Hybrid Binary Particle Swarm Optimization Differential Evolution-Based Feature Selection for EMG Signals Classification
Abstract
:1. Introduction
2. Preliminary
2.1. Binary Particle Swarm Optimization
2.2. Binary Differential Evolution
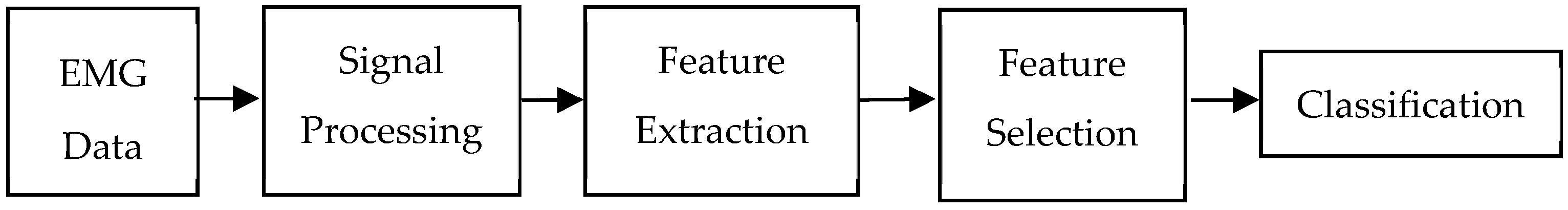
3. Materials and Methods
3.1. EMG Data
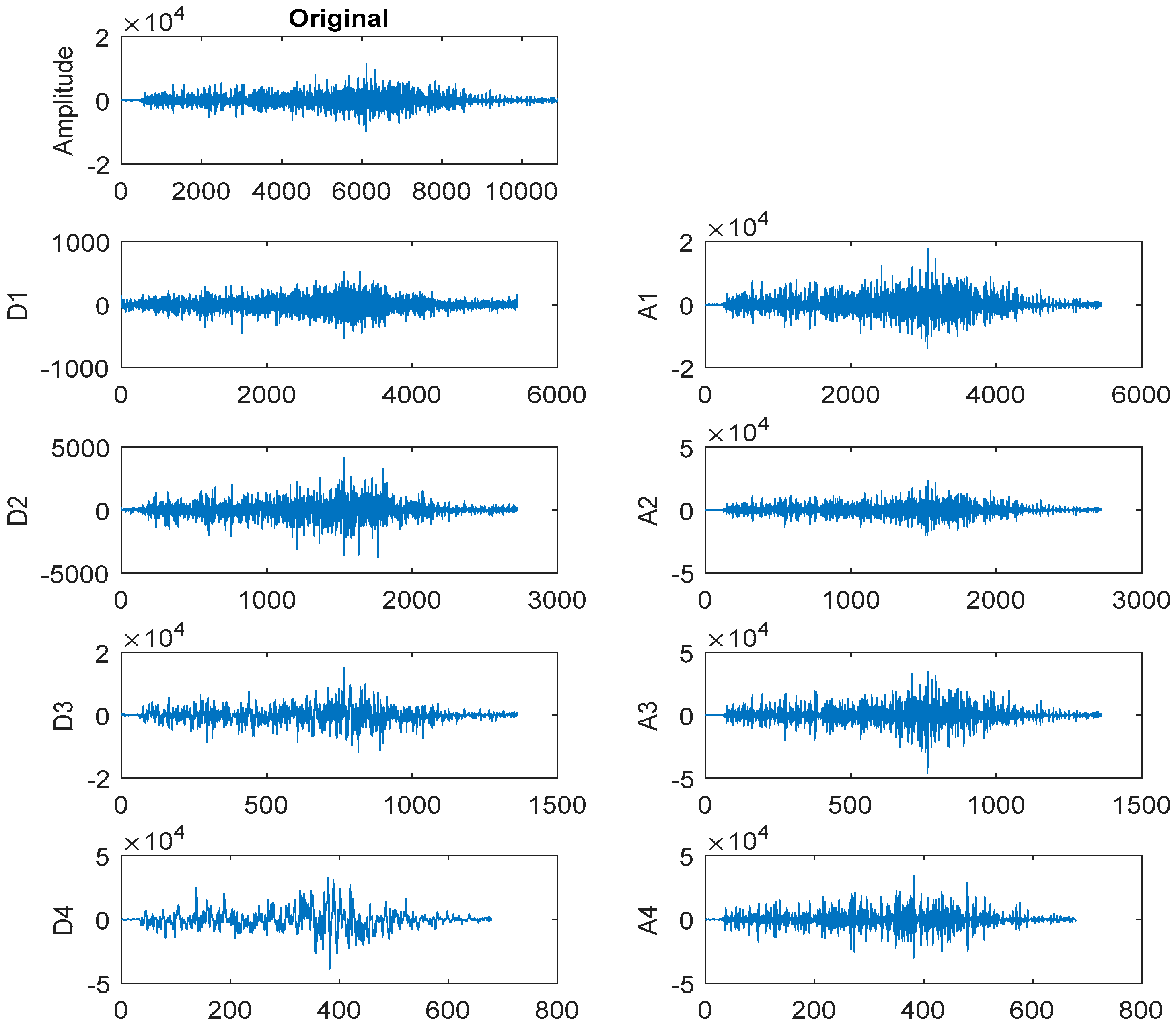
3.2. Discrete Wavelet Transform-Based Feature Extraction
3.3. Proposed Hybrid Binary Particle Swarm Optimization Differential Evolution
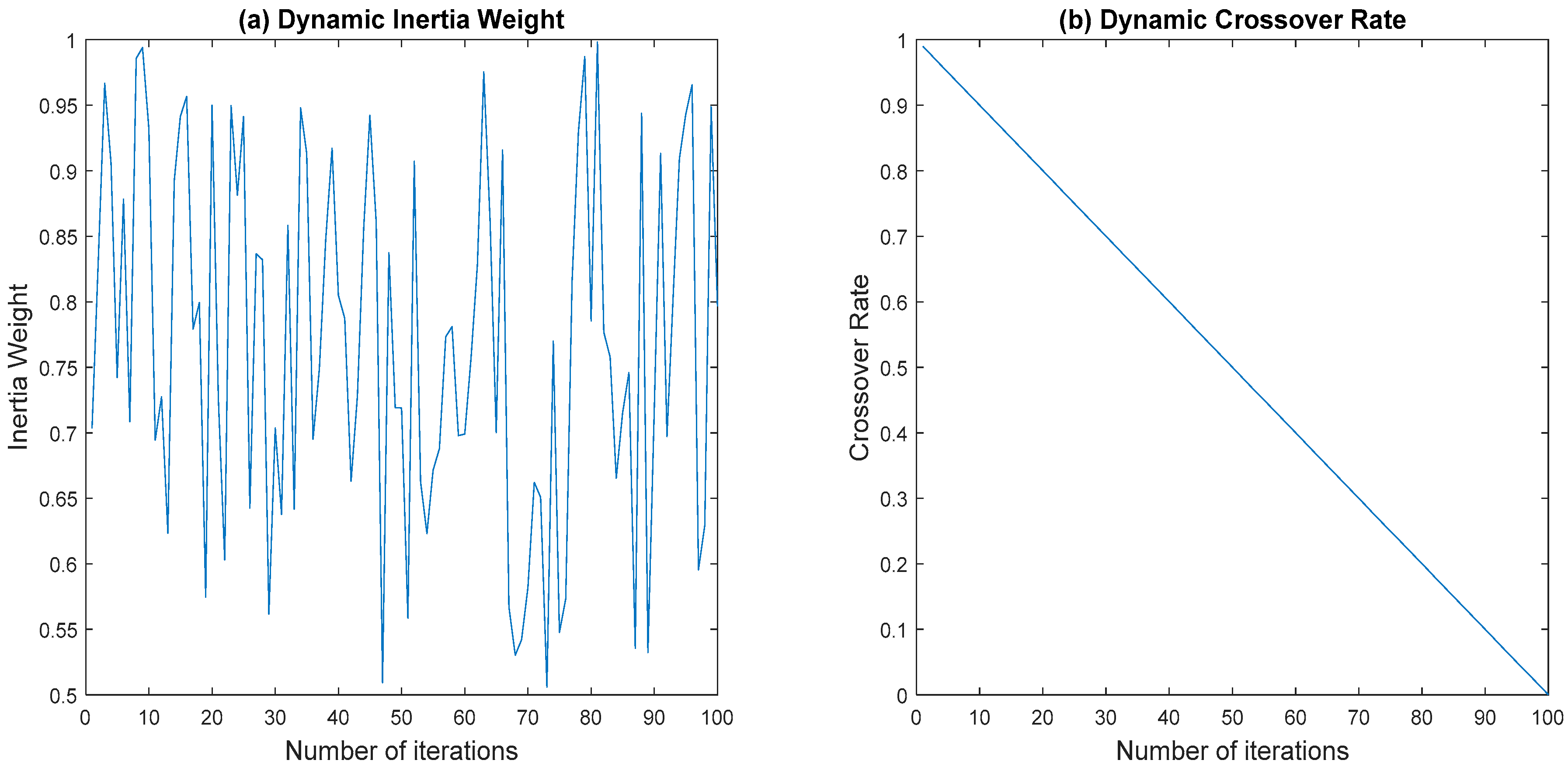
3.3.1. Dynamic Inertia Weight
3.3.2. Dynamic Crossover Rate
| Algorithm 1. Hybrid Binary Particle Swarm Optimization Differential Evolution |
| Input Parameters:N, T, c1, and c2 |
| (1) Randomly initialize a population of particles, x |
| (2) Evaluate the fitness of particles, F(x) |
| (3) Set Pbest and Gbest |
| (4) for t = 1 to maximum number of iterations, T |
| // BPSO Algorithm // |
| (5) if mod(t,2) = 1 |
| (6) |
| (7) for i = 1 to number of particles, N |
| (8) for d = 1 to number of dimension, D |
| (9) |
| (10) |
| (11) if |
| (12) |
| (13) else |
| (14) |
| (15) end if |
| (16) end for |
| (17) Evaluate the fitness of new particle, |
| (18) end for |
| // BDE Algorithm // |
| (19) else |
| (20) |
| (21) for i = 1 to number of particles, N |
| (22) Random select vectors and |
| (23) for d = 1 to number of dimension, D |
| (24) if |
| (25) |
| (26) else |
| (27) |
| (28) end if |
| (29) if |
| (30) |
| (31) else |
| (32) |
| (33) end if |
| (34) if |
| (35) |
| (36) else |
| (37) |
| (38) end if |
| (39) end for |
| (40) Evaluate the fitness of trial vector, |
| (41) Perform greedy selection between current particle and trial vector |
| (42) end for |
| (43) end if |
