Prospects and Challenges in the Development of Universal Influenza Vaccines
Abstract
1. Introduction
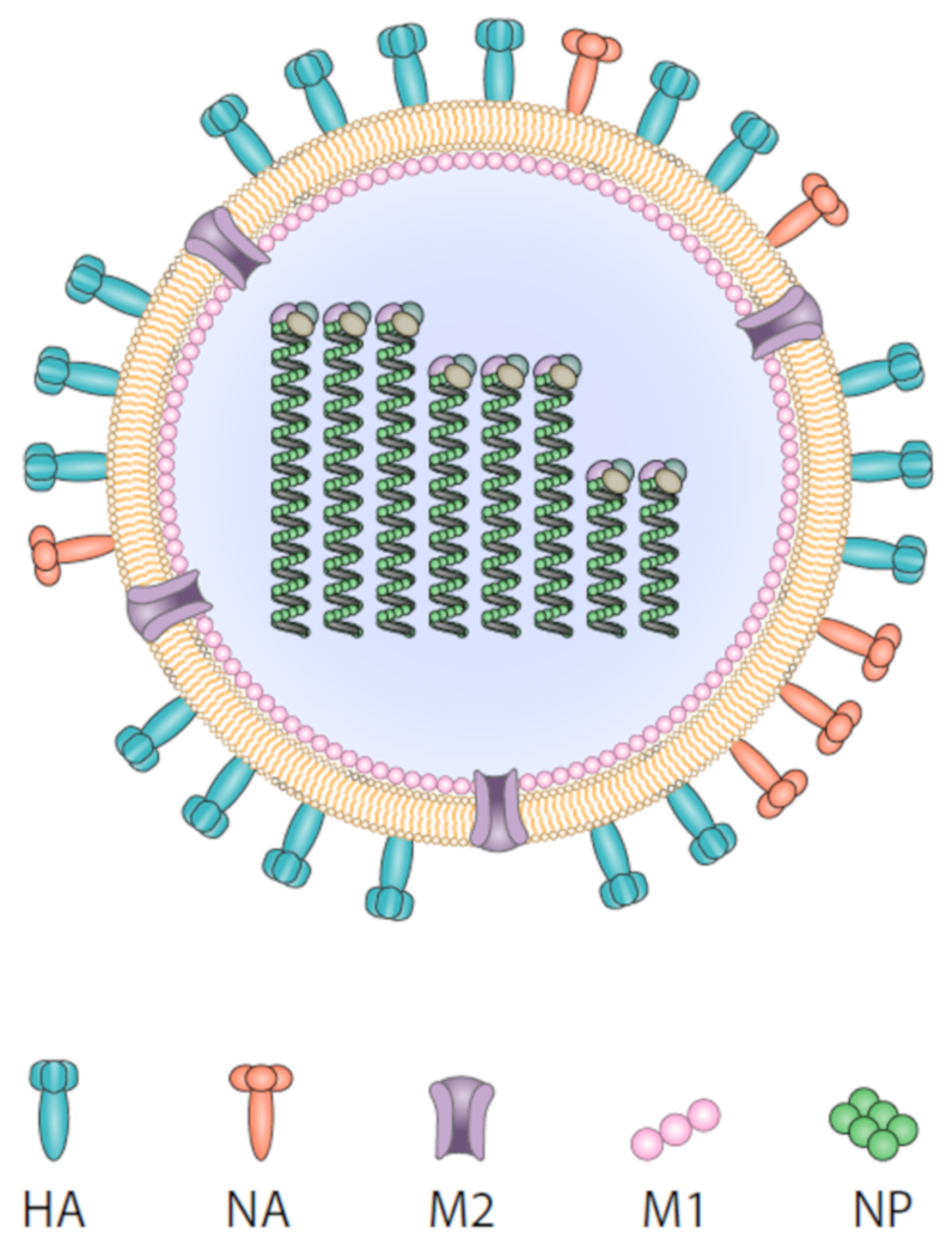
2. Influenza Virus and Target Epitopes
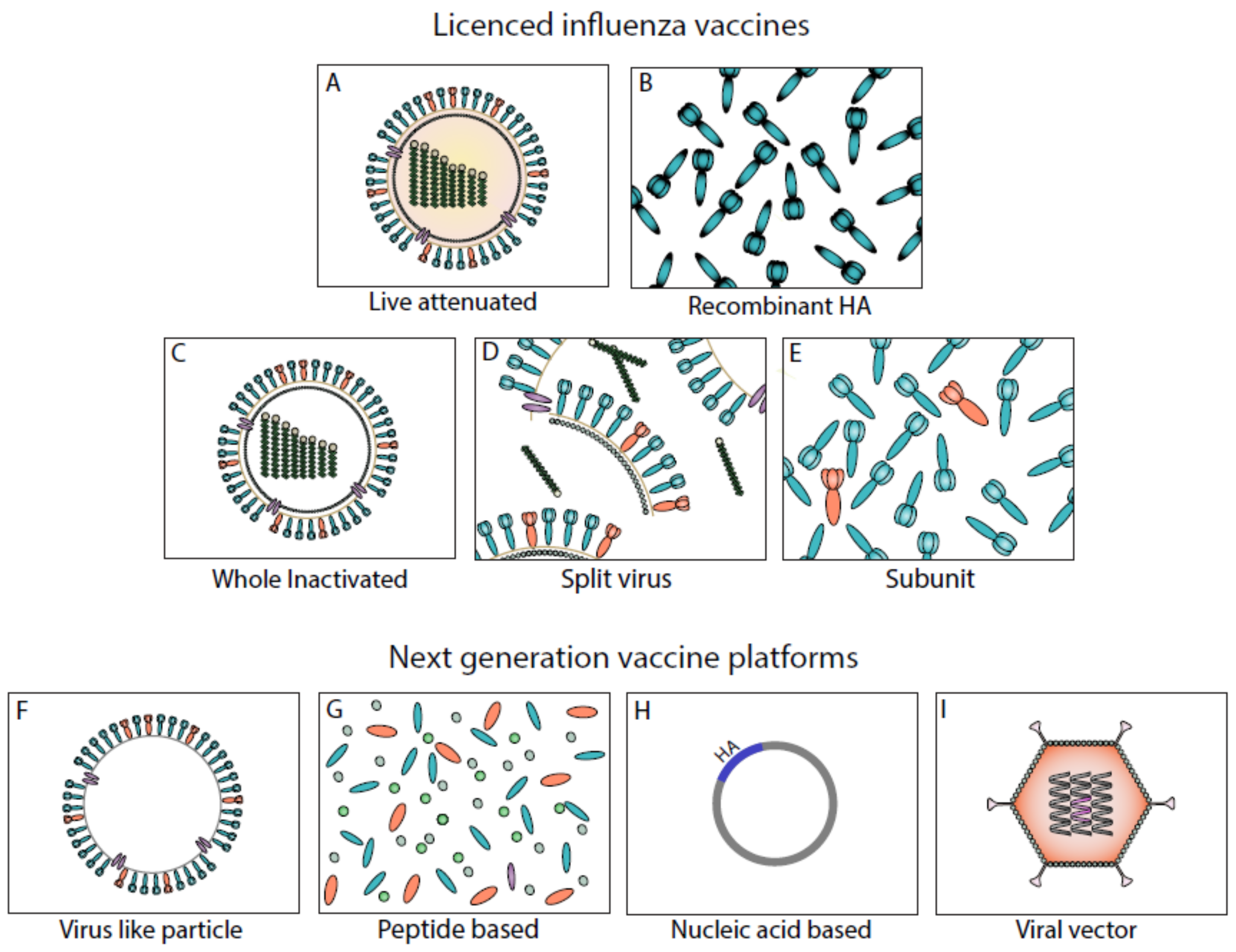
3. Current Influenza Vaccines
4. Next-Generation Vaccine Platforms
5. Universal Vaccine Strategies
6. Challenges in the Development of Universal Vaccines
- (1)
- There is no uniform evaluation of novel influenza vaccines in preclinical trials. This is partly due to the variety of vaccine platforms and targets. In addition, there is no consensus on which animal model is best suited to accurately reflect human infection [3,88]. This is partly due to the existence of pre-existing immunity in humans, which is difficult to translate into animal models. Other host factors to be considered include age, gender, and chronic diseases, which could affect vaccine efficacy.
- (2)
- There are no established correlates of protection for broadly cross-reactive immune responses to influenza. Assays measuring non-HA-head immune responses, such as HA stalk antibodies, NA antibodies, and cellular immunity, are inadequately standardized. Optimization and standardization of such assays are needed in order to assess the protective immunity contributed by broadly reactive or universal vaccines.
- (3)
- The funding for universal vaccine development is limited. This leads to down selection of promising candidates before they advance to clinical trials. Regulatory initiatives are needed to enhance advocacy for moving from an annually reformulated vaccine to a universal vaccine, despite the financial risks and inadequate incentives [6].
7. Lessons Learned Moving Forward with a SARS-CoV-2 Vaccine Design
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Iuliano, A.D.; Roguski, K.M.; Chang, H.H.; Muscatello, D.J.; Palekar, R.; Tempia, S.; Cohen, C.; Gran, J.M.; Schanzer, D.; Cowling, B.J. Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet 2018, 391, 1285–1300. [Google Scholar] [CrossRef]
- Itoh, Y.; Shinya, K.; Kiso, M.; Watanabe, T.; Sakoda, Y.; Hatta, M.; Muramoto, Y.; Tamura, D.; Sakai-Tagawa, Y.; Noda, T. In vitro and in vivo characterization of new swine-origin H1N1 influenza viruses. Nature 2009, 460, 1021–1025. [Google Scholar] [CrossRef] [PubMed]
- Bouvier, N.M. The future of influenza vaccines: A historical and clinical perspective. Vaccines 2018, 6, 58. [Google Scholar] [CrossRef] [PubMed]
- Estrada, L.D.; Schultz-Cherry, S. Development of a universal influenza vaccine. J. Immunol. 2019, 202, 392–398. [Google Scholar] [CrossRef] [PubMed]
- Erbelding, E.J.; Post, D.J.; Stemmy, E.J.; Roberts, P.C.; Augustine, A.D.; Ferguson, S.; Paules, C.I.; Graham, B.S.; Fauci, A.S. A universal influenza vaccine: The strategic plan for the national institute of allergy and infectious diseases. J. Infect. Dis. 2018, 218, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Osterholm, M.T.; Kelley, N.S.; Manske, J.M.; Ballering, K.S.; Leighton, T.R. The Compelling Need for Game-Changing Influenza Vaccines an Analysis of the Influenza Vaccine Enterpris; Center for Infectious Disease Research and Policy: Minneapolis, MN, USA, 2013. [Google Scholar]
- Bournazos, S.; DiLillo, D.J.; Ravetch, J.V. The role of Fc–FcγR interactions in IgG-mediated microbial neutralization. J. Exp. Med. 2015, 212, 1361–1369. [Google Scholar] [CrossRef] [PubMed]
- DiLillo, D.J.; Palese, P.; Wilson, P.C.; Ravetch, J.V. Broadly neutralizing anti-influenza antibodies require Fc receptor engagement for in vivo protection. J. Clin. Investig. 2016, 126, 605–610. [Google Scholar] [CrossRef]
