Cyanide Bioremediation by Bacillus subtilis under Alkaline Conditions
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Bacterial Isolation
2.3. Sequencing and Molecular Identification
2.4. Bacterial Growth Kinetics
2.5. Cyanide Biodegradation
2.6. Kinetic Biodegradation Models
2.7. Analytical Methods
3. Results
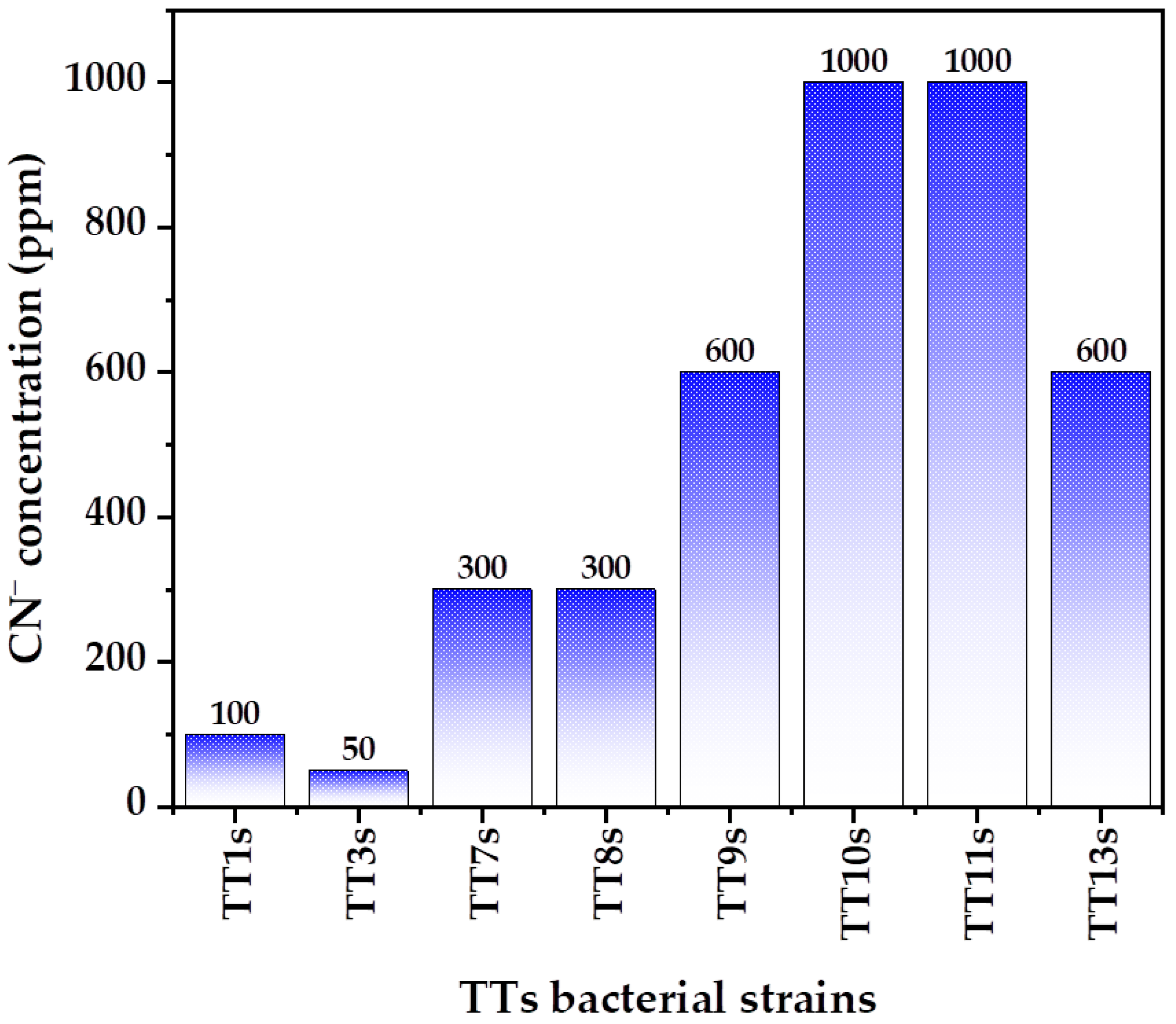
3.1. Bacterial Isolation and Selection
3.2. Molecular Identification
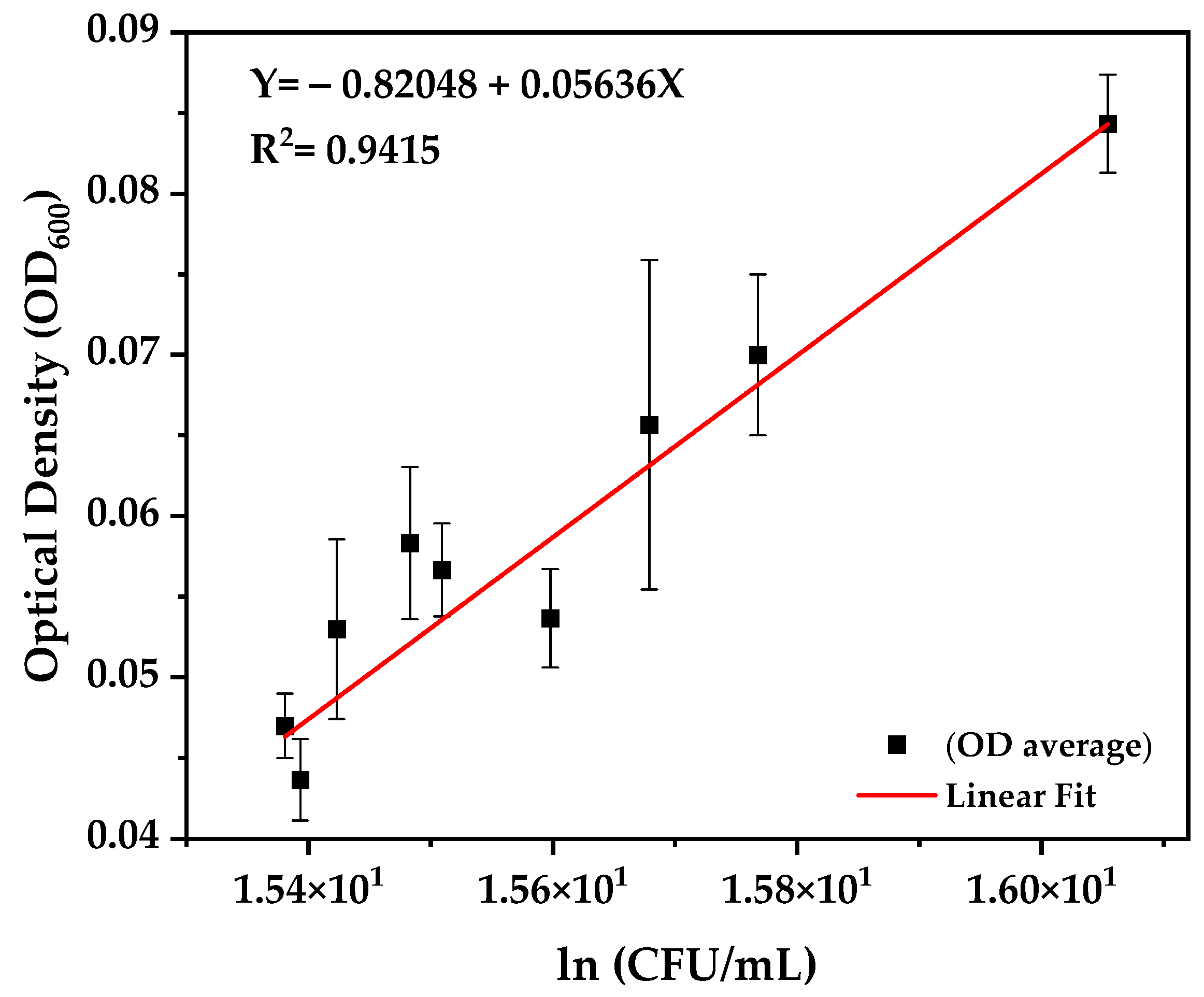
3.3. Growth Kinetics
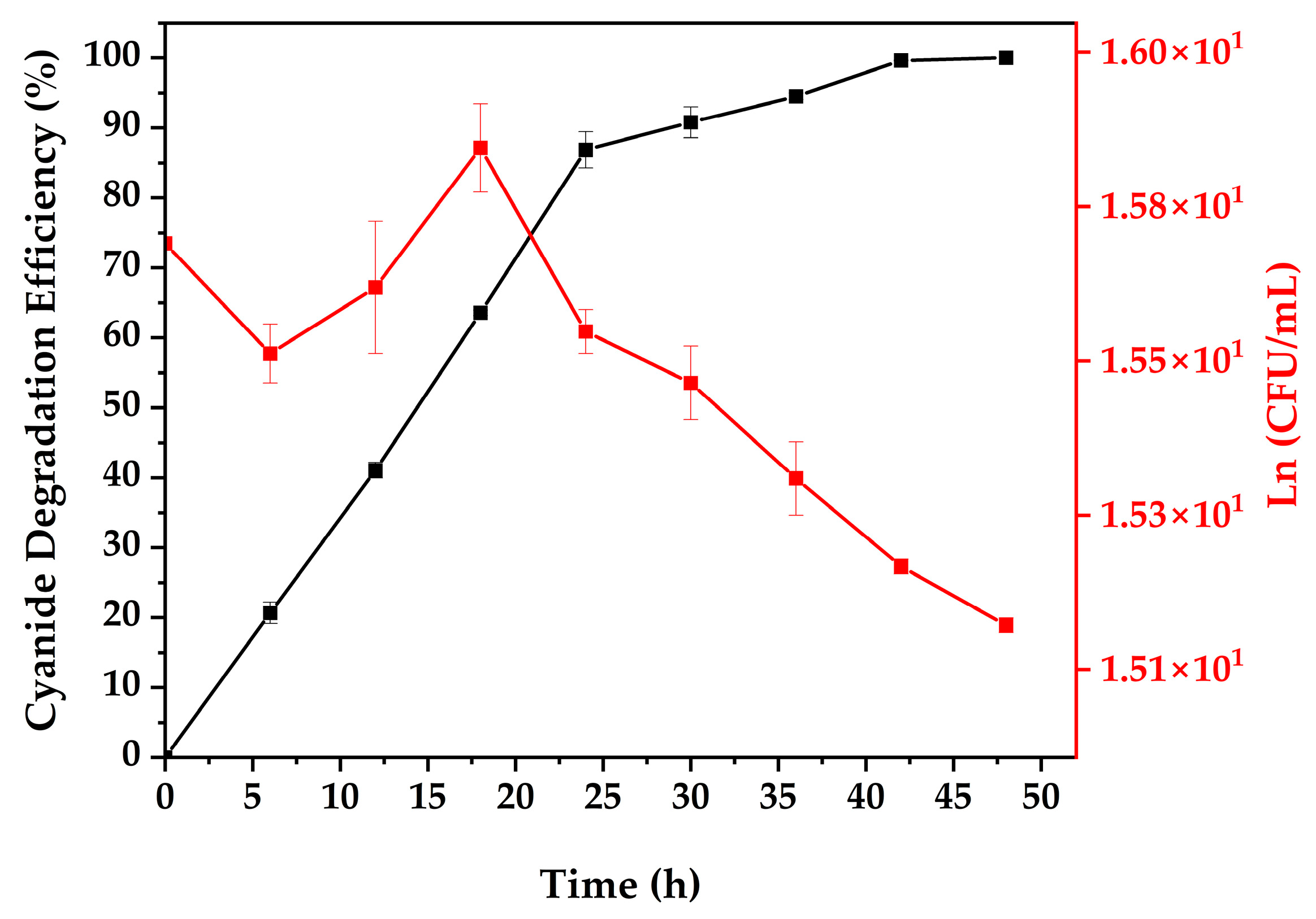
3.4. Cyanide Biodegradation
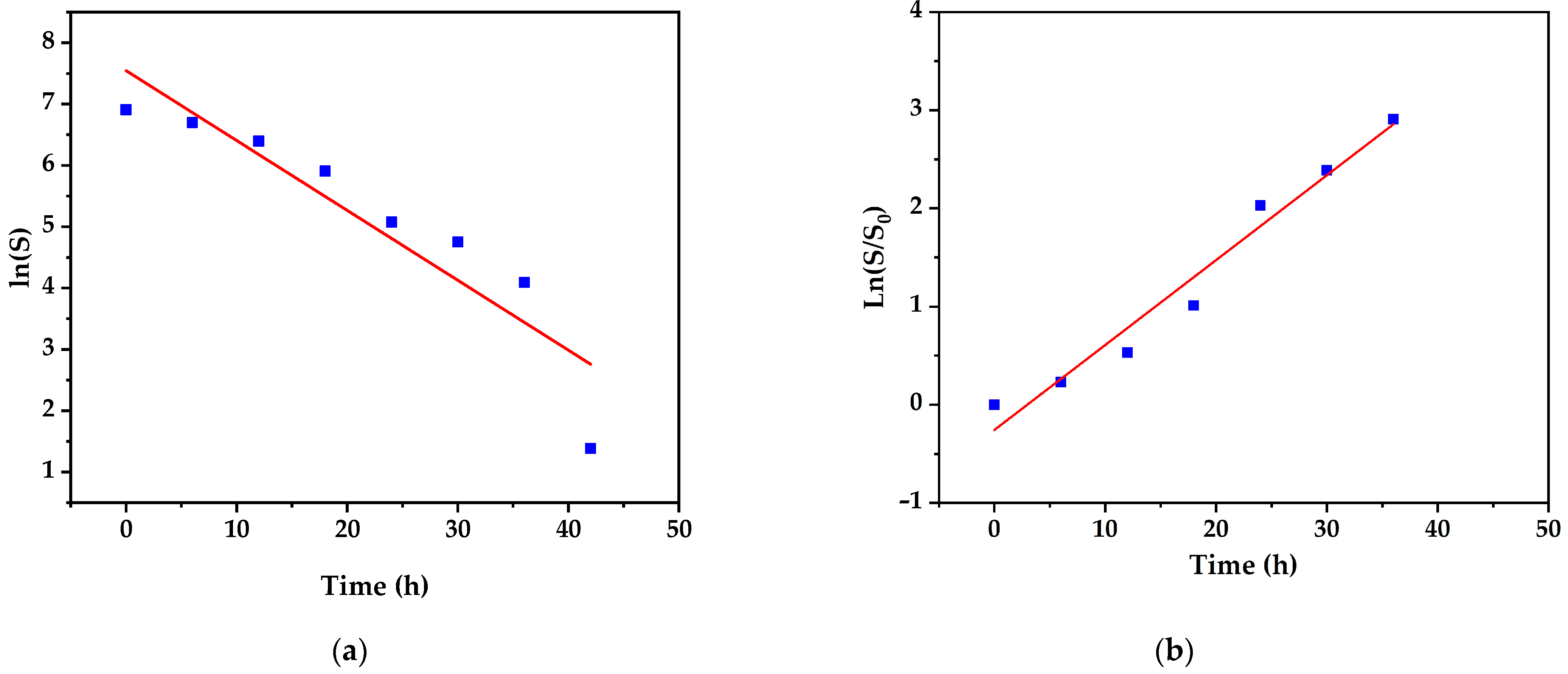
3.5. Kinetic Models
