Automatic Recognition of Colon and Esophagogastric Cancer with Machine Learning and Hyperspectral Imaging
Abstract
:1. Introduction
1.1. Clinical Context
1.2. Hyperspectral Imaging and Medical Applications
1.3. Contribution Summary
2. Material and Methods
2.1. Data Collection and Annotation
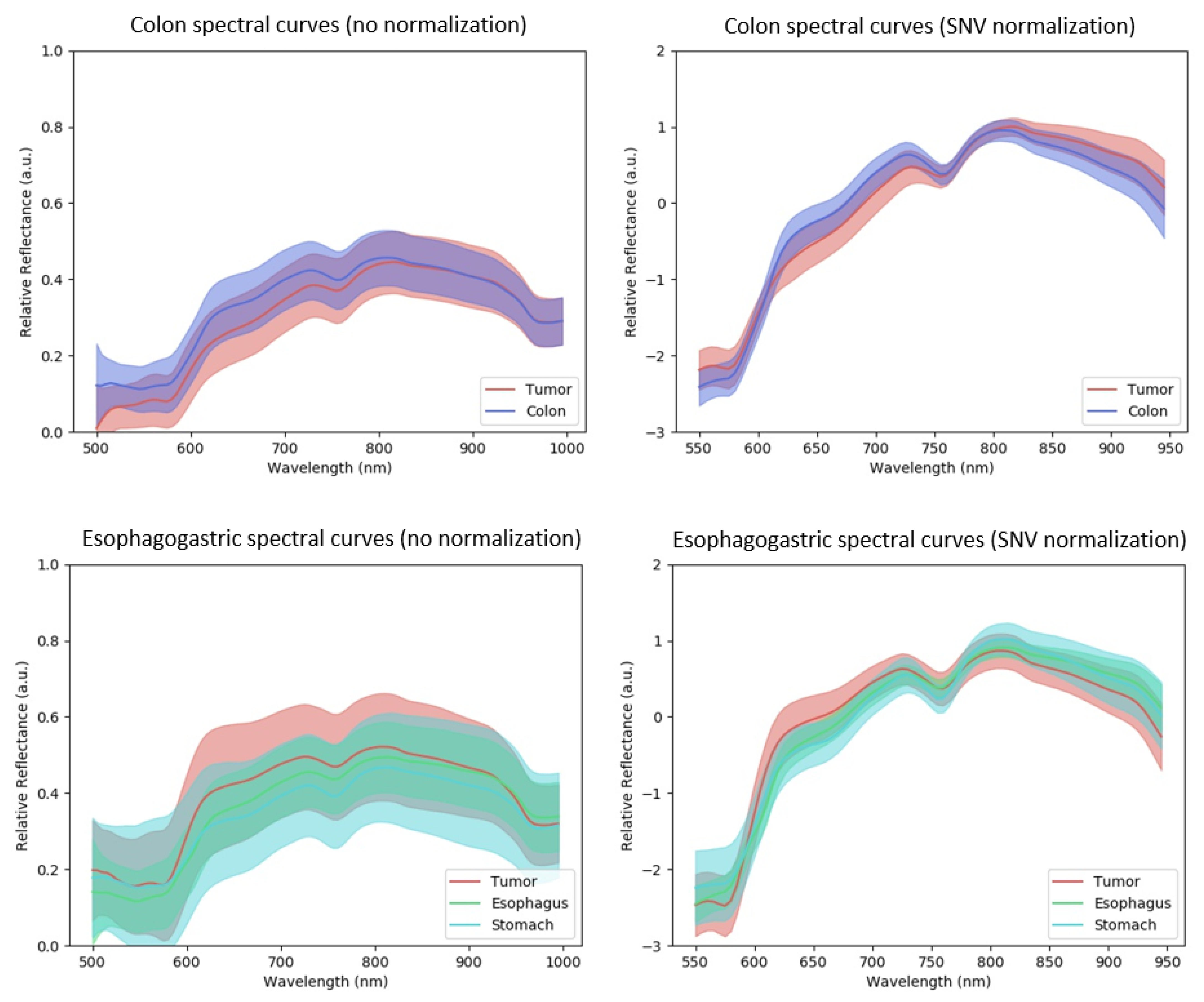
2.2. Dataset Analysis with Spectral Curves
2.3. Models, Training Processes and Performance Metrics
2.3.1. Classical Machine Learning Models
2.3.2. Convolutional Neural Network Models
2.3.3. Leave-One-Patient-Out Cross Validation (LOPOCV)
2.3.4. Evaluation with the Colon, Esophagogastric and Combined Datasets
2.3.5. Training Implementation Details
3. Results and Discussion
3.1. Evaluation Metrics
3.2. Statistical Tests
3.3. ROC-AUC Results
3.4. MCC and DICE Results
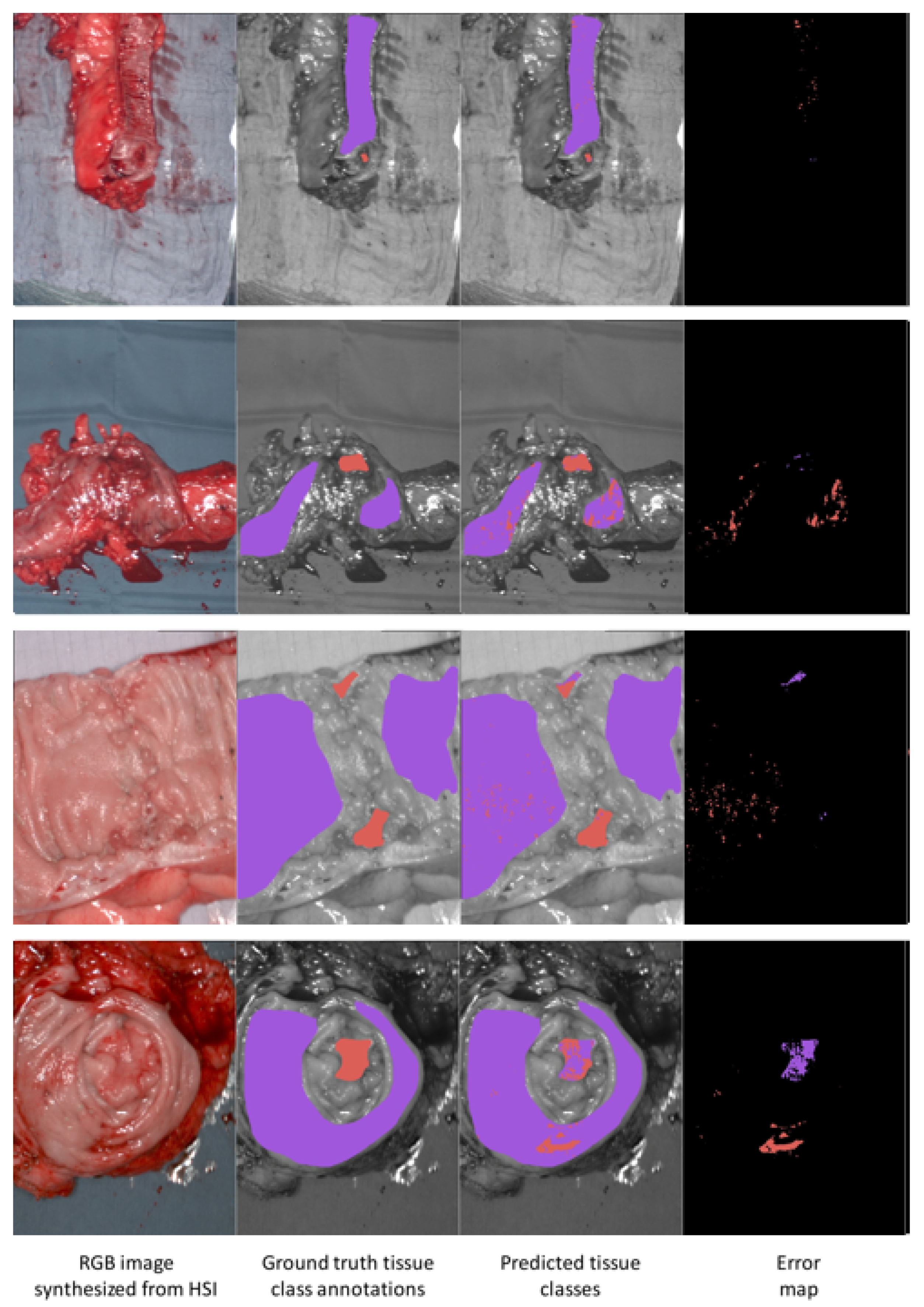
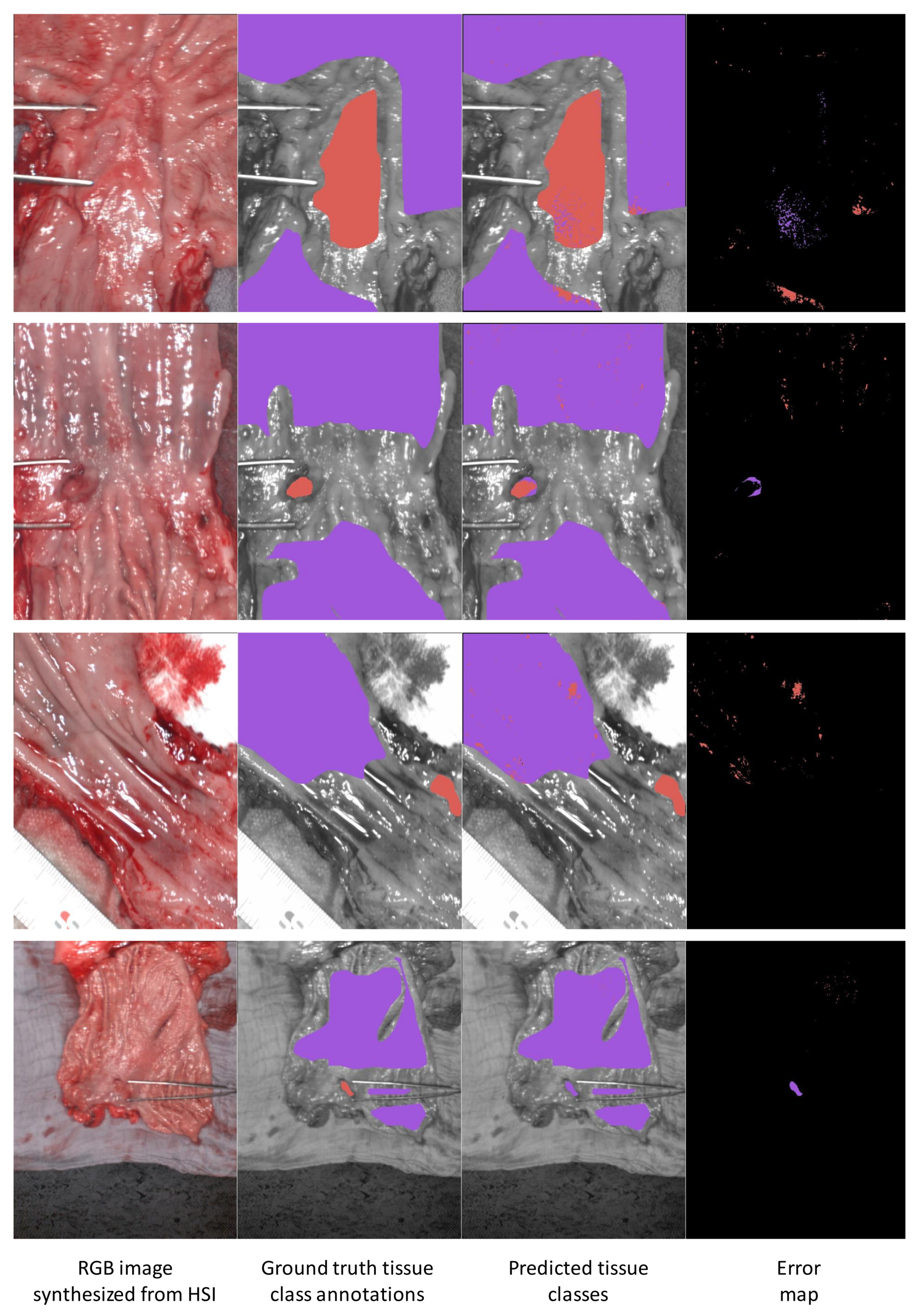
3.5. Results Visualization
3.6. Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ML | Machine Learning |
| SVM | Support Vector Machine |
| RBF-SVM | Radial basis function Support Vector Machine |
| MLP | Multi-Layer Perceptron |
| CNN | Convolutional Neural Network |
| 3DCNN | Three Dimensional Convolutional Neural Network |
| SNV | Standard Normal Variate |
| HSI | Hyperspectral imaging |
| LOPOCV | Leave-one-patient-out cross-validation |
| EG | esophagogastric |
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [Green Version]
- Tai, F.W.D.; Wray, N.; Sidhu, R.; Hopper, A.; McAlindon, M. Factors associated with oesophagogastric cancers missed by gastroscopy: A case–control study. Frontline Gastroenterol. 2020, 11, 194–201. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, P.; Pante, A.; Gross, S.A. Artificial intelligence in endoscopy. Gastrointest. Endosc. 2020, 91, 925–931. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Fei, B. Medical hyperspectral imaging: A review. J. Biomed. Opt. 2014, 19, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.J.; Khan, H.S.; Yousaf, A.; Khurshid, K.; Abbas, A. Modern Trends in Hyperspectral Image Analysis: A Review. IEEE Access 2018, 6, 14118–14129. [Google Scholar] [CrossRef]
- Li, Q.; He, X.; Wang, Y.; Liu, H.; Xu, D.; Guo, F. Review of spectral imaging technology in biomedical engineering: Achievements and challenges. J. Biomed. Opt. 2013, 18, 100901. [Google Scholar] [CrossRef] [Green Version]
- Ortega, S.; Fabelo, H.; Iakovidis, D.K.; Koulaouzidis, A.; Callico, G.M. Use of Hyperspectral/Multispectral Imaging in Gastroenterology. Shedding Some—Different—Light into the Dark. J. Clin. Med. 2019, 8, 36. [Google Scholar] [CrossRef] [Green Version]
