DNA-Based Tracers for the Characterization of Hydrogeological Systems—Recent Advances and New Frontiers
Abstract
1. Introduction
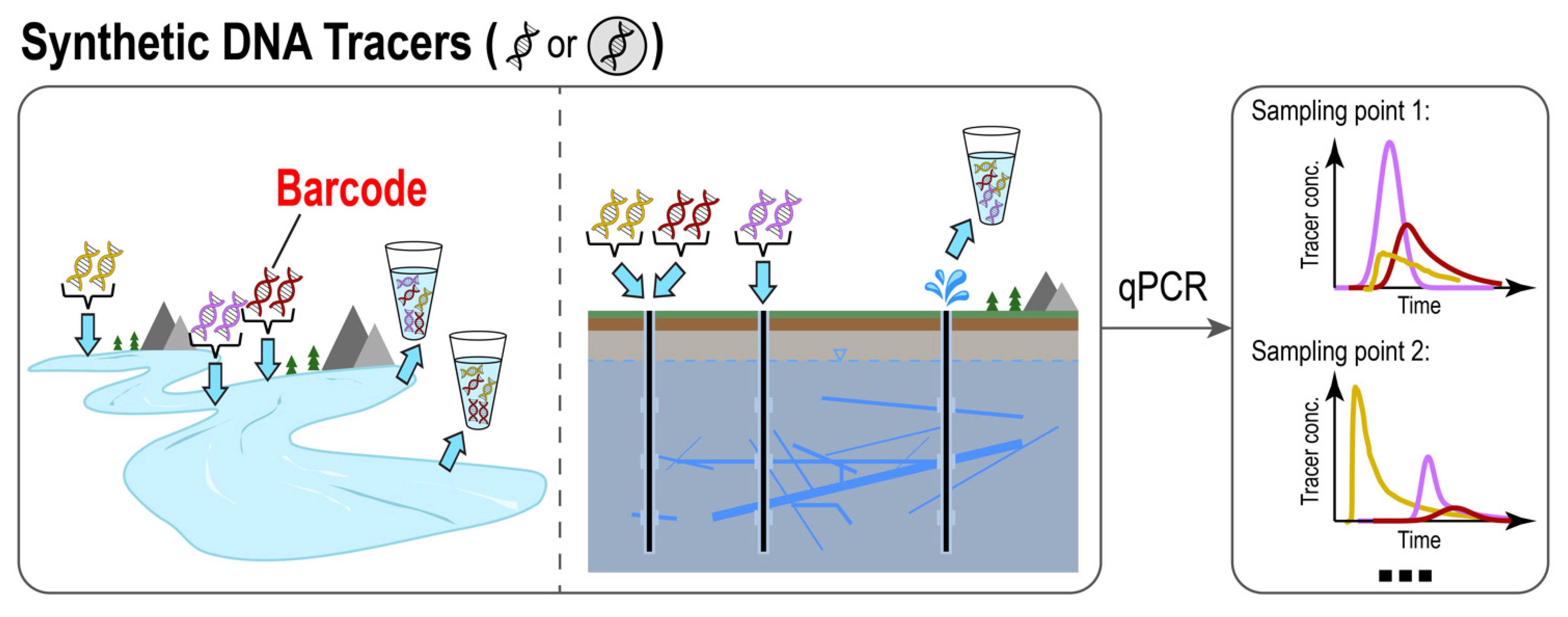
2. Naked DNA: Synthetic DNA in Its Free Form as an Artificial Tracer
2.1. Basic Methods: Design, Synthesis, and Quantification
2.2. Recent Advances
| Authors | Year (Country a) | ss-/ds- | Length b | Medium | Scale c | Details |
|---|---|---|---|---|---|---|
| Amplicon /Total | ||||||
| Sabir, I.H., et al. | 1999 [47] (Norway) | ss | 72/72 | Aquifer with mostly sand (0.06–2 mm) and 5–10% silt (0.002–0.06 mm) | Field, 10 m (<10 m deep) | Presence/absence detection (PCR); sequenced to decode the encoded text. |
| 2000 [53] (Norway) | NM | NM/NM | 1) Aquifer with course glaciofluvial sand and gravel sediments; 2) fractured gneiss bedrock; 3) fractured amphibolite bedrock. | 1) Field, 100 m (<20 m deep); 2) Field, 100 m; 3) Field, 10 m. | Qualitative measurements (PCR); demonstrated the application of synthetic DNA tracers in different field situations. | |
| Ptak, T., et al. | 2004 [18] | ss | NM/90 | 1) Column packed with aquifer media (see below); 2) ~5 m thick saturated aquifer with clay, sand, and gravel. | 1) Lab, L = 50 cm, D = 10 cm; 2) Field, 10 m | Two DNA sequences were tested; the first quantitatively measured (qPCR) DNA breakthrough curves at the field scale. DNA had an earlier peak arrival than bromide in laboratory tests but not necessarily in field tests; tracer recovery NM. |
| Foppen, J.W., et al. | 2011 [27] (Netherlands) | ss | 80/80 | Two streams. | Field, 100–1000 m | Six DNA sequences were used in two injection tests; breakthrough curves were measured at multiple locations downstream. DNA and NaCl had similar peak arrivals; recovery: 94–115% for NaCl and 19–122% for DNA. |
| 2013 [51] (Luxembourg and the Netherlands) | ss | 80/80 | Four streams. | Field, 100 m | One DNA sequence was used in six injection tests; breakthrough curves were measured at multiple locations downstream. DNA and NaCl had similar peak arrivals; recovery: 66.7–106.1% for NaCl and 2.9–52.6% for DNA. | |
| Aquilanti, L., et al. | 2013 [54] (Italy) | ss | 72/72 | 1) Two columns packed with aquifer material (crushed limestone, ~5.1 mm grain size); 2) fractured limestone aquifer | 1) Lab, L = 20 cm, D = 1.8 cm & L = 44.6 cm, D = 5.2 cm; 2) Field, 1 km | One DNA sequence was injected into a sinkhole, and breakthrough curves were measured at a spring downstream. DNA peak arrival occurred earlier than the reference tracer in all lab/field tests; tracer recovery NM. |
| 2016 [55] (Italy) | ss | 72/72 | Two karstic springs. | Field, 1 km | One DNA sequence was tested. DNA peak arrival occurred earlier or at similar time as the fluorescein tracer; the DNA breakthrough curve had sharp pulses rather than a Gaussian shape; recovery: 0–63% for fluorescein; NM for DNA. | |
| Bovolin, V., et al. | 2014 [56] (Italy) | NM | NM/NM | Karstic spring. | Field, 100 m. | One DNA sequence was injected, and the breakthrough curve was measured ~300 m downstream. DNA and salt had similar peak arrivals; recovery: 33.5% for salt and 87% for DNA. |
| Dahlke, H.E., et al. | 2015 [26] (Sweden) | ss | 95/99, 88/90 | Subglacial drainage system. | Field, 1 km | Three DNA sequences were injected into different moulins, and breakthrough curves were measured along proglacial streams. Recovery: 99% for uranine and 1–57% for free DNA. |
| Pang, L., et al. | 2017 [32] (New Zealand) | ds | 198/302 | 1) Column packed with aquifer gravels (94% > 2 mm, 3% 0.5–2 mm, 3% < 0.5 mm) from field site; 2) alluvial gravel aquifer; 3) lysimeter with stony soil. | 1) Lab, L = 2 m, D = 19 cm; 2) Field, 10 m 3) Lab, L = 0.7 m, D = 0.5 m. | Ten DNA sequences were tested. DNA typically had earlier breakthrough, less dispersion, and less recovery than the bromide reference tracer. |
| 2020 [52] (New Zealand) | ds | 198/302, 200/352 | 1) Surface stream; 2) alluvial gravel aquifer; 3) coastal sand aquifer; 4) soil lysimeter. | 1) Field, 1 km; 2) Field, 10 m (16-m below ground); 3) Field, 1 m; 4) Lab, L = 0.7 m, D = 0.5 m. | Free DNA was tested and compared with DNA encapsulated in polymer microparticles. Free DNA had greater mass recoveries than encapsulated DNA in the gravel aquifer but lower recoveries in the surface water experiments. | |