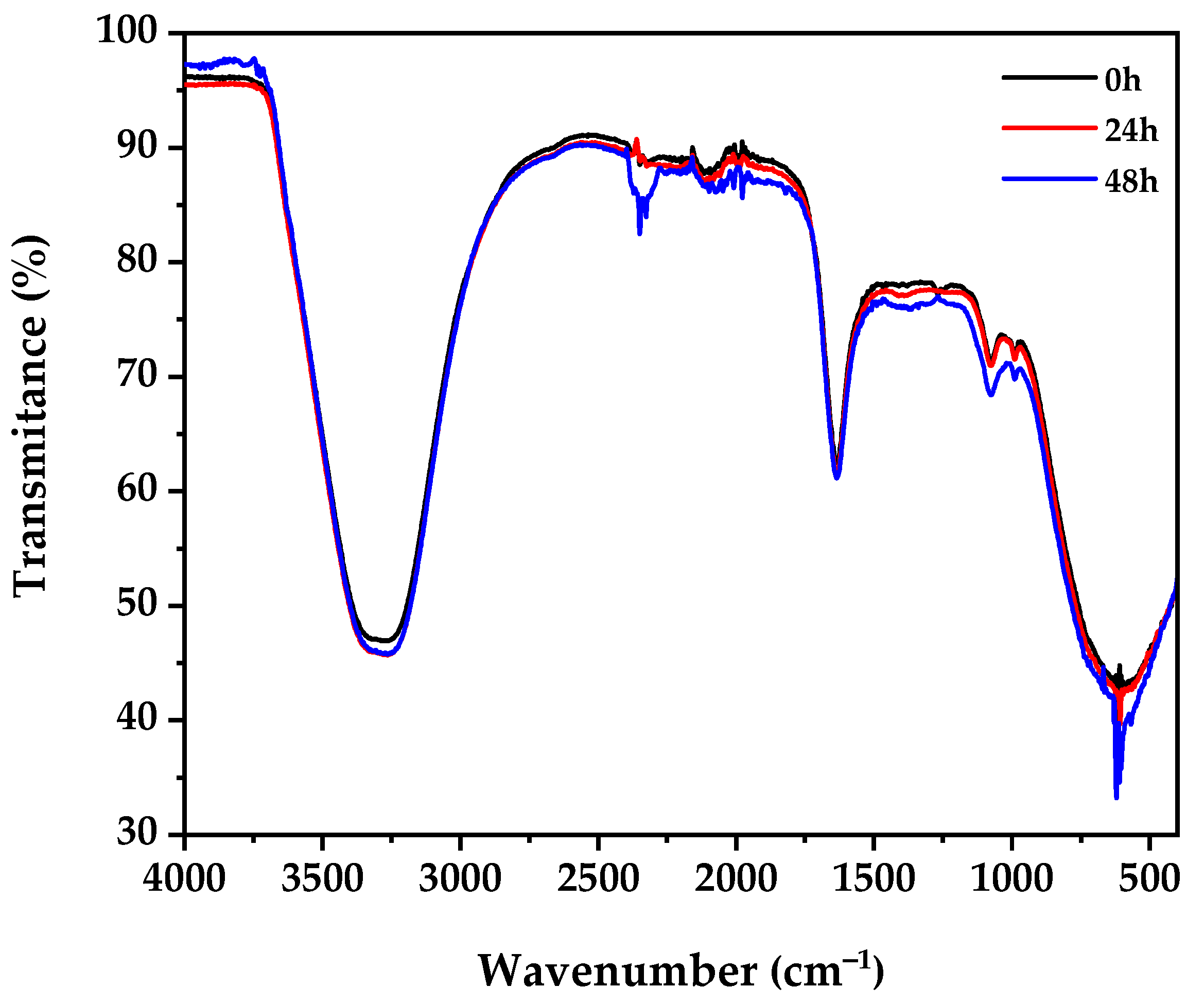
3.6. FTIR Spectroscopy Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wang, Z.-S.; Cho, Y.-C.; Lin, Y.-P. Removal of Cyanide from Electroplating Wastewater by Persulfate Oxidation Process. 2023. Available online: https://ssrn.com/abstract=4427783 (accessed on 28 September 2023).