- Dreyfus, C.; Laursen, N.S.; Kwaks, T.; Zuijdgeest, D.; Khayat, R.; Ekiert, D.C.; Lee, J.H.; Metlagel, Z.; Bujny, M.V.; Jongeneelen, M. Highly conserved protective epitopes on influenza B viruses. Science 2012, 337, 1343–1348. [Google Scholar] [CrossRef]
- Throsby, M.; van den Brink, E.; Jongeneelen, M.; Poon, L.L.; Alard, P.; Cornelissen, L.; Bakker, A.; Cox, F.; van Deventer, E.; Guan, Y. Heterosubtypic neutralizing monoclonal antibodies cross-protective against H5N1 and H1N1 recovered from human IgM+ memory B cells. PLoS ONE 2008, 3, e3942. [Google Scholar] [CrossRef]
- Eichelberger, M.C.; Morens, D.M.; Taubenberger, J.K. Neuraminidase as an influenza vaccine antigen: A low hanging fruit, ready for picking to improve vaccine effectiveness. Curr. Opin. Immunol. 2018, 53, 38–44. [Google Scholar] [CrossRef]
- Krammer, F.; Fouchier, R.A.; Eichelberger, M.C.; Webby, R.J.; Shaw-Saliba, K.; Wan, H.; Wilson, P.C.; Compans, R.W.; Skountzou, I.; Monto, A.S. NAction! How can neuraminidase-based immunity contribute to better influenza virus vaccines? MBio 2018, 9, e02332-17. [Google Scholar] [CrossRef] [PubMed]
- Couch, R.B.; Atmar, R.L.; Franco, L.M.; Quarles, J.M.; Wells, J.; Arden, N.; Niño, D.; Belmont, J.W. Antibody correlates and predictors of immunity to naturally occurring influenza in humans and the importance of antibody to the neuraminidase. J. Infect. Dis. 2013, 207, 974–981. [Google Scholar] [CrossRef] [PubMed]
- Memoli, M.J.; Shaw, P.A.; Han, A.; Czajkowski, L.; Reed, S.; Athota, R.; Bristol, T.; Fargis, S.; Risos, K.; Powers, J.H. Evaluation of antihemagglutinin and antineuraminidase antibodies as correlates of protection in an influenza A/H1N1 virus healthy human challenge model. MBio 2016, 7, e00417-16. [Google Scholar] [CrossRef]
- Monto, A.S.; Petrie, J.G.; Cross, R.T.; Johnson, E.; Liu, M.; Zhong, W.; Levine, M.; Katz, J.M.; Ohmit, S.E. Antibody to influenza virus neuraminidase: An independent correlate of protection. J. Infect. Dis. 2015, 212, 1191–1199. [Google Scholar] [CrossRef] [PubMed]
- Stadlbauer, D.; Zhu, X.; McMahon, M.; Turner, J.S.; Wohlbold, T.J.; Schmitz, A.J.; Strohmeier, S.; Yu, W.; Nachbagauer, R.; Mudd, P.A. Broadly protective human antibodies that target the active site of influenza virus neuraminidase. Science 2019, 366, 499–504. [Google Scholar] [CrossRef] [PubMed]
- Wohlbold, T.J.; Podolsky, K.A.; Chromikova, V.; Kirkpatrick, E.; Falconieri, V.; Meade, P.; Amanat, F.; Tan, J.; Tan, G.S.; Subramaniam, S. Broadly protective murine monoclonal antibodies against influenza B virus target highly conserved neuraminidase epitopes. Nat. Microbiol. 2017, 2, 1415–1424. [Google Scholar] [CrossRef]
- Jegerlehner, A.; Schmitz, N.; Storni, T.; Bachmann, M.F. Influenza A vaccine based on the extracellular domain of M2: Weak protection mediated via antibody-dependent NK cell activity. J. Immunol. 2004, 172, 5598–5605. [Google Scholar] [CrossRef] [PubMed]
- Simhadri, V.R.; Dimitrova, M.; Mariano, J.L.; Zenarruzabeitia, O.; Zhong, W.; Ozawa, T.; Muraguchi, A.; Kishi, H.; Eichelberger, M.C.; Borrego, F. A human anti-M2 antibody mediates antibody-dependent cell-mediated cytotoxicity (ADCC) and cytokine secretion by resting and cytokine-preactivated natural killer (NK) cells. PLoS ONE 2015, 10, e0124677. [Google Scholar] [CrossRef]
- Wang, R.; Song, A.; Levin, J.; Dennis, D.; Zhang, N.J.; Yoshida, H.; Koriazova, L.; Madura, L.; Shapiro, L.; Matsumoto, A. Therapeutic potential of a fully human monoclonal antibody against influenza A virus M2 protein. Antivir. Res. 2008, 80, 168–177. [Google Scholar] [CrossRef]
