Hydrogen Peroxide and Redox Regulation of Developments
Abstract
1. Introduction
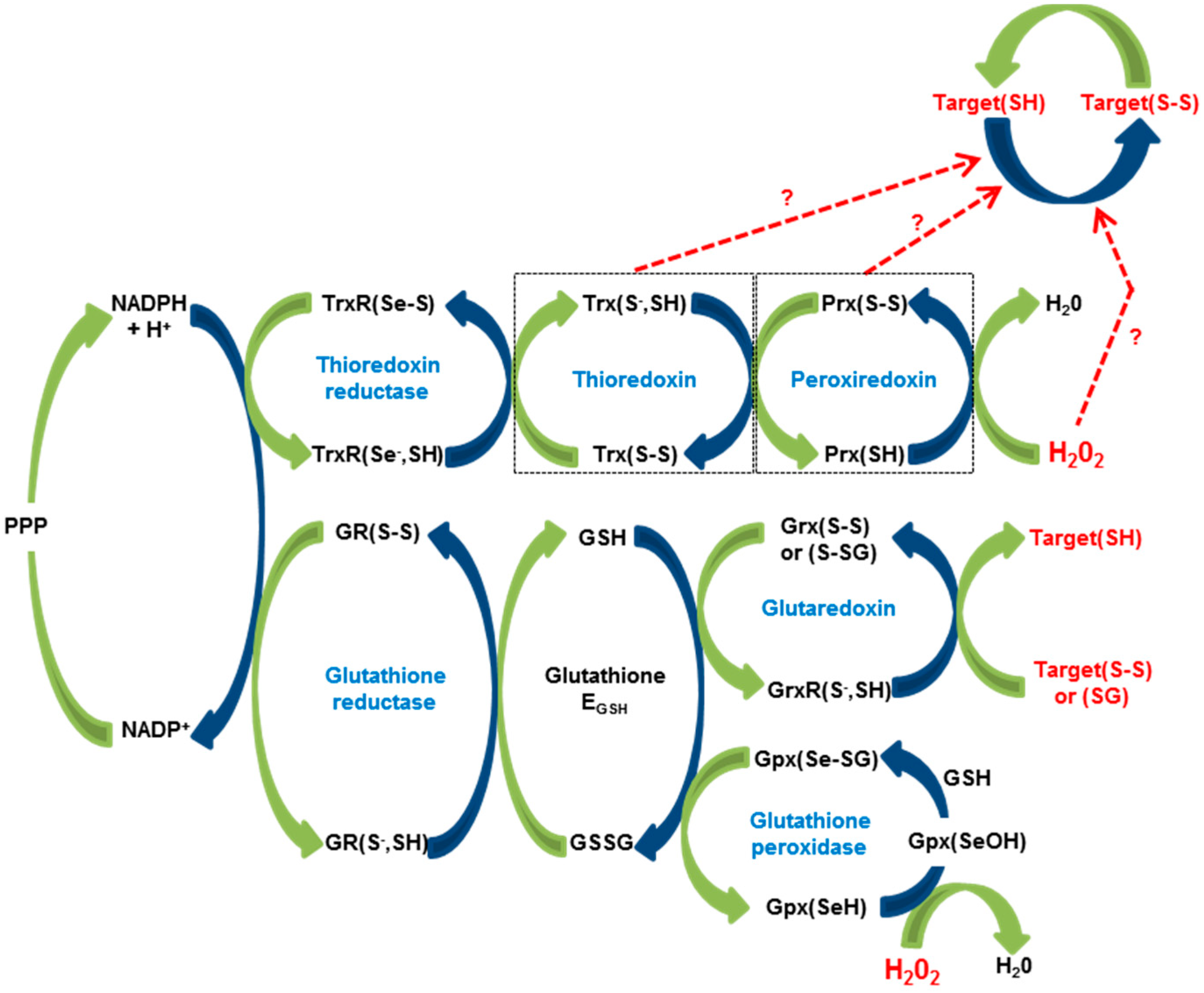
2. The Redox Machine
3. Seeing Is Believing
3.1. H2O2 Sensors
3.2. Glutathione Redox Potential Sensors
3.3. NADPH Sensor: iNap
4. Redox Signalling in Animal Development and Regeneration
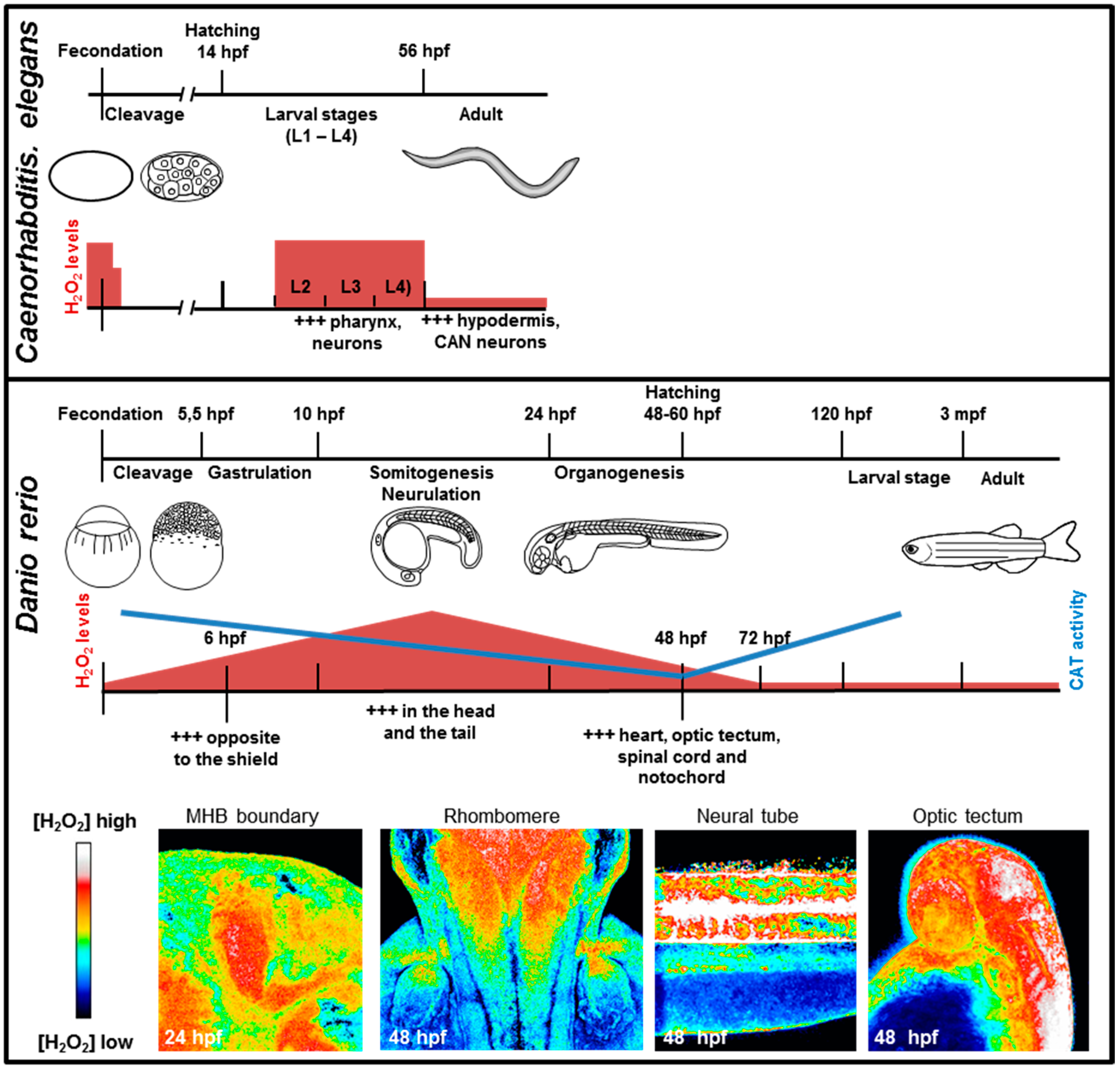
4.1. Embryonic Development
4.1.1. NADPH Oxidase Complexes in Embryonic Development
4.1.2. Catalase, Superoxide Dismutases and Glutathione Systems in Embryonic Development
4.1.3. Thioredoxin System in Embryonic Development
4.1.4. Peroxiredoxin Systems in Embryonic Development
4.2. Adult Stem Cells and Tissue Homeostasis
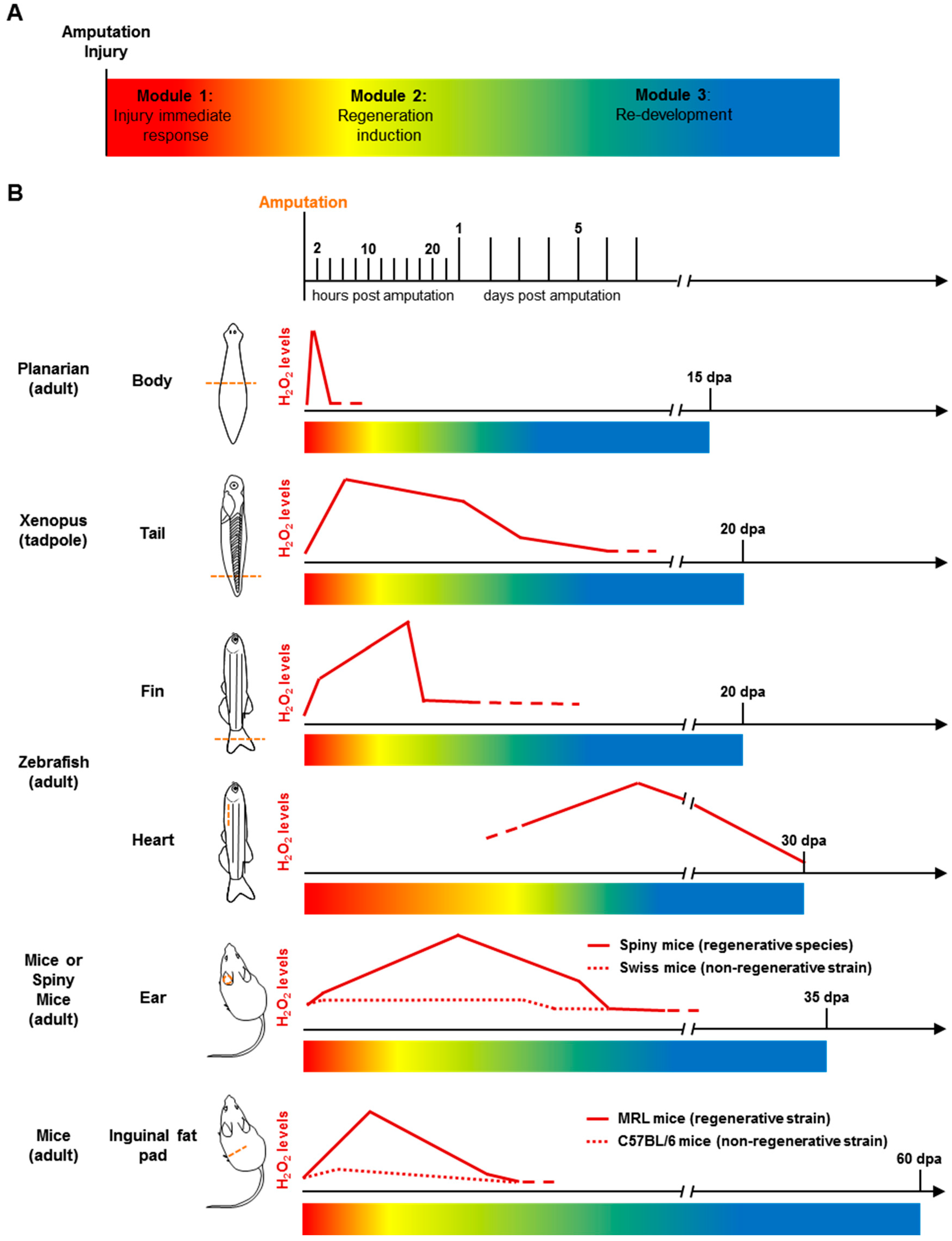
4.3. Regeneration
5. Conclusions: Towards the Redox Code
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Manda-Handzlik, A.; Demkow, U. Neutrophils: The Role of Oxidative and Nitrosative Stress in Health and Disease. Adv. Exp. Med. Biol. 2015, 857, 51–60. [Google Scholar] [PubMed]
- Liou, G.Y.; Storz, P. Reactive oxygen species in cancer. Free Radic. Res 2010, 44, 479–496. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, W.; Jiang, Z.G.; Ghanbari, H.A. Oxidative stress and neurodegenerative disorders. Int. J. Mol. Sci. 2013, 14, 24438–24475. [Google Scholar] [CrossRef] [PubMed]
- Chaturvedi, R.K.; Flint Beal, M. Mitochondrial diseases of the brain. Free Radic. Biol. Med. 2013, 63, 1–29. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.J.; Forman, H.J.; Sevanian, A. Oxidants as stimulators of signal transduction. Free Radic. Biol. Med. 1997, 22, 269–285. [Google Scholar] [CrossRef]
- D’Autreaux, B.; Toledano, M.B. ROS as signalling molecules: Mechanisms that generate specificity in ROS homeostasis. Nat. Rev. Mol. Cell Biol. 2007, 8, 813–824. [Google Scholar] [CrossRef] [PubMed]
- Forman, H.J.; Maiorino, M.; Ursini, F. Signaling functions of reactive oxygen species. Biochemistry 2010, 49, 835–842. [Google Scholar] [CrossRef] [PubMed]
- Forman, H.J.; Ursini, F.; Maiorino, M. An overview of mechanisms of redox signaling. J. Mol. Cell Cardiol. 2014, 73, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Covarrubias, L.; Hernandez-Garcia, D.; Schnabel, D.; Salas-Vidal, E.; Castro-Obregon, S. Function of reactive oxygen species during animal development: Passive or active? Dev. Biol. 2008, 320, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Garcia, D.; Wood, C.D.; Castro-Obregon, S.; Covarrubias, L. Reactive oxygen species: A radical role in development? Free Radic. Biol. Med. 2010, 49, 130–143. [Google Scholar] [CrossRef] [PubMed]
- Sies, H. Hydrogen peroxide as a central redox signaling molecule in physiological oxidative stress: Oxidative eustress. Redox Biol. 2017, 11, 613–619. [Google Scholar] [CrossRef] [PubMed]
- Nikolaidis, M.G.; Margaritelis, N.V. Same Redox Evidence But Different Physiological “Stories”: The Rashomon Effect in Biology. Bioessays 2018, 40, e1800041. [Google Scholar] [CrossRef] [PubMed]
- Di Meo, S.; Reed, T.T.; Venditti, P.; Victor, V.M. Role of ROS and RNS Sources in Physiological and Pathological Conditions. Oxid Med. Cell Longev. 2016, 2016, 1245049. [Google Scholar] [CrossRef] [PubMed]
- Tyurina, Y.Y.; Shrivastava, I.; Tyurin, V.A.; Mao, G.; Dar, H.H.; Watkins, S.; Epperly, M.; Bahar, I.; Shvedova, A.A.; Pitt, B.; et al. Only a Life Lived for Others Is Worth Living: Redox Signaling by Oxygenated Phospholipids in Cell Fate Decisions. Antioxid. Redox Signal. 2018, 29, 1333–1358. [Google Scholar] [CrossRef] [PubMed]
- Paulsen, C.E.; Carroll, K.S. Orchestrating redox signaling networks through regulatory cysteine switches. ACS Chem. Biol. 2010, 5, 47–62. [Google Scholar] [CrossRef] [PubMed]
- Marinho, H.S.; Real, C.; Cyrne, L.; Soares, H.; Antunes, F. Hydrogen peroxide sensing, signaling and regulation of transcription factors. Redox Biol. 2014, 2, 535–562. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.R.; Yang, S. Hydrogen peroxide: A signaling messenger. Antioxid. Redox Signal. 2006, 8, 243–270. [Google Scholar] [CrossRef] [PubMed]
- Holmstrom, K.M.; Finkel, T. Cellular mechanisms and physiological consequences of redox-dependent signalling. Nat. Rev. Mol. Cell Biol. 2014, 15, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Bedard, K.; Krause, K.H. The NOX family of ROS-generating NADPH oxidases: Physiology and pathophysiology. Physiol. Rev. 2007, 87, 245–313. [Google Scholar] [CrossRef] [PubMed]
- Brandes, R.P.; Weissmann, N.; Schroder, K. Nox family NADPH oxidases: Molecular mechanisms of activation. Free Radic. Biol. Med. 2014, 76, 208–226. [Google Scholar] [CrossRef] [PubMed]
- Miller, E.W.; Dickinson, B.C.; Chang, C.J. Aquaporin-3 mediates hydrogen peroxide uptake to regulate downstream intracellular signaling. Proc. Natl. Acad. Sci. USA 2010, 107, 15681–15686. [Google Scholar] [CrossRef] [PubMed]
- Bienert, G.P.; Chaumont, F. Aquaporin-facilitated transmembrane diffusion of hydrogen peroxide. Biochim. Biophys. Acta 2014, 1840, 1596–1604. [Google Scholar] [CrossRef] [PubMed]
- Bertolotti, M.; Farinelli, G.; Galli, M.; Aiuti, A.; Sitia, R. AQP8 transports NOX2-generated H2O2 across the plasma membrane to promote signaling in B cells. J. Leukoc. Biol. 2016, 100, 1071–1079. [Google Scholar] [CrossRef] [PubMed]
- Hara-Chikuma, M.; Watanabe, S.; Satooka, H. Involvement of aquaporin-3 in epidermal growth factor receptor signaling via hydrogen peroxide transport in cancer cells. Biochem. Biophys. Res. Commun. 2016, 471, 603–609. [Google Scholar] [CrossRef] [PubMed]
- Bechtel, W.; Bauer, G. Catalase protects tumor cells from apoptosis induction by intercellular ROS signaling. Anticancer Res. 2009, 29, 4541–4557. [Google Scholar] [PubMed]
- Heinzelmann, S.; Bauer, G. Multiple protective functions of catalase against intercellular apoptosis-inducing ROS signaling of human tumor cells. Biol. Chem. 2010, 391, 675–693. [Google Scholar] [CrossRef] [PubMed]
- Bohm, B.; Heinzelmann, S.; Motz, M.; Bauer, G. Extracellular localization of catalase is associated with the transformed state of malignant cells. Biol. Chem. 2015, 396, 1339–1356. [Google Scholar] [CrossRef] [PubMed]
- Moran, E.C.; Kamiguti, A.S.; Cawley, J.C.; Pettitt, A.R. Cytoprotective antioxidant activity of serum albumin and autocrine catalase in chronic lymphocytic leukaemia. Br. J. Haematol. 2002, 116, 316–328. [Google Scholar] [CrossRef] [PubMed]
- Poole, L.B. The basics of thiols and cysteines in redox biology and chemistry. Free Radic. Biol. Med. 2015, 80, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Delaunay, A.; Isnard, A.D.; Toledano, M.B. H2O2 sensing through oxidation of the Yap1 transcription factor. EMBO J. 2000, 19, 5157–5166. [Google Scholar] [CrossRef] [PubMed]
