Current and Novel Approaches in Influenza Management
Abstract
1. Introduction
2. Current Influenza Vaccines
3. Use of the Seasonal Influenza Vaccines
4. Novel Influenza Vaccine Platforms
5. Virus-Like Particle (VLP) Vaccines
6. Computationally Optimized Broadly Reactive Antigen Vaccines (COBRA) Vaccines
7. Synthetic Influenza Virus Vaccines
8. Epitope Vaccines
9. Antigen-Presenting Cell (APC) Inducible Vaccines
10. Nanoparticle-Based Influenza Vaccines
11. Viral-Vectored Vaccines
12. Current Influenza Managing Antivirals
13. Novel Influenza Management Therapies
13.1. Next-Generation Antivirals Against Influenza
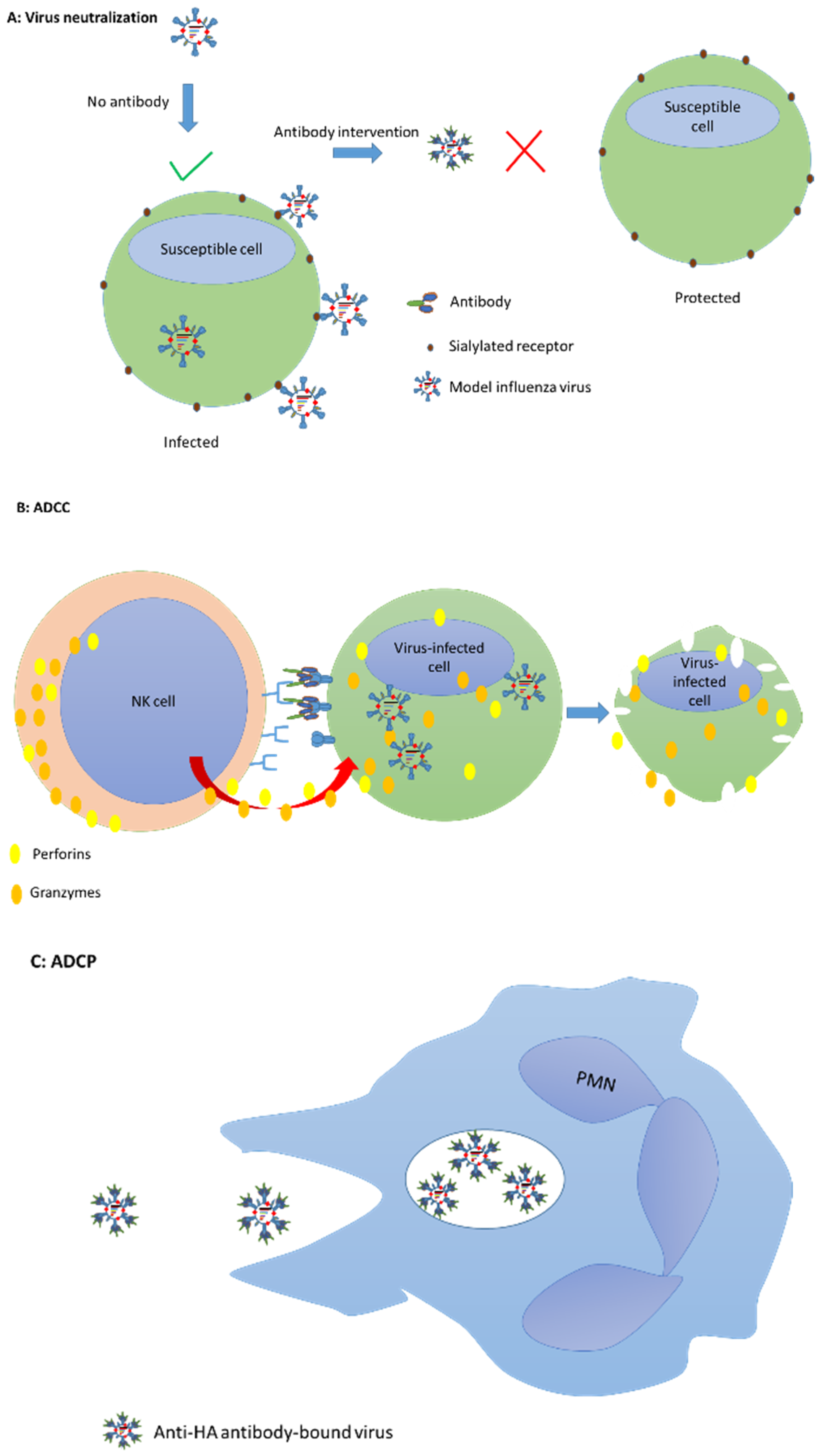
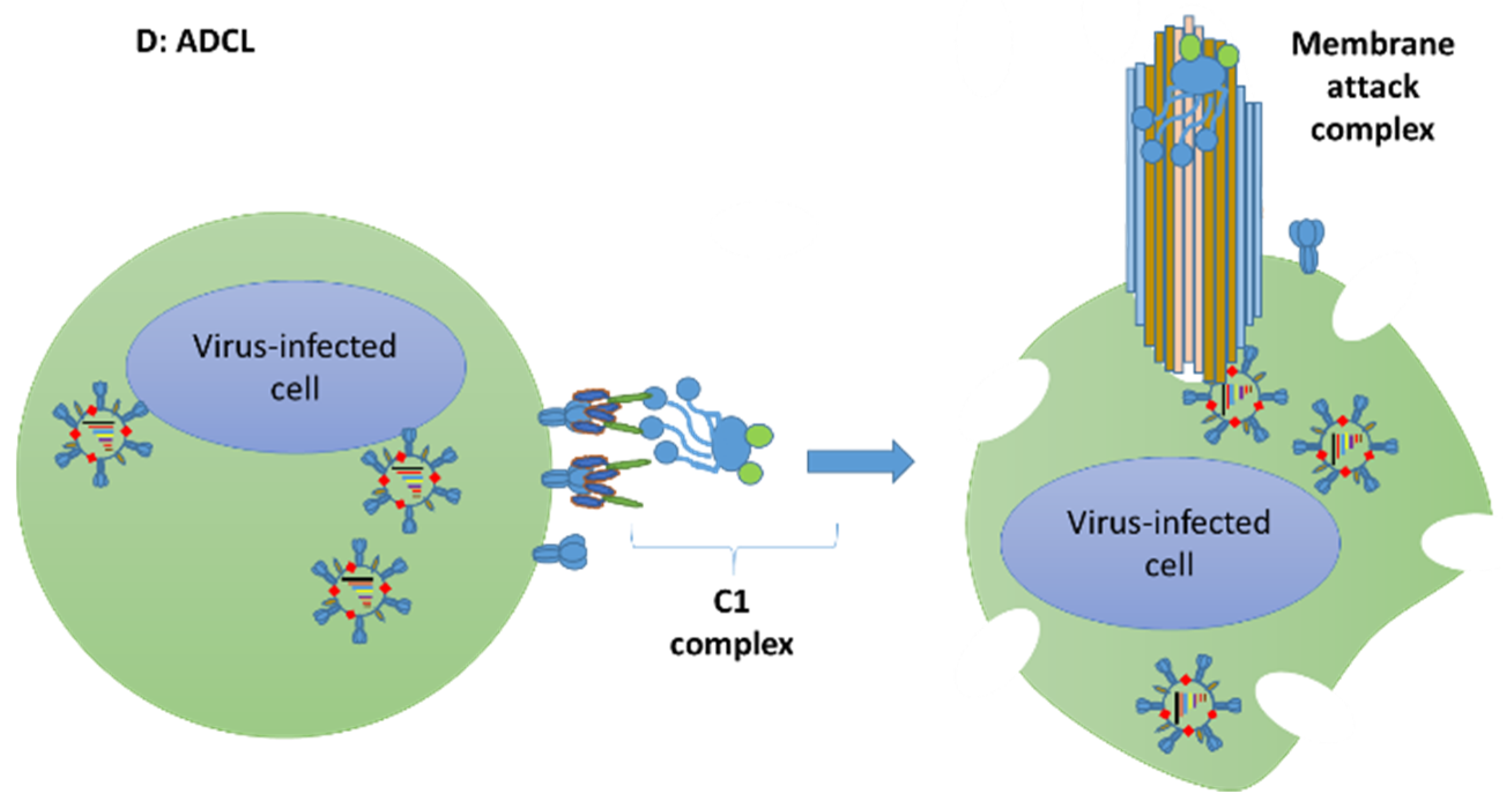
13.2. Passive Immunotherapeutics for Management of Influenza
14. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Su, S.; Fu, X.; Li, G.; Kerlin, F.; Veit, M. Novel Influenza D virus: Epidemiology, pathology, evolution and biological characteristics. Virulence 2017, 8, 1580–1591. [Google Scholar] [CrossRef] [PubMed]
- Kimura, H.; Abiko, C.; Peng, G.; Muraki, Y.; Sugawara, K.; Hongo, S.; Kitame, F.; Mizuta, K.; Numazaki, Y.; Suzuki, H.; et al. Interspecies transmission of influenza C virus between humans and pigs. Virus Res. 1997, 48, 71–79. [Google Scholar] [CrossRef]
- Hinshaw, V.S.; Webster, R.G.; Easterday, B.C.; Bean, W.J., Jr. Replication of avian influenza A viruses in mammals. Infect. Immun. 1981, 34, 354–361. [Google Scholar] [PubMed]
- Osterhaus, A.D.; Rimmelzwaan, G.F.; Martina, B.E.; Bestebroer, T.M.; Fouchier, R.A. Influenza B virus in seals. Science 2000, 288, 1051–1053. [Google Scholar] [CrossRef] [PubMed]
- Gerhard, W.; Webster, R.G. Antigenic drift in influenza A viruses. I. Selection and characterization of antigenic variants of A/PR/8/34 (HON1) influenza virus with monoclonal antibodies. J. Exp. Med. 1978, 148, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Yewdell, J.W.; Webster, R.G.; Gerhard, W.U. Antigenic variation in three distinct determinants of an influenza type A haemagglutinin molecule. Nature 1979, 279, 246–248. [Google Scholar] [CrossRef] [PubMed]
- Hensley, S.E.; Das, S.R.; Bailey, A.L.; Schmidt, L.M.; Hickman, H.D.; Jayaraman, A.; Viswanathan, K.; Raman, R.; Sasisekharan, R.; Bennink, J.R.; et al. Hemagglutinin Receptor Binding Avidity Drives Influenza A Virus Antigenic Drift. Science 2009, 326, 734–736. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, N.M.; Galvani, A.P.; Bush, R.M. Ecological and immunological determinants of influenza evolution. Nature 2003, 422, 428–433. [Google Scholar] [CrossRef]
- Carrat, F.; Flahault, A. Influenza vaccine: The challenge of antigenic drift. Vaccine 2007, 25, 6852–6862. [Google Scholar] [CrossRef]
- Skehel, J.J.; Stevens, D.J.; Daniels, R.S.; Douglas, A.R.; Knossow, M.; Wilson, I.A.; Wiley, D.C. A carbohydrate side chain on hemagglutinins of Hong Kong influenza viruses inhibits recognition by a monoclonal antibody. Proc. Natl. Acad. Sci. USA 1984, 81, 1779–1783. [Google Scholar] [CrossRef]
- Webster, R.G.; Govorkova, E.A. Continuing challenges in influenza. Ann. N. Y. Acad. Sci. 2014, 1323, 115–139. [Google Scholar] [CrossRef] [PubMed]
- Gething, M.J.; Bye, J.; Skehel, J.; Waterfield, M. Cloning and DNA sequence of double-stranded copies of haemagglutinin genes from H2 and H3 strains elucidates antigenic shift and drift in human influenza virus. Nature 1980, 287, 301–306. [Google Scholar] [CrossRef] [PubMed]
- Donatelli, I.; Castrucci, M.R.; De Marco, M.A.; Delogu, M.; Webster, R.G. Human–Animal Interface: The Case for Influenza Interspecies Transmission. Emerging and Re-emerging Viral Infections; Springer: Cham, Switzerland, 2016; pp. 17–33. [Google Scholar]
- Zambon, M.C. Epidemiology and pathogenesis of influenza. J. Antimicrob. Chemother. 1999, 44, 3–9. [Google Scholar] [CrossRef] [PubMed]
- WHO. Recommendations for Production and Control of Influenza Vaccine (Inactivated); WHO Technical Report Series; WHO: Geneva, Switzerland, 2003; pp. 99–134. [Google Scholar]
- Hoft, D.F.; Lottenbach, K.R.; Blazevic, A.; Turan, A.; Blevins, T.P.; Pacatte, T.P.; Yu, Y.; Mitchell, M.C.; Hoft, S.G.; Belshe, R.B. Comparisons of the Humoral and Cellular Immune Responses Induced by Live Attenuated Influenza Vaccine and Inactivated Influenza Vaccine in Adults. Clin. Vaccine Immunol. 2017, 24, e00414-16. [Google Scholar] [CrossRef] [PubMed]
- Shinjoh, M.; Sugaya, N.; Yamaguchi, Y.; Iibuchi, N.; Kamimaki, I.; Goto, A.; Kobayashi, H.; Kobayashi, Y.; Shibata, M.; Tamaoka, S.; et al. Inactivated influenza vaccine effectiveness and an analysis of repeated vaccination for children during the 2016/17 season. Vaccine 2018, 36, 5510–5518. [Google Scholar] [CrossRef] [PubMed]
- Osterholm, M.T.; Kelley, N.S.; Sommer, A.; Belongia, E.A. Efficacy and effectiveness of influenza vaccines: A systematic review and meta-analysis. Lancet Infect. Dis. 2012, 12, 36–44. [Google Scholar] [CrossRef]
- Beyer, W.E.P.; Palache, A.M.; de Jong, J.C.; Osterhaus, A.D.M.E. Cold-adapted live influenza vaccine versus inactivated vaccine: Systemic vaccine reactions, local and systemic antibody response, and vaccine efficacy A meta-analysis. Vaccine 2002, 20, 1340–1353. [Google Scholar] [CrossRef]