| // Pbest and Gbest Update // |
| (44) for i = 1 to number of particles, N |
| (45) Update Pbesti and Gbest |
| (46) end for |
| (47) end for |
| Output: Global best solution |
3.4. Application of BPSODE for Feature Selection
4. Results and Discussions
4.1. Comparison Algorithms and Evaluation Metrics
4.2. Experimental Results and Analysis
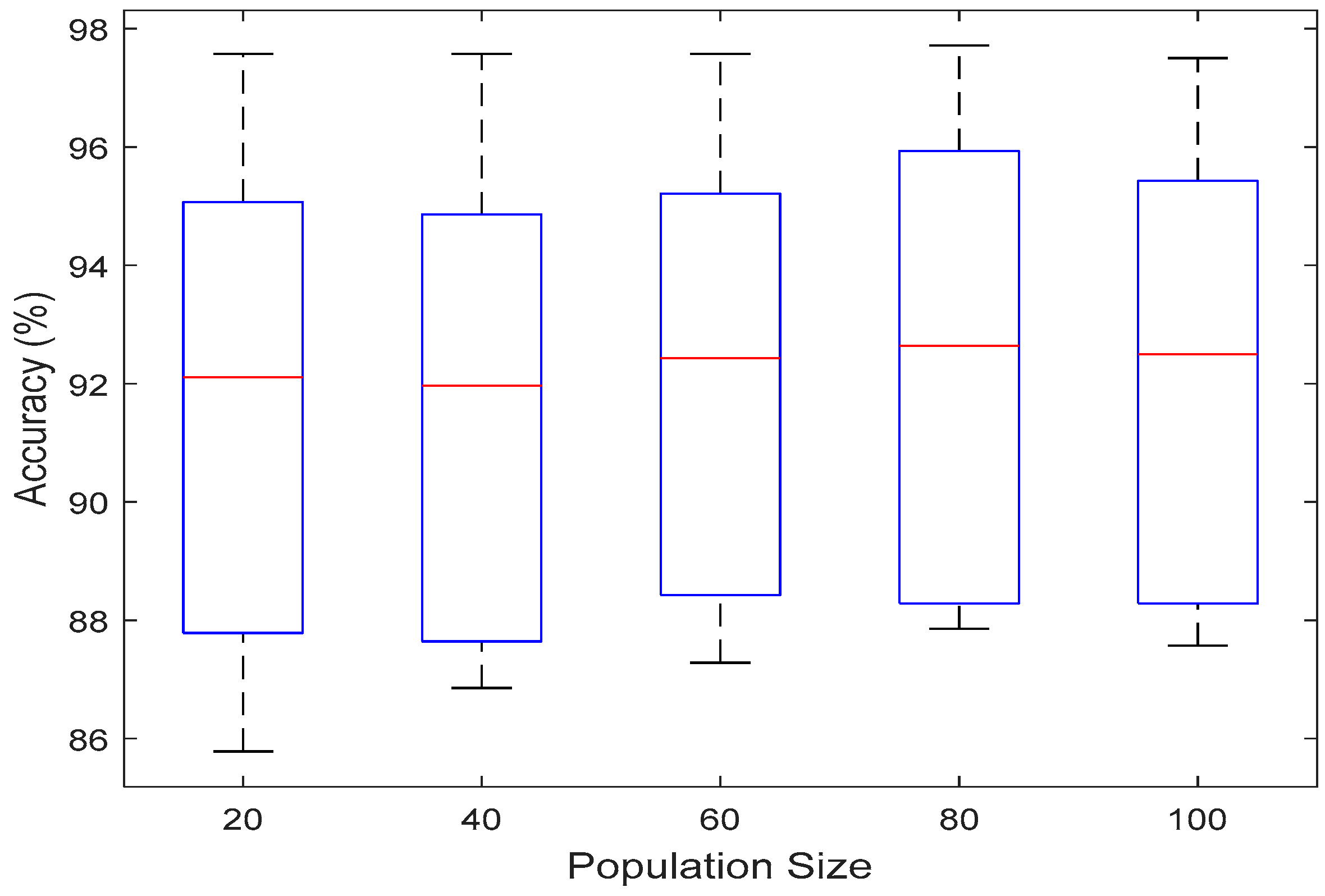
4.2.1. Effect of Population Size
4.2.2. Comparison Results
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Earley, E.J.; Hargrove, L.J.; Kuiken, T.A. Dual Window Pattern Recognition Classifier for Improved Partial-Hand Prosthesis Control. Front. Neurosci. 2016, 10. [Google Scholar] [CrossRef] [PubMed]
- Naik, G.R.; Kumar, D.K.; Palaniswami, M. Signal processing evaluation of myoelectric sensor placement in low-level gestures: sensitivity analysis using independent component analysis. Expert Syst. 2014, 31, 91–99. [Google Scholar] [CrossRef]
- Subasi, A. Classification of EMG signals using combined features and soft computing techniques. Appl. Soft Comput. 2012, 12, 2188–2198. [Google Scholar] [CrossRef]
- Phinyomark, A.; Limsakul, C.; Phukpattaranont, P. Application of Wavelet Analysis in EMG Feature Extraction for Pattern Classification. Meas. Sci. Rev. 2011, 11, 45–52. [Google Scholar] [CrossRef]
- Subasi, A. Classification of EMG signals using PSO optimized SVM for diagnosis of neuromuscular disorders. Comput. Biol. Med. 2013, 43, 576–586. [Google Scholar] [CrossRef] [PubMed]
- Xue, B.; Zhang, M.; Browne, W.N. Particle swarm optimisation for feature selection in classification: Novel initialisation and updating mechanisms. Appl. Soft Comput. 2014, 18, 261–276. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, H.; Liu, R.; Lv, W.; Wang, D. t-Test feature selection approach based on term frequency for text categorization. Pattern Recognit. Lett. 2014, 45, 1–10. [Google Scholar] [CrossRef]
