Abstract
UV-LEDs are a new method of disinfecting drinking water. Some viruses are very resistant to UV and the efficiency of UV-LEDs to disinfect them needs to be studied. Drinking water was disinfected with UV-LEDs after spiking the water with MS2 and four UV- and/or Cl-resistant coliphages belonging to RNA or DNA coliphages isolated from municipal wastewater. UV-LEDs operating at a wavelength of 270 nm for 2 min with 120 mW of irradiation caused 0.93–2.73 Log10-reductions of coliphages tested in a reactor of a 5.2 L volume. Irradiation time of 10 min in the same system increased the Log10-reductions to 4.30–5.16. Traditional mercury UV (Hg-UV) lamp at a 254 nm wavelength caused 0.67–4.08 Log10-reductions in 2 min and 4.56–7.21 Log10-reductions in 10 min in 10 mL of water. All coliphages tested except MS2 achieved 4 Log10-reductions with UV-LEDs at a dose that corresponded to 70 mWs/cm2 using Hg-UV. Thus, UV-LEDs are a promising method of disinfecting UV- and/or Cl-resistant viruses.
1. Introduction
Disinfection of water is essential, since contamination of water with pathogens, such as enteric viruses, may lead to diarrheal cases, the number of which is calculated to be more than 1.6 billion each year [1]. Many enteric viruses have similarities with coliphage viruses having RNA or DNA in their genome [2]. Male-specific RNA coliphages such as MS2 are often used as indicators for monitoring virological quality of water [3].
UV irradiation is one option when disinfecting microbes which may be resistant to the chemical disinfection of water [4]. The UV treatment needs only a short contact time compared to chemical disinfection [5,6], and it does not form disinfection by-products or change the taste or odor of water. In addition, UV irradiation does not cause corrosion in the water distribution system.
UV light emitting diodes (UV-LEDs) are an emerging technology, which may be a substitute for Hg-UV if they are shown to be as effective as traditional mercury lamps. The UV-LEDs consist of two semiconductors connected with a p (positive charge) –n (negative charge) junction to emit light in a narrow electromagnetic spectrum based on electroluminescence. The emitting wavelengths can be tailored with the choice of materials. Aluminum gallium nitride and aluminum nitride are the most used materials [7,8].
Advantages of UV-LEDs are that they do not contain any toxic mercury and the materials can be recycled. They consume much less energy than Hg-UV, can be repeatedly turned on and off without waiting times, and have a potential lifetime of approximately 100,000 h [8], i.e., 10 times longer than that of Hg-UV lamps [9]. In addition, they are small and fit in many places.
Disinfection with both Hg-UV and UV-LED relies on the ability of UV irradiation at wavelengths of 200–300 nm to damage nucleic acids and proteins [10,11,12]. So far, there are several studies on the effect of UV-LEDs within the range of 250 and 365 nm on microorganisms [8,13]. Most studies have focused on Escherichia coli [13,14,15,16,17,18,19,20,21,22,23], but there are fewer studies on viruses such as coliphages, which may better indicate the presence of pathogenic viruses than traditional indicator bacteria. For coliphages, wavelengths between 255 and 285 nm [8,13,14,15,16,17,18,21,23,24] have been tested.
The goal of this study was to compare efficiency of UV-LEDs at 120 mW and a traditional Hg-UV lamp at 30 W in disinfection of UV- and/or Cl-resistant DNA and RNA coliphages. The wavelength and volume of water for UV-LED were 270 nm and 5.2 L, respectively, and for Hg-UV, 254 nm and 10 mL, respectively. To our knowledge, the 270 nm wavelength has not been studied with UV-LEDs for virus disinfection before.
2. Materials and Methods
2.1. Isolation and Purification of Coliphages
In total, 18 coliphage strains were isolated and purified from the effluent of the Lehtoniemi municipal wastewater treatment plant and cultivated as described earlier [25]. The host bacteria were grown in TYG broth (tryptose 10 g, yeast extract 5 g, glucose 2 g, NaCl 5 g, and MgSO4·7H2O 0.25 g added to 1000 mL of deionized water). The bacteria were incubated in a shaker (Edison, NJ, USA) for 24 h at 37 °C, and then rejuvenated similarly for 2 h to reach the cell density of approximately 109 colony forming units/mL. Escherichia coli ATCC 13706 was used for detecting somatic coliphages, and E. coli ATCC 15597 was used for F-specific coliphages (Table 1). The isolated coliphages and MS2 (ATTC 15597-B1) as an indicator virus were rejuvenated in TYG broth as stock solutions of approximately 1011 PFU/mL and used in the disinfection experiments.