- Eisler, R.; Wiemeyer, S.N. Cyanide hazards to plants and animals from gold mining and related water issues. Rev. Environ. Contam. Toxicol. 2004, 183, 21–54. [Google Scholar] [CrossRef] [PubMed]
- Martínková, L.; Bojarová, P.; Sedova, A.; Křen, V. Recent trends in the treatment of cyanide-containing effluents: Comparison of different approaches. Crit. Rev. Environ. Sci. Technol. 2023, 53, 416–434. [Google Scholar] [CrossRef]
- Cacciuttolo, C.; Cano, D. Environmental Impact Assessment of Mine Tailings Spill Considering Metallurgical Processes of Gold and Copper Mining: Case Studies in the Andean Countries of Chile and Peru. Water 2022, 14, 3057. [Google Scholar] [CrossRef]
- Matlock, M.M.; Howerton, B.S.; van Aelstyn, M.A.; Nordstrom, F.L.; Atwood, D.A. Advanced mercury removal from gold leachate solutions prior to gold and silver extraction: A field study from an active gold mine in Peru. Environ. Sci. Technol. 2002, 36, 1636–1639. [Google Scholar] [CrossRef]
- Isla, A. The Guardians of Conga Lagoons: Defending Land Water and Freedom in Peru. Can. Woman Stud./Les Cah. De La Femme 2015, 30, 25–40. [Google Scholar]
- Presidencia del Concejo de Ministros de Perú. Decreto Supremo que Declara el Estado de Emergencia en los Distritos de Cocachacra, Dean Valdivia y Punta de Bombón de la Provincia de Islay, del Departamento de Arequipa, por Peligro Inminente Ante Contaminación Hídrica; Perú, 2021 (N° 106-2021-PCM). Available online: https://busquedas.elperuano.pe/dispositivo/NL/1957547-2 (accessed on 26 September 2023).
- Bebbington, A.; Williams, M. Water and Mining Conflicts in Peru. Mt. Res. Dev. 2008, 28, 190–195. [Google Scholar] [CrossRef]
- Salem, J.; Amonkar, Y.; Maennling, N.; Lall, U.; Bonnafous, L.; Thakkar, K. An analysis of Peru: Is water driving mining conflicts? Resour. Policy 2021, 74, 101270. [Google Scholar] [CrossRef]
- Ministerio del Ambiente. Modifican los Estándares Nacionales de Calidad Ambiental Para Agua y Establecen Disposiciones Complementarias Para su Aplicación; Diario el Peruano, 2015 (N° 015-2015-MINAM). Available online: https://www.minam.gob.pe/wp-content/uploads/2015/12/Decreto-Supremo-N%C2%B0-015-2015-MINAM.pdf (accessed on 28 September 2023).
- Marshall, B.G.; Veiga, M.M.; Da Silva, H.A.M.; Guimarães, J.R.D. Cyanide Contamination of the Puyango-Tumbes River Caused by Artisanal Gold Mining in Portovelo-Zaruma, Ecuador. Curr. Environ. Health Rep. 2020, 7, 303–310. [Google Scholar] [CrossRef]
- Copari Mamani, A.B.; Carpio Mamani, M.; Cáceda Quiroz, C.J. Optimización de factores fisicoquímicos en la biodegradación de cianuro por Klebsiella sp.ART1, en biorreactor aireado. Cienc. Desarro. 2020, 26, 20–31. [Google Scholar] [CrossRef]
- Quiroz, C.J.C.; Choque, G.J.M.; Mamani, M.C.; Quispe, G.D.L.F. Evaluation of the content of metals and contamination indices generated by environmental liabilities, in Tacna, Peru. Res. Sq. 2022. [Google Scholar] [CrossRef]
- Swenson, J.J.; Carter, C.E.; Domec, J.-C.; Delgado, C.I. Gold mining in the Peruvian Amazon: Global prices, deforestation, and mercury imports. PLoS ONE 2011, 6, e18875. [Google Scholar] [CrossRef]
- Veiga, M.M.; Angeloci, G.; Hitch, M.; Colon Velasquez-Lopez, P. Processing centres in artisanal gold mining. J. Clean. Prod. 2014, 64, 535–544. [Google Scholar] [CrossRef]
- Mayorca Clemente, S. Reducción de Cianuro Del Agua Industrial Contaminada Mediante Biopelícula Microbiana, Chala—Arequipa 2018; Universidad César Vallejo: Lima, Peru, 2018. [Google Scholar]
- Vuono, D.C.; Vanneste, J.; Figueroa, L.A.; Hammer, V.; Aguilar-Huaylla, F.N.; Malone, A.; Smith, N.M.; Garcia-Chevesich, P.A.; Bolaños-Sosa, H.G.; Alejo-Zapata, F.D.; et al. Photocatalytic Advanced Oxidation Processes for Neutralizing Free Cyanide in Gold Processing Effluents in Arequipa, Southern Peru. Sustainability 2021, 13, 9873. [Google Scholar] [CrossRef]
- Asner, G.P.; Tupayachi, R. Accelerated losses of protected forests from gold mining in the Peruvian Amazon. Environ. Res. Lett. 2016, 12, 94004. [Google Scholar] [CrossRef]
- Downey, J.; Basi, K.; DeFreytas, M.; Rockwood, G.; Chronic cyanide exposure. Case Studies and Animal Models in Toxicology of Cyanides and Cyanogens: Experimental, Applied and Clinical Aspects, 1st ed.; Hall, A.H., Isom, G.E., Rockwood, G.A., Eds.; John Wiley & Sons: Chichester, UK, 2015; pp. 21–40. ISBN 978-1119978534. [Google Scholar]
- Shifrin, N.S.; Beck, B.D.; Gauthier, T.D.; Chapnick, S.D.; Goodman, G. Chemistry, toxicology, and human health risk of cyanide compounds in soils at former manufactured gas plant sites. Regul. Toxicol. Pharmacol. 1996, 23, 106–116. [Google Scholar] [CrossRef]