| 2022 [61] (New Zealand) | ds | 198/302, 200/352 | 1) Incubation tests using stream water, groundwater, and wastewater; 2) batch adsorption tests using stream sediments, soils, and aquifer media with water from each site. | 1) Lab, 1.5 ml tubes; 2) Lab, 30 ml tubes. | Incubation tests were conducted for ~10 days, and adsorption tests were conducted for ~24 hr. With similar amplicon lengths, DNA with longer flanking regions degraded slower than DNA with shorter flanking regions. Longer DNA had similar or less adsorption than shorter DNA, depending on the type of solid phase. | |
| Zhang, Y., et al. | 2017 [62] (USA) | ds | 113/113, 141/~2-million | Batch heating test in Tris-EDTA buffer. | Lab, 5 ml vessels. | The heat degradation of short, synthetic DNA was compared with that of long, genomic DNA; the similar-length amplicon region was better preserved in the long DNA after one-hour heat treatment at 150 °C. |
| 2021 [28] (USA) | ds | 70/90, 90/110, 112/132, 114/134, 141/161, 160/180, 180/200 | 1) Column packed with glass beads (avg. 82.5 μm); 2) column packed with quartz sand (avg. 149 μm). | 1) Lab, L = 50 cm, D = 1 cm; 2) Lab, L = 50 cm, D = 1 cm. | The effect of DNA length and adsorption on DNA tracer transport was investigated (nine sequences with six lengths tested). DNA peaks occurred faster than bromide in the glass-bead column but slower in the sand column. DNA recovery increased with increasing length in the glass-bead column but decreased with increasing length in the sand column, which can be explained by the corresponding trends in adsorption partition coefficients. | |
| McCluskey, J., et al. | 2021 [48] (USA) | ds | 200/300 | 1) Incubation test using river water and distilled water; 2) column packed with sand and limestone; 3) surface stream. | 1) Lab, 2 ml tubes; 2) Lab, L = 25.4 cm, D = 6.35 cm; 3) Field, 10 m. | Two DNA sequences were tested; DNA concentrations were measured by droplet digital PCR (ddPCR). DNA had earlier peak arrival and lower mass recovery than uranine; different DNA sequences had the same peak arrival but different recoveries in the column test. |
| Wang, C., et al. | 2022 [57] (USA) | ss | 88/88 | Sloped lysimeter packed with crushed basaltic tephra (3.2% <2 μm, 12.2% 2–50 μm, 84.6% 50–2000 μm) | Lab, L = 2 m, W = 0.5 m, H = 1 m. | One DNA sequence was tested; DNA transport through the vadose zone under variably saturated transient flow conditions was studied both experimentally and numerically. Recovery: 97.78% for deuterium and 1.08% for DNA. |
3. Encapsulated DNA: Synthetic DNA Embedded in Nano/Microparticle Carriers as an Artificial Tracer
3.1. Basic Methods: Design, Synthesis, Release and Quantification
3.2. Recent Advances
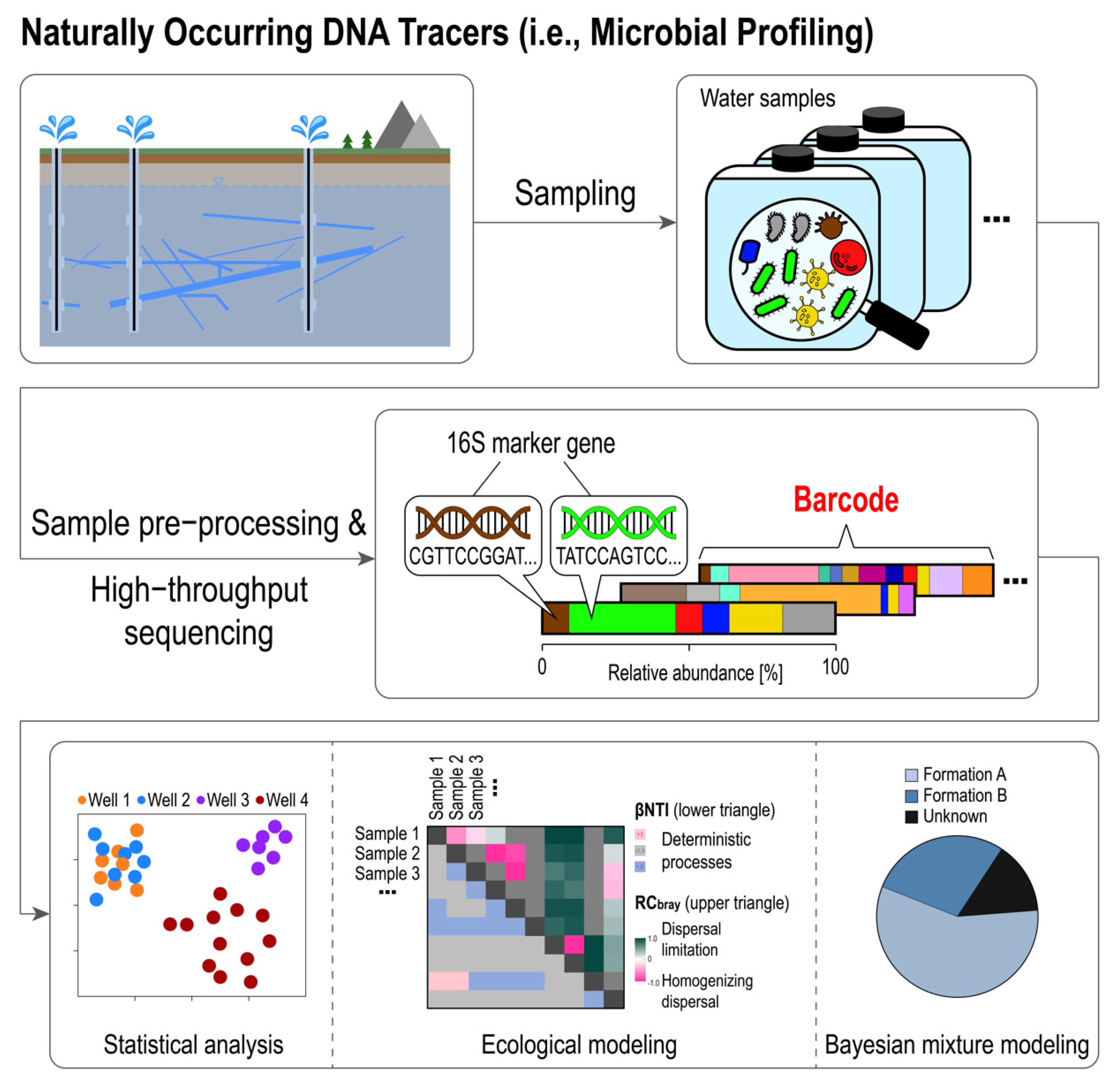
4. Barcoding Microbial Communities: The Composition of Reservoir Native Microbial Communities as a Natural Tracer
4.1. Basic Methods: High-Throughput Sequencing and Statistical Data Analysis
4.2. Recent Advances
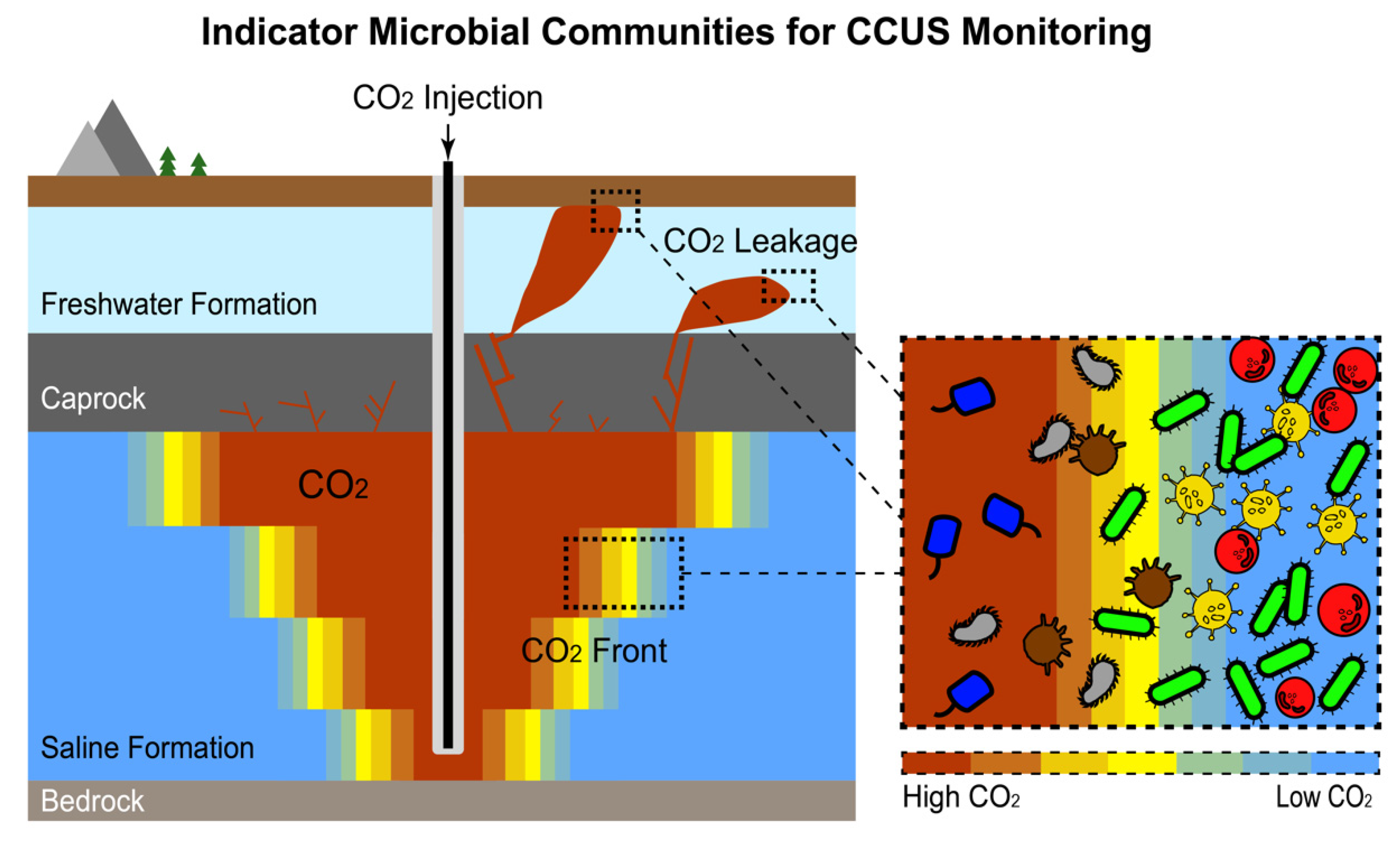
5. Indicator Microbial Communities: Microbial Response to Environmental Anomalies as an Indicator to Indirectly “Trace” the Seepage/Leakage of Hydrocarbon/CO2
6. Two Decades of Progress, Side-by-Side Comparisons and the Path Forward
7. Concluding Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Leibundgut, C.; Maloszewski, P.; Külls, C. Tracers in Hydrology; Wiley-Blackwell Chichester: Chichester, UK, 2009. [Google Scholar]
- Huang, T.; Pang, Z.; Yang, S.; Yin, L. Impact of Afforestation on Atmospheric Recharge to Groundwater in a Semiarid Area. J. Geophys. Res. Atmos. 2020, 125, e2019JD032185. [Google Scholar] [CrossRef]
- Medici, G.; Langman, J.B. Pathways and Estimate of Aquifer Recharge in a Flood Basalt Terrain; A Review from the South Fork Palouse River Basin (Columbia River Plateau, USA). Sustainability 2022, 14, 11349. [Google Scholar] [CrossRef]
- Li, J.; Pang, Z.; Tian, L.; Zhao, H.; Bai, G. Variations of Stable Isotopes in Daily Precipitation in a Monsoon Region. Water 2022, 14, 2891. [Google Scholar] [CrossRef]
