Differential Proteomics Analysis Unraveled Mechanisms of Arma chinensis Responding to Improved Artificial Diet
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
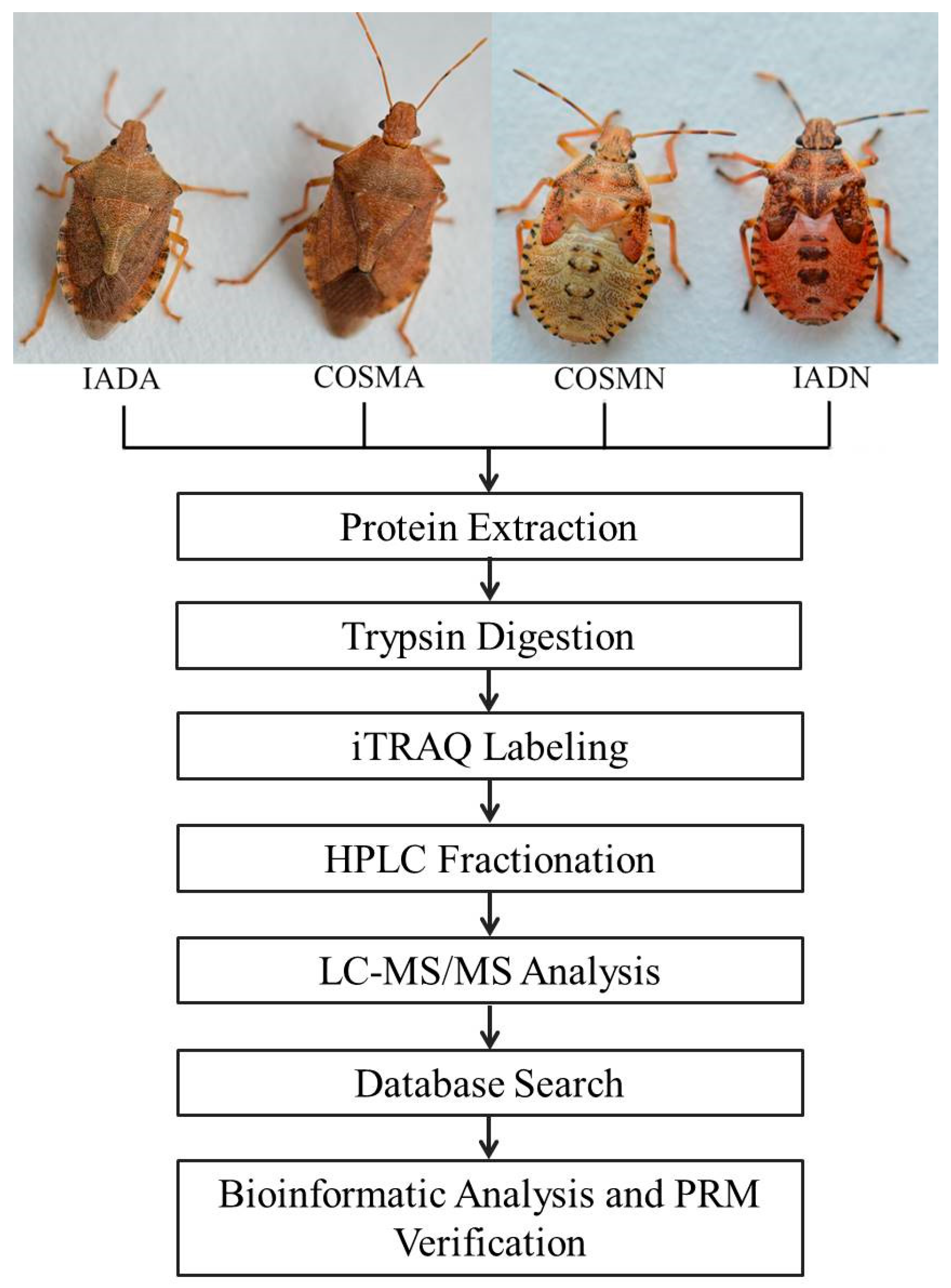
2.1. Experimental A. chinensis
2.2. Protein Extraction
2.3. Protein Digestion and iTRAQ Labeling
2.4. HPLC Fractionation
2.5. LC-MS/MS Analysis
2.6. Database Search
2.7. Bioinformatics and Statistical Analysis
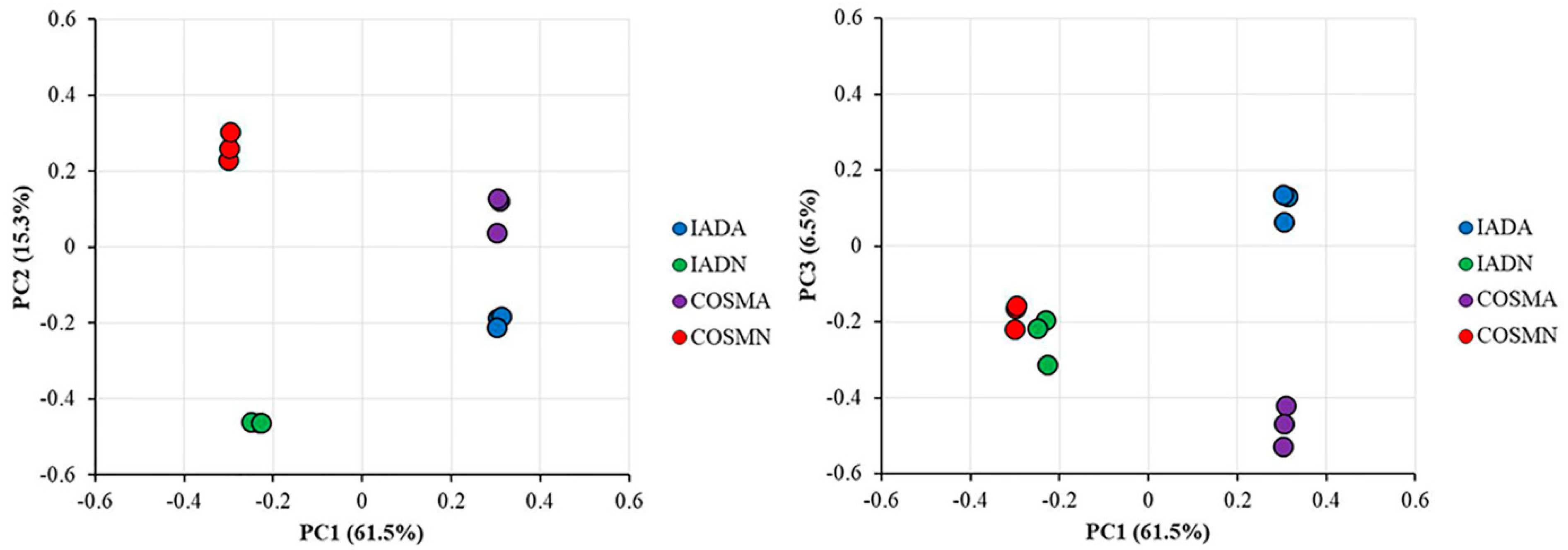
2.8. Principal Component Analysis
2.9. PRM Assays
3. Results and Discussion
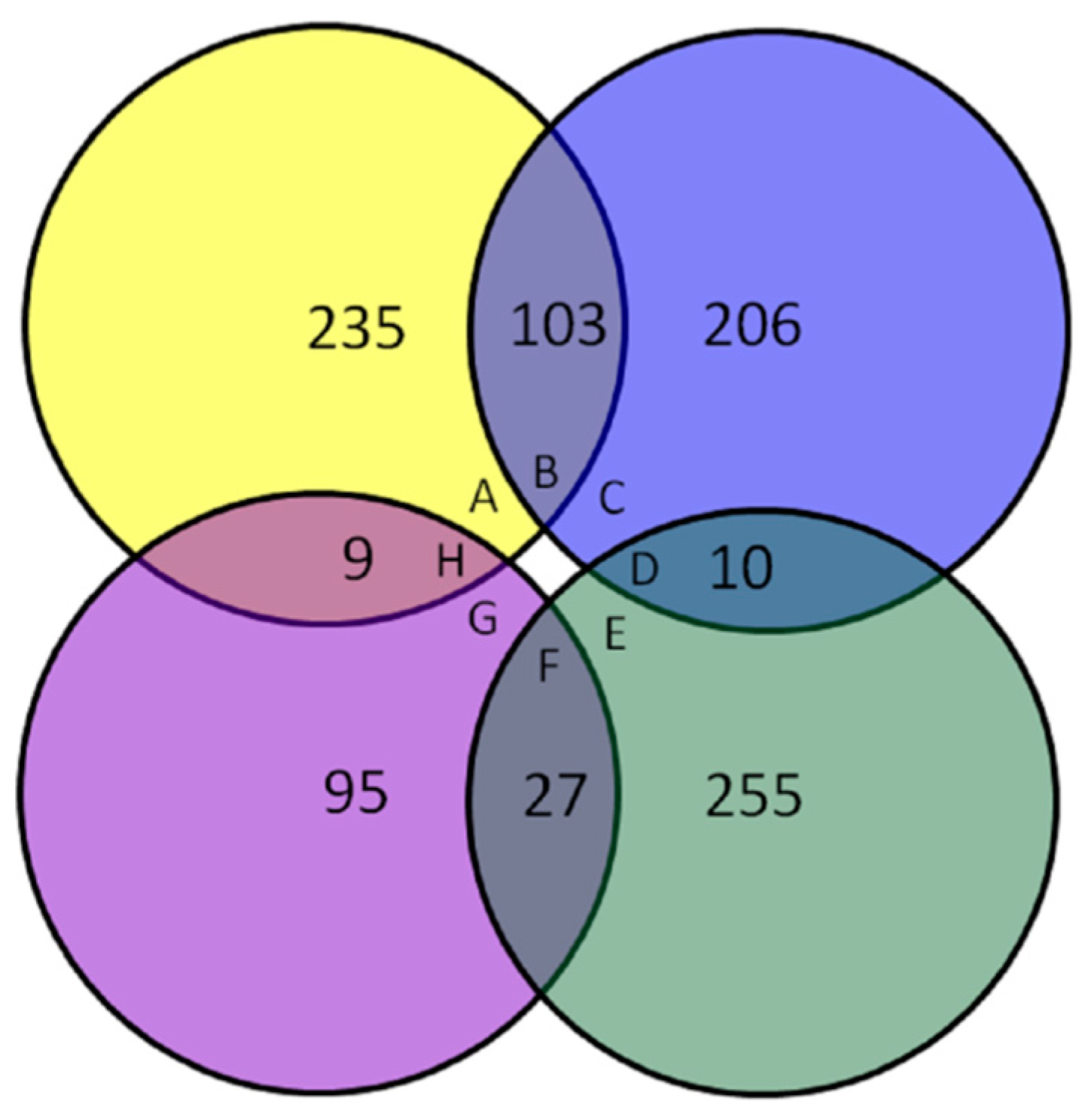
3.1. Protein Identification and Quantification
3.2. Expression Profile of Differentially Expressed Proteins
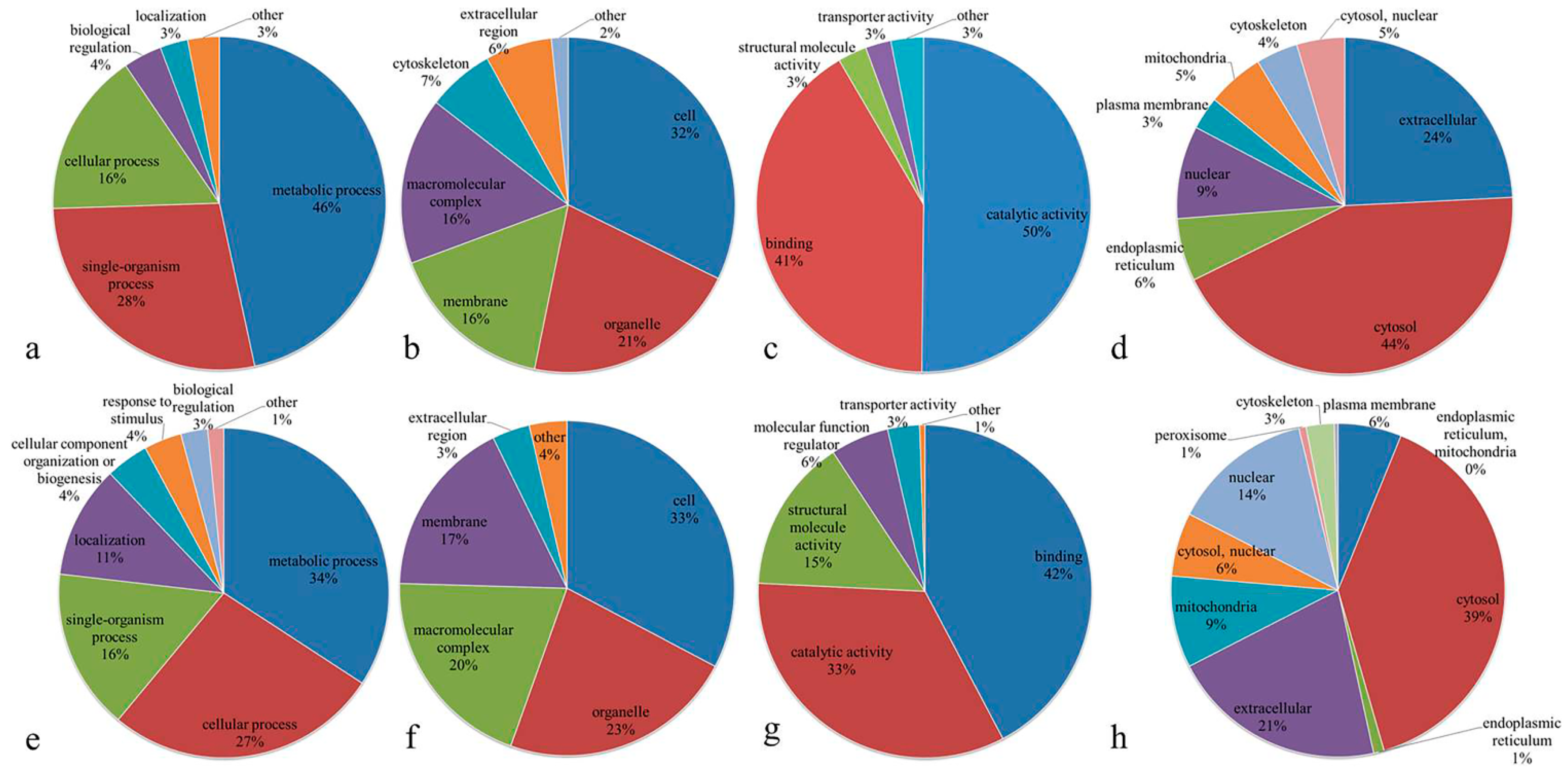
3.3. Functional Classification of Differentially Expressed Proteins
3.4. Functional Enrichment of Differentially Expressed Proteins
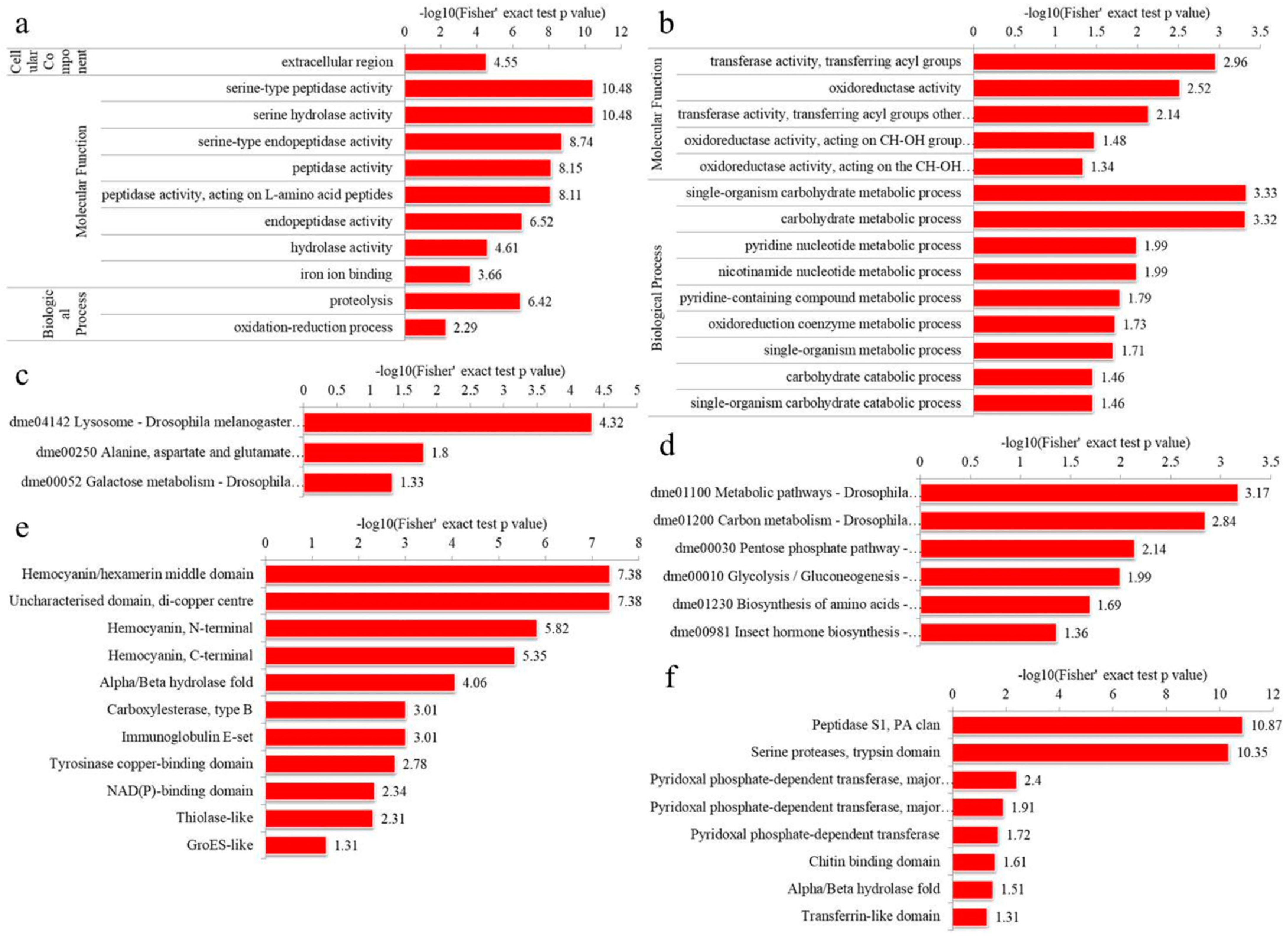
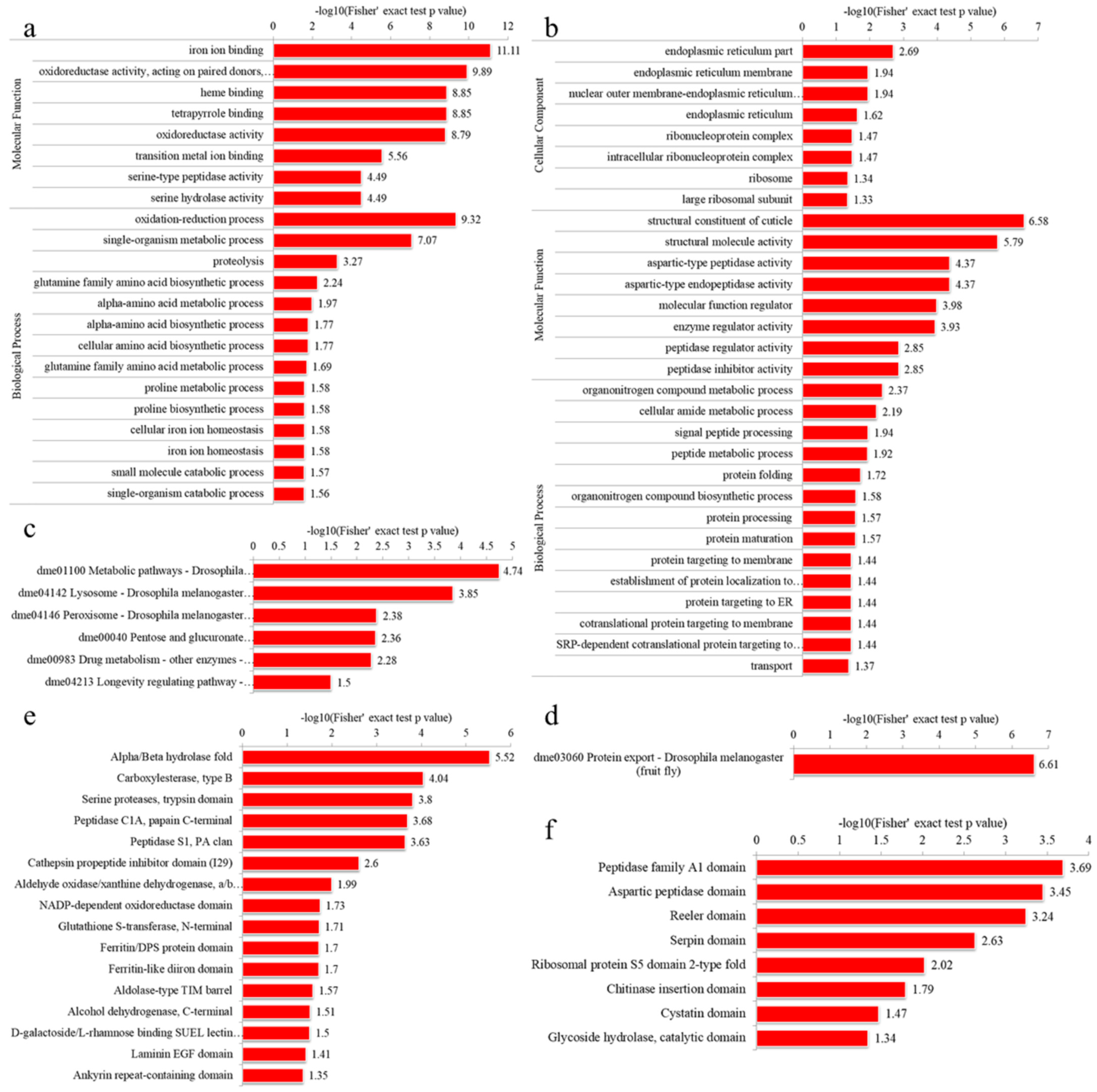
3.4.1. GO Enrichment
3.4.2. KEGG Pathway Enrichment
3.4.3. Domain Enrichment
3.5. Validation of DEPs by PRM
3.6. DEPs and KEGG Associated with Different Biological Parameters
3.7. DEPs and KEGG Associated with Artificial Diets
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Coudron, T.A.; Wittmeyer, J.; Kim, Y. Life history and cost analysis for continuous rearing of Podisus maculiventris (Say) (Heteroptera: Pentatomidae) on a zoophytophagous artificial diet. J. Econ. Entomol. 2002, 95, 1159–1168. [Google Scholar] [CrossRef] [PubMed]
- Coudron, T.A.; Kim, Y. Life history and cost analysis for continuous rearing of Perillus bioculatus (Heteroptera: Pentatomidae) on a zoophytogenous artificial diet. J. Econ. Entomol. 2004, 97, 807–812. [Google Scholar] [CrossRef]
- Glenister, C.S. Predatory heteropterans in augmentative biological control: An industry perspective. In Predatory Heteroptera: Their Ecology and Use in Biological Control; Coll, M., Ruberson, J.R., Eds.; Entomological Society of America: Lanham, MD, USA, 1998; pp. 199–208. [Google Scholar]
- Glenister, C.S.; Hoffmann, M.P. Mass-reared natural enemies: Scientific, technological, and informational needs and considerations. In Mass-Reared Natural Enemies: Application, Regulation, and Needs; Ridgway, R., Hoffmann, M.P., Inscoe, M.N., Glenister, C.S., Eds.; Entomological Society of America: Lanham, MD, USA, 1998; pp. 242–247. [Google Scholar]
- Ruberson, J.R.; Coll, M. Research needs for the predaceous Heteroptera. In Predatory Heteroptera: Their Ecology and Use in Biological Control; Coll, M., Ruberson, J.R., Eds.; Entomological Society of America: Lanham, MD, USA, 1998; pp. 225–233. [Google Scholar]
- Thompson, S.N. Nutrition and culture of entomophagous insects. Annu. Rev. Entomol. 1999, 44, 561–592. [Google Scholar] [CrossRef] [PubMed]
