A Deep Learning Model for Prostate Adenocarcinoma Classification in Needle Biopsy Whole-Slide Images Using Transfer Learning
Abstract
:1. Introduction
2. Materials and Methods
2.1. Clinical Cases and Pathological Records
2.2. Dataset
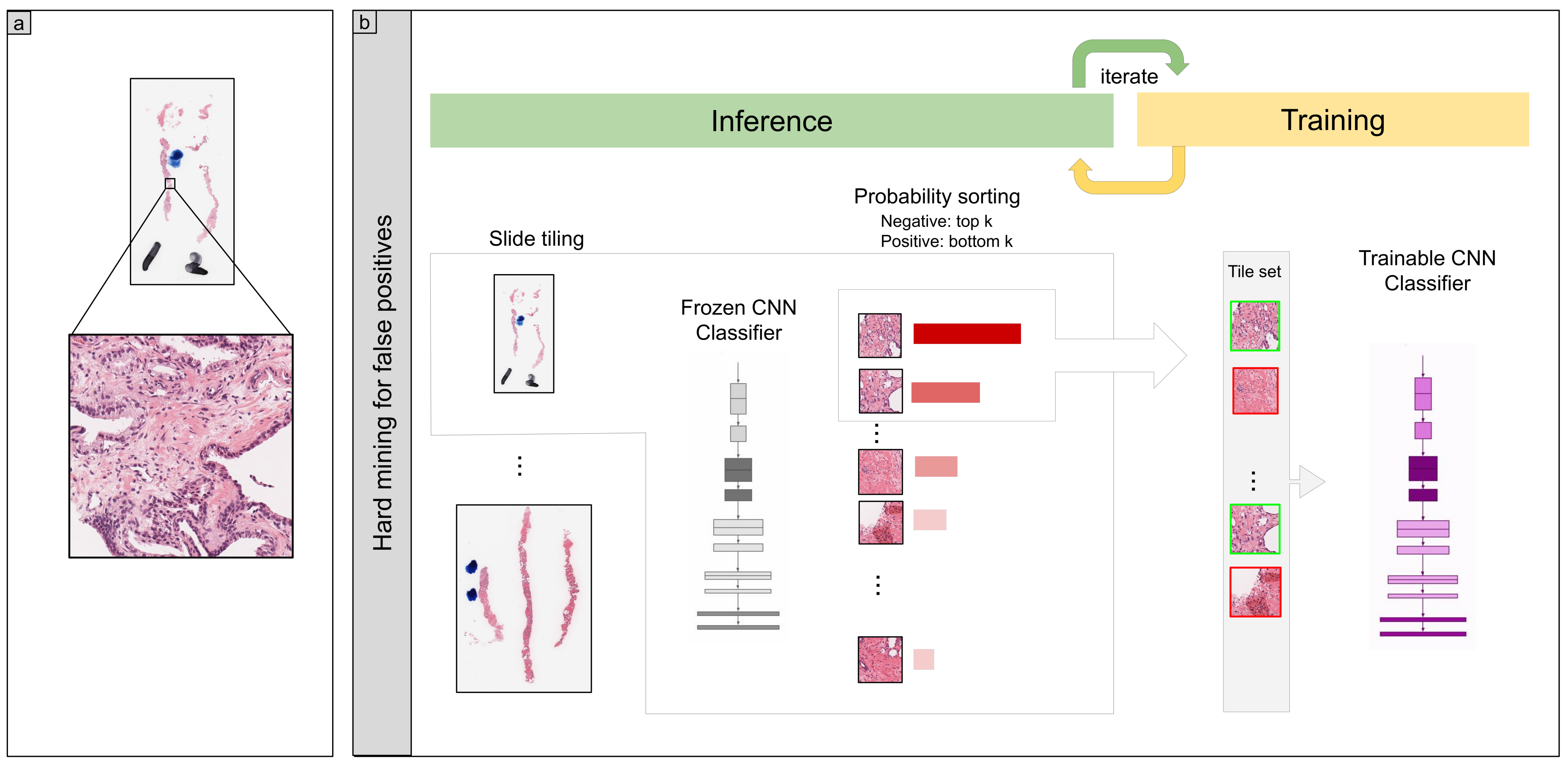
2.3. Deep Learning Models
2.4. Software and Statistical Analysis
2.5. Code Availability
3. Results
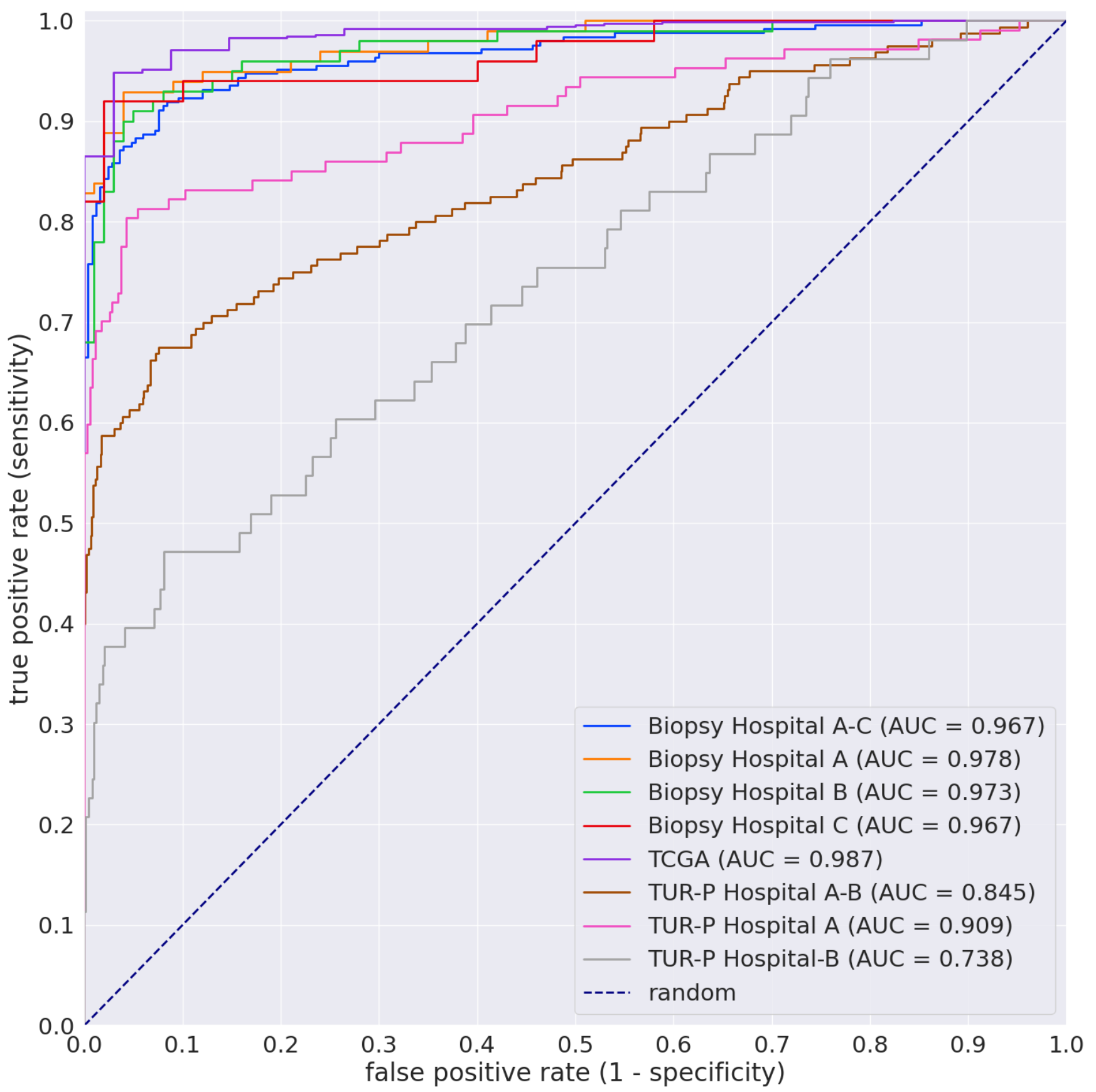
3.1. High AUC Performance of the WSI Evaluation of Prostate Adenocarcinoma Histopathology Images in the Needle Biopsy, TUR-P, and TCGA Test Sets
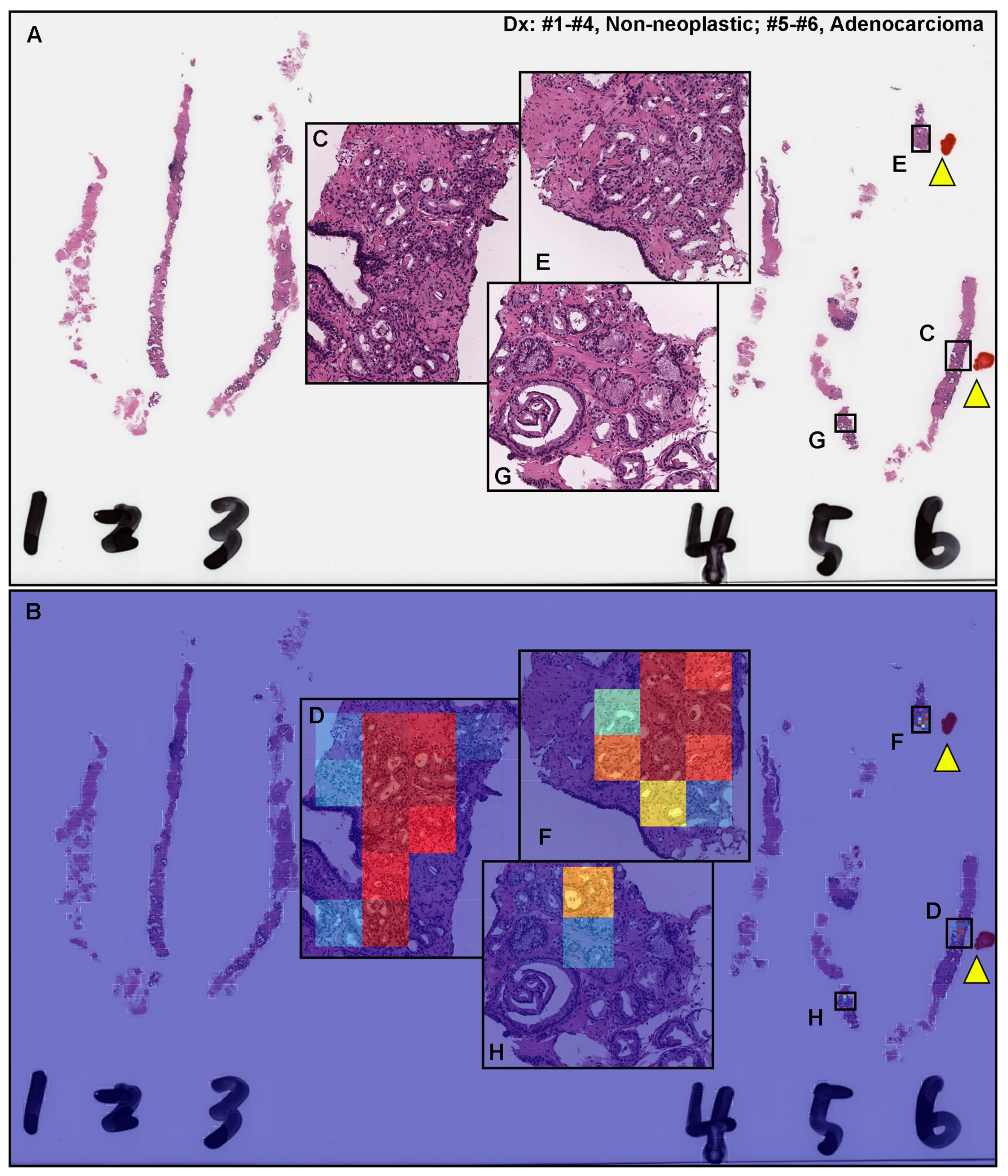
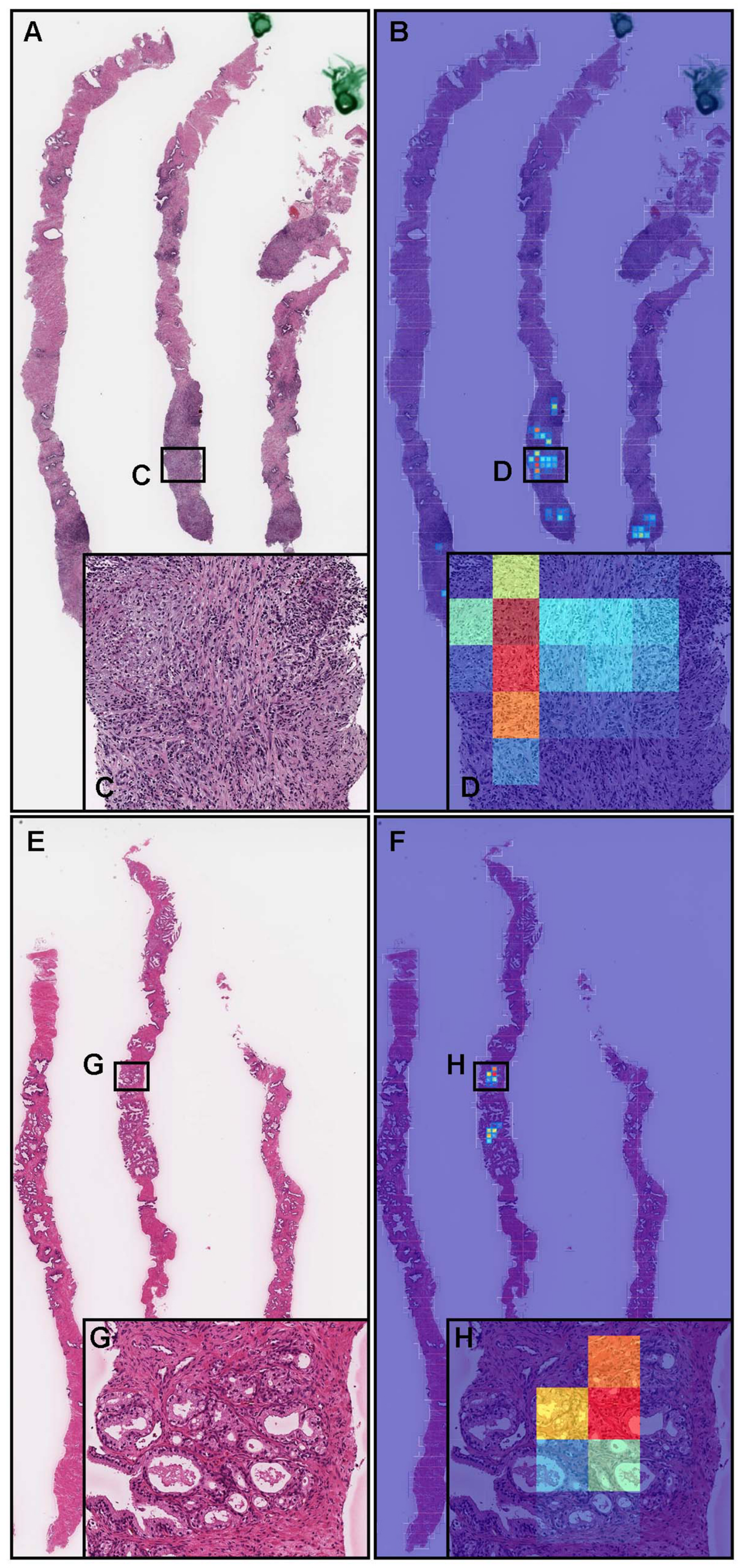
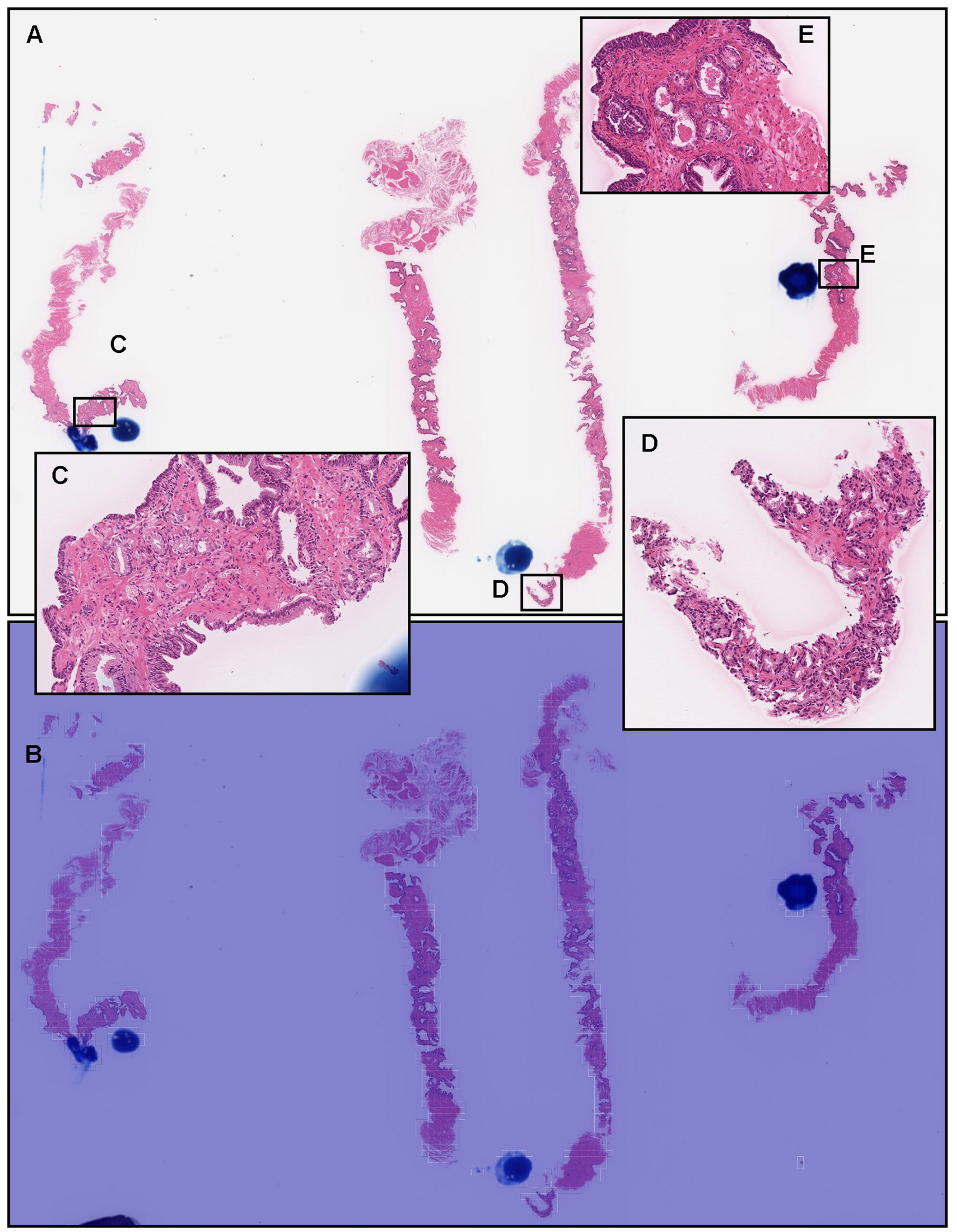
3.2. True Positive Prediction on Needle Biopsy Specimens
3.3. False Positive Prediction on Needle Biopsy Specimens
3.4. False Negative Prediction on the Needle Biopsy Specimens
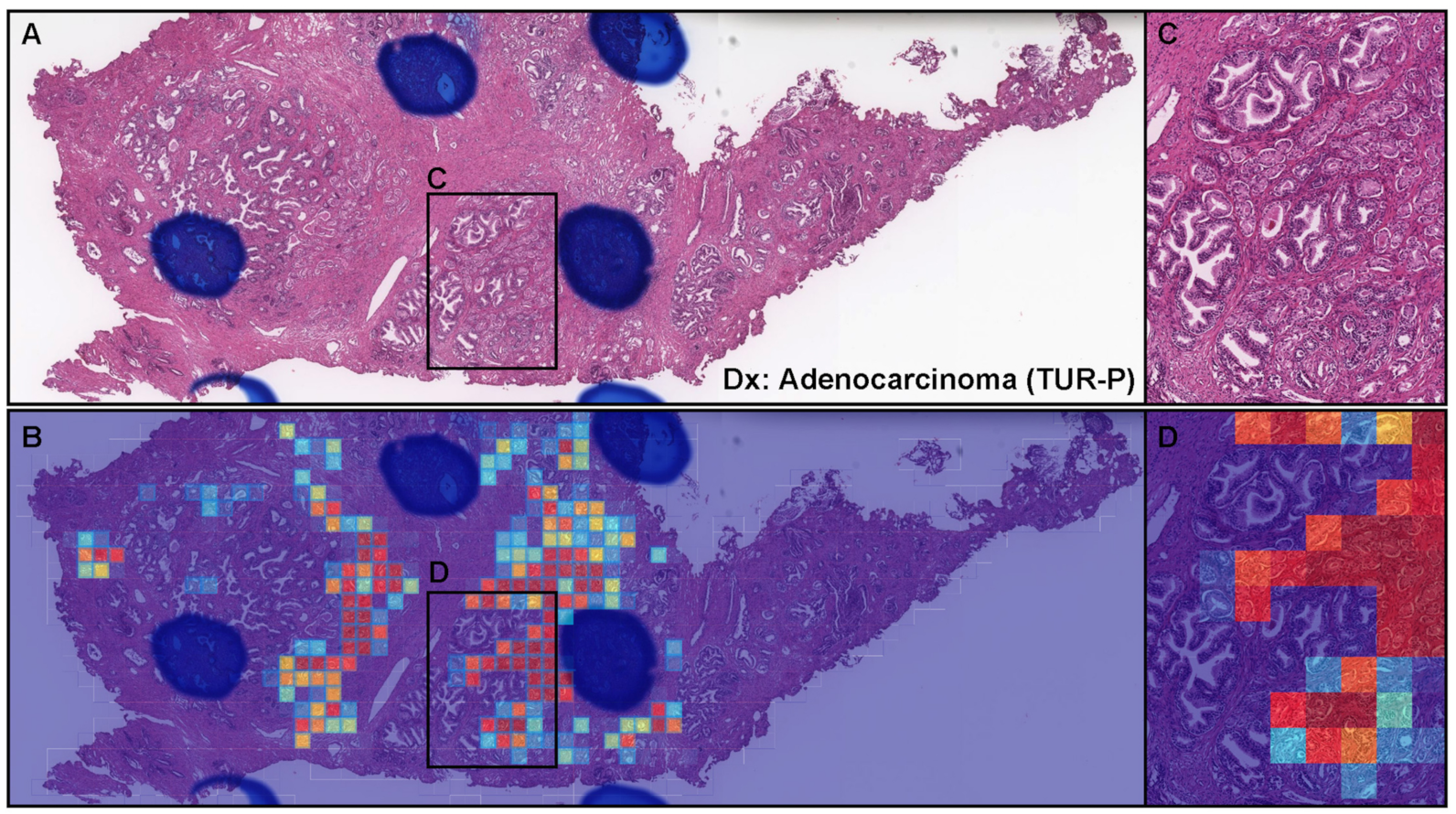
3.5. True Positive Prediction on the TUR-P Specimens
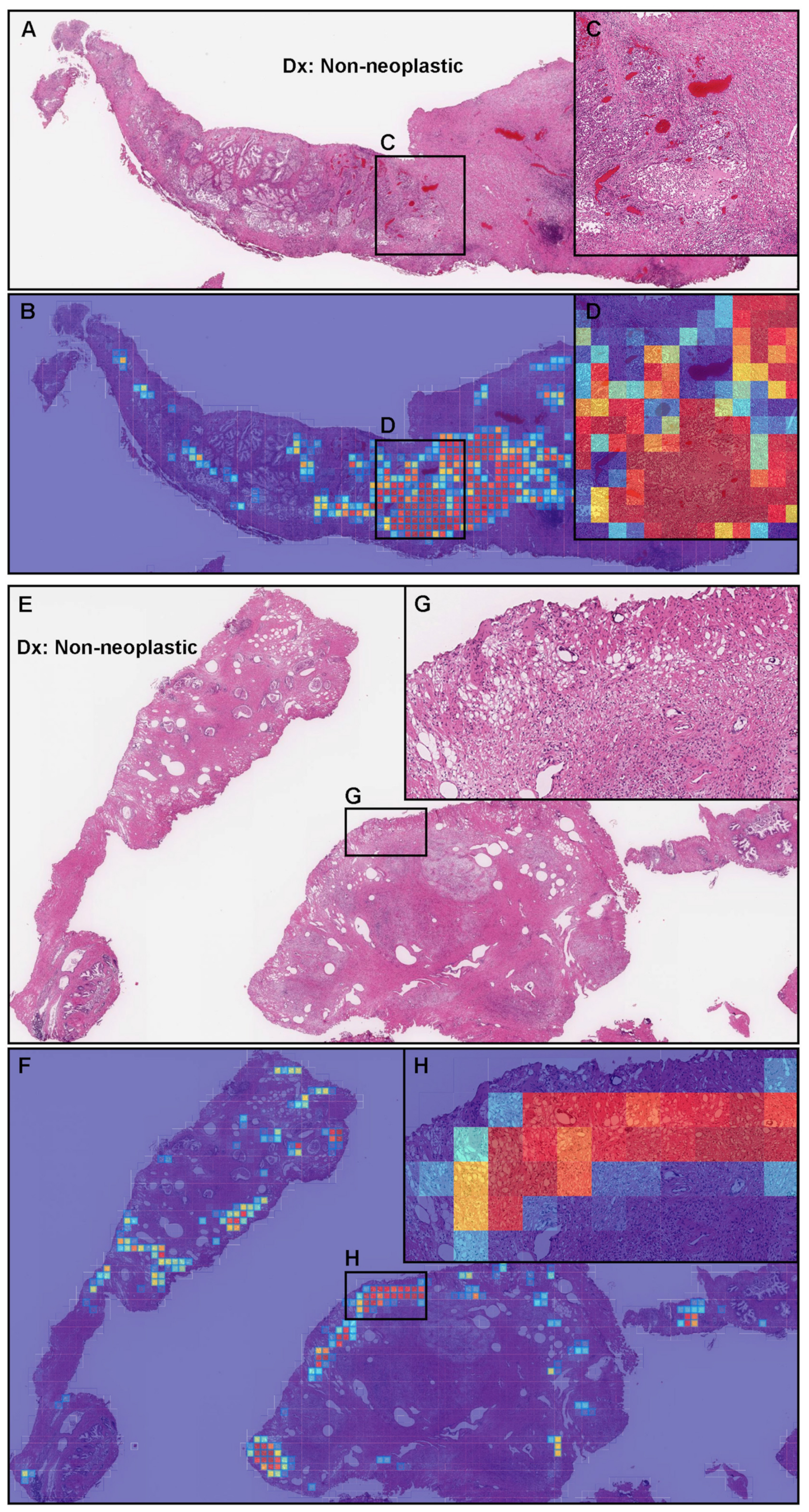
3.6. False Positive Prediction on TUR-P Specimens
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Montironi, R.; Mazzucchelli, R.; Algaba, F.; Bostwick, D.G.; Krongrad, A. Prostate-specific antigen as a marker of prostate disease. Virchows Arch. 2000, 436, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Sershon, P.D.; Barry, M.J.; Oesterling, J.E. Serum prostate-specific antigen discriminates weakly between men with benign prostatic hyperplasia and patients with organ-confined prostate cancer. Eur. Urol. 1994, 25, 281–287. [Google Scholar] [CrossRef] [PubMed]
- Nadler, R.B.; Humphrey, P.A.; Smith, D.S.; Catalona, W.J.; Ratliff, T.L. Effect of inflammation and benign prostatic hyperplasia on elevated serum prostate specific antigen levels. J. Urol. 1995, 154, 407–413. [Google Scholar] [CrossRef]
- Oesterling, J.E.; Jacobsen, S.J.; Chute, C.G.; Guess, H.A.; Girman, C.J.; Panser, L.A.; Lieber, M.M. Serum prostate-specific antigen in a community-based population of healthy men: Establishment of age-specific reference ranges. JAMA 1993, 270, 860–864. [Google Scholar] [CrossRef]
