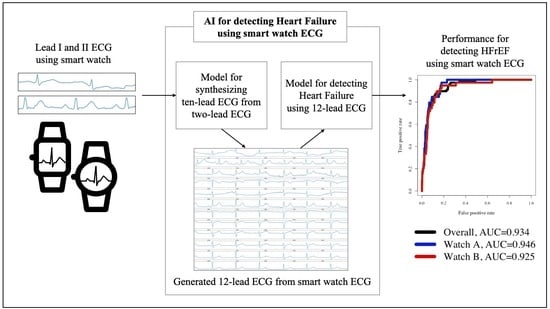
Artificial Intelligence-Enhanced Smartwatch ECG for Heart Failure-Reduced Ejection Fraction Detection by Generating 12-Lead ECG
Abstract
:1. Introduction
2. Methods
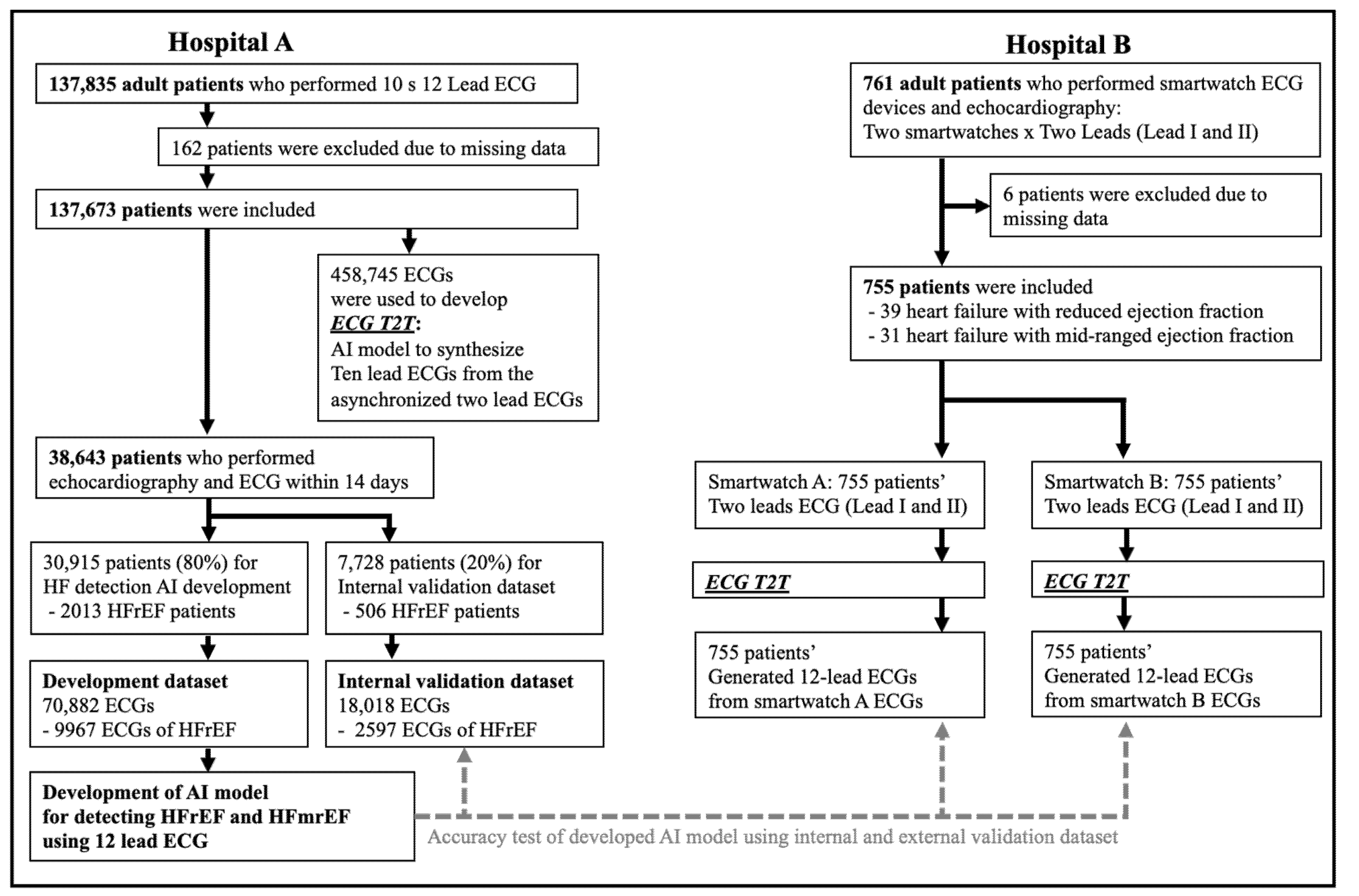
2.1. Data Source and Study Population
2.2. Outcomes and Predictive Variables
2.3. Data Preprocessing
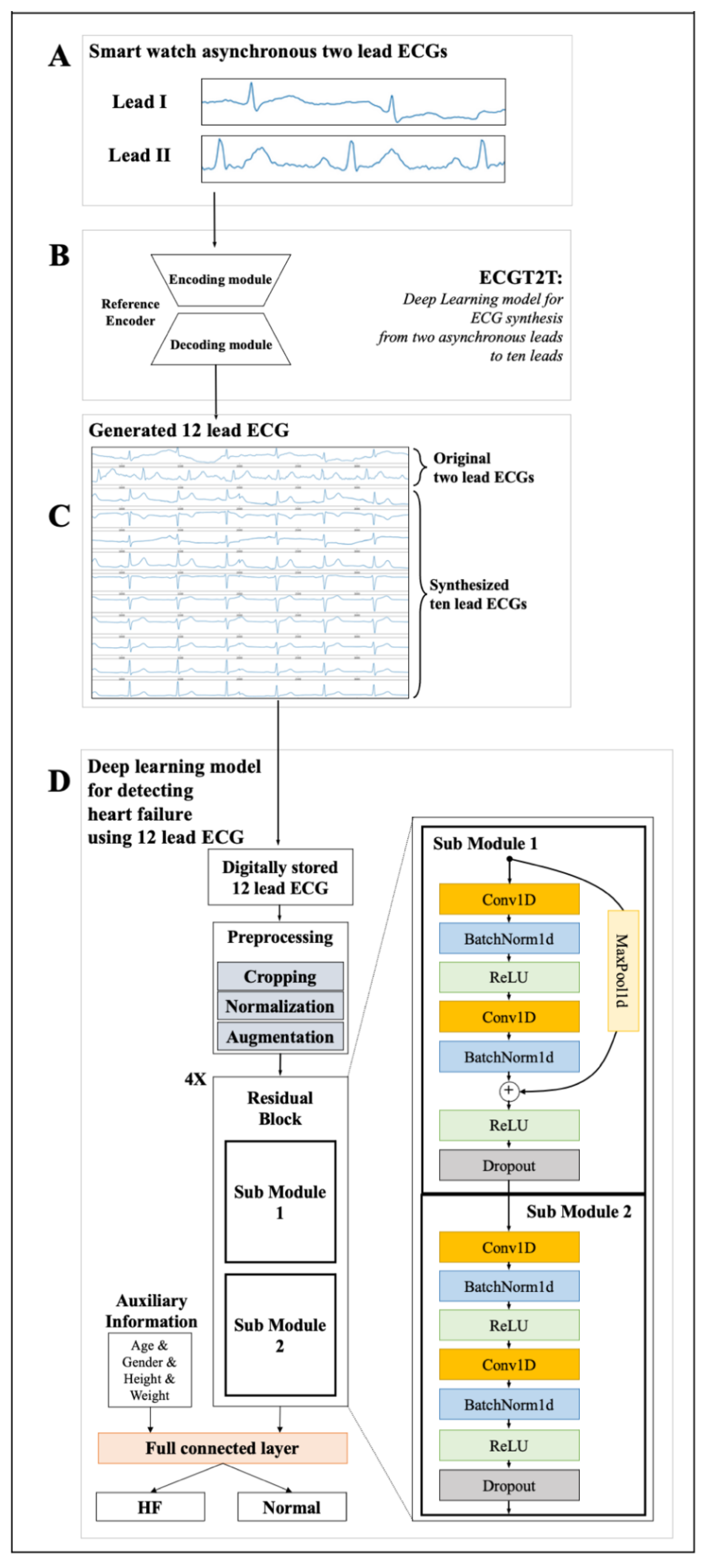
2.4. Development for a Platform Detecting HFrEF
2.5. Statistical Analysis
2.6. Role of the Funding Sources
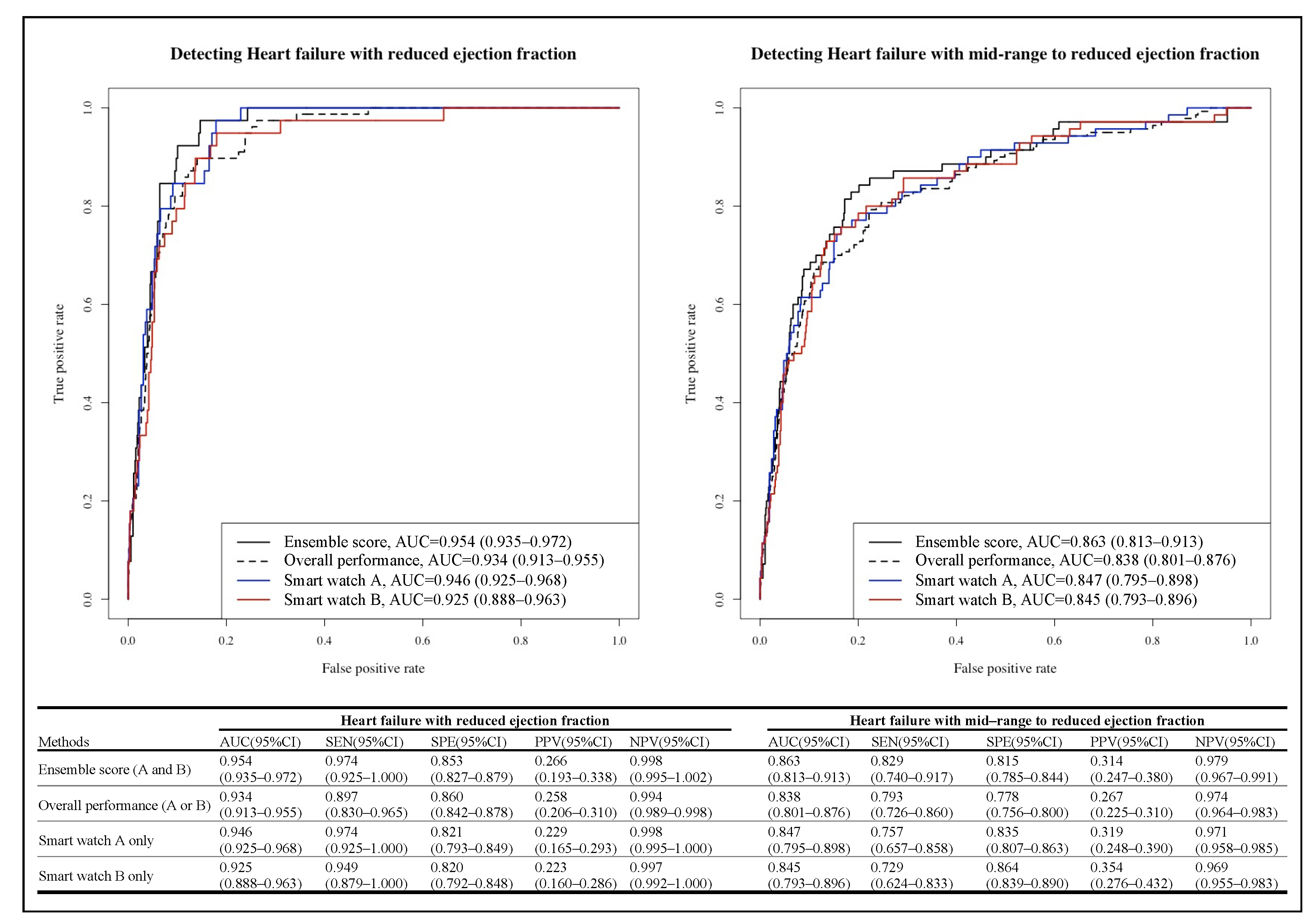
3. Results
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- GBD 2017 Disease and Injury Incidence and Prevalence Collaborators. Global, Regional, and National Incidence, Prevalence, and Years Lived with Disability for 354 Diseases and Injuries for 195 Countries and Territories, 1990–2017: A Systematic Analysis for the Global Burden of Disease Study. Lancet 2018, 392, 1789–1858. Available online: http://www.ncbi.nlm.nih.gov/pubmed/30496104 (accessed on 8 October 2021). [CrossRef] [Green Version]
- Lam, C.S.P.; Gamble, G.D.; Ling, L.H.; Sim, D.; Leong, K.T.G.; Yeo, P.S.D.; Ong, H.Y.; Jaufeerally, F.; Ng, T.P.; Cameron, V.A.; et al. Mortality associated with heart failure with preserved vs. reduced ejection fraction in a prospective international multi-ethnic cohort study. Eur. Heart J. 2018, 39, 1770–1780. Available online: http://www.ncbi.nlm.nih.gov/pubmed/29390051 (accessed on 8 October 2021). [CrossRef] [PubMed] [Green Version]
