New Benzimidazothiazolone Derivatives as Tyrosinase Inhibitors with Potential Anti-Melanogenesis and Reactive Oxygen Species Scavenging Activities
Abstract
:1. Introduction
2. Materials and Methods
2.1. Reagents and Antibodies
2.2. General Experimental Methods
2.2.1. Mushroom Tyrosinase Inhibitory Assay
2.2.2. Enzyme Kinetic Analysis of Mushroom Tyrosinase
2.2.3. Mushroom Tyrosinase Molecular Docking Simulations
2.3. Biological Evaluation
2.3.1. Cell Culture and Viability Analysis
2.3.2. Determination of Intracellular ROS Levels
2.3.3. Melanin Contents in B16F10 Cells
2.3.4. Cellular Tyrosinase Activity Assay in B16F10 Cells
2.3.5. RNA Isolation and Quantitative Real-Time PCR (qPCR)
2.3.6. Western Blot Analysis
2.4. Statistical Analysis
3. Results and Discussion
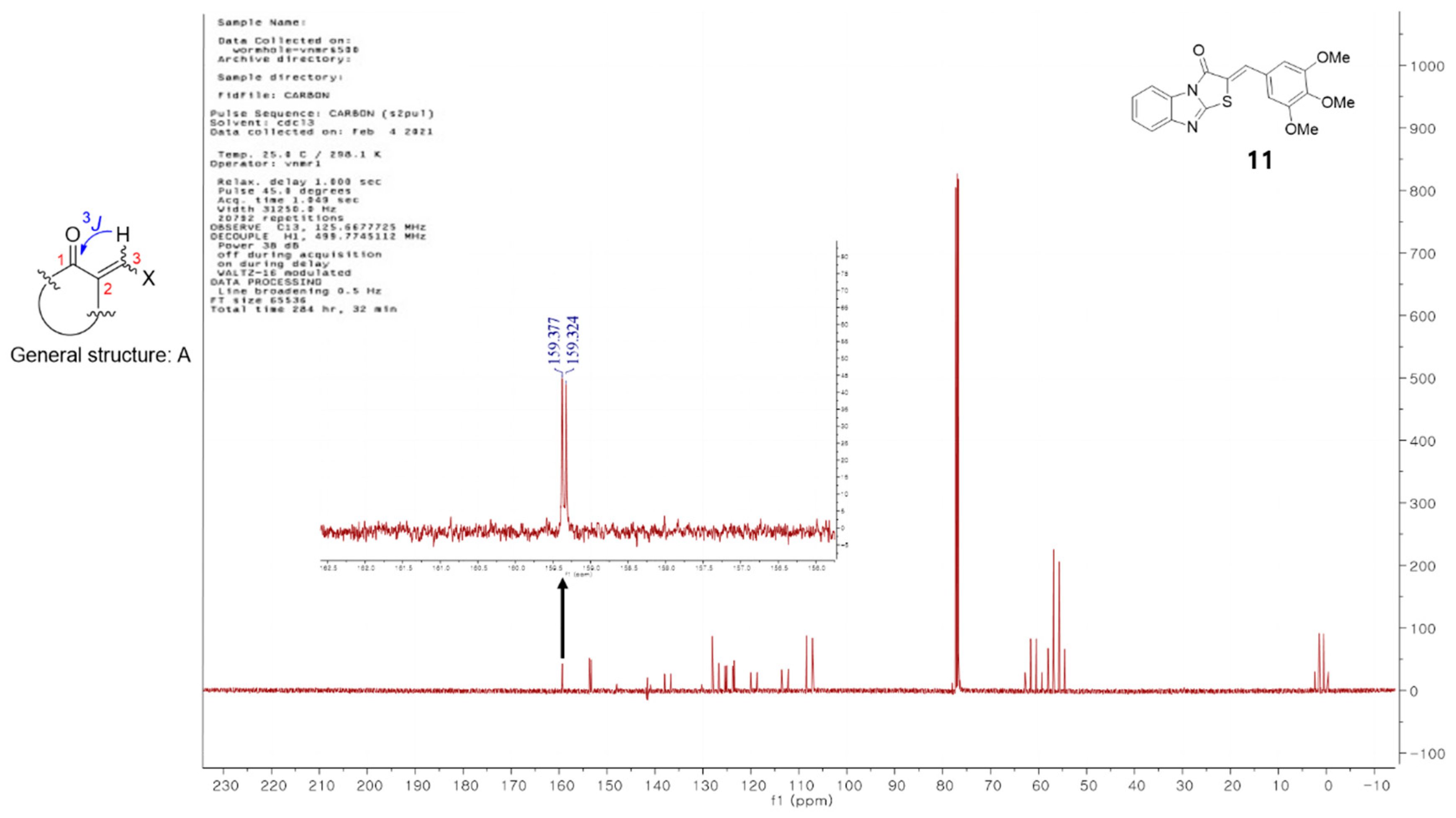
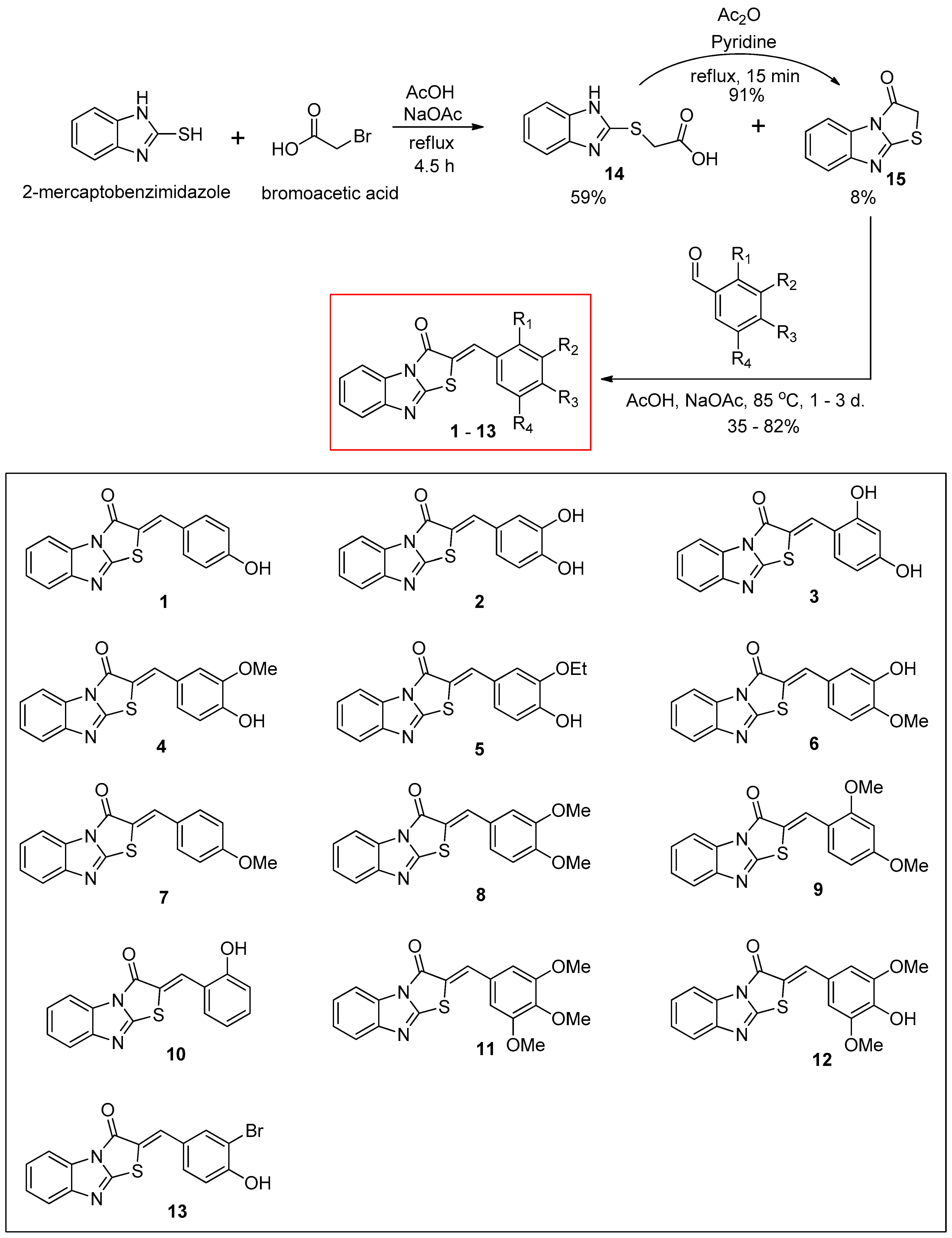
3.1. Synthesis of (Z)-2-(Substituted benzylidene)benzimidazothiazolone Derivatives 1–13
3.2. Inhibitory Activities of (Z)-2-(Substituted Benzylidene)benzimidazothiazolone Derivatives 1–13 against Mushroom Tyrosinase
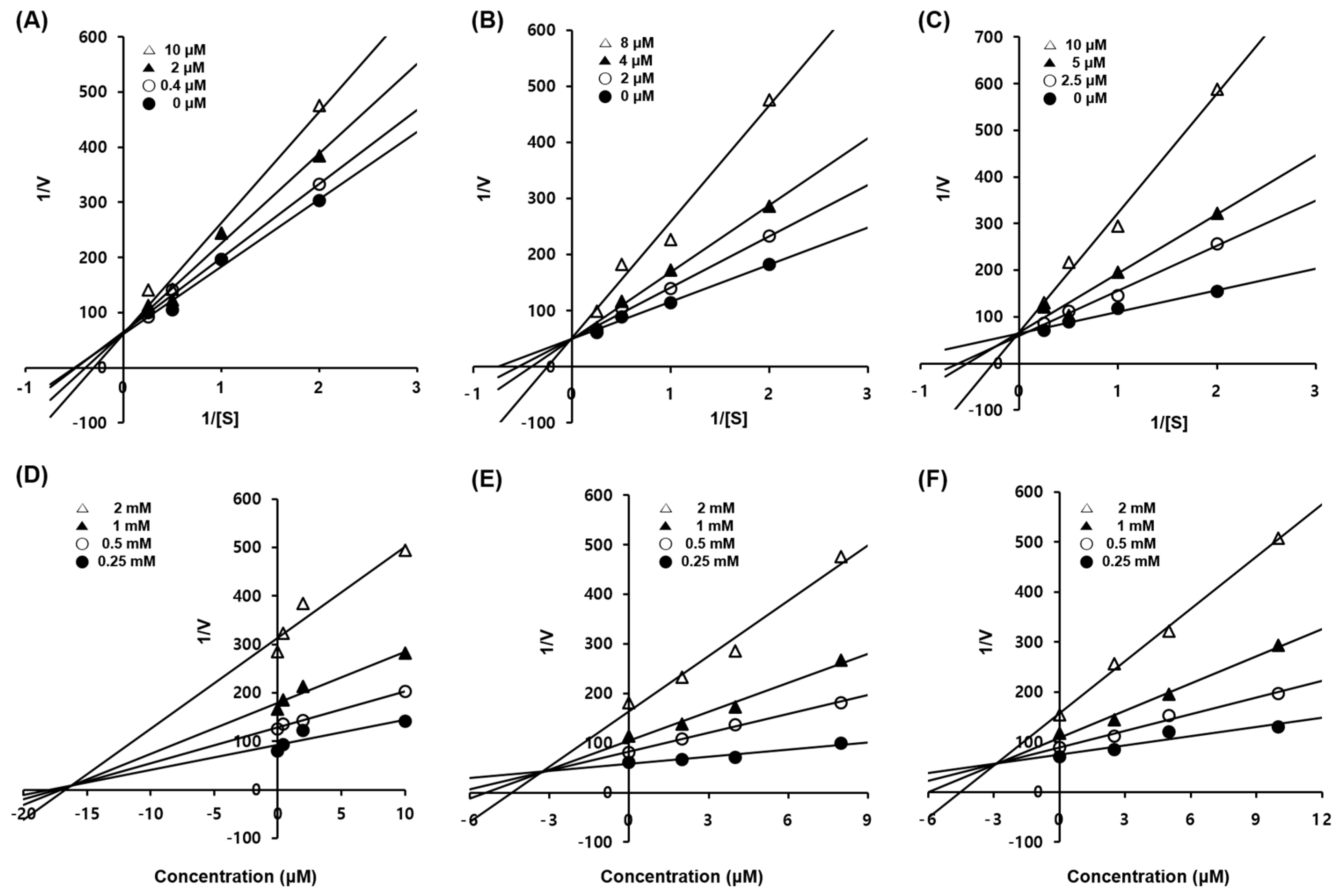
3.3. Enzyme Kinetic Studies of Compounds 1–3
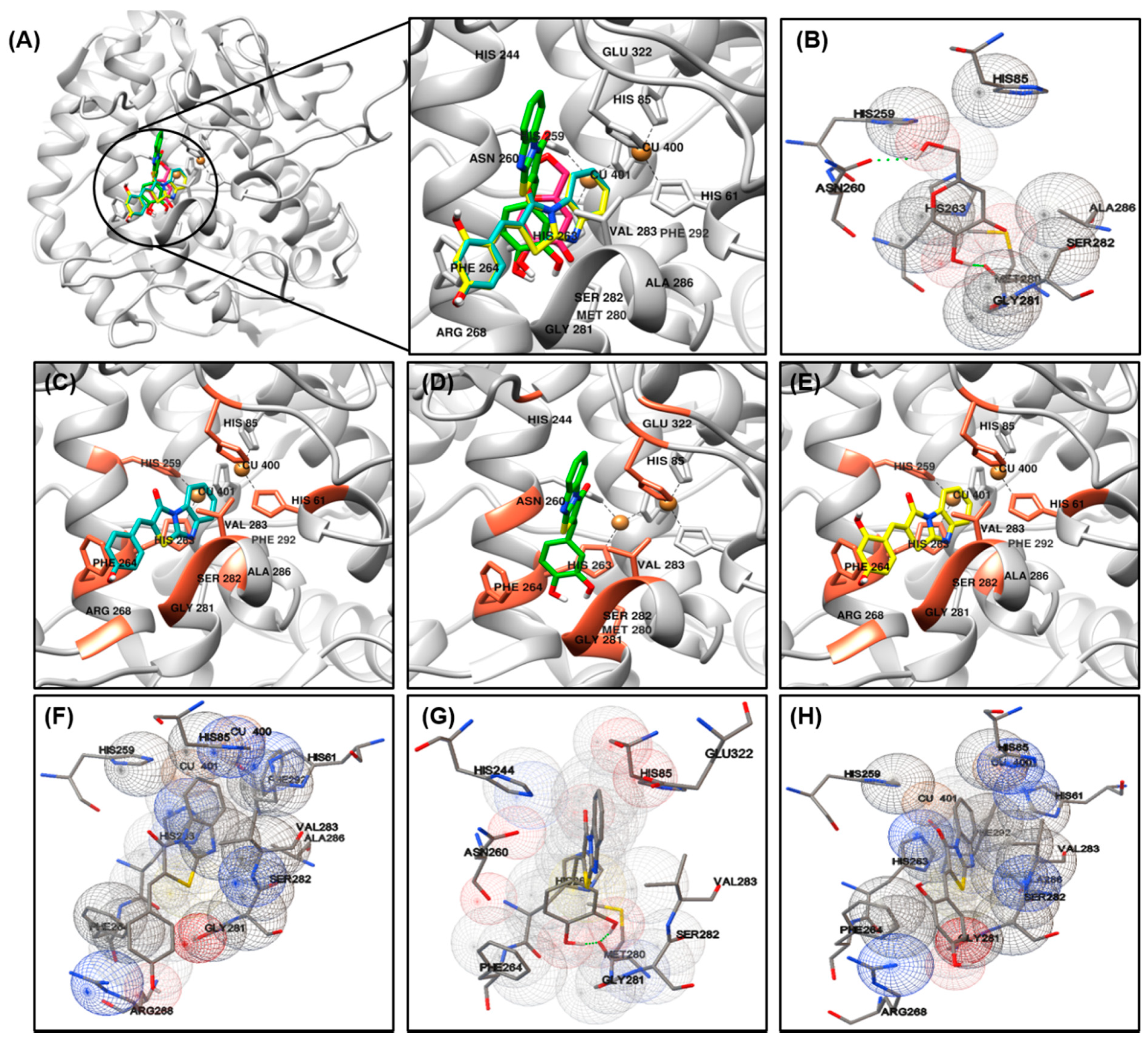
3.4. Docking Simulation and of Ligands against Tyrosinase
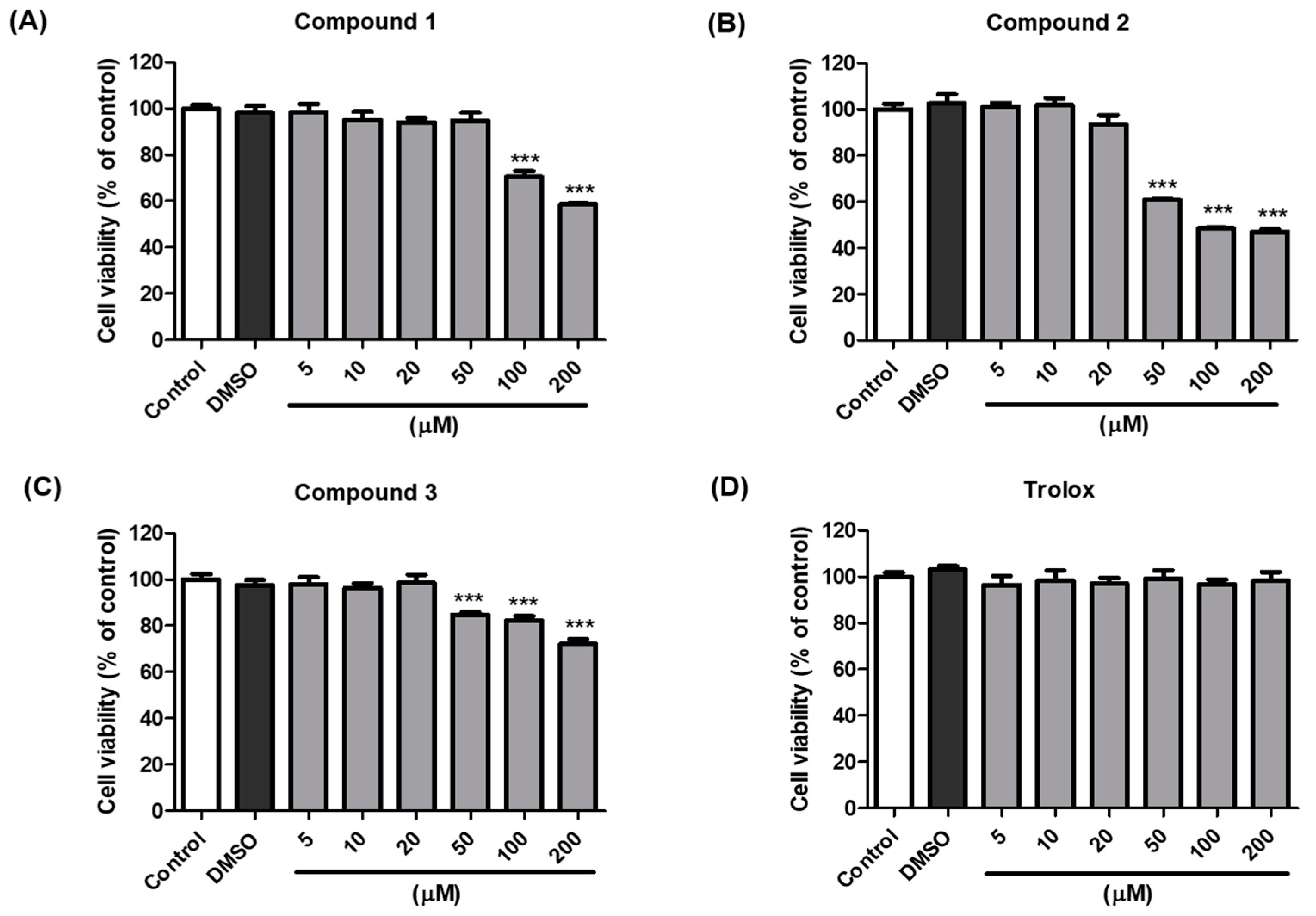
3.5. Cell Viability of Compounds 1–3
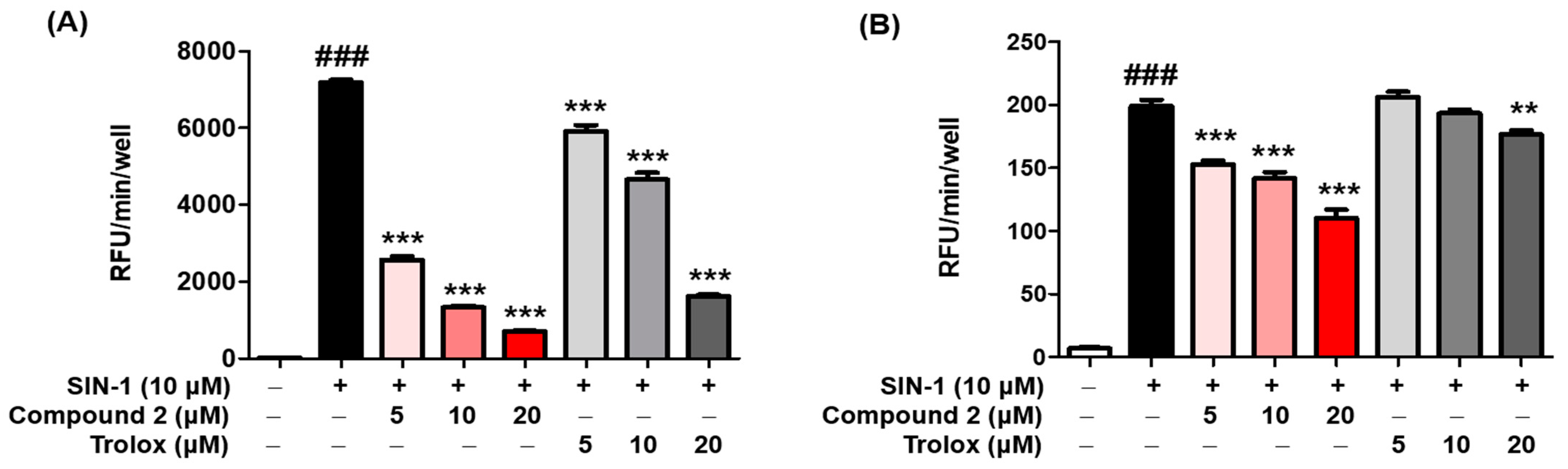
3.6. ROS Scavenging Activity
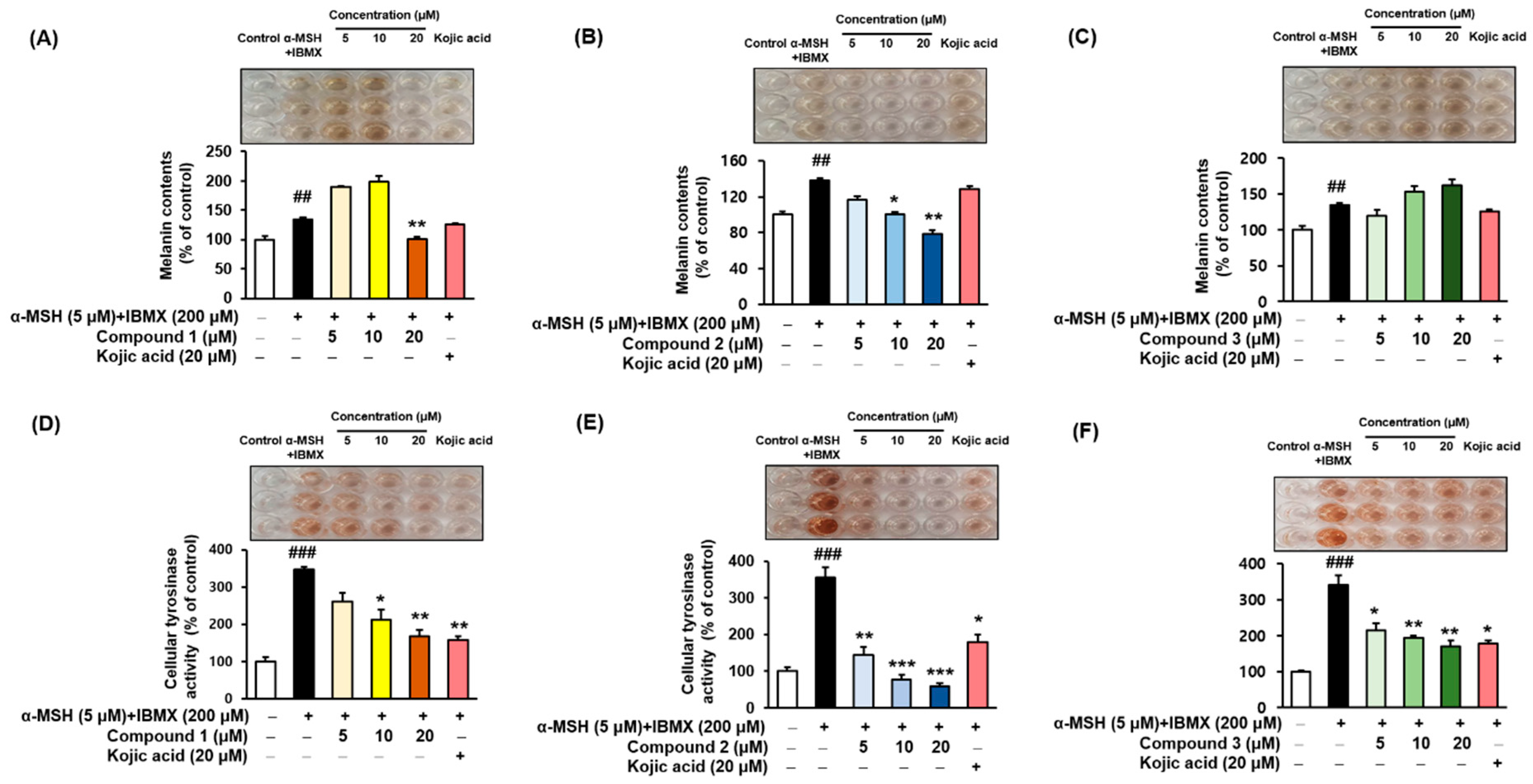
3.7. Melanin Contents and Cellular Tyrosinase Assay
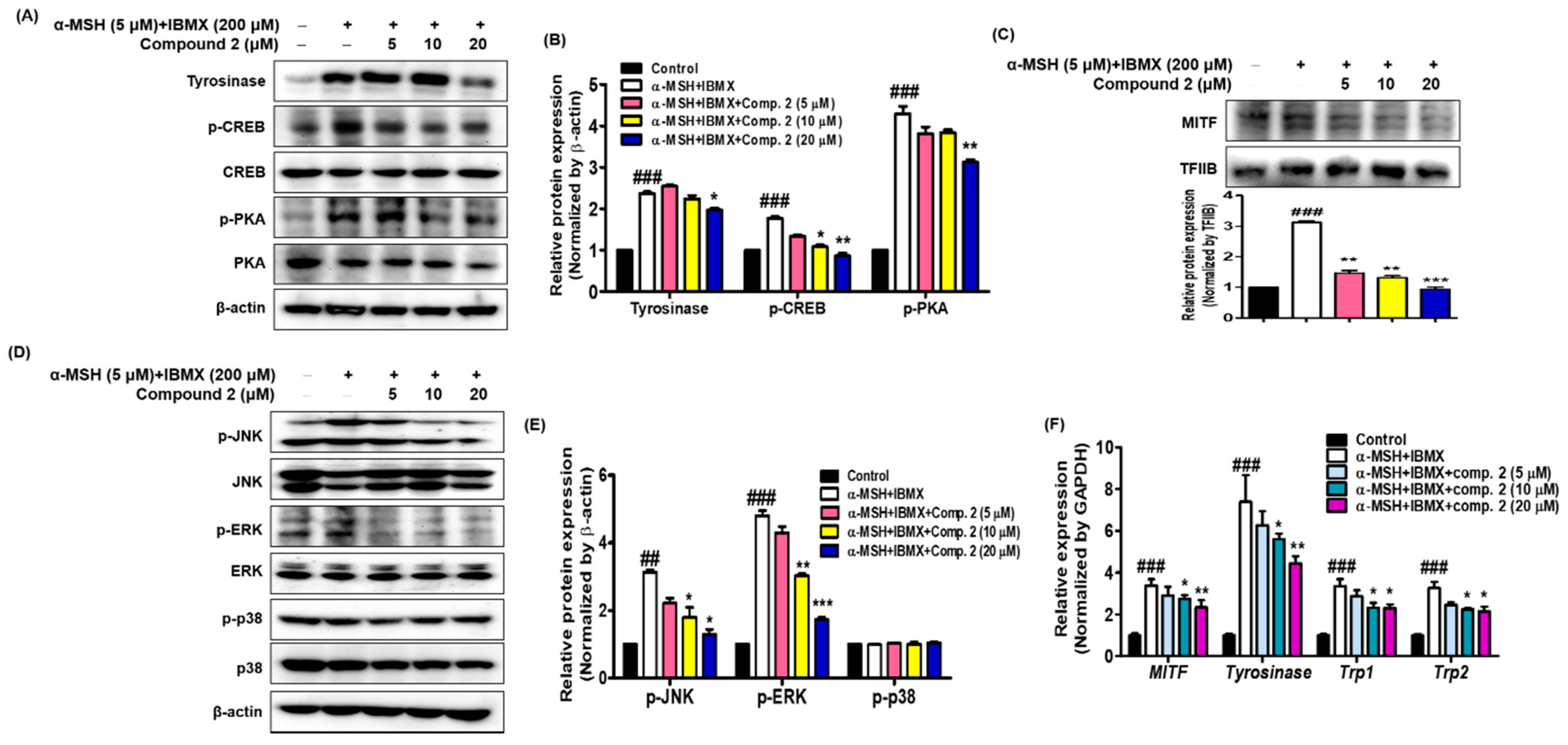
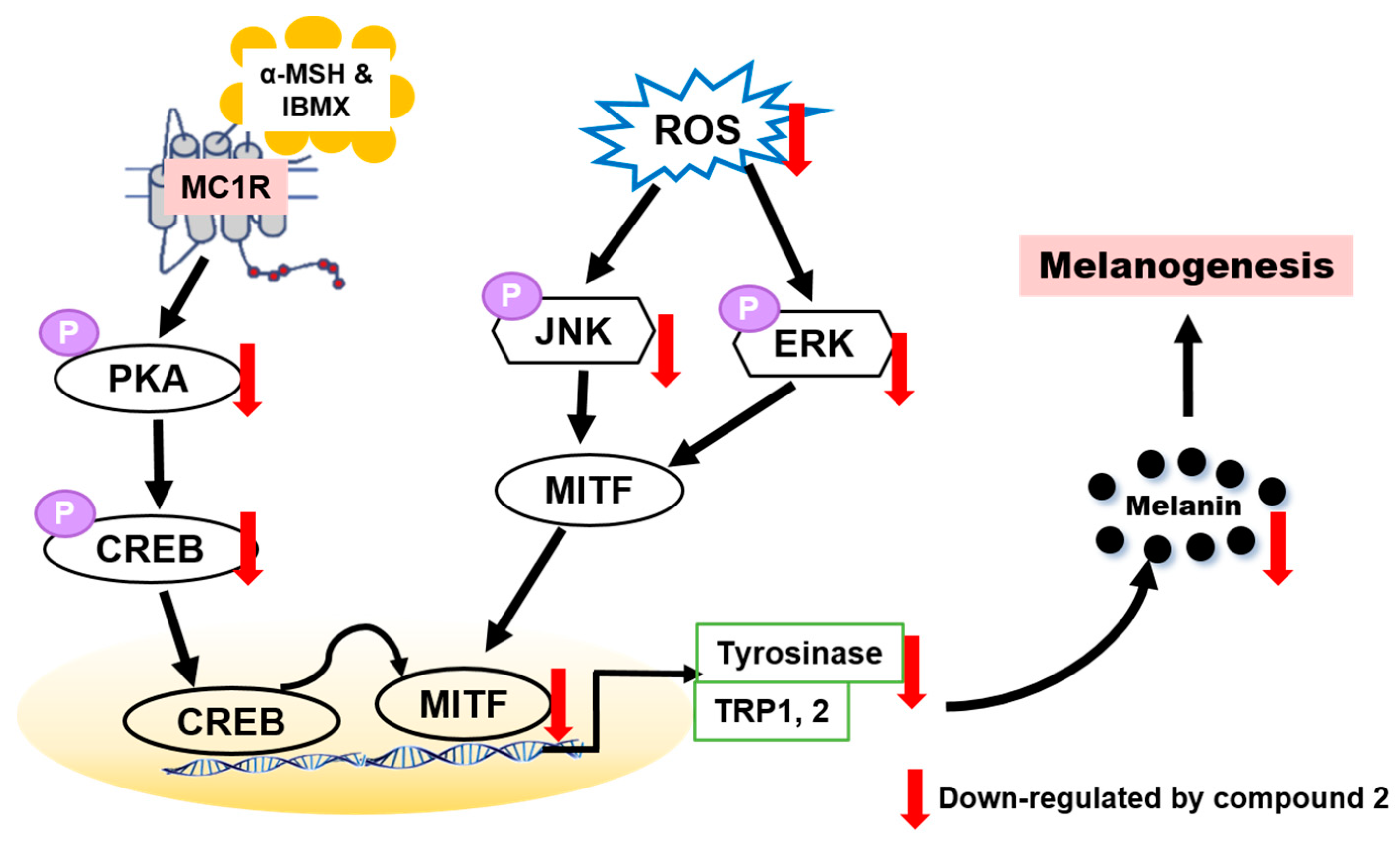
3.8. Effects of Compound 2 on Melanogenesis-Related Signaling
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Decker, H.; Tuczek, F. Tyrosinase/catecholoxidase activity of hemocyanins: Structural basis and molecular mechanism. Trends Biochem. Sci. 2000, 25, 392–397. [Google Scholar] [CrossRef]
- Kang, S.J.; Choi, B.R.; Lee, E.K.; Kim, S.H.; Yi, H.Y.; Park, H.R.; Song, C.H.; Lee, Y.J.; Ku, S.K. Inhibitory effect of dried pomegranate concentration powder on melanogenesis in B16F10 melanoma cells; involvement of p38 and PKA signaling pathways. Int. J. Mol. Sci. 2015, 16, 24219–24242. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakajima, M.; Shinoda, I.; Fukuwatary, Y.; Hayahawa, H. Arbutin increases the pigmentation of cultured human melanocytes through mechanisms other than the induction of tyrosinase activity. Pigment Cell Res. 1998, 11, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Takizawa, T.; Imai, T.; Onose, J.-I.; Ueda, M.; Tamura, T.; Mitsumori, K.; Izumi, K.; Hirose, M. Enhancement of hepatocarcinogenesis by kojic acid in rat two-stage models after initiation with N-bis(2-hydroxypropyl)nitrosamine or N-diethylnitrosamine. Toxicol. Sci. 2004, 81, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Arulmozhi, V.; Pandian, K.; Mirunalini, S. Ellagic acid encapsulated chitosan nanoparticles for drug delivery system in human oral cancer cell line (KB). Colloids Surf. B Biointerfaces 2013, 110, 313–320. [Google Scholar] [CrossRef]