- Peller, P.A.; Young, D.C.; Marmaduke, D.P.; Marsh, W.L.; Badalament, R.A. Sextant prostate biopsies. A histopathologic correlation with radical prostatectomy specimens. Cancer 1995, 75, 530–538. [Google Scholar] [CrossRef]
- Eichler, K.; Hempel, S.; Wilby, J.; Myers, L.; Bachmann, L.M.; Kleijnen, J. Diagnostic value of systematic biopsy methods in the investigation of prostate cancer: A systematic review. J. Urol. 2006, 175, 1605–1612. [Google Scholar] [CrossRef]
- Bjurlin, M.A.; Carter, H.B.; Schellhammer, P.; Cookson, M.S.; Gomella, L.G.; Troyer, D.; Wheeler, T.M.; Schlossberg, S.; Penson, D.F.; Taneja, S.S. Optimization of initial prostate biopsy in clinical practice: Sampling, labeling and specimen processing. J. Urol. 2013, 189, 2039–2046. [Google Scholar] [CrossRef] [Green Version]
- Yu, K.H.; Zhang, C.; Berry, G.J.; Altman, R.B.; Ré, C.; Rubin, D.L.; Snyder, M. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat. Commun. 2016, 7, 12474. [Google Scholar] [CrossRef] [Green Version]
- Hou, L.; Samaras, D.; Kurc, T.M.; Gao, Y.; Davis, J.E.; Saltz, J.H. Patch-based convolutional neural network for whole slide tissue image classification. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 2424–2433. [Google Scholar]
- Madabhushi, A.; Lee, G. Image analysis and machine learning in digital pathology: Challenges and opportunities. Med. Image Anal. 2016, 33, 170–175. [Google Scholar] [CrossRef] [Green Version]
- Litjens, G.; Sánchez, C.I.; Timofeeva, N.; Hermsen, M.; Nagtegaal, I.; Kovacs, I.; Hulsbergen-Van De Kaa, C.; Bult, P.; Van Ginneken, B.; Van Der Laak, J. Deep learning as a tool for increased accuracy and efficiency of histopathological diagnosis. Sci. Rep. 2016, 6, 26286. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kraus, O.Z.; Ba, J.L.; Frey, B.J. Classifying and segmenting microscopy images with deep multiple instance learning. Bioinformatics 2016, 32, i52–i59. [Google Scholar] [CrossRef] [PubMed]
- Korbar, B.; Olofson, A.M.; Miraflor, A.P.; Nicka, C.M.; Suriawinata, M.A.; Torresani, L.; Suriawinata, A.A.; Hassanpour, S. Deep learning for classification of colorectal polyps on whole-slide images. J. Pathol. Inform. 2017, 8, 30. [Google Scholar] [PubMed]
