1. Introduction
The DNA duplex, which is known as the main molecule of life, is a complex of two antiparallel complementary copolymers of four nucleotide units. Despite its simplicity, the chemical structure of DNA gives rise to a multitude of various distinct 3D structures including duplexes with different nucleotide pairing, triplexes, and quadruplexes. At the root of biologically important conformational variability of DNA is a nucleotide as a union of a rigid nitrogen base (purine, Pur, or pyrimidine, Pyr), which has nearly planar geometry, with conformationally flexible sugar (deoxyribose) and phosphate groups. The long chain of repeating sugar and phosphate subunits forms a chemically uniform and conformationally flexible sugar-phosphate backbone (SPB) that has four different bases uniformly attached to it. The nucleotide sequence unique for each living organism forms the DNA molecule, which enables the storage, replication, use, and evolutionary modification of genetic information.
The factors that allow the DNA double helix to act as a genetic information carrier have been the focus of many studies starting from the 1950s. Watson and Crick not only discovered the 3D structure of DNA in their groundbreaking early publications [
1,
2] but also described the biological significance of the double helix and discussed (though only conceptually at that time) the problems of copying genetic information. Their discovery gave impetus to intense research focusing on what Schrödinger [
3] called a marvel, that is, the reproduction and the very existence of life. Initial studies of the DNA molecule and its subunits have shown how surprisingly fit the molecular and 3D structures of DNA are to its biological functions. A lot of experimental and computational information obtained by various methods is now available in the literature. The wealth of quantitative structural information, including atom coordinates and mutual arrangement of subunits, is accumulated in the Nucleic Acid Database (NDB) [
4] and the Protein Data Bank (PDB) [
5]. Yet it is still unclear what properties of which subunits determine DNA characteristics that are responsible for its significant features essential for life. Deciphering the contribution of DNA subunits to the formation of the 3D structure and its variability as well as to biological functions is the important step toward the elucidation of DNA functions at the atomic level.
To approach this goal, we perform a computational study of minimal fragments of single-chain DNA and those of duplex DNA, as well as of their subunits by using quantum mechanics (QM) and molecular mechanics (MM) methods. Our previous studies [
6,
7,
8] revealed that the essential conformational characteristics of the widely studied and the most populated BI and AI conformational families of Watson–Crick duplexes (WCDs), including the sequence dependence of their 3D structure, preexist in the local energy minima of the elemental single-chain fragments, deoxydinucleoside monophosphates (dDMPs). Those computations uncovered the important sequence-dependent regularity in the superposition of neighbor bases, namely substantial superposition in Pur–Pur and Pur–Pyr sequences, and minor or negligible base overlap in Pyr–Pyr and Pyr–Pur ones. This regularity matches the experimental data; it can be reproduced by using various computational methods of QM and MM in spite of some inaccuracy inherent to each individual method, and it distinguishes the WCD from other polynucleotide structures, such as left-handed Z-form, Hoogsteen, and parallel-stranded duplexes, as well as from the fragments of WCD with mispairs [
9]. Based on limited data on optimized BI conformations of dDMPs [
6], we suggested that the uniform sugar-phosphate backbone plays a substantial role in the nucleotide sequence-dependence of the 3D structure of the WCD. More extended computations for BI and AI conformers [
8] have shown that the directionality and the preferable regions of the torsion angles of SPB, combined with the difference of purines from pyrimidines in ring shape, determine the above-mentioned regularity of sequence-dependence of WCD. We theorize that the correspondence of conformation regions of SPB torsion angles of BI and AI conformations to those of energy minima of separate minimal fragments of SPB (i.e., dDMPs without bases, which are substituted by hydrogen atoms) predetermine the regularities in the 3D structure formation. These conclusions have been extended to BII and AII conformation families as well as to minimal fragments of duplex, complementary dDMPs (cdDMPs) in our follow-up publications [
8,
9,
10].
Analysis of the conformational characteristics of DNA fragments deposited in NDB and PDB demonstrates the extreme variability of torsion angles of SPB. Schneider et al. [
11,
12,
13] perform an extensive analysis and classification of various dinucleotide steps of DNA fragments. Their recent publication [
13] defines 96 classes of dinucleotide conformers of DNA and RNA (designated as NtC, dinucleotide conformer class) grouped into 15 codes of the Conformational Alphabet of Nucleic Acids, CANA. The majority of NtCs for DNA correspond to the formation of a double helix. The well-known BI and AI conformations correspond to BB00 and AA00 NtCs, correspondingly; the first letter defines the 5′ nucleoside conformation type, and the second one defines the 3′ nucleoside conformation type. BA01 and BA05 NtCs denote different conformers with 5′-nucleoside of B-type (C2″endo or close to C2′-endo sugar) and 3′-nucleoside of A-type (c3′-endo sugar).