- Chang, T.S. An updated review of tyrosinase inhibitors. Int. J. Mol. Sci. 2009, 10, 2440–2475. [Google Scholar] [CrossRef] [Green Version]
- Pillaiyar, T.; Manickam, M.; Namasivayam, V. Skin whitening agents: Medicinal chemistry perspective of tyrosinase inhibitors. J. Enzym. Inhib. Med. Chem. 2017, 32, 403–425. [Google Scholar] [CrossRef] [Green Version]
- Han, H.J.; Park, S.K.; Kang, J.Y.; Kim, J.M.; Yoo, S.K.; Heo, H.J. Anti-melanogenic effect of ethanolic extract of Sorghum bicolor on IBMX-induced melanogenesis in B16/F10 melanoma cells. Nutrients 2020, 12, 832. [Google Scholar] [CrossRef] [Green Version]
- Rzepka, Z.; Buszman, E.; Beberok, A.; Wrześniok, D. From tyrosine to melanin: Signaling pathways and factors regulating melanogenesis. Postepy Hig. Med. Dosw. 2016, 70, 695–708. [Google Scholar] [CrossRef]
- Park, S.Y.; Jin, M.L.; Kim, Y.H.; Kim, Y.; Lee, S.J. Aromatic-turmerone inhibits α-MSH and IBMX-induced melanogenesis by inactivating CREB and MITF signaling pathways. Arch. Dermatol. Res. 2011, 303, 737–744. [Google Scholar] [CrossRef]
- Jiménez-Cervantes, C.; Solano, F.; Kobayashi, T.; Urabe, K.; Hearing, V.J.; Lozano, J.A.; García-Borrón, J.C. A new enzymatic function in the melanogenic pathway. The 5,6-dihydroxyindole-2-carboxylic acid oxidase activity of tyrosinase-related protein-1 (TRP1). J. Biol. Chem. 1994, 269, 17993–18000. [Google Scholar] [CrossRef]
- Tachibana, M. MITF: A stream flowing for pigment cells. Pigment Cell Res. 2000, 13, 230–240. [Google Scholar] [CrossRef]
- D’Mello, S.A.; Finlay, G.J.; Baguley, B.C.; Askarian-Amiri, M.E. Signaling pathways in melanogenesis. Int. J. Mol. Sci. 2016, 7, 1144. [Google Scholar] [CrossRef] [Green Version]
- Kang, H.Y.; Suzuki, I.; Lee, D.J.; Ha, J.; Reiniche, P.; Aubert, J.; Deret, S.; Zugaj, D.; Voegel, J.J.; Ortonne, J.P. Transcriptional profiling shows altered expression of wnt pathway- and lipid metabolism-related genes as well as melanogenesis-related genes in melasma. J. Investig. Dermatol. 2011, 131, 1692–1700. [Google Scholar] [CrossRef] [Green Version]
- Bang, S.; Won, K.H.; Moon, H.R.; Yoo, H.; Hong, A.; Song, Y.; Chang, S.E. Novel regulation of melanogenesis by adiponectin via the AMPK/CRTC pathway. Pigment Cell Melanoma Res. 2017, 30, 553–557. [Google Scholar] [CrossRef] [PubMed]
- Yun, C.Y.; Ko, S.M.; Choi, Y.P.; Kim, B.J.; Lee, J.; Kim, J.M.; Kim, J.Y.; Song, J.Y.; Kim, S.H.; Hwang, B.Y.; et al. α-Viniferin improves facial hyperpigmentation via accelerating feedback termination of cAMP/PKA-signaled phosphorylation circuit in facultative melanogenesis. Theranostics 2018, 8, 2031–2043. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Finkel, T.; Holbrook, N.J. Oxidants, oxidative stress and the biology of ageing. Nature 2000, 408, 239–247. [Google Scholar] [CrossRef]