- Eliasson, D.; Omokanye, A.; Schön, K.; Wenzel, U.; Bernasconi, V.; Bemark, M.; Kolpe, A.; El Bakkouri, K.; Ysenbaert, T.; Deng, L. M2e-tetramer-specific memory CD4 T cells are broadly protective against influenza infection. Mucosal Immunol. 2018, 11, 273–289. [Google Scholar] [CrossRef]
- Kim, M.-C.; Lee, Y.-N.; Ko, E.-J.; Lee, J.S.; Kwon, Y.-M.; Hwang, H.S.; Song, J.-M.; Song, B.-M.; Lee, Y.-J.; Choi, J.-G. Supplementation of influenza split vaccines with conserved M2 ectodomains overcomes strain specificity and provides long-term cross protection. Mol. Ther. 2014, 22, 1364–1374. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.N.; Lee, Y.T.; Kim, M.C.; Hwang, H.S.; Lee, J.S.; Kim, K.H.; Kang, S.M. Fc receptor is not required for inducing antibodies but plays a critical role in conferring protection after influenza M 2 vaccination. Immunology 2014, 143, 300–309. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Huang, J.-H.; Yuan, X.-Y.; Huang, W.-S.; Chen, Y.-H. Characterization of immunity induced by M2e of influenza virus. Vaccine 2007, 25, 8868–8873. [Google Scholar] [CrossRef] [PubMed]
- Doucet, J.-D.; Forget, M.-A.; Grange, C.; Rouxel, R.N.; Arbour, N.; von Messling, V.; Lapointe, R. Endogenously expressed matrix protein M1 and nucleoprotein of influenza A are efficiently presented by class I and class II major histocompatibility complexes. J. Gen. Virol. 2011, 92, 1162–1171. [Google Scholar] [CrossRef]
- Forrest, B.D.; Pride, M.W.; Dunning, A.J.; Capeding, M.R.Z.; Chotpitayasunondh, T.; Tam, J.S.; Rappaport, R.; Eldridge, J.H.; Gruber, W.C. Correlation of cellular immune responses with protection against culture-confirmed influenza virus in young children. Clin. Vaccine Immunol. 2008, 15, 1042–1053. [Google Scholar] [CrossRef]
- Hayward, A.C.; Wang, L.; Goonetilleke, N.; Fragaszy, E.B.; Bermingham, A.; Copas, A.; Dukes, O.; Millett, E.R.; Nazareth, I.; Nguyen-Van-Tam, J.S. Natural T cell–mediated protection against seasonal and pandemic influenza. Results of the flu watch cohort study. Am. J. Respir. Crit. Care Med. 2015, 191, 1422–1431. [Google Scholar] [CrossRef]
- McMichael, A.J.; Gotch, F.M.; Noble, G.R.; Beare, P.A. Cytotoxic T-cell immunity to influenza. N. Engl. J. Med. 1983, 309, 13–17. [Google Scholar] [CrossRef]
- Wilkinson, T.M.; Li, C.K.; Chui, C.S.; Huang, A.K.; Perkins, M.; Liebner, J.C.; Lambkin-Williams, R.; Gilbert, A.; Oxford, J.; Nicholas, B. Preexisting influenza-specific CD4+ T cells correlate with disease protection against influenza challenge in humans. Nat. Med. 2012, 18, 274–280. [Google Scholar] [CrossRef]
- Kim, H.; Webster, R.G.; Webby, R.J. Influenza virus: Dealing with a drifting and shifting pathogen. Viral Immunol. 2018, 31, 174–183. [Google Scholar] [CrossRef]
- Allen, J.D.; Ross, T.M. H3N2 influenza viruses in humans: Viral mechanisms, evolution, and evaluation. Hum. Vaccines Immunother. 2018, 14, 1840–1847. [Google Scholar] [CrossRef]
- Harding, A.T.; Heaton, N.S. Efforts to improve the seasonal influenza vaccine. Vaccines 2018, 6, 19. [Google Scholar] [CrossRef] [PubMed]
- Cox, M.M.; Izikson, R.; Post, P.; Dunkle, L. Safety, efficacy, and immunogenicity of Flublok in the prevention of seasonal influenza in adults. Ther. Adv. Vaccines 2015, 3, 97–108. [Google Scholar] [CrossRef] [PubMed]
- Tregoning, J.S.; Russell, R.F.; Kinnear, E. Adjuvanted influenza vaccines. Hum. Vaccines Immunother. 2018, 14, 550–564. [Google Scholar] [CrossRef] [PubMed]
