSR1 protein dynamics identifies two distinct states responsible for multi-timescale photosynthetic photoprotection. Nat. Chem. 2017, 9, 772. [Google Scholar] [CrossRef]
- Baroli, I.; Gutman, B.L.; Ledford, H.K.; Shin, J.W.; Chin, B.L.; Havaux, M.; Niyogi, K.K. Photo-oxidative stress in a xanthophyll-deficient mutant of Chlamydomonas. J. Biol. Chem. 2004, 279, 6337–6344. [Google Scholar] [CrossRef]
- Fiore, A.; Dall’Osto, L.; Cazzaniga, S.; Diretto, G.; Giuliano, G.; Bassi, R. A quadruple mutant of Arabidopsis reveals a β-carotene hydroxylation activity for LUT1/CYP97C1 and a regulatory role of xanthophylls on determination of the PSI/PSII ratio. BMC Plant Biol. 2012, 12, 50. [Google Scholar] [CrossRef] [PubMed]
- Ledford, H.K.; Chin, B.L.; Niyogi, K.K. Acclimation to singlet oxygen stress in Chlamydomonas reinhardtii. Eukaryot.Cell 2007, 6, 919–930. [Google Scholar] [CrossRef]
- Shao, N.; Krieger-Liszkay, A.; Schroda, M.; Beck, C.F. A reporter system for the individual detection of hydrogen peroxide and singlet oxygen: Its use for the assay of reactive oxygen species produced in vivo. Plant J. 2007, 50, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Fischer, B.B.; Dayer, R.; Schwarzenbach, Y.; Lemaire, S.D.; Behra, R.; Liedtke, A.; Eggen, R.I.L. Function and regulation of the glutathione peroxidase homologous gene GPXH/GPX5 in Chlamydomonas reinhardtii. Plant Mol. Biol. 2009, 71, 569–583. [Google Scholar] [CrossRef]
- Kleine, T.; Leister, D. Retrograde signaling: Organelles go networking. Biochim. Biophys. Acta Bioenerg. 2016, 1857, 1313–1325. [Google Scholar] [CrossRef]
- Ramel, F.; Birtic, S.; Ginies, C.; Soubigou-Taconnat, L.; Triantaphylides, C.; Havaux, M. Carotenoid oxidation products are stress signals that mediate gene responses to singlet oxygen in plants. Proc. Natl. Acad. Sci. USA 2012, 109, 5535–5540. [Google Scholar] [CrossRef]
- D’Alessandro, S.; Havaux, M. Sensing β-carotene oxidation in photosystem II to master plant stress tolerance. New Phytol. 2019, 223, 1776–1783. [Google Scholar] [CrossRef] [PubMed]
- García-Plazaola, J.I.; Portillo-Estrada, M.; Fernández-Marín, B.; Kännaste, A.; Niinemets, Ü. Emissions of carotenoid cleavage products upon heat shock and mechanical wounding from a foliose lichen. Environ. Exp. Bot. 2017, 133, 87–97. [Google Scholar] [CrossRef]
- Wase, N.; Black, P.N.; Stanley, B.A.; Dirusso, C.C. Integrated quantitative analysis of nitrogen stress response in Chlamydomonas reinhardtii using metabolite and protein profiling. J. Proteome Res. 2014, 13, 1373–1396. [Google Scholar] [CrossRef] [PubMed]
- Guarnieri, M.T.; Nag, A.; Smolinski, S.L.; Darzins, A.; Seibert, M.; Pienkos, P.T. Examination of triacylglycerol biosynthetic pathways via de novo transcriptomic and proteomic analyses in an unsequenced microalga. PLoS ONE 2011, 6, e25851. [Google Scholar] [CrossRef]
- Jacquier, N.; Choudhary, V.; Mari, M.; Toulmay, A.; Reggiori, F.; Schneiter, R. Lipid droplets are functionally connected to the endoplasmic reticulum in Saccharomyces cerevisiae. J. Cell Sci. 2011, 124, 2424–2437. [Google Scholar] [CrossRef] [PubMed]