- Jung, H.J.; Kim, S.M.; Kim, D.H.; Bang, E.; Kang, D.; Lee, S.; Chun, P.; Moon, H.R.; Chung, H.Y. 2,4-Dihydroxyphenyl-benzo[d]thiazole (MHY553), a synthetic PPARα agonist, decreases age-associated inflammatory responses through PPARα activation and RS scavenging in the skin. Exp. Gerontol. 2021, 143, 111153. [Google Scholar] [CrossRef]
- Masaki, H. Role of antioxidants in the skin: Anti-aging effects. J. Dermatol. Sci. 2010, 58, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.S.; Peshavariya, H.; Higuchi, M.; Brewer, A.C.; Chang, C.W.T.; Chan, E.C.; Dusting, G.J. Microphthalmia-associated transcription factor modulates expression of NADPH oxidase type 4: A negative regulator of melanogenesis. Free Radic. Biol. Med. 2012, 52, 1835–1843. [Google Scholar] [CrossRef] [PubMed]
- Roméro-Graillet, C.; Aberdam, E.; Clément, M.; Ortonne, J.P.; Ballotti, R. Nitric oxide produced by ultraviolet-irradiated keratinocytes stimulates melanogenesis. J. Clin. Invest. 1997, 99, 635–642. [Google Scholar] [CrossRef] [Green Version]
- Sasaki, M.; Horikoshi, T.; Uchiwa, H.; Miyachi, Y. Up-regulation of tyrosinase gene by nitric oxide in human melanocytes. Pigment Cell Res. 2000, 13, 248–252. [Google Scholar] [CrossRef]
- Zhou, S.; Sakamoto, K. Pyruvic acid/ethyl pyruvate inhibits melanogenesis in B16F10 melanoma cells through PI3K/AKT, GSK3β, and ROS-ERK signaling pathways. Genes Cells 2019, 24, 60–69. [Google Scholar] [CrossRef] [Green Version]
- Miao, F.; Su, M.Y.; Jiang, S.; Luo, L.F.; Shi, Y.; Lei, T.C. Intramelanocytic acidification plays a role in the antimelanogenic and antioxidative properties of vitamin C and its derivatives. Oxid. Med. Cell. Longev. 2019, 2019, 2084805. [Google Scholar] [CrossRef] [Green Version]
- Noha, R.M.; Abdelhameid, M.K.; Ismail, M.M.; Mohammed, M.R.; Salwa, E. Design, synthesis and screening of benzimidazole containing compounds with methoxylated aryl radicals as cytotoxic molecules on (HCT-116) colon cancer cells. Eur. J. Med. Chem. 2021, 209, 112870. [Google Scholar] [CrossRef]
- Fenichel, R.L.; Alburn, H.E.; Schreck, P.A.; Bloom, R.; Gregory, F.J. Immunomodulating and antimetastatic activity of 3-(p-chlorophenyl) thiazolo[3, 2-a]benzimidazole-2-acetic acid (Wy-18, 251, Nsc 310633). J. Immunopharmacol. 1980, 2, 491–508. [Google Scholar] [CrossRef] [PubMed]
- El-Kerdawy, M.M.; Ghaly, M.A.; Darwish, S.A.; Abdel-Aziz, H.A.; Elsheakh, A.R.; Abdelrahman, R.S.; Hassan, G.S. New benzimidazothiazole derivatives as anti-inflammatory, antitumor active agents: Synthesis, in-vitro and in-vivo screening and molecular modeling studies. Bioorg. Chem. 2019, 83, 250–261. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.R.; Lee, H.J.; Choi, Y.J.; Park, Y.J.; Woo, Y.; Kim, S.J.; Park, M.H.; Lee, H.W.; Chun, P.; Chung, H.Y.; et al. Benzylidene-linked thiohydantoin derivatives as inhibitors of tyrosinase and melanogenesis: Importance of the β-phenyl-α,β-unsaturated carbonyl functionality. MedChemComm 2014, 5, 1410–1417. [Google Scholar] [CrossRef]