- Galson, J.; Trück, J.; Kelly, D.; Van der Most, R. Investigating the effect of AS03 adjuvant on the plasma cell repertoire following pH1N1 influenza vaccination. Sci. Rep. 2016, 6, 37229. [Google Scholar] [CrossRef] [PubMed]
- Morel, S.; Didierlaurent, A.; Bourguignon, P.; Delhaye, S.; Baras, B.; Jacob, V.; Planty, C.; Elouahabi, A.; Harvengt, P.; Carlsen, H. Adjuvant System AS03 containing α-tocopherol modulates innate immune response and leads to improved adaptive immunity. Vaccine 2011, 29, 2461–2473. [Google Scholar] [CrossRef]
- Ghendon, Y.; Polezhaev, F.; Lisovskaya, K.; Medvedeva, T.; Alexandrova, G.; Klimov, A. Recombinant cold-adapted attenuated influenza A vaccines for use in children: Molecular genetic analysis of the cold-adapted donor and recombinants. Infect. Immun. 1984, 44, 730–733. [Google Scholar] [CrossRef]
- Jin, H.; Lu, B.; Zhou, H.; Ma, C.; Zhao, J.; Yang, C.-F.; Kemble, G.; Greenberg, H. Multiple amino acid residues confer temperature sensitivity to human influenza virus vaccine strains (FluMist) derived from cold-adapted A/Ann Arbor/6/60. Virology 2003, 306, 18–24. [Google Scholar] [CrossRef]
- Ambrose, C.S.; Wu, X.; Jones, T.; Mallory, R.M. The role of nasal IgA in children vaccinated with live attenuated influenza vaccine. Vaccine 2012, 30, 6794–6801. [Google Scholar] [CrossRef]
- Hoft, D.F.; Lottenbach, K.R.; Blazevic, A.; Turan, A.; Blevins, T.P.; Pacatte, T.P.; Yu, Y.; Mitchell, M.C.; Hoft, S.G.; Belshe, R.B. Comparisons of the humoral and cellular immune responses induced by live attenuated influenza vaccine and inactivated influenza vaccine in adults. Clin. Vaccine Immunol. 2017, 24, e00414-16. [Google Scholar] [CrossRef]
- Mohn, K.G.-I.; Smith, I.; Sjursen, H.; Cox, R.J. Immune responses after live attenuated influenza vaccination. Hum. Vaccines Immunother. 2018, 14, 571–578. [Google Scholar] [CrossRef]
- Belshe, R.B.; Edwards, K.M.; Vesikari, T.; Black, S.V.; Walker, R.E.; Hultquist, M.; Kemble, G.; Connor, E.M. Live attenuated versus inactivated influenza vaccine in infants and young children. N. Engl. J. Med. 2007, 356, 685–696. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Meldgaard, T.S.; Bertholet, S. Novel platforms for the development of a universal influenza vaccine. Front. Immunol. 2018, 9, 600. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.-M.J.; Chien, C.-Y.; Liu, M.-T.; Fang, Z.-S.; Chang, S.-Y.; Juang, R.-H.; Chang, S.-C.; Chen, H.-W. Multi-antigen avian influenza a (H7N9) virus-like particles: Particulate characterizations and immunogenicity evaluation in murine and avian models. BMC Biotechnol. 2017, 17, 2. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.-C.; Song, J.-M.; Eunju, O.; Kwon, Y.-M.; Lee, Y.-J.; Compans, R.W.; Kang, S.-M. Virus-like particles containing multiple M2 extracellular domains confer improved cross-protection against various subtypes of influenza virus. Mol. Ther. 2013, 21, 485–492. [Google Scholar] [CrossRef]
- Schwartzman, L.M.; Cathcart, A.L.; Pujanauski, L.M.; Qi, L.; Kash, J.C.; Taubenberger, J.K. An intranasal virus-like particle vaccine broadly protects mice from multiple subtypes of influenza A virus. mBio 2015, 6. [Google Scholar] [CrossRef]
- Steinhagen, F.; Kinjo, T.; Bode, C.; Klinman, D.M. TLR-based immune adjuvants. Vaccine 2011, 29, 3341–3355. [Google Scholar] [CrossRef]