- Wittmeyer, J.L.; Coudron, T.A. Life table parameters, reproductive rate, intrinsic rate of increase and estimated cost of rearing Podisus maculiventris (Heteroptera: Pentatomidae) on an artificial diet. J. Econ. Entomol. 2001, 94, 1344–1352. [Google Scholar] [CrossRef]
- Lapointe, S.L.; Evens, T.J.; Niedz, R.P. Insect diets as mixtures: Optimization for a polyphagous weevil. J. Insect Physiol. 2008, 54, 1157–1167. [Google Scholar] [CrossRef]
- Lapointe, S.L.; Evens, T.J.; Niedz, R.P.; Hall, D.G. Artificial diet optimized to produce normative adults of Diaprepes abbreviatus (Coleoptera: Curculionidae). Environ. Entomol. 2010, 39, 670–677. [Google Scholar] [CrossRef]
- Tan, X.L.; Wang, S.; Zhang, F. Optimization an optimal artificial diet for the predatory bug Orius sauteri (Hemiptera: Anthocoridae). PLoS ONE 2013, 8, e61129. [Google Scholar] [CrossRef]
- Morales-Ramos, J.A.; Rojas, M.G.; Shapiro-Ilan, D.I.; Tedders, W.L. Self-selection of two diet components by Tenebrio molitor (Coleoptera: Tenebrionidae) larvae and its impact on fitness. Environ. Entomol. 2011, 40, 1285–1294. [Google Scholar] [CrossRef]
- Morales-Ramos, J.A.; Rojas, M.G.; Shapiro-Ilan, D.I.; Tedders, W.L. Use of nutrient self-selection as a diet refining tool in Tenebrio molitor (Coleoptera: Tenebrionidae). J. Entomol. Sci. 2013, 48, 206–221. [Google Scholar] [CrossRef]
- Morales-Ramos, J.A.; Rojas, M.G.; Dossey, A.T.; Berhow, M. Self-selection of food ingredients and agricultural by-products by the house cricket, Acheta domesticus (Orthoptera: Gryllidae): A Holistic approach to develop optimized diets. PLoS ONE 2020, 15, e0227400. [Google Scholar] [CrossRef]
- Morales-Ramos, J.A.; Rojas, M.G.; Kelstrup, H.C.; Emery, V. Self-selection of agricultural by-products and food ingredients by Tenebrio molitor (Coleoptera: Tenebrionidae) and impact on food utilization and nutrient intake. Insects 2020, 11, 827. [Google Scholar] [CrossRef] [PubMed]
- Özdemir, V.; Kolker, E. Precision Nutrition 4.0: A big data and ethics foresight analysis-convergence of agrigenomics, nutrigenomics, nutriproteomics, and nutrimetabolomics. Omics A J. Integr. Biol. 2016, 20, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Sauer, S.; Luge, T. Nutriproteomics: Facts, concepts, and perspectives. Proteomics 2015, 15, 997–1013. [Google Scholar] [CrossRef] [PubMed]
- Korte, R.; Brockmeyer, J. Novel mass spectrometry approaches in food proteomics. TrAC Trend. Anal. Chem. 2017, 96, 99–106. [Google Scholar] [CrossRef]
- Gao, C.Q.; Wang, Z.M.; Yu, E.Y. Studies on artificial rearing of Arma chinensis Fallou. J. Jilin For. Sci. Technol. 1993, 2, 16–18. [Google Scholar]
- Chai, X.M.; He, Z.H.; Jiang, P.; Wu, Z.D.; Pan, C.R.; Hu, R.D.; Ruan, Z.M. Studies on natural enemies of Dendrolimus punctatus in Zhejiang Province. J. Zhejiang For. Sci. Technol. 2000, 20, 1–56. [Google Scholar]
- Liang, Z.P.; Zhang, X.X.; Song, A.D.; Peng, H.Y. Biology of Clostera anachoreta and its control methods. Chin. Bull. Entomol. 2006, 43, 147–152. [Google Scholar]