- Luo, X.; Zang, X.; Yang, L.; Huang, J.; Liang, F.; Rodriguez-Canales, J.; Wistuba, I.I.; Gazdar, A.; Xie, Y.; Xiao, G. Comprehensive computational pathological image analysis predicts lung cancer prognosis. J. Thorac. Oncol. 2017, 12, 501–509. [Google Scholar] [CrossRef] [Green Version]
- Coudray, N.; Ocampo, P.S.; Sakellaropoulos, T.; Narula, N.; Snuderl, M.; Fenyö, D.; Moreira, A.L.; Razavian, N.; Tsirigos, A. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat. Med. 2018, 24, 1559–1567. [Google Scholar] [CrossRef]
- Wei, J.W.; Tafe, L.J.; Linnik, Y.A.; Vaickus, L.J.; Tomita, N.; Hassanpour, S. Pathologist-level classification of histologic patterns on resected lung adenocarcinoma slides with deep neural networks. Sci. Rep. 2019, 9, 3358. [Google Scholar] [CrossRef] [Green Version]
- Gertych, A.; Swiderska-Chadaj, Z.; Ma, Z.; Ing, N.; Markiewicz, T.; Cierniak, S.; Salemi, H.; Guzman, S.; Walts, A.E.; Knudsen, B.S. Convolutional neural networks can accurately distinguish four histologic growth patterns of lung adenocarcinoma in digital slides. Sci. Rep. 2019, 9, 1483. [Google Scholar] [CrossRef]
- Bejnordi, B.E.; Veta, M.; Van Diest, P.J.; Van Ginneken, B.; Karssemeijer, N.; Litjens, G.; Van Der Laak, J.A.; Hermsen, M.; Manson, Q.F.; Balkenhol, M.; et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 2017, 318, 2199–2210. [Google Scholar] [CrossRef]
- Saltz, J.; Gupta, R.; Hou, L.; Kurc, T.; Singh, P.; Nguyen, V.; Samaras, D.; Shroyer, K.R.; Zhao, T.; Batiste, R.; et al. Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep. 2018, 23, 181–193. [Google Scholar] [CrossRef] [Green Version]
- Campanella, G.; Hanna, M.G.; Geneslaw, L.; Miraflor, A.; Silva, V.W.K.; Busam, K.J.; Brogi, E.; Reuter, V.E.; Klimstra, D.S.; Fuchs, T.J. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 2019, 25, 1301–1309. [Google Scholar] [CrossRef]