- Kuyucak, N.; Akcil, A. Cyanide and removal options from effluents in gold mining and metallurgical processes. Miner. Eng. 2013, 50–51, 13–29. [Google Scholar] [CrossRef]
- Sharma, M.; Akhter, Y.; Chatterjee, S. A review on remediation of cyanide containing industrial wastes using biological systems with special reference to enzymatic degradation. World J. Microbiol. Biotechnol. 2019, 35, 70. [Google Scholar] [CrossRef]
- Akcil, A. Destruction of cyanide in gold mill effluents: Biological versus chemical treatments. Biotechnol. Adv. 2003, 21, 501–511. [Google Scholar] [CrossRef]
- Luque-Almagro, V.M.; Moreno-Vivián, C.; Roldán, M.D. Biodegradation of cyanide wastes from mining and jewellery industries. Curr. Opin. Biotechnol. 2016, 38, 9–13. [Google Scholar] [CrossRef]
- Kunz, D.A.; Wang, C.S.; Chen, J.L. Alternative routes of enzymic cyanide metabolism in Pseudomonas fluorescens NCIMB 11764. Microbiology 1994, 140 Pt 7, 1705–1712. [Google Scholar] [CrossRef]
- Dash, R.R.; Gaur, A.; Balomajumder, C. Cyanide in industrial wastewaters and its removal: A review on biotreatment. J. Hazard. Mater. 2009, 163, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Castric, P.A.; Strobel, G.A. Cyanide Metabolism by Bacillus megaterium. J. Biol. Chem. 1969, 244, 4089–4094. [Google Scholar] [CrossRef]
- Rosario, C.G.A.; Vallenas-Arévalo, A.T.; Arévalo, S.J.; Espinosa, D.C.R.; Tenório, J.A.S. Biodegradation of cyanide using a Bacillus subtilis strain isolated from artisanal gold mining tailings. Braz. J. Chem. Eng. 2023, 40, 129–136. [Google Scholar] [CrossRef]
- Meyers, P.R.; Gokool, P.; Rawlings, D.E.; Woods, D.R. An efficient cyanide-degrading Bacillus pumilus strain. J. Gen. Microbiol. 1991, 137, 1397–1400. [Google Scholar] [CrossRef] [PubMed]
- Alvillo-Rivera, A.; Garrido-Hoyos, S.; Buitrón, G.; Thangarasu-Sarasvathi, P.; Rosano-Ortega, G. Biological treatment for the degradation of cyanide: A review. J. Mater. Res. Technol. 2021, 12, 1418–1433. [Google Scholar] [CrossRef]
- Liu, J.K.; Liu, C.H.; Lin, C.S. The role of nitrogenase in a cyanide-degrading Klebsiella oxytoca strain. Proc. Natl. Sci. Counc. Repub. China B 1997, 21, 37–42. [Google Scholar] [PubMed]
- Adjei, M.D.; Ohta, Y. Isolation and characterization of a cyanide-utilizing Burkholderia cepacia strain. World J. Microbiol. Biotechnol. 1999, 15, 699–704. [Google Scholar] [CrossRef]
- Nallapan Maniyam, M.; Sjahrir, F.; Ibrahim, A.L.; Cass, A.E.G. Biodegradation of cyanide by acetonitrile-induced cells of Rhodococcus sp. UKMP-5M. J. Gen. Appl. Microbiol. 2013, 59, 393–404. [Google Scholar] [CrossRef]
- Ebbs, S. Biological degradation of cyanide compounds. Curr. Opin. Biotechnol. 2004, 15, 231–236. [Google Scholar] [CrossRef]
- Nwokoro, O.; Dibua, M.E.U. Degradation of soil cyanide by single and mixed cultures of Pseudomonas stutzeri and Bacillus subtilis. Arh. Hig. Rada Toksikol. 2014, 65, 113–119. [Google Scholar] [CrossRef]
- Niu, H.; Volesky, B. Characteristics of gold biosorption from cyanide solution. J. Chem. Technol. Biotechnol. 1999, 74, 778–784. [Google Scholar] [CrossRef]
- Knowles, C.J. Cyanide utilization and degradation by microorganisms. Ciba Found. Symp. 1988, 140, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Patil, Y.; Paknikar, K. Development of a process for biodetoxification of metal cyanides from waste waters. Process Biochem. 2000, 35, 1139–1151. [Google Scholar] [CrossRef]
- Obed Ntwampe, S.K. Biodegradation of Free Cyanide Using Bacillus Sp. Consortium Dominated by Bacillus Safensis, Lichenformis and Tequilensis Strains: A Bioprocess Supported Solely with Whey. J. Bioremediation Biodegrad. 2014, 5. [Google Scholar] [CrossRef]
- Akcil, A.; Karahan, A.; Ciftci, H.; Sagdic, O. Biological treatment of cyanide by natural isolated bacteria (Pseudomonas sp.). Miner. Eng. 2003, 16, 643–649. [Google Scholar] [CrossRef]
- Nallapan Maniyam, M.; Sjahrir, F.; Ibrahim, A.L.; Cass, A.E.G. Cyanide degradation by immobilized cells of Rhodococcus UKMP-5M. Biologia 2012, 67, 837–844. [Google Scholar] [CrossRef]
- Ozkan, M.; Desai, S.G.; Zhang, Y.; Stevenson, D.M.; Beane, J.; White, E.A.; Guerinot, M.L.; Lynd, L.R. Characterization of 13 newly isolated strains of anaerobic, cellulolytic, thermophilic bacteria. J. Ind. Microbiol. Biotechnol. 2001, 27, 275–280. [Google Scholar] [CrossRef]
- Mirizadeh, S.; Yaghmaei, S.; Ghobadi Nejad, Z. Biodegradation of cyanide by a new isolated strain under alkaline conditions and optimization by response surface methodology (RSM). J. Environ. Health Sci. Eng. 2014, 12, 85. [Google Scholar] [CrossRef]
