Immunoinformatics Design and Assessment of a Multiepitope Antigen (OvMCBL02) for Onchocerciasis Diagnosis and Monitoring
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Consideration
2.2. Study Site, Population and Sample Collection
2.3. Proteomic Profile of O. volvulus
2.4. In Silico Screening of Signal Peptide, Homology and Antigenicity Predictions
2.5. Transmembrane Domain and Linear B-Epitope Prediction
2.6. Multiepitope Antigen Construction, Antigenicity, Physicochemical Properties and Solubility Prediction
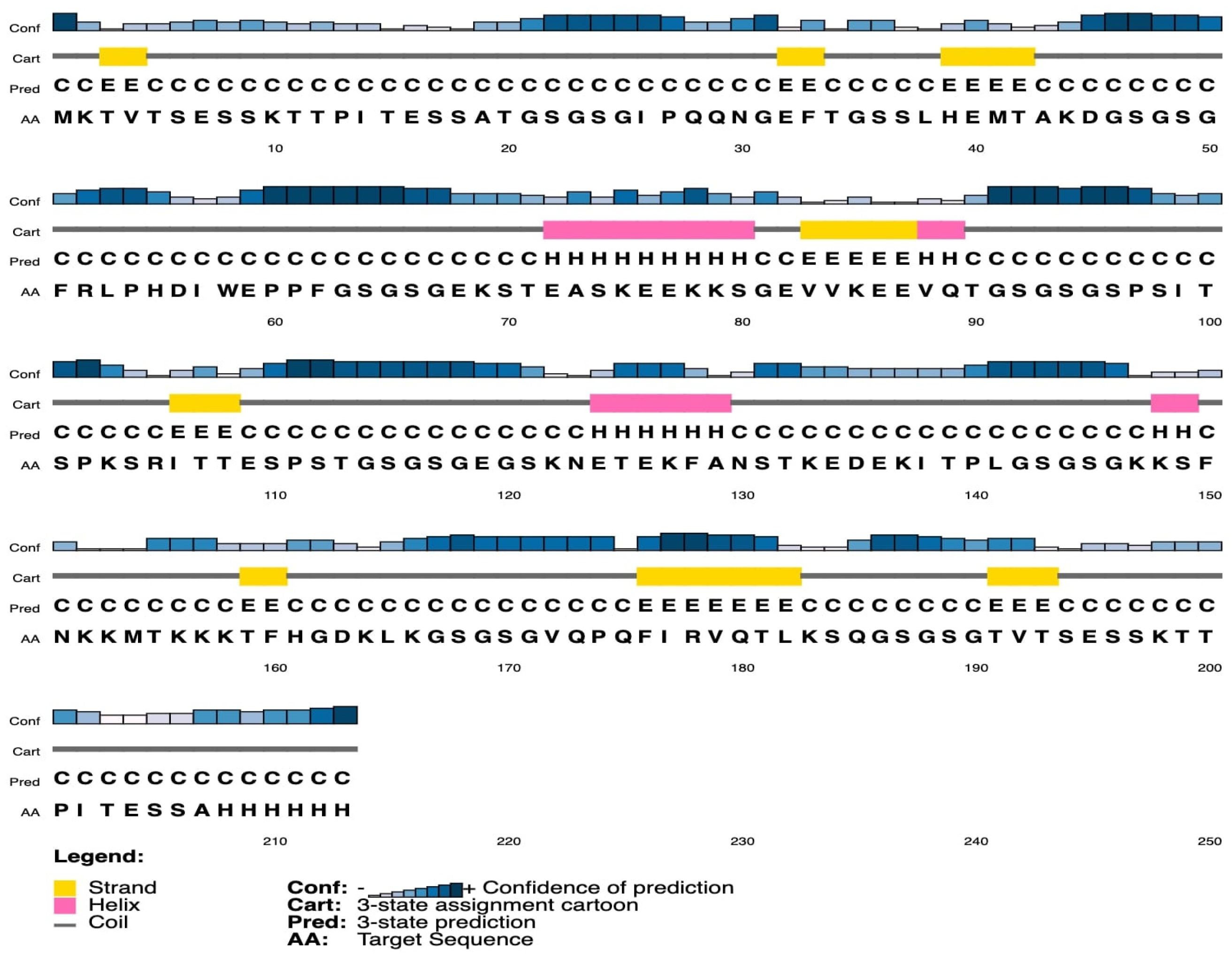
2.7. Secondary Structure Prediction
2.8. Serological Assessment of OvMCBL02 Multiepitope Antigen
2.9. Data Analysis
3. Results
3.1. Protein Selection
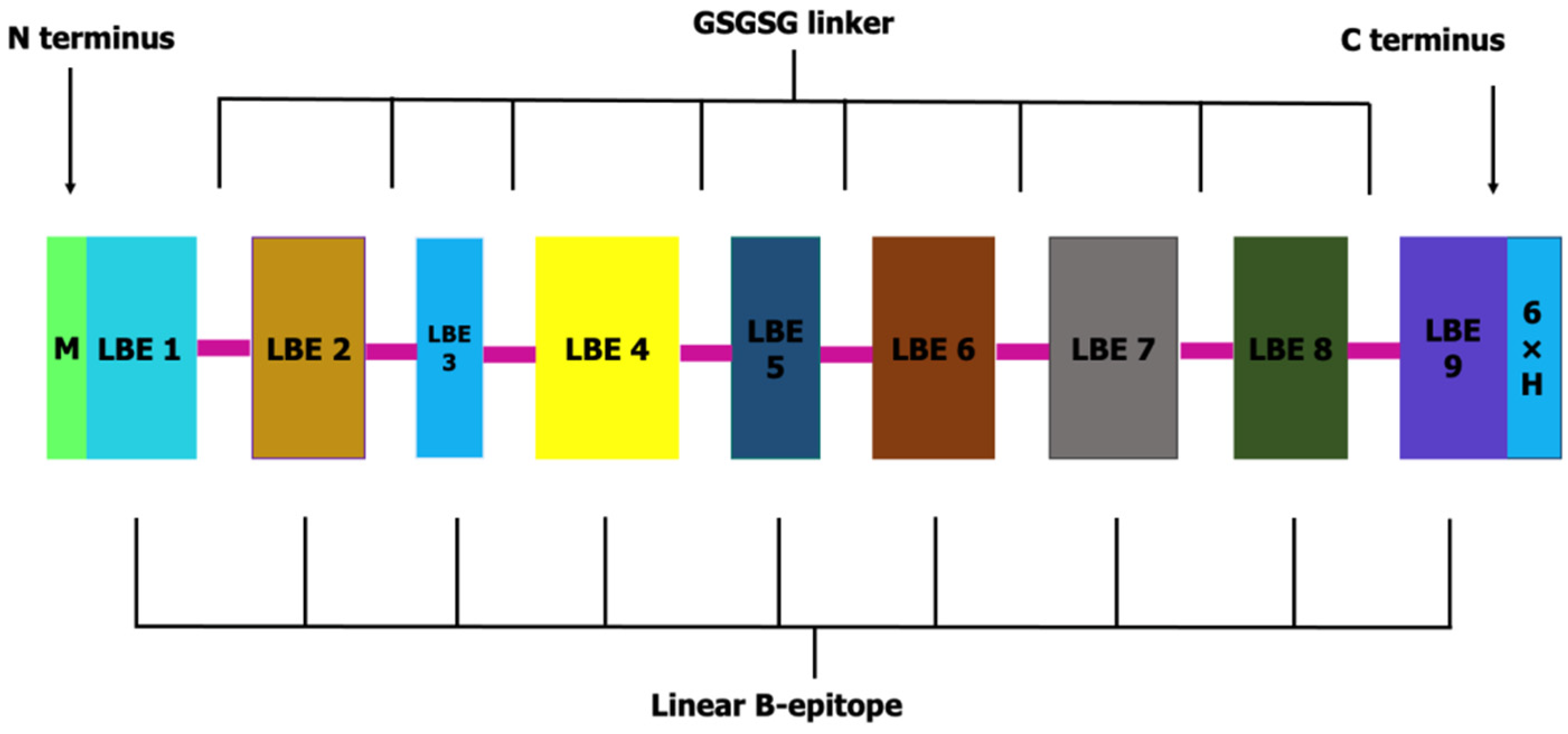
3.2. OvMCBL02 Multiepitope Antigen Construction