- Iizuka, O.; Kanavati, F.; Kato, K.; Rambeau, M.; Arihiro, K.; Tsuneki, M. Deep learning models for histopathological classification of gastric and colonic epithelial tumours. Sci. Rep. 2020, 10, 1504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lucas, M.; Jansen, I.; Savci-Heijink, C.D.; Meijer, S.L.; de Boer, O.J.; van Leeuwen, T.G.; de Bruin, D.M.; Marquering, H.A. Deep learning for automatic Gleason pattern classification for grade group determination of prostate biopsies. Virchows Arch. 2019, 475, 77–83. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, H.; Park, S.; Hwang, T.H. Computerized classification of prostate cancer gleason scores from whole slide images. IEEE/ACM Trans. Comput. Biol. Bioinform. 2019, 17, 1871–1882. [Google Scholar] [CrossRef] [PubMed]
- Raciti, P.; Sue, J.; Ceballos, R.; Godrich, R.; Kunz, J.D.; Kapur, S.; Reuter, V.; Grady, L.; Kanan, C.; Klimstra, D.S.; et al. Novel artificial intelligence system increases the detection of prostate cancer in whole slide images of core needle biopsies. Mod. Pathol. 2020, 33, 2058–2066. [Google Scholar] [CrossRef]
- Rana, A.; Lowe, A.; Lithgow, M.; Horback, K.; Janovitz, T.; Da Silva, A.; Tsai, H.; Shanmugam, V.; Bayat, A.; Shah, P. Use of deep learning to develop and analyze computational hematoxylin and eosin staining of prostate core biopsy images for tumor diagnosis. JAMA Netw. Open 2020, 3, e205111. [Google Scholar] [CrossRef]
- Silva-Rodríguez, J.; Colomer, A.; Dolz, J.; Naranjo, V. Self-learning for weakly supervised gleason grading of local patterns. IEEE J. Biomed. Health Inform. 2021, 25, 3094–3104. [Google Scholar] [CrossRef]
- da Silva, L.M.; Pereira, E.M.; Salles, P.G.; Godrich, R.; Ceballos, R.; Kunz, J.D.; Casson, A.; Viret, J.; Chandarlapaty, S.; Ferreira, C.G.; et al. Independent real-world application of a clinical-grade automated prostate cancer detection system. J. Pathol. 2021, 254, 147–158. [Google Scholar] [CrossRef]
- Otálora, S.; Marini, N.; Müller, H.; Atzori, M. Combining weakly and strongly supervised learning improves strong supervision in Gleason pattern classification. BMC Med. Imaging 2021, 21, 1–14. [Google Scholar] [CrossRef]
- Hammouda, K.; Khalifa, F.; El-Melegy, M.; Ghazal, M.; Darwish, H.E.; El-Ghar, M.A.; El-Baz, A. A Deep Learning Pipeline for Grade Groups Classification Using Digitized Prostate Biopsy Specimens. Sensors 2021, 21, 6708. [Google Scholar] [CrossRef]