- Sahni, P.; Aggarwal, P.; Sharma, S.; Singh, B. Nuances of microalgal technology in food and nutraceuticals: A review. Nutr. Food Sci. 2019, 49, 866–885. [Google Scholar] [CrossRef]
- Lafarga, T. Effect of microalgal biomass incorporation into foods: Nutritional and sensorial attributes of the end products. Algal Res. 2019, 41, 101566. [Google Scholar] [CrossRef]
- Zimny, T.; Sowa, S.; Tyczewska, A.; Twardowski, T. Certain new plant breeding techniques and their marketability in the context of EU GMO legislation—Recent developments. New Biotechnol. 2019, 51, 49–56. [Google Scholar] [CrossRef]
- Del Campo, J.A.; Moreno, J.; Rodríguez, H.; Angeles Vargas, M.; Rivas, J.; Guerrero, M.G. Carotenoid content of chlorophycean microalgae: Factors determining lutein accumulation in Muriellopsis sp. (Chlorophyta). J. Biotechnol. 2000, 76, 51–59. [Google Scholar] [CrossRef]
- Janßen, H.J.; Steinbüchel, A. Fatty acid synthesis in Escherichia coli and its applications towards the production of fatty acid based biofuels. Biotechnol. Biofuels 2014, 7. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Ye, L.; Zhao, D.; Li, S.; Li, Q.; Zhang, B.; Bi, C.; Zhang, X. Membrane engineering—A novel strategy to enhance the production and accumulation of β-carotene in Escherichia coli. Metab. Eng. 2017, 43, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Ling, Q.; Huang, W.; Baldwin, A.; Jarvis, P. Chloroplast biogenesis is regulated by direct action of the ubiquitin-proteasome system. Science 2012, 338, 655–659. [Google Scholar] [CrossRef]
- Ling, Q.; Jarvis, P. Regulation of chloroplast protein import by the ubiquitin E3 ligase SP1 is important for stress tolerance in plants. Curr. Biol. 2015, 25, 2527–2534. [Google Scholar] [CrossRef]
- Formighieri, C.; Cazzaniga, S.; Kuras, R.; Bassi, R. Biogenesis of photosynthetic complexes in the chloroplast of Chlamydomonas reinhardtii requires ARSA1, a homolog of prokaryotic arsenite transporter and eukaryotic TRC40 for guided entry of tail-anchored proteins. Plant J. 2012, 73, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Farkas, I.; Dombrádi, V.; Miskei, M.; Szabados, L.; Koncz, C. Arabidopsis PPP family of serine/threonine phosphatases. Trends Plant Sci. 2007, 12, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Park, J.J.; Wang, H.; Gargouri, M.; Deshpande, R.R.; Skepper, J.N.; Holguin, F.O.; Juergens, M.T.; Shachar-Hill, Y.; Hicks, L.M.; Gang, D.R. The response of Chlamydomonas reinhardtii to nitrogen deprivation: A systems biology analysis. Plant J. 2015, 81, 611–624. [Google Scholar] [CrossRef]
- Neily, M.H.; Matsukura, C.; Maucourt, M.; Bernillon, S.; Deborde, C.; Moing, A.; Yin, Y.G.; Saito, T.; Mori, K.; Asamizu, E.; et al. Enhanced polyamine accumulation alters carotenoid metabolism at the transcriptional level in tomato fruit over-expressing spermidine synthase. J. Plant Physiol. 2011, 168, 242–252. [Google Scholar] [CrossRef] [PubMed]
- Nakai, A.; Yamauchi, Y.; Sumi, S.; Tanaka, K. Role of acylamino acid-releasing enzyme/oxidized protein hydrolase in sustaining homeostasis of the cytoplasmic antioxidative system. Planta 2012, 236, 427–436. [Google Scholar] [CrossRef]
