1. Introduction
The discharge of nitrogen components in the environment is hazardous for both human and animal health and the nitrate pollution and remediation is at the same time a worldwide problem and a challenge. Nitrates cause the most serious problems when dispersed in water, since they cause the depletion of aquifers and the eutrophication of rivers. Sources of nitrate comprise natural cycle and human activities, mainly from uncontrolled land discharges of treated or raw domestic and industrial wastewaters, landfills, and animal wastes predominantly from animal farms [
1,
2]. Therefore, several studies focused on the nitrates removal from wastewater in order to achieve an acceptable concentration in treated waters to be discharged into the environment. Complementary techniques are often required for obtaining a free nitrate stream in treatment plants, being the most common methods either physicochemical or biological and sometimes a combination of both [
3,
4,
5]. The biological denitrification is recommended for the removal of relatively low concentration of nitrogen components and it is operated by the so called denitrifying bacteria in anoxic conditions, where they use nitrates as electron acceptors during their respiratory process in the place of the oxygen. Nitrates are efficiently removed when an external organic carbon source, generally methanol, ethanol or acetic acid, is added [
4,
6,
7,
8]. As demonstrated by the most recent literature, new bacterial strains are continuously isolated and tested for their NO
3− removal abilities [
9,
10,
11,
12,
13]. Depending on their characteristics, different bacteria are employed in different waste treatment facilities, with a preference towards those microorganisms capable of combined heterotrophic nitrification and aerobic denitrification. However, other characteristics are often desirable, for example bacteria with a marked resistance to high salinity, are generally employed in the treatment of polluted seawater [
14] and strains isolated from critically polluted environments are used for the treatment of special industrial wastes, such as tannery wastewater [
15].
In a previous work [
16] we studied an enrichment sample of a bacterial colony coming from a contaminated site and capable of reducing perchlorates and nitrates. The colony was isolated from the water of Sarno River, located in the south part of Italy and known to have high levels of both organic and inorganic pollutants, mainly due to the presence of tomato industries and leather tanneries [