- Kanavati, F.; Toyokawa, G.; Momosaki, S.; Rambeau, M.; Kozuma, Y.; Shoji, F.; Yamazaki, K.; Takeo, S.; Iizuka, O.; Tsuneki, M. Weakly-supervised learning for lung carcinoma classification using deep learning. Sci. Rep. 2020, 10, 9297. [Google Scholar] [CrossRef]
- Naito, Y.; Tsuneki, M.; Fukushima, N.; Koga, Y.; Higashi, M.; Notohara, K.; Aishima, S.; Ohike, N.; Tajiri, T.; Yamaguchi, H.; et al. A deep learning model to detect pancreatic ductal adenocarcinoma on endoscopic ultrasound-guided fine-needle biopsy. Sci. Rep. 2021, 11, 8454. [Google Scholar] [CrossRef] [PubMed]
- Kanavati, F.; Ichihara, S.; Rambeau, M.; Iizuka, O.; Arihiro, K.; Tsuneki, M. Deep learning models for gastric signet ring cell carcinoma classification in whole slide images. Technol. Cancer Res. Treat. 2021, 20, 15330338211027901. [Google Scholar] [CrossRef] [PubMed]
- Kanavati, F.; Tsuneki, M. A deep learning model for gastric diffuse-type adenocarcinoma classification in whole slide images. arXiv, 2021; arXiv:2104.12478. [Google Scholar]
- Kanavati, F.; Tsuneki, M. Breast invasive ductal carcinoma classification on whole slide images with weakly-supervised and transfer learning. Cancers 2021, 13, 5368. [Google Scholar] [CrossRef] [PubMed]
- Tsuneki, M.; Kanavati, F. Deep learning models for poorly differentiated colorectal adenocarcinoma classification in whole slide images using transfer learning. Diagnostics 2021, 11, 2074. [Google Scholar] [CrossRef]
- Kanavati, F.; Ichihara, S.; Tsuneki, M. A deep learning model for breast ductal carcinoma in situ classification in whole slide images. Virchows Arch. 2022, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Kanavati, F.; Tsuneki, M. Partial transfusion: On the expressive influence of trainable batch norm parameters for transfer learning. arXiv 2021, arXiv:2102.05543. [Google Scholar]
- Tan, M.; Le, Q. Efficientnet: Rethinking model scaling for convolutional neural networks. In Proceedings of the International Conference on Machine Learning, PMLR, Long Beach, CA, USA, 9–15 June 2019; pp. 6105–6114. [Google Scholar]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef] [Green Version]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Systems. 2015. Available online: tensorflow.org (accessed on 1 March 2020).