- Signoroni, A.; Savardi, M.; Baronio, A.; Benini, S. Deep Learning Meets Hyperspectral Image Analysis: A Multidisciplinary Review. J. Imaging 2019, 5, 52. [Google Scholar] [CrossRef] [Green Version]
- Baltussen, E.J.M.; Kok, E.N.D.; Brouwer de Koning, S.G.; Sanders, J.; Aalbers, A.G.J.; Kok, N.F.M.; Beets, G.L.; Flohil, C.C.; Bruin, S.C.; Kuhlmann, K.F.D.; et al. Hyperspectral imaging for tissue classification, a way toward smart laparoscopic colorectal surgery. J. Biomed. Opt. 2019, 24, 016002. [Google Scholar] [CrossRef]
- Hohmann, M.; Kanawade, R.; Klämpfl, F.; Douplik, A.; Mudter, J.; Neurath, M.F.; Albrecht, H. In-Vivo multispectral video endoscopy towards in-vivo hyperspectral video endoscopy. J. Biophotonics 2017, 10, 553–564. [Google Scholar] [CrossRef]
- Manni, F.; Fonollá, R.; van der Sommen, F.; Zinger, S.; Shan, C.; Kho, E.; de Koning, S.G.B.; Ruers, T.; de With, P.H.N. Hyperspectral imaging for colon cancer classification in surgical specimens: Towards optical biopsy during image-guided surgery. In Proceedings of the 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society, Montreal, QC, Canada, 20–24 July 2020; pp. 1169–1173. [Google Scholar]
- Liu, N.; Guo, Y.; Jiang, H.; Yi, W. Gastric cancer diagnosis using hyperspectral imaging with principal component analysis and spectral angle mapper. J. Biomed. Opt. 2020, 25, 1. [Google Scholar] [CrossRef]
- Maktabi, M.; Köhler, H.; Ivanova, M.; Jansen-Winkeln, B.; Takoh, J.; Niebisch, S.; Rabe, S.M.; Neumuth, T.; Gockel, I.; Chalopin, C. Tissue classification of oncologic esophageal resectates based on hyperspectral data. Int. J. Comput. Assist. Radiol. Surg. 2019, 14, 1651–1661. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wang, H.; Li, Q. Tongue Tumor Detection in Medical Hyperspectral Images. Sensors 2011, 12, 162–174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duann, J.R.; Jan, C.I.; Ou-Yang, M.; Lin, C.Y.; Mo, J.F.; Lin, Y.J.; Tsai, M.H.; Chiou, J.C. Separating spectral mixtures in hyperspectral image data using independent component analysis: Validation with oral cancer tissue sections. J. Biomed. Opt. 2013, 18, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hosking, A.M.; Coakley, B.J.; Chang, D.; Talebi-Liasi, F.; Lish, S.; Lee, S.W.; Zong, A.M.; Moore, I.; Browning, J.; Jacques, S.L.; et al. Hyperspectral imaging in automated digital dermoscopy screening for melanoma. Lasers Surg. Med. 2019, 51, 214–222. [Google Scholar] [CrossRef]
- Kho, E.; de Boer, L.L.; Van de Vijver, K.K.; van Duijnhoven, F.; Vrancken Peeters, M.J.T.; Sterenborg, H.J.; Ruers, T.J. Hyperspectral Imaging for Resection Margin Assessment during Cancer Surgery. Clin. Cancer Res. 2019, 25, 3572–3580. [Google Scholar] [CrossRef] [Green Version]