- Kong, Y.; Pang, Z. A positive altitude gradient of isotopes in the precipitation over the Tianshan Mountains: Effects of moisture recycling and sub-cloud evaporation. J. Hydrol. 2016, 542, 222–230. [Google Scholar] [CrossRef]
- Huang, T.; Pang, Z. Changes in groundwater induced by water diversion in the Lower Tarim River, Xinjiang Uygur, NW China: Evidence from environmental isotopes and water chemistry. J. Hydrol. 2010, 387, 188–201. [Google Scholar] [CrossRef]
- Huang, T.; Li, Z.; Ma, B.; Long, Y. Tracing the Origin of Groundwater Nitrate in an Area Affected by Acid Rain Using Dual Isotopic Composition of Nitrate. Geofluids 2019, 2019, 1–12. [Google Scholar] [CrossRef]
- Koletzko, B.; Sauerwald, T.; Demmelmair, H. Safety of stable isotope use. Eur. J. Pediatr. 1997, 156, S12–S17. [Google Scholar] [CrossRef]
- Yurtsever, Y. An overview of conceptual model formulations for evaluation of isotope data in hydrological systems. IAHS Publ.-Ser. Proc. Rep.-Intern Assoc. Hydrol. Sci. 1995, 229, 3–12. [Google Scholar]
- Zhang, Y.; Dekas, A.E.; Hawkins, A.J.; Parada, A.E.; Gorbatenko, O.; Li, K.; Horne, R.N. Microbial Community Composition in Deep-Subsurface Reservoir Fluids Reveals Natural Interwell Connectivity. Water Resour. Res. 2020, 56, e2019WR025916. [Google Scholar] [CrossRef]
- Aydin, H.; Nagabandi, N.; Jamal, D.; Temizel, C. A Comprehensive Review of Tracer Test Applications in Geothermal Reservoirs. In Proceedings of the SPE Western Regional Meeting, Bakersfield, CA, USA, 26–28 April 2022. [Google Scholar]
- Ursell, L.; Hale, M.; Menendez, E.; Zimmerman, J.; Dombroski, B.; Hoover, K.; Everman, Z.; Liu, J.K.; Shojaei, H.; Percak-Dennett, E.M. High Resolution Fluid Tracking from Verticals and Laterals Using Subsurface DNA Diagnostics in the Permian Basin. In Proceedings of the Unconventional Resources Technology Conference, Denver, CO, USA, 22–24 July 2019; pp. 1291–1302. [Google Scholar]
- Horne, R.N. Geothermal Reinjection Experience in Japan. J. Pet. Technol. 1982, 34, 495–503. [Google Scholar] [CrossRef]
- Tayyib, D.; Al-Qasim, A.; Kokal, S.; Huseby, O. Overview of tracer applications in oil and gas industry. In Proceedings of the SPE Kuwait Oil & Gas Show and Conference, Mishref, Kuwait, 14 October 2019. [Google Scholar]
- Hawkins, A.J.; Becker, M.W.; Tester, J.W. Inert and Adsorptive Tracer Tests for Field Measurement of Flow-Wetted Surface Area. Water Resour. Res. 2018, 54, 5341–5358. [Google Scholar] [CrossRef]
- Hawkins, A.J.; Fox, D.B.; Koch, D.L.; Becker, M.W.; Tester, J.W. Predictive Inverse Model for Advective Heat Transfer in a Short-Circuited Fracture: Dimensional Analysis, Machine Learning, and Field Demonstration. Water Resour. Res. 2020, 56, e2020WR027065. [Google Scholar] [CrossRef]
- Suzuki, A.; Cui, J.; Zhang, Y.; Uehara, S.; Li, K.; Horne, R.N.; Ito, T. Experimental Study on Nano-/Microparticles Transport to Characterize Structures in Fractured Porous Media. Rock Mech. Rock Eng. 2020, 53, 4357–4365. [Google Scholar] [CrossRef]
- Ptak, T.; Piepenbrink, M.; Martac, E. Tracer tests for the investigation of heterogeneous porous media and stochastic modelling of flow and transport—A review of some recent developments. J. Hydrol. 2004, 294, 122–163. [Google Scholar] [CrossRef]
- McKay, L.D.; Sanford, W.E.; Strong, J.M. Field-Scale Migration of Colloidal Tracers in a Fractured Shale Saprolite. Ground Water 2000, 38, 139–147. [Google Scholar] [CrossRef]
- Medici, G.; West, L.J. Groundwater flow velocities in karst aquifers; importance of spatial observation scale and hydraulic testing for contaminant transport prediction. Env. Sci. Pollut. Res. Int. 2021, 28, 43050–43063. [Google Scholar] [CrossRef]
- Worthington, S.R. Diagnostic tests for conceptualizing transport in bedrock aquifers. J. Hydrol. 2015, 529, 365–372. [Google Scholar] [CrossRef]
- Worthington, S.R.; Ford, D. Self-organized permeability in carbonate aquifers. Groundwater 2009, 47, 326–336. [Google Scholar] [CrossRef]
- Goldscheider, N.; Meiman, J.; Pronk, M.; Smart, C. Tracer tests in karst hydrogeology and speleology. Int. J. Speleol. 2008, 37, 27–40. [Google Scholar] [CrossRef]
- Brauchler, R.; Böhm, G.; Leven, C.; Dietrich, P.; Sauter, M. A laboratory study of tracer tomography. Hydrogeol. J. 2013, 21, 1265–1274. [Google Scholar] [CrossRef]