Table 1.
Genetic material of coliphages tested with RNase spot test and sensitivity of coliphages to Hg-UV and chlorine.
2.2. Identification of Genetic Material of Coliphages with RNase Spot Test
Coliphage isolates were tested with an RNase enzyme (Ribonuclease Type I-A, ≥50 Kunitz units/mg protein, Sigma-Aldrich, St. Louis, MO, USA) to distinguish if the viral nucleic acid content was RNA or DNA. Fresh host bacterium of 3 mL was added to 150 mL of TYG agars (TYG broth with 12 g agar for 1000 mL) melted and tempered in a water bath at 44.5 °C. RNase was added so that the final concentration was 100 μg/mL. Water instead of RNase was used as a negative control. The mixed solutions were poured into 150-mm-diameter Petri dishes. Five microliters of coliphage suspensions with dilutions from 10−1 to 10−4 were spotted onto both RNase-positive and RNase-negative plates. The plates were incubated overnight at 37 °C. If plaques were observed on plates with and without RNase enzyme, the phage was considered to be a DNA-coliphage. If the plaques were observed only on the plates without RNase, the phage was considered to be an RNA-coliphage [26].
2.3. UV Experiments
Four of the 18 strains and MS2 were selected for the UV comparison experiments based on their genetic material (DNA or RNA) and resistance against UV- and/or Cl. Two UV reactors were used in the experiments. The first reactor was a collimator (Figure 1a) using a low-pressure mercury arc lamp (Osram HNS 30 W, λ = 253.7 nm, Munich, Germany) with an average intensity of 0.1661 mW/cm2 taking into account reflection factor, Petri factor, water factor, and divergence factor according to Bolton and Linden [27]. Ten milliliters of the coliphage stock solution was diluted with deionized water to a density of approximately 107 PFU/mL. This was then pipetted to a sterile glass Petri dish (inner diameter 6.0 cm) and mixed with a magnetic stirrer. The UV irradiation beam was directly focused onto the Petri dish with exposure times of 2, 4, 7, and 10 min corresponding to the UV doses of 20, 40, 70, and 100 mWs/cm2, respectively, as described earlier in details [25]. The tests for each sampling time (each UV-dose) were done separately so that the water volume was always 10 mL at the start.
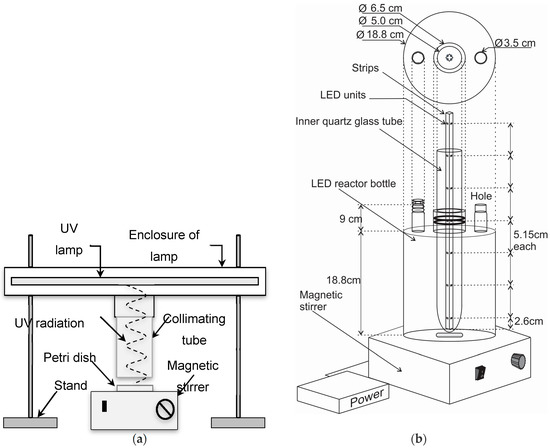
Figure 1.
(a) Reactor type of 10 mL for Hg-UV in collimator; (b) Reactor type of 5.2 L for UV-LED with four LEDs at three different heights inside the water.
The second reactor was a device using UV-LEDs manufactured by SETi (Columbia, SC, USA). The reactor consisted of a glass bottle (height 18.8 cm, inner diameter 18.8 cm) with a narrow neck (inner diameter of 6.5 cm) through which an inner quartz glass tube was set (outer diameter: 5.0 cm; inner diameter: 4.6 cm). The quartz glass tube was tightened to a vertical position with rubber rings. The LED-unit was inserted inside this quartz glass tube. The LED unit consisted of four strips that were fixed together in a square format and fastened tightly with plugs to a vertical position. Each strip had seven LEDs (5.15 cm from each other), three of which were immersed in to the water so that there were 12 LEDs disinfecting water (Figure 1b). Each strip was operated with a current of 120 mA and a voltage of 5.5 V. The wavelength of the LEDs was 270 nm, the power of each LED was 10 mW (totally 120 mW), and the opening angle was 120°, according to the manufacturer. During the experiments, the water in the reactor was mixed with a magnetic stirrer. The whole reactor was covered with a hood the inside of which was lined with aluminum foil.