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Efron, B.; Tibshirani, R.J. An Introduction to the Bootstrap; CRC Press: Boca Raton, FL, USA, 1994. [Google Scholar]
- Epstein, J.I.; Egevad, L.; Amin, M.B.; Delahunt, B.; Srigley, J.R.; Humphrey, P.A. The 2014 International Society of Urological Pathology (ISUP) consensus conference on Gleason grading of prostatic carcinoma. Am. J. Surg. Pathol. 2016, 40, 244–252. [Google Scholar] [CrossRef] [PubMed]
- Kweldam, C.; van Leenders, G.; van der Kwast, T. Grading of prostate cancer: A work in progress. Histopathology 2019, 74, 146–160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gaudin, P.; Reuter, V. Benign mimics of prostatic adenocarcinoma on needle biopsy. Anat. Pathol. 1997, 2, 111–134. [Google Scholar] [PubMed]
- Egan, A.M.; Lopez-Beltran, A.; Bostwick, D.G. Prostatic adenocarcinoma with atrophic features: Malignancy mimicking a benign process. Am. J. Surg. Pathol. 1997, 21, 931–935. [Google Scholar] [CrossRef]
| Adenocarcinoma | Benign | Total | ||
|---|---|---|---|---|
| Training set | Hospital A | 144 | 260 | 404 |
| Hospital B | 100 | 75 | 175 | |
| Hospital C | 115 | 159 | 274 | |
| Hospital D | 56 | 118 | 174 | |
| Hospital E | 23 | 72 | 95 | |
| Total | 438 | 684 | 1122 | |
| Validation set | Hospital A | 6 | 6 | 12 |
| Hospital B | 6 | 6 | 12 | |
| Hospital C | 6 | 6 | 12 | |
| Hospital D | 6 | 6 | 12 | |
| Hospital E | 6 | 6 | 12 | |
| Total | 30 | 30 | 60 |
| Adenocarcinoma | Benign | Total | ||
|---|---|---|---|---|
| Biopsy | Hospitals A–C | 250 | 250 | 500 |
| Hospital A | 100 | 100 | 200 | |
| Hospital B | 100 | 100 | 200 | |
| Hospital C | 50 | 50 | 100 | |
| TUR-P | Hospitals A–B | 162 | 1082 | 1244 |
| Hospital A | 109 | 352 | 461 | |
| Hospital B | 53 | 730 | 783 | |
| Public dataset | TCGA | 733 | 34 | 768 |
| Existing Models | ROC-AUC | Log Loss |
|---|---|---|
| Breast IDC (10×, 512) | 0.704 [0.659–0.751] | 0.947 [0.816–1.064] |
| Breast IDC, DCIS (10×, 224) | 0.692 [0.644–0.735] | 1.413 [1.282–1.566] |
| Colon ADC, AD (10×, 512) | 0.553 [0.507–0.611] | 1.525 [1.350–1.711] |