- Ullah, S.; Son, S.; Yun, H.Y.; Kim, D.H.; Chun, P.; Moon, H.R. Tyrosinase inhibitors: A patent review (2011–2015). Expert Opin. Ther. Pat. 2016, 26, 347–362. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.J.; Lee, M.J.; Park, Y.J.; Noh, S.G.; Lee, A.K.; Moon, K.M.; Lee, E.K.; Bang, E.J.; Park, Y.J.; Kim, S.J.; et al. A novel synthetic compound, (Z)-5-(3-hydroxy-4-methoxybenzylidene)-2-iminothiazolidin-4-one (MHY773) inhibits mushroom tyrosinase. Biosci. Biotechnol. Biochem. 2018, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.J.; Yang, J.; Lee, S.; Park, C.; Kang, D.; Akter, J.; Ullah, S.; Kim, Y.J.; Chun, P.; Moon, H.R. The tyrosinase inhibitory effects of isoxazolone derivatives with a (Z)-β-phenyl-α, β-unsaturated carbonyl scaffold. Bioorg. Med. Chem. 2018, 26, 3882–3889. [Google Scholar] [CrossRef]
- Hyun, S.K.; Lee, W.-H.; Jeong, D.M.; Kim, Y.; Choi, J.S. Inhibitory effects of kurarinol, kuraridinol, and trifolirhizin from Sophora flavescens on tyrosinase and melanin synthesis. Biol. Pharm. Bull. 2008, 31, 154–158. [Google Scholar] [CrossRef] [Green Version]
- Lineweaver, H.; Burk, D. The determination of enzyme dissociation constants. J. Am. Chem. Soc. 1934, 56, 658–666. [Google Scholar] [CrossRef]
- Cornish-Bowden, A. A simple graphical method for determining the inhibition constants of mixed, uncompetitive and non-competitive inhibitors. Biochem. J. 1974, 137, 143–144. [Google Scholar] [CrossRef]
- Ismaya, W.T.; Rozeboom, H.J.; Weijn, A.; Mes, J.J.; Fusetti, F.; Wichers, H.J.; Dijkstra, B.W. Crystal structure of Agaricus bisporus mushroom tyrosinase: Identity of the tetramer subunits and interaction with Tropolone. Biochemistry 2011, 50, 5477–5486. [Google Scholar] [CrossRef] [Green Version]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- No, J.K.; Soung, D.Y.; Kim, Y.J.; Shim, K.H.; Jun, Y.S.; Rhee, S.H.; Yokozawa, T.; Chung, H.Y. Inhibition of tyrosinase by green tea components. Life Sci. 1999, 65, PL241–PL246. [Google Scholar] [CrossRef]
- Ali, S.F.; LeBel, C.P.; Bondy, S.C. Reactive oxygen species formation as a biomarker of methylmercury and trimethyltin neurotoxicity. Neurotoxicology 1992, 13, 637–648. [Google Scholar]
- LeBel, C.P.; Bondy, S.C. Sensitive and rapid quantitation of oxygen reactive species formation in rat synaptosomes. Neurochem. Int. 1990, 17, 435–440. [Google Scholar] [CrossRef] [Green Version]
