The Power of First Impressions: Can Influenza Imprinting during Infancy Inform Vaccine Design?
Abstract
1. Introduction
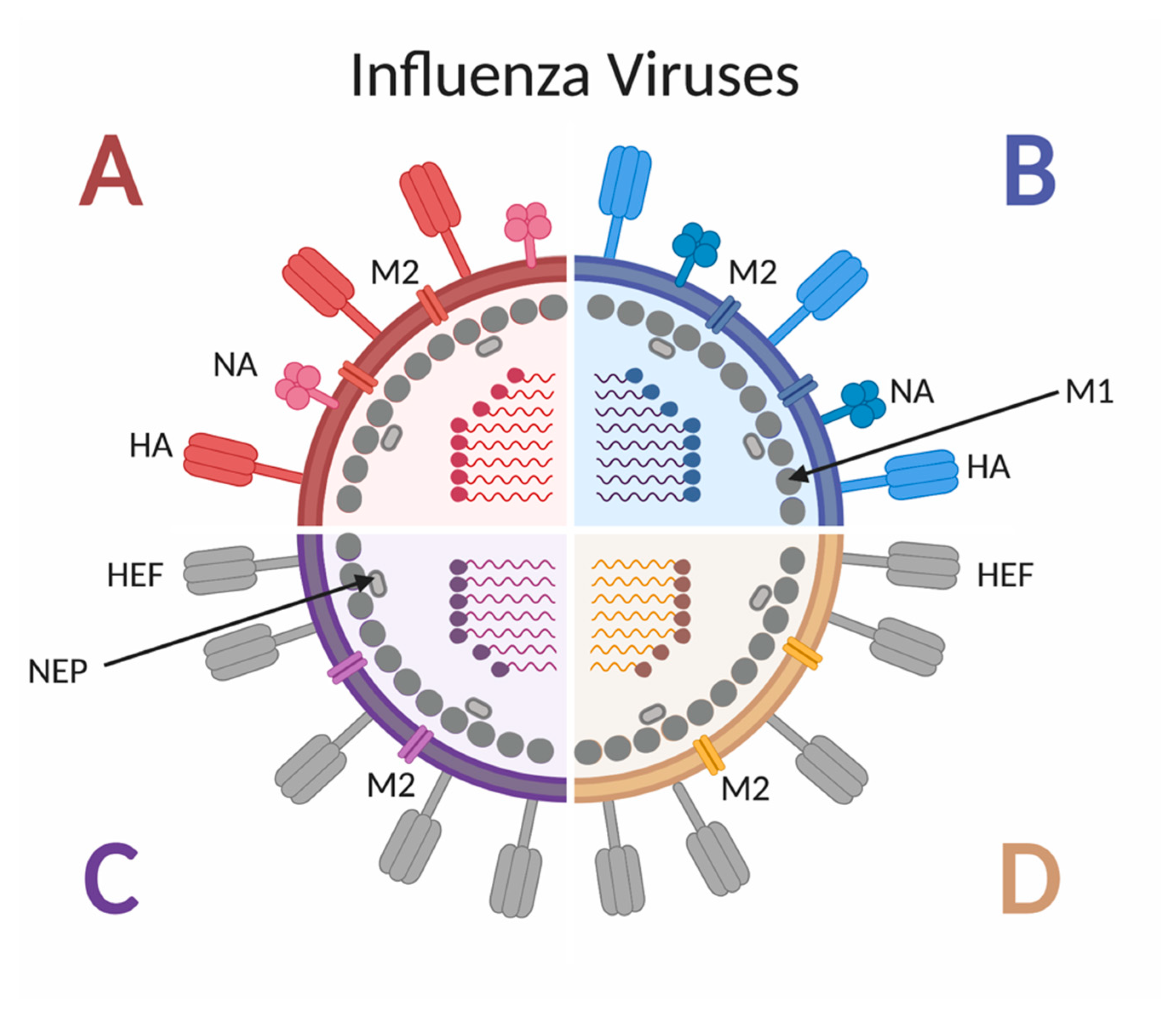
2. Influenza: The Virus, Disease, Vaccine and Children
3. The Predictive Power of Influenza Virus Immune Imprinting
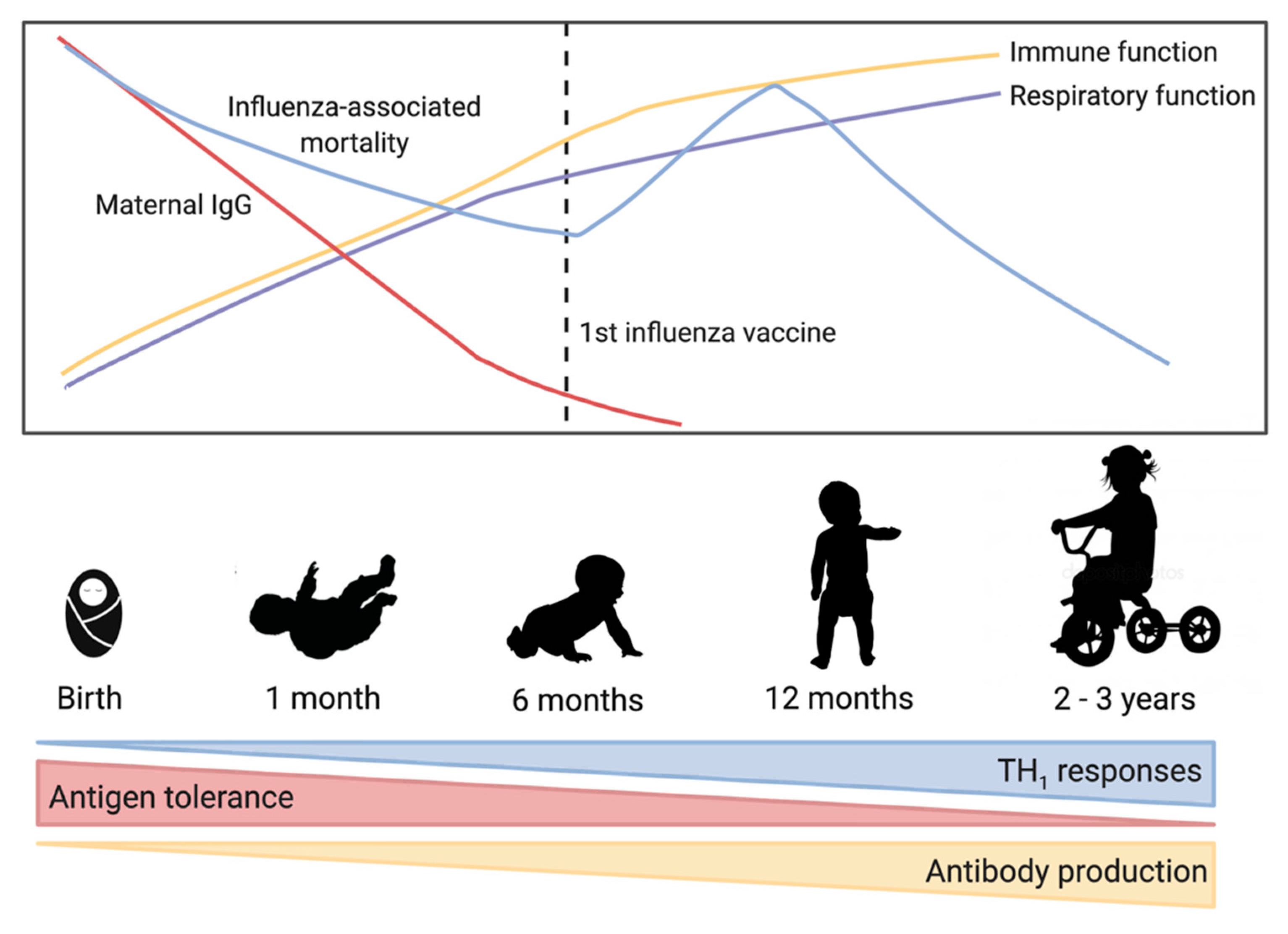
4. Infant Immune Development
4.1. Passive Immunity through Maternal Antibodies
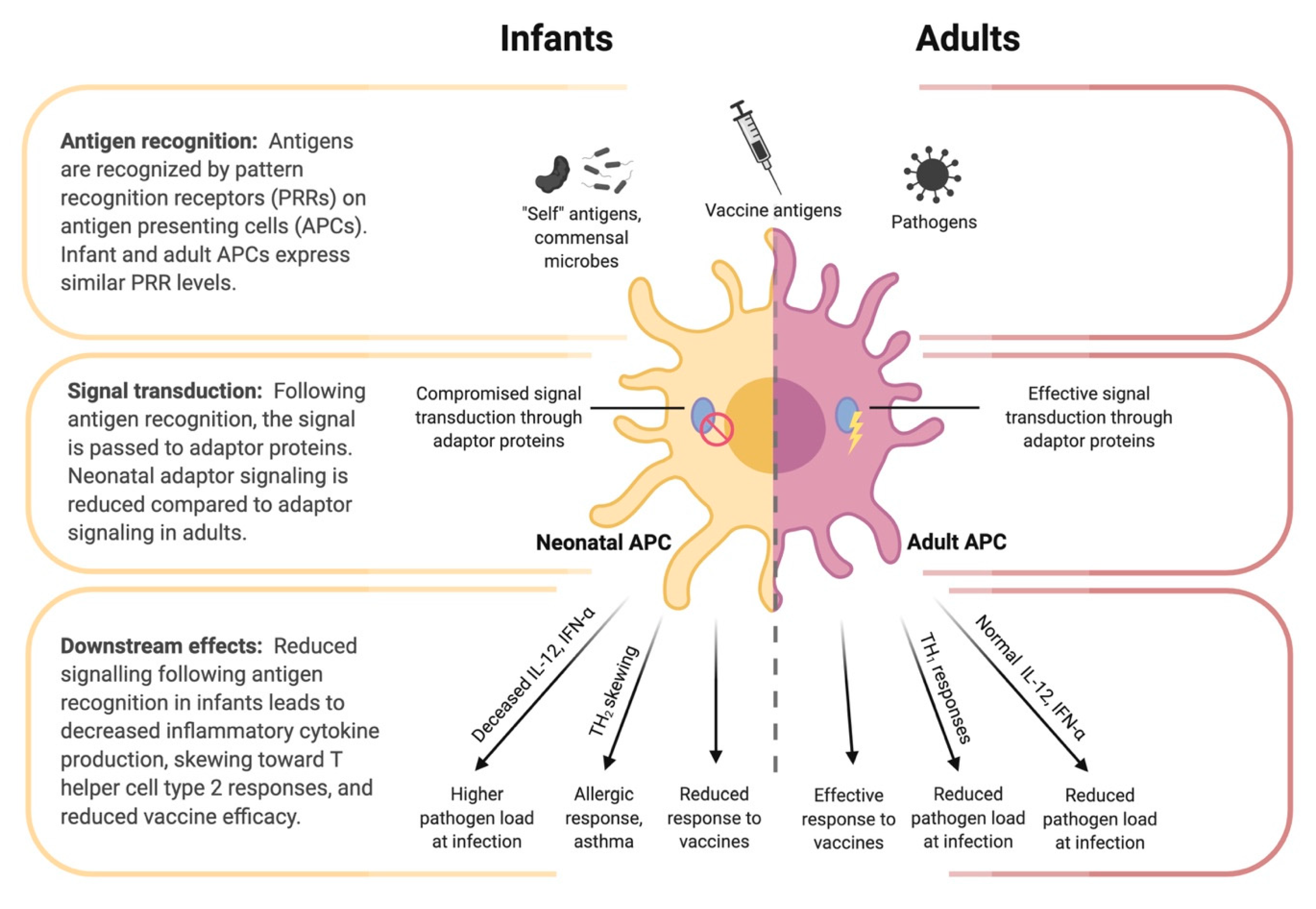
4.2. Innate Immunity of the Infant
4.3. Adaptive Immune Development in Early Life
5. Infant Immune Responses to Vaccination
6. Preclinical Modeling of Influenza Virus Infection and Vaccination
6.1. Mouse Models
6.2. Ferret Model
6.3. Non-Human Primates
7. Can We Use the Knowledge of the Infant Immune System and Influenza Imprinting to Design Better Vaccines?
8. Concluding Statements
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Krammer, F.; Smith, G.J.D.; Fouchier, R.A.M.; Peiris, M.; Kedzierska, K.; Doherty, P.C.; Palese, P.; Shaw, M.L.; Treanor, J.; Webster, R.G.; et al. Influenza. Nat. Rev. Dis. Primers 2018, 4, 1–21. [Google Scholar] [CrossRef]
- Kim, H.; Webster, R.G.; Webby, R.J. Influenza virus: Dealing with a drifting and shifting pathogen. Viral. Immunol. 2018, 31, 174–183. [Google Scholar] [CrossRef]
- WHO|Influenza. Available online: http://www.who.int/biologicals/vaccines/influenza/en/ (accessed on 25 June 2020).