- Ghosh, A.; Datta, A.; Ghosh, S. Self-adaptive differential evolution for feature selection in hyperspectral image data. Appl. Soft Comput. 2013, 13, 1969–1977. [Google Scholar] [CrossRef]
- Al-Ani, A.; Alsukker, A.; Khushaba, R.N. Feature subset selection using differential evolution and a wheel based search strategy. Swarm Evol. Comput. 2013, 9, 15–26. [Google Scholar] [CrossRef]
- Too, J.; Abdullah, A.R.; Mohd Saad, N. Binary Competitive Swarm Optimizer Approaches for Feature Selection. Computation 2019, 7, 31. [Google Scholar] [CrossRef]
- Banka, H.; Dara, S. A Hamming distance based binary particle swarm optimization (HDBPSO) algorithm for high dimensional feature selection, classification and validation. Pattern Recognit. Lett. 2015, 52, 94–100. [Google Scholar] [CrossRef]
- Bharti, K.K.; Singh, P.K. Opposition chaotic fitness mutation based adaptive inertia weight BPSO for feature selection in text clustering. Appl. Soft Comput. 2016, 43, 20–34. [Google Scholar] [CrossRef]
- Zorarpacı, E.; Özel, S.A. A hybrid approach of differential evolution and artificial bee colony for feature selection. Expert Syst. Appl. 2016, 62, 91–103. [Google Scholar] [CrossRef]
- Too, J.; Abdullah, A.R.; Mohd Saad, N. A New Co-Evolution Binary Particle Swarm Optimization with Multiple Inertia Weight Strategy for Feature Selection. Informatics 2019, 6, 21. [Google Scholar] [CrossRef]
- Das, S.; Konar, A.; Chakraborty, U.K. Improving Particle Swarm Optimization with Differentially Perturbed Velocity. Proceedings of Genetic and Evolutionary Computation, New York, NY, USA, 25–29 June 2005; ACM: New York, NY, USA, 2005; pp. 177–184. [Google Scholar] [CrossRef]