17]. The high resistance of the bacterial colony to extreme conditions and its ability to reduce both perchlorate and nitrates simultaneously, motivated a deeper study in order to employ bacteria in wastewater remediation. From the original colony a beta proteobacterium was isolated, which in batch experiments in anoxic conditions was able to efficiently use nitrates as electron acceptors and acetates as electron donors and carbons source. This strain was identified as
Azospira spp.,
Azospira sp. OGA 24, because from molecular analysis based on 16S rRNA sequence it showed 99.7%–100% identity with this genus (NCBI accession number, GU294119.1).
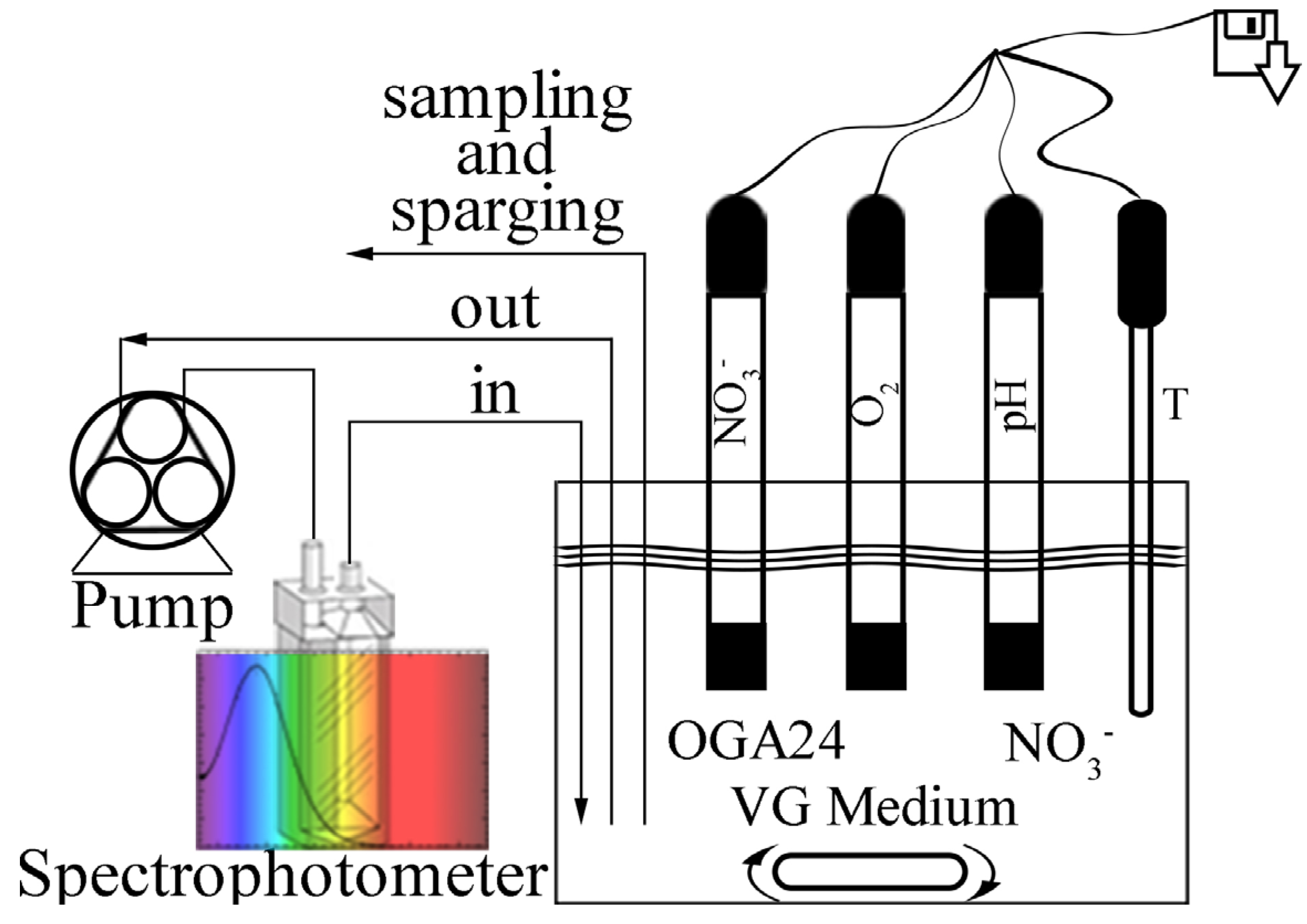
With this work we start a thorough characterization of the denitrifying properties of the Azospira sp. OGA 24. By using a specifically devised experimental setup, a sterile and anoxic environment was obtained where both the nitrates consumption and the bacterial growth could be easily monitored in real time. In our experimental conditions, the denitrification kinetics was found to be compatible with a zero- and a first-order kinetics, both reliable when an excess of nitrates and of organic substrate are present in the reactor. Moreover, an assessment of the denitrification potential of OGA 24 when used in real wastewaters treatment plants was done. A simple theoretical model was used to simulate the behaviour of OGA 24 during the anoxic stage of a denitrification process and the results were found to meet with the Italian legislation requirements.
3. Results and Discussion
Aerobically grown up to an optical density of 0.02 A.U. at 600 nm (roughly equal to 2 × 10
7 individuals/mL),
Azospira sp. OGA 24 was then inoculated in the batch bioreactor in the presence of the VG medium, acetate as the organic substrate (electron donor) and [NO
3−] = 350 mg/L (electron acceptor). The bioreactor was previously sparged with argon in order to obtain anaerobic conditions, so that the bacteria could use nitrates as the only final electron acceptors for their respiratory functions. Thanks to the
ad hoc experimental setup and to the analytical methods employed, the kinetics of the nitrates consumption and of the bacterial population growth could be recorded simultaneously and with a continuous sampling rate.