- Virani, S.S.; Alonso, A.; Benjamin, E.J.; Bittencourt, M.S.; Callaway, C.W.; Carson, A.P.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Delling, F.N.; et al. American Heart Association Council on Epidemiology and Prevention Statistics Committee and Stroke Statistics Subcommittee. Heart Disease and Stroke Statistics-2020 Update: A Report from the American Heart Association. Circulation 2020, 141, e139–e596. Available online: http://www.ncbi.nlm.nih.gov/pubmed/31992061 (accessed on 8 October 2021). [CrossRef] [PubMed]
- Braunwald, E. The war against heart failure: The Lancet lecture. Lancet 2015, 385, 812–824. [Google Scholar] [CrossRef]
- Groenewegen, A.; Rutten, F.H.; Mosterd, A.; Hoes, A.W. Epidemiology of heart failure. Eur. J. Heart Fail. 2020, 22, 1342–1356. Available online: http://www.ncbi.nlm.nih.gov/pubmed/32483830 (accessed on 8 October 2021). [CrossRef] [PubMed]
- Savarese, G.; Lund, L.H. Global Public Health Burden of Heart Failure. Card. Fail. Rev. 2017, 3, 7–11. [Google Scholar] [CrossRef]
- Flint, K.M.; Fairclough, D.L.; Spertus, J.A.; Bekelman, D.B. Does heart failure-specific health status identify patients with bothersome symptoms, depression, anxiety, and/or poorer spiritual well-being? Eur. Hear. J. Qual. Care Clin. Outcomes 2019, 5, 233–241. [Google Scholar] [CrossRef]
- Murphy, S.P.; Ibrahim, N.E.; Januzzi, J.L. Heart Failure with Reduced Ejection Fraction. JAMA 2020, 324, 488–504. Available online: https://jamanetwork.com/journals/jama/fullarticle/2768982 (accessed on 8 October 2021). [CrossRef]
- McDonagh, T.A.; Metra, M.; Adamo, M.; Gardner, R.S.; Baumbach, A.; Böhm, M.; Burri, H.; Butler, J.; Čelutkienė, J.; Chioncel, O.; et al. 2021 ESC Guidelines for the diagnosis and treatment of acute and chronic heart failure. Eur. Heart J. 2021, 42, 3599–3726. Available online: https://academic.oup.com/eurheartj/advance-article/doi/10.1093/eurheartj/ehab368/6358045 (accessed on 8 October 2021). [CrossRef]