- Delaunay, A.; Pflieger, D.; Barrault, M.B.; Vinh, J.; Toledano, M.B. A thiol peroxidase is an H2O2 receptor and redox-transducer in gene activation. Cell 2002, 111, 471–481. [Google Scholar] [CrossRef]
- Veal, E.A.; Ross, S.J.; Malakasi, P.; Peacock, E.; Morgan, B.A. Ybp1 is required for the hydrogen peroxide-induced oxidation of the Yap1 transcription factor. J. Biol. Chem. 2003, 278, 30896–30904. [Google Scholar] [CrossRef] [PubMed]
- Toledano, M.B.; Delaunay, A.; Monceau, L.; Tacnet, F. Microbial H2O2 sensors as archetypical redox signaling modules. Trends Biochem. Sci 2004, 29, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Gutscher, M.; Sobotta, M.C.; Wabnitz, G.H.; Ballikaya, S.; Meyer, A.J.; Samstag, Y.; Dick, T.P. Proximity-based protein thiol oxidation by H2O2-scavenging peroxidases. J. Biol. Chem. 2009, 284, 31532–31540. [Google Scholar] [CrossRef] [PubMed]
- Calvo, I.A.; Boronat, S.; Domenech, A.; Garcia-Santamarina, S.; Ayte, J.; Hidalgo, E. Dissection of a redox relay: H2O2-dependent activation of the transcription factor Pap1 through the peroxidatic Tpx1-thioredoxin cycle. Cell Rep. 2013, 5, 1413–1424. [Google Scholar] [CrossRef] [PubMed]
- Sobotta, M.C.; Liou, W.; Stocker, S.; Talwar, D.; Oehler, M.; Ruppert, T.; Scharf, A.N.; Dick, T.P. Peroxiredoxin-2 and STAT3 form a redox relay for H2O2 signaling. Nat. Chem. Biol. 2015, 11, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Veal, E.A.; Day, A.M.; Morgan, B.A. Hydrogen peroxide sensing and signaling. Mol. Cell 2007, 26, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Perkins, A.; Nelson, K.J.; Parsonage, D.; Poole, L.B.; Karplus, P.A. Peroxiredoxins: Guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem. Sci. 2015, 40, 435–445. [Google Scholar] [CrossRef] [PubMed]
- Netto, L.E.; Antunes, F. The Roles of Peroxiredoxin and Thioredoxin in Hydrogen Peroxide Sensing and in Signal Transduction. Mol. Cells 2016, 39, 65–71. [Google Scholar] [PubMed]
- Stocker, S.; Van Laer, K.; Mijuskovic, A.; Dick, T.P. The Conundrum of Hydrogen Peroxide Signaling and the Emerging Role of Peroxiredoxins as Redox Relay Hubs. Antioxid. Redox Signal. 2018, 28, 558–573. [Google Scholar] [CrossRef] [PubMed]
- Young, D.; Pedre, B.; Ezerina, D.; De Smet, B.; Lewandowska, A.; Tossounian, M.A.; Bodra, N.; Huang, J.; Astolfi Rosado, L.; Van Breusegem, F.; et al. Protein Promiscuity in H2O2 Signaling. Antioxid. Redox Signal. 2018. [Google Scholar] [CrossRef] [PubMed]
- Hanschmann, E.M.; Godoy, J.R.; Berndt, C.; Hudemann, C.; Lillig, C.H. Thioredoxins, glutaredoxins, and peroxiredoxins-molecular mechanisms and health significance: From cofactors to antioxidants to redox signaling. Antioxid. Redox Signal. 2013, 19, 1539–1605. [Google Scholar] [CrossRef] [PubMed]
- Espinosa-Diez, C.; Miguel, V.; Mennerich, D.; Kietzmann, T.; Sanchez-Perez, P.; Cadenas, S.; Lamas, S. Antioxidant responses and cellular adjustments to oxidative stress. Redox Biol. 2015, 6, 183–197. [Google Scholar] [CrossRef] [PubMed]
- Reczek, C.R.; Chandel, N.S. ROS-dependdent signal transduction. Curr. Opin. Cell Boil. 2015, 33, 8–13. [Google Scholar] [CrossRef] [PubMed]
- Sies, H.; Berndt, C.; Jones, D.P. Oxidative Stress. Ann. Rev Biochem. 2017, 86, 715–748. [Google Scholar] [CrossRef] [PubMed]
- Veal, E.A.; Underwood, Z.E.; Tomalin, L.E.; Morgan, B.A.; Pillay, C.S. Hyperoxidation of Peroxiredoxins: Gain or Loss of Function? Antioxid. Redox Signal. 2018, 28, 574–590. [Google Scholar] [CrossRef] [PubMed]
- Veal, E.A.; Findlay, V.J.; Day, A.M.; Bozonet, S.M.; Evans, J.M.; Quinn, J.; Morgan, B.A. A 2-Cys peroxiredoxin regulates peroxide-induced oxidation and activation of a stress-activated MAP kinase. Mol. Cell 2004, 15, 129–139. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, T.; Okazaki, S.; Murayama, A.; Naganuma, A.; Nomoto, A.; Kuge, S. A major peroxiredoxin-induced activation of Yap1 transcription factor is mediated by reduction-sensitive disulfide bonds and reveals a low level of transcriptional activation. J. Biol. Chem. 2009, 284, 4464–4472. [Google Scholar] [CrossRef] [PubMed]
- Iwai, K.; Naganuma, A.; Kuge, S. Peroxiredoxin Ahp1 acts as a receptor for alkylhydroperoxides to induce disulfide bond formation in the Cad1 transcription factor. J. Biol. Chem. 2010, 285, 10597–10604. [Google Scholar] [CrossRef] [PubMed]
- Fomenko, D.E.; Koc, A.; Agisheva, N.; Jacobsen, M.; Kaya, A.; Malinouski, M.; Rutherford, J.C.; Siu, K.L.; Jin, D.Y.; Winge, D.R.; et al. Thiol peroxidases mediate specific genome-wide regulation of gene expression in response to hydrogen peroxide. Proc. Natl. Acad. Sci. USA 2011, 108, 2729–2734. [Google Scholar] [CrossRef] [PubMed]
- Jarvis, R.M.; Hughes, S.M.; Ledgerwood, E.C. Peroxiredoxin 1 functions as a signal peroxidase to receive, transduce, and transmit peroxide signals in mammalian cells. Free Radic. Biol. Med. 2012, 53, 1522–1530. [Google Scholar] [CrossRef] [PubMed]
- Stocker, S.; Maurer, M.; Ruppert, T.; Dick, T.P. A role for 2-Cys peroxiredoxins in facilitating cytosolic protein thiol oxidation. Nat. Chem. Biol. 2018, 14, 148–155. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.; Fernandes, E.; Lima, J.L. Fluorescence probes used for detection of reactive oxygen species. J. Biochem. Biophys. Methods 2005, 65, 45–80. [Google Scholar] [CrossRef] [PubMed]
- Rhee, S.G.; Chang, T.S.; Jeong, W.; Kang, D. Methods for detection and measurement of hydrogen peroxide inside and outside of cells. Mol. Cells 2010, 29, 539–549. [Google Scholar] [CrossRef] [PubMed]
- Kalyanaraman, B.; Darley-Usmar, V.; Davies, K.J.; Dennery, P.A.; Forman, H.J.; Grisham, M.B.; Mann, G.E.; Moore, K.; Roberts, L.J., 2nd; Ischiropoulos, H. Measuring reactive oxygen and nitrogen species with fluorescent probes: Challenges and limitations. Free Radic. Biol. Med. 2012, 52, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Belousov, V.V.; Fradkov, A.F.; Lukyanov, K.A.; Staroverov, D.B.; Shakhbazov, K.S.; Terskikh, A.V.; Lukyanov, S. Genetically encoded fluorescent indicator for intracellular hydrogen peroxide. Nat. Methods 2006, 3, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Poburko, D.; Santo-Domingo, J.; Demaurex, N. Dynamic regulation of the mitochondrial proton gradient during cytosolic calcium elevations. J. Biol. Chem. 2011, 286, 11672–11684. [Google Scholar] [CrossRef] [PubMed]
- Bilan, D.S.; Belousov, V.V. HyPer Family Probes: State of the Art. Antioxid. Redox Signal. 2016, 24, 731–751. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Ishibashi, S.; Iglesias-Gonzalez, J.; Chen, Y.; Love, N.R.; Amaya, E. Ca2+-Induced Mitochondrial ROS Regulate the Early Embryonic Cell Cycle. Cell Rep. 2018, 22, 218–231. [Google Scholar] [CrossRef] [PubMed]