- Yan, J.H.; Li, H.J.; Peng, H.Y.; Zhou, X.Z.; Gao, B.D. Bionomics of Batrachomorphus dentatus and its control. Chin. Bull. Entomol. 2006, 43, 562–566. [Google Scholar]
- Yan, J.H.; Tang, W.Y.; Zhang, H.; Wang, H.Y. Bionomics of the leafhopper Macropsis matsudanis. Chin. Bull. Entomol. 2006, 43, 245–248. [Google Scholar]
- Chen, J.; Zhang, J.P.; Zhang, J.H.; Tian, Y.H.; Xu, Z.C.; Li, G.W. Study on functional response of Arma chinensis to the adults of Monolepta hieroglyphica. Nat. Enem. Insects 2007, 29, 149–154. [Google Scholar]
- Gao, Z. Studies on Biological Characteristic and Control Condition of Arma chinensis Fallou. Master’s Thesis, Heilongjiang University, Harbin, China, 7 May 2010. [Google Scholar]
- Zou, D.Y.; Wang, M.Q.; Zhang, L.S.; Zhang, Y.; Zhang, X.J.; Chen, H.Y. Taxonomic and bionomic notes on Arma chinensis (Fallou) (Hemiptera: Pentatomidae: Asopinae). Zootaxa 2012, 3382, 41–52. [Google Scholar] [CrossRef]
- Silver, A. China seeks predator to stop voracious caterpillar. Nature 2019, 570, 286. [Google Scholar] [CrossRef]
- Tang, Y.T.; Li, Y.Y.; Liu, C.X.; Mao, J.J.; Chen, H.Y.; Zhang, L.S.; Wang, M.Q. Predation and behavior of Arma chinensis to Spodoptera frugiperda. Plant Protec. 2019, 45, 65–68. [Google Scholar]
- Wang, Y.; Zhang, H.M.; Yin, Y.Q.; Li, X.Y.; Zhao, X.Q.; Tang, Y.T.; Wang, M.Q.; Chen, A.D.; Chen, F.S.; Zhang, L.S. Predation of adult of Arma chinensis to larvae of Spodoptera frugiperda. Plant Protec. 2019, 45, 42–46. [Google Scholar]
- Wu, K.M. Management strategies of fall armyworn (Spodoptera frugiperda) in China. Plant Protec. 2020, 46, 1–5. [Google Scholar]
- Zou, D.Y.; Wu, H.H.; Coudron, T.A.; Zhang, L.S.; Wang, M.Q.; Liu, C.X.; Chen, H.Y. A meridic diet for continuous rearing of Arma chinensis (Hemiptera: Pentatomidae: Asopinae). Biol. Control 2013, 67, 491–497. [Google Scholar] [CrossRef]
- Zou, D.Y.; Coudron, T.A.; Liu, C.X.; Zhang, L.S.; Wang, M.Q.; Chen, H.Y. Nutrigenomics in Arma chinensis: Transcriptome analysis of Arma chinensis fed on artificial diet and Chinese oak silk moth Antheraea pernyi pupae. PLoS ONE 2013, 8, e60881. [Google Scholar] [CrossRef]
- Zou, D.Y.; Coudron, T.A.; Zhang, L.S.; Gu, X.S.; Xu, W.H.; Liu, X.L.; Wu, H.H. Performance of Arma chinensis reared on an artificial diet formulated using transcriptomic methods. Bull. Entomol. Res. 2019, 109, 24–33. [Google Scholar] [CrossRef]
- Coudron, T.A.; Yocum, G.D.; Brandt, S.L. Nutrigenomics: A case study in the measurement of insect response to nutritional quality. Entomol. Exp. Appl. 2006, 121, 1–14. [Google Scholar] [CrossRef]
- Yocum, G.D.; Coudron, T.A.; Brandt, S.L. Differential gene expression in Perillus bioculatus nymphs fed a suboptimal artificial diet. J. Insect Physiol. 2006, 52, 586–592. [Google Scholar] [CrossRef]