- Kong, X.Z.; Deuber, C.A.; Kittila, A.; Somogyvari, M.; Mikutis, G.; Bayer, P.; Stark, W.J.; Saar, M.O. Tomographic Reservoir Imaging with DNA-Labeled Silica Nanotracers: The First Field Validation. Env. Sci. Technol. 2018, 52, 13681–13689. [Google Scholar] [CrossRef] [PubMed]
- Dahlke, H.E.; Williamson, A.G.; Georgakakos, C.; Leung, S.; Sharma, A.N.; Lyon, S.W.; Walter, M.T. Using concurrent DNA tracer injections to infer glacial flow pathways. Hydrol. Process. 2015, 29, 5257–5274. [Google Scholar] [CrossRef]
- Foppen, J.W.; Orup, C.; Adell, R.; Poulalion, V.; Uhlenbrook, S. Using multiple artificial DNA tracers in hydrology. Hydrol. Process. 2011, 25, 3101–3106. [Google Scholar] [CrossRef]
- Zhang, Y.; Hartung, M.B.; Hawkins, A.J.; Dekas, A.E.; Li, K.; Horne, R.N. DNA tracer transport through porous media—The effect of DNA length and adsorption. Water Resour. Res. 2021, 57, 2020WR028382. [Google Scholar] [CrossRef]
- Zhang, Y.; Manley, T.S.; Li, K.; Home, R.N. DNA-Encapsulated silica nanoparticle tracers for fracture characterization. In Proceedings of the 39th Geothermal Resources Council Annual Meeting—Geothermal: Always on, GRC 2015, Reno, NV, USA, 20–23 September 2015; pp. 967–974. [Google Scholar]
- Zhang, Y.; Dekas, A.E.; Hawkins, A.J.; Primo, J.C.; Gorbatenko, O.; Horne, R.N. Comparison of Microbial Profiling and Tracer Testing for the Characterization of Injector-Producer Interwell Connectivities. Water 2022, 14, 2921. [Google Scholar] [CrossRef]
- Zhang, Y.; Horne, R.N.; Hawkins, A.J.; Primo, J.C.; Gorbatenko, O.; Dekas, A.E. Geological activity shapes the microbiome in deep-subsurface aquifers by advection. Proc. Natl. Acad. Sci. USA 2022, 119, e2113985119. [Google Scholar] [CrossRef] [PubMed]
- Pang, L.; Robson, B.; Farkas, K.; McGill, E.; Varsani, A.; Gillot, L.; Li, J.; Abraham, P. Tracking effluent discharges in undisturbed stony soil and alluvial gravel aquifer using synthetic DNA tracers. Sci. Total Environ. 2017, 592, 144–152. [Google Scholar] [CrossRef] [PubMed]
- Lascelles, P.; Wan, J.; Robinson, L.; Allmon, R.; Evans, G.; Ursell, L.; Scott, N.M.; Chase, J.; Jablanovic, J.; Karimi, M. Applying Subsurface DNA Sequencing in Wolfcamp Shales, Midland Basin. In Proceedings of the SPE Hydraulic Fracturing Technology Conference and Exhibition, The Woodlands, TX, USA, 25 January 2017. [Google Scholar]
- Merino, N.; Jackson, T.R.; Campbell, J.H.; Kersting, A.B.; Sackett, J.; Fisher, J.C.; Bruckner, J.C.; Zavarin, M.; Hamilton-Brehm, S.D.; Moser, D.P. Subsurface microbial communities as a tool for characterizing regional-scale groundwater flow. Sci. Total Environ. 2022, 842, 156768. [Google Scholar] [CrossRef]
- Sawadogo, J.; Ursell, L.; Reeve, N.; Schlecht, M. Mature Fields-Optimizing Waterflood Management Through DNA Based Diagnostics. In Proceedings of the Abu Dhabi International Petroleum Exhibition & Conference, Abu Dhabi, UAE, 11 November 2020. [Google Scholar]
- Fukami, T. Historical contingency in community assembly: Integrating niches, species pools, and priority effects. Annu. Rev. Ecol. Evol. Syst. 2015, 46, 1–23. [Google Scholar] [CrossRef]
- Zhou, J.; Ning, D. Stochastic community assembly: Does it matter in microbial ecology? Microbiol. Mol. Biol. Rev. 2017, 81, e00002–e00017. [Google Scholar] [CrossRef]
- Schlecht, M.; Sawadogo, J.; Sadeghi, S.; Reeve, N.; Haggerty, M.; Liu, J.; Ursell, L. Improved Stacked Permian Development by Integrating DNA Diagnostics with Traditional Reservoir Analysis. In Proceedings of the SPE Annual Technical Conference and Exhibition, Virtual, 27 October 2020. [Google Scholar]
- Silva, J.; Ursell, L.; Percak-Dennett, E. Applying Subsurface DNA Diagnostics and Data Science in the Delaware Basin. In Proceedings of the SPE Hydraulic Fracturing Technology Conference and Exhibition, The Woodlands, TX, USA, 23–25 January 2018. [Google Scholar]
- Deepak, S.; Kottapalli, K.; Rakwal, R.; Oros, G.; Rangappa, K.; Iwahashi, H.; Masuo, Y.; Agrawal, G. Real-Time PCR: Revolutionizing Detection and Expression Analysis of Genes. Curr. Genom. 2007, 8, 234–251. [Google Scholar] [CrossRef] [PubMed]
- Wetterstrand, K.A. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP). Available online: www.genome.gov/sequencingcostsdata (accessed on 23 September 2022).