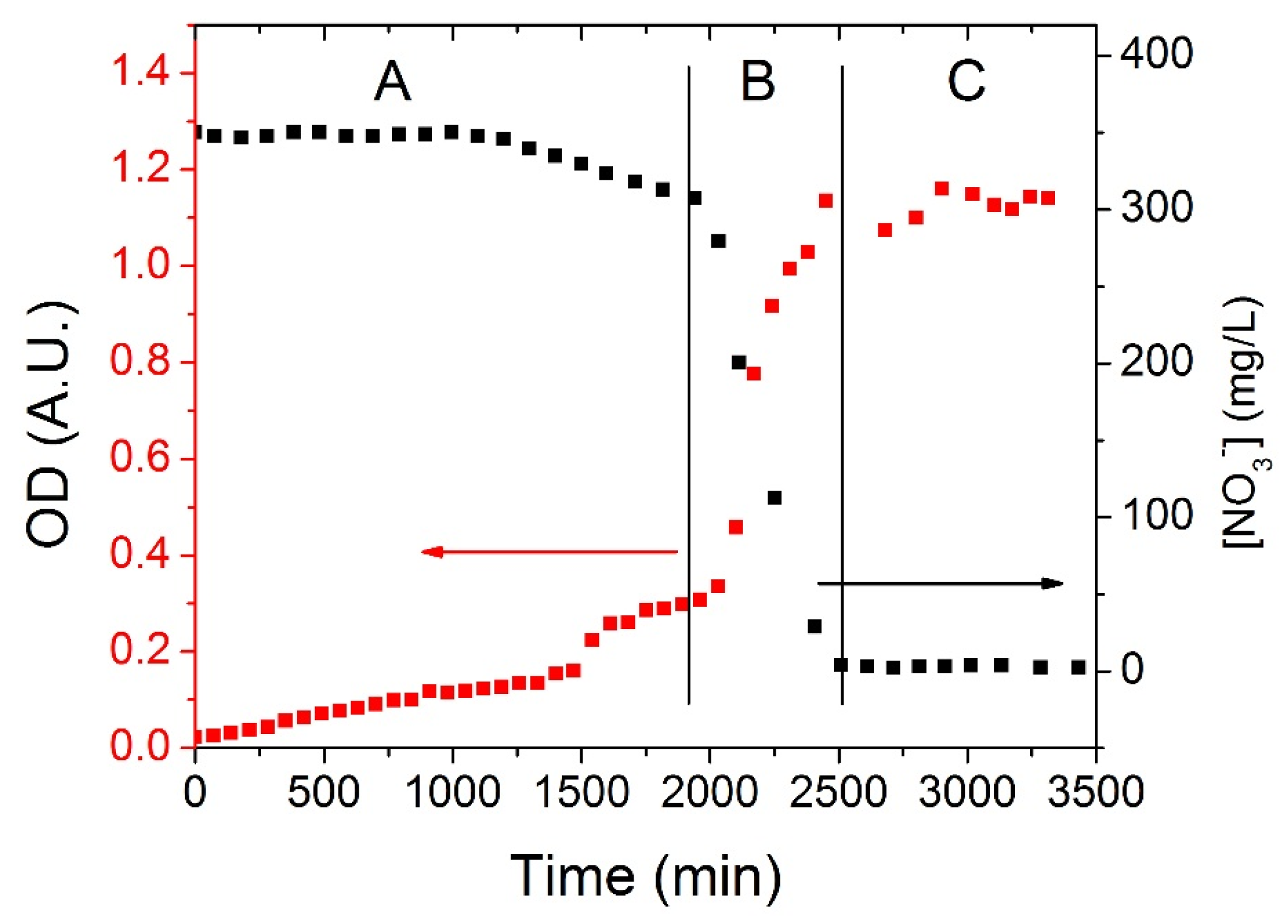
Figure 3 shows a typical time series of the nitrate consumption operated by OGA 24 (black trace, right axis) and the corresponding growth curve of the bacteria (red trace, left axis). The whole dynamics could be divided into three main regions for both the curves. The first region represents a rather long (0–~1800 min) acclimation period (A in the figure) where bacteria adapt to the anaerobic condition and to the high concentration of nitrates and their population slightly increases (OD 0.02→~0.3 A.U.); correspondingly the concentration of nitrates slowly decreases from 350 mg/L to ~320 mg/L. The quite long acclimation period could certainly be reduced by starting experiments with a larger population of bacteria,
i.e., by increasing the duration of the cells culture. The acclimation period is followed by the exponential region (B in the figure, ~1800–~2500 min) where nitrates are rapidly depleted until their concentration reaches a value of about 0 mg/L. The rapid consumption of the electron acceptor species corresponds to a fast increase of the bacteria population (OD ~0.3→~1.2 A.U.). Finally, a plateau region (C in the figure) indicates that the system reached a steady state where nitrates are completely consumed and the number of bacteria is stationary in time (~1.2 × 10
9 individuals/mL).