- Kumar, V.; Kumar, V.; Bhalla, T.C. Statistical Enhancement of Cyanide Degradation Using Microbial Consortium. J. Microb. Biochem. Technol. 2015, 7, 6. [Google Scholar] [CrossRef]
- Huertas, M.J.; Sáez, L.P.; Roldán, M.D.; Luque-Almagro, V.M.; Martínez-Luque, M.; Blasco, R.; Castillo, F.; Moreno-Vivián, C.; García-García, I. Alkaline cyanide degradation by Pseudomonas pseudoalcaligenes CECT5344 in a batch reactor. Influence of pH. J. Hazard. Mater. 2010, 179, 72–78. [Google Scholar] [CrossRef]
- Rice, E.W.; Baird, R.B.; Eaton, A.D. Standard Methods For the Examination of Water and Wastewater; American Public Health Association: Washington, DC, USA, 2017; ISBN 978-0-87553-299-8. [Google Scholar]
- Kandasamy, S.; Dananjeyan, B.; Krishnamurthy, K.; Benckiser, G. Aerobic cyanide degradation by bacterial isolates from cassava factory wastewater. Braz. J. Microbiol. 2015, 46, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Alvarado-López, M.J.; Garrido-Hoyos, S.E.; Raynal-Gutiérrez, M.E.; El-Kassis, E.G.; Luque-Almagro, V.M.; Rosano-Ortega, G. Cyanide Biodegradation by a Native Bacterial Consortium and Its Potential for Goldmine Tailing Biotreatment. Water 2023, 15, 1595. [Google Scholar] [CrossRef]
- Ibrahim, K.K.; Syed, M.A.; Shukor, M.Y.; Ahmad, S.A. Biological Remediation of Cyanide: A Review. Biotropia 2016, 22, 151–163. [Google Scholar] [CrossRef]
- Mekuto, L.; Alegbeleye, O.O.; Ntwampe, S.K.O.; Ngongang, M.M.; Mudumbi, J.B.; Akinpelu, E.A. Co-metabolism of thiocyanate and free cyanide by Exiguobacterium acetylicum and Bacillus marisflavi under alkaline conditions. 3 Biotech 2016, 6, 173. [Google Scholar] [CrossRef]
- Mekuto, L.; Ntwampe, S.K.O.; Jackson, V.A. Biodegradation of free cyanide and subsequent utilisation of biodegradation by-products by Bacillus consortia: Optimisation using response surface methodology. Environ. Sci. Pollut. Res. Int. 2015, 22, 10434–10443. [Google Scholar] [CrossRef]
- Wu, C.-F.; Xu, X.-M.; Zhu, Q.; Deng, M.-C.; Feng, L.; Peng, J.; Yuan, J.-P.; Wang, J.-H. An effective method for the detoxification of cyanide-rich wastewater by Bacillus sp. CN-22. Appl. Microbiol. Biotechnol. 2014, 98, 3801–3807. [Google Scholar] [CrossRef]
- Dubey, S.K.; Holmes, D.S. Biological cyanide destruction mediated by microorganisms. World J. Microbiol. Biotechnol. 1995, 11, 257–265. [Google Scholar] [CrossRef]
- Slepecky, R.A.; Hemphill, H.E. The Genus Bacillus—Nonmedical. In The Prokaryotes; Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.-H., Stackebrandt, E., Eds.; Springer: New York, NY, USA, 2006; pp. 530–562. ISBN 978-0-387-25494-4. [Google Scholar]
- Stülke, J.; Hillen, W. Regulation of carbon catabolism in Bacillus species. Annu. Rev. Microbiol. 2000, 54, 849–880. [Google Scholar] [CrossRef]
- Berkeley, R.; Logan, N.A.; Shute, L.A.; Capey, A.G. 12 Identification of Bacillus Species; Elsevier: Amsterdam, The Netherlands, 1984; pp. 291–328. ISBN 9780125215169. [Google Scholar]
- Bhattacharya, D.; de Los Santos Villalobos, S.; Ruiz, V.V.; Selvin, J.; Mukherjee, J. Bacillus rugosus sp. nov. producer of a diketopiperazine antimicrobial, isolated from marine sponge Spongia officinalis L. Antonie Van Leeuwenhoek 2020, 113, 1675–1687. [Google Scholar] [CrossRef]
- Gatson, J.W.; Benz, B.F.; Chandrasekaran, C.; Satomi, M.; Venkateswaran, K.; Hart, M.E. Bacillus tequilensis sp. nov., isolated from a 2000-year-old Mexican shaft-tomb, is closely related to Bacillus subtilis. Int. J. Syst. Evol. Microbiol. 2006, 56, 1475–1484. [Google Scholar] [CrossRef]
- Dunlap, C.A.; Bowman, M.J.; Zeigler, D.R. Promotion of Bacillus subtilis subsp. inaquosorum, Bacillus subtilis subsp. spizizenii and Bacillus subtilis subsp. stercoris to species status. Antonie Van Leeuwenhoek 2020, 113, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Muloiwa, M.; Nyende-Byakika, S.; Dinka, M. Comparison of unstructured kinetic bacterial growth models. S. Afr. J. Chem. Eng. 2020, 33, 141–150. [Google Scholar] [CrossRef]
- Malone, S.L.; Pearce, L.L.; Peterson, J. Environmental toxicology of cyanide. In Toxicology of Cyanides and Cyanogens: Experimental, Applied and Clinical Aspects, 1st ed.; Hall, A.H., Isom, G.E., Rockwood, G.A., Eds.; John Wiley & Sons: Chichester, UK, 2015; pp. 82–97. ISBN 978-1119978534. [Google Scholar]
- Kunz, D.A.; Nagappan, O.; Silva-Avalos, J.; Delong, G.T. Utilization of cyanide as nitrogenous substrate by Pseudomonas fluorescens NCIMB 11764: Evidence for multiple pathways of metabolic conversion. Appl. Environ. Microbiol. 1992, 58, 2022–2029. [Google Scholar] [CrossRef] [PubMed]
- Avcioglu, N.H.; Bilkay, I.S. Biological Treatment of Cyanide by Using Klebsiella pneumoniae Species. Food Technol. Biotechnol. 2016, 54, 450–454. [Google Scholar] [CrossRef] [PubMed]