| Colon poorly ADC-1 (20×, 512) | 0.795 [0.756–0.835] | 0.572 [0.513–0.637] |
| Colon poorly ADC-2 (20×, 512) | 0.817 [0.782–0.855] | 0.522 [0.475–0.569] |
| Stomach ADC, AD (10×, 512) | 0.706 [0.662–0.753] | 1.391 [1.248–1.569] |
| Stomach poorly ADC (20×, 224) | 0.724 [0.681–0.767] | 0.598 [0.565–0.629] |
| Stomach SRCC (10×, 224) | 0.804 [0.763–0.839] | 0.998 [0.894–1.114] |
| Pancreas EUS-FNA ADC (10×, 224) | 0.774 [0.735–0.817] | 0.587 [0.544–0.629] |
| Lung carcinoma (10×, 512) | 0.702 [0.659–0.751] | 1.398 [1.2560–1.546] |
| TL-Colon Poorly ADC-2 (20×, 512) | |||
|---|---|---|---|
| ROC-AUC | Log-Loss | ||
| Biopsy | Hospitals A–C | 0.967 [0.955–0.982] | 0.288 [0.210–0.354] |
| Hospital A | 0.978 [0.966–0.995] | 0.209 [0.117–0.276] | |
| Hospital B | 0.972 [0.948–0.988] | 0.378 [0.276–0.536] | |
| Hospital C | 0.967 [0.922–0.993] | 0.265 [0.117–0.512] | |
| TUR-P | Hospitals A–B | 0.845 [0.806–0.883] | 4.152 [4.047–4.253] |
| Hospital A | 0.909 [0.865–0.947] | 3.269 [3.089–3.451] | |
| Hospital B | 0.737 [0.657–0.810] | 4.672 [4.559–4.798] | |
| Public dataset | TCGA | 0.987 [0.977–0.995] | 0.074 [0.055–0.095] |
| EfficientNetB1 (20×, 512) | |||
| ROC-AUC | Log-Loss | ||
| Biopsy | Hospitals A–C | 0.971 [0.955–0.982] | 0.256 [0.188–0.349] |
| Hospital A | 0.979 [0.962–0.993] | 0.209 [0.110–0.322] | |
| Hospital B | 0.978 [0.963–0.992] | 0.279 [0.167–0.398] | |
| Hospital C | 0.977 [0.959–1.000] | 0.306 [0.037–0.406] | |
| TUR-P | Hospitals A–B | 0.803 [0.765–0.848] | 5.113 [4.976–5.252] |
| Hospital A | 0.875 [0.834–0.923] | 4.308 [4.059–4.550] | |
| Hospital B | 0.670 [0.597–0.753] | 5.588 [5.411–5.729] | |
| Public dataset | TCGA | 0.945 [0.912–0.973] | 0.101 [0.067–0.147] |
| EfficientNetB1 (10×, 224) | |||
| ROC-AUC | Log-Loss | ||
| Biopsy | Hospitals A–C | 0.739 [0.691–0.783] | 0.631 [0.545–0.724] |
| Hospital A | 0.751 [0.668–0.810] | 0.605 [0.511–0.744] | |
| Hospital B | 0.929 [0.885–0.970] | 0.335 [0.223–0.427] | |
| Hospital C | 0.472 [0.348–0.572] | 1.278 [0.979–1.501] | |
| TUR-P | Hospitals A–B | 0.804 [0.760–0.847] | 0.392 [0.369–0.417] |