- Knoefler, D.; Thamsen, M.; Koniczek, M.; Niemuth, N.J.; Diederich, A.K.; Jakob, U. Quantitative in vivo redox sensors uncover oxidative stress as an early event in life. Mol. Cell 2012, 47, 767–776. [Google Scholar] [CrossRef] [PubMed]
- Gauron, C.; Meda, F.; Dupont, E.; Albadri, S.; Quenech’Du, N.; Ipendey, E.; Volovitch, M.; Del Bene, F.; Joliot, A.; Rampon, C.; et al. Hydrogen peroxide (H2O2) controls axon pathfinding during zebrafish development. Dev. Biol. 2016, 414, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Back, P.; De Vos, W.H.; Depuydt, G.G.; Matthijssens, F.; Vanfleteren, J.R.; Braeckman, B.P. Exploring real-time in vivo redox biology of developing and aging Caenorhabditis elegans. Free Radic. Biol. Med. 2012, 52, 850–859. [Google Scholar] [CrossRef] [PubMed]
- Morgan, B.; Van Laer, K.; Owusu, T.N.; Ezerina, D.; Pastor-Flores, D.; Amponsah, P.S.; Tursch, A.; Dick, T.P. Real-time monitoring of basal H2O2 levels with peroxiredoxin-based probes. Nat. Chem. Biol. 2016, 12, 437–443. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, S.C.; Barata, A.G.; Grosshans, J.; Teleman, A.A.; Dick, T.P. In vivo mapping of hydrogen peroxide and oxidized glutathione reveals chemical and regional specificity of redox homeostasis. Cell Metab. 2011, 14, 819–829. [Google Scholar] [CrossRef] [PubMed]
- Braeckman, B.P.; Smolders, A.; Back, P.; De Henau, S. In Vivo Detection of Reactive Oxygen Species and Redox Status in Caenorhabditis elegans. Antioxid. Redox Signal. 2016, 25, 577–592. [Google Scholar] [CrossRef] [PubMed]
- Fujikawa, Y.; Roma, L.P.; Sobotta, M.C.; Rose, A.J.; Diaz, M.B.; Locatelli, G.; Breckwoldt, M.O.; Misgeld, T.; Kerschensteiner, M.; Herzig, S.; et al. Mouse redox histology using genetically encoded probes. Sci. Signal 2016, 9, rs1. [Google Scholar] [CrossRef] [PubMed]
- Panieri, E.; Millia, C.; Santoro, M.M. Real-time quantification of subcellular H2O2 and glutathione redox potential in living cardiovascular tissues. Free Radic. Biol. Med. 2017, 109, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Gutscher, M.; Pauleau, A.L.; Marty, L.; Brach, T.; Wabnitz, G.H.; Samstag, Y.; Meyer, A.J.; Dick, T.P. Real-time imaging of the intracellular glutathione redox potential. Nat. Methods 2008, 5, 553–559. [Google Scholar] [CrossRef] [PubMed]
- Tao, R.; Zhao, Y.; Chu, H.; Wang, A.; Zhu, J.; Chen, X.; Zou, Y.; Shi, M.; Liu, R.; Su, N.; et al. Genetically encoded fluorescent sensors reveal dynamic regulation of NADPH metabolism. Nat. Methods 2017, 14, 720–728. [Google Scholar] [CrossRef] [PubMed]
- Ufer, C.; Wang, C.C.; Borchert, A.; Heydeck, D.; Kuhn, H. Redox control in mammalian embryo development. Antioxid. Redox Signal. 2010, 13, 833–875. [Google Scholar] [CrossRef] [PubMed]
- Forman, H.J. Redox signaling: An evolution from free radicals to aging. Free Radic. Biol. Med. 2016, 97, 398–407. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.; Zou, L.; Zhang, X.; Branco, V.; Wang, J.; Carvalho, C.; Holmgren, A.; Lu, J. Redox Signaling Mediated by Thioredoxin and Glutathione Systems in the Central Nervous System. Antioxid. Redox Signal. 2017, 27, 989–1010. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.; Munoz-Palma, E.; Gonzalez-Billault, C. From birth to death: A role for reactive oxygen species in neuronal development. Semin. Cell Dev. Biol. 2018, 80, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Olguin-Albuerne, M.; Moran, J. Redox Signaling Mechanisms in Nervous System Development. Antioxid. Redox Signal. 2018, 28, 1603–1625. [Google Scholar] [CrossRef] [PubMed]
- Morishita, H.; Cabungcal, J.H.; Chen, Y.; Do, K.Q.; Hensch, T.K. Prolonged Period of Cortical Plasticity upon Redox Dysregulation in Fast-Spiking Interneurons. Biol. Psychiatry 2015, 78, 396–402. [Google Scholar] [CrossRef] [PubMed]
- Do, K.Q.; Cabungcal, J.H.; Frank, A.; Steullet, P.; Cuenod, M. Redox dysregulation, neurodevelopment, and schizophrenia. Curr. Opin. Neurobiol. 2009, 19, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Le Belle, J.E.; Sperry, J.; Ngo, A.; Ghochani, Y.; Laks, D.R.; Lopez-Aranda, M.; Silva, A.J.; Kornblum, H.I. Maternal inflammation contributes to brain overgrowth and autism-associated behaviors through altered redox signaling in stem and progenitor cells. Stem. Cell Rep. 2014, 3, 725–734. [Google Scholar] [CrossRef] [PubMed]
- Katsuyama, M. NOX/NADPH oxidase, the superoxide-generating enzyme: Its transcriptional regulation and physiological roles. J. Pharmacol. Sci. 2010, 114, 134–146. [Google Scholar] [CrossRef] [PubMed]
- Weaver, C.J.; Leung, Y.F.; Suter, D.M. Expression dynamics of NADPH oxidases during early zebrafish development. J. Comp. Neurol. 2016, 524, 2130–2141. [Google Scholar] [CrossRef] [PubMed]
- Milenkovic, M.; De Deken, X.; Jin, L.; De Felice, M.; Di Lauro, R.; Dumont, J.E.; Corvilain, B.; Miot, F. Duox expression and related H2O2 measurement in mouse thyroid: Onset in embryonic development and regulation by TSH in adult. J. Endocrinol. 2007, 192, 615–626. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.S.; Choi, H.W.; Yoon, H.E.; Kim, I.Y. Reactive oxygen species generated by NADPH oxidase 2 and 4 are required for chondrogenic differentiation. J. Biol. Chem. 2010, 285, 40294–40302. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Stouffs, M.; Serrander, L.; Banfi, B.; Bettiol, E.; Charnay, Y.; Steger, K.; Krause, K.H.; Jaconi, M.E. The NADPH oxidase NOX4 drives cardiac differentiation: Role in regulating cardiac transcription factors and MAP kinase activation. Mol. Biol. Cell 2006, 17, 3978–3988. [Google Scholar] [CrossRef] [PubMed]
- Buggisch, M.; Ateghang, B.; Ruhe, C.; Strobel, C.; Lange, S.; Wartenberg, M.; Sauer, H. Stimulation of ES-cell-derived cardiomyogenesis and neonatal cardiac cell proliferation by reactive oxygen species and NADPH oxidase. J. Cell Sci. 2007, 120, 885–894. [Google Scholar] [CrossRef] [PubMed]
- Ren, F.; Wang, K.; Zhang, T.; Jiang, J.; Nice, E.C.; Huang, C. New insights into redox regulation of stem cell self-renewal and differentiation. Biochim. Biophys. Acta 2015, 1850, 1518–1526. [Google Scholar] [CrossRef] [PubMed]
- Skonieczna, M.; Hejmo, T.; Poterala-Hejmo, A.; Cieslar-Pobuda, A.; Buldak, R.J. NADPH Oxidases: Insights into Selected Functions and Mechanisms of Action in Cancer and Stem Cells. Oxid. Med. Cell Longev. 2017, 2017, 9420539. [Google Scholar] [CrossRef] [PubMed]
- Thisse, B.; Thisse, C. Fast Release Clones: A High Throughput Expression Analysis. ZFIN Direct Data Submiss. 2004, in press. [Google Scholar]