- Jane, M.; Vidal, M.J.; Soldevila, N.; Romero, A.; Martinez, A.; Torner, N.; Godoy, P.; Launes, C.; Rius, C.; Marcos, M.A.; et al. Epidemiological and clinical characteristics of children hospitalized due to influenza A and B in the south of Europe, 2010–2016. Sci. Rep. 2019, 9, 12853. [Google Scholar] [CrossRef]
- Nair, H.; Brooks, W.A.; Katz, M.; Roca, A.; Berkley, J.A.; Madhi, S.A.; Simmerman, J.M.; Gordon, A.; Sato, M.; Howie, S.; et al. Global burden of respiratory infections due to seasonal influenza in young children: A systematic review and meta-analysis. Lancet 2011, 378, 1917–1930. [Google Scholar] [CrossRef]
- Neuzil, K.M.; Mellen, B.G.; Wright, P.F.; Mitchel, E.F.; Griffin, M.R. The effect of influenza on hospitalizations, outpatient visits, and courses of antibiotics in children. N. Engl. J. Med. 2000, 342, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Poehling, K.A.; Edwards, K.M.; Weinberg, G.A.; Szilagyi, P.; Staat, M.A.; Iwane, M.K.; Bridges, C.B.; Grijalva, C.G.; Zhu, Y.; Bernstein, D.I.; et al. The underrecognized burden of influenza in young children. N. Engl. J. Med. 2006, 355, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Adkins, B. Neonatal immunology: Responses to pathogenic microorganisms and epigenetics reveal an “immunodiverse” developmental state. Immunol. Res. 2013, 57, 246–257. [Google Scholar] [CrossRef]
- Krediet, T.G.; Beurskens, F.J.; van Dijk, H.; Gerards, L.J.; Fleer, A. Antibody responses and opsonic activity in sera of preterm neonates with coagulase-negative staphylococcal septicemia and the effect of the administration of fresh frozen plasma. Pediatr. Res. 1998, 43, 645–651. [Google Scholar] [CrossRef][Green Version]
- Yu, J.C.; Khodadadi, H.; Malik, A.; Davidson, B.; Salles, E.; Bhatia, J.; Hale, V.L.; Baban, B. Innate immunity of neonates and infants. Front. Immunol. 2018, 9, 1759. [Google Scholar] [CrossRef]
- Adkins, B.; Leclerc, C.; Marshall-Clarke, S. Neonatal adaptive immunity comes of age. Nat. Rev. Immunol. 2004, 4, 553–564. [Google Scholar] [CrossRef]
- Rijkers, G.T.; Dollekamp, E.G.; Zegers, B.J. The in vitro B-cell response to pneumococcal polysaccharides in adults and neonates. Scand. J. Immunol. 1987, 25, 447–452. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, S.C.A.; Roskin, K.M.; Jackson, K.J.L.; Joshi, S.A.; Nejad, P.; Lee, J.Y.; Wagar, L.E.; Pham, T.D.; Hoh, R.A.; Nguyen, K.D.; et al. Shaping of infant B cell receptor repertoires by environmental factors and infectious disease. Sci. Transl. Med. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Mazagatos, C.; Godoy, P.; Muñoz Almagro, C.; Pozo, F.; Larrauri, A. Effectiveness of influenza vaccination during pregnancy to prevent severe infection in children under 6 months of age, Spain, 2017–2019. Vaccine 2020. [Google Scholar] [CrossRef] [PubMed]
- Grohskopf, L.A.; Alyanak, E.; Broder, K.R.; Walter, E.B.; Fry, A.M.; Jernigan, D.B. Prevention and control of seasonal influenza with vaccines: Recommendations of the advisory committee on immunization practices—United States, 2019–2020 Influenza Season. MMWR Recomm. Rep. 2019, 68, 1–21. [Google Scholar] [CrossRef]
- Kelvin, A.A.; Zambon, M. Influenza imprinting in childhood and the influence on vaccine response later in life. Euro. Surveill. 2019, 24. [Google Scholar] [CrossRef]
- Rangel-Moreno, J.; Carragher, D.M.; de la Luz Garcia-Hernandez, M.; Hwang, J.Y.; Kusser, K.; Hartson, L.; Kolls, J.K.; Khader, S.A.; Randall, T.D. Induction of BALT in the absence of IL-17. Nat. Immunol. 2012, 13, 6392–6462. [Google Scholar] [CrossRef]
- Verhoeven, D.; Perry, S.; Pryharski, K. Control of influenza infection is impaired by diminished interferon-γ secretion by CD4 T cells in the lungs of toddler mice. J. Leukoc. Biol. 2016, 100, 203–212. [Google Scholar] [CrossRef]
- Paquette, S.G.; Banner, D.; Huang, S.S.H.; Almansa, R.; Leon, A.; Xu, L.; Bartoszko, J.; Kelvin, D.J.; Kelvin, A.A. Influenza transmission in the mother-infant dyad leads to severe disease, mammary gland infection, and pathogenesis by regulating host responses. PLoS Pathog. 2015, 11, e1005173. [Google Scholar] [CrossRef]
- Huang, S.S.H.; Banner, D.; Degousee, N.; Leon, A.J.; Xu, L.; Paquette, S.G.; Kanagasabai, T.; Fang, Y.; Rubino, S.; Rubin, B.; et al. Differential pathological and immune responses in newly weaned ferrets are associated with a mild clinical outcome of pandemic 2009 H1N1 infection. J. Virol. 2012, 86, 13187–13201. [Google Scholar] [CrossRef]
- Coates, D.M.; Husseini, R.H.; Rushton, D.I.; Sweet, C.; Smith, H. The role of lung development in the age-related susceptibility of ferrets to influenza virus. Br. J. Exp. Pathol. 1984, 65, 543–547. [Google Scholar] [CrossRef][Green Version]
- Rangel-Moreno, J.; Hartson, L.; Navarro, C.; Gaxiola, M.; Selman, M.; Randall, T.D. Inducible bronchus-associated lymphoid tissue (iBALT) in patients with pulmonary complications of rheumatoid arthritis. J. Clin. Investig. 2006, 116, 3183–3194. [Google Scholar] [CrossRef] [PubMed]
- Bouvier, N.M.; Palese, P. The biology of influenza viruses. Vaccine 2008, 26 (Suppl. 4), D49–53. [Google Scholar] [CrossRef] [PubMed]
- Influenza (Seasonal). Available online: https://www.who.int/news-room/fact-sheets/detail/influenza-(seasonal) (accessed on 22 June 2020).