- Fabelo, H.; Ortega, S.; Ravi, D.; Kiran, B.R.; Sosa, C.; Bulters, D.; Callicó, G.M.; Bulstrode, H.; Szolna, A.; Piñeiro, J.F.; et al. Spatio-spectral classification of hyperspectral images for brain cancer detection during surgical operations. PLoS ONE 2018, 13, e0193721. [Google Scholar] [CrossRef] [Green Version]
- Han, Z.; Zhang, A.; Wang, X.; Sun, Z.; Wang, M.D.; Xie, T. In vivo use of hyperspectral imaging to develop a noncontact endoscopic diagnosis support system for malignant colorectal tumors. J. Biomed. Opt. 2016, 21, 016001. [Google Scholar] [CrossRef] [Green Version]
- Laffers, W.; Westermann, S.; Regeling, B.; Martin, R.; Thies, B.; Gerstner, A.O.H.; Bootz, F.; Müller, N. Early recognition of cancerous lesions in the mouth and oropharynx: Automated evaluation of hyperspectral image stacks. HNO 2016, 64, 27–33. [Google Scholar] [CrossRef]
- Halicek, M.; Dormer, J.D.; Little, J.V.; Chen, A.Y.; Myers, L.; Sumer, B.D.; Fei, B. Hyperspectral Imaging of Head and Neck Squamous Cell Carcinoma for Cancer Margin Detection in Surgical Specimens from 102 Patients Using Deep Learning. Cancers 2019, 11, 1367. [Google Scholar] [CrossRef] [Green Version]
- Fabelo, H.; Ortega, S.; Kabwama, S.; Callico, G.M.; Bulters, D.; Szolna, A.; Pineiro, J.F.; Sarmiento, R. HELICoiD project: A new use of hyperspectral imaging for brain cancer detection in real-time during neurosurgical operations. In Hyperspectral Imaging Sensors: Innovative Applications and Sensor Standards 2016; Bannon, D.P., Ed.; SPIE: Bellingham, DC, USA, 2016. [Google Scholar]
- Halicek, M.; Lu, G.; Little, J.V.; Wang, X.; Patel, M.; Griffith, C.C.; El-Deiry, M.W.; Chen, A.Y.; Fei, B. Deep convolutional neural networks for classifying head and neck cancer using hyperspectral imaging. J. Biomed. Opt. 2017, 22, 060503. [Google Scholar] [CrossRef] [PubMed]
- Halicek, M.; Fabelo, H.; Ortega, S.; Callico, G.M.; Fei, B. In-Vivo and Ex-Vivo Tissue Analysis through Hyperspectral Imaging Techniques: Revealing the Invisible Features of Cancer. Cancers 2019, 11, 756. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clancy, N.T.; Jones, G.; Maier-Hein, L.; Elson, D.S.; Stoyanov, D. Surgical spectral imaging. Med. Image Anal. 2020, 63, 101699. [Google Scholar] [CrossRef] [PubMed]
- Köhler, H.; Kulcke, A.; Maktabi, M.; Moulla, Y.; Jansen-Winkeln, B.; Barberio, M.; Diana, M.; Gockel, I.; Neumuth, T.; Chalopin, C. Laparoscopic system for simultaneous high-resolution video and rapid hyperspectral imaging in the visible and near-infrared spectral range. J. Biomed. Opt. 2020, 25, 086004. [Google Scholar] [CrossRef] [PubMed]