Kuopio municipal drinking water was used for the UV-LED experiments. The quality of the water was described by Kuopion Vesi [28] to be 0.09 FTU for turbidity, 1.4 mg/L for CODMn, 7.7 for pH, and 0.44 mg Cl2/L for residual Cl. The transmittance of the spiked water used in the UV-LED experiments was calculated from the absorbance measured with a spectrophotometer (UV-2401PC, Shimadzu, Kyoto, Japan) at a wavelength of 270 nm, and was 90%. Drinking water was flushed from a tap for at least 3 min before it was used. In total, 15.8 L of water was collected in a beaker and left at room temperature for 24 h for the chlorine to evaporate. The concentrations of total and free chlorine in the evaporated water were measured with colorimetric analyses of DPD (N,N-diethyl-p-phenylenediamine) according to the Standard Method 4500-Cl G for drinking water and wastewater analyses [29] using Hach DR 2800 spectrophotometer and methods 8167 (total chlorine) and 8021 (free chlorine) according to the manufacturer’s instructions. The total and free chlorine concentrations were 0.03 ± 0.005 mg/L and 0.01 ± 0.01 mg/L, respectively; therefore, the residual chlorine was not quenched.
The volume of 5.2 L of dechlorinated drinking water was poured into the LED reactor bottle, and the coliphage stock solution was added with the initial concentration of approximately 107 PFU/mL. The suspension was stirred with a magnetic stirrer for 3 min after which the zero time sample was taken via a sampling hole at the top of the bottle. The LED irradiation was started, and water samples were taken after 2, 4, 5.5, 7, 10, 12, and 15 min without switching off the LED irradiation for the samplings. All samples were stored at 4 °C until coliphage analyses were performed within an hour. The experiments were done as three parallels on the same day using the same coliphage stock solution.
The densities of coliphages were analyzed with a double layer technique [30,31] to calculate the reductions after exposure times. The tests were carried out at room temperature (20–21 °C).
2.4. Calculations and Statistical Analyses
Inactivation values (average ± standard deviation) were calculated as the Log10 of N/N0, where N is the coliphage density after the treatment and N0 the density before the treatment. Non-parametric tests (related sample Friedman’s two-way analysis) using SPSS version 22 were performed in order to determine if the coliphage Log10-reductions after exposures differed significantly from the control (UV dose 0 mWs/cm2). Linear regression equations (average ± standard deviation) with Log10-reductions of coliphages and disinfection time for each coliphage and UV treatment were calculated with Excel 2013. Inactivation with Hg-UV and UV-LED disinfection was compared by analyzing the slopes of three parallel linear regression equations of both methods by Wilcoxon signed rank testing (SPSS 22).
3. Results
MS2 and two tested coliphages belonged to RNA phages and two others belonged to DNA phages (Table 1). The resistances of these strains to UV and/or chlorine were tested earlier and are shown in Table 1.
The UV-LEDs caused 0.93–2.73 Log10-reductions after 2 min of contact time for Strains 1, 5, 7, and 17, and 10 min of contact time increased the Log10-reductions to 4.30–5.16 (Figure 2). Hg-UV irradiation achieved 0.67–4.08 Log10-reductions within 2 min and 4.56–7.21 Log10-reductions within 10 min. Four Log10-reductions of Strains 1, 5, 7 and 17 were achieved with UV-LEDs within 7 min of contact time which corresponds to the dose of 70 mWs/cm2 using Hg-UV in a collimator. Log10-reductions achieved with contact times 7 min and longer differed statistically significantly (p < 0.05) from the controls. MS2 differed from other coliphages by being more UV-resistant (Figure 2). After 15 min of contact time, the Log10-reductions with UV-LEDs increased to 4.78–6.67 for Strains 1, 5, 7, and 17 and to 1.47 for MS2 (Figure 2).