- King, J.C.; Treanor, J.; Fast, P.E.; Wolff, M.; Yan, L.H.; Iacuzio, D.; Readmond, B.; O’Brien, D.; Mallon, K.; Highsmith, W.E.; et al. Comparison of the safety, vaccine virus shedding, and immunogenicity of influenza virus vaccine, trivalent, types A and B, live cold-adapted, administered to human immunodeficiency virus (HIV)-infected and non-HIV-infected adults. J. Infect. Dis. 2000, 181, 725–728. [Google Scholar] [CrossRef]
- Belshe, R.B.; Mendelman, P.M.; Treanor, J.; King, J.; Gruber, W.C.; Piedra, P.; Bernstein, D.I.; Hayden, F.G.; Kotloff, K.; Zangwill, K.; et al. The efficacy of live attenuated, cold-adapted, trivalent, intranasal influenzavirus vaccine in children. N. Engl. J. Med. 1998, 338, 1405–1412. [Google Scholar] [CrossRef]
- Keitel, W.A.; Couch, R.B.; Cate, T.R.; Maassab, H.F. Variability in infectivity of cold-adapted recombinant influenza virus vaccines in humans. J. Infect. Dis. 1994, 169, 477. [Google Scholar] [CrossRef]
- Galazka, A.M.; Lauer, B.A.; Henderson, R.H.; Keja, J. Indications and Contraindications for Vaccines Used in the Expanded Program on Immunization. Bull. World Health Organ. 1984, 62, 357–366. [Google Scholar] [PubMed]
- Kelso, J.M.; Greenhawt, M.J.; Li, J.T.; Nicklas, R.A.; Bernstein, D.I.; Blessing-Moore, J.; Cox, L.; Khan, D.; Lang, D.M.; Oppenheimer, J.; et al. Adverse reactions to vaccines practice parameter 2012 update. J. Allergy Clin. Immunol. 2012, 130, 25–43. [Google Scholar] [CrossRef] [PubMed]
- Owens, G.; MacGinnitie, A. Higher-ovalbumin-content influenza vaccines are well tolerated in children with egg allergy. J. Allergy Clin. Immunol. 2011, 127, 264–265. [Google Scholar] [CrossRef] [PubMed]
- Li, J.T.; Rank, M.A.; Squillace, D.L.; Kita, H. Ovalbumin content of influenza vaccines. J. Allergy Clin. Immunol. 2010, 125, 1412–1413. [Google Scholar] [CrossRef][Green Version]
- Grohskopf, L.A.; Olsen, S.J.; Sokolow, L.Z.; Bresee, J.S.; Cox, N.J.; Broder, K.R.; Karron, R.A.; Walter, E.B.; Centers for Disease Control and Prevention. Prevention and Control of Seasonal Influenza with Vaccines: Recommendations of the Advisory Committee on Immunization Practices (ACIP)—United States, 2014–2015 Influenza Season. Am. J. Transplant. 2014, 14, 2906–2913. [Google Scholar]
- Stephenson, I.; Nicholson, K.G.; Gluck, R.; Mischler, R.; Newman, R.W.; Palache, A.M.; Verlander, N.Q.; Warburton, F.; Wood, J.M.; Zambon, M.C. Safety and antigenicity of whole virus and subunit influenza A/Hong Kong/1073/99 (H9N2) vaccine in healthy adults: Phase I randomised trial. Lancet 2003, 362, 1959–1966. [Google Scholar] [CrossRef]
- Giezeman, K.M.; Nauta, J.; de Bruijn, I.A.; Palache, A.M. Trivalent inactivated subunit influenza vaccine Influvac (R): 25-Year experience of safety and immunogenicity. Vaccine 2009, 27, 2414–2417. [Google Scholar] [CrossRef]
- Ambrose, C.S.; Levin, M.J. The rationale for quadrivalent influenza vaccines. Hum. Vaccines Immunother. 2012, 8, 81–88. [Google Scholar] [CrossRef]
- Kumar, A.; Meldgaard, T.S.; Bertholet, S. Novel Platforms for the Development of a Universal influenza vaccine. Front. Immunol. 2018, 9, 600. [Google Scholar] [CrossRef]
- Fiore, A.E.; Uyeki, T.M.; Broder, K.; Finelli, L.; Euler, G.L.; Singleton, J.A.; Iskander, J.K.; Wortley, P.M.; Shay, D.K.; Bresee, J.S.; et al. Prevention and control of influenza with vaccines: Recommendations of the Advisory Committee on Immunization Practices (ACIP), 2010. MMWR Recomm. Rep. 2010, 59, 1–62. [Google Scholar]
- WHO. Who/Europe Recommendations on Influenza Vaccination during the 2011–2012 Winter Season; WHO: Geneva, Switzerland, 2011. [Google Scholar]
- Xu, C.; Thompson, M.G.; Cowling, B.J. Influenza vaccination in tropical and subtropical areas. Lancet Respir. Med. 2017, 5, 920–922. [Google Scholar] [CrossRef]
- Duque, J.; McMorrow, M.L.; Cohen, A.L. Influenza vaccines and influenza antiviral drugs in Africa: Are they available and do guidelines for their use exist? BMC Public Health 2014, 14, 41. [Google Scholar] [CrossRef] [PubMed]
- Grgacic, E.V.; Anderson, D.A. Virus-like particles: Passport to immune recognition. Methods 2006, 40, 60–65. [Google Scholar] [CrossRef] [PubMed]
- Roldao, A.; Mellado, M.C.M.; Castilho, L.R.; Carrondo, M.J.T.; Alves, P.M. Virus-like particles in vaccine development. Expert Rev. Vaccines 2010, 9, 1149–1176. [Google Scholar] [CrossRef]
- Zhao, Q.; Li, S.; Yu, H.; Xia, N.; Modis, Y. Virus-like particle-based human vaccines: Quality assessment based on structural and functional properties. Trends Biotechnol. 2013, 31, 654–663. [Google Scholar] [CrossRef]
- Gao, X.; Wang, W.; Li, Y.; Zhang, S.; Duan, Y.; Xing, L.; Zhao, Z.; Zhang, P.; Li, Z.; Li, R.; et al. Enhanced Influenza VLP vaccines comprising matrix-2 ectodomain and nucleoprotein epitopes protects mice from lethal challenge. Antiviral Res. 2013, 98, 4–11. [Google Scholar] [CrossRef]
- Kapczynski, D.R.; Tumpey, T.M.; Hidajat, R.; Zsak, A.; Chrzastek, K.; Tretyakova, I.; Pushko, P. Vaccination with virus-like particles containing H5 antigens from three H5N1 clades protects chickens from H5N1 and H5N8 influenza viruses. Vaccine 2016, 34, 1575–1581. [Google Scholar] [CrossRef]
- Wang, B.-Z.; Quan, F.-S.; Kang, S.-M.; Bozja, J.; Skountzou, I.; Compans, R.W. Incorporation of membrane-anchored flagellin into influenza virus-like particles enhances the breadth of immune responses. J. Virol. 2008, 82, 11813–11823. [Google Scholar] [CrossRef]