Torsion angles of dDMPs and cdDMPs continuously vary in relatively wide regions. Rather small and consistent variations of the torsions can result in large changes of the conformational parameters of the helices belonging to the same conformation family. As a result, each family contains a countless number of conformers. In this work, we extend our computations and analysis to conformational parameters of dDMPs corresponding to NtCs compatible with the formation of a double helix with Watson–Crick nucleoside pairs, i.e., with A:T and G:C complementary base pairs and an anti-orientation of base and deoxyribose in both nucleosides. These “non-canonical” (non-BI and non-AI) conformations of minimal fragments of DNA chains arise as a result of existence of various sets of regions of SPB torsions that satisfy Watson–Crick nucleoside pairing in the right-handed fragment of the double helix structure.
In our recent paper [
14], we performed computations of several minimal DNA fragments in which the sets of torsion angles are positioned in the regions different from classical BI and AI conformations. In it, we analyzed variations of torsion angles of SPB for different conformational classes and inspected changes in these angles after energy minimization. We found that the regularities of 3D structure formation, namely, the base–base superposition in dDMPs and cdDMPs, can differ from those in BI and AI conformation families depending on the dDMP class (NtC) that specifies the torsion set. Detailed consideration of these results enabled us to correlate the torsion angles in dDMP and conformations of the local energy minima of separate SPB with the appearance of those regularities. The analysis revealed that conformations of dDMPs and cdDMPs for those NtCs that have SPB torsion angles close to the conformations of the geometry-optimized local energy minima of separate minimal units of SPB exhibit sequence-dependent characteristics such as those observed in BI and AI families. In contrast, conformations of dDMPs and cdDMPs with SPB torsions are rather different from those of the local minima of separate SPB units and demonstrate more complex sequence-dependence regularities. For some of the NtC classes, the optimized geometries of dDMP or cdDMP do not structurally match the corresponding fragments of the longer duplex.
2. Materials and Methods
The molecular systems, the methods of computation, and the computational software used in this work closely resemble those used in our current publication series [
6,
7,
8,
9,
14]. We consider the minimal fragments of single-strand, deoxydinucleoside monophosphates (dDMP) neutralized with Na ion, complementary parts of a DNA duplex, cdDMPs, and the fragments of sugar-phosphate backbone (SPB) encountered in dDMPs and cdDMPs.
Figure 1 provides an example of dDMP accompanied by a designation of torsion angles. This system corresponds to dinucleotide fragments used by Cerny et al. in their analysis and classification of DNA fragments [
13]. dDMP encompasses a contiguous part of a DNA single strand between atom C5′ of the first nucleoside and atom O3′ of the second nucleoside followed by the termination of oxygen atoms at C5′ and C3′ ends by hydrogen atoms. The addition of Na ion neutralizes the phosphate group to maintain charge neutrality. The dDMP fragment includes all nearest-neighbor interactions of DNA subunits that are essential for the conformation of the entire polynucleotide strand. In addition to that, cdDMP as a minimal fragment of double helix includes all nearest-neighbor interactions happening between complementary strands. Conducting systematic analysis of larger fragments of various DNA conformations by using quantum mechanics (QM) methods would be computationally prohibitive. Such studies are limited to a few unique conformations and base sequences (see [
15] and references therein).
For each dDMP and cdDMP, an initial conformation was constructed using atomic coordinates of the corresponding fragments of a crystal structure obtained from the Nucleic Acid Database (NDB) [
4]. Hydrogen atoms and sodium ions were added in the positions most likely for a given atomic group. The minimal fragments of SPB were cut out from the respective dDMPs by substitution of the bases by hydrogen atoms.
The choice of computational method for the computer simulation of a 3D structure of nucleic acids is dictated by the need to balance the computational cost with achieving sufficiently accurate accounting of the contribution of various subunits. In our recent publication [
14], we discussed the problems in computer simulation of DNA 3D-structure formation and variability, and we explained the choice of computational methods. Here, we only mention a few of those problems and use the results reproducible by several methods to draw conclusions. The main QM method that we use for geometry optimization is based on the Density Functional Theory (DFT). This method provides reasonable compromise between the accuracy and computational cost. The main results have been obtained by using the Gaussian09 software package [
16] and PBEPBE [
17] and M05-2X [
18] functionals in combination with 6-31G* basis set, which is a valence double-zeta polarized basis set. All energy minima were confirmed by using frequency calculations. The first functional, PBEPBE, underestimates the dispersion interactions, which is common for many other DFT functionals. The second functional, M05-2X, is similar to M06-2X [
19] and many other functionals in the DFT-D group [
20]; it overestimates dispersion and leads to shortened atom–atom contacts of stacked bases. To minimize the possible errors in conclusions about various conformational classes of minimal fragments, we performed computations on select molecular systems by using the ADF (Amsterdam Density Functional) [
21,
22] package and a few additional functionals. The geometry optimization of all SPB fragments and selected dDMPs and cdDMPs has also been performed at the MP2/6-31G(d,p) level of theory. The MP2 method is insufficiently accurate for evaluation of base stacking even when using a large basis set, but it is suitable here, as we focus mainly on SPB conformations and their contribution to the variability of DNA 3D structure. We use the MM method for the purpose of preparation and preliminary optimization of several initial structures as well as for the evaluation of validity of force fields for reproducing QM results and for the identification of a suitable force field for the extended computation of many conformers of the simple fragments. For that purpose, we use the AMBER software package [
23] and three AMBER force fields, BSC1 [
24], OL15 [
25], and ff99 [
26], which are dubbed as MM, MM’, and MM”, respectively.