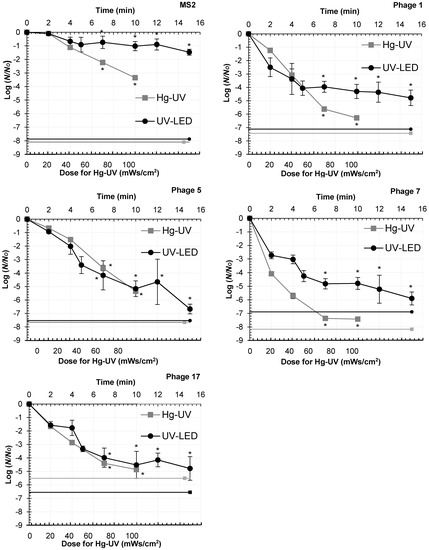
Figure 2.
Inactivation of MS2 and four UV- and/or Cl-resistant coliphages (1, 5, 7 and 17) with Hg-UV and UV-LED treatment. Statistically significant differences from the control (UV dose 0 mWs/cm2), assessed by related sample Friedman’s two-way analysis of variance, are indicated with asterisks, * p < 0.05. The lines show the theoretical maximal Log10-reductions for each coliphage. For  Hg-UV and for
Hg-UV and for  UV-LEDs.
UV-LEDs.
 Hg-UV and for
Hg-UV and for  UV-LEDs.
UV-LEDs.
The slopes for the linear regression equations (Table 2) were statistically similar between UV-LEDs and Hg-UV during 10 min of disinfection. The p-values for the slopes of all coliphage strains were > 0.1. The slopes of the linear regression equations for 15 min with UV-LEDs were lower than the corresponding slopes for 10 min indicating a tailing effect in disinfection (Table 2 and Figure 2).

Table 2.
Linear regression equations for inactivation of coliphages using UV-LEDs for 0–15 and 0–10 min contact times and Hg-UV for 0–10 min contact times (n = 3) (average slope ± standard deviation).
4. Discussion
We found that UV-LEDs at 270 nm were sufficient to inactivate DNA and RNA coliphages in water even when the water volume was as high as 5.2 L with water layer thicknesses of 6.7 cm. The inactivations of RNA Coliphage Strain 5 and DNA Coliphage Strain 17 in UV-LED treatment were similar to the inactivation in traditional Hg-UV treatment where the water volume was 10 mL and the thickness only 0.35 cm (Figure 2). For the other strains, inactivation with UV-LEDs seemed to be lower than with Hg-UV, as seen from the smaller slopes of linear regression equations, but statistically there was no difference between the UV-methods in these strains either (Table 2). More than 4 Log10-inactivations were reached for all strains except for MS2 (Figure 2). Our results thus support other results that show that inactivation kinetics are similar to both UV-LEDs and Hg-UV if tested with MS2 and T7 [18] or Escherichia coli, MS2, and Bacillus atrophaeus [23].
UV-LEDs achieved 3 Log10-reductions of Coliphage Strains 1, 5, 7, and 17 within 4 min of contact time and 4 Log10-reductions within 7 min, which corresponded to doses of 40 mWs/cm2 and 70 mWs/cm2 in the Hg-UV collimator, respectively. In earlier UV-LED studies more than 3 Log10-reductions of coliphages φX174 and T7 were achieved with doses of 6.4–20 mWs/cm2 at wavelengths of 255–285 nm [18,24]. Coliphages Qβ and MS2 were more resistant and they required above 40 mWs/cm2 to reach 1–3 Log10-reductions [13,16,18,21,23,24]. Bowker et al. [18] compared the wavelengths of 255 and 275 nm with coliphages T7 and MS2 and found rather similar inactivation rates at both wavelengths. Qβ and MS2 were studied by [13,14,16] at wavelengths of 260, 275, and 280 nm and a wavelength of 260 was found to be more effective compared with the other wavelengths. Aoyagi et al. [24] compared the wavelengths of 255 nm and 280 nm with different coliphages (φX174, Qβ, and MS2) and concluded that, even though the inactivation efficiency of 280 nm was lower than that of 255 nm, 280 nm LEDs are more suitable for practical applications due to their easier production at high-power output. It seems, regardless of this, that the tested wavelengths between 255 and 280 nm of UV-LEDs can efficiently inactivate coliphages.