3.3. Antigenicity Prediction, Physicochemical Properties, Solubility and Secondary Structure of OvMCBL02 Multiepitope Antigen
3.4. Humoral Immune Response to OvMCBL02 Multiepitope Antigen
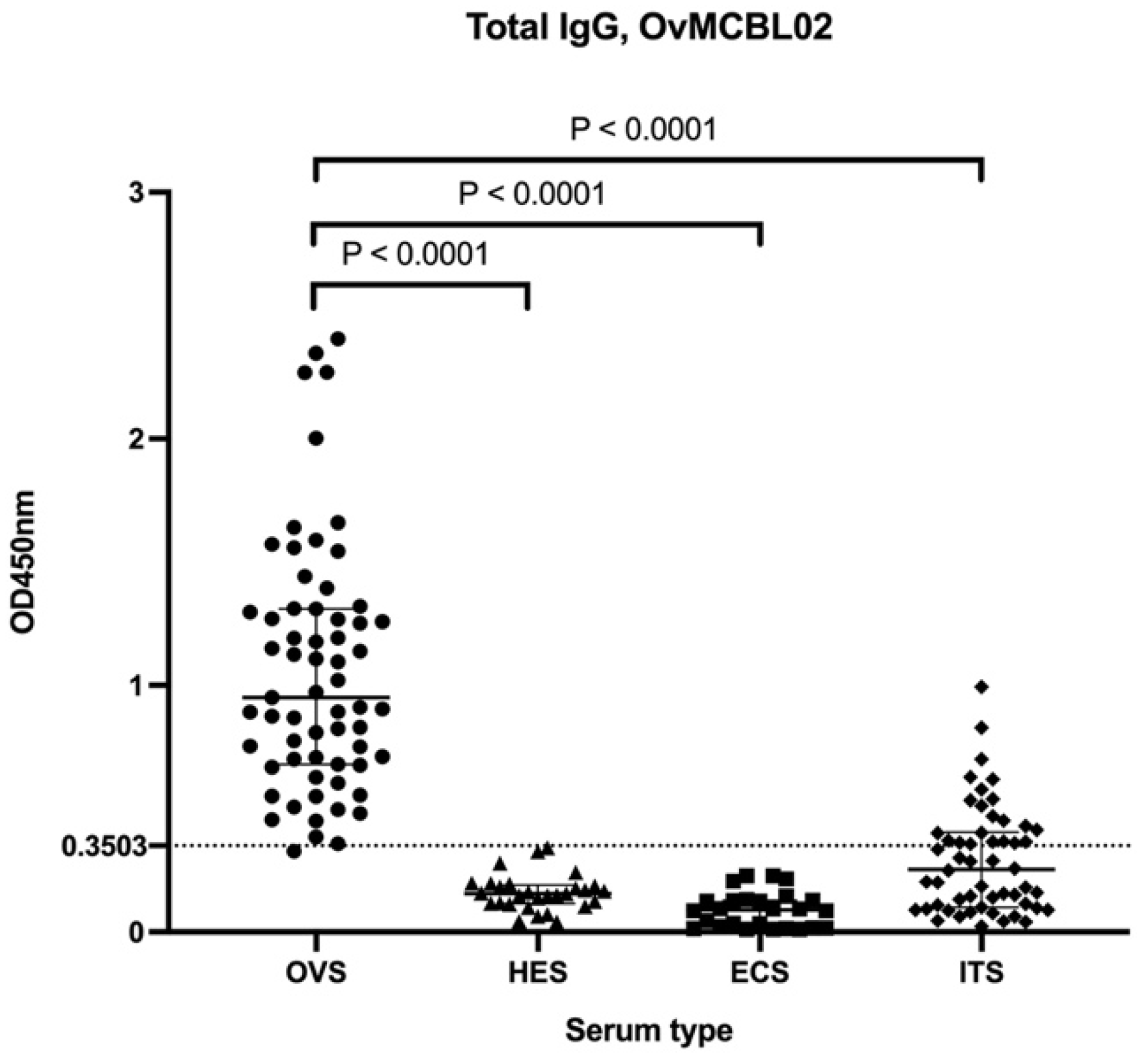
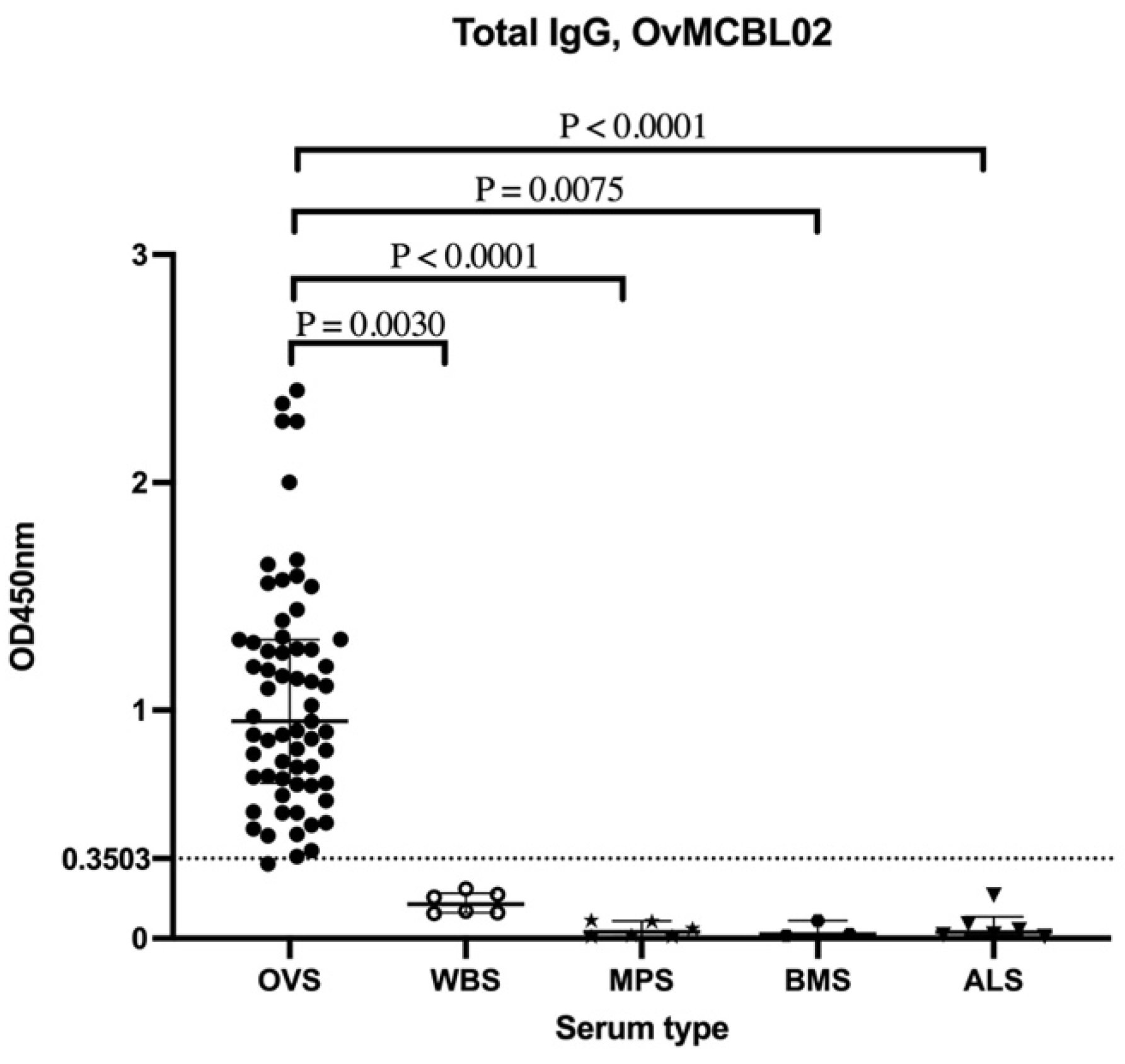
3.5. Assessment of Humoral Immune Response of Related Nematode Sera to OvMCBL02 Multiepitope Antigen
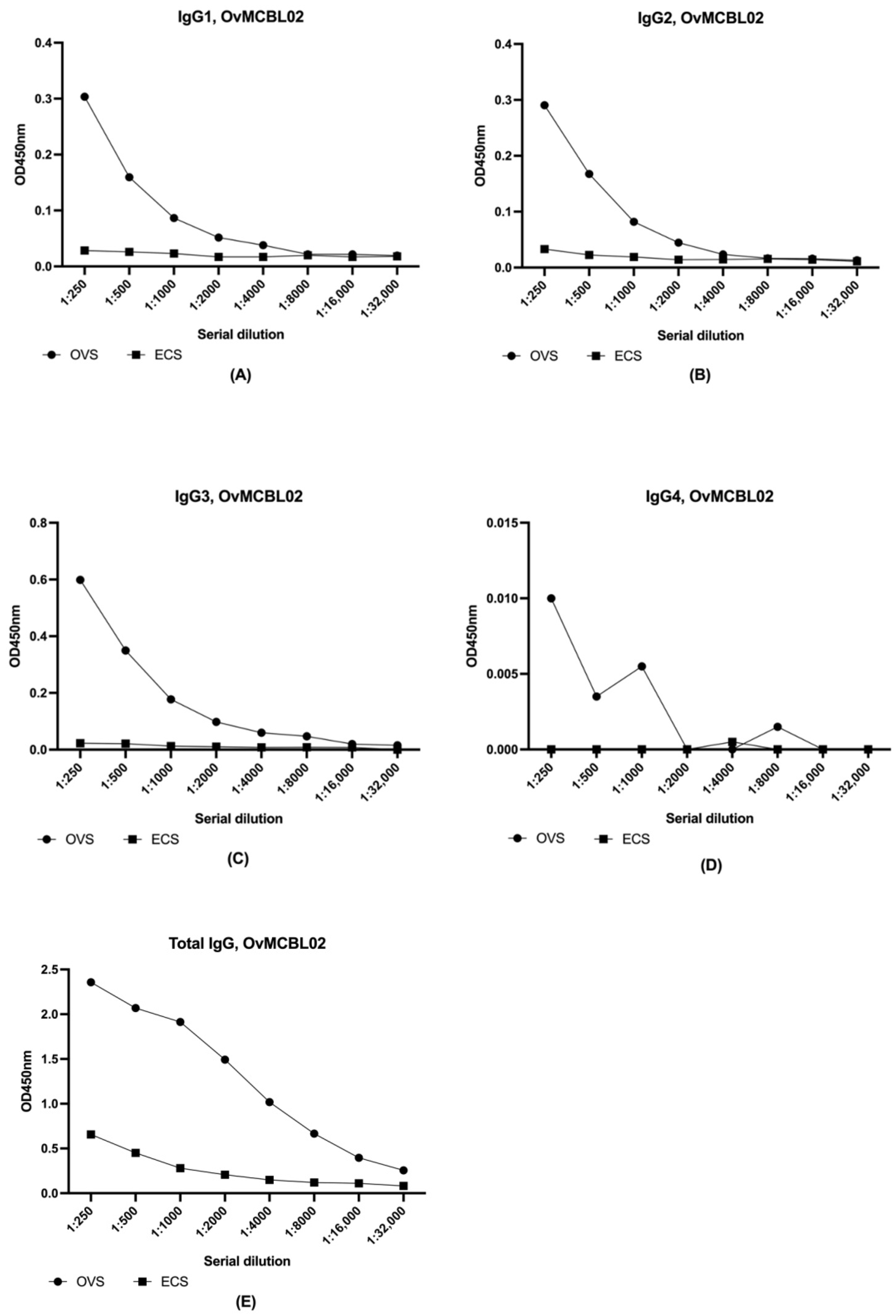
3.6. Total IgG, IgG1, IgG2 and IgG3 but Not IgG4 Subclass Responded Positively to OvMCBL02 Multiepitope Antigen
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organisation. Onchocerciasis Key Facts Geneva 2022. Available online: https://www.who.int/news-room/fact-sheets/detail/onchocerciasis#:~:text=Key%20facts,the%20parasitic%20worm%20Onchocerca%20volvulus.&text=Symptoms%20include%20severe%20itching%2C%20disfiguring,live%20in%2031%20African%20countries (accessed on 4 March 2022).