- Dwivedi, N.; Balomajumder, C.; Mondal, P. Comparative evaluation of cyanide removal by adsorption, biodegradation, and simultaneous adsorption and biodegradation (SAB) process using Bacillus cereus and almond shell. J. Environ. Biol. 2016, 37, 551. [Google Scholar]
- Justo Arevalo, S.; Zapata Sifuentes, D.; Cuba Portocarrero, A.; Brescia Reátegui, M.; Monge Pimentel, C.; Farage Martins, L.; Marques Pierry, P.; Morais Piroupo, C.; Guerra Santa Cruz, A.; Quiñones Aguilar, M.; et al. Genomic Characterization of Bacillus safensis Isolated from Mine Tailings in Peru and Evaluation of Its Cyanide-Degrading Enzyme CynD. Appl. Environ. Microbiol. 2022, 88, e0091622. [Google Scholar] [CrossRef]
- Watanabe, A.; Yano, K.; Ikebukuro, K.; Karube, I. Cyanide hydrolysis in a cyanide-degrading bacterium, Pseudomonas stutzeri AK61, by cyanidase. Microbiology 1998, 144 Pt 6, 1677–1682. [Google Scholar] [CrossRef]
- Panay, A.J.; Vargas-Serna, C.L.; Carmona-Orozco, M.L. Biodegradation of cyanide using recombinant Escherichia coli expressing Bacillus pumilus cyanide dihydratase. Rev. Colomb. Biotecnol. 2020, 22, 27–35. [Google Scholar] [CrossRef]
- Dursun, A.Y.; Uslu, G.; Cuci, Y.; Aksu, Z. Bioaccumulation of copper(II), lead(II) and chromium(VI) by growing Aspergillus niger. Process Biochem. 2003, 38, 1647–1651. [Google Scholar] [CrossRef]
- Maciel, A.C.; Da Silva Pena, R.; do Nascimento, L.D.; de Oliveira, T.A.; Chagas-Junior, G.C.A.; Lopes, A.S. Health exposure risks and bioremediation of cyanide in cassava processing effluents: An overview. J. Water Process Eng. 2023, 55, 104079. [Google Scholar] [CrossRef]
- Monga, I.; Paul, V.; Muniyasamy, S.; Zinyemba, O. Green Synthesis of Sodium Cyanide Using Hydrogen Cyanide Extracted under Vacuum from Cassava (Manihot esculenta Crantz) Leaves. Sustain. Chem. 2022, 3, 312–333. [Google Scholar] [CrossRef]
- Shivanoor, S.M.; David, M. Fourier transform infrared (FT-IR) study on cyanide induced biochemical and structural changes in rat sperm. Toxicol. Rep. 2015, 2, 1347–1356. [Google Scholar] [CrossRef]
- Nandiyanto, A.B.D.; Oktiani, R.; Ragadhita, R. How to Read and Interpret FTIR Spectroscope of Organic Material. Indones. J. Sci. Technol. 2019, 4, 97. [Google Scholar] [CrossRef]
- Ojaghi, A.; Shafaie Tonkaboni, S.Z.; Shariati, P.; Doulati Ardejani, F. Novel cyanide electro-biodegradation using Bacillus pumilus ATCC 7061 in aqueous solution. J. Environ. Health Sci. Eng. 2018, 16, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Razanamahandry, L.C.; Andrianisa, H.A.; Karoui, H.; Kouakou, K.M.; Yacouba, H. Biodegradation of free cyanide by bacterial species isolated from cyanide-contaminated artisanal gold mining catchment area in Burkina Faso. Chemosphere 2016, 157, 71–78. [Google Scholar] [CrossRef]
- Abouian Jahromi, M.; Jamshidi-Zanjani, A.; Khodadadi Darban, A. Heavy metal pollution and human health risk assessment for exposure to surface soil of mining area: A comprehensive study. Environ. Earth Sci. 2020, 79, 365. [Google Scholar] [CrossRef]
- Celandroni, F.; Vecchione, A.; Cara, A.; Mazzantini, D.; Lupetti, A.; Ghelardi, E. Identification of Bacillus species: Implication on the quality of probiotic formulations. PLoS ONE 2019, 14, e0217021. [Google Scholar] [CrossRef]
- Durairaju Nisshanthini, S.; Teresa Infanta S, A.K.; Raja, D.S.; Natarajan, K.; Palaniswamy, M.; Angayarkanni, J. Spectral characterization of a pteridine derivative from cyanide-utilizing bacterium Bacillus subtilis—JN989651. J. Microbiol. 2015, 53, 262–271. [Google Scholar] [CrossRef]
- Dangi, A.K.; Sharma, B.; Hill, R.T.; Shukla, P. Bioremediation through microbes: Systems biology and metabolic engineering approach. Crit. Rev. Biotechnol. 2019, 39, 79–98. [Google Scholar] [CrossRef]
- Luque-Almagro, V.M.; Huertas, M.-J.; Martínez-Luque, M.; Moreno-Vivián, C.; Roldán, M.D.; García-Gil, L.J.; Castillo, F.; Blasco, R. Bacterial degradation of cyanide and its metal complexes under alkaline conditions. Appl. Environ. Microbiol. 2005, 71, 940–947. [Google Scholar] [CrossRef]
- Satyanarayana, T.; Johri, B.N. Microorganisms in Environmental Management; Springer: Dordrecht, The Netherlands, 2012; ISBN 978-94-007-2228-6. [Google Scholar]
- Kaspar, F.; Neubauer, P.; Gimpel, M. Bioactive Secondary Metabolites from Bacillus subtilis: A Comprehensive Review. J. Nat. Prod. 2019, 82, 2038–2053. [Google Scholar] [CrossRef] [PubMed]
- Luque Almagro, V.M. Metabolismo Del Cianuro y Del Cianato en “Pseudomonas pseudoalcaligenes” CECT5344: Aplicaciones Biotecnológicas: (Accésit); Analistas Económicos de Andalucía: Málaga, Spain, 2006; ISBN 978-84-95191-81-6. [Google Scholar]