- Angstenberger, M.; De Signori, F.; Vecchi, V.; Dall’Osto, L.; Bassi, R. Cell Synchronization Enhances Nuclear Transformation and Genome Editing via Cas9 Enabling Homologous Recombination in Chlamydomonas reinhardtii. ACS Synth. Biol. 2020, 9, 2840–2850. [Google Scholar] [CrossRef]
- Li, X.; Patena, W.; Fauser, F.; Jinkerson, R.E.; Saroussi, S.; Meyer, M.T.; Ivanova, N.; Robertson, J.M.; Yue, R.; Zhang, R.; et al. A genome-wide algal mutant library and functional screen identifies genes required for eukaryotic photosynthesis. Nat. Genet. 2019, 51, 627–635. [Google Scholar] [CrossRef]
- Kropat, J.; Hong-Hermesdorf, A.; Casero, D.; Ent, P.; Castruita, M.; Pellegrini, M.; Merchant, S.S.; Malasarn, D. A revised mineral nutrient supplement increases biomass and growth rate in Chlamydomonas reinhardtii. Plant J. 2011, 66, 770–780. [Google Scholar] [CrossRef] [PubMed]
- Allen, M.M. Simple conditions for growth of unicellular blue-green algae on plates. J.Phycol. 1968, 4, 1–4. [Google Scholar] [CrossRef]
- Gilmore, A.M.; Yamamoto, H.Y. Zeaxanthin formation and energy-dependent fluorescence quenching in pea chloroplasts under artificially mediated linear and cyclic electron transport. Plant Physiol. 1991, 96, 635–643. [Google Scholar] [CrossRef] [PubMed]
- Croce, R.; Canino, G.; Ros, F.; Bassi, R. Chromophore organization in the higher-plant photosystem II antenna protein CP26. Biochemistry 2002, 41, 7334–7343. [Google Scholar] [CrossRef]
- Schagger, H.; von Jagow, G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal. Biochem. 1987, 166, 368–379. [Google Scholar] [CrossRef]
- Towbin, H.; Staehelin, T.; Gordon, J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: Procedure and some applications. Proc. Natl. Acad. Sci. USA 1979, 76, 4350–4354. [Google Scholar] [CrossRef]
- Rappaport, F.; Beal, D.; Joliot, A.; Joliot, P. On the advantages of using green light to study fluorescence yield changes in leaves. Biochim. Biophys. Acta 2007, 1767, 56–65. [Google Scholar] [CrossRef] [PubMed]
- Maxwell, K.; Johnson, G.N. Chlorophyll fluorescence—A practical guide. J. Exp. Bot. 2000, 51, 659–668. [Google Scholar] [CrossRef]
- Havaux, M.; Eymery, F.; Porfirova, S.; Rey, P.; Dormann, P. Vitamin E protects against photoinhibition and photooxidative stress in Arabidopsis thaliana. Plant Cell 2005, 17, 3451–3469. [Google Scholar] [CrossRef] [PubMed]
- Axelsson, M.; Gentili, F. A single-step method for rapid extraction of total lipids from green microalgae. PLoS ONE 2014, 9, e89643. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Cecchin, M.; Marcolungo, L.; Rossato, M.; Girolomoni, L.; Cosentino, E.; Cuine, S.; Li-Beisson, Y.; Delledonne, M.; Ballottari, M. Chlorella vulgaris genome assembly and annotation reveals the molecular basis for metabolic acclimation to high light conditions. Plant J. 2019, 100, 1289–1305. [Google Scholar] [CrossRef]
- García-Alcalde, F.; Okonechnikov, K.; Carbonell, J.; Cruz, L.M.; Götz, S.; Tarazona, S.; Dopazo, J.; Meyer, T.F.; Conesa, A. Qualimap: Evaluating next-generation sequencing alignment data. Bioinformatics 2012, 28, 2678–2679. [Google Scholar] [CrossRef] [PubMed]