In our study, MS2 was shown to be resistant to UV irradiation and it achieved a maximum 1.5 and 3.5 Log10-reductions with UV-LEDs and Hg-UV, respectively, with doses corresponding to 70 mWs/cm2 in the Hg-UV collimator. MS2 has also proved to be UV-resistant in other studies. Doses from 41 to 60 mWs/cm2 were needed to reach 2–3 Log10-reductions of MS2 at wavelengths of 255–280 nm (excluding 270 nm) [13,18,24], while approximately 110 mWs/cm2 was needed at 285 nm to reach 3 Log10-reduction [21]. However, a tunable laser at 270 nm has yielded 1.5 and at least 3 Log10-reductions of MS2 with doses of 20–30 mWs/cm2 and 60 mWs/cm2, respectively [33,34]. The tunable laser at a lower wavelength of 253.7 nm yielded approximately 3.5 Log10-reduction of MS2 with a dose of 70 mWs/cm2 [33], which corresponds to our result.
MS2 and other UV-resistant coliphages tested by [13,16,18,21,24] were RNA viruses. In our study, however, either RNA or DNA coliphages could be UV-resistant or UV-sensitive. Resistance of MS2 makes it a good surrogate for human enteric viruses when the efficiency, wavelength, and energy in UV disinfection are studied. Human pathogenic adenovirus 5, which is a DNA virus, seems to be slightly more resistant to UV-LED at 285 nm than MS2 [21].
It is difficult to compare the efficiency of different wavelengths to disinfect coliphages since the reactor configurations, water volumes, and types of water are different in different UV-LED studies. Earlier studies have used sterilized water in a vertical laboratory system [14,16,24], saline-calcium or phosphate buffer saline in a collimated beam system [18,23], or 50 mL of phosphate buffer saline in a ring-shaped reactor [21]. We used 5.2 L of real municipal drinking water in the reactor where UV-LEDs were placed centrally facing in four directions. The longest distance to the wall of the LED, i.e., the thickness of the water layer, was rather high, approximately 6.7 cm, while the distance was 0.158 cm in a collimated beam system [23]. Oguma et al. [21] used 20 LEDs on the inner side walls of a ring with a diameter of 5.3 cm (a radius of 2.65 cm). Opening angle of our LEDs was 120°, but there may have been a shadow somewhere in the reactor, e.g., in the bottom under the inner glass tube. A reactor where LEDs have been set in a ring position to the walls or a tube might overcome these problems and reduce standard deviations and tailing of the inactivation.
A ring-shaped reactor operating with 285 nm LEDs [35] and a rectangular point-of-use reactor operating with 260 or 275 nm [14,16] have been tested with flowing water. Compared to the batch reactors, the inactivations of MS2 and Qβ have been lower in both systems (maximum 1.2 Log10 of coliphage Qβ with a flow rate of 400 mL/min [35] and 1.6 Log10 of coliphage Qβ with a flow rate of 109 mL/min [14,16]). In the future, inactivation may be improved by modifying the geometry of the UV reactor [35]. The output powers of the current UV-LEDs are rather low but they are increasing. In previous works, 0.3 and 0.5 mW LEDs [18] and 1.3 mW LEDs [21,35] have been used, and we used 10 mW LEDs. These output powers are still very moderate compared with traditional low-pressure UV lamps, the power of which can be on the level of 30–40 W. The development in LED technology, however, will enable higher power and hence higher doses of UV-LEDs. Thus, this technology shows promise for the future disinfection of high water flows.
LED technology enables simultaneous use of LEDs emitting different wavelengths. It is therefore possible to affect different molecules, which have different absorption peaks for UV, and cause damage that the cells cannot repair. Nucleotides have UV absorbance peaks between 240 and 280 nm and absorption maximum of DNA is considered to be near 260 nm [36]. UV-absorption of proteins generally peaks at 280 nm [37] although proteins of some viruses may efficiently be affected by wavelengths below 240 nm [33,34]. In addition, wavelengths on UVA region (above 315 nm) may produce reactive intermediates and oxidative damage to DNA and other components of the bacterial cell [38,39]. Compared to a single wavelength, simultaneous treatment with multiple wavelengths of UV-C and UV-A have yielded higher reductions of fecal enterococci and total and fecal coliforms in wastewater and in pure cultures [40,41] and Vibrio parahaemolyticus in pure culture [42]. On the other hand, the UV-LEDs at wavelengths 260 nm and 280 nm irradiated separately or simultaneously yielded statistically similar reductions in E. coli, Bacillus pumilus spores, MS2, or adenovirus when the UV-doses were the same [13]. More studies are obviously needed on the effect of different wavelength combinations on viruses and other resistant microorganisms.