The computation of conformational characteristics of DNA fragments employs 3DNA software [
27]. In addition to torsion angles and parameters determining the mutual position of bases, we consider the superposition area of the rings of two bases of dDMP. It is quantified in 3DNA by the overlap area (in Å
2) of two base rings, which corresponds to the overlapped polygon of the two base rings projected onto the mean plane of the base normal. We do not discuss quantitative values that depend on the method used and consider the superposition areas qualitatively, i.e., small superposition (less than 1 Å
2, frequently negligible ring overlap) or substantial overlap (more than 2 Å
2). The assignment of NtCs to optimized dDMPs and cdDMPs uses DNATCO software [
28].
3. Results
3.1. Selection of Conformational Classes of DNA Minimal Fragments
From the variety of conformational classes of minimal units of the DNA chain (NtCs), we selected several NtCs compatible with the formation of Watson–Crick duplexes (WCDs), which have canonic A:T and G:C base pairs and both nucleosides in anti-conformation, for geometry optimization and for detailed analysis of structural characteristics. These classes differ one from another by the values of torsion angles of sugar-phosphate backbone (SPB), which (including glycoside torsions) are reproduced in
Table 1.
The list of selected NtCs includes BI, AI, BII, and AII conformation families that we considered earlier as well as a few additional classes for which we had preliminary data. In there, a few NtCs differ from BI (BB00) by no more than one torsion. For some of the NtCs from
Table 1, we previously obtained optimized geometries of dDMP and cdDMP with SPB torsions being close or different to the energy minimum of the separate SPB fragment. In general, an NtC in
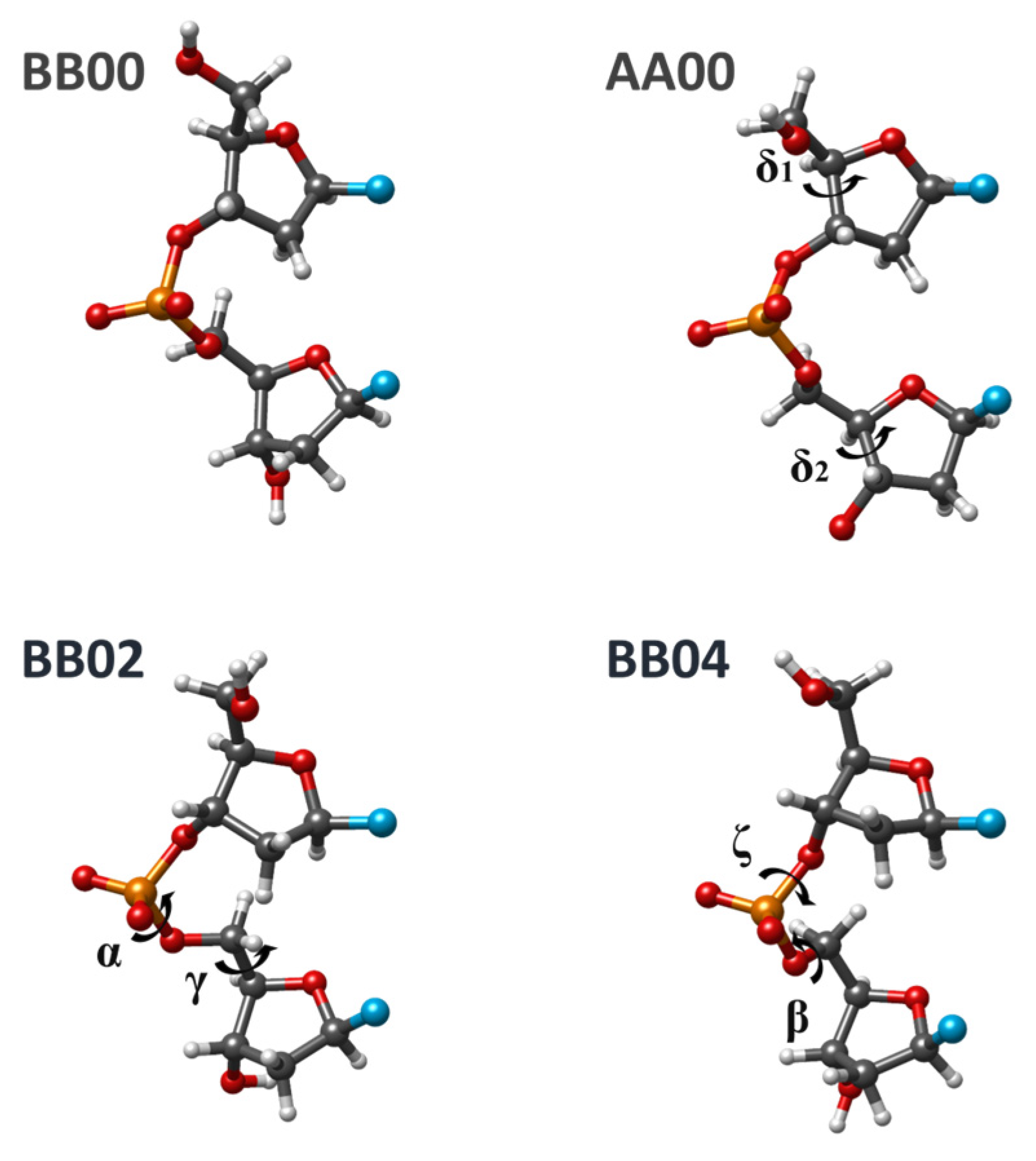
Table 1 differs from the BI form by more than 30° for one, two, three, four, or six torsion angles of SPB. Most of them are frequently encountered in the crystal structures of DNA fragments or DNA complexes with proteins. Four examples of conformation classes are shown in
Figure 2.