- Li, H.S.; Huang, Y.H.; Chen, M.L.; Ren, Z.; Qiu, B.Y.; De Clercq, P.; Heckel, G.; Pang, H. Genomic insight into diet adaptation in the biological control agent Cryptolaemus montrouzieri. BMC Genom. 2021, 22, 135. [Google Scholar] [CrossRef] [PubMed]
- Greenbaum, D.; Colangelo, C.; Williams, K.; Gerstein, M. Comparing protein abundance and mRNA expression levels on a genomic scale. Genome Biol. 2003, 4, 117. [Google Scholar] [CrossRef] [PubMed]
- de Godoy, L.M.F.; Olsen, J.V.; Cox, J.; Nielsen, M.L.; Hubner, N.C.; Fröhlich, F.; Walther, T.C.; Mann, M. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature 2008, 455, 1251–1254. [Google Scholar] [CrossRef] [PubMed]
- Maier, T.; Güell, M.; Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef] [PubMed]
- Clayton, C.E. Life without transcriptional control? From fly to man and back again. EMBO J. 2002, 21, 1881–1888. [Google Scholar] [CrossRef]
- Karp, N.A.; Huber, W.; Sadowski, P.G.; Charles, P.D.; Hester, S.V.; Lilley, K.S. Addressing accuracy and precision issues in iTRAQ quantitation. Mol. Cell. Proteom. 2010, 9, 1885–1897. [Google Scholar] [CrossRef] [PubMed]
- Becker, C.H.; Bern, M. Recent developments in quantitative proteomics. Mutat. Res.-Genet. Toxicol. Environ. Mutagenesis 2011, 722, 171–182. [Google Scholar] [CrossRef]
- Barnouin, K. Special issue in quantitative mass spectrometric proteomics. Amino Acids 2012, 43, 1005–1007. [Google Scholar] [CrossRef][Green Version]
- Bonnett, T.R.; Robert, J.A.; Pitt, C.; Fraser, J.D.; Keeling, C.I.; Bohlmann, J.; Huber, D.P.W. Global and comparitive proteomic profiling of overwintering and developing mountain pine beetle, Dendroctonus ponderosae (Coleoptera: Curculionidae), larvae, Insect Biochem. Mol. Biol. 2012, 42, 890–901. [Google Scholar]
- Tu, X.; Wang, J.; Hao, K.; Whitman, D.W.; Fan, Y.; Cao, G.; Zhang, Z. Transcriptomic and proteomic analysis of pre-diapause and non-diapause eggs of migratory locust, Locusta migratoria L. (Orthoptera: Acridoidea). Sci. Rep. 2015, 5, 11402. [Google Scholar] [CrossRef]
- Liu, L.; Li, G.; Sun, P.; Lei, C.; Huang, Q. Experimental verification and molecular basis of active immunization against fungal pathogens in termites. Sci. Rep. 2015, 5, 15106. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.Z.; Li, Y.Y.; An, T.; Huang, F.X.; Wang, M.Q.; Liu, C.X.; Mao, J.J.; Zhang, L.S. Comparative transcriptome and iTRAQ proteome analyses reveal the mechanisms of diapause in Aphidius gifuensis Ashmead (Hymenoptera: Aphidiidae). Front. Physiol. 2018, 9, 1697. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Gao, Y.; Li, J.; Ban, L. iTRAQ-based comparative proteomic analysis reveals molecular mechanisms underlying wing dimorphism of the pea aphid, Acyrthosiphon pisum. Front. Physiol. 2018, 9, 1016. [Google Scholar] [CrossRef] [PubMed]
- Du, C.; Liu, H.F.; Lin, Y.Z.; Wang, X.F.; Ma, J.; Li, Y.J.; Wang, X.; Zhou, J.H. Proteomic alteration of equine monocyte-derived macrophages infected with equine infectious anemia virus. Proteomics 2015, 15, 1843–1858. [Google Scholar] [CrossRef] [PubMed]