- Metzker, M.L. Sequencing technologies—the next generation. Nat. Rev. Genet. 2010, 11, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Rothmel, R.; Onstott, T.C.; Fuller, M.E.; DeFlaun, M.F.; Streger, S.H.; Dunlap, R.; Fletcher, M. Simultaneous transport of two bacterial strains in intact cores from Oyster, Virginia: Biological effects and numerical modeling. Appl Env. Microbiol 2002, 68, 2120–2132. [Google Scholar] [CrossRef] [PubMed]
- Paruch, L.; Paruch, A.M. An Overview of Microbial Source Tracking Using Host-Specific Genetic Markers to Identify Origins of Fecal Contamination in Different Water Environments. Water 2022, 14, 1809. [Google Scholar] [CrossRef]
- Pang, L.; Farkas, K.; Bennett, G.; Varsani, A.; Easingwood, R.; Tilley, R.; Nowostawska, U.; Lin, S. Mimicking filtration and transport of rotavirus and adenovirus in sand media using DNA-labeled, protein-coated silica nanoparticles. Water Res. 2014, 62, 167–179. [Google Scholar] [CrossRef]
- Grass, R.N.; Schälchli, J.; Paunescu, D.; Soellner, J.O.; Kaegi, R.; Stark, W.J. Tracking trace amounts of submicrometer silica particles in wastewaters and activated sludge using silica-encapsulated DNA barcodes. Environ. Sci. Technol. Lett. 2014, 1, 484–489. [Google Scholar] [CrossRef]
- Sabir, I.H.; Torgersen, J.; Haldorsen, S.; Aleström, P. DNA tracers with information capacity and high detection sensitivity tested in groundwater studies. Hydrogeol. J. 1999, 7, 264–272. [Google Scholar] [CrossRef]
- McCluskey, J.; Flores, M.E.; Hinojosa, J.; Jafarzadeh, A.; Moghadam, S.V.; Phan, D.C.; Green, R.T.; Kapoor, V. Tracking Water with Synthetic DNA Tracers Using Droplet Digital PCR. ACS EST Water 2021, 1, 1177–1183. [Google Scholar] [CrossRef]
- Anderson, C.F.; Record Jr, M.T. Polyelectrolyte theories and their applications to DNA. Annu. Rev. Phys. Chem. 1982, 33, 191–222. [Google Scholar] [CrossRef]
- Cooper, G.M.; Hausman, R.E.; Hausman, R.E. The Cell: A Molecular Approach; ASM Press: Washington, DC, USA, 2007; Volume 4. [Google Scholar]
- Foppen, J.W.; Seopa, J.; Bakobie, N.; Bogaard, T. Development of a methodology for the application of synthetic DNA in stream tracer injection experiments. Water Resour. Res. 2013, 49, 5369–5380. [Google Scholar] [CrossRef]
- Pang, L.; Abeysekera, G.; Hanning, K.; Premaratne, A.; Robson, B.; Abraham, P.; Sutton, R.; Hanson, C.; Hadfield, J.; Heiligenthal, L.; et al. Water tracking in surface water, groundwater and soils using free and alginate-chitosan encapsulated synthetic DNA tracers. Water Res. 2020, 184, 116192. [Google Scholar] [CrossRef] [PubMed]
- Sabir, I.H.; Haldorsen, S.; Torgersen, J.; Alestrom, P.; Gaut, S.; Colleuille, H.; Pedersen, T.S.; Kitterod, N.-O.; Alestrom, P. Synthetic DNA tracers: Examples of their application in water related studies. IAHS Publ. (Int. Assoc. Hydrol. Sci. ) 2000, 262, 159–165. [Google Scholar]
- Aquilanti, L.; Clementi, F.; Landolfo, S.; Nanni, T.; Palpacelli, S.; Tazioli, A. A DNA tracer used in column tests for hydrogeology applications. Environ. Earth Sci. 2013, 70, 3143–3154. [Google Scholar] [CrossRef]
- Aquilanti, L.; Clementi, F.; Nanni, T.; Palpacelli, S.; Tazioli, A.; Vivalda, P.M. DNA and fluorescein tracer tests to study the recharge, groundwater flowpath and hydraulic contact of aquifers in the Umbria-Marche limestone ridge (central Apennines, Italy). Environ. Earth Sci. 2016, 75, 1–17. [Google Scholar] [CrossRef]
- Bovolin, V.; Cuomo, A.; Guida, D.; Foppen, J.W. Using artificial DNA as tracer in a bedrock river of the Middle Bussento Karst System (Cilento, Vallo Diano and Alburni European & Global Geopark, southern Italy). In Proceedings of the the 7th International Conference on Engineering Mechanics, Structures, Engineering Geology (EMESEG14), Salerno, Italy, 3–5 June 2014; pp. 105–112. [Google Scholar]
- Wang, C.; Liu, G.; McNew, C.P.; Volkmann, T.H.M.; Pangle, L.; Troch, P.A.; Lyon, S.W.; Kim, M.; Huo, Z.; Dahlke, H.E. Simulation of experimental synthetic DNA tracer transport through the vadose zone. Water Res. 2022, 223, 119009. [Google Scholar] [CrossRef]
- Viswanathan, H.S.; Ajo-Franklin, J.; Birkholzer, J.T.; Carey, J.W.; Guglielmi, Y.; Hyman, J.; Karra, S.; Pyrak-Nolte, L.; Rajaram, H.; Srinivasan, G. From fluid flow to coupled processes in fractured rock: Recent advances and new frontiers. Rev. Geophys. 2022, 60, e2021RG000744. [Google Scholar] [CrossRef]
- Huang, T.; Li, Z.; Long, Y.; Zhang, F.; Pang, Z. Role of desorption-adsorption and ion exchange in isotopic and chemical (Li, B, and Sr) evolution of water following water–rock interaction. J. Hydrol. 2022, 610, 127800. [Google Scholar] [CrossRef]
- Huang, T.; Li, Z.; Mayer, B.; Nightingale, M.; Li, X.; Li, G.; Long, Y.; Pang, Z. Identification of Geochemical Processes During Hydraulic Fracturing of a Shale Gas Reservoir: A Controlled Field and Laboratory Water-Rock Interaction Experiment. Geophys. Res. Lett. 2020, 47, e2020GL090420. [Google Scholar] [CrossRef]
- Pang, L.; Heiligenthal, L.; Premaratne, A.; Hanning, K.R.; Abraham, P.; Sutton, R.; Hadfield, J.; Billington, C. Degradation and adsorption of synthetic DNA water tracers in environmental matrices. Sci Total Environ. 2022, 844, 157146. [Google Scholar] [CrossRef]
- Zhang, Y.; Zeng, Z.; Li, K.; Horne, R. DNA Barcoding for Fractured Reservoir Analysis—An Initial Investigation. In Proceedings of the 42nd Workshop on Geothermal Reservoir Engineering, Stanford University, Stanford, CA, USA, 13–15 February 2017. [Google Scholar]
- Alaskar, M.; Ames, M.; Liu, C.; Li, K.; Horne, R. Temperature nanotracers for fractured reservoirs characterization. J. Pet. Sci. Eng. 2015, 127, 212–228. [Google Scholar] [CrossRef]
- Zhang, Y.; Manley, T.S.; Li, K.; Horne, R. Uniquely Identifiable DNA-Embedded Silica Nanotracer for Fractured Reservoir Characterization. In Proceedings of the 41st Workshop on Geothermal Reservoir Engineering, Stanford University, Stanford, CA, USA, 22–24 February 2016. [Google Scholar]
- Mikutis, G.; Deuber, C.A.; Schmid, L.; Kittila, A.; Lobsiger, N.; Puddu, M.; Asgeirsson, D.O.; Grass, R.N.; Saar, M.O.; Stark, W.J. Silica-Encapsulated DNA-Based Tracers for Aquifer Characterization. Environ. Sci. Technol. 2018, 52, 12142–12152. [Google Scholar] [CrossRef] [PubMed]
- Kittilä, A.; Jalali, M.R.; Evans, K.F.; Willmann, M.; Saar, M.O.; Kong, X.Z. Field Comparison of DNA-Labeled Nanoparticle and Solute Tracer Transport in a Fractured Crystalline Rock. Water Resour. Res. 2019, 55, 6577–6595. [Google Scholar] [CrossRef]
- Tang, Y.; Foppen, J.W.; Bogaard, T.A. Transport of silica encapsulated DNA microparticles in controlled instantaneous injection open channel experiments. J. Contam. Hydrol. 2021, 242, 103880. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Foppen, J.W.; Schijven, J.F. Effect of concentration of silica encapsulated ds-DNA colloidal microparticles on their transport through saturated porous media. Colloids Surf. A Physicochem. Eng. Asp. 2022, 651, 129625. [Google Scholar] [CrossRef]
- Kianfar, B.; Tian, J.; Rozemeijer, J.; van der Zaan, B.; Bogaard, T.A.; Foppen, J.W. Transport characteristics of DNA-tagged silica colloids as a colloidal tracer in saturated sand columns; role of solution chemistry, flow velocity, and sand grain size. J. Contam. Hydrol. 2022, 246, 103954. [Google Scholar] [CrossRef]
- Sharma, A.N.; Luo, D.; Walter, M.T. Hydrological tracers using nanobiotechnology: Proof of concept. Environ. Sci. Technol. 2012, 46, 8928–8936. [Google Scholar] [CrossRef]