- Mohan, T.; Berman, Z.; Luo, Y.; Wang, C.; Wang, S.; Compans, R.W.; Wang, B.Z. Chimeric virus-like particles containing influenza HA antigen and GPI-CCL28 induce long-lasting mucosal immunity against H3N2 viruses. Sci. Rep. 2017, 7, 40226. [Google Scholar] [CrossRef]
- Giles, B.M.; Ross, T.M. A computationally optimized broadly reactive antigen (COBRA) based H5N1 VLP vaccine elicits broadly reactive antibodies in mice and ferrets. Vaccine 2011, 29, 3043–3054. [Google Scholar] [CrossRef]
- Giles, B.M.; Bissel, S.J.; Dealmeida, D.R.; Wiley, C.A.; Ross, T.M. Antibody breadth and protective efficacy are increased by vaccination with computationally optimized hemagglutinin but not with polyvalent hemagglutinin-based H5N1 virus-like particle vaccines. Clin. Vaccine Immunol. 2012, 19, 128–139. [Google Scholar] [CrossRef] [PubMed]
- Crevar, C.J.; Carter, D.M.; Lee, K.Y.; Ross, T.M. Cocktail of H5N1 COBRA HA vaccines elicit protective antibodies against H5N1 viruses from multiple clades. Hum. Vaccine Immunol. 2015, 11, 572–583. [Google Scholar] [CrossRef] [PubMed]
- Carter, D.M.; Darby, C.A.; Lefoley, B.C.; Crevar, C.J.; Alefantis, T.; Oomen, R.; Anderson, S.F.; Strugnell, T.; Cortés-Garcia, G.; Vogel, T.U.; et al. Design and Characterization of a Computationally Optimized Broadly Reactive Hemagglutinin Vaccine for H1N1 Influenza Viruses. J. Virol. 2016, 90, 4720–4734. [Google Scholar] [CrossRef] [PubMed]
- Wong, T.M.; Allen, J.D.; Bebin-Blackwell, A.G.; Carter, D.M.; Alefantis, T.; DiNapoli, J.; Kleanthous, H.; Ross, T.M. Computationally Optimized Broadly Reactive Hemagglutinin Elicits Hemagglutination Inhibition Antibodies against a Panel of H3N2 Influenza Virus Cocirculating Variants. J. Virol. 2017, 91, e01581-17. [Google Scholar] [CrossRef] [PubMed]
- Fan, R.L.Y.; Valkenburg, S.A.; Wong, C.K.S.; Li, O.T.W.; Nicholls, J.M.; Rabadan, R.; Peiris, J.S.; Poon, L.L. Generation of Live Attenuated Influenza Virus by Using Codon Usage Bias. J. Virol. 2015, 89, 10762–10773. [Google Scholar] [CrossRef] [PubMed]
- Pica, N.; Langlois, R.A.; Krammer, F.; Margine, I.; Palese, P. NS1-truncated live attenuated virus vaccine provides robust protection to aged mice from viral challenge. J. Virol. 2012, 86, 10293–102301. [Google Scholar] [CrossRef] [PubMed]
- Mossler, C.; Groiss, F.; Wolzt, M.; Wolschek, M.; Seipelt, J.; Muster, T. Phase I/II trial of a replication-deficient trivalent influenza virus vaccine lacking NS1. Vaccine 2013, 31, 6194–6200. [Google Scholar] [CrossRef] [PubMed]
- Baz, M.; Boonnak, K.; Paskel, M.; Santos, C.; Powell, T.; Townsend, A.; Subbarao, K. Nonreplicating influenza A virus vaccines confer broad protection against lethal challenge. MBio 2015, 6, e01487-15. [Google Scholar] [CrossRef]
- Holzer, B.; Morgan, S.B.; Matsuoka, Y.; Edmans, M.; Salguero, F.J.; Everett, H.; Brookes, S.M.; Porter, E.; MacLoughlin, R.; Charleston, B.; et al. Comparison of Heterosubtypic Protection in Ferrets and Pigs Induced by a Single-Cycle Influenza Vaccine. J. Immunol. 2018, 200, 4068–4077. [Google Scholar] [CrossRef]
- Nachbagauer, R.; Liu, W.C.; Choi, A.; Wohlbold, T.J.; Atlas, T.; Rajendran, M.; Solórzano, A.; Berlanda-Scorza, F.; García-Sastre, A.; Palese, P.; et al. A universal influenza virus vaccine candidate confers protection against pandemic H1N1 infection in preclinical ferret studies. Npj Vaccines 2017, 2, 26. [Google Scholar] [CrossRef]
- Impagliazzo, A.; Milder, F.; Kuipers, H.; Wagner, M.V.; Zhu, X.Y.; Hoffman, R.M.B.; van Meersbergen, R.; Huizingh, J.; Wanningen, P.; Verspuij, J.; et al. A stable trimeric influenza hemagglutinin stem as a broadly protective immunogen. Science 2015, 349, 1301–1306. [Google Scholar] [CrossRef] [PubMed]
- Correia, B.E.; Bates, J.T.; Loomis, R.J.; Baneyx, G.; Carrico, C.; Jardine, J.G.; Rupert, P.; Correnti, C.; Kalyuzhniy, O.; Vittal, V.; et al. Proof of principle for epitope-focused vaccine design. Nature 2014, 507, 201–206. [Google Scholar] [CrossRef] [PubMed]
- McLellan, J.S.; Chen, M.; Joyce, M.G.; Sastry, M.; Stewart-Jones, G.B.; Yang, Y.; Zhang, B.; Chen, L.; Srivatsan, S.; Zheng, A.; et al. Structure-based design of a fusion glycoprotein vaccine for respiratory syncytial virus. Science 2013, 342, 592–598. [Google Scholar] [CrossRef] [PubMed]
- Thompson, C.P.; Lourenco, J.; Walters, A.A.; Obolski, U.; Edmans, M.; Palmer, D.S.; Kooblall, K.; Carnell, G.W.; O’Connor, D.; Bowden, T.A.; et al. A naturally protective epitope of limited variability as an influenza vaccine target. Nat. Commun. 2018, 9, 3859. [Google Scholar] [CrossRef] [PubMed]
- Atsmon, J.; Kate-Ilovitz, E.; Shaikevich, D.; Singer, Y.; Volokhov, I.; Haim, K.Y.; Ben-Yedidia, T. Safety and immunogenicity of multimeric-001—A novel universal influenza vaccine. J. Clin. Immunol. 2012, 32, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Fonteneau, J.-F.; Gilliet, M.; Larsson, M.; Dasilva, I.; Münz, C.; Liu, Y.-J.; Bhardwaj, N. Activation of influenza virus–specific CD4+ and CD8+ T cells: A new role for plasmacytoid dendritic cells in adaptive immunity. Blood 2003, 101, 3520–3526. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Motal, U.M.; Guay, H.M.; Wigglesworth, K.; Welsh, R.M.; Galili, U. Immunogenicity of influenza virus vaccine is increased by anti-gal-mediated targeting to antigen-presenting cells. J. Virol. 2007, 81, 9131–9141. [Google Scholar] [CrossRef] [PubMed]
- Grodeland, G.; Mjaaland, S.; Tunheim, G.; Fredriksen, A.B.; Bogen, B. The specificity of targeted vaccines for APC surface molecules influences the immune response phenotype. PLoS ONE 2013, 8, e80008. [Google Scholar] [CrossRef]