- Peterson, A.C.; Russell, J.D.; Bailey, D.J.; Westphall, M.S.; Coon, J.J. Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol. Cell. Proteom. 2012, 11, 1475–1488. [Google Scholar] [CrossRef] [PubMed]
- Tsuchiya, H.; Tanaka, K.; Saeki, Y. The parallel reaction monitoring method contributes to a highly sensitive polyubiquitin chain quantification. Biochem. Biophys. Res. Commun. 2013, 436, 223–229. [Google Scholar] [CrossRef]
- Yu, Q.; Liu, B.; Ruan, D.; Niu, C.; Shen, J.; Ni, M.; Cong, W.; Lu, X.; Jin, L. A novel targeted proteomics method for identification and relative quantitation of difference in nitration degree of OGDH between healthy and diabetic mouse. Proteomics 2014, 14, 2417–2426. [Google Scholar] [CrossRef]
- Wang, Z.X.; Shang, P.; Li, Q.G.; Wang, L.Y.; Chamba, Y.Z.; Zhang, H.; Zhang, B.; Wu, C.X. iTRAQ-based proteomic analysis reveals key proteins affecting muscle growth and lipid deposition in pigs. Sci. Rep. 2017, 7, 46717. [Google Scholar] [CrossRef]
- Wu, X.X.; Yan, J.X.; Wu, Y.H.; Zhang, H.B.; Mo, S.R.; Xu, X.Y.; Zhou, F.C.; Ding, H.D. Proteomic analysis by iTRAQ-PRM provides integrated insight into mechanisms of resistance in pepper to Bemisia tabaci (Gennadius). BMC Plant Biol. 2019, 19, 270. [Google Scholar] [CrossRef]
- Zeng, W.; Sun, Z.; Cai, Z.; Chen, H.; Lai, Z.; Yang, S.; Tang, X. Proteomic analysis by iTRAQ-MRM of soybean resistance to Lamprosema indicate. BMC Genom. 2017, 18, 444. [Google Scholar] [CrossRef]
- Bownes, M.; Rembold, H. The titre of juvenile hormone during the pupal and adult stages of the life-cycle of Drosophila melanogaster. Eur. J. Biochem. 1987, 164, 709–712. [Google Scholar] [CrossRef] [PubMed]
- Wyatt, G.R. Gene regulation in insect reproduction. Invert. Reprod. Dev. 1991, 20, 1–35. [Google Scholar] [CrossRef]
- Grewal, S.S. Insulin/TOR signaling in growth and homeostasis: A view from the fly world. Int. J. Biochem. Cell Biol. 2009, 41, 1006–1010. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Edgar, B.A.; Grewal, S.S. Nutritional control of gene expression in Drosophila larvae via TOR, Myc and a novel cis-regulatory element. BMC Cell Biol. 2010, 11, 7. [Google Scholar] [CrossRef] [PubMed]
- Hay, N.; Sonenberg, N. Upstream and downstream of mTOR. Genes Dev. 2004, 18, 1926–1945. [Google Scholar] [CrossRef]
- Grewal, S.S.; Evans, J.R.; Edgar, B.A. Drosophila TIF-IA is required for ribosome synthesis and cell growth and is regulated by the TOR pathway. J. Cell Biol. 2007, 179, 1105–1113. [Google Scholar] [CrossRef]
- Guertin, D.A.; Guntur, K.V.; Bell, G.W.; Thoreen, C.C.; Sabatini, D.M. Functional genomics identifies TOR-regulated genes that control growth and division. Curr. Biol. 2006, 16, 958–970. [Google Scholar] [CrossRef]