- Wang, C.; McNew, C.P.; Lyon, S.W.; Walter, M.T.; Volkman, T.H.M.; Abramson, N.; Sengupta, A.; Wang, Y.; Meira Neto, A.A.; Pangle, L.; et al. Particle tracer transport in a sloping soil lysimeter under periodic, steady state conditions. J. Hydrol. 2019, 569, 61–76. [Google Scholar] [CrossRef]
- Liao, R.; Zhang, J.; Li, T.; Luo, D.; Yang, D. Biopolymer/plasmid DNA microspheres as tracers for multiplexed hydrological investigation. Chem. Eng. J. 2020, 401, 126035. [Google Scholar] [CrossRef]
- McNew, C.P.; Wang, C.; Walter, M.T.; Dahlke, H.E. Fabrication, detection, and analysis of DNA-labeled PLGA particles for environmental transport studies. J. Colloid. Interface Sci. 2018, 526, 207–219. [Google Scholar] [CrossRef]
- Georgakakos, C.B.; Richards, P.L.; Walter, M.T. Tracing Septic Pollution Sources Using Synthetic DNA Tracers: Proof of Concept. Air Soil Water Res. 2019, 12. [Google Scholar] [CrossRef]
- Paunescu, D.; Puddu, M.; Soellner, J.O.; Stoessel, P.R.; Grass, R.N. Reversible DNA encapsulation in silica to produce ROS-resistant and heat-resistant synthetic DNA ‘fossils’. Nat. Protoc. 2013, 8, 2440–2448. [Google Scholar] [CrossRef] [PubMed]
- Alexander, G.B.; Heston, W.M.; Iler, R.K. The Solubility of Amorphous Silica in Water. J. Phys. Chem. 1954, 58, 453–455. [Google Scholar] [CrossRef]
- Liao, H.; Yu, K.; Duan, Y.; Ning, Z.; Li, B.; He, L.; Liu, C. Profiling microbial communities in a watershed undergoing intensive anthropogenic activities. Sci. Total Environ. 2019, 647, 1137–1147. [Google Scholar] [CrossRef] [PubMed]
- Semler, A.C.; Fortney, J.L.; Fulweiler, R.W.; Dekas, A.E. Cold Seeps on the Passive Northern U.S. Atlantic Margin Host Globally Representative Members of the Seep Microbiome with Locally Dominant Strains of Archaea. Appl Environ. Microbiol. 2022, 88, e0046822. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, A.; Hashida, T.; Li, K.; Horne, R.N. Experimental tests of truncated diffusion in fault damage zones. Water Resour. Res. 2016, 52, 8578–8589. [Google Scholar] [CrossRef]
- Nguyen, L.H.; Holmes, S. Ten quick tips for effective dimensionality reduction. PLoS Comput. Biol. 2019, 15, e1006907. [Google Scholar] [CrossRef]
- Callahan, B.J.; Sankaran, K.; Fukuyama, J.A.; McMurdie, P.J.; Holmes, S.P. Bioconductor Workflow for Microbiome Data Analysis: From raw reads to community analyses. F1000Res 2016, 5, 1492. [Google Scholar] [CrossRef]
- Knights, D.; Kuczynski, J.; Charlson, E.S.; Zaneveld, J.; Mozer, M.C.; Collman, R.G.; Bushman, F.D.; Knight, R.; Kelley, S.T. Bayesian community-wide culture-independent microbial source tracking. Nat. Methods 2011, 8, 761–763. [Google Scholar] [CrossRef]
- Griebler, C.; Lueders, T. Microbial biodiversity in groundwater ecosystems. Freshw. Biol. 2009, 54, 649–677. [Google Scholar] [CrossRef]
- Hubalek, V.; Wu, X.; Eiler, A.; Buck, M.; Heim, C.; Dopson, M.; Bertilsson, S.; Ionescu, D. Connectivity to the surface determines diversity patterns in subsurface aquifers of the Fennoscandian shield. ISME J. 2016, 10, 2447–2458. [Google Scholar] [CrossRef]
- Gringarten, E.; Deutsch, C. Methodology for variogram interpretation and modeling for improved reservoir characterization. In Proceedings of the Spe annual technical conference and exhibition, Houston, TX, USA, 3–6 October 1999. [Google Scholar]
- Mukerji, T.; Avseth, P.; Mavko, G.; Takahashi, I.; González, E.F. Statistical rock physics: Combining rock physics, information theory, and geostatistics to reduce uncertainty in seismic reservoir characterization. Lead. Edge 2001, 20, 313–319. [Google Scholar] [CrossRef]
- Percak-Dennett, E.; Liu, J.; Shojaei, H.; Luke, U.; Thomas, I. High Resolution Dynamic Drainage Height Estimations using Subsurface DNA Diagnostics. In Proceedings of the SPE Western Regional Meeting, San Jose, CA, USA, 24 April 2019. [Google Scholar]
- Sawadogo, J.; Haggerty, M.; Mallory, C.; Huchton, J.; DeAngelis, W.; Price, C. Impact of Completion Design and Interwell Communication on Well Performance in Full Section Development: A STACK Case Study Using DNA Based Diagnostics. In Proceedings of the SPE Annual Technical Conference and Exhibition, Virtual, 27 October 2020. [Google Scholar]
- Kneafsey, T.J.; Blankenship, D.; Dobson, P.F.; Morris, J.P.; White, M.D.; Fu, P.; Schwering, P.C.; Ajo-Franklin, J.B.; Huang, L.; Schoenball, M.; et al. The EGS Collab Project -Learnings from Experiment 1. In Proceedings of the 45th Workshop on Geothermal Reservoir Engineering, Stanford University, Stanford, CA, USA, 10–12 February 2020. [Google Scholar]
- Fu, P.; Schoenball, M.; Ajo-Franklin, J.B.; Chai, C.; Maceira, M.; Morris, J.P.; Wu, H.; Knox, H.; Schwering, P.C.; White, M.D.; et al. Close Observation of Hydraulic Fracturing at EGS Collab Experiment 1: Fracture Trajectory, Microseismic Interpretations, and the Role of Natural Fractures. J. Geophys. Res. Solid Earth 2021, 126, e2020JB020840. [Google Scholar] [CrossRef]
- Wu, H.; Fu, P.; Frone, Z.; White, M.D.; Ajo-Franklin, J.B.; Morris, J.P.; Knox, H.A.; Schwering, P.C.; Strickland, C.E.; Roberts, B.Q.; et al. Modeling heat transport processes in enhanced geothermal systems: A validation study from EGS Collab Experiment 1. Geothermics 2021, 97, 102254. [Google Scholar] [CrossRef]
- Wu, H.; Fu, P.; Morris, J.P.; Mattson, E.D.; Neupane, G.; Smith, M.M.; Hawkins, A.J.; Zhang, Y.; Kneafsey, T. Characterization of flow and transport in a fracture network at the EGS Collab field experiment through stochastic modeling of tracer recovery. J. Hydrol. 2021, 593, 125888. [Google Scholar] [CrossRef]
- Gupta, I.; Rai, C.; Devegowda, D.; Sondergeld, C.H. Fracture hits in unconventional reservoirs: A critical review. SPE J. 2021, 26, 412–434. [Google Scholar] [CrossRef]
- Gao, P.K.; Li, G.Q.; Tian, H.M.; Wang, Y.S.; Sun, H.W.; Ma, T. Differences in microbial community composition between injection and production water samples of water flooding petroleum reservoirs. Biogeosciences 2015, 12, 3403–3414. [Google Scholar] [CrossRef]
- Mouser, P.J.; Rizzo, D.M.; Druschel, G.K.; Morales, S.E.; Hayden, N.; O’Grady, P.; Stevens, L. Enhanced detection of groundwater contamination from a leaking waste disposal site by microbial community profiles. Water Resour. Res. 2010, 46, 2010wr009459. [Google Scholar] [CrossRef]
- McElhinney, J.; Catacutan, M.K.; Mawart, A.; Hasan, A.; Dias, J. Interfacing Machine Learning and Microbial Omics: A Promising Means to Address Environmental Challenges. Front. Microbiol. 2022, 13, 851450. [Google Scholar] [CrossRef]
- Hicks, N.; Vik, U.; Taylor, P.; Ladoukakis, E.; Park, J.; Kolisis, F.; Jakobsen, K.S. Using Prokaryotes for Carbon Capture Storage. Trends Biotechnol. 2017, 35, 22–32. [Google Scholar] [CrossRef]
- Chiţu, A.G.; Zijp, M.H.; Zwaan, J. A novel exploration technique using the microbial fingerprint of shallow sediment to detect hydrocarbon microseepage and predict hydrocarbon charge—An Argentinian case study. Interpretation 2022, 10, SB77–SB92. [Google Scholar] [CrossRef]
- Zijp, M.; Mallinson, T.; Zwaan, J.; Chitu, A.; David, P. Eagle Ford and Bakken productivity prediction using soil microbial fingerprinting and machine learning. In Proceedings of the SPE/AAPG/SEG Unconventional Resources Technology Conference, Online, 16–18 November 2021. [Google Scholar]
- Te Stroet, C.; Zwaan, J.; de Jager, G.; Montijn, R.; Schuren, F. Predicting sweet spots in shale plays by DNA fingerprinting and machine learning. In Proceedings of the SPE/AAPG/SEG Unconventional Resources Technology Conference, Austin, TX, USA, 24–26 July 2017. [Google Scholar]