- Jonas, D.E.; Reddy, S.; Middleton, J.C.; Barclay, C.; Green, J.; Baker, C.; Asher, G.N. Screening for Cardiovascular Disease Risk with Resting or Exercise Electrocardiography. JAMA 2018, 319, 2315–2328. [Google Scholar] [CrossRef] [Green Version]
- Kwon, J.-M.; Kim, K.-H.; Jeon, K.-H.; Kim, H.M.; Kim, M.J.; Lim, S.-M.; Song, P.S.; Park, J.; Choi, R.K.; Oh, B.-H. Development and Validation of Deep-Learning Algorithm for Electrocardiography-Based Heart Failure Identification. Korean Circ. J. 2019, 49, 629–639. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.; Lee, B.; Kwon, J.-M.; Lee, Y.; Park, H.; Oh, B.-H.; Jeon, K.-H.; Park, J.; Kim, K.-H. Artificial Intelligence Algorithm for Screening Heart Failure with Reduced Ejection Fraction Using Electrocardiography. ASAIO J. 2020, 67, 314–321. [Google Scholar] [CrossRef] [PubMed]
- Spaccarotella, C.A.M.; Polimeni, A.; Migliarino, S.; Principe, E.; Curcio, A.; Mongiardo, A.; Sorrentino, S.; De Rosa, S.; Indolfi, C. Multichannel Electrocardiograms Obtained by a Smartwatch for the Diagnosis of ST-Segment Changes. JAMA Cardiol. 2020, 5, 1176–1180. [Google Scholar] [CrossRef] [PubMed]
- Bozkurt, B.; Coats, A.J.; Tsutsui, H.; Abdelhamid, M.; Adamopoulos, S.; Albert, N.; Anker, S.D.; Atherton, J.; Böhm, M.; Butler, J.; et al. Universal Definition and Classification of Heart Failure. J. Card. Fail. 2021, 27, 387–413. Available online: https://linkinghub.elsevier.com/retrieve/pii/S1071916421000506 (accessed on 8 October 2021). [CrossRef] [PubMed]