Combined treatments of traditional Hg-UV and chlorine have also shown synergistic disinfection efficiency of coliphages [25,43,44]. Therefore, combining UV-LEDs with a low concentration of chlorine would be worth studying.
5. Conclusions
Our results showed that UV-LEDs at 270 nm are effective for the inactivation of most DNA and RNA viruses tested. Regardless of the results, the efficiency of different wavelengths and wavelength combinations should be further studied. UV-LEDs with low output power showed high efficiency even though we used real unsterilized municipal drinking water with high water volume and water thickness. Therefore, we conclude that UV-LEDs are a promising method for future water disinfection when the output power of the LEDs becomes more powerful.
Acknowledgments
The authors acknowledge The University of Baghdad for financial support of Alyaa M. Zyara and the Academy of Finland (decision number 277639) for other financial support. We thank LED Suutari Oy for providing the LEDs and for cooperation in LED technology. We also thank Sirpa Martikainen for assistance in laboratory work, and Daniel Blande (M. Sc.) for the revision of the English language.
Author Contributions
Alyaa M. Zyara performed the experiments and analyzed the data under the supervision of Helvi Heinonen-Tanski, Anna-Maria Veijalainen and Eila Torvinen. All authors contributed to the writing of the paper.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Water Health Organization (WHO). Diarrhoeal Disease; World Health Organization: Geneva, Switzerland, 2015. [Google Scholar]
- Love, D.; Vinje, J.; Khalil, S.; Murphy, J.; Lovelace, G.; Sobsey, M. Evaluation of RT-PCR and reverse line blot hybridization for detection and genotyping F+ RNA coliphages from estuarine waters and molluscan shellfish. J. Appl. Microbiol. 2008, 104, 1203–1212. [Google Scholar] [CrossRef] [PubMed]
- Long, S.C.; El-Khoury, S.S.; Oudejans, S.J.G.; Sobsey, M.D.; Vinjé, J. Assessment of sources and diversity of male-specific coliphage for source tracking. Environ. Eng. Sci. 2005, 22, 367–377. [Google Scholar] [CrossRef]
- Hijnen, W.A.M.; Beerendonk, E.F.; Medema, G.J. Inactivation credit of UV radiation for viruses, bacteria and protozoan (oo) cysts in water: A review. Water Res. 2006, 40, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Wright, H.B.; Cairns, W.L. Ultraviolet Light. In Proceedings of the Regional Symposium on Water Quality: Effective Disinfection, Lima, Peru, 27–29 October 1998; pp. 1–26.
- U.S. Environmental Protection Agency (EPA). Alternative Disinfectants and Oxidants Guidance Manual; EPA Office of Water: Washington, DC, USA, 1999.
- Tamulaitis, G. Ultraviolet light emitting diodes: Review. LITH J. Phys. 2011, 51, 177–193. [Google Scholar] [CrossRef]
- Song, K.; Mohseni, M.; Taghipour, F. Application of ultraviolet light-emitting diodes (UV-LEDs) for water disinfection: A review. Water Res. 2016, 94, 341–349. [Google Scholar] [CrossRef] [PubMed]
- Heinonen-Tanski, H.; Juntunen, P.; Rajala, R.; Haume, E.; Niemelä, A. Costs of tertiary treatment of municipal wastewater by rapid sand filter with coagulants and UV. Water Sci. Technol. Water Supply 2003, 3, 145–152. [Google Scholar]
- Environmental Protection Agency (EPA). Challenge Organisms for Inactivation of Viruses by Ultraviolet Treatment; Water Research Foundation: Denver, CO, USA, 2010.