- Noormahomed, E.V.; Akrami, K.; Mascaró-Lazcano, C. Onchocerciasis, an Undiagnosed Disease in Mozambique: Identifying Research Opportunities. Parasites Vectors 2016, 9, 180. [Google Scholar] [CrossRef] [PubMed]
- Melchers, N.V.S.V.; Stolk, W.A.; van Loon, W.; Pedrique, B.; Bakker, R.; Murdoch, M.E.; de Vlas, S.J.; Coffeng, L.E. The Burden of Skin Disease and Eye Disease Due to Onchocerciasis in Countries Formerly under the African Programme for Onchocerciasis Control Mandate for 1990, 2020, and 2030. PLoS Neglected Trop. Dis. 2021, 15, e0009604. [Google Scholar] [CrossRef]
- Ubachukwu, P.O. Socio-Economic Impact Of Onchocerciasis With Particular Reference To Females And Children: A Review. Anim. Res. Int. 2008, 3, 494–504. [Google Scholar] [CrossRef]
- Gopinath, R.; Ostrowski, M.; Justement, S.J.; Fauci, A.S.; Nutman, T.B. Filarial Infections Increase Susceptibility to Human Immunodeficiency Virus Infection in Peripheral Blood Mononuclear Cells In Vitro. J. Infect. Dis. 2000, 182, 1804–1808. [Google Scholar] [CrossRef] [PubMed]
- Colebunders, R.; Njamnshi, A.K.; Menon, S.; Newton, C.R.; Hotterbeekx, A.; Preux, P.-M.; Hopkins, A.; Vaillant, M.; Fodjo, J.N.S. Onchocerca Volvulus and Epilepsy: A Comprehensive Review Using the Bradford Hill Criteria for Causation. PLOS Neglected Trop. Dis. 2021, 15, e0008965. [Google Scholar] [CrossRef] [PubMed]
- Egbert, P.R.; Jacobson, D.W.; Fiadoyor, S.; Dadzie, P.; Ellingson, K.D. Onchocerciasis: A Potential Risk Factor for Glaucoma. Br. J. Ophthalmol. 2005, 89, 796–798. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, A.D. Neglected Tropical Diseases in Africa: A New Paradigm. Int. Health 2016, 8, i28–i33. [Google Scholar] [CrossRef]
- Eberhard, M.L.; Cupp, E.W.; Katholi, C.R.; Richards, F.O.; Unnasch, T.R. Skin Snips Have No Role in Programmatic Evaluations for Onchocerciasis Elimination: A Reply to Bottomley et al. Parasites Vectors 2017, 10, 154. [Google Scholar] [CrossRef][Green Version]
- Denery, J.R.; Nunes, A.A.K.; Hixon, M.S.; Dickerson, T.J.; Janda, K.D. Metabolomics-Based Discovery of Diagnostic Biomarkers for Onchocerciasis. PLOS Neglected Trop. Dis. 2010, 4, e834. [Google Scholar] [CrossRef]
- Bennuru, S.; Lustigman, S.; Abraham, D.; Nutman, T.B. Metabolite Profiling of Infection-Associated Metabolic Markers of Onchocerciasis. Mol. Biochem. Parasitol. 2017, 215, 58–69. [Google Scholar] [CrossRef]
- Quintana, J.F.; Makepeace, B.L.; Babayan, S.A.; Ivens, A.; Pfarr, K.M.; Blaxter, M.; Debrah, A.; Wanji, S.; Ngangyung, H.F.; Bah, G.S.; et al. Extracellular Onchocerca-Derived Small RNAs in Host Nodules and Blood. Parasites Vectors 2015, 8, 58. [Google Scholar] [CrossRef] [PubMed]
- Tritten, L.; Burkman, E.; Moorhead, A.; Satti, M.; Geary, J.; MacKenzie, C.; Geary, T. Detection of Circulating Parasite-Derived MicroRNAs in Filarial Infections. PLoS Neglected Trop. Dis. 2014, 8, e2971. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, C.; Quek, S.; Pionnier, N.; Turner, J.D.; Wanji, S.; Wagstaff, S.C.; Taylor, M.J. The Insufficiency of Circulating miRNA and DNA as Diagnostic Tools or as Biomarkers of Treatment Efficacy for Onchocerca Volvulus. Sci. Rep. 2020, 10, 6672. [Google Scholar] [CrossRef] [PubMed]
- Lagatie, O.; Debrah, L.B.; Debrah, A.; Stuyver, L.J. Plasma-Derived Parasitic MicroRNAs Have Insufficient Concentrations to Be Used as Diagnostic Biomarker for Detection of Onchocerca Volvulus Infection or Treatment Monitoring using LNA-Based RT-qPCR. Parasitol. Res. 2017, 116, 1013–1022. [Google Scholar] [CrossRef]
- Bennuru, S.; Cotton, J.A.; Ribeiro, J.M.C.; Grote, A.; Harsha, B.; Holroyd, N.; Mhashilkar, A.; Molina, D.M.; Randall, A.Z.; Shandling, A.D.; et al. Stage-Specific Transcriptome and Proteome Analyses of the Filarial Parasite Onchocerca volvulus and Its Wolbachia endosymbiont. mBio 2016, 7, e02028-16. [Google Scholar] [CrossRef]
- McNulty, S.N.; Rosa, B.A.; Fischer, P.U.; Rumsey, J.M.; Erdmann-Gilmore, P.; Curtis, K.C.; Specht, S.; Townsend, R.R.; Weil, G.J.; Mitreva, M. An Integrated Multiomics Approach to Identify Candidate Antigens for Serodiagnosis of Human Onchocerciasis*. Mol. Cell. Proteom. 2015, 14, 3224–3233. [Google Scholar] [CrossRef]