- Jo, Y.-Y.; Kwon, J.-M. Electrocardiogram synthesis from two asynchronoous leads to Ten leads. arXiv 2021, arXiv:2103.00006. Available online: http://arxiv.org/abs/2103.00006 (accessed on 8 October 2021).
- Schisterman, E.F.; Perkins, N.J.; Liu, A.; Bondell, H. Optimal Cut-point and Its Corresponding Youden Index to Discriminate Individuals Using Pooled Blood Samples. Epidemiology 2005, 16, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Lokuge, A.; Lam, L.; Cameron, P.; Krum, H.; Smit, D.V.; Bystrzycki, A.; Naughton, M.T.; Eccleston, D.; Flannery, G.; Federman, J.; et al. B-Type Natriuretic Peptide Testing and the Accuracy of Heart Failure Diagnosis in the Emergency Department. Circ. Hear. Fail. 2010, 3, 104–110. [Google Scholar] [CrossRef] [Green Version]
- Jackson, S.L.; Tong, X.; King, R.J.; Loustalot, F.; Hong, Y.; Ritchey, M.D. National Burden of Heart Failure Events in the United States, 2006 to Circ. Hear. Fail. 2018, 11, e004873. [Google Scholar] [CrossRef]
- Cowie, M.R.; Anker, S.D.; Cleland, J.G.F.; Felker, G.M.; Filippatos, G.; Jaarsma, T.; Jourdain, P.; Knight, E.; Massie, B.; Ponikowski, P.; et al. Improving care for patients with acute heart failure: Before, during and after hospitalization. ESC Hear. Fail. 2014, 1, 110–145. [Google Scholar] [CrossRef]
- Perez, M.V.; Mahaffey, K.W.; Hedlin, H.; Rumsfeld, J.S.; Garcia, A.; Ferris, T.; Balasubramanian, V.; Russo, A.M.; Rajmane, A.; Cheung, L.; et al. Large-Scale Assessment of a Smartwatch to Identify Atrial Fibrillation. N. Engl. J. Med. 2019, 381, 1909–1917. [Google Scholar] [CrossRef]