- Brautigam, L.; Schutte, L.D.; Godoy, J.R.; Prozorovski, T.; Gellert, M.; Hauptmann, G.; Holmgren, A.; Lillig, C.H.; Berndt, C. Vertebrate-specific glutaredoxin is essential for brain development. Proc. Natl. Acad. Sci. USA 2011, 108, 20532–20537. [Google Scholar] [CrossRef] [PubMed]
- Haunhorst, P.; Hanschmann, E.M.; Brautigam, L.; Stehling, O.; Hoffmann, B.; Muhlenhoff, U.; Lill, R.; Berndt, C.; Lillig, C.H. Crucial function of vertebrate glutaredoxin 3 (PICOT) in iron homeostasis and hemoglobin maturation. Mol. Biol. Cell 2013, 24, 1895–1903. [Google Scholar] [CrossRef] [PubMed]
- Wingert, R.A.; Galloway, J.L.; Barut, B.; Foott, H.; Fraenkel, P.; Axe, J.L.; Weber, G.J.; Dooley, K.; Davidson, A.J.; Schmid, B.; et al. Deficiency of glutaredoxin 5 reveals Fe-S clusters are required for vertebrate haem synthesis. Nature 2005, 436, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Thisse, B.; Pflumio, S.; Fürthauer, M.; Loppin, B.; Heyer, V. Expression of the zebrafish genome during embryogenesis (NIH R01 RR15402). ZFIN Direct Data Submiss. 2001, in press. [Google Scholar]
- Yang, L.; Kemadjou, J.R.; Zinsmeister, C.; Bauer, M.; Legradi, J.; Muller, F.; Pankratz, M.; Jakel, J.; Strahle, U. Transcriptional profiling reveals barcode-like toxicogenomic responses in the zebrafish embryo. Genome Biol. 2007, 8, R227. [Google Scholar] [CrossRef] [PubMed]
- Seiler, C.; Davuluri, G.; Abrams, J.; Byfield, F.J.; Janmey, P.A.; Pack, M. Smooth muscle tension induces invasive remodeling of the zebrafish intestine. PLoS Biol. 2012, 10, e1001386. [Google Scholar] [CrossRef] [PubMed]
- Mendieta-Serrano, M.A.; Schnabel, D.; Lomeli, H.; Salas-Vidal, E. Spatial and temporal expression of zebrafish glutathione peroxidase 4 a and b genes during early embryo development. Gene Expr. Patterns 2015, 19, 98–107. [Google Scholar] [CrossRef] [PubMed]
- Rong, X.; Zhou, Y.; Liu, Y.; Zhao, B.; Wang, B.; Wang, C.; Gong, X.; Tang, P.; Lu, L.; Li, Y.; et al. Glutathione peroxidase 4 inhibits Wnt/beta-catenin signaling and regulates dorsal organizer formation in zebrafish embryos. Development 2017, 144, 1687–1697. [Google Scholar] [CrossRef] [PubMed]
- Flores, M.V.; Crawford, K.C.; Pullin, L.M.; Hall, C.J.; Crosier, K.E.; Crosier, P.S. Dual oxidase in the intestinal epithelium of zebrafish larvae has anti-bacterial properties. Biochem. Biophys. Res. Commun. 2010, 400, 164–168. [Google Scholar] [CrossRef] [PubMed]
- Opitz, R.; Maquet, E.; Zoenen, M.; Dadhich, R.; Costagliola, S. TSH receptor function is required for normal thyroid differentiation in zebrafish. Mol. Endocrinol. 2011, 25, 1579–1599. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.C.; Chiu, C.C.; Chang, H.W.; Wang, Y.S.; Syue, H.H.; Song, Y.C.; Weng, Z.H.; Tai, M.H.; Wu, C.Y. Prdx1-encoded peroxiredoxin is important for vascular development in zebrafish. FEBS Lett. 2017, 591, 889–902. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, H.; Nakajima-Takagi, Y.; Tsujita, T.; Akiyama, S.; Wakasa, T.; Mukaigasa, K.; Kaneko, H.; Tamaru, Y.; Yamamoto, M.; Kobayashi, M. Tissue-restricted expression of Nrf2 and its target genes in zebrafish with gene-specific variations in the induction profiles. PLoS ONE 2011, 6, e26884. [Google Scholar] [CrossRef] [PubMed]
- Hegde, A.; Qiu, N.C.; Qiu, X.; Ho, S.H.; Tay, K.Q.; George, J.; Ng, F.S.; Govindarajan, K.R.; Gong, Z.; Mathavan, S.; et al. Genomewide expression analysis in zebrafish mind bomb alleles with pancreas defects of different severity identifies putative Notch responsive genes. PLoS ONE 2008, 3, e1479. [Google Scholar] [CrossRef] [PubMed]
- Bazzini, A.A.; Lee, M.T.; Giraldez, A.J. Ribosome profiling shows that miR-430 reduces translation before causing mRNA decay in zebrafish. Science 2012, 336, 233–237. [Google Scholar] [CrossRef] [PubMed]
- Peterman, E.M.; Sullivan, C.; Goody, M.F.; Rodriguez-Nunez, I.; Yoder, J.A.; Kim, C.H. Neutralization of mitochondrial superoxide by superoxide dismutase 2 promotes bacterial clearance and regulates phagocyte numbers in zebrafish. Infect. Immun. 2015, 83, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Priyadarshini, M.; Tuimala, J.; Chen, Y.C.; Panula, P. A zebrafish model of PINK1 deficiency reveals key pathway dysfunction including HIF signaling. Neurobiol. Dis. 2013, 54, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Weaver, C.J.; Terzi, A.; Roeder, H.; Gurol, T.; Deng, Q.; Leung, Y.F.; Suter, D.M. nox2/cybb Deficiency Affects Zebrafish Retinotectal Connectivity. J. Neurosci. 2018, 38, 5854–5871. [Google Scholar] [CrossRef] [PubMed]
- Chopra, K.; Ishibashi, S.; Amaya, E. Zebrafish duox mutations provide a model for human congenital hypothyroidism. bioRxiv 2018. [Google Scholar] [CrossRef]
- Al-Sabbagh, M.; Fusi, L.; Higham, J.; Lee, Y.; Lei, K.; Hanyaloglu, A.C.; Lam, E.W.; Christian, M.; Brosens, J.J. NADPH oxidase-derived reactive oxygen species mediate decidualization of human endometrial stromal cells in response to cyclic AMP signaling. Endocrinology 2011, 152, 730–740. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.; Nunez, M.T.; Gonzalez-Billault, C. Contribution of NADPH oxidase to the establishment of hippocampal neuronal polarity in culture. J. Cell Sci. 2015, 128, 2989–2995. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.; Munoz-Palma, E.; Henriquez, D.R.; Palmisano, I.; Nunez, M.T.; Di Giovanni, S.; Gonzalez-Billault, C. A Feed-Forward Mechanism Involving the NOX Complex and RyR-Mediated Ca2+ Release During Axonal Specification. J. Neurosci. 2016, 36, 11107–11119. [Google Scholar] [CrossRef] [PubMed]
- Olguin-Albuerne, M.; Moran, J. ROS produced by NOX2 control in vitro development of cerebellar granule neurons development. ASN Neuro 2015, 7. [Google Scholar] [CrossRef] [PubMed]
- Munnamalai, V.; Suter, D.M. Reactive oxygen species regulate F-actin dynamics in neuronal growth cones and neurite outgrowth. J. Neurochem. 2009, 108, 644–661. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.F.; Forscher, P. Rac1 modulates stimulus-evoked Ca2+ release in neuronal growth cones via parallel effects on microtubule/endoplasmic reticulum dynamics and reactive oxygen species production. Mol. Biol. Cell 2009, 20, 3700–3712. [Google Scholar] [CrossRef] [PubMed]
- Munnamalai, V.; Weaver, C.J.; Weisheit, C.E.; Venkatraman, P.; Agim, Z.S.; Quinn, M.T.; Suter, D.M. Bidirectional interactions between NOX2-type NADPH oxidase and the F-actin cytoskeleton in neuronal growth cones. J. Neurochem. 2014, 130, 526–540. [Google Scholar] [CrossRef] [PubMed]