- Tiefenbock, S.K.; Baltzer, C.; Egli, N.A.; Frei, C. The Drosophila PGC-1 homologue Spargel coordinates mitochondrial activity to insulin signalling. EMBO J. 2009, 29, 171–183. [Google Scholar] [CrossRef]
- Baltzer, C.; Tiefenbock, S.K.; Marti, M.; Frei, C. Nutrition controls mitochondrial biogenesis in the Drosophila adipose tissue through Delg and cyclin D/Cdk4. PLoS ONE 2009, 4, e6935. [Google Scholar] [CrossRef]
- Keith, C.T.; Schreiber, S.L. PIK-related kinases: DNA repair, recombination, and cell cycle checkpoints. Science 1995, 270, 50–51. [Google Scholar] [CrossRef]
- Stanfel, M.N.; Shamieh, L.S.; Kaeberlein, M.; Kennedy, B.K. The TOR pathway comes of age. Biochim. Biophys. Acta 2009, 1790, 1067–1074. [Google Scholar] [CrossRef] [PubMed]
- Laplante, M.; Sabatini, D.M. mTOR signaling in growth control and disease. Cell 2012, 149, 274–293. [Google Scholar] [CrossRef]
- Shimobayashi, M.; Hall, M.N. Making new contacts: The mTOR network in metabolism and signalling crosstalk. Nat. Rev. Mol. Cell. Biol. 2014, 15, 155–162. [Google Scholar] [CrossRef]
- Johnson, S.C.; Sangesland, M.; Kaeberlein, M.; Rabinovitch, P.S. Modulating mTOR in aging and health. Interdiscip. Top. Gerontol. 2015, 40, 107–127. [Google Scholar]
- Vellai, T.; Takacs-Vellai, K.; Zhang, Y.; Kovacs, A.L.; Orosz, L.; Muller, F. Genetics: Influence of TOR kinase on lifespan in C. elegans. Nature 2003, 426, 620. [Google Scholar] [CrossRef] [PubMed]
- Jia, K.; Chen, D.; Riddle, D.L. The TOR pathway interacts with the insulin signaling pathway to regulate C. elegans larval development, metabolism and life span. Development 2004, 131, 3897–3906. [Google Scholar] [CrossRef] [PubMed]
- Kapahi, P.; Zid, B.M.; Harper, T.; Koslover, D.; Sapin, V.; Benzer, S. Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway. Curr. Biol. 2004, 14, 885–890. [Google Scholar] [CrossRef] [PubMed]
- Schinaman, J.M.; Rana, A.; Ja, W.W.; Clark, R.I.; Walker, D.W. Rapamycin modulates tissue aging and lifespan independently of the gut microbiota in Drosophila. Sci. Rep. 2019, 9, 7824. [Google Scholar] [CrossRef]
- Takeuchi, A.; Miyamoto, T.; Yamaji, K.; Masuho, Y.; Hayashi, M.; Hayashi, H.; Onozaki, K. A human erythrocyte-derived growth-promoting factor with a wide target cell spectrum: Identification as catalase. Cancer Res. 1995, 55, 1586–1589. [Google Scholar]
- Taub, J.; Lau, J.F.; Ma, C.; Hahn, J.H.; Hoque, R.; Rothblatt, J.; Chalfie, M. A cytosolic catalase is needed to extend adult lifespan in C. elegans daf-C and clk-1 mutants. Nature 2003, 421, 764. [Google Scholar] [CrossRef]
- Martins, D.; English, A.M. Catalase activity is stimulated by H2O2 in rich culture medium and is required for H2O2 resistance and adaptation in yeast. Redox Biol. 2014, 2, 308–313. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Tower, J. FLP recombinase-mediated induction of Cu/Zn-superoxide dismutase transgene expression can extend the life span of adult Drosophila melanogaster flies. Mol. Cell. Biol. 1999, 19, 216–228. [Google Scholar] [CrossRef] [PubMed]