- Lin, G.-H.; Zhang, J.; Liu, Z.-H. Hybrid particle swarm optimization with differential evolution for numerical and engineering optimization. Int. J. Autom. Comput. 2018, 15, 103–114. [Google Scholar] [CrossRef]
- Mirjalili, S.; Mirjalili, S.M.; Yang, X.-S. Binary bat algorithm. Neural Comput. Appl. 2014, 25, 663–681. [Google Scholar] [CrossRef]
- Rodrigues, D.; Yang, X.-S.; Souza, A.N. de; Papa, J.P. Binary Flower Pollination Algorithm and Its Application to Feature Selection. In Recent Advances in Swarm Intelligence and Evolutionary Computation; Studies in Computational Intelligence; Springer: Cham, Switzerland, 2015; pp. 85–100. [Google Scholar] [CrossRef]
- Kennedy, J.; Eberhart, R.C. A discrete binary version of the particle swarm algorithm. In Proceedings of the Computational Cybernetics and Simulation, Orlando, FL, USA, 12–15 October 1997; pp. 4104–4108. [Google Scholar]
- Behera, H.S.; Dash, P.K.; Biswal, B. Power quality time series data mining using S-transform and fuzzy expert system. Appl. Soft Comput. 2010, 10, 945–955. [Google Scholar] [CrossRef]
- Unler, A.; Murat, A. A discrete particle swarm optimization method for feature selection in binary classification problems. Eur. J. Oper. Res. 2010, 206, 528–539. [Google Scholar] [CrossRef]
- Storn, R.; Price, K. Differential Evolution—A Simple and Efficient Heuristic for global Optimization over Continuous Spaces. J. Glob. Optim. 1997, 11, 341–359. [Google Scholar] [CrossRef]
- NinaPro Database—Non-Invasive Adaptive Hand Prosthetics. Available online: https://www.idiap.ch/project/ninapro/database (accessed on 31 March 2019).