- Boudreau, H.E.; Casterline, B.W.; Rada, B.; Korzeniowska, A.; Leto, T.L. Nox4 involvement in TGF-beta and SMAD3-driven induction of the epithelial-to-mesenchymal transition and migration of breast epithelial cells. Free Radic. Biol. Med. 2012, 53, 1489–1499. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.H.; Lee, Y.H.; Shih, H.Y.; Chen, S.H.; Cheng, Y.C.; Tsun-Yee Chiu, D. Glucose-6-phosphate dehydrogenase is indispensable in embryonic development by modulation of epithelial-mesenchymal transition via the NOX/Smad3/miR-200b axis. Cell Death Dis. 2018, 9, 10. [Google Scholar] [CrossRef] [PubMed]
- el-Hage, S.; Singh, S.M. Temporal expression of genes encoding free radical-metabolizing enzymes is associated with higher mRNA levels during in utero development in mice. Dev. Genet. 1990, 11, 149–159. [Google Scholar] [CrossRef] [PubMed]
- Radyuk, S.N.; Klichko, V.I.; Orr, W.C. Catalase expression in Drosophila melanogaster is responsive to ecdysone and exhibits both transcriptional and post-transcriptional regulation. Arch. Insect Biochem. Physiol. 2000, 45, 79–93. [Google Scholar] [CrossRef]
- Hansen, J.M.; Harris, C. Glutathione during embryonic development. Biochim. Biophys. Acta 2015, 1850, 1527–1542. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.Z.; Osei-Frimpong, J.; Kala, G.; Kala, S.V.; Barrios, R.J.; Habib, G.M.; Lukin, D.J.; Danney, C.M.; Matzuk, M.M.; Lieberman, M.W. Glutathione synthesis is essential for mouse development but not for cell growth in culture. Proc. Natl. Acad. Sci. USA 2000, 97, 5101–5106. [Google Scholar] [CrossRef] [PubMed]
- Winkler, A.; Njalsson, R.; Carlsson, K.; Elgadi, A.; Rozell, B.; Abraham, L.; Ercal, N.; Shi, Z.Z.; Lieberman, M.W.; Larsson, A.; et al. Glutathione is essential for early embryogenesis—Analysis of a glutathione synthetase knockout mouse. Biochem. Biophys Res. Commun. 2011, 412, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Timme-Laragy, A.R.; Goldstone, J.V.; Imhoff, B.R.; Stegeman, J.J.; Hahn, M.E.; Hansen, J.M. Glutathione redox dynamics and expression of glutathione-related genes in the developing embryo. Free Radic. Biol. Med. 2013, 65, 89–101. [Google Scholar] [CrossRef] [PubMed]
- Jurado, J.; Prieto-Alamo, M.J.; Madrid-Risquez, J.; Pueyo, C. Absolute gene expression patterns of thioredoxin and glutaredoxin redox systems in mouse. J. Biol. Chem. 2003, 278, 45546–45554. [Google Scholar] [CrossRef] [PubMed]
- Karunakaran, S.; Saeed, U.; Ramakrishnan, S.; Koumar, R.C.; Ravindranath, V. Constitutive expression and functional characterization of mitochondrial glutaredoxin (Grx2) in mouse and human brain. Brain Res. 2007, 1185, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Lonn, M.E.; Hudemann, C.; Berndt, C.; Cherkasov, V.; Capani, F.; Holmgren, A.; Lillig, C.H. Expression pattern of human glutaredoxin 2 isoforms: Identification and characterization of two testis/cancer cell-specific isoforms. Antioxid. Redox Signal. 2008, 10, 547–557. [Google Scholar] [CrossRef] [PubMed]
- Hudemann, C.; Lonn, M.E.; Godoy, J.R.; Zahedi Avval, F.; Capani, F.; Holmgren, A.; Lillig, C.H. Identification, expression pattern, and characterization of mouse glutaredoxin 2 isoforms. Antioxid. Redox Signal. 2009, 11, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Choi, C.H.; Kim, B.J.; Jeong, S.Y.; Lee, C.H.; Kim, J.S.; Park, S.J.; Yim, H.S.; Kang, S.O. Reduced glutathione levels affect the culmination and cell fate decision in Dictyostelium discoideum. Dev. Biol. 2006, 295, 523–533. [Google Scholar] [CrossRef] [PubMed]
- Gellert, M.; Venz, S.; Mitlohner, J.; Cott, C.; Hanschmann, E.M.; Lillig, C.H. Identification of a dithiol-disulfide switch in collapsin response mediator protein 2 (CRMP2) that is toggled in a model of neuronal differentiation. J. Biol. Chem. 2013, 288, 35117–35125. [Google Scholar] [CrossRef] [PubMed]
- Brautigam, L.; Jensen, L.D.; Poschmann, G.; Nystrom, S.; Bannenberg, S.; Dreij, K.; Lepka, K.; Prozorovski, T.; Montano, S.J.; Aktas, O.; et al. Glutaredoxin regulates vascular development by reversible glutathionylation of sirtuin 1. Proc. Natl. Acad. Sci. USA 2013, 110, 20057–20062. [Google Scholar] [CrossRef] [PubMed]
- Berndt, C.; Poschmann, G.; Stuhler, K.; Holmgren, A.; Brautigam, L. Zebrafish heart development is regulated via glutaredoxin 2 dependent migration and survival of neural crest cells. Redox Biol. 2014, 2, 673–678. [Google Scholar] [CrossRef] [PubMed]
- Cheng, N.H.; Zhang, W.; Chen, W.Q.; Jin, J.; Cui, X.; Butte, N.F.; Chan, L.; Hirschi, K.D. A mammalian monothiol glutaredoxin, Grx3, is critical for cell cycle progression during embryogenesis. FEBS J. 2011, 278, 2525–2539. [Google Scholar] [CrossRef] [PubMed]
- Pham, K.; Dong, J.; Jiang, X.; Qu, Y.; Yu, H.; Yang, Y.; Olea, W.; Marini, J.C.; Chan, L.; Wang, J.; et al. Loss of glutaredoxin 3 impedes mammary lobuloalveolar development during pregnancy and lactation. Am. J. Physiol. Endocrinol. Metab. 2017, 312, E136–E149. [Google Scholar] [CrossRef] [PubMed]
- Lillig, C.H.; Berndt, C.; Holmgren, A. Glutaredoxin systems. Biochim. Biophys. Acta 2008, 1780, 1304–1317. [Google Scholar] [CrossRef] [PubMed]
- Fujii, S.; Nanbu, Y.; Konishi, I.; Mori, T.; Masutani, H.; Yodoi, J. Immunohistochemical localization of adult T-cell leukaemia-derived factor, a human thioredoxin homologue, in human fetal tissues. Virchows Arch. A Pathol. Anat. Histopathol. 1991, 419, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Matsui, M.; Oshima, M.; Oshima, H.; Takaku, K.; Maruyama, T.; Yodoi, J.; Taketo, M.M. Early embryonic lethality caused by targeted disruption of the mouse thioredoxin gene. Dev. Biol. 1996, 178, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Nonn, L.; Williams, R.R.; Erickson, R.P.; Powis, G. The absence of mitochondrial thioredoxin 2 causes massive apoptosis, exencephaly, and early embryonic lethality in homozygous mice. Mol. Cell. Biol. 2003, 23, 916–922. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Cui, X.; Wang, L.; Liu, F.; Jiang, T.; Li, C.; Li, D.; Huang, M.; Liao, S.; Wang, J.; et al. The mitochondrial thioredoxin is required for liver development in zebrafish. Curr. Mol. Med. 2014, 14, 772–782. [Google Scholar] [CrossRef] [PubMed]
- Pirson, M.; Debrulle, S.; Clippe, A.; Clotman, F.; Knoops, B. Thioredoxin-2 Modulates Neuronal Programmed Cell Death in the Embryonic Chick Spinal Cord in Basal and Target-Deprived Conditions. PLoS ONE 2015, 10, e0142280. [Google Scholar] [CrossRef] [PubMed]
- Jakupoglu, C.; Przemeck, G.K.; Schneider, M.; Moreno, S.G.; Mayr, N.; Hatzopoulos, A.K.; de Angelis, M.H.; Wurst, W.; Bornkamm, G.W.; Brielmeier, M.; et al. Cytoplasmic thioredoxin reductase is essential for embryogenesis but dispensable for cardiac development. Mol. Cell Biol. 2005, 25, 1980–1988. [Google Scholar] [CrossRef] [PubMed]