- Parkes, T.L.; Elia, A.J.; Dickson, D.; Hilliker, A.J.; Phillips, J.P.; Boulianne, G.L. Extension of Drosophila lifespan by overexpression of human SOD1 in motorneurons. Nat. Genet. 1998, 19, 171–174. [Google Scholar] [CrossRef] [PubMed]
- Mayer, M.P.; Bukau, B. Hsp70 chaperones: Cellular functions and molecular mechanism. Cell. Mol. Life Sci. 2005, 62, 670. [Google Scholar] [CrossRef] [PubMed]
- Benoit, J.B.; Lopez-Martinez, G.; Teets, N.M.; Phillips, S.A.; Denlinger, D.L. Responses of the bed bug, Cimex lectularius, to temperature extremes and dehydration: Levels of tolerance, rapid cold hardening and expression of heat shock proteins. Med. Vet. Entomol. 2009, 23, 418–425. [Google Scholar] [CrossRef] [PubMed]
- Kalies, K.-U.; Hartmann, E. Membrane topology of the 12- and the 25-kDa subunits of the mammalian signal peptidase complex. J. Biol. Chem. 1996, 271, 3925–3929. [Google Scholar] [CrossRef]
- Haase Gilbert, E.; Kwak, S.J.; Chen, R.; Mardon, G. Drosophila signal peptidase complex member Spase12 is required for development and cell differentiation. PLoS ONE 2013, 8, e60908. [Google Scholar] [CrossRef]
- YaDeau, J.T.; Blobel, G. Solubilization and characterization of yeast signal peptidase. J. Biol. Chem. 1989, 264, 2928–2934. [Google Scholar] [CrossRef]
- Mullins, C.; Lu, Y.; Campbell, A.; Fang, H.; Green, N. A mutation affecting signal peptidase inhibits degradation of an abnormal membrane protein in Saccharomyces cerevisiae. J. Biol. Chem. 1995, 270, 17139–17147. [Google Scholar] [CrossRef]
- VanValkenburgh, C.; Chen, X.; Mullins, C.; Fang, H.; Green, N. The catalytic mechanism of endoplasmic reticulum signal peptidase appears to be distinct from most eubacterial signal peptidases. J. Biol. Chem. 1999, 274, 11519–11525. [Google Scholar] [CrossRef]
- Chen, X.; VanValkenburgh, C.; Liang, H.; Fang, H.; Green, N. Signal peptidase and oligosaccharyltransferase interact in a sequential and dependent manner within the endoplasmic reticulum. J. Biol. Chem. 2001, 276, 2411–2416. [Google Scholar] [CrossRef] [PubMed]
- Amdam, G.V.; Fennern, E.; Havukainen, H. Vitellogenin in honey bee behavior and lifespan. In Honeybee Neurobiology and Behavior: A Tribute to Randolf Menzel; Galizia, C., Eisenhardt, D., Giurfa, M., Eds.; Springer: Dordrecht, The Netherlands, 2012; pp. 17–30. [Google Scholar]
- Chapman, T. Seminal fluid-mediated fitness traits in Drosophila. Heredity 2001, 87, 511–521. [Google Scholar] [CrossRef] [PubMed]
- Poiani, A. Complexity of seminal fluid: A review. Behav. Ecol. Sociobiol. 2006, 60, 289–310. [Google Scholar] [CrossRef]
- Ravi Ram, K.; Wolfner, M.F. Seminal influences: Drosophila Acps and the molecular interplay between males and females during reproduction. Integr. Comp. Biol. 2007, 47, 427–445. [Google Scholar] [CrossRef]