- Kanekiyo, M.; Wei, C.J.; Yassine, H.M.; McTamney, P.M.; Boyington, J.C.; Whittle, J.R.; Rao, S.S.; Kong, W.P.; Wang, L.; Nabel, G.J. Self-assembling influenza nanoparticle vaccines elicit broadly neutralizing H1N1 antibodies. Nature 2013, 499, 102–106. [Google Scholar] [CrossRef]
- Deng, L.; Cho, K.J.; Fiers, W.; Saelens, X. M2e-Based Universal Influenza A Vaccines. Vaccines 2015, 3, 105–136. [Google Scholar] [CrossRef]
- Tao, W.Q.; Gill, H.S. M2e-immobilized gold nanoparticles as influenza A vaccine: Role of soluble M2e and longevity of protection. Vaccine 2015, 33, 2307–2315. [Google Scholar] [CrossRef]
- Hiremath, J.; Kang, K.I.; Xia, M.; Elaish, M.; Binjawadagi, B.; Ouyang, K.; Dhakal, S.; Arcos, J.; Torrelles, J.B.; Jiang, X. Entrapment of H1N1 Influenza Virus Derived Conserved Peptides in PLGA Nanoparticles Enhances T Cell Response and Vaccine Efficacy in Pigs. PLoS ONE 2016, 11, e0151922. [Google Scholar] [CrossRef] [PubMed]
- Chahal, J.S.; Khan, O.F.; Cooper, C.L.; McPartlan, J.S.; Tsosie, J.K.; Tilley, L.D.; Sidik, S.M.; Lourido, S.; Langer, R.; Bavari, S.; et al. Dendrimer-RNA nanoparticles generate protective immunity against lethal Ebola, H1N1 influenza, and Toxoplasma gondii challenges with a single dose. Proc. Natl. Acad. Sci. USA 2016, 113, 4133–4142. [Google Scholar] [CrossRef]
- Deng, L.; Mohan, T.; Chang, T.Z.; Gonzalez, G.X.; Wang, Y.; Kwon, Y.M.; Kang, S.M.; Compans, R.W.; Champion, J.A.; Wang, B.Z. Double-layered protein nanoparticles induce broad protection against divergent influenza A viruses. Nat. Commun. 2018, 24, 359. [Google Scholar] [CrossRef] [PubMed]
- Harding, A.T.; Heaton, N.S. Efforts to Improve the Seasonal Influenza Vaccine. Vaccines 2018, 6, 19. [Google Scholar] [CrossRef] [PubMed]
- Draper, S.J.; Moore, A.C.; Goodman, A.L.; Long, C.A.; Holder, A.A.; Gilbert, S.C.; Hill, F.; Hill, A.V. Effective induction of high-titer antibodies by viral vector vaccines. Nat. Med. 2008, 14, 819–821. [Google Scholar] [CrossRef]
- Berthoud, T.K.; Hamill, M.; Lillie, P.J.; Hwenda, L.; Collins, K.A.; Ewer, K.J.; Milicic, A.; Poyntz, H.C.; Lambe, T.; Fletcher, H.A.; et al. Potent CD8+ T-cell immunogenicity in humans of a novel heterosubtypic influenza A vaccine, MVA-NP+M1. Clin. Infect. Dis. 2011, 52, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Antrobus, R.D.; Berthoud, T.K.; Mullarkey, C.E.; Hoschler, K.; Coughlan, L.; Zambon, M.; Hill, A.V.; Gilbert, S.C. Coadministration of seasonal influenza vaccine and MVA-NP+M1 simultaneously achieves potent humoral and cell-mediated responses. Mol. Ther. 2014, 22, 233–238. [Google Scholar] [CrossRef]
- Kim, E.H.; Han, G.Y.; Nguyen, H. An Adenovirus-Vectored Influenza Vaccine Induces Durable Cross-Protective Hemagglutinin Stalk Antibody Responses in Mice. Viruses 2017, 9, 234. [Google Scholar] [CrossRef]
- Lingel, A.; Bullard, B.L.; Weaver, E.A. Efficacy of an Adenoviral Vectored Multivalent Centralized Influenza Vaccine. Sci. Rep. 2017, 7, 14912. [Google Scholar] [CrossRef]
- Tripp, R.A.; Tompkins, S.M. Virus-vectored influenza virus vaccines. Viruses 2014, 6, 3055–3079. [Google Scholar] [CrossRef] [PubMed]
- Gubareva, L.V.; Kaiser, L.; Hayden, F.G. Influenza virus neuraminidase inhibitors. Lancet 2000, 355, 827–835. [Google Scholar] [CrossRef]
- Pinto, L.H.; Dieckmann, G.R.; Gandhi, C.S.; Papworth, C.G.; Braman, J.; Shaughnessy, M.A.; Lear, J.D.; Lamb, R.A.; DeGrado, W.F. A functionally defined model for the M2 proton channel of influenza A virus suggests a mechanism for its ion selectivity. Proc. Natl. Acad. Sci. USA 1997, 94, 11301–11306. [Google Scholar] [CrossRef] [PubMed]
- Pinto, L.H.; Holsinger, L.J.; Lamb, R.A. Influenza virus M2 protein has ion channel activity. Cell 1992, 69, 517–528. [Google Scholar] [CrossRef]
- Ma, C.; Polishchuk, A.L.; Ohigashi, Y.; Stouffer, A.L.; Schön, A.; Magavern, E.; Jing, X.; Lear, J.D.; Freire, E.; Lamb, R.A. Identification of the functional core of the influenza A virus A/M2 proton-selective ion channel. Proc. Natl. Acad. Sci. USA 2009, 106, 12283–12288. [Google Scholar] [CrossRef] [PubMed]
- Van Voris, L.P.; Betts, R.F.; Hayden, F.G.; Christmas, W.A.; Douglas, R.G., Jr. Successful treatment of naturally occurring influenza A/USSR/77 H1N1. JAMA 1981, 245, 1128–1231. [Google Scholar] [CrossRef] [PubMed]
- Bean, W.J.; Threlkeld, S.C.; Webster, R.G. Biologic potential of amantadine-resistant influenza A virus in an avian model. J. Infect. Dis. 1989, 159, 1050–1056. [Google Scholar] [CrossRef] [PubMed]
- Deyde, V.M.; Xu, X.Y.; Bright, R.A.; Shaw, M.; Smith, C.B.; Zhang, Y.; Shu, Y.; Gubareva, L.V.; Cox, N.J.; Klimov, A.I. Surveillance of resistance to adamantanes among influenza A(H3N2) and A(H1N1) viruses isolated worldwide. J. Infect. Dis. 2007, 196, 249–257. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Galvin, H.D.; Haw, T.Y.; Nutsford, A.N.; Husain, M. Drug resistance in influenza A virus: The epidemiology and management. Infect. Drug Resist. 2017, 10, 121–134. [Google Scholar] [CrossRef]
- McKimm-Breschkin, J.L. Neuraminidase inhibitors for the treatment and prevention of influenza. Expert Opin. Pharmacother. 2002, 3, 103–112. [Google Scholar] [CrossRef]