- Kumashiro, R.; Konishi, K.; Chiba, T.; Akahoshi, T.; Nakamura, S.; Murata, M.; Tomikawa, M.; Matsumoto, T.; Maehara, Y.; Hashizume, M. Integrated Endoscopic System Based on Optical Imaging and Hyperspectral Data Analysis for Colorectal Cancer Detection. Anti-Cancer Res. 2016, 36, 3925–3932. [Google Scholar]
- Regeling, B.; Laffers, W.; Gerstner, A.O.H.; Westermann, S.; Müller, N.A.; Schmidt, K.; Bendix, J.; Thies, B. Development of an image pre-processor for operational hyperspectral laryngeal cancer detection. J. Biophotonics 2016, 9, 235–245. [Google Scholar] [CrossRef]
- Yoon, J.; Joseph, J.; Waterhouse, D.J.; Luthman, A.S.; Gordon, G.S.D.; di Pietro, M.; Januszewicz, W.; Fitzgerald, R.C.; Bohndiek, S.E. A clinically translatable hyperspectral endoscopy (HySE) system for imaging the gastrointestinal tract. Nat. Commun. 2019, 10, 1902. [Google Scholar] [CrossRef] [Green Version]
- Beaulieu, R.J.; Goldstein, S.D.; Singh, J.; Safar, B.; Banerjee, A.; Ahuja, N. Automated diagnosis of colon cancer using hyperspectral sensing. Int. J. Med Robot. Comput. Assist. Surg. 2018, 14, e1897. [Google Scholar] [CrossRef]
- Jansen-Winkeln, B.; Barberio, M.; Chalopin, C.; Schierle, K.; Diana, M.; Köhler, H.; Gockel, I.; Maktabi, M. Feedforward Artificial Neural Network-Based Colorectal Cancer Detection Using Hyperspectral Imaging: A Step towards Automatic Optical Biopsy. Cancers 2021, 13, 967. [Google Scholar] [CrossRef]
- Barnes, R.J.; Dhanoa, M.S.; Lister, S.J. Standard Normal Variate Transformation and De-Trending of Near-Infrared Diffuse Reflectance Spectra. Appl. Spectrosc. 1989, 43, 772–777. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Chen, Y.; Jiang, H.; Li, C.; Jia, X.; Ghamisi, P. Deep Feature Extraction and Classification of Hyperspectral Images Based on Convolutional Neural Networks. IEEE Trans. Geosci. Remote Sens. 2016, 54, 6232–6251. [Google Scholar] [CrossRef] [Green Version]
- Audebert, N.; Le Saux, B.; Lefevre, S. Deep Learning for Classification of Hyperspectral Data: A Comparative Review. IEEE Geosci. Remote Sens. Mag. 2019, 7, 159–173. [Google Scholar] [CrossRef] [Green Version]
- Hu, W.; Huang, Y.; Wei, L.; Zhang, F.; Li, H. Deep Convolutional Neural Networks for Hyperspectral Image Classification. J. Sensors 2015, 2015, 258619:1–258619:12. [Google Scholar] [CrossRef] [Green Version]
- He, M.; Li, B.; Chen, H. Multi-scale 3D deep convolutional neural network for hyperspectral image classification. In Proceedings of the 2017 IEEE International Conference on Image Processing (ICIP), Beijing, China, 17–20 September 2017; pp. 3904–3908. [Google Scholar]