- Depristo, M.A.; Banks, E.; Poplin, R.; Garimella, K.V.; Maguire, J.R.; Hartl, C.; Philippakis, A.A.; Del Angel, G.; Rivas, M.A.; Hanna, M.; et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 2011, 43, 491–501. [Google Scholar] [CrossRef] [PubMed]
- Thorvaldsdóttir, H.; Robinson, J.T.; Mesirov, J.P. Integrative Genomics Viewer (IGV): High-performance genomics data visualization and exploration. Brief. Bioinform. 2013, 14, 178–192. [Google Scholar] [CrossRef] [PubMed]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Götz, S.; García-Gómez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef]
- Mitchell, A.L.; Attwood, T.K.; Babbitt, P.C.; Blum, M.; Bork, P.; Bridge, A.; Brown, S.D.; Chang, H.Y.; El-Gebali, S.; Fraser, M.I.; et al. InterPro in 2019: Improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res. 2019, 47, D351–D360. [Google Scholar] [CrossRef]
- Vaser, R.; Adusumalli, S.; Leng, S.N.; Sikic, M.; Ng, P.C. SIFT missense predictions for genomes. Nat. Protoc. 2016, 11, 1–9. [Google Scholar] [CrossRef]
- Armenteros, J.J.A.; Salvatore, M.; Emanuelsson, O.; Winther, O.; Von Heijne, G.; Elofsson, A.; Nielsen, H. Detecting sequence signals in targeting peptides using deep learning. Life Sci. Alliance 2019, 2. [Google Scholar] [CrossRef] [PubMed]
- Emanuelsson, O.; Nielsen, H.; Heijne, G. Von ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci. 1999, 8, 978–984. [Google Scholar] [CrossRef] [PubMed]

| Genotype | Chl/Cell (pg) | Car/Cell (fg) | Chl a/b | Fv/Fm | PSII Antenna Size (T2/3−1·103, ms−1) |
|---|---|---|---|---|---|
| WT | 0.26 ± 0.03 a | 57 ± 3 a | 2.67 ± 0.22 a | 0.67 ± 0.03 a | 6.74 ± 0.49 a |
| NFR-3 | 0.11 ± 0.02 b | 71 ± 3 b | 4.14 ± 0.35 b | 0.66 ± 0.03 a | 4.32 ± 0.22 b |
| NFR-13 | 0.07 ± 0.01 c | 65 ± 2 c | 4.92 ± 0.46 c | 0.65 ± 0.04 a | 3.19 ± 0.42 c |
| mol/100 mol Chl | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Genotype | Car/Cell (fg) | Chl a/b | Chl/Car | Neo | Viola | Anthera | Lute | Zea | α-Car | β-Car | |
| Control | WT | 57 ± 3 a | 2.7 ± 0.2 a | 4.3 ± 0.1 a | 2.8 ± 0.8 a | 2.8 ± 0.7 a | 0.1 ± 0.1 a | 13.5 ± 1.7 a | 1.3 ± 0.1 a | 0.5 ± 0.1 a | 1.9 ± 0.3 a |
| NFR-3 | 71 ± 3 b | 4.1 ± 0.3 b | 1.6 ± 0.1 c | 5.3 ± 1.3 b | 8.9 ± 1.7 b | 1.4 ± 0.2 c | 37.3 ± 8.7 c | 5.8 ± 1.1 c | 0.8 ± 0.2 a | 4.9 ± 1.0 b,c | |
| NFR-13 | 65 ± 2 c | 4.9 ± 0.4 b | 1.0 ± 0.3 d | 4.6 ± 0.5 b | 15.5 ± 3.0 c | 2.0 ± 0.5 c,d | 54.9 ± 5.0 d | 7.9 ± 1.6 c | 1.5 ± 0.2 b | 9.5 ± 1.6 c,d | |
| N-depleted | WT | 89 ± 4 d | 2.8 ± 0.1 a | 2.8 ± 0.1 e | 3.9 ± 0.5 b | 3.1 ± 0.4 a | 1.1 ± 0.2 c | 21.5 ± 0.3 b | 2.9 ± 0.3 b | 0.7 ± 0.1 a | 2.9 ± 0.1 b |
| NFR-3 | 121 ± 2 e | 4.1 ± 0.1 b | 0.9 ± 0.1 d | 14.7 ± 1.1 d | 18.8 ± 3.9 e | 3.1 ± 0.8 d | 52.9 ± 1.1 d | 11.2 ± 0.8 d | 1.8 ± 0.1 b | 8.4 ± 0.1 d | |
| NFR-13 | 160 ± 8 f | 4.8 ± 0.2 b | 0.4 ± 0.1 d | 27.7 ± 7.1 e | 40.0 ± 0.6 f | 4.7 ± 1.7 d | 116.3 ± 2.6 e | 21.9 ± 0.4 e | 2.9 ± 0.2 c | 19.8 ± 0.2 e | |
| Parameters. | WT | NFR-3 | NFR-13 |
|---|---|---|---|
| Half-saturation intensity (µmol photons m−2 s−1) | 110 ± 24 a | 128 ± 12 a | 241 ± 73 b |
| Pmax (µmol O2 mg Chl−1 h−1) | 96 ± 5 a | 225 ± 2 b | 323 ± 45 c |