- Pizzolato, S.; Tagliapietra, L.; Cognolato, M.; Reggiani, M.; Müller, H.; Atzori, M. Comparison of six electromyography acquisition setups on hand movement classification tasks. PLoS ONE 2017, 12, e0186132. [Google Scholar] [CrossRef]
- Ahila, R.; Sadasivam, V.; Manimala, K. An integrated PSO for parameter determination and feature selection of ELM and its application in classification of power system disturbances. Appl. Soft Comput. 2015, 32, 23–37. [Google Scholar] [CrossRef]
- Omari, F.A.; Hui, J.; Mei, C.; Liu, G. Pattern Recognition of Eight Hand Motions Using Feature Extraction of Forearm EMG Signal. Proc. Natl. Acad. Sci. India Sect. Phys. Sci. 2014, 84, 473–480. [Google Scholar] [CrossRef]
- Chowdhury, R.H.; Reaz, M.B.I.; Ali, M.A.B.M.; Bakar, A.A.A.; Chellappan, K.; Chang, T.G. Surface electromyography signal processing and classification techniques. Sensors 2013, 13, 12431–12466. [Google Scholar] [CrossRef] [PubMed]
- Phinyomark, A.; Phukpattaranont, P.; Limsakul, C. Fractal analysis features for weak and single-channel upper-limb EMG signals. Expert Syst. Appl. 2012, 39, 11156–11163. [Google Scholar] [CrossRef]
- Shi, Y.; Eberhart, R. A modified particle swarm optimizer. In Proceedings of the 1998 IEEE International Conference on Evolutionary Computation Proceedings, IEEE World Congress on Computational Intelligence, Anchorage, AK, USA, 4–9 May 1998; pp. 69–73. [Google Scholar]
- Jiao, B.; Lian, Z.; Gu, X. A dynamic inertia weight particle swarm optimization algorithm. Chaos Solitons Fractals 2008, 37, 698–705. [Google Scholar] [CrossRef]
- Taherkhani, M.; Safabakhsh, R. A novel stability-based adaptive inertia weight for particle swarm optimization. Appl. Soft Comput. 2016, 38, 281–295. [Google Scholar] [CrossRef]
- Khushaba, R.N.; Al-Ani, A.; Al-Jumaily, A. Feature subset selection using differential evolution and a statistical repair mechanism. Expert Syst. Appl. 2011, 38, 11515–11526. [Google Scholar] [CrossRef]
- Chuang, L.-Y.; Chang, H.-W.; Tu, C.-J.; Yang, C.-H. Improved binary PSO for feature selection using gene expression data. Comput. Biol. Chem. 2008, 32, 29–38. [Google Scholar] [CrossRef]
- Tawhid, M.A.; Dsouza, K.B. Hybrid Binary Bat Enhanced Particle Swarm Optimization Algorithm for solving feature selection problems. Appl. Comput. Inform. 2018. [Google Scholar] [CrossRef]
- Gokgoz, E.; Subasi, A. Comparison of decision tree algorithms for EMG signal classification using DWT. Biomed. Signal Process. Control 2015, 18, 138–144. [Google Scholar] [CrossRef]