- Deng, L.; Chang, T.Z.; Wang, Y.; Li, S.; Wang, S.; Matsuyama, S.; Yu, G.; Compans, R.W.; Li, J.-D.; Prausnitz, M.R. Heterosubtypic influenza protection elicited by double-layered polypeptide nanoparticles in mice. Proc. Natl. Acad. Sci. USA 2018, 115, E7758–E7767. [Google Scholar] [CrossRef]
- Stoloff, G.A.; Caparros-Wanderley, W. Synthetic multi-epitope peptides identified in silico induce protective immunity against multiple influenza serotypes. Eur. J. Immunol. 2007, 37, 2441–2449. [Google Scholar] [CrossRef]
- Hekele, A.; Bertholet, S.; Archer, J.; Gibson, D.G.; Palladino, G.; Brito, L.A.; Otten, G.R.; Brazzoli, M.; Buccato, S.; Bonci, A. Rapidly produced SAM® vaccine against H7N9 influenza is immunogenic in mice. Emerg. Microbes Infect. 2013, 2, 1–7. [Google Scholar] [CrossRef]
- Martin, J.E.; Louder, M.K.; Holman, L.A.; Gordon, I.J.; Enama, M.E.; Larkin, B.D.; Andrews, C.A.; Vogel, L.; Koup, R.A.; Roederer, M. A SARS DNA vaccine induces neutralizing antibody and cellular immune responses in healthy adults in a Phase I clinical trial. Vaccine 2008, 26, 6338–6343. [Google Scholar] [CrossRef]
- de Vries, R.D.; Rimmelzwaan, G.F. Viral vector-based influenza vaccines. Hum. Vaccines Immunother. 2016, 12, 2881–2901. [Google Scholar] [CrossRef] [PubMed]
- Gurwith, M.; Lock, M.; Taylor, E.M.; Ishioka, G.; Alexander, J.; Mayall, T.; Ervin, J.E.; Greenberg, R.N.; Strout, C.; Treanor, J.J. Safety and immunogenicity of an oral, replicating adenovirus serotype 4 vector vaccine for H5N1 influenza: A randomised, double-blind, placebo-controlled, phase 1 study. Lancet Infect. Dis. 2013, 13, 238–250. [Google Scholar] [CrossRef]
- Lillie, P.J.; Berthoud, T.K.; Powell, T.J.; Lambe, T.; Mullarkey, C.; Spencer, A.J.; Hamill, M.; Peng, Y.; Blais, M.-E.; Duncan, C.J. Preliminary assessment of the efficacy of a t-cell–based influenza vaccine, MVA-NP+ M1, in humans. Clin. Infect. Dis. 2012, 55, 19–25. [Google Scholar] [CrossRef]
- Van Kampen, K.R.; Shi, Z.; Gao, P.; Zhang, J.; Foster, K.W.; Chen, D.-T.; Marks, D.; Elmets, C.A.; De-chu, C.T. Safety and immunogenicity of adenovirus-vectored nasal and epicutaneous influenza vaccines in humans. Vaccine 2005, 23, 1029–1036. [Google Scholar] [CrossRef] [PubMed]
- Swayne, D.E.; Beck, J.R.; Kinney, N. Failure of a recombinant fowl poxvirus vaccine containing an avian influenza hemagglutinin gene to provide consistent protection against influenza in chickens preimmunized with a fowl pox vaccine. Avian Dis. 2000, 44, 132–137. [Google Scholar] [CrossRef] [PubMed]
- Krammer, F.; Palese, P. Universal influenza virus vaccines that target the conserved hemagglutinin stalk and conserved sites in the head domain. J. Infect. Dis. 2019, 219, S62–S67. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Welsh, J.P.; Swartz, J.R. Production and stabilization of the trimeric influenza hemagglutinin stem domain for potentially broadly protective influenza vaccines. Proc. Natl. Acad. Sci. USA 2014, 111, 125–130. [Google Scholar] [CrossRef] [PubMed]
- Impagliazzo, A.; Milder, F.; Kuipers, H.; Wagner, M.V.; Zhu, X.; Hoffman, R.M.; van Meersbergen, R.; Huizingh, J.; Wanningen, P.; Verspuij, J. A stable trimeric influenza hemagglutinin stem as a broadly protective immunogen. Science 2015, 349, 1301–1306. [Google Scholar] [CrossRef]
- Steel, J.; Lowen, A.C.; Wang, T.T.; Yondola, M.; Gao, Q.; Haye, K.; García-Sastre, A.; Palese, P. Influenza virus vaccine based on the conserved hemagglutinin stalk domain. mBio 2010, 1, e00018-10. [Google Scholar] [CrossRef]