3.2. The Most Known DNA Conformations Revisited
Since the beginning of this work series, we detected several exceptions from the general conclusions about 3D structure formation related to sequence dependence for BI, AI, AII, and BII conformations. These observations refer to the superposition of neighbor bases in both experimental structures of duplexes and optimized dDMPs and cdDMPs. For BI and AI conformation families (BB00 and AA00 NtC), exceptions are rather rare; the BII family (BB07 NtC) includes many cases contradicting our previous conclusions. To refine our previous conclusions on the regularities of 3D structure formation, we perform extended DFT and MP2 optimizations of dDMPs, cdDMPs, and SPB fragments of these most known DNA structures. Optimizations with three AMBER force fields have been performed in order to evaluate the ability of the molecular mechanics (MM) method to reproduce the sequence-dependent regularities in the formation of a DNA 3D structure.
Examinations of dozens of experimental dDMPs and the energy optimization of corresponding dDMPs, cdDMPs, and SPB fragments pertaining to BB00 NtC by using QM methods demonstrate that a great majority of both experimental and calculated structures confirm the regularities in the 3D structure formation obtained earlier. The geometry optimization of separate SPBs of various dDMPs pertaining to BB00 NtC by using three QM methods produces three minimal energy structures, which differ one from another in energy and in each torsion angle by about 1–3 kcal/mol and by no more than a few degrees, respectively. The torsion angles in these structures match the averaged values for this NtC with the difference being mostly in the range of a single digit.
Table 2 lists the torsion angles for two QM optimized structures. Three tested AMBER force fields also reproduce the structural data for BB00 SPB (
Table 2). The puckering of both sugar rings remains in the C2′-endo region in all QM optimized structures and also in the structure optimized by using the BSC1 force field. Two other AMBER force fields produce somewhat distorted sugar rings and δ torsions, although the values remain in the vicinity of those for experimental structures. Distances between the C1′ atoms of two sugars in PBE and BSC1-optimized structures are rather close to the averaged experimental value. Variations in the computed distances are smaller than those in the experimental structures by about ±0.5 Å. The differences due to the other two force fields do not exceed 0.9 Å. The limited accuracy of both experimental and theoretical data makes a more detailed analysis of these variations impractical. The principal result that follows from the undertaken analysis of theoretical and experimental data for BB00 conformers is that it upholds our previous conclusions about the sequence dependence of base superposition and about the correspondence of SPB torsion angles to the local energy minima of separate SPB fragment. Exceptions from the general rule will be discussed further down.
Similar conclusions follow from an extended analysis of experimental and theoretical data for the AA00 NtC class, which represents the most populated A-family of DNA. Analysis of the AA01 NtC class (AII family) leads to minor changes to the previous conclusions. The geometry optimization of separate SPB fragments pertaining to this NtC produces two types of energy minima, those having torsion angles close to the experimental values (
Table 2) and the minima where some torsions differ from the averaged experimental values for AA01 NtC by more than 30°. AMBER force fields reproduce the tosion angles for the AA01 class except angle α, which differs from the experimental value by 61°.
The main changes that we need to make in our previous conclusions pertain to the BII conformation family. The recently compiled list of BB07 conformations includes many structures with torsion angles being rather different from the structures of energy minimum of a separate SPB fragment (
Table 2). The base superposition patterns observed in this family can be quite different from the patterns characteristic to the other three conformation classes considered above. Based on that, one may conclude that this NtC contains a mix of structures corresponding to regularities for BI, AI, and AII conformations intertwined with some other structures. The limited accuracy of our previous conclusions pertaining to the BII family was due to a restricted set of structures being available at that time. Some nucleotide sequences of BII conformations did not exist in the Nucleic Acid Database at the time of the analysis, and we manually constructed the missing structures by interchanging the base in the available sequences.