- Botz, M.M.; Mudder, T.I.; Akcil, A.U. Cyanide Treatment. In Gold Ore Processing; Elsevier: Amsterdam, The Netherlands, 2016; pp. 619–645. ISBN 9780444636584. [Google Scholar]
- Benedik, M.J.; Sewell, B.T. Cyanide-degrading nitrilases in nature. J. Gen. Appl. Microbiol. 2018, 64, 90–93. [Google Scholar] [CrossRef] [PubMed]
- Crum, M.A.; Sewell, B.T.; Benedik, M.J. Bacillus pumilus Cyanide Dihydratase Mutants with Higher Catalytic Activity. Front. Microbiol. 2016, 7, 1264. [Google Scholar] [CrossRef] [PubMed]
- Jandhyala, D.; Berman, M.; Meyers, P.R.; Sewell, B.T.; Willson, R.C.; Benedik, M.J. CynD, the cyanide dihydratase from Bacillus pumilus: Gene cloning and structural studies. Appl. Environ. Microbiol. 2003, 69, 4794–4805. [Google Scholar] [CrossRef]
- Guamán Guadalima, M.P.; Nieto Monteros, D.A. Evaluation of the rotational speed and carbon source on the biological removal of free cyanide present on gold mine wastewater, using a rotating biological contactor. J. Water Process Eng. 2018, 23, 84–90. [Google Scholar] [CrossRef]
- Terada, A.; Komatsu, D.; Ogawa, T.; Flamandita, D.; Sahlan, M.; Nishimura, M.; Yohda, M. Isolation of cyanide-degrading bacteria and molecular characterization of its cyanide-degrading nitrilase. Biotechnol. Appl. Biochem. 2022, 69, 183–189. [Google Scholar] [CrossRef]
- Oudjehani, K.; Zagury, G.J.; Deschênes, L. Natural attenuation potential of cyanide via microbial activity in mine tailings. Appl. Microbiol. Biotechnol. 2002, 58, 409–415. [Google Scholar] [CrossRef]
- Deloya, A. Tratamiento de desechos del cianuro por biorremediación. Tecnología en Marcha 2012, 25, 61–72. [Google Scholar] [CrossRef]
- Baxter, J.; Cummings, S.P. The current and future applications of microorganism in the bioremediation of cyanide contamination. Antonie Van Leeuwenhoek 2006, 90, 1–17. [Google Scholar] [CrossRef]
| Bacterial Code | Gram Reaction | Cell Morphology | Spore Formation | Motility |
|---|---|---|---|---|
| TTs1 | Positive | Rod chain | + | + |
| TTs3 | Negative | Rods chain | − | − |
| TT7s | Negative | Rod | − | + |
| TT8s | Positive | Coco | − | − |
| TT9s | Positive | Rods | + | + |
| TT10s | Positive | Rods | + | + |
| TT11s | Positive | Rods chain | + | + |
| TT13s | Negative | Rods | − | + |
| Bacterial | Catalase | Oxidase | MR | VP | Citrate | Urease | Nitrate | Glucose | Maltose | Mannitol | Lactose/N2 | D-Xylose | Sucrose/Indole |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TT10s | + | + | − | + | + | − | + | + | − | + | + | + | + |
| Time (h) | CN− Degradation Efficiency (%) | SD | Plate Count Ln (CFU/mL) | SD |
|---|---|---|---|---|
| 0 | 0.00 | 0.00 | 15.70 | 0.00 |
| 6 | 20.67 | 1.53 | 15.55 | 0.04 |
| 12 | 41.07 | 1.10 | 15.64 | 0.09 |
| 18 | 63.60 | 0.40 | 15.83 | 0.06 |
| 24 | 86.87 | 2.58 | 15.58 | 0.03 |
| 30 | 90.80 | 2.23 | 15.51 | 0.05 |
| 36 | 94.53 | 0.46 | 15.38 | 0.05 |
| 42 | 99.63 | 0.06 | 15.26 | 0.01 |
| 48 | 100.00 | 0.00 | 15.18 | 0.01 |
| Bacterium | Experimental Conditions | Cyanide Degradation | Reference |
|---|---|---|---|
| Bacillus subtilis | 48 h, pH 10.5, 30 °C, 1000 mg/L initial cyanide | 100% | Present work |
| Pseudomonas putida | 24 h, pH 9, 30 °C, 100 mg/L initial cyanide | 90% | [26] |
| Pseudomonas fluorescens | 48 h, pH 7, 30 °C, 50 mg/L initial cyanide | 80% | [62] |
| Bacillus sp. | 96 h, pH 9.88, 33.6 °C, 500 mg/L initial cyanide | 99% | [51] |
| Klebsiella pneumoniae | 72 h, pH 7, 25 °C, 25 mg/L initial cyanide | 87% | [63] |
| Bacillus cereus | 48 h, pH 7, 37 °C, 100 mg/L initial cyanide | 95.87% | [64] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cáceda Quiroz, C.J.; Fora Quispe, G.d.L.; Carpio Mamani, M.; Maraza Choque, G.J.; Sacari Sacari, E.J. Cyanide Bioremediation by Bacillus subtilis under Alkaline Conditions. Water 2023, 15, 3645. https://doi.org/10.3390/w15203645
Cáceda Quiroz CJ, Fora Quispe GdL, Carpio Mamani M, Maraza Choque GJ, Sacari Sacari EJ. Cyanide Bioremediation by Bacillus subtilis under Alkaline Conditions. Water. 2023; 15(20):3645. https://doi.org/10.3390/w15203645
Chicago/Turabian StyleCáceda Quiroz, César Julio, Gabriela de Lourdes Fora Quispe, Milena Carpio Mamani, Gisela July Maraza Choque, and Elisban Juani Sacari Sacari. 2023. "Cyanide Bioremediation by Bacillus subtilis under Alkaline Conditions" Water 15, no. 20: 3645. https://doi.org/10.3390/w15203645
APA StyleCáceda Quiroz, C. J., Fora Quispe, G. d. L., Carpio Mamani, M., Maraza Choque, G. J., & Sacari Sacari, E. J. (2023). Cyanide Bioremediation by Bacillus subtilis under Alkaline Conditions. Water, 15(20), 3645. https://doi.org/10.3390/w15203645