- Hotterbeekx, A.; Perneel, J.; Mandro, M.; Abhafule, G.; Fodjo, J.N.S.; Dusabimana, A.; Abrams, S.; Kumar-Singh, S.; Colebunders, R. Comparison of Diagnostic Tests for Onchocerca volvulus in the Democratic Republic of Congo. Pathogens 2020, 9, 435. [Google Scholar] [CrossRef]
- Joshi, V.G.; Dighe, V.D.; Thakuria, D.; Malik, Y.S.; Kumar, S. Multiple Antigenic Peptide (MAP): A Synthetic Peptide Dendrimer for Diagnostic, Antiviral and Vaccine Strategies for Emerging and Re-Emerging Viral Diseases. Indian J. Virol. 2013, 24, 312–320. [Google Scholar] [CrossRef]
- Nkouawa, A.; Sako, Y.; Itoh, S.; Kouojip-Mabou, A.; Nganou, C.N.; Saijo, Y.; Knapp, J.; Yamasaki, H.; Nakao, M.; Nakaya, K.; et al. Serological Studies of Neurologic Helminthic Infections in Rural Areas of Southwest Cameroon: Toxocariasis, Cysticercosis and Paragonimiasis. PLOS Neglected Trop. Dis. 2010, 4, e732. [Google Scholar] [CrossRef]
- Shintouo, C.M.; Shey, R.A.; Nebangwa, D.N.; Esoh, K.K.; Nongley, N.F.; Nguve, J.E.; Giron, P.; Mutesa, L.; Vanhamme, L.; Souopgui, J.; et al. In Silico Design and Validation of OvMANE1, a Chimeric Antigen for Human Onchocerciasis Diagnosis. Pathogens 2020, 9, 495. [Google Scholar] [CrossRef]
- Yasin, N.; Laxmanappa, H.S.; Muddapur, U.M.; Cheruvathur, J.; Prakash, S.U.; Thulasiram, H.V. Design, Expression, and Evaluation of Novel Multiepitope Chimeric Antigen of Wuchereria Bancrofti for the Diagnosis of Lymphatic Filariasis—A Structure-Based Strategy. Int. Immunopharmacol. 2020, 83, 106431. [Google Scholar] [CrossRef] [PubMed]
- Gómara, M.J.; Fernández, L.; Pérez, T.; Ercilla, G.; Haro, I. Assessment of Synthetic Chimeric Multiple Antigenic Peptides for Diagnosis of GB Virus C Infection. Anal. Biochem. 2010, 396, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Hajissa, K.; Zakaria, R.; Suppian, R.; Mohamed, Z. An Evaluation of a Recombinant Multiepitope Based Antigen for Detection of Toxoplasma Gondii Specific Antibodies. BMC Infect. Dis. 2017, 17, 807. [Google Scholar] [CrossRef] [PubMed]
- Hajissa, K.; Zakaria, R.; Suppian, R.; Mohamed, Z. Design and Evaluation of a Recombinant Multi-Epitope Antigen for Serodiagnosis of Toxoplasma Gondii Infection in Humans. Parasites Vectors 2015, 8, 315. [Google Scholar] [CrossRef] [PubMed]
- Hernández, M.; Pozo, L.; Gómez, I.; Melchor, A. Chimeric Synthetic Peptide as Antigen for Immunodiagnosis of HIV-1 Infection. Biochem. Biophys. Res. Commun. 2000, 272, 259–262. [Google Scholar] [CrossRef]
- Marin, M.H.; Peña, L.P.; Tanty, C.R.; Clarke, D.H.; Arenas, M.A.; Noguerol, K.R.; León, C.S. Antigenic Activity of Three Chimeric Synthetic Peptides of the Transmembrane (Gp41) and the Envelope (Gp120) Glycoproteins of HIV-1 Virus. Prep. Biochem. Biotechnol. 2004, 34, 227–237. [Google Scholar] [CrossRef]
- Santos, F.L.N.; Celedon, P.A.F.; Zanchin, N.; Souza, W.; Da Silva, E.D.; Foti, L.; Krieger, M.A.; Gomes, Y.D.M. Accuracy of Chimeric Proteins in the Serological Diagnosis of Chronic Chagas Disease—A Phase II Study. PLoS Neglected Trop. Dis. 2017, 11, e0005433. [Google Scholar] [CrossRef]
- Schulz-Key, H.; Albiez, E.J.; Büttner, D.W. Isolation of Living Adult Onchocerca volvulus from Nodules. Tropenmedizin Parasitol. 1977, 28, 428–430. [Google Scholar]
- Kamga, G.R.G.R.; Dissak-Delon, F.N.F.N.; Nana-Djeunga, H.H.C.; Biholong, B.D.B.D.; Ghogomu, S.M.; Souopgui, J.; Kamgno, J.J.; Robert, A. Important Progress Towards Elimination of Onchocerciasis in the West Region of Cameroon. Parasites Vectors 2017, 10, 373. [Google Scholar] [CrossRef]
- Noma, M.; Zouré, H.G.M.; Tekle, A.H.; Enyong, P.A.I.; Nwoke, B.E.B.; Remme, J.H.F. The Geographic Distribution of Onchocerciasis in the 20 Participating Countries of the African Programme for Onchocerciasis Control: (1) Priority Areas for Ivermectin Treatment. Parasites Vectors 2014, 7, 325. [Google Scholar] [CrossRef]
- Zhang, Y.; Fonslow, B.R.; Shan, B.; Baek, M.-C.; Yates, J.R., 3rd. Protein Analysis by Shotgun/Bottom-up Proteomics. Chem. Rev. 2013, 113, 2343–2394. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-W.; Pai, T.-W. Machine Learning-Based Methods for Prediction of Linear B-Cell Epitopes. Immunoinformatics 2014, 1184, 217–236. [Google Scholar] [CrossRef]
- Faria, A.R.; Veloso, L.D.C.; Coura-Vital, W.; Reis, A.B.; Damasceno, L.M.; Gazzinelli, R.; Andrade, H.M. Novel Recombinant Multiepitope Proteins for the Diagnosis of Asymptomatic Leishmania Infantum-Infected Dogs. PLoS Neglected Trop. Dis. 2015, 9, e3429. [Google Scholar] [CrossRef] [PubMed]