- Luo, Y.; Zou, J.; Yao, C.; Zhao, X.; Li, T.; Bai, G. HSI-CNN: A Novel Convolution Neural Network for Hyperspectral Image. In Proceedings of the 2018 International Conference on Audio, Language and Image Processing (ICALIP), Shanghai, China, 16–17 July 2018; pp. 464–469. [Google Scholar]
- Li, Y.; Zhang, H.; Shen, Q. Spectral—Spatial Classification of Hyperspectral Imagery with 3D Convolutional Neural Network. Remote Sens. 2017, 9, 67. [Google Scholar] [CrossRef] [Green Version]
- Hamida, A.; Benoit, A.; Lambert, P.; Ben Amar, C. 3-D Deep Learning Approach for Remote Sensing Image Classification. IEEE Trans. Geosci. Remote Sens. 2018, 56, 4420–4434. [Google Scholar] [CrossRef] [Green Version]
- Iandola, F.N.; Moskewicz, M.W.; Ashraf, K.; Han, S.; Dally, W.J.; Keutzer, K. SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and <1MB model size. arXiv 2016, arXiv:1602.07360. [Google Scholar]
- Ronneberger, O.; Fischer., P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention (MICCAI), Munich, Germany, 5–9 October 2015; Volume 9351, pp. 234–241. [Google Scholar]
- Li, Y.; Xie, X.; Yang, X.; Guo, L.; Liu, Z.; Zhao, X.; Luo, Y.; Jia, W.; Huang, F.; Zhu, S.; et al. Diagnosis of early gastric cancer based on fluorescence hyperspectral imaging technology combined with partial-least-square discriminant analysis and support vector machine. J. Biophotonics 2019, 12, e201800324. [Google Scholar] [CrossRef]
- Mandrekar, J. Receiver Operating Characteristic Curve in Diagnostic Test Assessment. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2010, 5, 1315–1316. [Google Scholar] [CrossRef] [Green Version]
- Kamruzzaman, M.; Sun, D.W. Chapter 5—Introduction to Hyperspectral Imaging Technology. In Computer Vision Technology for Food Quality Evaluation, 2nd ed.; Academic Press: San Diego, CA, USA, 2016; pp. 111–139. [Google Scholar]
- Chen, P.C.; Lin, W.C. Spectral-profile-based algorithm for hemoglobin oxygen saturation determination from diffuse reflectance spectra. Biomed. Opt. Express 2011, 2, 1082–1096. [Google Scholar] [CrossRef]
- Welch, A.J.; van Gemert, M.J.C.; Star, W.M.; Wilson, B.C. Definitions and Overview of Tissue Optics. In Optical-Thermal Response of Laser-Irradiated Tissue; Welch, A.J., Van Gemert, M.J.C., Eds.; Lasers, Photonics, and Electro-Optics; Springer: Boston, MA, USA, 1995; pp. 15–46. [Google Scholar]
- Sensing of Optical Properties and Spectroscopy. In Biomedical Optics; John Wiley & Sons, Ltd: New York, NY, USA, 2009; pp. 135–151.