- Jang, J.; Bae, S.E. Comparative co-evolution analysis between the HA and NA genes of influenza A virus. Virology (Auckl) 2018, 9, 1178122X18788328. [Google Scholar] [CrossRef] [PubMed]
- Gamblin, S.J.; Skehel, J.J. Influenza hemagglutinin and neuraminidase membrane glycoproteins. J. Biol. Chem. 2010, 285, 28403–28409. [Google Scholar] [CrossRef]
- Chen, C.-J.; Ermler, M.E.; Tan, G.S.; Krammer, F.; Palese, P.; Hai, R. Influenza A viruses expressing intra- or intergroup chimeric hemagglutinins. J. Virol. 2016, 90, 3789–3793. [Google Scholar] [CrossRef]
- Francis, M.E.; King, M.L.; Kelvin, A.A. Back to the future for influenza preimmunity-looking back at influenza virus history to infer the outcome of future infections. Viruses 2019, 11, 122. [Google Scholar] [CrossRef]
- Kalil, A.C.; Thomas, P.G. Influenza virus-related critical illness: Pathophysiology and epidemiology. Crit. Care 2019, 23, 258. [Google Scholar] [CrossRef]
- Taubenberger, J.K.; Morens, D.M. The pathology of influenza virus infections. Annu. Rev. Pathol. 2008, 3, 499–522. [Google Scholar] [CrossRef]
- Garg, S.; Jain, S.; Dawood, F.S.; Jhung, M.; Pérez, A.; D’Mello, T.; Reingold, A.; Gershman, K.; Meek, J.; Arnold, K.E.; et al. Pneumonia among adults hospitalized with laboratory-confirmed seasonal influenza virus infection—United States, 2005–2008. BMC Infect. Dis. 2015, 15. [Google Scholar] [CrossRef]
- Mauad, T.; Hajjar, L.A.; Callegari, G.D.; da Silva, L.F.F.; Schout, D.; Galas, F.R.B.G.; Alves, V.A.F.; Malheiros, D.M.A.C.; Auler, J.O.C.; Ferreira, A.F.; et al. Lung pathology in fatal novel human influenza A (H1N1) infection. Am. J. Respir. Crit. Care Med. 2010, 181, 72–79. [Google Scholar] [CrossRef]
- Vemula, S.V.; Sayedahmed, E.E.; Sambhara, S.; Mittal, S.K. Vaccine approaches conferring cross-protection against influenza viruses. Expert Rev. Vaccines 2017, 16, 1141–1154. [Google Scholar] [CrossRef] [PubMed]
- Fiore, A.E.; Shay, D.K.; Broder, K.; Iskander, J.K.; Uyeki, T.M.; Mootrey, G.; Bresee, J.S.; Cox, N.J.; Centers for Disease Control and Prevention. Prevention and control of seasonal influenza with vaccines: Recommendations of the Advisory Committee on Immunization Practices (ACIP), 2009. MMWR Recomm. Rep. 2009, 58, 1–52. [Google Scholar] [PubMed]
- Wong, S.-S.; Webby, R.J. Traditional and new influenza vaccines. Clin. Microbiol. Rev. 2013, 26, 476–492. [Google Scholar] [CrossRef] [PubMed]
- Rajao, D.S.; Perez, D.R. Universal vaccines and vaccine platforms to protect against influenza viruses in humans and agriculture. Front. Microbiol. 2018, 9, 123. [Google Scholar] [CrossRef]
- Adjuvanted Influenza Vaccines. Available online: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5861793/ (accessed on 28 July 2020).
- Del Giudice, G.; Rappuoli, R. Inactivated and adjuvanted influenza vaccines. In Influenza Pathogenesis and Control—Volume II; Oldstone, M.B.A., Compans, R.W., Eds.; Current Topics in Microbiology and Immunology; Springer International Publishing: Cham, Switzerland, 2015; pp. 151–180. ISBN 978-3-319-11158-2. [Google Scholar]
- Public Health Agency of Canada. Canadian Immunization Guide Chapter on Influenza and Statement on Seasonal Influenza Vaccine for 2019–2020. Available online: https://www.canada.ca/en/public-health/services/publications/vaccines-immunization/canadian-immunization-guide-statement-seasonal-influenza-vaccine-2019–2020.html#IV (accessed on 28 July 2020).
- Influenza Vaccination: A Summary for Clinicians|CDC. Available online: https://www.cdc.gov/flu/professionals/vaccination/vax-summary.htm (accessed on 7 July 2020).
- Hancock, K.; Veguilla, V.; Lu, X.; Zhong, W.; Butler, E.N.; Sun, H.; Liu, F.; Dong, L.; DeVos, J.R.; Gargiullo, P.M.; et al. Cross-reactive antibody responses to the 2009 pandemic H1N1 influenza virus. N. Engl. J. Med. 2009, 361, 1945–1952. [Google Scholar] [CrossRef]
- Francis, T. On the doctrine of original antigenic sin. Proc. Am. Philos. Soc. 1960, 104, 572–578. [Google Scholar]
- Fonville, J.M.; Wilks, S.H.; James, S.L.; Fox, A.; Ventresca, M.; Aban, M.; Xue, L.; Jones, T.C.; Le, N.M.H.; Pham, Q.T.; et al. Antibody landscapes after influenza virus infection or vaccination. Science 2014, 346, 996–1000. [Google Scholar] [CrossRef]
- Monto, A.S.; Malosh, R.E.; Petrie, J.G.; Martin, E.T. The doctrine of original antigenic sin: Separating good from evil. J. Infect. Dis. 2017, 215, 1782–1788. [Google Scholar] [CrossRef]
- Lessler, J.; Riley, S.; Read, J.M.; Wang, S.; Zhu, H.; Smith, G.J.D.; Guan, Y.; Jiang, C.Q.; Cummings, D.A.T. Evidence for antigenic seniority in influenza A (H3N2) antibody responses in southern China. PLoS Pathog. 2012, 8, e1002802. [Google Scholar] [CrossRef]
- Skowronski, D.M.; Chambers, C.; De Serres, G.; Sabaiduc, S.; Winter, A.L.; Dickinson, J.A.; Gubbay, J.B.; Fonseca, K.; Drews, S.J.; Charest, H.; et al. Serial vaccination and the antigenic distance hypothesis: Effects on influenza vaccine effectiveness during A(H3N2) epidemics in Canada, 2010–2011 to 2014–2015. J. Infect. Dis. 2017, 215, 1059–1099. [Google Scholar] [CrossRef]
- Gostic, K.M.; Ambrose, M.; Worobey, M.; Lloyd-Smith, J.O. Potent protection against H5N1 and H7N9 influenza via childhood hemagglutinin imprinting. Science 2016, 354, 722–726. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S.F.; Huang, Y.; Kaur, K.; Popova, L.I.; Ho, I.Y.; Pauli, N.T.; Henry Dunand, C.J.; Taylor, W.M.; Lim, S.; Huang, M.; et al. Immune history profoundly affects broadly protective B cell responses to influenza. Sci. Transl. Med. 2015, 7, 316ra192. [Google Scholar] [CrossRef] [PubMed]
- Ellebedy, A.H.; Ahmed, R. Re-engaging cross-reactive memory B cells: The influenza puzzle. Front. Immunol. 2012, 3, 53. [Google Scholar] [CrossRef] [PubMed]
- Worobey, M.; Han, G.-Z.; Rambaut, A. Genesis and pathogenesis of the 1918 pandemic H1N1 influenza A virus. Proc. Natl. Acad. Sci. USA 2014, 111, 8107–8112. [Google Scholar] [CrossRef]
- Arevalo, C.P.; Le Sage, V.; Bolton, M.J.; Eilola, T.; Jones, J.E.; Kormuth, K.A.; Nturibi, E.; Balmaseda, A.; Gordon, A.; Lakdawala, S.S.; et al. Original antigenic sin priming of influenza virus hemagglutinin stalk antibodies. Proc. Natl. Acad. Sci. USA 2020, 117, 17221–17227. [Google Scholar] [CrossRef]