An example of a minimal fragment of the DNA duplex, which deviates from the general regularities, is cdDMP with two thymines in one strand (BB00 NtC) and two adenines in a complementary strand pertaining to BB07 NtC extracted from the DNA–protein complex, NDB ID PD0192. This cdDMP has a considerable superposition of two Thy bases (2.84 Å2) and zero overlap of two Ade bases. In it, the torsion angles of the thymine chain are rather close to the averaged experimental values of BB00 NtC. Geometry optimization of the SPB fragment representing the dTpdT portion of this cdDMP performed by using various computational methods produces one local energy minima matching the SPB structure previously obtained in the analysis of other structures of the BI conformation family. The geometry optimization of cdDMP and dTpdT dDMP quantitatively changes the respective torsion angles but preserves the substantial superposition in the dTpdT fragment. This example demonstrates that some sets of torsion angles characteristic for the BI DNA family are compatible with the dDMP structure that has a substantial overlap of two pyrimidines (and that may possibly extend to the Pyr–Pur sequence). Such structures can arise as a result of a simultaneous consistent variation in all or several torsions. These are rare exceptions from the general rule for BI, AI, and other conformational classes, which have torsion values positioned in the vicinity of the local energy minima of the SPB fragment.
3.3. Conformations of dDMP with a Sugar-Phosphate Backbone Corresponding to the Energy Minimum of a Separate SPB Fragment
In addition to the BI, AI, and AII conformations considered in the previous subsection, there are a few other NtCs with their SPB torsions being close to those in the energy minimum of the corresponding separate SPB minimal fragment. The values of SPB torsion angles representing a selected set of such fragments that are optimized by different computational methods are presented in
Table 3. This set includes BB01 NtC, which has conformational characteristics being very close to those of BB00. The optimized SPB fragments for this NtC correspond to the energy minima of BB00 NtC. Two other NtCs correspond to BA-type dDMPs; i.e., these structures have B-like 5′-end sugar puckering and A-like 3′-end sugar. Their SPB torsions differ from BB00 NtC only in δ2 value. A few optimized SPB structures from these NtCs correspond to one of the BB00 minima. Two more NtCs included in our study set, BB12 and BA09, correspond to a substantial change in α, β, and γ torsions. The BA09 NtC additionally differs in the value of the δ2 torsion. Several QM optimized BA09 SPB correspond to the energy-minimized BB12 structures. We added a subclass X of the structures “closely corresponding” to BB10 NtC but differ from it only by one torsion γ (
Table 3) to the present set of NtCs. The BB10 structures have angles ζ, α, and β being different from the BI family of conformations. We obtained several optimized structures of dDMPs and SPBs corresponding to this minimum two years before the publication of the modern version of the NtC classification. After the publication of the paper [
13], we added a few structures selected from the list of BB10 NtCs with γ torsion being close to this minimum. AMBER force fields do not reproduce this minimum for SPB fragments as well as the QM minima for BA09 SPBs. We realize that
Table 3 does not contain an exhaustive list of NtC classes corresponding to the energy minima of separate SPB fragments. The selected structures are considered as examples of sufficiently populated NtCs compatible with Watson–Crick duplexes.
The puckering of both sugar rings and C1′–C1′ distances in these fragments optimized by QM methods are close to those of the corresponding initial conformation. The changes in SPB conformation due to the PBE optimization of BB00 and BB12 classes are displayed in
Figure 3. For the most of the considered examples, SPB optimization with AMBER force fields produces the energy minimum structure being rather close to the starting one. In a few cases, geometry optimization by using an AMBER force field fails to reproduce the sequence-dependent regularity seen in QM calculations.
Analysis of the conformational characteristics of randomly selected initial dDMP structures upholds, for the vast majority of cases, our previous conclusion about the sequence-dependent regularities in the canonic BI and AI forms. The QM-optimized structures remain in the same NtC class as that of the initial structures and have the torsion angles changed due to optimization by no more than 30°. For these NtCs, important characteristics of the sequence dependence of DNA minimal fragments include the difference of Pur–Pur and Pur–Pyr sequences from Pyr–Pur and Pyr–Pyr ones.