| Hospital A | 0.771 [0.705–0.820] | 0.424 [0.384–0.474] | |
| Hospital B | 0.928 [0.859–0.980] | 0.373 [0.347–0.408] | |
| Public dataset | TCGA | 0.578 [0.497–0.661] | 1.575 [1.481–1.657] |
| Accuracy | Sensitivity | Specificity | F1-Score | ||
|---|---|---|---|---|---|
| Biopsy | Hospitals A–C | 0.918 [0.894–0.942] | 0.912 [0.878–0.946] | 0.924 [0.888–0.955] | 0.918 [0.889–0.941] |
| Hospital A | 0.945 [0.920–0.980] | 0.930 [0.897–0.989] | 0.960 [0.915–0.991] | 0.944 [0.920–0.981] | |
| Hospital B | 0.925 [0.885–0.955] | 0.890 [0.824–0.944] | 0.960 [0.912–0.991] | 0.922 [0.878–0.955] | |
| Hospital C | 0.940 [0.880–0.980] | 0.900 [0.796–0.964] | 0.980 [0.921–1.000] | 0.938 [0.870–0.978] | |
| TUR-P | Hospitals A–B | 0.894 [0.866–0.922] | 0.700 [0.603–0.813] | 0.926 [0.896–0.950] | 0.618 [0.561–0.675] |
| Hospital A | 0.918 [0.889–0.939] | 0.798 [0.712–0.867] | 0.955 [0.930–0.975] | 0.821 [0.749–0.871] | |
| Hospital B | 0.890 [0.867–0.909] | 0.415 [0.265–0.529] | 0.925 [0.906–0.940] | 0.339 [0.212–0.424] | |
| Public dataset | TCGA | 0.949 [0.934–0.965] | 0.948 [0.932–0.965] | 0.971 [0.906–1.000] | 0.973 [0.964–0.981] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsuneki, M.; Abe, M.; Kanavati, F. A Deep Learning Model for Prostate Adenocarcinoma Classification in Needle Biopsy Whole-Slide Images Using Transfer Learning. Diagnostics 2022, 12, 768. https://doi.org/10.3390/diagnostics12030768
Tsuneki M, Abe M, Kanavati F. A Deep Learning Model for Prostate Adenocarcinoma Classification in Needle Biopsy Whole-Slide Images Using Transfer Learning. Diagnostics. 2022; 12(3):768. https://doi.org/10.3390/diagnostics12030768
Chicago/Turabian StyleTsuneki, Masayuki, Makoto Abe, and Fahdi Kanavati. 2022. "A Deep Learning Model for Prostate Adenocarcinoma Classification in Needle Biopsy Whole-Slide Images Using Transfer Learning" Diagnostics 12, no. 3: 768. https://doi.org/10.3390/diagnostics12030768
APA StyleTsuneki, M., Abe, M., & Kanavati, F. (2022). A Deep Learning Model for Prostate Adenocarcinoma Classification in Needle Biopsy Whole-Slide Images Using Transfer Learning. Diagnostics, 12(3), 768. https://doi.org/10.3390/diagnostics12030768