- Rameix-Welti, M.A.; Enouf, V.; Cuvelier, F.; Jeannin, P.; van der Werf, S. Enzymatic properties of the neuraminidase of seasonal H1N1 influenza viruses provide insights for the emergence of natural resistance to oseltamivir. PLoS Pathog 2008, 4, e1000103. [Google Scholar] [CrossRef] [PubMed]
- Aoki, F.Y.; Boivin, G. Influenza virus shedding: Excretion patterns and effects of antiviral treatment. J. Clin. Virol. 2009, 44, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Samson, M.; Pizzorno, A.; Abed, Y.; Boivin, G. Influenza virus resistance to neuraminidase inhibitors. Antivir. Res. 2013, 98, 174–185. [Google Scholar] [CrossRef] [PubMed]
- Noah, D.L.; Noah, J.W. Adapting global influenza management strategies to address emerging viruses. Am. J. Physiol.-Lung Cell. Mol. Physiol. 2013, 305, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.T.; Nguyen, T.; Mishin, V.P.; Sleeman, K.; Balish, A.; Jones, J.; Creanga, A.; Marjuki, H.; Uyeki, T.M.; Nguyen, D.H.; et al. Antiviral susceptibility of highly pathogenic avian influenza A(H5N1) viruses isolated from poultry, Vietnam, 2009–2011. Emerg. Infect. Dis. 2013, 19, 1963–1971. [Google Scholar] [CrossRef] [PubMed]
- Malakhov, M.P.; Aschenbrenner, L.M.; Smee, D.F.; Wandersee, M.K.; Sidwell, R.W.; Gubareva, L.V.; Mishin, V.P.; Hayden, F.G.; Kim, D.H.; Ing, A.; et al. Sialidase fusion protein as a novel broad-spectrum inhibitor of influenza virus infection. Antimicrob. Agents Chemother. 2006, 50, 1470–1479. [Google Scholar] [CrossRef] [PubMed]
- Baum, L.G.; Paulson, J.C. Sialyloligosaccharides of the respiratory epithelium in the selection of human influenza virus receptor specificity. Acta Histochem. Suppl. 1990, 40, 35–38. [Google Scholar] [PubMed]
- Belser, J.A.; Lu, X.; Szretter, K.J.; Jin, X.; Aschenbrenner, L.M.; Lee, A.; Hawley, S.; Kim, D.H.; Malakhov, M.P.; Yu, M.; et al. DAS181, a novel sialidase fusion protein, protects mice from lethal avian influenza H5N1 virus infection. J. Infect. Dis. 2007, 196, 1493–1499. [Google Scholar] [CrossRef]
- Chan, R.W.; Chan, M.C.; Wong, A.C.; Karamanska, R.; Dell, A.; Haslam, S.M.; Sihoe, A.D.; Chui, W.H.; Triana-Baltzer, G.; Li, Q.; et al. DAS181 inhibits H5N1 influenza virus infection of human lung tissues. Antimicrob. Agents Chemother. 2009, 53, 3935–3941. [Google Scholar] [CrossRef]
- Triana-Baltzer, G.B.; Babizki, M.; Chan, M.C.; Wong, A.C.; Aschenbrenner, L.M.; Campbell, E.R.; Li, Q.X.; Chan, R.W.; Peiris, J.S.; Nicholls, J.M.; et al. DAS181, a sialidase fusion protein, protects human airway epithelium against influenza virus infection: An in vitro pharmacodynamic analysis. J. Antimicrob. Chemother. 2010, 65, 275–284. [Google Scholar] [CrossRef]
- Thammawat, S.; Sadlon, T.A.; Adamson, P.; Gordon, D.L. Effect of sialidase fusion protein (DAS 181) on human metapneumovirus infection of Hep-2 cells. Antivir. Chem. Chemother. 2015, 24, 161–165. [Google Scholar] [CrossRef] [PubMed]
- Rossignol, J.F. Nitazoxanide: A first-in-class broad-spectrum antiviral agent. Antiviral Res. 2014, 110, 94–103. [Google Scholar] [CrossRef] [PubMed]
- Rossignol, J.F.; La Frazia, S.; Chiappa, L.; Ciucci, A.; Santoro, M.G. Thiazolides, a new class of anti-influenza molecules targeting viral hemagglutinin at the post-translational level. J. Biol. Chem. 2009, 284, 29798–29808. [Google Scholar] [CrossRef] [PubMed]
- Byrn, R.A.; Jones, S.M.; Bennett, H.B.; Bral, C.; Clark, M.P.; Jacobs, M.D.; Kwong, A.D.; Ledeboer, M.W.; Leeman, J.R.; McNeil, C.F.; et al. Preclinical activity of VX-787, a first-in-class, orally bioavailable inhibitor of the influenza virus polymerase PB2 subunit. Antimicrob. Agents Chemother. 2015, 59, 1569–1582. [Google Scholar] [CrossRef] [PubMed]
- Smee, D.F.; Barnard, D.L.; Jones, S.M. Activities of JNJ63623872 and oseltamivir against influenza A H1N1pdm and H3N2 virus infections in mice. Antivir. Res. 2016, 136, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Gaelings, L.; Soderholm, S.; Belanov, S.; Nandania, J.; Nyman, T.A.; Matikainen, S.; Anders, S.; Velagapudi, V.; Kainov, D.E. JNJ872 inhibits influenza A virus replication without altering cellular antiviral responses. Antivir. Res. 2016, 133, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Finberg, R.W.; Lanno, R.; Anderson, D.; Fleischhackl, R.; van Duijnhoven, W.; Kauffman, R.S.; Kosoglou, T.; Vingerhoets, J.; Leopold, L. Phase 2b Study of Pimodivir (JNJ-63623872) as Monotherapy or in Combination With Oseltamivir for Treatment of Acute Uncomplicated Seasonal Influenza A: TOPAZ Trial. J. Infect. Dis. 2019, 219, 1026–1034. [Google Scholar] [CrossRef] [PubMed]
- Furuta, Y.; Takahashi, K.; Shiraki, K.; Sakamoto, K.; Smee, D.F.; Barnard, D.L.; Gowen, B.B.; Julander, J.G.; Morrey, J.D. T-705 (favipiravir) and related compounds: Novel broad-spectrum inhibitors of RNA viral infections. Antivir. Res. 2009, 82, 95–102. [Google Scholar] [CrossRef]
- Furuta, Y.; Takahashi, K.; Kuno-Maekawa, M.; Sangawa, H.; Uehara, S.; Kozaki, K.; Nomura, N.; Egawa, H.; Shiraki, K. Mechanism of action of T-705 against influenza virus. Antimicrob. Agents Chemother. 2005, 49, 981–986. [Google Scholar] [CrossRef]
- Baranovich, T.; Wong, S.S.; Armstrong, J.; Marjuki, H.; Webby, R.J.; Webster, R.G.; Govorkova, E.A. T-705 (favipiravir) induces lethal mutagenesis in influenza A H1N1 viruses in vitro. J. Virol. 2013, 87, 3741–3751. [Google Scholar] [CrossRef]