- Aathmanathan, V.S.; Jothi, N.; Prajapati, V.K.; Krishnan, M. Investigation of Immunogenic Properties of Hemolin from Silkworm, Bombyx Mori as Carrier Protein: An Immunoinformatic Approach. Sci. Rep. 2018, 8, 6957. [Google Scholar] [CrossRef] [PubMed]
- Shiwang, L.; Li, W.; Liu, S.; Xu, J. RaptorX-Property: A Web Server for Protein Structure Property Prediction. Nucleic Acids Res. 2016, 44, W430–W435. [Google Scholar] [CrossRef]
- Zhang, H.-T.; Jiang, J.-Q.; Wang, Z.-L.; Chang, X.-Y.; Liu, X.-Y.; Wang, S.-H.; Zhao, K.; Chen, J.-S. Development of an Indirect Competitive ELISA for Simultaneous Detection of Enrofloxacin and Ciprofloxacin. J. Zhejiang Univ. Sci. B 2011, 12, 884–891. [Google Scholar] [CrossRef]
- Smith, S.M. Strategies for the Purification of Membrane Proteins. Methods Mol. Biol. 2017, 1485, 389–400. [Google Scholar] [CrossRef]
- Shams, M.; Nourmohammadi, H.; Asghari, A.; Basati, G.; Majidiani, H.; Naserifar, R.; Irannejad, H. Construction of a Multi-Epitope Protein for Human Toxocara Canis Detection: Immunoinformatics Approach Multi-Epitope Construct for T. Canis Serodiagnosis. Informatics Med. Unlocked 2021, 26, 100732. [Google Scholar] [CrossRef]
- Ottesen, E.A.; Skvaril, F.; Tripathy, S.P.; Poindexter, R.W.; Hussain, R. Prominence of IgG4 in the IgG Antibody Response to Human Filariasis. J. Immunol. 1985, 134, 2707–2712. [Google Scholar]
- Banla, M.; Tchalim, S.; Karabou, P.K.; Gantin, R.G.; Agba, A.I.; Kére-Banla, A.; Helling-Giese, G.; Heuschkel, C.; Schulz-Key, H.; Soboslay, P.T. Sustainable Control of Onchocerciasis: Ocular Pathology in Onchocerciasis Patients Treated Annually with Ivermectin for 23 Years: A Cohort Study. PLoS ONE 2014, 9, e98411. [Google Scholar] [CrossRef]
- Thiele, E.A.; Cama, V.A.; Sleshi, M.; Abanyie, F.; Lakwo, T.; Mekasha, S.; Cantey, P.T.; Kebede, A. Detection of Onchocerca volvulus in Skin Snips by Microscopy and Real-Time Polymerase Chain Reaction: Implications for Monitoring and Evaluation Activities. Am. J. Trop. Med. Hyg. 2016, 94, 906–911. [Google Scholar] [CrossRef] [PubMed]
- Macé, J.M.; Boussinesq, M.; Ngoumou, P.; Oye, J.E.; Koéranga, A.; Godin, C. Country-Wide Rapid Epidemiological Mapping of Onchocerciasis (REMO) in Cameroon. Ann. Trop. Med. Parasitol. 1997, 91, 379–391. [Google Scholar] [CrossRef] [PubMed]
- Cama, V.A.; Feleke, S.M.; McDonald, C.; Wiegand, R.E.; Cantey, P.T.; Arcury-Quandt, A.; Eberhard, M.; Abanyie, F.; Smith, J.; Jenks, M.H.; et al. Evaluation of an OV-16 IgG4 Enzyme-Linked Immunosorbent Assay in Humans and Its Application to Determine the Dynamics of Antibody Responses in a Non-Human Primate Model of Onchocerca volvulus Infection. Am. J. Trop. Med. Hyg. 2018, 99, 1041–1048. [Google Scholar] [CrossRef] [PubMed]
- Dieye, Y.; Storey, H.L.; Barrett, K.L.; Gerth-Guyette, E.; Di Giorgio, L.; Golden, A.; Faulx, D.; Kalnoky, M.; Ndiaye, M.K.N.; Sy, N.; et al. Feasibility of Utilizing the SD BIOLINE Onchocerciasis IgG4 Rapid Test in Onchocerciasis Surveillance in Senegal. PLOS Neglected Trop. Dis. 2017, 11, e0005884. [Google Scholar] [CrossRef]
- Shintouo, C.M.; Ghogomu, S.M.; Shey, R.A.; Hotterbeekx, A.; Yagmur, E.; Mets, T.; Vanhamme, L.; Colebunders, R.; Souopgui, J.; Njemini, R. Tandem Use of OvMANE1 and Ov-16 ELISA Tests Increases the Sensitivity for the Diagnosis of Human Onchocerciasis. Life 2021, 11, 1284. [Google Scholar] [CrossRef]
- Mackenzie, C.D.; Al-Qubati, Y.; Al-Kubati, A.-S.; Nutman, T.B.; Kubofcik, J.; Hopkins, A.; Behan-Braman, A. A Serological Survey of Human Onchocerciasis in Yemen. Am. J. Trop. Med. Hyg. 2018, 99, 1049–1052. [Google Scholar] [CrossRef]
- Van Regenmortel, M.H. Antigenicity and Immunogenicity of Synthetic Peptides. Biologicals 2001, 29, 209–213. [Google Scholar] [CrossRef]
- Weiss, N.; Karam, M. Evaluation of a Specific Enzyme Immunoassay for Onchocerciasis using a Low Molecular Weight Antigen Fraction of Onchocerca volvulus. Am. J. Trop. Med. Hyg. 1989, 40, 261–267. [Google Scholar] [CrossRef]
- Corradin, G.; Villard, V.; Kajava, A. Protein Structure Based Strategies for Antigen Discovery and Vaccine Development Against Malaria and Other Pathogens. Endocr. Metab. Immune Disord.—Drug Targets 2007, 7, 259–265. [Google Scholar] [CrossRef]