- Tsuboi, T.; Kondoh, H.; Hiratsuka, J.; Mishima, Y. Enhanced melanogenesis induced by tyrosinase gene-transfer increases boron-uptake and killing effect of boron neutron capture therapy for amelanotic melanoma. Pigment Cell Res. 1998, 11, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.G.; Chang, W.L.; Lee, C.J.; Lee, L.T.; Shih, C.M.; Wang, C.C. Melanogenesis inhibition by gallotannins from Chinese galls in B16 mouse melanoma cells. Biol. Pharm. Bull. 2009, 32, 1447–1452. [Google Scholar] [CrossRef] [Green Version]
- Vögeli, U.; von Philipsborn, W.; Nagarajan, K.; Nair, M.D. Structures of addition products of acetylenedicarboxylic acid esters with various dinucleophiles. An application of C, H-spin-coupling constants. Helv. Chim. Acta 1987, 61, 607–617. [Google Scholar] [CrossRef]
- Ashooriha, M.; Khoshneviszadeh, M.; Khoshneviszadeh, M.; Rafiei, A.; Kardan, M.; Yazdian-Robati, R.; Emami, S. Kojic acid-natural product conjugates as mushroom tyrosinase inhibitors. Eur. J. Med. Chem. 2020, 201, 112480. [Google Scholar] [CrossRef]
- Larik, F.A.; Saeed, A.; Channar, P.A.; Muqadar, U.; Abbas, Q.; Hassan, M.; Seo, S.Y.; Bolte, M. Design, synthesis, kinetic mechanism and molecular docking studies of novel 1-pentanoyl-3-arylthioureas as inhibitors of mushroom tyrosinase and free radical scavengers. Eur. J. Med. Chem. 2017, 141, 273–281. [Google Scholar] [CrossRef] [PubMed]
- Choi, I.; Park, Y.; Ryu, I.Y.; Jung, H.J.; Ullah, S.; Choi, H.; Park, C.; Kang, D.; Lee, S.; Chun, P.; et al. In silico and in vitro insights into tyrosinase inhibitors with a 2-thioxooxazoline-4-one template. Comput. Struct. Biotechnol. J. 2021, 19, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Chung, K.W.; Jeong, H.O.; Jang, E.J.; Choi, Y.J.; Kim, D.H.; Kim, S.R.; Lee, K.J.; Lee, H.J.; Chun, P.; Byun, Y.; et al. Characterization of a small molecule inhibitor of melanogenesis that inhibits tyrosinase activity and scavenges nitric oxide (NO). Biochim. Biophys. Acta 2013, 1830, 4752–4761. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [Green Version]
- Heitz, M.P.; Rupp, J.W. Determining mushroom tyrosinase inhibition by imidazolium ionic liquids: A spectroscopic and molecular docking study. Int. J. Biol. Macromol. 2018, 107, 1971–1981. [Google Scholar] [CrossRef]
- Battaini, G.; Monzani, E.; Casella, L.; Santagostini, L.; Pagliarin, R. Inhibition of the catecholase activity of biomimetic dinuclear copper complexes by kojic acid. J. Biol. Inorg. Chem. 2000, 5, 262–268. [Google Scholar] [CrossRef]
- Ujan, R.; Saeed, A.; Ashraf, S.; Channar, P.A.; Abbas, Q.; Rind, M.A.; Hassan, M.; Raza, H.; Seo, S.Y.; El-Seedi, H.R. Synthesis, computational studies and enzyme inhibitory kinetics of benzothiazole-linked thioureas as mushroom tyrosinase inhibitors. J. Biomol. Struct. Dyn. 2020, 1–9. [Google Scholar] [CrossRef]