- Tesini, B.L.; Kanagaiah, P.; Wang, J.; Hahn, M.; Halliley, J.L.; Chaves, F.A.; Nguyen, P.Q.T.; Nogales, A.; DeDiego, M.L.; Anderson, C.S.; et al. Broad Hemagglutinin-specific memory b cell expansion by seasonal influenza virus infection reflects early-life imprinting and adaptation to the infecting virus. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Skowronski, D.M.; Sabaiduc, S.; Leir, S.; Rose, C.; Zou, M.; Murti, M.; Dickinson, J.A.; Olsha, R.; Gubbay, J.B.; Croxen, M.A.; et al. Paradoxical clade- and age-specific vaccine effectiveness during the 2018/19 influenza A(H3N2) epidemic in Canada: Potential imprint-regulated effect of vaccine (I-REV). Euro. Surveill. 2019, 24. [Google Scholar] [CrossRef]
- Francis, M.E.; McNeil, M.; Dawe, N.J.; Foley, M.K.; King, M.L.; Ross, T.M.; Kelvin, A.A. Historical H1N1 influenza virus imprinting increases vaccine protection by influencing the activity and sustained production of antibodies elicited at vaccination in ferrets. Vaccines 2019, 7, 133. [Google Scholar] [CrossRef]
- Strunk, T.; Currie, A.; Richmond, P.; Simmer, K.; Burgner, D. Innate immunity in human newborn infants: Prematurity means more than immaturity. J. Matern. Fetal. Neonatal. Med. 2011, 24, 25–31. [Google Scholar] [CrossRef]
- Fadel, S.; Sarzotti, M. Cellular immune responses in neonates. Int. Rev. Immunol. 2000, 19, 173–193. [Google Scholar] [CrossRef]
- Sarzotti, M.; Robbins, D.S.; Hoffman, P.M. Induction of protective CTL responses in newborn mice by a murine retrovirus. Science 1996, 271, 1726–1728. [Google Scholar] [CrossRef] [PubMed]
- Ridge, J.P.; Fuchs, E.J.; Matzinger, P. Neonatal tolerance revisited: Turning on newborn T cells with dendritic cells. Science 1996, 271, 1723–1726. [Google Scholar] [CrossRef] [PubMed]
- Forsthuber, T.; Yip, H.C.; Lehmann, P.V. Induction of TH1 and TH2 immunity in neonatal mice. Science 1996, 271, 1728–1730. [Google Scholar] [CrossRef]
- Offit, P.A.; Quarles, J.; Gerber, M.A.; Hackett, C.J.; Marcuse, E.K.; Kollman, T.R.; Gellin, B.G.; Landry, S. Addressing parents’ concerns: Do multiple vaccines overwhelm or weaken the infant’s immune system? Pediatrics 2002, 109, 124–129. [Google Scholar] [CrossRef] [PubMed]
- Siegrist, C.A.; Cordova, M.; Brandt, C.; Barrios, C.; Berney, M.; Tougne, C.; Kovarik, J.; Lambert, P.H. Determinants of infant responses to vaccines in presence of maternal antibodies. Vaccine 1998, 16, 1409–1414. [Google Scholar] [CrossRef]
- Glezen, W.P.; Taber, L.H.; Frank, A.L.; Gruber, W.C.; Piedra, P.A. Influenza virus infections in infants. Pediatr. Infect. Dis. J. 1997, 16, 1065–1068. [Google Scholar] [CrossRef]
- Eick, A.A.; Uyeki, T.M.; Klimov, A.; Hall, H.; Reid, R.; Santosham, M.; O’Brien, K.L. Maternal influenza vaccination and effect on influenza virus infection in young infants. Arch. Pediatr. Adolesc. Med. 2011, 165, 104–111. [Google Scholar] [CrossRef]
- Yachie, A.; Takano, N.; Ohta, K.; Uehara, T.; Fujita, S.; Miyawaki, T.; Taniguchi, N. Defective production of interleukin-6 in very small premature infants in response to bacterial pathogens. Infect. Immun. 1992, 60, 749–753. [Google Scholar] [CrossRef]
- Angelone, D.F.; Wessels, M.R.; Coughlin, M.; Suter, E.E.; Valentini, P.; Kalish, L.A.; Levy, O. Innate immunity of the human newborn is polarized toward a high ratio of IL-6/TNF-alpha production in vitro and in vivo. Pediatr. Res. 2006, 60, 205–209. [Google Scholar] [CrossRef]
- Schaller-Bals, S.; Schulze, A.; Bals, R. Increased levels of antimicrobial peptides in tracheal aspirates of newborn infants during infection. Am. J. Respir. Crit. Care Med. 2002, 165, 992–995. [Google Scholar] [CrossRef]
- Drew, J.H.; Arroyave, C.M. The complement system of the newborn infant. Biol. Neonate. 1980, 37, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Diefenbach, A.; Colonna, M.; Romagnani, C. The ILC world revisited. Immunity 2017, 46, 327–332. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Eberl, G.; Colonna, M.; Di Santo, J.P.; McKenzie, A.N. Innate lymphoid cells. Innate lymphoid cells: A new paradigm in immunology. Science 2015, 348, aaa6566. [Google Scholar] [CrossRef] [PubMed]
- Artis, D.; Spits, H. The biology of innate lymphoid cells. Nature 2015, 517, 293–301. [Google Scholar] [CrossRef]
- Goldblatt, D. Immunisation and the maturation of infant immune responses. Dev. Biol. Stand. 1998, 95, 125–132. [Google Scholar]
- Fadel, S.A.; Cowell, L.G.; Cao, S.; Ozaki, D.A.; Kepler, T.B.; Steeber, D.A.; Sarzotti, M. Neonate-primed CD8+ memory cells rival adult-primed memory cells in antigen-driven expansion and anti-viral protection. Int. Immunol. 2006, 18, 249–257. [Google Scholar] [CrossRef][Green Version]
- Siegrist, C.A. Neonatal and early life vaccinology. Vaccine 2001, 19, 3331–3346. [Google Scholar] [CrossRef]
- Mackie, R.I.; Sghir, A.; Gaskins, H.R. Developmental microbial ecology of the neonatal gastrointestinal tract. Am. J. Clin. Nutr. 1999, 69, 1035S–1045S. [Google Scholar] [CrossRef]
- Mellander, L.; Carlsson, B.; Jalil, F.; Soderstrom, T.; Hanson, L.A. Secretory IgA antibody response against Escherichia coli antigens in infants in relation to exposure. J. Pediatr. 1985, 107, 430–433. [Google Scholar] [CrossRef]
- Zhang, X.; Deriaud, E.; Jiao, X.; Braun, D.; Leclerc, C.; Lo-Man, R. Type I interferons protect neonates from acute inflammation through interleukin 10-producing B cells. J. Exp. Med. 2007, 204, 1107–1118. [Google Scholar] [CrossRef]
- Belderbos, M.E.; Levy, O.; Stalpers, F.; Kimpen, J.L.; Meyaard, L.; Bont, L. Neonatal plasma polarizes TLR4-mediated cytokine responses towards low IL-12p70 and high IL-10 production via distinct factors. PLoS ONE 2012, 7, e33419. [Google Scholar] [CrossRef] [PubMed]
- Siegrist, C.-A.; Aspinall, R. B-cell responses to vaccination at the extremes of age. Nat. Rev. Immunol. 2009, 9, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Katz, J.; Englund, J.A.; Steinhoff, M.C.; Khatry, S.K.; Shrestha, L.; Kuypers, J.; Mullany, L.C.; Chu, H.Y.; LeClerq, S.C.; Kozuki, N.; et al. Impact of timing of influenza vaccination in pregnancy on transplacental antibody transfer, influenza incidence, and birth outcomes: A randomized trial in rural nepal. Clin. Infect. Dis. 2018, 67, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Vesikari, T.; Knuf, M.; Wutzler, P.; Karvonen, A.; Kieninger-Baum, D.; Schmitt, H.-J.; Baehner, F.; Borkowski, A.; Tsai, T.F.; Clemens, R. Oil-in-Water Emulsion Adjuvant with Influenza Vaccine in Young Children. Available online: https://www.nejm.org/doi/10.1056/NEJMoa1010331 (accessed on 29 August 2020).