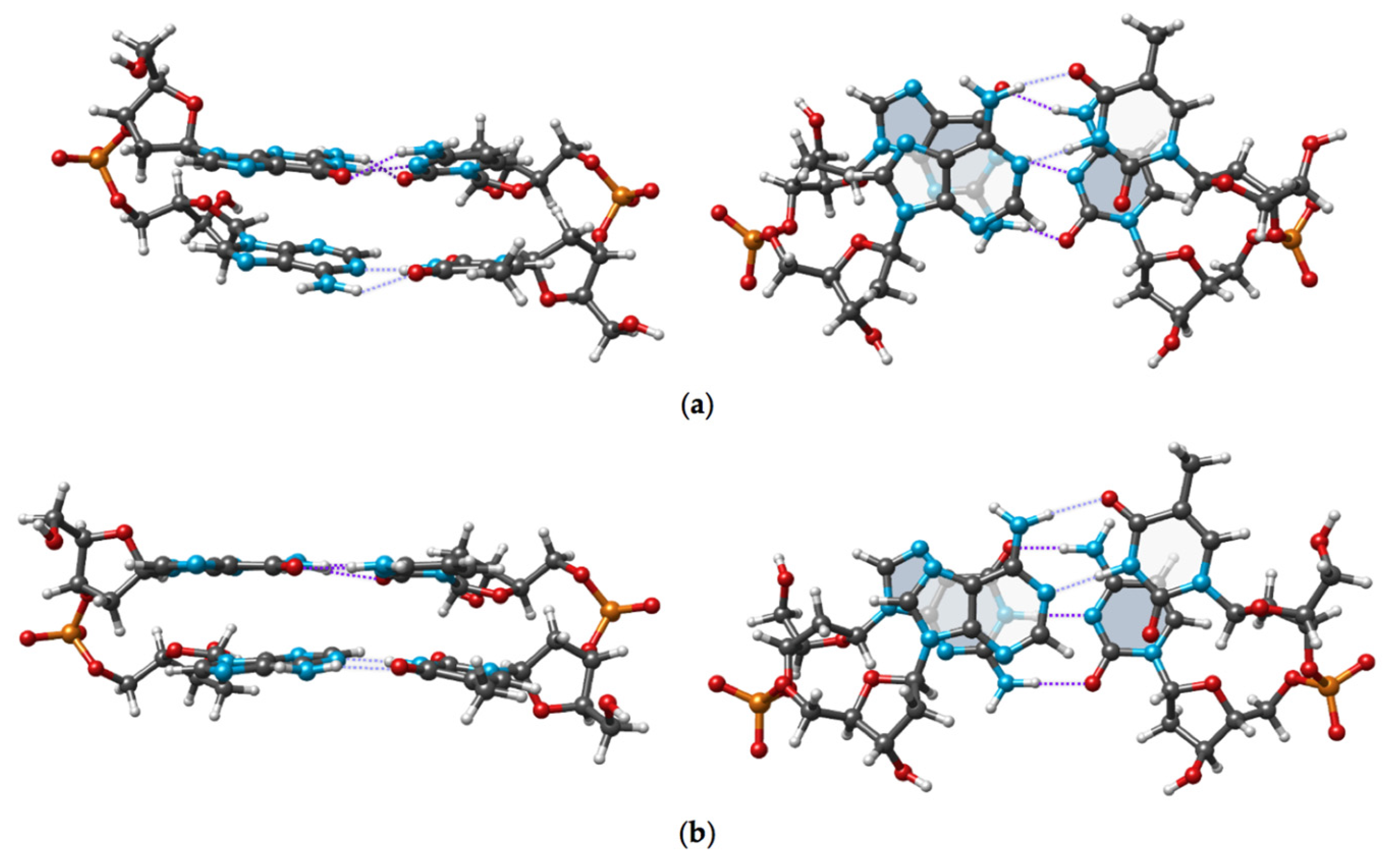
The experimental and DFT optimized conformations of cdDMP extracted from a DNA–nuclease complex with both complementary dDMPs being in the BB12 NtC class are displayed in
Figure 4. Geometry optimization retains the substantial overlap of Ade base rings and the negligible overlap of Thy rings, which is characteristic of the experimental structure.
Despite the overall uniform agreement taking place between the experiment and theory, for several of these NtCs, we encounter exceptions with a major superposition happening in a Pyr–Pyr sequence, e.g., in dCpdC belonging to BA09 NtC from the protein–DNA–DNTP ternary complex (NDB ID: PD1271) and dTpdT BA05 sequence from all-AT DNA dodecamer (NDB ID: 2122). This situation resembles that for a rather rare example from BB00 NtC considered in the previous subsection. Elucidation of the mechanism of such exceptions requires conducting an extended search for relevant examples and possibly involving more accurate computation methods.
3.4. Conformations of dDMPs with Sugar-Phosphate Backbone Deviating from the Energy Minima of a Separate SPB Fragment
It happens that a great number of dDMPs in Watson–Crick duplexes with nucleosides in anti-conformations have SPB torsion angles that are considerably different from the values of torsions in the nearest minima of a separate SPB fragment. The majority of dDMPs from the BII conformation family (BB07 NtC) pertain to this type of fragment. As we revealed earlier, the regularities of the sequence-dependent properties in these dDMPs and corresponding cdDMPs are rather different from those for BI, AI, and other dDMPs, which have torsion angles being close to the local energy minima of a separate SPB. In this work, we consider some examples of such NtCs of BB and BA types, i.e., dDMPs with both nucleosides having B-like sugar (BB type) and dDMPs with the first nucleoside having B-like sugar and the second one having A-like sugar (BA type). These NtCs differ from BI conformations by two, three, or six torsions. The torsion angles of such SPBs optimized by various computational methods as well as the averaged experimental data are presented in
Table 4.