- Jin, Z.; Smith, L.K.; Rajwanshi, V.K.; Kim, B.; Deval, J. The ambiguous base-pairing and high substrate efficiency of T-705 (Favipiravir) Ribofuranosyl 5′-triphosphate towards influenza A virus polymerase. PLoS ONE 2013, 8, e68347. [Google Scholar] [CrossRef] [PubMed]
- Naesens, L.; Guddat, L.W.; Keough, D.T.; van Kuilenburg, A.B.; Meijer, J.; Vande Voorde, J.; Balzarini, J. Role of human hypoxanthine guanine phosphoribosyltransferase in activation of the antiviral agent T-705 (favipiravir). Mol. Pharmacol. 2013, 84, 615–629. [Google Scholar] [CrossRef] [PubMed]
- Safronetz, D.; Falzarano, D.; Scott, D.P.; Furuta, Y.; Feldmann, H.; Gowen, B.B. Antiviral efficacy of favipiravir against two prominent etiological agents of hantavirus pulmonary syndrome. Antimicrob. Agents Chemother. 2013, 57, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Gowen, B.B.; Wong, M.H.; Jung, K.H.; Sanders, A.B.; Mendenhall, M.; Bailey, K.W.; Furuta, Y.; Sidwell, R.W. In vitro and in vivo activities of T-705 against arenavirus and bunyavirus infections. Antimicrob. Agents Chemother. 2007, 51, 3168–3176. [Google Scholar] [CrossRef] [PubMed]
- Morrey, J.D.; Taro, B.S.; Siddharthan, V.; Wang, H.; Smee, D.F.; Christensen, A.J.; Furuta, Y. Efficacy of orally administered T-705 pyrazine analog on lethal West Nile virus infection in rodents. Antivir. Res. 2008, 80, 377–379. [Google Scholar] [CrossRef] [PubMed]
- Rocha-Pereira, J.; Jochmans, D.; Dallmeier, K.; Leyssen, P.; Nascimento, M.S.; Neyts, J. Favipiravir (T-705) inhibits in vitro norovirus replication. Biochem. Biophys. Res. Commun. 2012, 424, 777–780. [Google Scholar] [CrossRef] [PubMed]
- Bai, C.Q.; Mu, J.S.; Kargbo, D.; Song, Y.B.; Niu, W.K.; Nie, W.M.; Kanu, A.; Liu, W.W.; Wang, Y.P.; Dafae, F.; et al. Clinical and Virological Characteristics of Ebola Virus Disease Patients Treated With Favipiravir (T-705)-Sierra Leone, 2014. Clin. Infect. Dis. 2016, 63, 1288–1294. [Google Scholar] [CrossRef] [PubMed]
- Goldhill, D.H.; Te Velthuis, A.J.W.; Fletcher, R.A.; Langat, P.; Zambon, M.; Lackenby, A.; Barclay, W.S. The mechanism of resistance to favipiravir in influenza. Proc. Natl. Acad. Sci. USA 2018, 115, 11613–11618. [Google Scholar] [CrossRef] [PubMed]
- Furuta, Y.; Gowen, B.B.; Takahashi, K.; Shiraki, K.; Smee, D.F.; Barnard, D.L. Favipiravir (T-705), a novel viral RNA polymerase inhibitor. Antivir. Res. 2013, 100, 446–454. [Google Scholar] [CrossRef] [PubMed]
- Koshimichi, H.; Ishibashi, T.; Kawaguchi, N.; Sato, C.; Kawasaki, A.; Wajima, T. Safety, Tolerability, and Pharmacokinetics of the Novel Anti-influenza Agent Baloxavir Marboxil in Healthy Adults: Phase I Study Findings. Clin. Drug Investig. 2018, 38, 1189–1196. [Google Scholar] [CrossRef] [PubMed]
- Hayden, F.G.; Sugaya, N.; Hirotsu, N.; Lee, N.; de Jong, M.D.; Hurt, A.C.; Ishida, T.; Sekino, H.; Yamada, K.; Portsmouth, S.; et al. Baloxavir Marboxil for Uncomplicated Influenza in Adults and Adolescents. N. Engl. J. Med. 2018, 379, 913–923. [Google Scholar] [CrossRef] [PubMed]
- O’Hanlon, R.; Shaw, M.L. Baloxavir marboxil: The new influenza drug on the market. Curr. Opin. Virol. 2019, 35, 14–18. [Google Scholar] [CrossRef] [PubMed]
- Gagarinova, V.M.; Ignat’eva, G.S.; Sinitskaia, L.V.; Ivanova, A.M.; Rodina, M.A.; Tur’eva, A.V. The new chemical preparation arbidol: Its prophylactic efficacy during influenza epidemics. Zh Mikrobiol. Epidemiol. Immunobiol. 1993, 5, 40–43. [Google Scholar]
- Boriskin, Y.S.; Leneva, I.A.; Pecheur, E.I.; Polyak, S.J. Arbidol: A broad-spectrum antiviral compound that blocks viral fusion. Curr. Med. Chem. 2008, 15, 997–1005. [Google Scholar] [CrossRef] [PubMed]
- Brooks, M.J.; Burtseva, E.I.; Ellery, P.J.; Marsh, G.A.; Lew, A.M.; Slepushkin, A.N.; Crowe, S.M.; Tannock, G.A. Antiviral activity of arbidol, a broad-spectrum drug for use against respiratory viruses, varies according to test conditions. J. Med. Virol. 2012, 84, 170–181. [Google Scholar] [CrossRef] [PubMed]
- Blaising, J.; Polyak, S.J.; Pecheur, E.I. Arbidol as a broad-spectrum antiviral: An update. Antivir. Res. 2014, 107, 84–94. [Google Scholar] [CrossRef] [PubMed]
- Pecheur, E.I.; Polyak, S.J. The synthetic antiviral drug arbidol inhibits globally prevalent pathogenic viruses. M S-Med. Sci. 2016, 32, 1056–1059. [Google Scholar] [CrossRef] [PubMed]
- Haviernik, J.; Stefanik, M.; Fojtikova, M.; Kali, S.; Tordo, N.; Rudolf, I.; Hubálek, Z.; Eyer, L.; Ruzek, D. Arbidol (Umifenovir): A Broad-Spectrum Antiviral Drug That Inhibits Medically Important Arthropod-Borne Flaviviruses. Viruses 2018, 10, 184. [Google Scholar] [CrossRef]
- Loginova, S.; Borisevich, S.V.; Maksimov, V.A.; Bondarev, V.P.; Nebol’sin, V.E. Therapeutic efficacy of Ingavirin, a new domestic formulation against influenza A virus (H3N2). Antibiot Khimioter 2008, 53, 27–30. [Google Scholar]
- Galegov, G.A.; Andronova, V.L.; Nebol’sin, V.E. Antiviral effect of Ingavirin against seasonal influenza virus A/H1N1 in MDCK cell culture. Antibiot Khimioter 2009, 54, 19–22. [Google Scholar]
- Zarubaev, V.V.; Beliaevskaia, S.V.; Sirotkin, A.K.; Anfimov, P.M.; Nebol’sin, V.E.; Kiselev, O.I.; Reĭkhart, D.V. In vitro and in vivo effects of ingavirin on the ultrastructure and infectivity of influenza virus. Vopr. Virusol. 2011, 56, 21–25. [Google Scholar] [PubMed]