- Xiong, S.L.; Lim, G.T.; Yin, S.J.; Lee, J.; Si, Y.X.; Yang, J.M.; Park, Y.D.; Qian, G.Y. The inhibitory effect of pyrogallol on tyrosinase activity and structure: Integration study of inhibition kinetics with molecular dynamics simulation. Int. J. Biol. Macromol. 2019, 121, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Saeed, A.; Mahesar, P.A.; Channar, P.A.; Abbas, Q.; Larik, F.A.; Hassan, M.; Raza, H.; Seo, S.Y. Synthesis, molecular docking studies of coumarinyl-pyrazolinyl substituted thiazoles as non-competitive inhibitors of mushroom tyrosinase. Bioorg. Chem. 2017, 74, 187–196. [Google Scholar] [CrossRef]
- Raza, H.; Abbasi, M.A.; Siddiqui, S.Z.; Hassan, M.; Abbas, Q.; Hong, H.; Shah, S.A.A.; Shahid, M.; Seo, S.Y. Synthesis, molecular docking, dynamic simulations, kinetic mechanism, cytotoxicity evaluation of N-(substituted-phenyl)-4-{(4-[(E)-3-phenyl-2-propenyl]-1-piperazinyl} butanamides as tyrosinase and melanin inhibitors: In vitro, in vivo and in silico approaches. Bioorg. Chem. 2020, 94, 103445. [Google Scholar]
- Pelle, E.; Mammone, T.; Maes, D.; Frenkel, K. Keratinocytes act as a source of reactive oxygen species by transferring hydrogen peroxide to melanocytes. J. Investig. Dermatol. 2005, 124, 793–797. [Google Scholar] [CrossRef] [Green Version]
- Hu, S.; Huang, J.; Pei, S.; Ouyang, Y.; Ding, Y.; Jiang, L.; Lu, J.; Kang, L.; Huang, L.; Xiang, H.; et al. Ganoderma lucidum polysaccharide inhibits UVB-induced melanogenesis by antagonizing cAMP/PKA and ROS/MAPK signaling pathways. J. Cell. Physiol. 2019, 234, 7330–7340. [Google Scholar] [CrossRef]
- Caldefie-Chézet, F.; Walrand, S.; Moinard, C.; Tridon, A.; Chassagne, J.; Vasson, M.P. Is the neutrophil reactive oxygen species production measured by luminol and lucigenin chemiluminescence intra or extracellular? Comparison with DCFH-DA flow cytometry and cytochrome c reduction. Clin. Chim. Acta 2002, 319, 9–17. [Google Scholar] [CrossRef]
- Becquet, F.; Courtois, Y.; Goureau, O. Nitric oxide decreases in vitro phagocytosis of photoreceptor outer segments by bovine retinal pigmented epithelial cells. J. Cell. Physiol. 1994, 159, 256–262. [Google Scholar] [CrossRef] [PubMed]
- Feelisch, M.; Ostrowski, J.; Noack, E. On the mechanism of NO release from sydnonimines. J. Cardiovasc. Pharmacol. 1989, 14, S13–S22. [Google Scholar] [CrossRef] [Green Version]
- Anouar, E. A Quantum chemical and statistical study of phenolic schiff bases with antioxidant activity against DPPH free radical. Antioxidants 2014, 3, 309–322. [Google Scholar] [CrossRef] [Green Version]
- Marc, G.; Atana, A.; Oniga, S.D.; Pirnau, A.; Vlase, L.; Oniga, O. New phenolic derivatives of thiazolidine-2,4-dione with antioxidant and antiradical properties: Synthesis, characterization, in vitro evaluation, and quantum studies. Molecules 2019, 24, 2060. [Google Scholar] [CrossRef] [Green Version]
- Corre, S.; Primot, A.; Sviderskaya, E.; Bennett, D.C.; Vaulont, S.; Goding, C.R.; Galibert, M.D. UV-induced expression of key component of the tanning process, the POMC and MC1R genes, is dependent on the p-38-activated upstream stimulating factor-1 (USF-1). J. Biol. Chem. 2004, 279, 51226–51233. [Google Scholar] [CrossRef] [Green Version]