- Shintouo, C.M.; Nguve, J.E.; Asa, F.B.; Shey, R.A.; Kamga, J.; Souopgui, J.; Ghogomu, S.M.; Njemini, R. Entomological Assessment of Onchocerca Species Transmission by Black Flies in Selected Communities in the West Region of Cameroon. Pathogens 2020, 9, 722. [Google Scholar] [CrossRef]
- Aza’Ah, R.A.; Sumo, L.; Ntonifor, N.H.; Bopda, J.; Bamou, R.H.; Nana-Djeunga, H.C. Point Prevalence Mapping Reveals Hotspot for Onchocerciasis Transmission in the Ndikinimeki Health District, Centre Region, Cameroon. Parasites Vectors 2020, 13, 519. [Google Scholar] [CrossRef] [PubMed]
- Chandrashekar, R.; Ogunrinade, A.F.; Weil, G.J. Use of Recombinant Onchocerca volvulus Antigens for Diagnosis and Surveillance of Human Onchocerciasis. Trop. Med. Int. Health 2007, 1, 575–580. [Google Scholar] [CrossRef] [PubMed]
- Mpagi, J.L.; Buttner, D.W.; Tischendorf, F.W.; Erttmann, K.D.; Brattig, N.W. Humoral Responses to a Secretory Onchocerca volvulus Protein: Differences in the Pattern of Antibody Isotypes to Recombinant Ov20/OvS1 in Generalized and Hyperreactive Onchocerciasis. Parasite Immunol. 2000, 22, 455–460. [Google Scholar] [CrossRef] [PubMed]
- Hussain, R.; Ottesen, E.A. IgE Responses in Human Filariasis. IV. Parallel Antigen Recognition by IgE and IgG4 Subclass Antibodies. J. Immunol. 1986, 136, 1859–1863. [Google Scholar] [PubMed]

| S/N | Protein ID | Antigenicity (Cut off > 0.4999) | Linear B-Epitopes Selected | |||
|---|---|---|---|---|---|---|
| ANTIGENpro | Vaxijen 2.0 | Remarks | Linear B-Epitopes | Antigenicity on Vaxijen 2.0 | ||
| 1 | OVOC7606 | 0.493181 | 0.3129 | Non antigenic | None | - |
| 2 | OVOC9989 | 0.247895 | 0.4514 | Non antigenic | None | - |
| 3 | OVOC10207 | 0.377321 | 0.2670 | Non antigenic | None | - |
| 4 | OVOC5574 | 0.936181 | 0.4886 | May be antigenic | IPQQNGEFTGSSLHEMTAKD (LBE 2) | 0.7725 |
| 5 | OVOC5909 | 0.605038 | 0.7636 | Antigenic | TVTSESSKTTPITESSA (LBE 9) | 1.0945 |
| KTVTSESSKTTPITESSAT (LBE 1) | 1.0717 | |||||
| SPSITSPKSRITTESPST (LBE 5) | 1.1604 | |||||
| 6 | OVOC8498 | 0.678848 | 0.8364 | Antigenic | FRLPHDIWEPPF (LBE 3) | 0.8433 |
| 7 | OVOC8529 | 0.844335 | 0.5419 | Antigenic | KKSFNKKMTKKKTFHGDKLK (LBE 7) | 0.7077 |
| 8 | OVOC8936 | 0.732818 | 0.5409 | Antigenic | VQPQFIRVQTLKSQ (LBE 8) | 0.6061 |
| 9 | OVOC10037 | 0.844335 | 1.1807 | Antigenic | EGSKNETEKFANSTKEDEKITPL (LBE 6) | 1.1520 |
| EKSTEASKEEKKSGEVVKEEVQT (LBE 4) | 1.4237 | |||||
| Total IgG | ||
|---|---|---|
| ROC Curve Analysis | ROC curve area (AUC) | 0.9995 |
| 95% CI of AUC | 0.9976 to 1.000 | |
| p-value (against AUC = 0.5) | <0.0001 | |
| Diagnostic Accuracy Parameter | Cut off value | 0.3503 |
| Sensitivity (%) (95% CI) | 98.4 (91.54% to 99.92%) | |
| Specificity (%) (95% CI) | 100.0 (88.30% to 100.00%) | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yengo, B.N.; Shintouo, C.M.; Hotterbeekx, A.; Yaah, N.E.; Shey, R.A.; Quanico, J.; Baggerman, G.; Ayong, L.; Vanhamme, L.; Njemini, R.; et al. Immunoinformatics Design and Assessment of a Multiepitope Antigen (OvMCBL02) for Onchocerciasis Diagnosis and Monitoring. Diagnostics 2022, 12, 1440. https://doi.org/10.3390/diagnostics12061440
Yengo BN, Shintouo CM, Hotterbeekx A, Yaah NE, Shey RA, Quanico J, Baggerman G, Ayong L, Vanhamme L, Njemini R, et al. Immunoinformatics Design and Assessment of a Multiepitope Antigen (OvMCBL02) for Onchocerciasis Diagnosis and Monitoring. Diagnostics. 2022; 12(6):1440. https://doi.org/10.3390/diagnostics12061440
Chicago/Turabian StyleYengo, Bernis Neneyoh, Cabirou Mounchili Shintouo, An Hotterbeekx, Ntang Emmaculate Yaah, Robert Adamu Shey, Jusal Quanico, Geert Baggerman, Lawrence Ayong, Luc Vanhamme, Rose Njemini, and et al. 2022. "Immunoinformatics Design and Assessment of a Multiepitope Antigen (OvMCBL02) for Onchocerciasis Diagnosis and Monitoring" Diagnostics 12, no. 6: 1440. https://doi.org/10.3390/diagnostics12061440
APA StyleYengo, B. N., Shintouo, C. M., Hotterbeekx, A., Yaah, N. E., Shey, R. A., Quanico, J., Baggerman, G., Ayong, L., Vanhamme, L., Njemini, R., Souopgui, J., Colebunders, R., & Ghogomu, S. M. (2022). Immunoinformatics Design and Assessment of a Multiepitope Antigen (OvMCBL02) for Onchocerciasis Diagnosis and Monitoring. Diagnostics, 12(6), 1440. https://doi.org/10.3390/diagnostics12061440