- Mishra, T.; Wang, M.; Metwally, A.A.; Bogu, G.K.; Brooks, A.W.; Bahmani, A.; Alavi, A.; Celli, A.; Higgs, E.; Dagan-Rosenfeld, O.; et al. Pre-symptomatic detection of COVID-19 from smartwatch data. Nat. Biomed. Eng. 2020, 4, 1208–1220. [Google Scholar] [CrossRef] [PubMed]
- Breiman, L. Statistical Modeling: The Two Cultures. Stat. Sci. 2001, 16, 199–215. [Google Scholar] [CrossRef]
- Kwon, J.-M.; Jo, Y.-Y.; Lee, S.Y.; Kim, K.-H. Artificial intelligence using electrocardiography: Strengths and pitfalls. Eur. Hear. J. 2021, 42, 2896–2898. [Google Scholar] [CrossRef] [PubMed]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. Available online: http://arxiv.org/abs/1603.05691 (accessed on 8 October 2021). [CrossRef]
- Tison, G.; Sanchez, J.M.; Ballinger, B.; Singh, A.; Olgin, J.E.; Pletcher, M.J.; Vittinghoff, E.; Lee, E.S.; Fan, S.M.; Gladstone, R.A.; et al. Passive Detection of Atrial Fibrillation Using a Commercially Available Smartwatch. JAMA Cardiol. 2018, 3, 409–416. [Google Scholar] [CrossRef] [Green Version]
| Hospital A (38,643 Patients) Development and Interval Validation Data | Hospital B (755 Patients) External Validation Smart Watch Data | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Characteristics | HFrEF | HFmrEF | Normal | p † | HFrEF | HFmrEF | Normal | p † | p ‡ |
| Total Patients | 2519 (6.5) | 1755 (4.5) | 34,369 (88.9) | 39 (5.2) | 31 (4.1) | 685 (90.7) | 0.241 | ||
| Age (year) | 64.82 (13.52) | 64.98 (13.47) | 58.60 (15.45) | <0.001 | 60.69 (13.84) | 58.97 (14.46) | 55.61 (15.16) | 0.067 | <0.001 |
| Male | 1665 (65.7) | 1083 (61.3) | 16,341 (47.6) | <0.001 | 29 (74.4) | 20 (64.5) | 325 (47.4) | 0.001 | 0.969 |
| Weight (Kg) | 64.68 (13.90) | 65.25 (13.34) | 64.87 (12.60) | 0.341 | 69.29 (14.02) | 66.80 (12.68) | 66.03 (14.07) | 0.358 | 0.004 |