- Avila, F.W.; Sirot, L.K.; LaFlamme, B.A.; Rubinstein, C.D.; Wolfner, M.F. Insect seminal fluid proteins: Identification and function. Annu. Rev. Entomol. 2011, 56, 21–40. [Google Scholar] [CrossRef]
- Robinson, P.K. Enzymes: Principles and biotechnological applications. Essays Biochem. 2015, 59, 1–41. [Google Scholar] [CrossRef]
- Islinger, M.; Voelkl, A.; Fahimi, H.D.; Schrader, M. The peroxisome: An update on mysteries 2.0. Histochem. Cell Biol. 2018, 150, 443–471. [Google Scholar] [CrossRef]
- Bonekamp, N.A.; Völkl, A.; Fahimi, H.D.; Schrader, M. Reactive oxygen species and peroxisomes: Struggling for balance. BioFactors 2009, 35, 346–355. [Google Scholar] [CrossRef]
- Zinke, I.; Schütz, C.S.; Katzenberger, J.D.; Bauer, M.; Pankratz, M.J. Nutrient control of gene expression in Drosophila: Microarray analysis of starvation and sugar-dependent response. EMBO J. 2002, 21, 6162–6173. [Google Scholar]


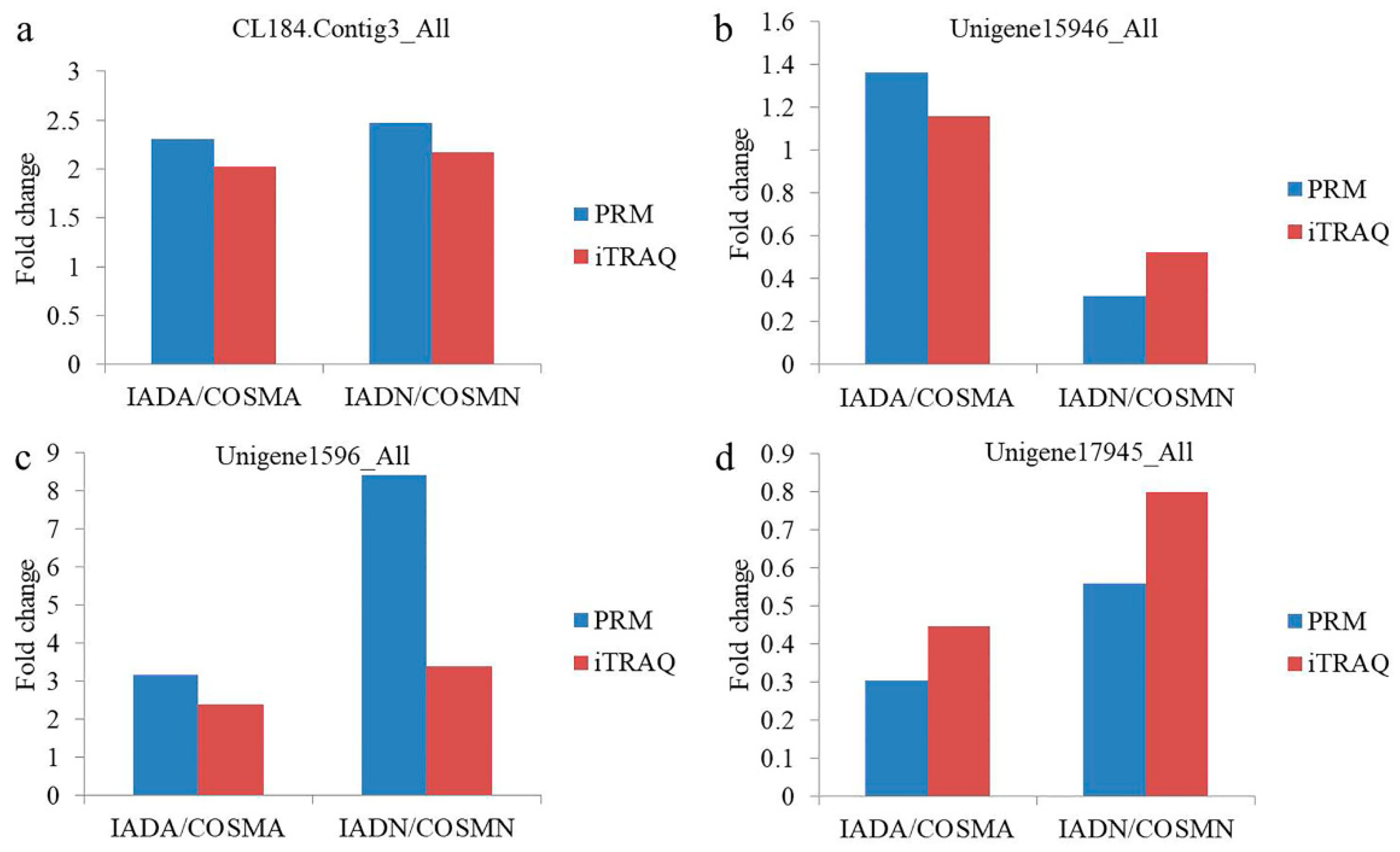
| Protein Accession | Protein Description | Peptide Sequence | Fold Change (IADA/COSMA) in PRM | Fold Change (IADN/COSMN) in PRM |
|---|---|---|---|---|
| CL184.Contig3_All | minus strand secreted salivary trypsin | EVSCPCGWTNK | 2.303 (p = 0.0007) | 2.476 (p = 0.0058) |
| Unigene15946_All | seminal fluid protein CSSFP042 | ELFYISNSQR | 1.363 (p = 0.0863) | 0.32 (p = 0.0023) |
| FQPESTSK | ||||
| TNFAGDSAGALK | ||||
| Unigene1596_All | ferritin | AVEASLQLEK | 3.177 (p = 0.0157) | 8.417 (p = 0.0637) |
| SLGDLLTNVR | ||||
| Unigene17945_All | minus strand odorant-binding protein RproOBP2 precursor | ELQVFGK | 0.305 (p = 0.0254) | 0.56 (p = 0.0555) |
| VPENLGNPCEVAFAVTK a | ||||
| VPENLGNPCEVAFAVTK b |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zou, D.; Coudron, T.A.; Wu, H.; Zhang, L.; Wang, M.; Xu, W.; Xu, J.; Song, L.; Xiao, X. Differential Proteomics Analysis Unraveled Mechanisms of Arma chinensis Responding to Improved Artificial Diet. Insects 2022, 13, 605. https://doi.org/10.3390/insects13070605
Zou D, Coudron TA, Wu H, Zhang L, Wang M, Xu W, Xu J, Song L, Xiao X. Differential Proteomics Analysis Unraveled Mechanisms of Arma chinensis Responding to Improved Artificial Diet. Insects. 2022; 13(7):605. https://doi.org/10.3390/insects13070605
Chicago/Turabian StyleZou, Deyu, Thomas A. Coudron, Huihui Wu, Lisheng Zhang, Mengqing Wang, Weihong Xu, Jingyang Xu, Liuxiao Song, and Xuezhuang Xiao. 2022. "Differential Proteomics Analysis Unraveled Mechanisms of Arma chinensis Responding to Improved Artificial Diet" Insects 13, no. 7: 605. https://doi.org/10.3390/insects13070605
APA StyleZou, D., Coudron, T. A., Wu, H., Zhang, L., Wang, M., Xu, W., Xu, J., Song, L., & Xiao, X. (2022). Differential Proteomics Analysis Unraveled Mechanisms of Arma chinensis Responding to Improved Artificial Diet. Insects, 13(7), 605. https://doi.org/10.3390/insects13070605