- Coelingh, K.; Olajide, I.R.; MacDonald, P.; Yogev, R. Efficacy and effectiveness of live attenuated influenza vaccine in school-age children. Expert Rev. Vaccines 2015, 14, 1331–1346. [Google Scholar] [CrossRef]
- Blanco-Lobo, P.; Nogales, A.; Rodríguez, L.; Martínez-Sobrido, L. Novel approaches for the development of live attenuated influenza vaccines. Viruses 2019, 11, 190. [Google Scholar] [CrossRef]
- Restori, K.H.; Srinivasa, B.T.; Ward, B.J.; Fixman, E.D. Neonatal immunity, respiratory virus infections, and the development of asthma. Front. Immunol. 2018, 9, 1249. [Google Scholar] [CrossRef]
- Bryda, E.C. The mighty mouse: The impact of rodents on advances in biomedical research. Mo. Med. 2013, 110, 207–211. [Google Scholar]
- Richter, S.H.; Kästner, N.; Loddenkemper, D.-H.; Kaiser, S.; Sachser, N. A time to wean? Impact of weaning age on anxiety-like behaviour and stability of behavioural traits in full adulthood. PLoS ONE 2016, 11, e0167652. [Google Scholar] [CrossRef]
- Holladay, S.D.; Smialowicz, R.J. Development of the murine and human immune system: Differential effects of immunotoxicants depend on time of exposure. Environ. Health Perspect. 2000, 108 (Suppl. 3), 463–473. [Google Scholar] [CrossRef]
- Garcia, A.M.; Fadel, S.A.; Cao, S.; Sarzotti, M. T cell immunity in neonates. Immunol. Res. 2000, 22, 177–190. [Google Scholar] [CrossRef]
- Campion, A.L.; Bourgeois, C.; Lambolez, F.; Martin, B.; Léaument, S.; Dautigny, N.; Tanchot, C.; Pénit, C.; Lucas, B. Naive T cells proliferate strongly in neonatal mice in response to self-peptide/self-MHC complexes. Proc. Natl. Acad. Sci. USA 2002, 99, 4538–4543. [Google Scholar] [CrossRef]
- Fike, A.J.; Nguyen, L.T.; Kumova, O.K.; Carey, A.J. Characterization of CD31 expression on murine and human neonatal T lymphocytes during development and activation. Pediatr. Res. 2017, 82, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Carey, A.J.; Gracias, D.T.; Thayer, J.L.; Boesteanu, A.C.; Kumova, O.K.; Mueller, Y.M.; Hope, J.L.; Fraietta, J.A.; van Zessen, D.B.H.; Katsikis, P.D. Rapid evolution of the CD8+ TCR repertoire in neonatal mice. J. Immunol. 2016, 196, 2602–2613. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.-Z.J.; Dash, P.; Crawford, J.C.; Allen, E.K.; Zamora, A.E.; Boyd, D.F.; Duan, S.; Bajracharya, R.; Awad, W.A.; Apiwattanakul, N.; et al. Lung γδ T cells mediate protective responses during neonatal influenza infection that are associated with type 2 immunity. Immunity 2018, 49, 531–544.e6. [Google Scholar] [CrossRef] [PubMed]
- Lines, J.L.; Hoskins, S.; Hollifield, M.; Cauley, L.S.; Garvy, B.A. The migration of T Cells in response to influenza virus is altered in neonatal mice. J. Immunol. 2010, 185, 2980–2988. [Google Scholar] [CrossRef]
- Fike, A.J.; Kumova, O.K.; Tardif, V.J.; Carey, A.J. Neonatal influenza-specific effector CTLs retain elevated CD31 levels at the site of infection and have decreased IFN-γ production. J. Leukoc. Biol. 2019, 105, 539–549. [Google Scholar] [CrossRef]
- Oliphant, S.; Lines, J.L.; Hollifield, M.L.; Garvy, B.A. Regulatory T cells are critical for clearing influenza A Virus in neonatal mice. Viral Immunol. 2015, 28, 580–589. [Google Scholar] [CrossRef]
- Zens, K.D.; Chen, J.K.; Guyer, R.S.; Wu, F.L.; Cvetkovski, F.; Miron, M.; Farber, D.L. Reduced generation of lung tissue–resident memory T cells during infancy. J. Exp. Med. 2017, 214, 2915–2932. [Google Scholar] [CrossRef]
- Zens, K.D.; Chen, J.K.; Farber, D.L. Vaccine-generated lung tissue–resident memory T cells provide heterosubtypic protection to influenza infection. JCI Insight 2016, 1. [Google Scholar] [CrossRef]
- Teijaro, J.R.; Turner, D.; Pham, Q.; Wherry, E.J.; Lefrançois, L.; Farber, D.L. Tissue-retentive lung memory CD4 T cells mediate optimal protection to respiratory virus infection. J. Immunol. 2011, 187, 5510–5514. [Google Scholar] [CrossRef]
- Paquette, S.G.; Huang, S.S.H.; Banner, D.; Xu, L.; Leόn, A.; Kelvin, A.A.; Kelvin, D.J. Impaired heterologous immunity in aged ferrets during sequential influenza A H1N1 infection. Virology 2014, 464–465, 177–183. [Google Scholar] [CrossRef]
- Smith, H.; Sweet, C. Lessons for human influenza from pathogenicity studies with ferrets. Rev. Infect. Dis. 1988, 10, 56–75. [Google Scholar] [CrossRef] [PubMed]
- Bissel, S.J.; Carter, C.E.; Wang, G.; Johnson, S.K.; Lashua, L.P.; Kelvin, A.A.; Wiley, C.A.; Ghedin, E.; Ross, T.M. Age-related pathology associated with H1N1 A/California/07/2009 influenza virus infection. Am. J. Pathol. 2019, 189, 2389–2399. [Google Scholar] [CrossRef] [PubMed]
- Jakeman, K.J.; Smith, H.; Sweet, C. Mechanism of immunity to influenza: Maternal and passive neonatal protection following immunization of adult ferrets with a live vaccinia-influenza virus haemagglutinin recombinant but not with recombinants containing other influenza virus proteins. J. Gen. Virol. 1989, 70, 1523–1531. [Google Scholar] [CrossRef] [PubMed]