- Eischeid, A.C.; Linden, K.G. Molecular indications of protein damage in adenovirus after UV disinfection. Appl. Environ. Microbiol. 2011, 77, 1145–1147. [Google Scholar] [CrossRef] [PubMed]
- National Water Research Institute (NWRI). Ultraviolet Disinfection Guidelines for Drinking Water and Water Reuse, 3rd ed.; National Water Research Institute in Collaboration with Water Research Foundation: Fountain Valley, CA, USA, 2012. [Google Scholar]
- Beck, S.E.; Ryu, H.; Boczek, L.A.; Cashdollar, J.L.; Jeanis, K.M.; Rosenblum, J.S.; Lawal, O.R.; Linden, K.G. Evaluating UV-C LED disinfection performance and investigating potential dual wavelength synergy. Water Res. 2017, 109, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Jenny, R.M.; Jasper, M.N.; Simmons, O.D., III; Shatalov, M.; Ducoste, J.J. Heuristic optimization of a continuous flow point-of-use UV-LED disinfection reactor using computational fluid dynamics. Water Res. 2015, 83, 310–318. [Google Scholar] [CrossRef] [PubMed]
- Vilhunen, S.; Särkkä, H.; Sillanpää, M. Ultraviolet light-emitting diodes in water disinfection. Environ. Sci. Pollut. Res. 2009, 16, 439–442. [Google Scholar] [CrossRef] [PubMed]
- Jenny, R.M.; Simmons, O.D., III; Shatalov, M.; Ducoste, J.J. Modeling a continuous flow ultraviolet light emitting diode reactor using computational fluid dynamics. Chem. Eng. Sci. 2014, 116, 524–535. [Google Scholar] [CrossRef]
- Chatterley, C.; Linden, K. Demonstration and evaluation of germicidal UV-LEDs for point-of-use water disinfection. J. Water Health 2010, 8, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Bowker, C.; Sain, A.; Shatalov, M.; Ducoste, J. Microbial UV fluence-response assessment using a novel UV-LED collimated beam system. Water Res. 2011, 45, 2011–2019. [Google Scholar] [CrossRef] [PubMed]
- Nelson, K.Y.; McMartin, D.W.; Yost, C.K.; Runtz, K.J.; Ono, T. Point-of-use water disinfection using UV light-emitting diodes to reduce bacterial contamination. Environ. Sci. Pollut. Res. 2013, 20, 5441–5448. [Google Scholar] [CrossRef] [PubMed]
- Oguma, K.; Kita, R.; Sakai, H.; Murakami, M.; Takizawa, S. Application of UV light emitting diodes to batch and flow-through water disinfection system. Desalination 2013, 328, 24–30. [Google Scholar] [CrossRef]
- Oguma, K.; Rattanakul, S.; Bolton, J.R. Application of UV light-emitting diodes to adenovirus in water. J. Environ. Eng. 2016, 142, 1215–1218. [Google Scholar] [CrossRef]
- Lui, G.Y.; Roser, D.; Corkish, R.; Ashbolt, N.J.; Stuetz, R. Point-of-use water disinfection using ultraviolet and visible light-emitting diodes. Sci. Total Environ. 2016, 553, 626–635. [Google Scholar] [CrossRef] [PubMed]
- Sholtes, K.A.; Lowe, K.; Walters, G.W.; Sobsey, M.D.; Linden, K.G.; Casanova, L.M. Comparison of ultraviolet light-emitting diodes and low-pressure mercury-arc lamps for disinfection of water. Environ. Technol. 2016, 37, 2183–2188. [Google Scholar] [CrossRef] [PubMed]
- Aoyagi, Y.; Takeuchi, M.; Yoshida, K.; Kurouchi, M.; Yasui, N.; Kamiko, N.; Araki, T.; Nanishi, Y. Inactivation of bacterial viruses in water using deep ultraviolet semiconductor light-emitting diode. J. Environ. Eng. 2011, 137, 1215–1218. [Google Scholar] [CrossRef]
- Zyara, A.M.; Torvinen, E.; Veijalainen, A.-M.; Heinonen-Tanski, H. The effect of UV and combined Chlorine/UV treatment on coliphages in drinking water disinfection. Water 2016, 8, 130. [Google Scholar] [CrossRef]
- Hsu, F.-C.; Shieh, Y.S.; Duin, J.V.; Beekwilder, M.J.; Sobsey, M.D. Genotyping male-specific RNA coliphages by hybridization with oligonucleotide probes. Appl. Environ. Microbiol. 1995, 61, 3960–3966. [Google Scholar] [PubMed]
- Bolton, J.R.; Linden, K.G. Standardization of methods for fluence (UV Dose) determination in Bench-Scale UV experiments. J. Environ. Eng. 2003. [Google Scholar] [CrossRef]
- Kuopion Vesi. Drinking Water Quality in Kuopio. Available online: http//www.kuopionvesi.fi/c/document_library/get_file?uuid=e6c27e13-472e-4496-8f99-9fb7933e89b4&groupId=518539 (accessed on 27 January 2016).