- Conrad, M.; Jakupoglu, C.; Moreno, S.G.; Lippl, S.; Banjac, A.; Schneider, M.; Beck, H.; Hatzopoulos, A.K.; Just, U.; Sinowatz, F.; et al. Essential role for mitochondrial thioredoxin reductase in hematopoiesis, heart development, and heart function. Mol. Cell Biol. 2004, 24, 9414–9423. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Holmgren, A. The thioredoxin antioxidant system. Free Radic. Biol. Med. 2014, 66, 75–87. [Google Scholar] [CrossRef] [PubMed]
- Hurd, T.R.; DeGennaro, M.; Lehmann, R. Redox regulation of cell migration and adhesion. Trends Cell Biol. 2012, 22, 107–115. [Google Scholar] [CrossRef] [PubMed]
- DeGennaro, M.; Hurd, T.R.; Siekhaus, D.E.; Biteau, B.; Jasper, H.; Lehmann, R. Peroxiredoxin stabilization of DE-cadherin promotes primordial germ cell adhesion. Dev. Cell 2011, 20, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Shafer, M.E.; Willson, J.A.; Damjanovski, S. Expression analysis of the peroxiredoxin gene family during early development in Xenopus laevis. Gene Expr. Patterns 2011, 11, 511–516. [Google Scholar] [CrossRef] [PubMed]
- Shan, S.W.; Tang, M.K.; Cai, D.Q.; Chui, Y.L.; Chow, P.H.; Grotewold, L.; Lee, K.K. Comparative proteomic analysis identifies protein disulfide isomerase and peroxiredoxin 1 as new players involved in embryonic interdigital cell death. Dev. Dyn. 2005, 233, 266–281. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Sabharwal, P.; Rao, M.; Sockanathan, S. The antioxidant enzyme Prdx1 controls neuronal differentiation by thiol-redox-dependent activation of GDE2. Cell 2009, 138, 1209–1221. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Wladyka, C.; Fujii, J.; Sockanathan, S. Prdx4 is a compartment-specific H2O2 sensor that regulates neurogenesis by controlling surface expression of GDE2. Nat. Commun. 2015, 6, 7006. [Google Scholar] [CrossRef] [PubMed]
- Morimoto, H.; Iwata, K.; Ogonuki, N.; Inoue, K.; Atsuo, O.; Kanatsu-Shinohara, M.; Morimoto, T.; Yabe-Nishimura, C.; Shinohara, T. ROS are required for mouse spermatogonial stem cell self-renewal. Cell Stem Cell 2013, 12, 774–786. [Google Scholar] [CrossRef] [PubMed]
- Conrad, M.; Ingold, I.; Buday, K.; Kobayashi, S.; Angeli, J.P. ROS, thiols and thiol-regulating systems in male gametogenesis. Biochim. Biophys. Acta 2015, 1850, 1566–1574. [Google Scholar] [CrossRef] [PubMed]
- Noblanc, A.; Kocer, A.; Chabory, E.; Vernet, P.; Saez, F.; Cadet, R.; Conrad, M.; Drevet, J.R. Glutathione peroxidases at work on epididymal spermatozoa: An example of the dual effect of reactive oxygen species on mammalian male fertilizing ability. J. Androl. 2011, 32, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Bigarella, C.L.; Liang, R.; Ghaffari, S. Stem cells and the impact of ROS signaling. Development 2014, 141, 4206–4218. [Google Scholar] [CrossRef] [PubMed]
- Tatapudy, S.; Aloisio, F.; Barber, D.; Nystul, T. Cell fate decisions: Emerging roles for metabolic signals and cell morphology. EMBO Rep. 2017, 18, 2105–2118. [Google Scholar] [CrossRef] [PubMed]
- Owusu-Ansah, E.; Banerjee, U. Reactive oxygen species prime Drosophila haematopoietic progenitors for differentiation. Nature 2009, 461, 537–541. [Google Scholar] [CrossRef] [PubMed]
- Su, B.; Mitra, S.; Gregg, H.; Flavahan, S.; Chotani, M.A.; Clark, K.R.; Goldschmidt-Clermont, P.J.; Flavahan, N.A. Redox regulation of vascular smooth muscle cell differentiation. Circ. Res. 2001, 89, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Jang, Y.Y.; Sharkis, S.J. A low level of reactive oxygen species selects for primitive hematopoietic stem cells that may reside in the low-oxygenic niche. Blood 2007, 110, 3056–3063. [Google Scholar] [CrossRef] [PubMed]
- Hochmuth, C.E.; Biteau, B.; Bohmann, D.; Jasper, H. Redox regulation by Keap1 and Nrf2 controls intestinal stem cell proliferation in Drosophila. Cell Stem. Cell 2011, 8, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Le Belle, J.E.; Orozco, N.M.; Paucar, A.A.; Saxe, J.P.; Mottahedeh, J.; Pyle, A.D.; Wu, H.; Kornblum, H.I. Proliferative neural stem cells have high endogenous ROS levels that regulate self-renewal and neurogenesis in a PI3K/Akt-dependant manner. Cell Stem. Cell 2011, 8, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Dickinson, B.C.; Peltier, J.; Stone, D.; Schaffer, D.V.; Chang, C.J. Nox2 redox signaling maintains essential cell populations in the brain. Nat. Chem. Biol. 2011, 7, 106–112. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Meng, S.; Li, Y.; Ghebre, Y.T.; Cooke, J.P. Optimal ROS Signaling Is Critical for Nuclear Reprogramming. Cell Rep. 2016, 15, 919–925. [Google Scholar] [CrossRef] [PubMed]
- Kang, X.; Wei, X.; Wang, X.; Jiang, L.; Niu, C.; Zhang, J.; Chen, S.; Meng, D. Nox2 contributes to the arterial endothelial specification of mouse induced pluripotent stem cells by upregulating Notch signaling. Sci. Rep. 2016, 6, 33737. [Google Scholar] [CrossRef] [PubMed]
- Sanchez Alvarado, A. Regeneration in the metazoans: Why does it happen? Bioessays 2000, 22, 578–590. [Google Scholar] [CrossRef]
- Brockes, J.P.; Kumar, A. Comparative aspects of animal regeneration. Ann. Rev. Cell Dev. Biol. 2008, 24, 525–549. [Google Scholar] [CrossRef] [PubMed]
- Galliot, B.; Ghila, L. Cell plasticity in homeostasis and regeneration. Mol. Reprod. Dev. 2010, 77, 837–855. [Google Scholar] [CrossRef] [PubMed]
- Poss, K.D. Advances in understanding tissue regenerative capacity and mechanisms in animals. Nat. Rev. Genet. 2010, 11, 710–722. [Google Scholar] [CrossRef] [PubMed]
- Sandoval-Guzman, T.; Wang, H.; Khattak, S.; Schuez, M.; Roensch, K.; Nacu, E.; Tazaki, A.; Joven, A.; Tanaka, E.M.; Simon, A. Fundamental differences in dedifferentiation and stem cell recruitment during skeletal muscle regeneration in two salamander species. Cell Stem Cell 2014, 14, 174–187. [Google Scholar] [CrossRef] [PubMed]
- Meda, F.; Rampon, C.; Dupont, E.; Gauron, C.; Mourton, A.; Queguiner, I.; Thauvin, M.; Volovitch, M.; Joliot, A.; Vriz, S. Nerves, H2O2 and Shh: Three players in the game of regeneration. Semin. Cell Dev. Biol. 2018, 80, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Pirotte, N.; Stevens, A.S.; Fraguas, S.; Plusquin, M.; Van Roten, A.; Van Belleghem, F.; Paesen, R.; Ameloot, M.; Cebria, F.; Artois, T.; et al. Reactive Oxygen Species in Planarian Regeneration: An Upstream Necessity for Correct Patterning and Brain Formation. Oxid Med. Cell Longev. 2015, 2015, 392476. [Google Scholar] [CrossRef] [PubMed]
- Love, N.R.; Chen, Y.; Ishibashi, S.; Kritsiligkou, P.; Lea, R.; Koh, Y.; Gallop, J.L.; Dorey, K.; Amaya, E. Amputation-induced reactive oxygen species are required for successful Xenopus tadpole tail regeneration. Nat. Cell Biol. 2013, 15, 222–228. [Google Scholar] [CrossRef] [PubMed]