- Li, Q.; Chen, H.; Huang, H.; Zhao, X.; Cai, Z.; Tong, C.; Liu, W.; Tian, X. An Enhanced Grey Wolf Optimization Based Feature Selection Wrapped Kernel Extreme Learning Machine for Medical Diagnosis. Comput. Math. Methods Med. 2017, 2017. [Google Scholar] [CrossRef] [PubMed]
- Adeli, A.; Broumandnia, A. Image steganalysis using improved particle swarm optimization based feature selection. Appl. Intell. 2018, 48, 1609–1622. [Google Scholar] [CrossRef]
- Taradeh, M.; Mafarja, M.; Heidari, A.A.; Faris, H.; Aljarah, I.; Mirjalili, S.; Fujita, H. An evolutionary gravitational search-based feature selection. Inf. Sci. 2019, 497, 219–239. [Google Scholar] [CrossRef]
| Algorithm | Parameter | Value |
|---|---|---|
| BPSODE | Acceleration coefficient, c1 and c2 | 2 |
| Bound on velocity | (−6,6) | |
| BDE | Crossover rate, CR | 1 |
| BPSO | Acceleration coefficient, c1 and c2 | 2 |
| Inertia weight, w | 0.9–0.4 | |
| Bound on velocity | (−6,6) | |
| BFPA | Switch probability, P | 0.8 |
| Levy component, λ | 1.5 | |
| BBA | Maximum frequency, fmax | 2 |
| Minimum frequency, fmin | 0 | |
| Control coefficient, α and γ | 0.9 | |
| Loudness, A | (1,2) | |
| Pulse rate, r | (0,1) |
| Subject | Metrics | Feature Selection Method | ||||
|---|---|---|---|---|---|---|
| BPSODE | BBA | BDE | BFPA | BPSO | ||
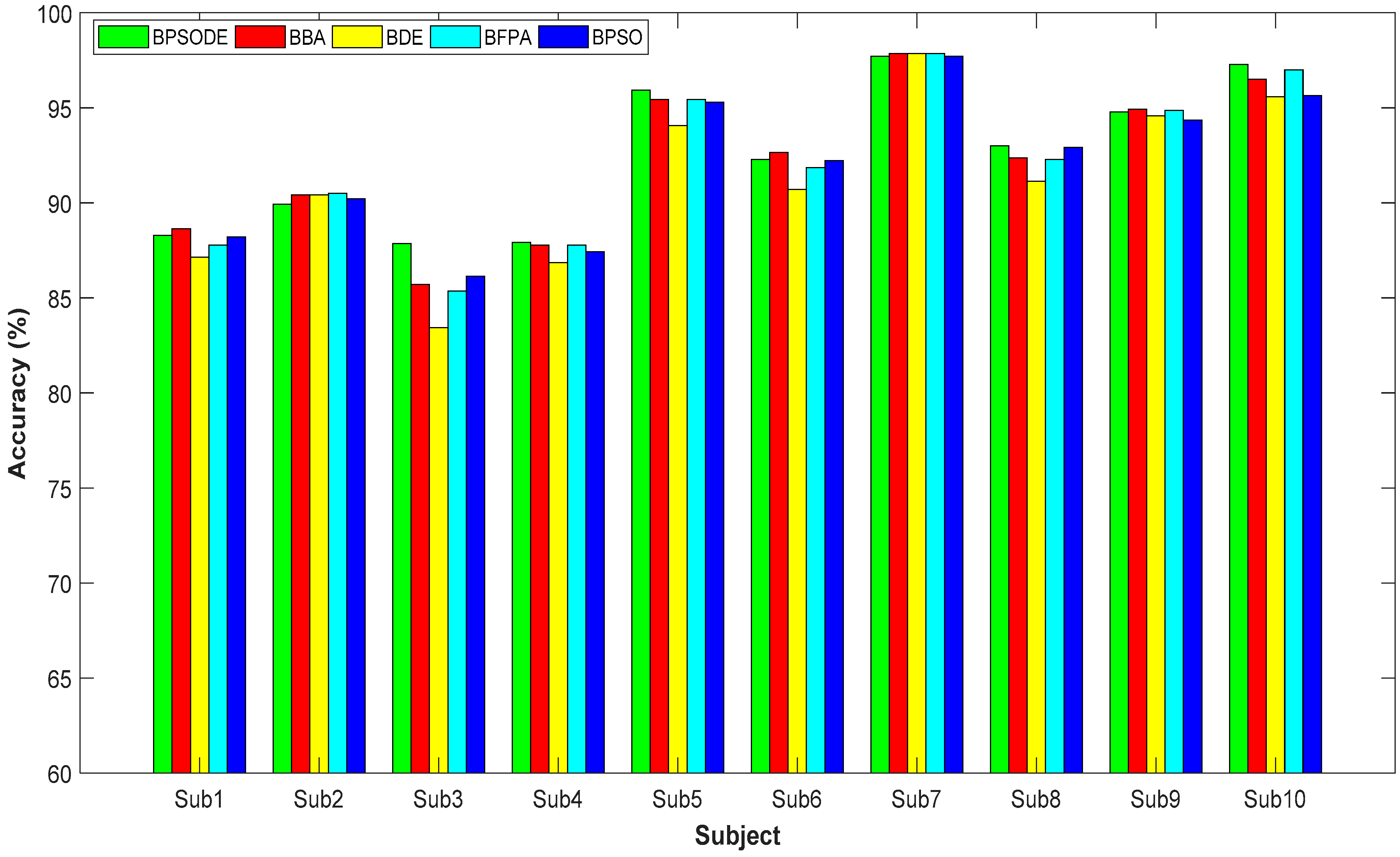
| 1 | Accuracy (%) | 88.29 ± 1.58 | 88.64 ± 1.64 | 87.14 ± 1.67 | 87.79 ± 1.70 | 88.21 ± 1.60 |
| Feature selection ratio (FSR) | 0.4196 ± 0.0283 | 0.4462 ± 0.0260 | 0.4926 ± 0.0329 | 0.5525 ± 0.0420 | 0.4486 ± 0.0250 | |
| Precision | 0.9034 ± 0.0108 | 0.9049 ± 0.0128 | 0.8941 ± 0.0156 | 0.9001 ± 0.0130 | 0.9023 ± 0.0120 | |
| F-measure | 0.8852 ± 0.0148 | 0.8880 ± 0.0165 | 0.8732 ± 0.0180 | 0.8798 ± 0.0169 | 0.8843 ± 0.0152 | |
| 2 | Accuracy (%) | 89.93 ± 1.35 | 90.43 ± 1.54 | 90.43 ± 1.24 | 90.50 ± 1.25 | 90.21 ± 1.16 |
| FSR | 0.4079 ± 0.0324 | 0.4424 ± 0.0167 | 0.4920 ± 0.0342 | 0.5594 ± 0.0550 | 0.4467 ± 0.0184 | |
| Precision | 0.9143 ± 0.0098 | 0.9182 ± 0.0132 | 0.9169 ± 0.0101 | 0.9190 ± 0.0106 | 0.9153 ± 0.0108 | |
| F-measure | 0.9018 ± 0.0120 | 0.9063 ± 0.0146 | 0.9061 ± 0.0111 | 0.9069 ± 0.0115 | 0.9042 ± 0.0105 | |
| 3 | Accuracy (%) | 87.86 ± 3.09 | 85.71 ± 2.22 | 83.43 ± 1.70 | 85.36 ± 1.60 | 86.14 ± 1.68 |
| FSR | 0.4386 ± 0.0289 | 0.4528 ± 0.0210 | 0.4981 ± 0.0489 | 0.5811 ± 0.0488 | 0.4636 ± 0.0185 | |
| Precision | 0.8949 ± 0.0303 | 0.8727 ± 0.0245 | 0.8519 ± 0.0181 | 0.8709 ± 0.0188 | 0.8779 ± 0.0186 | |
| F-measure | 0.8810 ± 0.0326 | 0.8588 ± 0.0227 | 0.8359 ± 0.0168 | 0.8555 ± 0.0160 | 0.8632 ± 0.0182 | |