- Oh, T.I.; Jung, H.J.; Lee, Y.M.; Lee, S.; Kim, G.H.; Kan, S.Y.; Kang, H.; Oh, T.; Ko, H.M.; Kwak, K.C.; et al. Zerumbone, a tropical ginger sesquiterpene of Zingiber officinale Roscoe, attenuates α-MSH-induced melanogenesis in B16F10 Cells. Int. J. Mol. Sci. 2018, 19, 3149. [Google Scholar] [CrossRef] [Green Version]
- Sun, L.; Guo, Y.; Zhang, Y.; Zhuang, Y. Antioxidant and anti-tyrosinase activities of phenolic extracts from rape bee pollen and inhibitory melanogenesis by cAMP/MITF/TYR pathway in B16 mouse melanoma cells. Front. Pharmacol. 2017, 8, 104. [Google Scholar] [CrossRef] [Green Version]
- Hirata, N.; Naruto, S.; Ohguchi, K.; Akao, Y.; Nozawa, Y.; Iinuma, M.; Matsuda, H. Mechanism of the melanogenesis stimulation activity of (−)-cubebin in murine B16 melanoma cells. Bioorg. Med. Chem. 2007, 15, 4897–4902. [Google Scholar] [CrossRef]
- Dong, Y.; Cao, J.; Wang, H.; Zhang, J.; Zhu, Z.; Bai, R.; Hao, H.; He, X.; Fan, R.; Dong, C. Nitric oxide enhances the sensitivity of alpaca melanocytes to respond to alpha-melanocyte-stimulating hormone by up-regulating melanocortin-1 receptor. Biochem. Biophys. Res. Commun. 2010, 396, 849–853. [Google Scholar] [CrossRef] [PubMed]
- Horikoshi, T.; Nakahara, M.; Kaminaga, H.; Sasaki, M.; Uchiwa, H.; Miyachi, Y. Involvement of nitric oxide in UVB-induced pigmentation in guinea pig skin. Pigment Cell Res. 2000, 13, 358–363. [Google Scholar] [CrossRef]
- Smalley, K.; Eisen, T. The involvement of p38 mitogen-activated protein kinase in the α-melanocyte stimulating hormone (α-MSH)-induced melanogenic and anti-proliferative effects in B16 murine melanoma cells. FEBS Lett. 2000, 476, 198–202. [Google Scholar] [CrossRef] [Green Version]
- Park, H.Y.; Kosmadaki, M.; Yaar, M.; Gilchrest, B.A. Cellular mechanisms regulating human melanogenesis. Cell. Mol. Life Sci. 2009, 66, 1493–1506. [Google Scholar] [CrossRef]
- Videira, I.F.; Moura, D.F.; Magina, S. Mechanisms regulating melanogenesis. An. Bras. Dermatol. 2013, 88, 76–83. [Google Scholar] [CrossRef] [Green Version]
- Hsu, K.D.; Chen, H.J.; Wang, C.S.; Lum, C.C.; Wu, S.P.; Lin, S.P.; Cheng, K.C. Extract of Ganoderma formosamum mycelium as a highly potent tyrosinase inhibitor. Sci. Rep. 2016, 6, 32584. [Google Scholar] [CrossRef]
| Gene | Ensembl Version a | Primers Sequence (Forward; 5′→3′, Reverse; 3′→5′) |
|---|---|---|
| MITF | ENSMUSG00000035158.16 | Forward = GGAACAGCAACGAGCTAAGG Reverse = TGATGATCCGATTCACCAGA |
| Tyrosinase | ENSMUSG00000004651.7 | Forward = GGAACAGCAACGAGCTAAGG Reverse = TGATGATCCGATTCACCAGA |
| TRP1 | ENSMUSG00000005994.15 | Forward = GGAACAGCAACGAGCTAAGG Reverse = TGATGATCCGATTCACCAGA |
| TRP2 | ENSMUSG00000022129.5 | Forward = GGAACAGCAACGAGCTAAGG Reverse = TGATGATCCGATTCACCAGA |