- Husseini, R.H.; Sweet, C.; Overton, H.; Smith, H. Role of maternal immunity in the protection of newborn ferrets against infection with a virulent influenza virus. Immunology 1984, 52, 389–394. [Google Scholar]
- Husseini, R.H.; Collie, M.H.; Rushton, D.I.; Sweet, C.; Smith, H. The role of naturally-acquired bacterial infection in influenza-related death in neonatal ferrets. Br. J. Exp. Pathol. 1983, 64, 559–569. [Google Scholar]
- Rushton, D.I.; Collie, M.H.; Sweet, C.; Husseini, R.H.; Smith, H. The effects of maternal influenzal viraemia in late gestation on the conceptus of the pregnant ferret. J. Pathol. 1983, 140, 181–191. [Google Scholar] [CrossRef]
- Collie, M.H.; Rushton, D.I.; Sweet, C.; Smith, H. Studies of influenza virus infection in newborn ferrets. J. Med. Microbiol. 1980, 13, 561–571. [Google Scholar] [CrossRef]
- Huang, S.S.H.; Banner, D.; Paquette, S.G.; Leon, A.J.; Kelvin, A.A.; Kelvin, D.J. Pathogenic influenza B virus in the ferret model establishes lower respiratory tract infection. J. Gen. Virol. 2014, 95, 2127–2139. [Google Scholar] [CrossRef]
- Huang, S.S.H.; Banner, D.; Fang, Y.; Ng, D.C.K.; Kanagasabai, T.; Kelvin, D.J.; Kelvin, A.A. Comparative analyses of pandemic H1N1 and seasonal H1N1, H3N2, and influenza B infections depict distinct clinical pictures in ferrets. PLoS ONE 2011, 6. [Google Scholar] [CrossRef]
- Coates, D.M.; Husseini, R.H.; Collie, M.H.; Sweet, C.; Smith, H. The role of cellular susceptibility in the declining severity of respiratory influenza of ferrets with age. Br. J. Exp. Pathol. 1984, 65, 29–39. [Google Scholar] [PubMed]
- Skarlupka, A.L.; Ross, T.M. Immune imprinting in the influenza ferret model. Vaccines 2020, 8, 173. [Google Scholar] [CrossRef]
- Davis, A.S.; Taubenberger, J.K.; Bray, M. The use of nonhuman primates in research on seasonal, pandemic and avian influenza, 1893–2014. Antivir. Res. 2015, 117, 75–98. [Google Scholar] [CrossRef]
- Moncla, L.H.; Ross, T.M.; Dinis, J.M.; Weinfurter, J.T.; Mortimer, T.D.; Schultz-Darken, N.; Brunner, K.; Iii, S.V.C.; Boettcher, C.; Post, J.; et al. A novel nonhuman primate model for influenza transmission. PLoS ONE 2013, 8, e78750. [Google Scholar] [CrossRef] [PubMed]
- Rimmelzwaan, G.F.; Kuiken, T.; van Amerongen, G.; Bestebroer, T.M.; Fouchier, R.A.M.; Osterhaus, A.D.M.E. Pathogenesis of influenza A (H5N1) virus infection in a primate model. J. Virol. 2001, 75, 6687–6691. [Google Scholar] [CrossRef] [PubMed]
- Baskin, C.R.; Bielefeldt-Ohmann, H.; Tumpey, T.M.; Sabourin, P.J.; Long, J.P.; García-Sastre, A.; Tolnay, A.-E.; Albrecht, R.; Pyles, J.A.; Olson, P.H.; et al. Early and sustained innate immune response defines pathology and death in nonhuman primates infected by highly pathogenic influenza virus. Proc. Natl. Acad. Sci. USA 2009, 106, 3455–3460. [Google Scholar] [CrossRef] [PubMed]
- Rimmelzwaan, G.F.; Baars, M.; van Beek, R.; van Amerongen, G.; Lövgren-Bengtsson, K.; Claas, E.C.; Osterhaus, A.D. Induction of protective immunity against influenza virus in a macaque model: Comparison of conventional and iscom vaccines. J. Gen. Virol. 1997, 78, 757–765. [Google Scholar] [CrossRef] [PubMed]
- Batchelder, C.A.; Duru, N.; Lee, C.I.; Baker, C.A.R.; Swainson, L.; Mccune, J.M.; Tarantal, A.F. Myeloid-lymphoid ontogeny in the rhesus monkey (Macaca mulatta). Anat. Rec. (Hoboken) 2014, 297, 1392–1406. [Google Scholar] [CrossRef]
- Merino, K.M.; Slisarenko, N.; Taylor, J.M.; Falkenstein, K.P.; Gilbert, M.H.; Bohm, R.P.; Blanchard, J.L.; Ardeshir, A.; Didier, E.S.; Kim, W.-K.; et al. Clinical and immunological metrics during pediatric rhesus macaque development. Front. Pediatr. 2020, 8. [Google Scholar] [CrossRef]
- Oxford, K.L.; Dela Pena-Ponce, M.G.A.; Jensen, K.; Eberhardt, M.K.; Spinner, A.; Van Rompay, K.K.; Rigdon, J.; Mollan, K.R.; Krishnan, V.V.; Hudgens, M.G.; et al. The interplay between immune maturation, age, chronic viral infection and environment. Immun. Ageing 2015, 12, 3. [Google Scholar] [CrossRef]
- van Riel, D.; Munster, V.J.; de Wit, E.; Rimmelzwaan, G.F.; Fouchier, R.A.M.; Osterhaus, A.D.M.E.; Kuiken, T. Human and avian influenza viruses target different cells in the lower respiratory tract of humans and other mammals. Am. J. Pathol. 2007, 171, 1215–1223. [Google Scholar] [CrossRef] [PubMed]
- Weinfurter, J.T.; Brunner, K.; Iii, S.V.C.; Li, C.; Broman, K.W.; Kawaoka, Y.; Friedrich, T.C. Cross-reactive T cells are involved in rapid clearance of 2009 pandemic H1N1 influenza virus in nonhuman primates. PLoS Pathog. 2011, 7, e1002381. [Google Scholar] [CrossRef] [PubMed]
- Cillóniz, C.; Shinya, K.; Peng, X.; Korth, M.J.; Proll, S.C.; Aicher, L.D.; Carter, V.S.; Chang, J.H.; Kobasa, D.; Feldmann, F.; et al. Lethal Influenza virus infection in macaques is associated with early dysregulation of inflammatory related genes. PLoS Pathog. 2009, 5, e1000604. [Google Scholar] [CrossRef]