| 4 | Accuracy (%) | 87.93 ± 1.35 | 87.79 ± 1.43 | 86.86 ± 1.89 | 87.79 ± 1.35 | 87.43 ± 1.36 |
| FSR | 0.4196 ± 0.0343 | 0.4393 ± 0.0207 | 0.4859 ± 0.0233 | 0.5581 ± 0.0523 | 0.4444 ± 0.0179 | |
| Precision | 0.8891 ± 0.0139 | 0.8870 ± 0.0146 | 0.8782 ± 0.0205 | 0.8880 ± 0.0136 | 0.8854 ± 0.0136 | |
| F-measure | 0.8754 ± 0.0135 | 0.8736 ± 0.0146 | 0.8653 ± 0.0194 | 0.8742 ± 0.0140 | 0.8705 ± 0.0136 | |
| 5 | Accuracy (%) | 95.93 ± 1.41 | 95.43 ± 1.19 | 94.07 ± 1.49 | 95.43 ± 0.99 | 95.29 ± 1.32 |
| FSR | 0.4157 ± 0.0318 | 0.4333 ± 0.0189 | 0.4820 ± 0.0168 | 0.5546 ± 0.0402 | 0.4426 ± 0.0221 | |
| Precision | 0.9654 ± 0.0103 | 0.9603 ± 0.0101 | 0.9505 ± 0.0121 | 0.9607 ± 0.0086 | 0.9593 ± 0.0111 | |
| F-measure | 0.9593 ± 0.0141 | 0.9544 ± 0.0119 | 0.9411 ± 0.0144 | 0.9546 ± 0.0099 | 0.9525 ± 0.0134 | |
| 6 | Accuracy (%) | 92.29 ± 1.42 | 92.64 ± 1.56 | 90.71 ± 1.50 | 91.86 ± 1.24 | 92.21 ± 1.57 |
| FSR | 0.4225 ± 0.0295 | 0.4436 ± 0.0216 | 0.4876 ± 0.0176 | 0.5663 ± 0.0434 | 0.4507 ± 0.0158 | |
| Precision | 0.9355 ± 0.0113 | 0.9389 ± 0.0121 | 0.9234 ± 0.0098 | 0.9338 ± 0.0087 | 0.9351 ± 0.0121 | |
| F-measure | 0.9258 ± 0.0136 | 0.9289 ± 0.0147 | 0.9111 ± 0.0152 | 0.9215 ± 0.0123 | 0.9251 ± 0.0151 | |
| 7 | Accuracy (%) | 97.71 ± 0.97 | 97.86 ± 0.87 | 97.86 ± 0.98 | 97.86 ± 0.98 | 97.71 ± 1.17 |
| FSR | 0.3824 ± 0.0361 | 0.4031 ± 0.0165 | 0.4627 ± 0.0111 | 0.4717 ± 0.0348 | 0.4149 ± 0.0200 | |
| Precision | 0.9788 ± 0.0087 | 0.9798 ± 0.0079 | 0.9799 ± 0.0092 | 0.9799 ± 0.0092 | 0.9789 ± 0.0102 | |
| F-measure | 0.9774 ± 0.0098 | 0.9785 ± 0.0092 | 0.9786 ± 0.0104 | 0.9786 ± 0.0104 | 0.9772 ± 0.0121 | |
| 8 | Accuracy (%) | 93.00 ± 1.60 | 92.36 ± 1.33 | 91.14 ± 1.44 | 92.29 ± 1.34 | 92.93 ± 1.43 |
| FSR | 0.4426 ± 0.0285 | 0.4536 ± 0.0272 | 0.5169 ± 0.0522 | 0.5676 ± 0.0485 | 0.4593 ± 0.0202 | |
| Precision | 0.9338 ± 0.0143 | 0.9278 ± 0.0125 | 0.9164 ± 0.0132 | 0.9268 ± 0.0121 | 0.9323 ± 0.0131 | |
| F-measure | 0.9295 ± 0.0160 | 0.9229 ± 0.0133 | 0.9110 ± 0.0143 | 0.9225 ± 0.0135 | 0.9287 ± 0.0141 | |
| 9 | Accuracy (%) | 94.79 ± 1.25 | 94.93 ± 1.57 | 94.57 ± 1.94 | 94.86 ± 2.24 | 94.36 ± 1.64 |
| FSR | 0.4065 ± 0.0411 | 0.4139 ± 0.0203 | 0.4813 ± 0.0270 | 0.5217 ± 0.0557 | 0.4332 ± 0.0207 | |
| Precision | 0.9563 ± 0.0097 | 0.9575 ± 0.0115 | 0.9543 ± 0.0161 | 0.9566 ± 0.0180 | 0.9518 ± 0.0143 | |
| F-measure | 0.9472 ± 0.0132 | 0.9483 ± 0.0165 | 0.9453 ± 0.0197 | 0.9478 ± 0.0235 | 0.9428 ± 0.0172 | |
| 10 | Accuracy (%) | 97.29 ± 1.13 | 96.5 ± 1.64 | 95.57 ± 1.38 | 97.00 ± 0.92 | 95.64 ± 1.35 |