| GAPDH | ENSMUSG00000057666.19 | Forward = GGAACAGCAACGAGCTAAGG Reverse = TGATGATCCGATTCACCAGA |
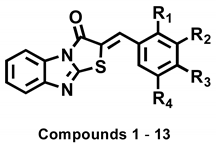
| Compounds | R1 | R2 | R3 | R4 | IC50 Values a (µM) |
|---|---|---|---|---|---|
| 1 | H | H | OH | H | 3.70 ± 0.51 |
| 2 | H | OH | OH | H | 3.05 ± 0.95 |
| 3 | OH | H | OH | H | 5.00 ± 0.38 |
| 4 | H | OMe | OH | H | 42.06 ± 4.28 |
| 5 | H | OEt | OH | H | 21.03 ± 0.82 |
| 6 | H | OH | OMe | H | NI b |
| 7 | H | H | OMe | H | NI |
| 8 | H | OMe | OMe | H | 41.40 ± 0.68 |
| 9 | OMe | H | OMe | H | NI |
| 10 | OH | H | H | H | 27.96 ± 1.32 |
| 11 | H | OMe | OMe | OMe | 34.10 ± 4.06 |
| 12 | H | OMe | OH | OMe | 53.55 ± 4.71 |
| 13 | H | Br | OH | H | 34.60 ± 2.32 |
| Kojic acid c | 18.27 ± 0.89 |
| Compounds | Inhibition Type a | Kic Values (µM) b |
|---|---|---|
| 1 | Competitive | 16.55 |
| 2 | Competitive | 3.21 |
| 3 | Competitive | 3.01 |
| Kojic acid c | – | – |
| Compounds | Binding Energy (kcal/mol) a | Interaction with Amino Acid Residues b | |
|---|---|---|---|
| AutoDock 4.2 | AutoDock Vina | ||
| 1 | –6.73 | –6.3 | HIS61, HIS85, HIS259, HIS263, PHE264, ARG268, GLY281, SER282, VAL283, ALA286, PHE292. |
| 2 | –6.92 | –6.5 | HIS85, HIS244, ASN260, HIS263, PHE264, MET280, GLY281, SER282, VAL283, GLU322. |
| 3 | –6.35 | –6.4 | HIS61, HIS85, HIS259, HIS263, PHE264, ARG268, GLY281, SER282, VAL283, ALA286, PHE292. |
| Kojic acid c | –4.21 | –5.6 | HIS85, HIS259, ASN260, HIS263, MET280, GLY281, SER282, ALA286. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jung, H.J.; Choi, D.C.; Noh, S.G.; Choi, H.; Choi, I.; Ryu, I.Y.; Chung, H.Y.; Moon, H.R. New Benzimidazothiazolone Derivatives as Tyrosinase Inhibitors with Potential Anti-Melanogenesis and Reactive Oxygen Species Scavenging Activities. Antioxidants 2021, 10, 1078. https://doi.org/10.3390/antiox10071078
Jung HJ, Choi DC, Noh SG, Choi H, Choi I, Ryu IY, Chung HY, Moon HR. New Benzimidazothiazolone Derivatives as Tyrosinase Inhibitors with Potential Anti-Melanogenesis and Reactive Oxygen Species Scavenging Activities. Antioxidants. 2021; 10(7):1078. https://doi.org/10.3390/antiox10071078
Chicago/Turabian StyleJung, Hee Jin, Dong Chan Choi, Sang Gyun Noh, Heejeong Choi, Inkyu Choi, Il Young Ryu, Hae Young Chung, and Hyung Ryong Moon. 2021. "New Benzimidazothiazolone Derivatives as Tyrosinase Inhibitors with Potential Anti-Melanogenesis and Reactive Oxygen Species Scavenging Activities" Antioxidants 10, no. 7: 1078. https://doi.org/10.3390/antiox10071078
APA StyleJung, H. J., Choi, D. C., Noh, S. G., Choi, H., Choi, I., Ryu, I. Y., Chung, H. Y., & Moon, H. R. (2021). New Benzimidazothiazolone Derivatives as Tyrosinase Inhibitors with Potential Anti-Melanogenesis and Reactive Oxygen Species Scavenging Activities. Antioxidants, 10(7), 1078. https://doi.org/10.3390/antiox10071078