- Sahani, D.V.; Kalva, S.P.; Hamberg, L.M.; Hahn, P.F.; Willett, C.G.; Saini, S.; Mueller, P.R.; Lee, T.Y. Assessing Tumor Perfusion and Treatment Response in Rectal Cancer with Multisection CT: Initial Observations. Radiology 2005, 234, 785–792. [Google Scholar] [CrossRef] [PubMed]
- Stolik, S.; Delgado, J.A.; Pérez, A.; Anasagasti, L. Measurement of the penetration depths of red and near infrared light in human “ex vivo” tissues. J. Photochem. Photobiol. B Biol. 2000, 57, 90–93. [Google Scholar] [CrossRef]
- Jacques, S.L. Optical properties of biological tissues: A review. Phys. Med. Biol. 2013, 58, R37–R61. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.V.; Wu, H.I. Biomedical Optics: Principles and Imaging; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2009. [Google Scholar]
- Singh, A.; Sengupta, S.; Lakshminarayanan, V. Explainable Deep Learning Models in Medical Image Analysis. J. Imaging 2020, 6, 52. [Google Scholar] [CrossRef] [PubMed]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar]
- Caruana, R. Multitask Learning. In Learning to Learn; Thrun, S., Pratt, L., Eds.; Springer: Boston, MA, USA, 1998; pp. 95–133. [Google Scholar]
- Misra, I.; Shrivastava, A.; Gupta, A.; Hebert, M. Cross-Stitch Networks for Multi-Task Learning. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, Nevada, 26 June–1 July 2016. [Google Scholar]
- Kiyotoki, S.; Nishikawa, J.; Okamoto, T.; Hamabe, K.; Saito, M.; Goto, A.; Fujita, Y.; Hamamoto, Y.; Takeuchi, Y.; Satori, S.; et al. New method for detection of gastric cancer by hyperspectral imaging: A pilot study. J. Biomed. Opt. 2013, 18, 7. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Zhou, S.; Han, W.; Liu, W.; Qiu, Z.; Li, C. Convolutional neural network for hyperspectral data analysis and effective wavelengths selection. Anal. Chim. Acta 2019, 1086, 46–54. [Google Scholar] [CrossRef]
- Vohra, R.; Tiwari, K.C. Spatial shape feature descriptors in classification of engineered objects using high spatial resolution remote sensing data. Evol. Syst. 2020, 11, 647–660. [Google Scholar] [CrossRef]
- Al-Sarayreh, M.; Reis, M.M.; Yan, W.Q.; Klette, R. Potential of deep learning and snapshot hyperspectral imaging for classification of species in meat. Food Control 2020, 117, 107332. [Google Scholar] [CrossRef]

| Patient | Age | Gender | Proportion of Healthy Tissue Pixels (%) | Proportion of Cancer Tissue Pixels (%) | Tissue Location | T Classification | Tumor Grade |
|---|---|---|---|---|---|---|---|
| 1 | 67 | f | 93.57 | 6.43 | colon | pT3b | G2 |
| 2 | 68 | m | 100 | 0 | colon | adenoma | |
| 3 | 52 | m | 91.27 | 8.73 | colon | pT2 | G2 |
| 4 | 67 | f | 98.68 | 1.32 | colorectal | pT2 | G2 |
| 5 | 80 | m | 92.55 | 7.45 | colorectal | pT2 | G2 |
| 6 | 80 | m | 100 | 0 | colon | adenoma | |
| 7 | 81 | m | 56.16 | 43.84 | colorectal | ypT3 | G1 (mainly cancer) |
| 8 | 66 | m | 97.12 | 2.88 | colorectal | rpT2 | G2 |
| 9 | 66 | m | 94.62 | 5.38 | colorectal | ypT2 | G1 (mainly cancer) |
| 10 | 60 | m | 90.50 | 9.50 | sigma | ypT3 | G3 |
| 11 | 79 | f | 89.63 | 10.37 | ascending colon | pT3 | G2 |