Figure 3.
Experimental curves of nitrates reduction operated by the azospira OGA 24. Black squares represent nitrates concentration over time, measured by means of the NO3− ISE; red squares show the increase of the solution optical density over time, i.e., the growth of the bacteria population. Three different regions can be identified: A: acclimation zone; B: exponential consumption of nitrates (increase of the bacterial population); C: steady state.
Figure 3.
Experimental curves of nitrates reduction operated by the azospira OGA 24. Black squares represent nitrates concentration over time, measured by means of the NO3− ISE; red squares show the increase of the solution optical density over time, i.e., the growth of the bacteria population. Three different regions can be identified: A: acclimation zone; B: exponential consumption of nitrates (increase of the bacterial population); C: steady state.
In order to understand the potentiality of the
Azospira sp. OGA 24 as denitrifying bacterium in wastewater treatment plants, the kinetics of the exponential region B was characterized. The bio-denitrification process is generally described by a Monod or Michaelis-Menten type kinetics [
18,
19], however, when the initial concentration of nitrates is high (generally above 200 mg/L) and the organic substrate is in large excess, the nitrates consumption dynamics can be approximated to a linear zero- or exponential first-order kinetics [
18,
20,
21,
22,
23].
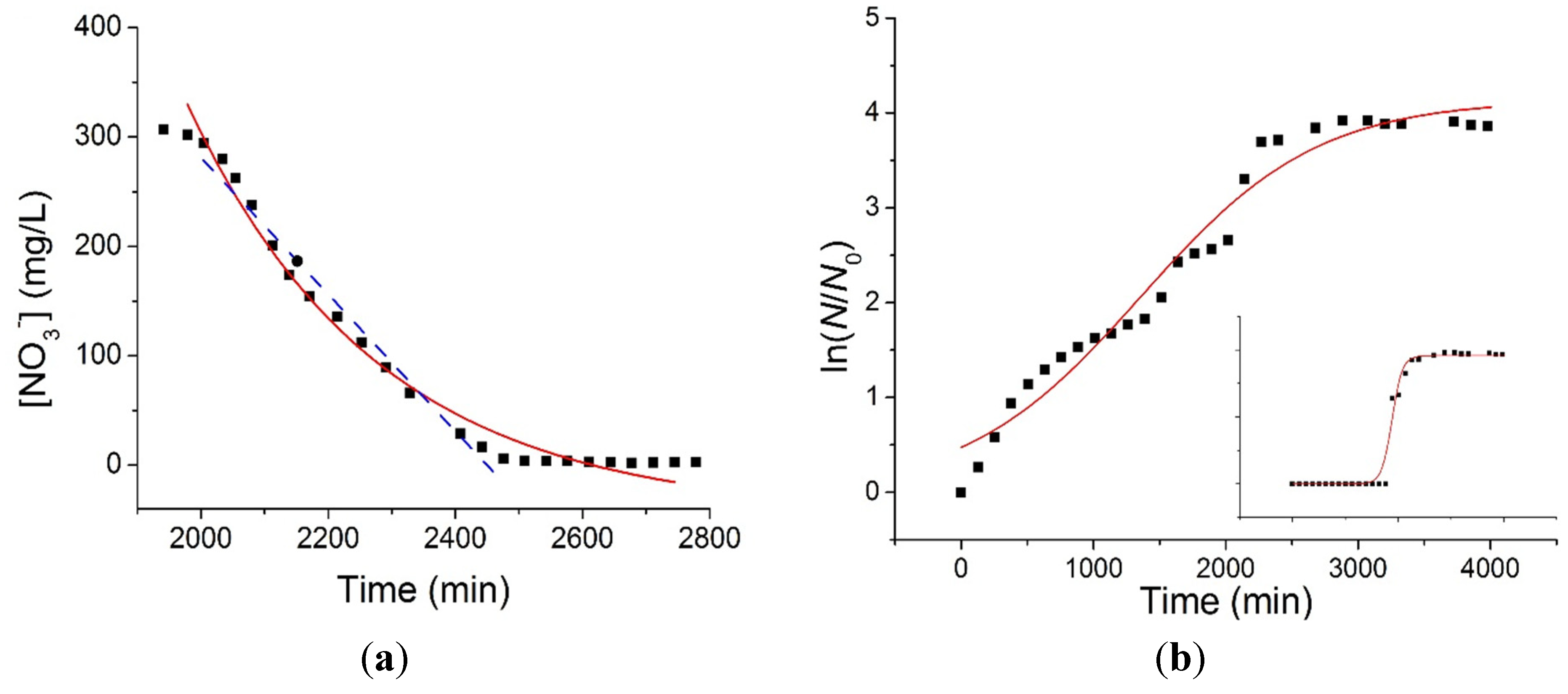
Figure 4a shows the detail of the exponential region B relative to the [NO
3−] time series of
Figure 3, where the rate of consumption of the electron acceptor is maximum. In order to characterize the kinetics of the denitrification process, data from different experiments where fitted both to a linear equation (blue dashed line, fitting interval 2000–2400 min) having the form
where
(mg L
−1·min
−1) is the kinetic constant of the zero-order kinetics, and to an exponential equation (red line, fitting interval 2000–2800 min) having the form
where
(min
−1) is the kinetic constant of the first-order kinetics. Data where fitted by means of a Levenberg–Marquardt [
24] nonlinear regression algorithm, which yielded a value of
= 0.63 ± 0.03 mg L
−1 min
−1 and
= 4 ± 0.5 × 10
−3 min
−1. For both the fitting procedures the value of the R-squared index was higher than 0.99, meaning that both the mechanisms proposed for the denitrification process were plausible.