- Kobasa, D.; Jones, S.M.; Shinya, K.; Kash, J.C.; Copps, J.; Ebihara, H.; Hatta, Y.; Hyun Kim, J.; Halfmann, P.; Hatta, M.; et al. Aberrant innate immune response in lethal infection of macaques with the 1918 influenza virus. Nature 2007, 445, 319–323. [Google Scholar] [CrossRef] [PubMed]
- Carroll, T.D.; Matzinger, S.R.; Genescà, M.; Fritts, L.; Colòn, R.; McChesney, M.B.; Miller, C.J. Interferon-induced expression of MxA in the Respiratory tract of rhesus macaques is suppressed by influenza virus replication. J. Immunol. 2008, 180, 2385–2395. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.R.; Holbrook, B.C.; Hayward, S.L.; Blevins, L.K.; Jorgensen, M.J.; Kock, N.D.; Paris, K.D.; D’Agostino, R.B.; Aycock, S.T.; Mizel, S.B.; et al. Inclusion of flagellin during vaccination against influenza enhances recall responses in nonhuman primate neonates. J. Virol. 2015, 89, 7291–7303. [Google Scholar] [CrossRef] [PubMed]
- Holbrook, B.C.; Hayward, S.L.; Blevins, L.K.; Kock, N.; Aycock, T.; Parks, G.D.; Alexander-Miller, M.A. Nonhuman primate infants have an impaired respiratory but not systemic IgG antibody response following influenza virus infection. Virology 2015, 476, 124–133. [Google Scholar] [CrossRef]
- Holbrook, B.C.; D’Agostino, R.B.; Parks, G.D.; Alexander-Miller, M.A. Adjuvanting an inactivated influenza vaccine with flagellin improves the function and quantity of the long-term antibody response in a nonhuman primate neonate model. Vaccine 2016, 34, 4712–4717. [Google Scholar] [CrossRef]
- Holbrook, B.C.; Aycock, S.T.; Machiele, E.; Clemens, E.; Gries, D.; Jorgensen, M.J.; Hadimani, M.B.; King, S.B.; Alexander-Miller, M.A. An R848 adjuvanted influenza vaccine promotes early activation of B cells in the draining lymph nodes of non-human primate neonates. Immunology 2018, 153, 357–367. [Google Scholar] [CrossRef]
- Krammer, F.; Palese, P. Influenza virus hemagglutinin stalk-based antibodies and vaccines. Curr. Opin. Virol. 2013, 3, 521–530. [Google Scholar] [CrossRef]
- Bui, H.-H.; Peters, B.; Assarsson, E.; Mbawuike, I.; Sette, A. Ab and T cell epitopes of influenza A virus, knowledge and opportunities. Proc. Natl. Acad. Sci. USA 2007, 104, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Nachbagauer, R.; Wohlbold, T.J.; Hirsh, A.; Hai, R.; Sjursen, H.; Palese, P.; Cox, R.J.; Krammer, F. Induction of broadly reactive anti-hemagglutinin stalk antibodies by an H5N1 vaccine in humans. J. Virol. 2014, 88, 13260–13268. [Google Scholar] [CrossRef] [PubMed]
- Jazayeri, S.D.; Poh, C.L. Development of universal influenza vaccines targeting conserved viral proteins. Vaccines 2019, 7, 169. [Google Scholar] [CrossRef]
- Kirchenbaum, G.A.; Carter, D.M.; Ross, T.M. Sequential infection in ferrets with antigenically distinct seasonal H1N1 influenza viruses boosts hemagglutinin stalk-specific antibodies. J. Virol. 2016, 90, 1116–1128. [Google Scholar] [CrossRef] [PubMed]
| Age Group | Recommended Vaccine | Live Vaccine (Yes/No) | Reason for Recommendation |
|---|---|---|---|
| Infants aged 0–6 months | None; maternal vaccination during pregnancy recommended | No | No licensed influenza vaccines for infants less than 6 months of age |
| Infants aged 6 months–2 years | TIV or QIV | No | LAIV is not authorized for use in children less than two years of age |
| Children aged 2–17 years | TIV, QIV or LAIV | Yes | All vaccines authorized for age group |
| Adults aged 18–59 years | TIV, QIV or LAIV | Yes | All vaccines authorized for age group |
| Adults 60–64 years | TIV and QIV | No | LAIV not recommended |
| Adults aged 65 and over | Adjuvanted TIV or high dose TIV | No | Unadjuvanted vaccines are poorly immunogenic in elderly populations |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rioux, M.; McNeil, M.; Francis, M.E.; Dawe, N.; Foley, M.; Langley, J.M.; Kelvin, A.A. The Power of First Impressions: Can Influenza Imprinting during Infancy Inform Vaccine Design? Vaccines 2020, 8, 546. https://doi.org/10.3390/vaccines8030546
Rioux M, McNeil M, Francis ME, Dawe N, Foley M, Langley JM, Kelvin AA. The Power of First Impressions: Can Influenza Imprinting during Infancy Inform Vaccine Design? Vaccines. 2020; 8(3):546. https://doi.org/10.3390/vaccines8030546
Chicago/Turabian StyleRioux, Melissa, Mara McNeil, Magen E. Francis, Nicholas Dawe, Mary Foley, Joanne M. Langley, and Alyson A. Kelvin. 2020. "The Power of First Impressions: Can Influenza Imprinting during Infancy Inform Vaccine Design?" Vaccines 8, no. 3: 546. https://doi.org/10.3390/vaccines8030546
APA StyleRioux, M., McNeil, M., Francis, M. E., Dawe, N., Foley, M., Langley, J. M., & Kelvin, A. A. (2020). The Power of First Impressions: Can Influenza Imprinting during Infancy Inform Vaccine Design? Vaccines, 8(3), 546. https://doi.org/10.3390/vaccines8030546