- van der Lubbe, J.E.; Huizingh, J.; Verspuij, J.W.; Tettero, L.; Schmit-Tillemans, S.P.; Mooij, P.; Mortier, D.; Koopman, G.; Bogers, W.M.; Dekking, L. Mini-hemagglutinin vaccination induces cross-reactive antibodies in pre-exposed NHP that protect mice against lethal influenza challenge. NPJ Vaccines 2018, 3, 1–9. [Google Scholar] [CrossRef]
- Yassine, H.M.; Boyington, J.C.; McTamney, P.M.; Wei, C.-J.; Kanekiyo, M.; Kong, W.-P.; Gallagher, J.R.; Wang, L.; Zhang, Y.; Joyce, M.G. Hemagglutinin-stem nanoparticles generate heterosubtypic influenza protection. Nat. Med. 2015, 21, 1065–1070. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, D.I.; Guptill, J.; Naficy, A.; Nachbagauer, R.; Berlanda-Scorza, F.; Feser, J.; Wilson, P.C.; Solórzano, A.; Van der Wielen, M.; Walter, E.B. Immunogenicity of chimeric haemagglutinin-based, universal influenza virus vaccine candidates: Interim results of a randomised, placebo-controlled, phase 1 clinical trial. Lancet Infect. Dis. 2020, 20, 80–91. [Google Scholar] [CrossRef]
- Ekiert, D.C.; Kashyap, A.K.; Steel, J.; Rubrum, A.; Bhabha, G.; Khayat, R.; Lee, J.H.; Dillon, M.A.; O’Neil, R.E.; Faynboym, A.M. Cross-neutralization of influenza A viruses mediated by a single antibody loop. Nature 2012, 489, 526–532. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.S.; Ohshima, N.; Stanfield, R.L.; Yu, W.; Iba, Y.; Okuno, Y.; Kurosawa, Y.; Wilson, I.A. Receptor mimicry by antibody F045–092 facilitates universal binding to the H3 subtype of influenza virus. Nat. Commun. 2014, 5, 1–9. [Google Scholar] [CrossRef]
- Lee, P.S.; Yoshida, R.; Ekiert, D.C.; Sakai, N.; Suzuki, Y.; Takada, A.; Wilson, I.A. Heterosubtypic antibody recognition of the influenza virus hemagglutinin receptor binding site enhanced by avidity. Proc. Natl. Acad. Sci. USA 2012, 109, 17040–17045. [Google Scholar] [CrossRef]
- Carter, D.M.; Darby, C.A.; Lefoley, B.C.; Crevar, C.J.; Alefantis, T.; Oomen, R.; Anderson, S.F.; Strugnell, T.; Cortés-Garcia, G.; Vogel, T.U. Design and characterization of a computationally optimized broadly reactive hemagglutinin vaccine for H1N1 influenza viruses. J. Virol. 2016, 90, 4720–4734. [Google Scholar] [CrossRef]
- Giles, B.M.; Ross, T.M. A computationally optimized broadly reactive antigen (COBRA) based H5N1 VLP vaccine elicits broadly reactive antibodies in mice and ferrets. Vaccine 2011, 29, 3043–3054. [Google Scholar] [CrossRef] [PubMed]
- Sautto, G.A.; Kirchenbaum, G.A.; Abreu, R.B.; Ecker, J.W.; Pierce, S.R.; Kleanthous, H.; Ross, T.M. A computationally optimized broadly reactive antigen subtype–specific influenza vaccine strategy elicits unique potent broadly neutralizing antibodies against hemagglutinin. J. Immunol. 2020, 204, 375–385. [Google Scholar] [CrossRef]
- Kilbourne, E.D.; Couch, R.B.; Kasel, J.A.; Keitel, W.A.; Cate, T.R.; Quarles, J.H.; Grajower, B.; Pokorny, B.A.; Johansson, B.E. Purified influenza A virus N2 neuraminidase vaccine is immunogenic and non-toxic in humans. Vaccine 1995, 13, 1799–1803. [Google Scholar] [CrossRef]
- Wohlbold, T.J.; Nachbagauer, R.; Xu, H.; Tan, G.S.; Hirsh, A.; Brokstad, K.A.; Cox, R.J.; Palese, P.; Krammer, F. Vaccination with adjuvanted recombinant neuraminidase induces broad heterologous, but not heterosubtypic, cross-protection against influenza virus infection in mice. mBio 2015, 6, e02556. [Google Scholar] [CrossRef]
- Mooney, A.J.; Gabbard, J.D.; Li, Z.; Dlugolenski, D.A.; Johnson, S.K.; Tripp, R.A.; He, B.; Tompkins, S.M. Vaccination with recombinant parainfluenza virus 5 expressing neuraminidase protects against homologous and heterologous influenza virus challenge. J. Virol. 2017, 91, e01579-17. [Google Scholar] [CrossRef] [PubMed]