| Respiration (µmol O2 mg Chl−1 h−1) | 25 ± 3 a | 52 ± 7 b | 73 ± 6 c |
| Respiration (fmol oxygen cell−1 h−1) | 6.4 ± 0.8 a | 5.7 ± 0.7 a | 5.1 ± 0.4 a |
| Pmax/respiration (relative units) | 3.9 ± 0.4 a | 4.4 ± 0.5 b | 4.4 ± 0.4 b |
| Genotype | Mean Increase of Biomass (g L−1 day−1) | µ (day−1) |
|---|---|---|
| WT | 0.43 ± 0.03 a | 1.87 ± 0.08 a |
| NFR-3 | 0.56 ± 0.03 b | 1.99 ± 0.04 b |
| NFR-13 | 0.56 ± 0.01 b | 1.99 ± 0.02 b |
| Sample | Total Number of Reads Produced | Total Number of Reads after Trimming | Mapped Reads | Mean Fold Coverage |
|---|---|---|---|---|
| WT | 38,406,276 | 25,308,558 | 25,157,713 | 65.9× |
| NFR-3 | 37,378,884 | 28,743,282 | 20,555,916 | 55.2× |
| NFR-13 | 17,628,716 | 11,931,178 | 7,249,676 | 18.6× |
| Mutant | Variant pos. | Ref/Alt | Region | CDS Effect [SIFT Prediction] | TargetP | ChloroP | Putative Gene Function (Gene ID) | |
|---|---|---|---|---|---|---|---|---|
| NFR-13 | SS_1:3929152 | G | A | Exon | Missense sub. (Arg598Gln) [SIFT: Tolerated] | noTP | - | Predicted protein (g380) |
| NFR-13 | SS_2:14375 | G | A | Exon | Missense sub. (Arg150Cys) [SIFT: Deleterious] | noTP | - | RNA helicase, activating signal cointegrator 1 (g5804) |
| NFR-13/NFR-3 | SS_2:5043375 | G | C | Exon | Missense sub. (Gln67His) [SIFT: Tolerated] | noTP | - | Predicted protein (g2438) |
| NFR-13 | SS_3:393571 | G | A | Exon | Missense sub. (Arg470Cys) [SIFT: Deleterious *] | noTP | - | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A beta isoform-like (g2544) |
| NFR-13/NFR-3 | SS_5:2569375 | G | C | Exon | Missense sub. (Glu178Asp) [SIFT: Deleterious] | noTP | cTP | Glycerol-3-phosphate acyltransferase 3 (g4271) |
| NFR-13 | SS_5:2624939 | C | T | Exon | Missense sub. (Ser119Leu) [SIFT: Deleterious] | SP | - | Serine threonine- kinase receptor R831 (g4284) |
| NFR-3 | SS_10:492953 | G | A | Exon | Missense sub. (Arg233Cys) [SIFT: Deleterious] | noTP | - | S49 family peptidase (g6424) |
| NFR-13 | SS_13:1005023 | G | A | Exon | Missense sub. (Ala80Thr) [SIFT: Tolerated] | mTP | cTP | Acylamino-acid-releasing enzyme-like (g7937) |
| NFR-3 | SS_14:552899 | G | A | Exon | Missense sub. (Gly391Asp) [SIFT: Deleterious] | noTP | - | Biosynthetic arginine decarboxylase (g9015) |
| NFR-13 | SS_14:1569725 | C | T | Exon | Missense sub. (Ala6Thr) [SIFT: Deleterious *] | noTP | - | Hypothetical protein (g8357) |
| NFR-13/NFR-3 | SS_18:1233529 | G | A | Exon | Missense sub. (Gly1235Arg) [SIFT: Deleterious *] | noTP | - | Hypothetical protein (g9528) |
| NFR-13/NFR-3 | SS_1:4651287 | G | A | Exon | Synonymous sub. | noTP | - | cold shock domain-containing protein (g211) |
| NFR-13/NFR-3 | SS_2:4256144 | C | T | Exon | Synonymous sub. | noTP | - | haloalkane dehalogenase (g2220) |
| NFR-13/NFR-3 | SS_2:4392343 | C | G | Exon | Synonymous sub. | noTP | - | solute carrier family 25 member 44 (g2251) |
| NFR-13/NFR-3 | SS_5:1173035 | G | A | Exon | Synonymous sub. | noTP | - | Large subunit GTPase 1 (g3878) |
| NFR-13/NFR-3 | PO_26F:57866 | C | T | Exon | Synonymous sub. | noTP | - | Predicted protein (g10631) |
| NFR-3 | SS _1:3944123 | C | T | Intron | No | mTP | cTP | Methionyl-tRNA synthetase (g376) |