The majority of the energy minima of separate SPBs in these NtCs are characterized by a considerable change happening in the C1′–C1′ distance (up to 2.5 Å) and in the mutual orientation of glycoside bonds as compared to experimental structures. Nevertheless, some energy minima have a C1′–C1′ distance that is rather close to the experimental value (despite great variation happening in one or two torsion angles). Two examples of conformational changes happening in separate SPB fragments upon geometry optimization are displayed in
Figure 5.
The geometry optimization of several SPB fragments collected in
Table 4 produces energy minima already seen with other NtCs considered in the previous subsection, while for other fragments of the same NtC, we obtain energy minima having the torsions in the regions rather different from those of other NtCs. For example, the SPB optimization data for three QM methods for several dDMPs pertaining to the BA08 conformation class match the energy minima obtained for BB00 and BA05 NtCs as well as introduce a new energy minimum. The geometry optimization of SPB of various dDMPs from BB04 NtC leads to a minimum matching that for BB00 (not included in
Table 4), another one matching the minimum for BB07, and it uncovers other minimal energy conformations that do not resemble any NtC for the considered conformations (
Table 4).
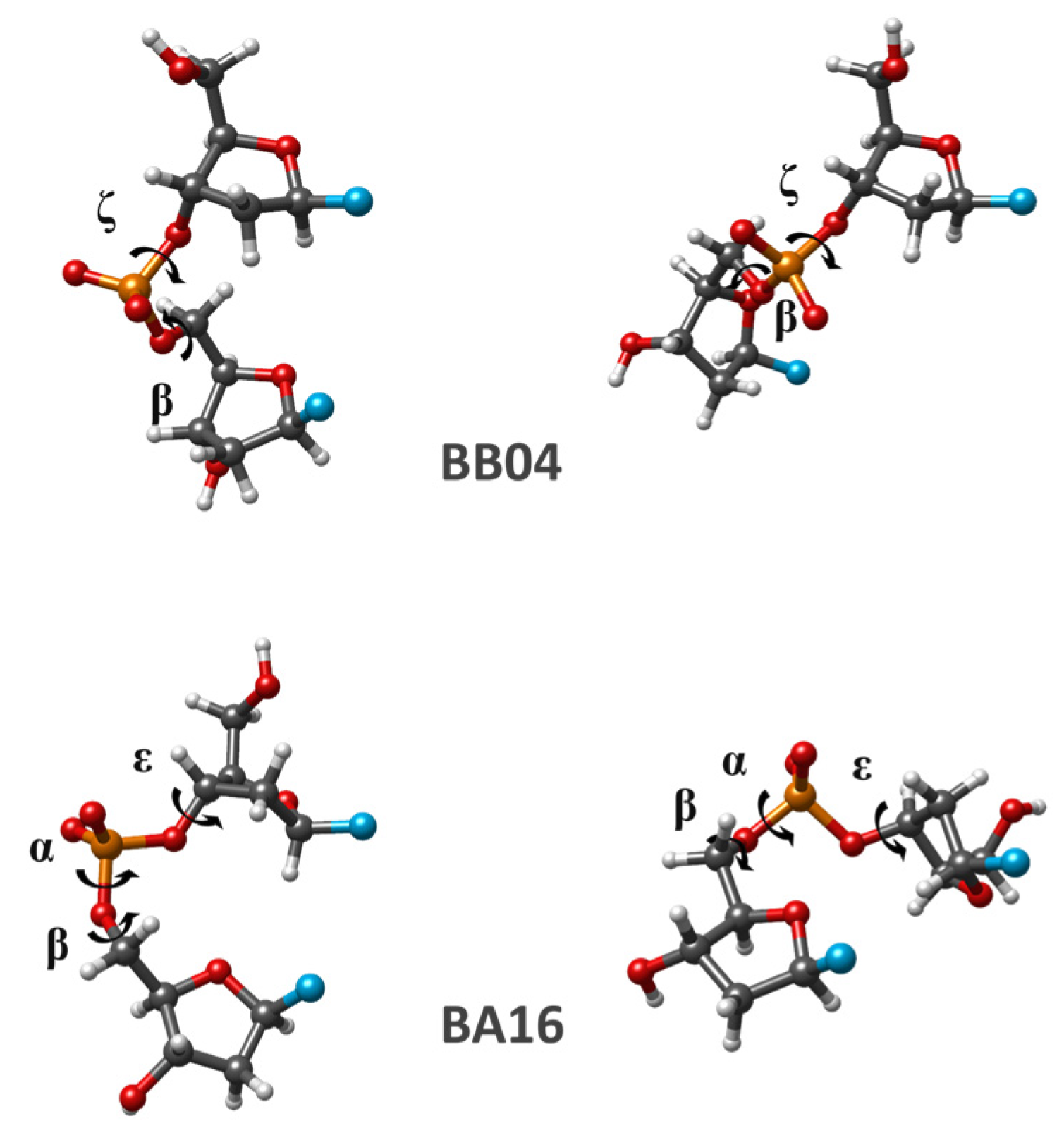
It is interesting to mention that for many dDMPs considered in this section, from our random selection of experimental structures, we observe the same regularity of the base superposition as that revealed for dDMPs when torsion angles correspond to the energy minima of separate SPBs. The geometry optimization of dDMP and cdDMP may retain or considerably change the structure. One example of the drastic change in the base superposition happening upon geometry optimization is displayed in
Figure 6. The conformation of dTpdT:dApdA with dTpdT pertaining to BB02 NtC and dApdA corresponding to BB04 NtC is extracted from the crystal structures of nucleosome core particles in a complex with minor groove DNA-binding ligands (NDB ID: PD0329). It has a substantial superposition of thymine rings and a minor superposition of adenine bases. DFT optimizations each using a different functional drastically change this conformation. Both optimized conformations (as well as many other optimized dDMP and cdDMP structures) have some deficiencies characterizing the methods, such as too short atom–atom contacts due to the overestimation of dispersion contribution by M05-2X functional and underestimation of these interactions in PBE functional, resulting in some distortion of the parallelity of base pairs. In any case, either computation clearly demonstrates the absence of Thy overlap and the presence of a substantial superposition of Ade rings. Since both very different DFT functionals consistently produce the same pattern of base superposition, we hypothesize that the disagreement between the theoretical and experimental data points to external factors existing outside of cdDMP that govern the specific superposition seen in the experimental data of the larger molecular system.