- Graziani, S.; Beaubien, S.E.; Ciotoli, G.; Bigi, S. Development and testing of a rapid, sensitive, high-resolution tool to improve mapping of CO2 leakage at the ground surface. Appl. Geochem. 2022, 145, 105424. [Google Scholar] [CrossRef]
- Fernández-Montiel, I.; Pedescoll, A.; Bécares, E. Microbial communities in a range of carbon dioxide fluxes from a natural volcanic vent in Campo de Calatrava, Spain. Int. J. Greenh. Gas Control 2016, 50, 70–79. [Google Scholar] [CrossRef]
- Sáenz de Miera, L.E.; Arroyo, P.; de Luis Calabuig, E.; Falagán, J.; Ansola, G. High-throughput sequencing of 16S RNA genes of soil bacterial communities from a naturally occurring CO2 gas vent. Int. J. Greenh. Gas Control 2014, 29, 176–184. [Google Scholar] [CrossRef]
- Gulliver, D.M.; Lowry, G.V.; Gregory, K.B. Effect of CO2(aq) Exposure on a Freshwater Aquifer Microbial Community from Simulated Geologic Carbon Storage Leakage. Environ. Sci. Technol. Lett. 2014, 1, 479–483. [Google Scholar] [CrossRef]
- Gulliver, D.M.; Lowry, G.V.; Gregory, K.B. Comparative study of effects of CO2 concentration and pH on microbial communities from a saline aquifer, a depleted oil reservoir, and a freshwater aquifer. Environ. Eng. Sci. 2016, 33, 806–816. [Google Scholar] [CrossRef]
- Tait, K.; Stahl, H.; Taylor, P.; Widdicombe, S. Rapid response of the active microbial community to CO2 exposure from a controlled sub-seabed CO2 leak in Ardmucknish Bay (Oban, Scotland). Int. J. Greenh. Gas Control 2015, 38, 171–181. [Google Scholar] [CrossRef]
- Fernández-Montiel, I.; Touceda, M.; Pedescoll, A.; Gabilondo, R.; Prieto-Fernández, A.; Bécares, E. Short-term effects of simulated below-ground carbon dioxide leakage on a soil microbial community. Int. J. Greenh. Gas Control 2015, 36, 51–59. [Google Scholar] [CrossRef]
- Chen, F.; Yang, Y.; Ma, Y.; Hou, H.; Zhang, S.; Ma, J. Effects of CO2 leakage on soil bacterial communities from simulated CO2-EOR areas. Environ. Sci. Process. Impacts 2016, 18, 547–554. [Google Scholar] [CrossRef]
- Trautz, R.C.; Pugh, J.D.; Varadharajan, C.; Zheng, L.; Bianchi, M.; Nico, P.S.; Spycher, N.F.; Newell, D.L.; Esposito, R.A.; Wu, Y.; et al. Effect of dissolved CO2 on a shallow groundwater system: A controlled release field experiment. Env. Sci Technol 2013, 47, 298–305. [Google Scholar] [CrossRef]
- Taylor, P.; Stahl, H.; Vardy, M.E.; Bull, J.M.; Akhurst, M.; Hauton, C.; James, R.H.; Lichtschlag, A.; Long, D.; Aleynik, D.; et al. A novel sub-seabed CO2 release experiment informing monitoring and impact assessment for geological carbon storage. Int. J. Greenh. Gas Control 2015, 38, 3–17. [Google Scholar] [CrossRef]
- Ma, J.; Zhang, W.; Zhang, S.; Zhu, Q.; Feng, Q.; Chen, F. Short-term effects of CO2 leakage on the soil bacterial community in a simulated gas leakage scenario. PeerJ 2017, 5, e4024. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Chen, Y. Effects of elevated carbon dioxide on environmental microbes and its mechanisms: A review. Sci. Total Environ. 2019, 655, 865–879. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Foppen, J.W.; Banerjee, A.; Sawssen, S.; Bachhar, N.; Peddis, D.; Bandyopadhyay, S. Magnetic Nanoparticles to Unique DNA Tracers: Effect of Functionalization on Physico-chemical Properties. Nanoscale Res. Lett. 2021, 16, 24. [Google Scholar] [CrossRef] [PubMed]
| Authors | Year (Country a) | Type of Particle Carrier | Average Particle Size | Solvent for DNA Release | Medium | Scale b | Details |
|---|---|---|---|---|---|---|---|
| Sharma, A.N., et al. | 2012 [70] (USA) | Polylactic acid (PLA) microspheres | 400−500 nm | Chloroform | 1) Column packed with quartz sand; 2) asphalt surface with manually established stream; 3) a stream. | 1) Lab, L = 30.5 cm, D = 2.54 cm; 2) field, 1 m; 3) field, 10 m. | Proof-of-concept experiments; particles were synthesized using the double emulsion method; iron oxide nanoparticles were embedded into the PLA microspheres for magnetic separation. The DNA microtracers exhibited advective–dispersive behavior, with recovery rates similar to those of a conservative tracer. |
| Dahlke, H.E., et al. | 2015 [26] (Sweden) | PLA microspheres | NM | Methylene chloride | Subglacial drainage system. | Field, 1 km | Six DNA microtracers were injected into moulins/crevasses, and breakthrough curves were measured along proglacial streams. Recovery: 99% for uranine and 15–66% for DNA microtracers. |
| Alaskar, M., et al. | 2015 [63] (USA) | Silica (SiO2) nanoparticles | 175 nm | Buffered hydrofluoric acid (HF) | Column packed with quartz sand (150–180 μm). | Lab, L = 30.48 cm, D = 0.46 cm. | Concept proposal in the context of geothermal reservoir characterization; a high-temperature (up to 180 °C) column test on the DNA nanotracer showed particle dissolution and aggregation in the effluent. |
| Zhang, Y., et al. | 2015 [29] (USA) | Silica nanoparticles | 160 nm | Buffered HF | Column packed with quartz sand. | Lab, <1 m. | Concept proposal; DNA nanotracers were synthesized according to the literature, with agglomeration observed. A preliminary column test showed successful but limited particle breakthrough. |
| 2016 [64] (USA) | Silica nanoparticles | 160 nm | Buffered HF | Column packed with quartz sand. | Lab, <1 m. | Injection test on a self-synthesized DNA nanotracer. Particles were retained in the sand column after transport; DNA could not be quantified in the effluent particles. | |
| Mikutis, G., et al. | 2018 [65] (Switzerland) | Silica nanoparticles | 159 nm, 410 nm, 848 nm | Buffered HF | 1) Column packed with sand (0.20–0.63 mm); 2) unconsolidated aquifer with sandy gravels. | 1) Lab, L = 29.6 cm, D = 6.3 cm; 2) field, 1 m (<10 m deep). | DNA nanotracers had earlier and sharper breakthrough than uranine in all cases; 159, 410, and 848 nm particles were compared in the column test; transport velocity increased, and recovery (39.3–85.9%) decreased with increasing particle size. The 159 nm nanotracer was tested in the field, with peak concentrations of 10−4–10−7 that of the injection concentration; recovery NM. |
| Kong, X., et al. | 2018 [25] (Switzerland) | Silica nanoparticles | ~150 nm | Buffered HF | Unconsolidated aquifer with sandy gravels. | Field, 1 m (<10 m deep). | Multilevel DNA nanotracer injection tests; three unique DNA nanotracers were injected, one of which successfully generated breakthrough and recovery data (recovery was larger than that of the sulforhodamine B reference tracer); tomographic inversion yielded a hydraulic conductivity tomogram. |
| McNew C.P., et al. | 2018 [73] (USA) | Poly(lactic-co-glycolic acid) (PLGA) microspheres | ~300 nm | Dichloromethane | 1) A hillslope (soil); 2) batch test with deionized, NaCl, and stream water. | 1) Field, 1 m (0–65 cm deep); 2) lab. | Protocols were optimized for particle preparation and quantification; field tests demonstrated lack of signal interference among unique DNA tracers or with background environmental DNA; batch tests showed temperature-dependent biodegradability of the PLGA particles. |
| Kittilä, A., et al. | 2019 [66] (Switzerland) | Silica nanoparticles | 166 nm | Buffered HF | Fractured crystalline rock. | Field, 10 m (400–500 m overburden). | Seven unique DNA nanotracers were tested, along with solute dye tracers; breakthrough curves were obtained and analyzed by temporal moments. DNA nanotracers had smaller mean residence times, lower mass recoveries, less dispersion, and lower swept volume than the dye tracers and tended to settle, especially at low flow velocities. |