| 12 | 59 | m | 96.59 | 3.41 | colon ascendens | pT2 | G2 |
| Mean | 91.72 | 8.28 |
| Patient | Age | Gender | Proportion of Healthy Stomach Tissue Pixels (%) | Proportion of Healthy Esophagus Tissue Pixels (%) | Proportion of Cancer Tissue Pixels (%) | Tumor~ Type | Tissue Location | T Classification |
|---|---|---|---|---|---|---|---|---|
| 13 | 83 | m | 39.65 | 36.16 | 24.19 | Not determined | G-E junction | ypT0 |
| 14 | 71 | m | 0 | 96.38 | 3.62 | AC | G-E junction | ypT3 |
| 15 | 72 | m | 31.77 | 61.88 | 6.35 | AC | G-E junction | ypT1b |
| 16 | 67 | m | 60.59 | 31.11 | 8.30 | AC | G-E junction | ypT1b |
| 17 | 73 | m | 32.71 | 48.31 | 18.98 | AC | G-E junction | ypT0 |
| 18 | 67 | m | 6.79 | 87.02 | 6.20 | AC | G-E junction | ypT3 |
| 19 | 54 | m | 36.66 | 62.15 | 1.19 | AC | G-E junction | ypT1b |
| 20 | 65 | f | 99.13 | 0 | 0.87 | AC | G-E junction | pT1b |
| 21 | 56 | m | 61.64 | 16.31 | 22.06 | AC | G-E junction | ypT2 |
| 22 | 60 | m | 30.34 | 67.09 | 2.57 | AC | G-E junction | ypT0 |
| Mean | 39.93 | 50.64 | 9.43 |
| Train Dataset: Colon, Test Dataset: Colon | Train Dataset: Combined, Test Dataset: Colon | |||||
|---|---|---|---|---|---|---|
| Patient ID | RBF-SVM | MLP | 3DCNN | RBF-SVM | MLP | 3DCNN |
| 1 | 0.97 | 0.98 | 1.0 | 0.98 | 0.98 | 0.99 |
| 2 | / | / | / | / | / | / |
| 3 | 0.93 | 0.96 | 0.96 | 0.93 | 0.97 | 0.85 |
| 4 | 0.98 | 1.0 | 0.99 | 0.98 | 1.0 | 0.87 |
| 5 | 0.86 | 0.87 | 0.95 | 0.87 | 0.93 | 0.94 |
| 6 | / | / | / | / | / | / |
| 7 | 0.69 | 0.66 | 0.89 | 0.78 | 0.87 | 0.72 |
| 8 | 0.56 | 0.77 | 0.93 | 0.67 | 0.75 | 0.96 |
| 9 | 0.95 | 0.90 | 0.99 | 0.94 | 0.74 | 0.99 |
| 10 | 0.91 | 0.92 | 0.77 | 0.78 | 0.68 | 0.89 |
| 11 | 0.98 | 0.98 | 0.93 | 0.98 | 0.99 | 0.98 |
| 12 | 0.94 | 0.88 | 0.88 | 0.89 | 0.90 | 1.0 |
| Mean ± S.D. | 0.88 ± 0.12 | 0.89 ± 0.11 | 0.93 ± 0.069 | 0.88 ± 0.11 | 0.88 ± 0.12 | 0.92 ± 0.088 |
| Train Dataset: EG, Test Dataset: EG | Train Dataset: Combined, Test Dataset: EG | |||||
|---|---|---|---|---|---|---|
| Patient ID | RBF-SVM | MLP | 3DCNN | RBF-SVM | MLP | 3DCNN |
| 13 | 0.96 | 0.96 | 0.91 | 0.98 | 0.99 | 0.99 |
| 14 | 0.81 | 0.85 | 0.99 | 0.99 | 0.98 | 0.98 |
| 15 | 0.91 | 0.92 | 0.92 | 0.87 | 0.92 | 0.95 |
| 16 | 0.91 | 0.87 | 0.98 | 0.91 | 0.96 | 0.96 |
| 17 | 0.59 | 0.67 | 0.89 | 0.73 | 0.48 | 0.89 |
| 18 | 0.53 | 0.47 | 0.90 | 0.80 | 0.87 | 0.85 |
| 19 | 0.81 | 0.93 | 0.93 | 0.95 | 0.99 | 0.97 |
| 20 | 0.68 | 0.55 | 0.71 | 0.85 | 0.77 | 0.79 |
| 21 | 0.91 | 0.94 | 0.92 | 0.99 | 1.0 | 0.96 |
| 22 | 0.80 | 0.79 | 0.99 | 0.95 | 0.98 | 0.98 |
| Mean ± S.D. | 0.79 ± 0.15 | 0.80 ± 0.17 | 0.91 ± 0.081 | 0.90 ± 0.082 | 0.89 ± 0.124 | 0.93 ± 0.067 |
| Train Dataset: EG, Test Dataset: Colon | Train Dataset: Colon, Test Dataset: EG | ||||
|---|---|---|---|---|---|
| Patient ID | MLP | 3DCNN | Patient ID | MLP | 3DCNN |
| 1 | 0.64 | 0.96 | 13 | 0.80 | 0.99 |
| 2 | / | / | 14 | 0.75 | 0.77 |
| 3 | 0.34 | 0.59 | 15 | 0.91 | 0.83 |
| 4 | 0.67 | 0.61 | 16 | 0.84 | 0.79 |
| 5 | 0.64 | 0.90 | 17 | 0.72 | 0.68 |
| 6 | / | / | 18 | 0.48 | 0.78 |
| 7 | 0.79 | 0.43 | 19 | 0.82 | 0.99 |