- Meseda, C.A.; Atukorale, V.; Soto, J.; Eichelberger, M.C.; Gao, J.; Wang, W.; Weiss, C.D.; Weir, J.P. Immunogenicity and protection against influenza H7N3 in mice by modified vaccinia virus Ankara vectors expressing influenza virus hemagglutinin or neuraminidase. Sci. Rep. 2018, 8, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.-H.; Lee, Y.-T.; Park, S.; Jung, Y.-J.; Lee, Y.; Ko, E.-J.; Kim, Y.-J.; Li, X.; Kang, S.-M. Neuraminidase expressing virus-like particle vaccine provides effective cross protection against influenza virus. Virology 2019, 535, 179–188. [Google Scholar] [CrossRef]
- Mezhenskaya, D.; Isakova-Sivak, I.; Rudenko, L. M2e-based universal influenza vaccines: A historical overview and new approaches to development. J. Biomed. Sci. 2019, 26, 76. [Google Scholar] [CrossRef]
- Talbot, H.K.; Rock, M.T.; Johnson, C.; Tussey, L.; Kavita, U.; Shanker, A.; Shaw, A.R.; Taylor, D.N. Immunopotentiation of trivalent influenza vaccine when given with VAX102, a recombinant influenza M2e vaccine fused to the TLR5 ligand flagellin. PLoS ONE 2010, 5, e14442. [Google Scholar] [CrossRef] [PubMed]
- Turley, C.B.; Rupp, R.E.; Johnson, C.; Taylor, D.N.; Wolfson, J.; Tussey, L.; Kavita, U.; Stanberry, L.; Shaw, A. Safety and immunogenicity of a recombinant M2e–flagellin influenza vaccine (STF2. 4xM2e) in healthy adults. Vaccine 2011, 29, 5145–5152. [Google Scholar] [CrossRef]
- Lee, Y.-T.; Kim, K.-H.; Ko, E.-J.; Kim, M.-C.; Lee, Y.-N.; Hwang, H.-S.; Lee, Y.; Jung, Y.-J.; Kim, Y.J.; Santos, J. Enhancing the cross protective efficacy of live attenuated influenza virus vaccine by supplemented vaccination with M2 ectodomain virus-like particles. Virology 2019, 529, 111–121. [Google Scholar] [CrossRef]
- Kang, H.-J.; Chu, K.-B.; Lee, D.-H.; Lee, S.-H.; Park, B.R.; Kim, M.-C.; Kang, S.-M.; Quan, F.-S. Influenza M2 virus-like particle vaccination enhances protection in combination with avian influenza HA VLPs. PLoS ONE 2019, 14, e0216871. [Google Scholar] [CrossRef]
- Atsmon, J.; Kate-Ilovitz, E.; Shaikevich, D.; Singer, Y.; Volokhov, I.; Haim, K.Y.; Ben-Yedidia, T. Safety and immunogenicity of multimeric-001—A novel universal influenza vaccine. J. Clin. Immunol. 2012, 32, 595–603. [Google Scholar] [CrossRef]
- Antrobus, R.D.; Lillie, P.J.; Berthoud, T.K.; Spencer, A.J.; McLaren, J.E.; Ladell, K.; Lambe, T.; Milicic, A.; Price, D.A.; Hill, A.V. AT cell-inducing influenza vaccine for the elderly: Safety and immunogenicity of MVA-NP+ M1 in adults aged over 50 years. PLoS ONE 2012, 7, e48322. [Google Scholar] [CrossRef]
- Takeuchi, A.; Badr, M.E.S.G.; Miyauchi, K.; Ishihara, C.; Onishi, R.; Guo, Z.; Sasaki, Y.; Ike, H.; Takumi, A.; Tsuji, N.M. CRTAM determines the CD4+ cytotoxic T lymphocyte lineage. J. Exp. Med. 2016, 213, 123–138. [Google Scholar] [CrossRef]
- Marshall, N.B.; Vong, A.M.; Devarajan, P.; Brauner, M.D.; Kuang, Y.; Nayar, R.; Schutten, E.A.; Castonguay, C.H.; Berg, L.J.; Nutt, S.L. NKG2C/E marks the unique cytotoxic CD4 T cell subset, ThCTL, generated by influenza infection. J. Immunol. 2017, 198, 1142–1155. [Google Scholar] [CrossRef] [PubMed]
- Selin, L.K.; Varga, S.M.; Wong, I.C.; Welsh, R.M. Protective heterologous antiviral immunity and enhanced immunopathogenesis mediated by memory T cell populations. J. Exp. Med. 1998, 188, 1705–1715. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.E.; Knudson, C.J.; Hartwig, S.M.; Pewe, L.L.; Meyerholz, D.K.; Langlois, R.A.; Harty, J.T.; Varga, S.M. Memory CD8 T cells mediate severe immunopathology following respiratory syncytial virus infection. PLoS Pathog. 2018, 14, e1006810. [Google Scholar] [CrossRef] [PubMed]