| Height (cm) | 162.68 (9.56) | 162.37 (10.01) | 162.20 (9.58) | 0.044 | 168.41 (10.55) | 164.48 (9.36) | 163.38 (9.39) | 0.005 | <0.001 |
| Body surface area (m2) | 1.70 (0.22) | 1.71 (0.21) | 1.70 (0.20) | 0.505 | 1.79 (0.22) | 1.74 (0.20) | 1.72 (0.22) | 0.145 | 0.001 |
| Heart rate (bpm) | 84.37 (24.54) | 78.26 (20.53) | 73.14 (15.82) | <0.001 | 78.31 (19.78) | 70.03 (14.55) | 69.95 (12.56) | 0.001 | <0.001 |
| PR interval (ms) | 175.83 (36.69) | 176.83 (37.36) | 167.99 (29.01) | <0.001 | 122.00 (60.53) | 156.74 (144.40) | 149.46 (97.07) | 0.225 | <0.001 |
| QRS duration (ms) | 111.81 (27.97) | 104.85 (23.55) | 95.47 (15.88) | <0.001 | 155.74 (63.39) | 139.42 (31.17) | 138.95 (31.03) | 0.010 | <0.001 |
| QT interval (ms) | 407.78 (57.74) | 408.08 (51.59) | 398.94 (40.13) | <0.001 | 421.90 (99.88) | 417.63 (45.29) | 425.56 (50.21) | 0.681 | <0.001 |
| Atrial fibrillation of flutter | 296 (11.7) | 170 (9.6) | 1172 (3.4) | <0.001 | 3 (7.7) | 1 (3.2) | 14 (2.0) | 0.076 | 0.015 |
| P wave axis | 45.58 (39.72) | 44.48 (35.84) | 45.32 (28.89) | 0.585 | NA | NA | NA | NA | |
| R wave axis | 27.71 (65.00) | 31.15 (53.61) | 39.80 (42.07) | <0.001 | NA | NA | NA | NA | |
| T wave axis | 83.07 (85.26) | 58.82 (72.34) | 42.57 (44.37) | <0.001 | NA | NA | NA | NA | |
| Ejection fraction (%) | 32.03 (9.44) | 46.08 (5.98) | 60.64 (6.33) | <0.001 | 31.23 (7.21) | 45.97 (2.50) | 64.63 (5.19) | <0.001 | <0.001 |
| Left atrial dimension (mm) | 45.66 (8.97) | 44.05 (9.48) | 38.98 (7.84) | <0.001 | 43.76 (7.48) | 42.48 (9.31) | 36.48 (6.90) | <0.001 | <0.001 |