- Simoes, E.A.; Groothuis, J.R.; Carbonell-Estrany, X.; Rieger, C.H.; Mitchell, I.; Fredrick, L.M.; Kimpen, J.L.; Palivizumab Long-Term Respiratory Outcomes Study Group. Palivizumab prophylaxis, respiratory syncytial virus, and subsequent recurrent wheezing. J. Pediatr. 2007, 151, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, J.M.; Kuritzkes, D.R.; Godofsky, E.; DeJesus, E.; Larson, J.A.; Weinheimer, S.P.; Lewis, S.T. Safety, pharmacokinetics, and antiretroviral activity of multiple doses of ibalizumab (formerly TNX-355), an anti-CD4 monoclonal antibody, in human immunodeficiency virus type 1-infected adults. Antimicrob. Agents Chemother. 2009, 53, 450–457. [Google Scholar] [CrossRef] [PubMed]
- Okuno, Y.; Isegawa, Y.; Sasao, F.; Ueda, S. A Common Neutralizing Epitope Conserved between the Hemagglutinins of Influenza—A Virus H1 and H2 Strains. J. Virol. 1993, 67, 2552–2558. [Google Scholar] [PubMed]
- Dreyfus, C.; Ekiert, D.C.; Wilson, I.A. Structure of a classical broadly neutralizing stem antibody in complex with a pandemic H2 influenza virus hemagglutinin. J. Virol. 2013, 87, 7149–7154. [Google Scholar] [CrossRef] [PubMed]
- Throsby, M.; van den Brink, E.; Jongeneelen, M.; Poon, L.L.M.; Alard, P.; Cornelissen, L.; Bakker, A.; Cox, F.; van Deventer, E.; Guan, Y.; et al. Heterosubtypic Neutralizing Monoclonal Antibodies Cross-Protective against H5N1 and H1N1 Recovered from Human IgM(+) Memory B Cells. PLoS ONE 2008, 3, e3942. [Google Scholar] [CrossRef] [PubMed]
- Kallewaard, N.L.; Corti, D.; Collins, P.J.; Neu, U.; McAuliffe, J.M.; Benjamin, E.; Wachter-Rosati, L.; Palmer-Hill, F.J.; Yuan, A.Q.; Walker, P.A.; et al. Structure and Function Analysis of an Antibody Recognizing All Influenza A Subtypes. Cell 2016, 166, 596–608. [Google Scholar] [CrossRef]
- Ali, S.O.; Takas, T.; Nyborg, A.; Shoemaker, K.; Kallewaard, N.L.; Chiong, R.; Dubovsky, F.; Mallory, R.M. Evaluation of MEDI8852, an Anti-Influenza A Monoclonal Antibody, in Treating Acute Uncomplicated Influenza. Antimicrob. Agents Chemother. 2018, 62, e00694-18. [Google Scholar] [CrossRef]
- Pappas, L.; Foglierini, M.; Piccoli, L.; Kallewaard, N.L.; Turrini, F.; Silacci, C.; Fernandez-Roadriguez, B.; Agatic, G.; Giacchetto-Sasselli, I.; Pellicciotta, G.; et al. Rapid development of broadly influenza neutralizing antibodies through redundant mutations. Nature 2014, 516, 418–422. [Google Scholar] [CrossRef]
- Lu, J.; Guo, Z.; Pan, X.; Wang, G.; Zhang, D.; Li, Y.; Tan, B.; Ouyang, L.; Yu, X. Passive immunotherapy for influenza A H5N1 virus infection with equine hyperimmune globulin F (ab’) 2 in mice. Respir. Res. 2006, 7, 43. [Google Scholar] [CrossRef][Green Version]
- Sui, J.; Sheehan, J.; Hwang, W.C.; Bankston, L.A.; Burchett, S.K.; Huang, C.-Y.; Liddington, R.C.; Beigel, J.H.; Marasco, W.A. Wide prevalence of heterosubtypic broadly neutralizing human anti–influenza A antibodies. Clin. Infect. Dis. 2011, 52, 1003–1009. [Google Scholar] [CrossRef] [PubMed]
- Corti, D.; Suguitan, A.L.; Pinna, D.; Silacci, C.; Fernandez-Rodriguez, B.M.; Vanzetta, F.; Santos, C.; Luke, C.J.; Torres-Velez, F.J.; Temperton, N.J.; et al. Heterosubtypic neutralizing antibodies are produced by individuals immunized with a seasonal influenza vaccine. J. Clin. Investig. 2010, 120, 1663–1673. [Google Scholar] [CrossRef] [PubMed]
- Margine, I.; Krammer, F.; Hai, R.; Heaton, N.; Tan, G.; Andrews, S.; Runstadler, J.A.; Wilson, P.C.; Albrecht, R.A.; Garcia-Sastre, A.; et al. Hemagglutinin stalk-based universal vaccine constructs protect against group 2 influenza A viruses. J. Virol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Nachbagauer, R.; Miller, M.S.; Hai, R.; Ryder, A.B.; Rose, J.K.; Palese, P.; García-Sastre, A.; Krammer, F.; Albrecht, R.A. Hemagglutinin stalk immunity reduces influenza virus replication and transmission in ferrets. J. Virol. 2016, 90, 3268–3273. [Google Scholar] [CrossRef]
- Wohlbold, T.J.; Nachbagauer, R.; Margine, I.; Tan, G.S.; Hirsh, A.; Krammer, F. Vaccination with soluble headless hemagglutinin protects mice from challenge with divergent influenza viruses. Vaccine 2015, 33, 3314–3321. [Google Scholar] [CrossRef]
- El Bakkouri, K.; Descamps, F.; De Filette, M.; Smet, A.; Festjens, E.; Birkett, A.; Van Rooijen, N.; Verbeek, S.; Fiers, W.; Saelens, X. Universal vaccine based on ectodomain of matrix protein 2 of influenza A: Fc receptors and alveolar macrophages mediate protection. J. Immunol. 2011, 186, 1022–1031. [Google Scholar] [CrossRef] [PubMed]
- Wilson, J.R.; Belser, J.A.; DaSilva, J.; Guo, Z.; Sun, X.; Gansebom, S.; Bai, Y.; Stark, T.J.; Chang, J.; Carney, P.; et al. An influenza A virus (H7N9) anti-neuraminidase monoclonal antibody protects mice from morbidity without interfering with the development of protective immunity to subsequent homologous challenge. Virology 2017, 511, 214–221. [Google Scholar] [CrossRef]
- Zhao, J.; Perera, R.A.; Kayali, G.; Meyerholz, D.; Perlman, S.; Peiris, M. Passive immunotherapy with dromedary immune serum in an experimental animal model for Middle East respiratory syndrome coronavirus infection. J. Virol. 2015, 89, 6117–6120. [Google Scholar] [CrossRef]
- Audet, J.; Wong, G.; Wang, H.; Lu, G.; Gao, G.F.; Kobinger, G.; Qiu, X. Molecular characterization of the monoclonal antibodies composing ZMAb: A protective cocktail against Ebola virus. Sci. Rep. 2014, 4, 6881. [Google Scholar] [CrossRef]
- Qiu, X.; Wong, G.; Audet, J.; Bello, A.; Fernando, L.; Alimonti, J.B.; Fausther-Bovendo, H.; Wei, H.; Aviles, J.; Hiatt, E.; et al. Reversion of advanced Ebola virus disease in nonhuman primates with ZMapp. Nature 2014, 514, 47–53. [Google Scholar] [CrossRef]