Figure 4.
(a) Exponential fitting (red line, Equation (3)) and linear fitting (blue dashed line, Equation (2)) of the [NO3−] consumption during the exponential period; (b) Fitting of the bacterial growth curve by means of the logistic Equation (4). The inset shows the bacterial growth curve when the bacterial population is considered constant during the acclimation period.
Figure 4.
(a) Exponential fitting (red line, Equation (3)) and linear fitting (blue dashed line, Equation (2)) of the [NO3−] consumption during the exponential period; (b) Fitting of the bacterial growth curve by means of the logistic Equation (4). The inset shows the bacterial growth curve when the bacterial population is considered constant during the acclimation period.
The growth dynamics of the bacterial population was also characterized by fitting OD data (
Figure 4b) to a logistic equation having the form
where
N/N0 is the relative bacterial population size,
A = ln(
N∞/N0) is the maximum value reached, λ is the lag time,
i.e., the time of the acclimation period (region A in
Figure 3) and μ
m is the maximum specific growth rate [
25]. The slight increase of the population size during the acclimation period is amplified in the logarithmic scale and it makes difficult a good fitting to the Equation (4), as showed in the main graphic of
Figure 4b. The values of the fitting parameters were found to be
A = 4.14 ± 0.04 (which roughly corresponds to a population increase of 60 times,
i.e., 10 times larger than the value obtained from OD measurements), μ
m = 1.6 ± 0.04 × 10
−3 min
−1 and an unrealistic lag time λ = 28 ± 10 min, with R-squared = 0.95. In order to obtain a better fitting and a more realistic value for the lag time, the population size was considered as constant during the acclimation period (inset in
Figure 4). In this case, the fitting procedure yielded
A = 3.83 ± 0.04 (which roughly corresponds to a population increase of 50 times in accordance with the OD measurements), μ
m = 9.9 ± 0.4 × 10
−3 min
−1 and λ = 1689 ± 12 min, with R-squared = 0.99. The specific growth rate differs by almost one order of magnitude in the two cases and it is difficult at this stage to obtain a proper characterization of the growth dynamics. However, the value of
was found to be in between these two extremes, therefore the value of μ
m can be reliable in both cases.