- Gauron, C.; Rampon, C.; Bouzaffour, M.; Ipendey, E.; Teillon, J.; Volovitch, M.; Vriz, S. Sustained production of ROS triggers compensatory proliferation and is required for regeneration to proceed. Sci. Rep. 2013, 3, 2084. [Google Scholar] [CrossRef] [PubMed]
- Han, P.; Zhou, X.H.; Chang, N.; Xiao, C.L.; Yan, S.; Ren, H.; Yang, X.Z.; Zhang, M.L.; Wu, Q.; Tang, B.; et al. Hydrogen peroxide primes heart regeneration with a derepression mechanism. Cell Res 2014, 29, 1091–1107. [Google Scholar] [CrossRef] [PubMed]
- Simkin, J.; Gawriluk, T.R.; Gensel, J.C.; Seifert, A.W. Macrophages are necessary for epimorphic regeneration in African spiny mice. eLife 2017, 6, e24623. [Google Scholar] [CrossRef] [PubMed]
- Labit, E.; Rabiller, L.; Rampon, C.; Guissard, C.; Andre, M.; Barreau, C.; Cousin, B.; Carriere, A.; Eddine, M.A.; Pipy, B.; et al. Opioids prevent regeneration in adult mammals through inhibition of ROS production. Sci. Rep. 2018, 8, 12170. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Love, N.R.; Amaya, E. Tadpole tail regeneration in Xenopus. Biochem. Soc. Trans. 2014, 42, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Khanna, S.; Nallu, K.; Hunt, T.K.; Sen, C.K. Dermal wound healing is subject to redox control. Mol. Ther. 2006, 13, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Niethammer, P.; Grabher, C.; Look, A.T.; Mitchison, T.J. A tissue-scale gradient of hydrogen peroxide mediates rapid wound detection in zebrafish. Nature 2009, 459, 996–999. [Google Scholar] [CrossRef] [PubMed]
- Moreira, S.; Stramer, B.; Evans, I.; Wood, W.; Martin, P. Prioritization of competing damage and developmental signals by migrating macrophages in the Drosophila embryo. Curr. Biol. 2010, 20, 464–470. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Chisholm, A.D. C. elegans epidermal wounding induces a mitochondrial ROS burst that promotes wound repair. Dev. Cell. 2014, 31, 48–60. [Google Scholar] [CrossRef] [PubMed]
- Leon, J.; Rojo, E.; Sanchez-Serrano, J.J. Wound signalling in plants. J. Exp. Bot. 2001, 52, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Orozco-Cardenas, M.L.; Narvaez-Vasquez, J.; Ryan, C.A. Hydrogen peroxide acts as a second messenger for the induction of defense genes in tomato plants in response to wounding, systemin, and methyl jasmonate. Plant Cell 2001, 13, 179–191. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Mittler, R. Reactive oxygen species-dependent wound responses in animals and plants. Free Radic. Biol. Med. 2012, 53, 2269–2276. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.K.; Starnes, T.W.; Deng, Q.; Huttenlocher, A. Lyn is a redox sensor that mediates leukocyte wound attraction in vivo. Nature 2011, 480, 109–112. [Google Scholar] [CrossRef] [PubMed]
- Sanchez Alvarado, A.; Newmark, P.A. The use of planarians to dissect the molecular basis of metazoan regeneration. Wound Rep. Regen. 1998, 6, 413–420. [Google Scholar]
- Vriz, S.; Reiter, S.; Galliot, B. Cell death: A program to regenerate. Curr. Top. Dev. Biol. 2014, 108, 121–151. [Google Scholar] [PubMed]
- Santabarbara-Ruiz, P.; Lopez-Santillan, M.; Martinez-Rodriguez, I.; Binagui-Casas, A.; Perez, L.; Milan, M.; Corominas, M.; Serras, F. ROS-Induced JNK and p38 Signaling Is Required for Unpaired Cytokine Activation during Drosophila Regeneration. PLoS Genet. 2015, 11, e1005595. [Google Scholar] [CrossRef] [PubMed]
- Fogarty, C.E.; Diwanji, N.; Lindblad, J.L.; Tare, M.; Amcheslavsky, A.; Makhijani, K.; Bruckner, K.; Fan, Y.; Bergmann, A. Extracellular Reactive Oxygen Species Drive Apoptosis-Induced Proliferation via Drosophila Macrophages. Curr. Biol. 2016, 26, 575–584. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Wu, X.; Luo, H.; Zhao, L.; Ji, X.; Qiao, X.; Jin, Y.; Liu, W. Acute exposure of mercury chloride stimulates the tissue regeneration program and reactive oxygen species production in the Drosophila midgut. Environ. Toxicol. Pharmacol. 2016, 41, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.J.; Abidi, S.N.F.; Skinner, A.; Tian, Y.; Smith-Bolton, R.K. The Drosophila Duox maturation factor is a key component of a positive feedback loop that sustains regeneration signaling. PLoS Genet. 2017, 13, e1006937. [Google Scholar] [CrossRef] [PubMed]
- Brock, A.R.; Seto, M.; Smith-Bolton, R.K. Cap-n-Collar Promotes Tissue Regeneration by Regulating ROS and JNK Signaling in the Drosophila melanogaster Wing Imaginal Disc. Genetics 2017, 206, 1505–1520. [Google Scholar] [CrossRef] [PubMed]
- Diwanji, N.; Bergmann, A. The beneficial role of extracellular reactive oxygen species in apoptosis-induced compensatory proliferation. Fly (Austin) 2017, 11, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Amcheslavsky, A.; Wang, S.; Fogarty, C.E.; Lindblad, J.L.; Fan, Y.; Bergmann, A. Plasma Membrane Localization of Apoptotic Caspases for Non-apoptotic Functions. Dev. Cell. 2018, 45, 450–464.e3. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.; Takagi, C.; Miura, S.; Sakane, Y.; Suzuki, M.; Sakuma, T.; Sakamoto, N.; Endo, T.; Kamei, Y.; Sato, Y.; et al. In vivo tracking of histone H3 lysine 9 acetylation in Xenopus laevis during tail regeneration. Genes Cells 2016, 21, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Hameed, L.S.; Berg, D.A.; Belnoue, L.; Jensen, L.D.; Cao, Y.; Simon, A. Environmental changes in oxygen tension reveal ROS-dependent neurogenesis and regeneration in the adult newt brain. eLife 2015, 4, e08422. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, F.; Luxardi, G.; Reid, B.; Zhao, M. Early bioelectric activities mediate redox-modulated regeneration. Development 2016, 143, 4582–4594. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Wang, Y.; Man, L.; Zhu, Z.; Bai, X.; Wei, S.; Liu, Y.; Liu, M.; Wang, X.; Gu, X.; et al. Reactive oxygen species generated from skeletal muscles are required for gecko tail regeneration. Sci. Rep. 2016, 6, 20752. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.H.; Puliafito, A.; Cox, B.D.; Primo, L.; Fang, Y.; Di Talia, S.; Poss, K.D. Multicolor Cell Barcoding Technology for Long-Term Surveillance of Epithelial Regeneration in Zebrafish. Dev. Cell. 2016, 36, 668–680. [Google Scholar] [CrossRef] [PubMed]
- Meda, F.; Gauron, C.; Rampon, C.; Teillon, J.; Volovitch, M.; Vriz, S. Nerves Control Redox Levels in Mature Tissues Through Schwann Cells and Hedgehog Signaling. Antioxid. Redox Signal. 2016, 24, 299–311. [Google Scholar] [CrossRef] [PubMed]