- Balazs, A.B.; Bloom, J.D.; Hong, C.M.; Rao, D.S.; Baltimore, D. Broad protection against influenza infection by vectored immunoprophylaxis in mice. Nat. Biotechnol. 2013, 31, 647–652. [Google Scholar] [CrossRef] [PubMed]
- Berry, C.M.; Penhale, W.J.; Sangster, M.Y. Passive broad-spectrum influenza immunoprophylaxis. Influenza. Res. Treat. 2014, 2014, 267594. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Zhong, N.; Guan, Y. Treatment with convalescent plasma for influenza A (H5N1) infection. N. Engl. J. Med. 2007, 357, 1450–1451. [Google Scholar] [CrossRef] [PubMed]
- Jegerlehner, A.; Schmitz, N.; Storni, T.; Bachmann, M.F. Influenza A vaccine based on the extracellular domain of M2: Weak protection mediated via antibody-dependent NK cell activity. J. Immunol. 2004, 172, 5598–5605. [Google Scholar] [CrossRef] [PubMed]
- Fujisawa, H. Neutrophils play an essential role in cooperation with antibody in both protection against and recovery from pulmonary infection with influenza virus in mice. J. Virol. 2008, 82, 2772–2783. [Google Scholar] [CrossRef]
- Terajima, M.; Co, M.D.T.; Cruz, J.; Ennis, F.A. High Antibody-Dependent Cellular Cytotoxicity Antibody Titers to H5N1 and H7N9 Avian Influenza A Viruses in Healthy US Adults and Older Children. J. Infect. Dis. 2015, 212, 1052–1060. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Tsibane, T.; McGraw, P.A.; House, F.S.; Keefer, C.J.; Hicar, M.D.; Tumpey, T.M.; Pappas, C.; Perrone, L.A.; Martinez, O.; et al. Neutralizing antibodies derived from the B cells of 1918 influenza pandemic survivors. Nature 2008, 455, 532. [Google Scholar] [CrossRef]
- Krammer, F.; García-Sastre, A.; Palese, P. Is it possible to develop a “universal” influenza virus vaccine? Toward a universal influenza virus vaccine: Potential target antigens and critical aspects for vaccine development. Cold Spring Harb. Perspect. Biol. 2017, 10, a028845. [Google Scholar] [CrossRef]
| Vaccines | Design | References |
|---|---|---|
| Virus-like particle vaccines (VLP) | Self-assembling viral matrices that express a single or multivalent viral surface proteins | [36,37,38,39,40,41,42] |
| Computationally optimized broadly reactive antigen vaccines (COBRA) | VLPs bearing computationally optimized viral surface proteins | [43,44,45,46,47,48,49] |
| Synthetic virus | Generation of replication-incompetent viruses bearing genetically attenuated genomic sequences | [18,49,50,51,52,53,54] |
| Epitope | Epitope-rich proteins of viruses, designed to induce protective epitope targeted antibodies | [55,56,57,58,59] |
| antigen-presenting cell (APC) inducible | APC-targeted delivery of immunogenic viral proteins to induce quicker and T cell responses | [60,61,62] |
| Nanoparticle-based | Self-assembling nano-molecules that carry a single or multivalent viral surface protein | [63,64,65,66,67,68,69] |
| Viral-vectored | Mainly involves use of dissimilar viral matrices as carriers of specific viral protein | [70,71,72,73,74,75] |
| Antiviral | Mechanism of Action | Clinical Phase and Status | Country of Development/Trial |
|---|---|---|---|
| Das181 (Fludase) | Sialic acid removal in the respiratory airways | II (IFV), III (PIV) not yet recruiting | USA |
| Nitazoxanide | HA maturation inhibition | III completed | USA |
| JNJ-63623872 (Pimodivir) | Small molecule inhibitor of influenza A virus PB2 | III recruiting | Belgium |
| T705 (Favipiravir) | RNA-dependent RNA polymerase inhibitor | IV | Japan |
| Baloxavir marboxil | Small molecule inhibitor of cap-dependent endonuclease (PA) | III recruiting children <1 year 1 | Japan |
| Arbidol (Umifenovir) | HA resistance to conformational changes triggered by pH | III recruiting in China/IV unknown status in Russia | China; Russia |
| Ingavirin | Interaction with NP and inhibition of viral genome release | IV completed | Russia |
| Antibody | Target/Mechanism of Action | Clinical Phase Status | Country |
|---|---|---|---|
| MEDI8852 | HA stem | IIa completed | USA |
| MHAA4549A | HA stem | II completed | USA |
| VIS410 | HA stem | II completed | USA |
| Intravenous hyper-immune immunoglobulin (IVIG) | Antigen specific antibody pool with neutralizing potential | III recruiting | USA |
| Ergoferon | Suppression of non-specific immune activation (by any virus) | IV completed | Russia |
| CR6261 | “highly conserved membrane-proximal stem of H1 and H5 viruses’ HA1 and HA2” | II completed | USA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kotey, E.; Lukosaityte, D.; Quaye, O.; Ampofo, W.; Awandare, G.; Iqbal, M. Current and Novel Approaches in Influenza Management. Vaccines 2019, 7, 53. https://doi.org/10.3390/vaccines7020053
Kotey E, Lukosaityte D, Quaye O, Ampofo W, Awandare G, Iqbal M. Current and Novel Approaches in Influenza Management. Vaccines. 2019; 7(2):53. https://doi.org/10.3390/vaccines7020053
Chicago/Turabian StyleKotey, Erasmus, Deimante Lukosaityte, Osbourne Quaye, William Ampofo, Gordon Awandare, and Munir Iqbal. 2019. "Current and Novel Approaches in Influenza Management" Vaccines 7, no. 2: 53. https://doi.org/10.3390/vaccines7020053
APA StyleKotey, E., Lukosaityte, D., Quaye, O., Ampofo, W., Awandare, G., & Iqbal, M. (2019). Current and Novel Approaches in Influenza Management. Vaccines, 7(2), 53. https://doi.org/10.3390/vaccines7020053