| E | 67.69 (27.32) | 63.05 (25.71) | 63.50 (19.49) | <0.001 | 72.00 (22.11) | 68.65 (27.26) | 66.55 (19.21) | 0.37 | <0.001 |
| A | 68.40 (23.50) | 71.03 (21.03) | 70.06 (20.23) | 0.002 | 71.28 (22.05) | 74.00 (25.71) | 66.74 (20.64) | 0.251 | <0.001 |
| E′ | 5.04 (1.91) | 5.72 (2.09) | 7.10 (2.67) | <0.001 | 5.80 (3.81) | 6.06 (2.54) | 7.64 (4.62) | 0.044 | <0.001 |
| E/E′ | 14.90 (7.84) | 12.04 (6.27) | 9.88 (4.58) | <0.001 | 15.37 (6.90) | 13.13 (7.88) | 9.85 (4.22) | <0.001 | 0.534 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kwon, J.-m.; Jo, Y.-Y.; Lee, S.Y.; Kang, S.; Lim, S.-Y.; Lee, M.S.; Kim, K.-H. Artificial Intelligence-Enhanced Smartwatch ECG for Heart Failure-Reduced Ejection Fraction Detection by Generating 12-Lead ECG. Diagnostics 2022, 12, 654. https://doi.org/10.3390/diagnostics12030654
Kwon J-m, Jo Y-Y, Lee SY, Kang S, Lim S-Y, Lee MS, Kim K-H. Artificial Intelligence-Enhanced Smartwatch ECG for Heart Failure-Reduced Ejection Fraction Detection by Generating 12-Lead ECG. Diagnostics. 2022; 12(3):654. https://doi.org/10.3390/diagnostics12030654
Chicago/Turabian StyleKwon, Joon-myoung, Yong-Yeon Jo, Soo Youn Lee, Seonmi Kang, Seon-Yu Lim, Min Sung Lee, and Kyung-Hee Kim. 2022. "Artificial Intelligence-Enhanced Smartwatch ECG for Heart Failure-Reduced Ejection Fraction Detection by Generating 12-Lead ECG" Diagnostics 12, no. 3: 654. https://doi.org/10.3390/diagnostics12030654
APA StyleKwon, J.-m., Jo, Y.-Y., Lee, S. Y., Kang, S., Lim, S.-Y., Lee, M. S., & Kim, K.-H. (2022). Artificial Intelligence-Enhanced Smartwatch ECG for Heart Failure-Reduced Ejection Fraction Detection by Generating 12-Lead ECG. Diagnostics, 12(3), 654. https://doi.org/10.3390/diagnostics12030654