- Bely, A.E.; Nyberg, K.G. Evolution of animal regeneration: Re-emergence of a field. Trends Ecol. Evol. 2010, 25, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Bai, H.; Zhang, W.; Qin, X.J.; Zhang, T.; Wu, H.; Liu, J.Z.; Hai, C.X. Hydrogen peroxide modulates the proliferation/quiescence switch in the liver during embryonic development and posthepatectomy regeneration. Antioxid. Redox Signal. 2015, 22, 921–937. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.K.; Freisinger, C.M.; Lebert, D.C.; Huttenlocher, A. Early redox, Src family kinase, and calcium signaling integrate wound responses and tissue regeneration in zebrafish. J. Cell. Biol. 2012, 199, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Vriz, S. Redox signalling in development and regeneration. Semin. Cell. Dev. Biol. 2018, 80, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Go, Y.M.; Chandler, J.D.; Jones, D.P. The cysteine proteome. Free Radic. Biol. Med. 2015, 84, 227–245. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, J.M.; Dick, T.P. Redox Biology on the rise. Biol. Chem. 2012, 393, 999–1004. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.P.; Sies, H. The Redox Code. Antioxid. Redox Signal. 2015, 23, 734–746. [Google Scholar] [CrossRef] [PubMed]
| Enzyme | Gene | Cellular Localization | Gene Expression | Reference |
|---|---|---|---|---|
| Catalase | cat | mitochondria, peroxisome | brain, digestive system, gill, muscle, sensory system | [86] |
| Glutaredoxin | glrx2 | cytoplasm | whole organism | [87] |
| glrx3 | cytoplasm | brain, heart, sensory system | [86,88] | |
| glrx5 | cytoplasm | blood island, digestive system, heart, sensory system | [89] | |
| Glutathione Peroxidase | gpx1a | nd | digestive system, muscle, sensory system | [90,91] |
| gpx1b | cytoplasm | digestive system, sensory system | [86,92] | |
| gpx4a | nd | digestive system, peridermis | [90,93] | |
| gpx4b | nd | blastoderm, digestive system, epidermis, epiphysis, muscle, pharyngeal arch, pronephric duct, sensory system | [90,93,94] | |
| gpx7 | nd | notochord, splanchnocranium | [86] | |
| gpx8 | membrane | notochord, pharyngeal arch, sensory system | [86] | |
| Glutathione Reductase | gsr | cytoplasm | digestive system, macrophage | [86] |
| NADPH Oxidase | nox1 | membrane | brain, spinal cord, sensory system | [79] |
| nox2/Cybb | membrane | blood, brain, spinal cord, sensory system | [79,86] | |
| nox5 | membrane | brain, spinal cord, sensory system | [79] | |
| duox | membrane | brain, digestive system, epidermis, spinal cord, sensory system, swim bladder, thyroid, | [79,95,96] | |
| Peroxiredoxin | prdx1 | cytoplasm | brain, neural crest derivatives, vessels | [97,98] |
| prdx2 | nd | blood, CNS, digestive system, pharyngeal arch, sensory system | [90] | |
| prdx3 | nd | blood, digestive system, myotome, pharyngeal arch, sensory system | [90] | |
| prdx4 | nd | digestive system, hatching gland, pharyngeal arch, sensory system | [86,99] | |
| prdx5 | nd | macrophage, pronephric duct, sensory system | [86] | |
| prdx6 | nd | digestive system, rhombomere, sensory system | [86] | |
| Superoxide Dismutase | sod1 | cytoplasm | whole organism | [86,100] |
| sod2 | mitochondria | blood, brain, digestive system, gill, kidney, muscle, sensory system | [90,101] | |
| sod3b | cytoplasm | whole organism | [102] | |
| Thioredoxin | txn | nd | digestive system, Hypophysis, spinal cord, sensory system, tegmentum | [86,91] |
| txn2 | mitochondria | whole organism | [86] | |
| Thioredoxin Reductase | txnrd3 | mitochondria | blood, CNS, digestive system, muscle, pharyngeal arch, spinal cord, sensory system | [86,90] |
| Classification | Animals/Species | Stage | ROS Detection | Profil | ROS Modulation | Organ/Appendage | ROS Targets | Reference |
|---|---|---|---|---|---|---|---|---|
| Cnidaria | Hydra | DMPO | wound edge | [180] | ||||
| Platyhelminthes | Planarian (Schmidtea mediterranea) | H2DCFDA | burst at the wound site | DPI, APO | central nervous system | neuroregeneration | [164] | |
| Arthropoda | Drosophila (Drosophila melanogaster) | Larvae | CellRox green | burst after apoptosis induction | NAC, vitamin C, Trolox, SOD, CAT | wing imaginal disc | p38 pathway | [181] |
| JNK pathway | ||||||||
| DHE, H2DCFDA | up to 24 h after apoptosis induction | misexpression of extracell.CAT | eye and wing imaginal disc | macrophages | [182] | |||
| Adult | H2DCFDA | burst after oral admin. of HgCl2 | Vitamin E | midgut | [183] | |||
| Amphibians | Xenopus (Xenopus laevis or tropicalis) | Tadpole | HyPer | production 6 h-4 dpa | DPI, APO, MCI-186 | Tail | Wnt/b-catenin pathway | [165] |
| FGF pathway | ||||||||
| DHE | nd | DPI, MCI, VAS, H2O2 | bioelectric activity | [190] | ||||
| H2DCFDA | nd | APO | Notochord in tail | acetylation of H3K9 | [188] | |||
| Squamata | Gecko (Gekko japonicus) | Adult | H2DCFDA | production (0-7 dpa), skeletal muscles | DPI, APO | tail | autophagy in skeletal muscles (ULK, MAPK) | [191] |
| Teleost fish | Zebrafish (Danio rerio) | Larvae | HyPer | nd | DPI | caudal fin | Src family kinase | [196] |
| PFBS-F | ptch1, tcf7, raldh2, pea3, ihhb | [91] | ||||||
| Adult | H2DCFDA | production 0-16 hpa | VAS2870, DPI | Apoptosis | [166] | |||
| JNK pathway | ||||||||
| H2DCFDA, HyPer | VAS2870, H2O2 | Hh pathway | [193] | |||||
| nerve | ||||||||
| Myl7:HyPer, Redox sensor cc-1 | production 3-14 dpa epicardium and adjacent myocardium, max 7 dpa | DPI, APO, CAT | heart | ERK pathway | [167] | |||
| CellRox green | production 2 h-12 hpa | VAS2870 | superficial epithelial cells (SECs) in caudal fin | [192] | ||||
| Mammals | Rat (Rattus norvegicus) | Adult | H2DCFDA, red H2O2 assay kit, amplex red H2O2 assay kit | production 1 h-3 dpa | GOX, CAT | liver | ERK pathway | [195] |
| p38 pathway | ||||||||
| Mice (Mus musculus) | Adult | luminol | MRL mice (production 0-72 h pa, max 12 h), C57Bl6 (n.s. production) | inguinal fat pad | [169] | |||
| lucigenin | n.s. production 3 h-10 dpa | ear | [168] | |||||
| luminol | production 3h-4 dpa | |||||||
| Spiny mice (Acomys cahirinus) | Adult | lucigenin | production 3 h-5 dpa | |||||
| luminol | n.s. production 3h-10 dpa |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rampon, C.; Volovitch, M.; Joliot, A.; Vriz, S. Hydrogen Peroxide and Redox Regulation of Developments. Antioxidants 2018, 7, 159. https://doi.org/10.3390/antiox7110159
Rampon C, Volovitch M, Joliot A, Vriz S. Hydrogen Peroxide and Redox Regulation of Developments. Antioxidants. 2018; 7(11):159. https://doi.org/10.3390/antiox7110159
Chicago/Turabian StyleRampon, Christine, Michel Volovitch, Alain Joliot, and Sophie Vriz. 2018. "Hydrogen Peroxide and Redox Regulation of Developments" Antioxidants 7, no. 11: 159. https://doi.org/10.3390/antiox7110159
APA StyleRampon, C., Volovitch, M., Joliot, A., & Vriz, S. (2018). Hydrogen Peroxide and Redox Regulation of Developments. Antioxidants, 7(11), 159. https://doi.org/10.3390/antiox7110159