4. Discussion
This work continues our series of publications on the contribution of DNA subunits to its 3D structure formation and the regularities of the sequence dependence of the DNA double helix. The presence of new experimental data on the structure of DNA fragments and the availability of more extended quantum mechanics and molecular mechanics computations of minimal fragments of the DNA chain and DNA duplex lead to the need to refine our previous generalizations and conclusions. The new findings fully support our former conclusion about the important role of a chemically monotonous sugar-phosphate backbone (SPB) in the formation of a sequence-dependent WCD structure. The SPB predetermines important structural properties of the minimal fragments of the DNA single chain, dDMP. The correspondence of SPB torsions of dDMP (and NtCs) to the energy minima of a separate SPB fragment determines the sequence-dependent patterns of conformational classes and provides the foundation for conformational families of DNA fragments. The difference in the character of base-ring superposition in Pur–Pur and Pur–Pyr sequences from that in Pyr–Pur and Pyr–Pyr ones is a common regularity that goes beyond the most known BI and AI conformation families and extends to all classes of conformations that have torsion angles corresponding to the energy minima of separate SPB fragments. With that, all dDMP conformations (and NtC conformers) compatible with WCDs that have both nucleosides of the complementary pair in anti-conformation can be divided in two groups: namely, those with SPB torsions being close to the local energy minima of a separate SPB fragment and those having torsions of SPB being rather different from the energy minima of a separate SPB fragment. For the first type of dDMPs, the regularities of base-ring overlap are the same as those for the canonic BI form. For the second type, these regularities do not necessarily uphold, and the reason for that is not completely known. Some exceptions from the first type exist, they are rather rare, and will be discussed below.
The most notable change to the previously reported regularities comes from the present analysis of BII conformation family. All dDMPs pertaining to this family fall into two groups depending on the outcome of geometry optimization of a separate SPB fragment. Geometry optimization of the SPB fragment extracted from a dDMP belonging to the first group produces conformations that have all torsion angles being rather close to the initial values and to those averaged for this NtC. At the same time, the geometry optimization of separate SPB fragments extracted from the second group of dDMPs, which is the largest one, lead to one or more torsion angles significantly deviating from the target values by more than 50°. It shows that the SPB conformation in dDMPs from the first group, which is in the minority, corresponds to the energy minima of a separate SPB, while the majority of structures combined into the second group do not correspond to the energy minima of a separate SPB. This result corroborates with the observation that for the majority of dDMPs from our set of experimental data for this family, we see a minor overlap of base rings, including dDMPs with Pur–Pur and Pur–Pyr sequences.
An important detail coming from this work is that geometry optimization, by using any computational method, of several separate SPB fragments extracted from the dDMPs pertaining to the same NtC typically leads to two or more local energy minima. This observation applies to all NtCs studied in this work, and it sheds some light on the existence of the two dDMP types mentioned above. The latter two types diverge one from another due to the difference between the values of corresponding angles of various minima. The minima for the first dDMP type can differ in a few degrees only; this difference can be less than the difference between torsions of the same minimum calculated by different methods and between the initial and optimized conformations. The BI conformation family represents this particular case. For the second dDMP type, the difference for one or two torsions can exceed 50°. An example of that appears in BB04 NtC, which is a conformation class intermediate between those for BI and BII conformation families. For this NtC, we found three minima, one being the same as that for BB07, another having a unique torsion set, and the third minimum matching that for BB00 (not included in
Table 4). Another class exhibiting significant differences in torsion angles between the different minima is BA08 NtC. One of its minima corresponds to BB00 NtC, another one to BA05, and the third minimum is unique (
Table 4).