- Brehm, M.A.; Pinto, A.K.; Daniels, K.A.; Schneck, J.P.; Welsh, R.M.; Selin, L.K. T cell immunodominance and maintenance of memory regulated by unexpectedly cross-reactive pathogens. Nat. Immunol. 2002, 3, 627–634. [Google Scholar] [CrossRef] [PubMed]
- Souquette, A.; Thomas, P.G. Past life and future effects—How heterologous infections alter immunity to influenza viruses. Front. Immunol. 2018, 9, 1071. [Google Scholar] [CrossRef]
- D’Alessio, F.; Koopman, G.; Houard, S.; Remarque, E.J.; Stockhofe, N.; Engelhardt, O.G. Workshop report: Experimental animal models for universal influenza vaccines. Vaccine 2018, 36, 6895–6901. [Google Scholar] [CrossRef]
- Amanat, F.; Krammer, F. SARS-CoV-2 vaccines: Status report. Immunity 2020, 52, 583–589. [Google Scholar] [CrossRef]
- Kim, E.; Okada, K.; Kenniston, T.; Raj, V.S.; AlHajri, M.M.; Farag, E.A.; AlHajri, F.; Osterhaus, A.D.; Haagmans, B.L.; Gambotto, A. Immunogenicity of an adenoviral-based Middle East Respiratory Syndrome coronavirus vaccine in BALB/c mice. Vaccine 2014, 32, 5975–5982. [Google Scholar] [CrossRef]
- Guo, X.; Deng, Y.; Chen, H.; Lan, J.; Wang, W.; Zou, X.; Hung, T.; Lu, Z.; Tan, W. Systemic and mucosal immunity in mice elicited by a single immunization with human adenovirus type 5 or 41 vector-based vaccines carrying the spike protein of Middle East respiratory syndrome coronavirus. Immunology 2015, 145, 476–484. [Google Scholar] [CrossRef]
- Yong, C.Y.; Ong, H.K.; Yeap, S.K.; Ho, K.L.; Tan, W.S. Recent advances in the vaccine development against Middle East respiratory syndrome-coronavirus. Front. Microbiol. 2019, 10, 1781. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Trieu, M.-C.; Davies, R.; Cox, R.J. Improving influenza vaccines: Challenges to effective implementation. Curr. Opin. Immunol. 2018, 53, 88–95. [Google Scholar] [CrossRef] [PubMed]

| Critical | Important | Desired |
|---|---|---|
| Provide protection against all HA subtypes and at a minimum protection against H1, H2, H3, H5, H7 and H9 subtypes. | Provide a decade or more of protection. | Use manufacturing technology that can be readily transferred to developing-world countries. |
| Provide immunologic protection for those populations most at risk for severe disease and increased mortality | Use inexpensive manufacturing technology that permits rapid and highly scalable production, particularly to address emergence of a pandemic virus. | Offer heat stability, thereby eliminating the need to maintain a cold chain. |
| Rarely cause adverse events, and any adverse events are mild and temporary. | Do not require injection for administration. | Do not require injection for administration. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Madsen, A.; Cox, R.J. Prospects and Challenges in the Development of Universal Influenza Vaccines. Vaccines 2020, 8, 361. https://doi.org/10.3390/vaccines8030361
Madsen A, Cox RJ. Prospects and Challenges in the Development of Universal Influenza Vaccines. Vaccines. 2020; 8(3):361. https://doi.org/10.3390/vaccines8030361
Chicago/Turabian StyleMadsen, Anders, and Rebecca Jane Cox. 2020. "Prospects and Challenges in the Development of Universal Influenza Vaccines" Vaccines 8, no. 3: 361. https://doi.org/10.3390/vaccines8030361
APA StyleMadsen, A., & Cox, R. J. (2020). Prospects and Challenges in the Development of Universal Influenza Vaccines. Vaccines, 8(3), 361. https://doi.org/10.3390/vaccines8030361