| NFR-13/NFR-3 | SS_1:5342472 | G | A | Intron | No | noTP | cTP | Cleavage and polyadenylation specificity factor subunit 1 (g15) |
| NFR-13/NFR-3 | SS_2:4545066 | G | A | Intron | No | noTP | - | Putative Xaa-Pro aminopeptidase 3 (g2293) |
| NFR-13/NFR-3 | SS_2:4545075 | G | A | Intron | No | noTP | - | Putative Xaa-Pro aminopeptidase 3 (g2293) |
| NFR-13/NFR-3 | SS _3:1002678 | A | G | Intron | No | SP | - | Hypothetical protein (g2664) |
| NFR-13/NFR-3 | SS_5: 1971361 | AG | A | Intron | No | noTP | - | MAU2 chromatid cohesion factor-like protein (g4126) |
| NFR-3 | SS _5:2687539 | G | C | Intron | No | noTP | cTP | Serine/threonine-protein kinase sepA (g4294) |
| NFR-13/NFR-3 | SS_6: 1727681 | TC | T | Intron | No | cTP | cTP | Water chloroplastic-like (g4763) |
| NFR-13/NFR-3 | SS _13:303225 | G | A | Intron | No | noTP | cTP | Allophanate hydrolase (g7748) |
| NFR-3 | SS_13:1920503 | G | A | Intron | No | noTP | - | U3 small nucleolar ribonucleoprotein protein IMP4 (g8749) |
| NFR-3 | SS_7:737237 | G | C | Intron | No | noTP | - | Sodium:proline symporter (g5015) |
| NFR-3 | SS_8:332225 | C | T | Intron | No | noTP | cTP | MFS general substrate transporter (g5411) |
| NFR-13 | SS_1:269018 | C | A | Splice region | No | noTP | - | Preprotein translocase subunit SecA (g1481) |
| NFR-13/NFR-3 | SS_3:344676 | G | A | Splice donor | Yes | noTP | - | E3 Ubiquitin-protein Ligase SP1 related [PTHR47568:SF2] (g2532) |
| NFR-13/NFR-3 | SS_13:2646657 | G | A | Splice region | No | cTP | cTP | S1 motif domain-containing protein (g8549) |
| NFR-13 | SS_1:2991332 | G | T | Intergenic | No | noTP | - | Upstream of: Nuclear/nucleolar GTPase 2 (g672); Nuclear transport receptor (g671) |
| NFR-13/NFR-3 | SS_5:1304829 | C | T | Intergenic | No | noTP | - | Upstream of predicted protein ( g3913). Downstream of predicted protein (g3914) |
| NFR-3 | SS1:3529923 | A | AT | Intergenic | No | noTP | - | Downstream of transcription initiation factor IIB-2 (g501). Downstream of putative phosphatase 2C 35 (g500) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guardini, Z.; Dall’Osto, L.; Barera, S.; Jaberi, M.; Cazzaniga, S.; Vitulo, N.; Bassi, R. High Carotenoid Mutants of Chlorella vulgaris Show Enhanced Biomass Yield under High Irradiance. Plants 2021, 10, 911. https://doi.org/10.3390/plants10050911
Guardini Z, Dall’Osto L, Barera S, Jaberi M, Cazzaniga S, Vitulo N, Bassi R. High Carotenoid Mutants of Chlorella vulgaris Show Enhanced Biomass Yield under High Irradiance. Plants. 2021; 10(5):911. https://doi.org/10.3390/plants10050911
Chicago/Turabian StyleGuardini, Zeno, Luca Dall’Osto, Simone Barera, Mehrdad Jaberi, Stefano Cazzaniga, Nicola Vitulo, and Roberto Bassi. 2021. "High Carotenoid Mutants of Chlorella vulgaris Show Enhanced Biomass Yield under High Irradiance" Plants 10, no. 5: 911. https://doi.org/10.3390/plants10050911
APA StyleGuardini, Z., Dall’Osto, L., Barera, S., Jaberi, M., Cazzaniga, S., Vitulo, N., & Bassi, R. (2021). High Carotenoid Mutants of Chlorella vulgaris Show Enhanced Biomass Yield under High Irradiance. Plants, 10(5), 911. https://doi.org/10.3390/plants10050911