In order to assess the performances of the
Azospira sp. OGA 24 for its employment in a real wastewater treatment plant, the kinetic data obtained in the batch bioreactor were used in a simple theoretical model of the denitrification process. For the sake of simplicity only the anoxic stage, which is generally the core of the biological wastewater treatments [
6,
8,
26], was considered. In this stage the bacteria are continuously supplied with an organic substrate and they reduce the nitrates to nitrogen in the absence of oxygen. The process can be modeled by considering a CSTR (Continuously Stirred Tank Reactor) with an excess of organic substrate and with a flux of incoming wastewater,
Fin, equals to the outgoing denitrified water,
Fout,
i.e., the volume of the reactor,
V(
t), is in stationary state:
Fin =
Fout =
F (m
3/d),
V(
t) =
V0 (m
3). Moreover, the bacterial population is constant and in the exponential growth stage.
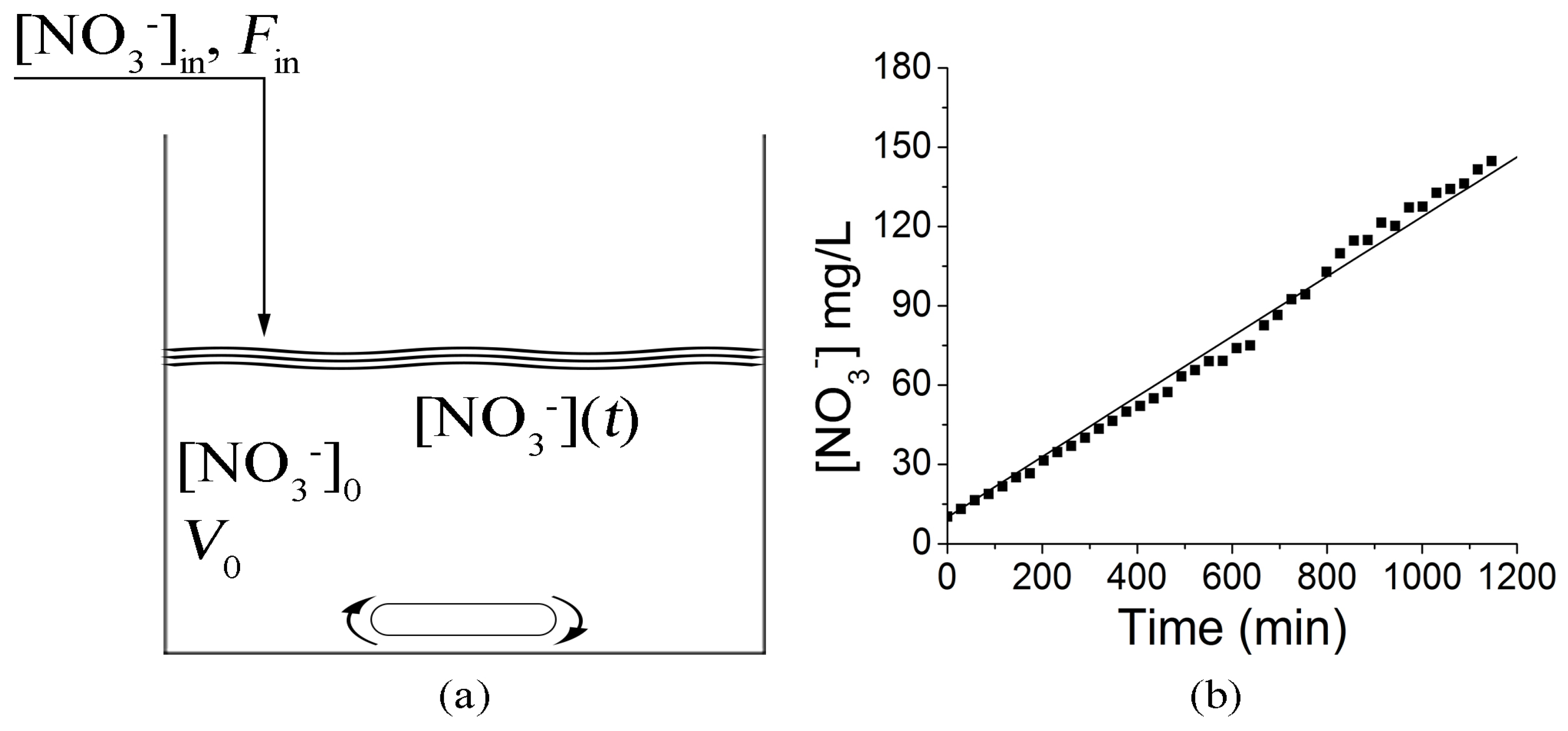
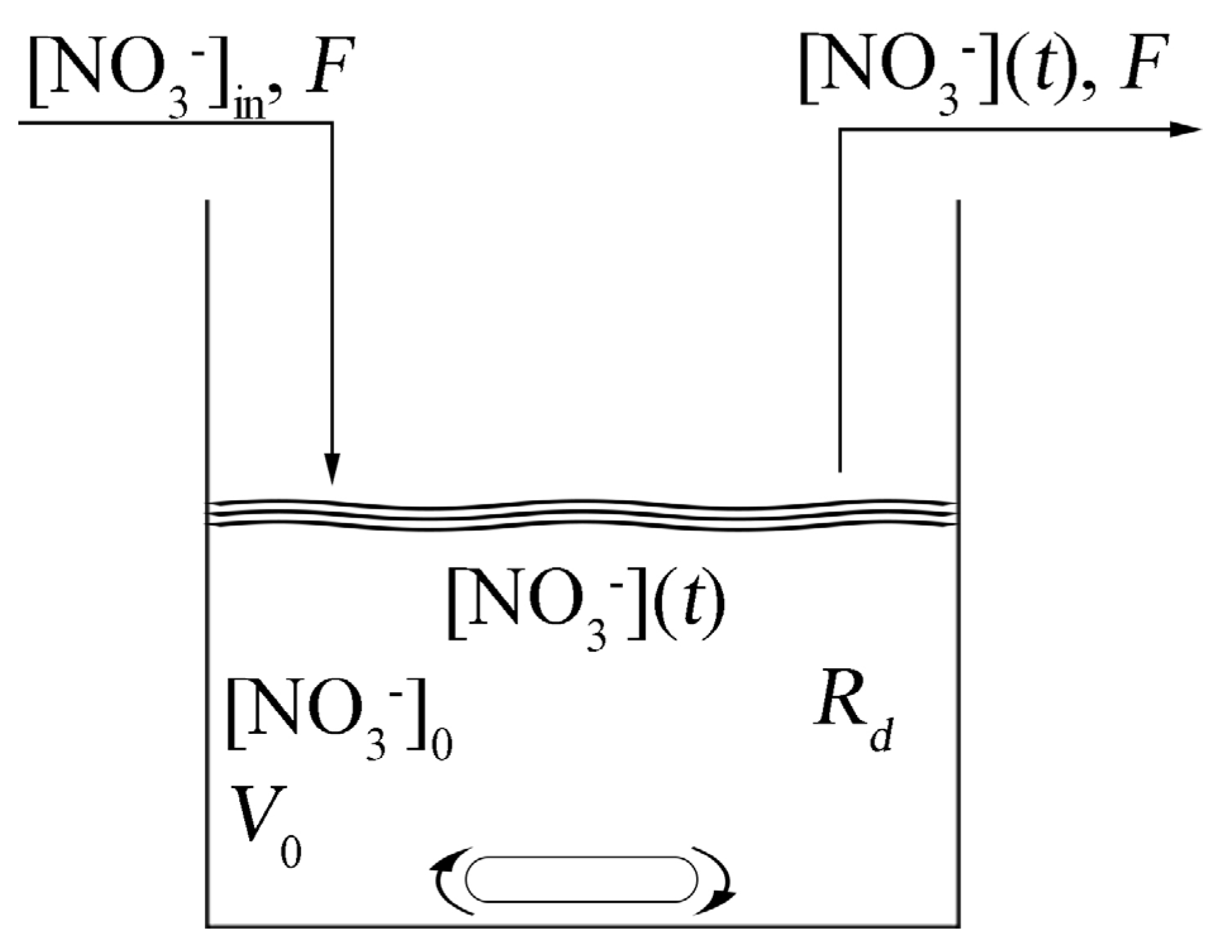
Figure 5 shows a scheme of the CSTR where [NO
3−]
in is the concentration of nitrates in the wastewater, [NO
3−]
0 is the initial concentration of nitrates at time
t = 0,
Rd represents the kinetics of the denitrification reactions and [NO
3−](
t) is the concentration of nitrates in time, which corresponds to the concentration of nitrates in the outgoing wastewater. When we consider the zero-order kinetics for the nitrates consumption, the system is described by a differential equation having the form
where [NO
3−] ≡ [NO
3−](
t). Being both
F and
V0 constant,
k0 =
F/
V0 (d
−1) can be defined as the reciprocal of the mean residence time of a molecule in the reactor and the integral form of the Equation (5) yields the variation in time of nitrates
The stationary state value of nitrates [NO3−]ss, i.e., the concentration in the outgoing wastewater when the system is in continuous working regime, can be found by Equating (5) to zero or by calculating the limit of Equation (6) for t → ∞ and obtain