| FSR | 0.4214 ± 0.0380 | 0.4343 ± 0.0177 | 0.4985 ± 0.0433 | 0.5785 ± 0.0314 | 0.4408 ± 0.0199 | |
| Precision | 0.9758 ± 0.0101 | 0.9680 ± 0.0154 | 0.9598 ± 0.0132 | 0.9731 ± 0.0087 | 0.9612 ± 0.0117 | |
| F-measure | 0.9731 ± 0.0114 | 0.9652 ± 0.0167 | 0.9561 ± 0.0142 | 0.9705 ± 0.0094 | 0.9570 ± 0.0129 | |
| Subject | p-Value | |||
|---|---|---|---|---|
| BBA | BDE | BFPA | BPSO | |
| 1 | 0.506287 | 0.056945 | 0.413356 | 0.847362 |
| 2 | 0.217023 | 0.109897 | 0.088031 | 0.384724 |
| 3 | 0.010163 | 2.00 × 10−5 | 0.018040 | 0.001725 |
| 4 | 0.629456 | 0.031698 | 0.666264 | 0.049260 |
| 5 | 0.109897 | 0.000358 | 0.129670 | 0.058264 |
| 6 | 0.425133 | 0.000462 | 0.249168 | 0.870789 |
| 7 | 0.605826 | 0.605826 | 0.605826 | 1.000000 |
| 8 | 0.131348 | 0.000265 | 0.135088 | 0.803685 |
| 9 | 0.693922 | 0.651311 | 0.894854 | 0.186411 |
| 10 | 0.085574 | 0.000499 | 0.329877 | 0.000490 |
| Win (w)/tie (t)/lose(l) | 1/9/0 | 6/4/0 | 1/9/0 | 3/7/0 |
| Subject | Average Computational Time(s) | ||||
|---|---|---|---|---|---|
| BPSODE | BBA | BDE | BFPA | BPSO | |
| 1 | 11.2170 | 9.6904 | 11.0745 | 10.9240 | 13.9385 |
| 2 | 11.4169 | 9.5965 | 11.3315 | 10.9586 | 13.6228 |
| 3 | 11.5064 | 9.7207 | 11.0580 | 11.4189 | 13.6346 |
| 4 | 11.6714 | 9.3344 | 11.2869 | 10.6936 | 13.4868 |
| 5 | 11.3360 | 9.3501 | 11.5490 | 11.0549 | 13.3068 |
| 6 | 11.5847 | 9.3415 | 11.2253 | 11.3795 | 13.2607 |
| 7 | 11.5799 | 9.2611 | 11.5731 | 11.1553 | 13.0535 |
| 8 | 11.8117 | 9.2501 | 11.9112 | 11.3934 | 13.3610 |
| 9 | 11.5575 | 9.1336 | 11.8026 | 11.2800 | 13.4043 |
| 10 | 11.7899 | 9.2060 | 11.7631 | 11.4669 | 13.3526 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Too, J.; Abdullah, A.R.; Mohd Saad, N. Hybrid Binary Particle Swarm Optimization Differential Evolution-Based Feature Selection for EMG Signals Classification. Axioms 2019, 8, 79. https://doi.org/10.3390/axioms8030079
Too J, Abdullah AR, Mohd Saad N. Hybrid Binary Particle Swarm Optimization Differential Evolution-Based Feature Selection for EMG Signals Classification. Axioms. 2019; 8(3):79. https://doi.org/10.3390/axioms8030079
Chicago/Turabian StyleToo, Jingwei, Abdul Rahim Abdullah, and Norhashimah Mohd Saad. 2019. "Hybrid Binary Particle Swarm Optimization Differential Evolution-Based Feature Selection for EMG Signals Classification" Axioms 8, no. 3: 79. https://doi.org/10.3390/axioms8030079
APA StyleToo, J., Abdullah, A. R., & Mohd Saad, N. (2019). Hybrid Binary Particle Swarm Optimization Differential Evolution-Based Feature Selection for EMG Signals Classification. Axioms, 8(3), 79. https://doi.org/10.3390/axioms8030079