| Wang, C., et al. | 2019 [71] (USA) | PLA microspheres | ~900 nm | Dichloromethane | Sloped lysimeter packed with crushed basaltic tephra (0.17% <2 μm, 13.5% 2–50 μm, 86.5% > 50 μm) | Lab, L = 2 m, W = 0.5 m, H = 1 m. | Four unique DNA microtracers were injected sequentially and in parallel. Complex interplay was observed between fast transport (e.g., size exclusion effect) and slow retention–release (e.g., straining and adsorption) mechanisms; recovery: 97.78% for deuterium and 1–2% for DNA microtracers. |
| Georgakakos C.B., et al. | 2019 [74] (USA) | PLGA microspheres | NM | Dichloromethane | 1) Silt loam underlain by clay; 2) gravel loam and stream. | 1) Field, 10 m; 2) field, 100 m. | Proof-of-concept experiments confirming the applicability of DNA microtracers in septic systems at both the local scale and watershed scale. |
| Pang, L., et al. | 2020 [52] (New Zealand) | Alginate-chitosan microparticles | 312 nm | Chelex® treatment and 5 min heating at 95°C | 1) Surface stream; 2) alluvial gravel aquifer; 3) soil lysimeter. | 1) Field, 1 km; 2) field, 10 m (16-m below ground); 3) lab, L = 0.7 m, D = 0.5 m. | DNA microtracers were compared with free DNA in flow experiments; encapsulated DNA had greater recoveries than free DNA in surface water experiments but lower recoveries in the gravel aquifer. |
| Liao, R., et al. | 2020 [72] (China) | PLA microspheres | 246 μm | NM | Open channel with tap water. | Lab, L = 5 m, W = 0.6 m, H = 1 m. | Magnetite nanoparticles attached with plasmid DNA (3500–6500 bp) were encapsulated; microspheres were buoyant on the water surface; DNA was detected at the outlet. |
| Tang, Y., et al. | 2021 [67] (Netherlands) | Silica nanoparticles | 237–299 nm | Buffered HF | Open channel with six types of environmental water. | Lab, L = 20 cm, W = 10 cm, H = 3 cm. | DNA nanotracers had similar hydrodynamic dispersion compared with NaCl under surface water conditions but more scattered breakthrough curve datapoints. |
| Chakraborty, S., et al. | 2022 [68] (Netherlands) | Silica nanoparticles | 280–300 nm | Buffered HF | Column packed with acid-washed quartz sand (~400 μm). | Lab, L = 15 cm, D = 2.1 cm. | A DNA nanotracer with varying particle concentrations was flowed through a saturated column and modeled; particle retention depended on the injection concentration; the kinetic attachment rate coefficient decreased with increasing injection concentration. |
| Kianfar, B., et al. | 2022 [69] (Netherlands) | Silica nanoparticles | ~270 nm | Buffered HF | Column packed with acid-washed quartz sand (1–1.4 mm) or silver sand (0.5–0.63 mm) | Lab, L = 6.5 cm, D = 2.5 cm. | The effect of solution chemistry, flow velocity, and sand grain size on DNA nanotracer transport was investigated. Higher ionic strength enhanced particle sticking after collision with sand grains; the DNA nanotracer was most applicable for coarse-grained aquifers with high flow rates. |
| Authors | Year, Reference (Country a) | Reservoir Type | Microbiome Type | Scale b | Primary Data Interpretation Method | Proposed Utility | Validation of Proposed Utility |
|---|---|---|---|---|---|---|---|
| Lascelles, P., et al. | 2017 [33] (USA) | Hydrocarbon (unconventional) | Drill cuttings and produced oil. | Field, 1 km. | “Percentage contribution of DNA signature”. | Effective drainage height; well-to-well communication; production contribution by fracturing stage. | Blinded analyses; well log data. |
| Silva, J., et al. | 2018 [39] (USA) | Hydrocarbon (unconventional) | Drill cuttings and produced hydrocarbon/water. | Field, 33 wells in the Delaware Basin. | Composition; ordination c; Bayesian mixture modeling. | Production contribution by formation. | NM. |
| Percak-Dennett, E., et al. | 2019 [87] (USA) | Hydrocarbon (unconventional) | Drill cuttings and produced fluids. | Field, >1 km (to a depth of >3 km). | Composition; “percentage contribution”; Bayesian mixture modeling. | Production contribution by formation. | Blinded analyses; tracer, geochemistry, and microseismic data. |
| Ursell, L., et al. | 2019 [12] (USA) | Hydrocarbon (unconventional) | Drill cuttings and produced fluids. | Field, >1 km (to a depth of >3 km). | Bayesian mixture modeling. | Contribution of fracturing fluid to produced fluids; effective drainage height; production contribution by lateral stage. | Oil production rate; oil-based chemical tracers. |
| Zhang, Y., et al. | 2020 [10] (USA) | Groundwater | Groundwater (0.2 and 0.02 μm d). | Field, 10 m (1478 m deep). | Composition; ordination. | Indicator for the natural connectivities among artesian wells. | Fracture model informed by core log and sewer camera footages. |
| 2022 [31] (USA) | Groundwater | Groundwater (0.2 μm). | Field, 10 m (1478 m deep). | Composition; ordination; ecological modeling. | Monitoring of changes in a permeable fracture network over time, e.g., fracture opening/closure. | Data-informed fracture network model; conservative tracer and flow rate data; ecological modeling results. | |
| 2022 [30] (USA) | Groundwater | Groundwater (0.2 μm). | Field, 10 m (1478 m deep). | Percentage of injectate ASVs detected in each producer. | Indicator of relative connectivities across several producers with an injector. | Conservative tracer data (recoveries). | |
| Schlecht, M., et al. | 2020 [38] (USA) | Hydrocarbon (unconventional) | Drill cuttings, cores, injected fluids, and produced fluids. | Field, >1 km (to a depth of >3 km). | Composition; Bayesian mixture modeling. | Effective drainage height; well-to-well communication; fracture height and half length; production contribution by formation. | Petrophysical logs (gamma ray, effective porosity, and water saturation); geochemical data. |
| Sawadogo, J., et al. | 2020 [88] (USA) | Hydrocarbon (unconventional) | Drill cuttings and produced fluids. | Field, >1 km (to a depth of >2 km). | Composition; Bayesian mixture modeling. | Production contribution by formation. | Oil and water tracer data. |
| 2020 [35] (USA) | Hydrocarbon (conventional) | Injected fluids (for waterflooding) and produced fluids. | Field, 1 km (to a depth of 1 km). | Composition; ordination; abundance of injector DNA markers in the produced fluids. | Injector–producer communication; waterflood monitoring. | Interference test results and historical field observations. | |
| Merino, N., et al. | 2022 [34] (USA) | Groundwater | Groundwater (0.2 μm). | Field, 100 km. | Ordination; network analysis; ecological modeling. | Tracing of regional-scale groundwater flow. | Prior knowledge of regional groundwater flowpaths; ecological modeling results. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Huang, T. DNA-Based Tracers for the Characterization of Hydrogeological Systems—Recent Advances and New Frontiers. Water 2022, 14, 3545. https://doi.org/10.3390/w14213545
Zhang Y, Huang T. DNA-Based Tracers for the Characterization of Hydrogeological Systems—Recent Advances and New Frontiers. Water. 2022; 14(21):3545. https://doi.org/10.3390/w14213545
Chicago/Turabian StyleZhang, Yuran, and Tianming Huang. 2022. "DNA-Based Tracers for the Characterization of Hydrogeological Systems—Recent Advances and New Frontiers" Water 14, no. 21: 3545. https://doi.org/10.3390/w14213545
APA StyleZhang, Y., & Huang, T. (2022). DNA-Based Tracers for the Characterization of Hydrogeological Systems—Recent Advances and New Frontiers. Water, 14(21), 3545. https://doi.org/10.3390/w14213545