We then substituted in the equation the value for
found in the experiments (~ 907 mg·L
−1·d
−1) and the values for
k0 and [NO
3−]
in of a real wastewater treatment plant with a potential of < 100,000 A.E. (
F = 210 m
3/d,
V0 = 93 m
3,
k0 = 2.25 d
−1) and [NO
3−]
in = 450 mg/L to find that [NO
3−]
ss ~47 mg/L which corresponds to a reduction of nitrate of about 90%. This value meets the requests of Italian legislation that requires a minimum reduction of nitrate of 70% for this kind of treatment plant. The process was also modeled considering a first-order kinetics and using
in the place of
, in this case
and the efficiency of the process was found to be about 70%. A comparison with the literature confirms that the performance of the
Azospira sp. OGA 24 are in line (sometimes better) of the most common bacteria employed in the biological treatment of polluted waters [
7,
12,
23,
27,
28,
29].
Figure 5.
Scheme of the anoxic denitrification process in a Continuously Stirred Tank Reactor (CSTR).
Figure 5.
Scheme of the anoxic denitrification process in a Continuously Stirred Tank Reactor (CSTR).
4. Conclusions
With this work we started a thorough investigation of the denitrifying properties of the bacterium Azospira sp. OGA 24, recently isolated from the highly polluted waters of the Sarno river in the south of Italy. Thanks to the specifically devised bioreactor, it was possible to follow in real time both the concentration of nitrates, through an ion-selective electrode, and the optical density of the solution (a measure of the bacteria population size) through an in-line flow-thru spectrophotometer. The fitting of the experimental data allowed us to extract important parameters, which characterize the denitrification ability (kinetics constants) and the population growth (maximum specific growth rate) of the bacteria in anoxic conditions. In particular, it was found