| 8 | 0.59 | 0.89 | 20 | 0.49 | 0.90 |
| 9 | 0.57 | 0.92 | 21 | 0.84 | 0.95 |
| 10 | 0.87 | 0.58 | 22 | 0.71 | 0.94 |
| 11 | 0.65 | 0.98 | - | - | - |
| 12 | 0.91 | 0.96 | - | - | - |
| Mean ± S.D. | 0.67 ± 0.16 | 0.78 ± 0.20 | 0.74 ± 0.15 | 0.86 ± 0.11 | |
| Mean MCC ± S.D. | Patient-Generic Decision Threshold | Patient-Specific Decition Threshold | ||||
|---|---|---|---|---|---|---|
| RBF-SVM | MLP | 3DCNN | RBF-SVM | MLP | 3DCNN | |
| Train dataset: colon, test dataset: colon | 0.37 ± 0.22 | 0.22 ± 0.26 | 0.49 ± 0.22 | 0.57 ± 0.31 | 0.53 ± 0.25 | 0.58 ± 0.23 |
| Train dataset: combined, test dataset: colon | 0.35 ± 0.23 | 0.29 ± 0.24 | 0.42 ± 0.16 | 0.57 ± 0.31 | 0.53± 0.28 | 0.55 ± 0.20 |
| Train dataset: EG, test dataset: EG | 0.27 ± 0.27 | 0.26 ± 0.26 | 0.41 ± 0.18 | 0.39 ± 0.30 | 0.34 ± 0.26 | 0.60 ± 0.25 |
| Train dataset: combined, test dataset: EG | 0.37 ± 0.23 | 0.33 ± 0.22 | 0.41 ± 0.22 | 0.63 ± 0.28 | 0.54 ± 0.29 | 0.51 ± 0.25 |
| Mean DICE ± S.D. | Patient-Generic Decision Threshold | Patient-Specific Decision Threshold | ||||
|---|---|---|---|---|---|---|
| RBF-SVM | MLP | 3DCNN | RBF-SVM | MLP | 3DCNN | |
| Train dataset: colon, test dataset: colon | 0.39 ± 0.24 | 0.36 ± 0.22 | 0.50 ± 0.24 | 0.52 ± 0.25 | 0.58 ± 0.24 | 0.61 ± 0.24 |
| Train dataset: combined, test dataset: colon | 0.38 ± 0.24 | 0.32 ± 0.25 | 0.44 ± 0.18 | 0.56 ± 0.25 | 0.57 ± 0.28 | 0.59 ± 0.20 |
| Train dataset: EG, test dataset: EG | 0.30 ± 0.29 | 0.29 ± 0.26 | 0.41 ± 0.20 | 0.49 ± 0.31 | 0.38 ± 0.26 | 0.62 ± 0.26 |
| Train dataset: combined, test dataset: EG | 0.38 ± 0.25 | 0.34 ± 0.24 | 0.40 ± 0.13 | 0.56 ± 0.30 | 0.60 ± 0.24 | 0.52 ± 0.26 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Collins, T.; Maktabi, M.; Barberio, M.; Bencteux, V.; Jansen-Winkeln, B.; Chalopin, C.; Marescaux, J.; Hostettler, A.; Diana, M.; Gockel, I. Automatic Recognition of Colon and Esophagogastric Cancer with Machine Learning and Hyperspectral Imaging. Diagnostics 2021, 11, 1810. https://doi.org/10.3390/diagnostics11101810
Collins T, Maktabi M, Barberio M, Bencteux V, Jansen-Winkeln B, Chalopin C, Marescaux J, Hostettler A, Diana M, Gockel I. Automatic Recognition of Colon and Esophagogastric Cancer with Machine Learning and Hyperspectral Imaging. Diagnostics. 2021; 11(10):1810. https://doi.org/10.3390/diagnostics11101810
Chicago/Turabian StyleCollins, Toby, Marianne Maktabi, Manuel Barberio, Valentin Bencteux, Boris Jansen-Winkeln, Claire Chalopin, Jacques Marescaux, Alexandre Hostettler, Michele Diana, and Ines Gockel. 2021. "Automatic Recognition of Colon and Esophagogastric Cancer with Machine Learning and Hyperspectral Imaging" Diagnostics 11, no. 10: 1810. https://doi.org/10.3390/diagnostics11101810
APA StyleCollins, T., Maktabi, M., Barberio, M., Bencteux, V., Jansen-Winkeln, B., Chalopin, C., Marescaux, J., Hostettler, A., Diana, M., & Gockel, I. (2021). Automatic Recognition of Colon and Esophagogastric Cancer with Machine Learning and Hyperspectral Imaging. Diagnostics, 11(10), 1810. https://doi.org/10.3390/diagnostics11101810