- American Public Health Association (APHA); American Water Works Association (AWWA); Water Environment Federation (WEF). Standard Methods for the Examination of Water and Wastewater, 21st ed.; APHA, AWWA and WEF: Washington, DC, USA, 2005. [Google Scholar]
- Adams, M.H. Bacteriophages; Interscience: New York, NY, USA, 1959. [Google Scholar]
- Rajala-Mustonen, R.L.; Heinonen-Tanski, H. Sensitivity of host strains and host range of coliphages isolated from Finnish and Nicaraguan wastewater. Water Res. 1994, 28, 1811–1815. [Google Scholar] [CrossRef]
- Zyara, A.M.; Torvinen, E.; Veijalainen, A.-M.; Heinonen-Tanski, H. The effect of chlorine and combined chlorine/UV treatment on coliphages in drinking water disinfection. J. Water Health 2016, 14, 640–649. [Google Scholar] [CrossRef] [PubMed]
- Beck, S.E.; Wright, H.B.; Hargy, T.M.; Larason, T.C.; Linden, K.G. Action spectra for validation of pathogen disinfection in medium-pressure ultraviolet (UV) systems. Water Res. 2015, 70, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Beck, S.E.; Rodriguez, R.A.; Hawkins, M.A.; Hargy, T.M.; Larason, T.C.; Linden, K.G. Comparison of UV-Induced Inactivation and RNA Damage in MS2 Phage across the Germicidal UV Spectrum. Appl. Environ. Microbiol. 2016, 82, 1468–1474. [Google Scholar] [CrossRef] [PubMed]
- Oguma, K.; Kita, R.; Takizawa, S. Effects of arrangement of UV light-emitting diodes on the inactivation efficiency of microorganisms in water. J. Photochem. Photobiol. 2016, 92, 314–317. [Google Scholar] [CrossRef] [PubMed]
- U.S. Environmental Protection Agency (EPA). Ultraviolet Disinfection Guidance Manual for the Final Long Term 2 Enhanced Surface Water Treatment Rule; EPA Office of Water: Washington, DC, USA, 2006.
- Kalisvaart, B.F. Re-use of wastewater: Preventing the recovery of pathogens by using medium-pressure UV lamp technology. Water Sci. Technol. 2004, 50, 337–344. [Google Scholar] [PubMed]
- Oppezzo, O.J.; Pizarro, R.A. Sublethal effects of ultraviolet A radiation on Enterobacter cloacae. J. Photochem. Photobiol. B 2001, 62, 158–165. [Google Scholar] [CrossRef]
- Hamamoto, A.; Mori, M.; Takahashi, A.; Nakano, M.; Wakikawa, N.; Akutagawa, M.; Ikehara, T.; Nakaya, Y.; Kinouchi, Y. New water disinfection system using UVA light-emitting diodes. J. Appl. Microbiol. 2007, 103, 2291–2298. [Google Scholar] [CrossRef] [PubMed]
- Chevremont, A.C.; Farnet, A.M.; Coulomb, B.; Boudenne, J.L. Effect of coupled UV-A and UV-C LEDs on both microbiological and chemical pollution of urban wastewater. Sci. Total Environ. 2012, 426, 304–310. [Google Scholar] [CrossRef] [PubMed]
- Chevremont, A.C.; Farnet, A.M.; Sergent, M.; Coulomb, B.; Boudenne, J.L. Multivariate optimization of fecal bioindicator inactivation by coupling UV-A and UV-C LEDs. Desalination 2012, 285, 219–225. [Google Scholar] [CrossRef]
- Nakahashi, M.; Mawatari, K.; Hirata, A.; Maetani, M.; Shimohata, T.; Uebanso, T.; Hamada, Y.; Akutagawa, M.; Kinouchi, Y.; Takahashi, A. Simultaneous irradiation with different wavelengths of ultraviolet light has synergistic bactericidal effect on Vibrio Parahaemolyticus. Photochem. Photobiol. 2014, 90, 1397–1403. [Google Scholar] [CrossRef] [PubMed]
- Rattanakul, S.; Oguma, K.; Sakai, H.; Takizawa, S. Inactivation of viruses by combination processes of UV and chlorine. J. Water Environ. Technol. 2014, 12, 511–523. [Google Scholar] [CrossRef]
- Rattanakul, S.; Oguma, K.; Sakai, H.; Takizawa, S. Sequential and simultaneous applications of UV and chlorine for adenovirus inactivation. Food Environ. Virol. 2015, 7, 295–304. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).