that in the presence of an excess of the organic substrate and at a relatively high [NO3−], the stage with the maximum rate of nitrates consumption can be described by both a zero- and a first-order kinetics. The main motivation of the present work was to assess the effectiveness of OGA 24 as denitrifying agent; therefore a better characterization of the kinetic mechanism will be done in future works.
Kinetic constants were finally used in a simplified model of the anoxic stage of a real wastewater biological treatment plant, in order to assess the denitrifying capability of OGA 24 in the presence of a continuous influx of wastewater with a high concentration of nitrates. In particular, we looked for the stationary state solutions of the model where the characteristic parameters were taken from a real treatment plant and from our experiments. Despite many simplifications we introduced, our simulations could give a clear idea of the effectiveness of OGA 24 in depleting the nitrates concentration in polluted wastewater and the stationary state solutions were found to match the Italian law requirements, both when a zero- and a first-order denitrification kinetics were considered. Although many bacteria or consortium are known and used for the denitrification process, our results might provide an alternative microbial resource for biological removal of nitrogen compounds from wastewater. In particular, the extreme conditions where the bacteria colony was isolated favor the employment of the strain in those environments that generally inhibit biological treatments, such as tannery wastewater. Moreover, the OGA 24 effectiveness in reducing both nitrates and perchlorates simultaneously, make this strain very versatile and useful in several environmental conditions. Further investigations are still in progress to study the influence of bacterial concentration, temperature and pH variations on the denitrification efficiency of OGA 24.