Denitrification-Potential Evaluation and Nitrate-Removal-Pathway Analysis of Aerobic Denitrifier Strain Marinobacter hydrocarbonoclasticus RAD-2
Abstract
:1. Introduction
2. Materials and Methods
2.1. Culture Media
2.2. Bacteria Isolation, Screening, and Identification
2.3. Nitrogen-Removal Performance Evaluation
2.4. RT-qPCR Analysis
2.5. Analytical Methods
3. Results
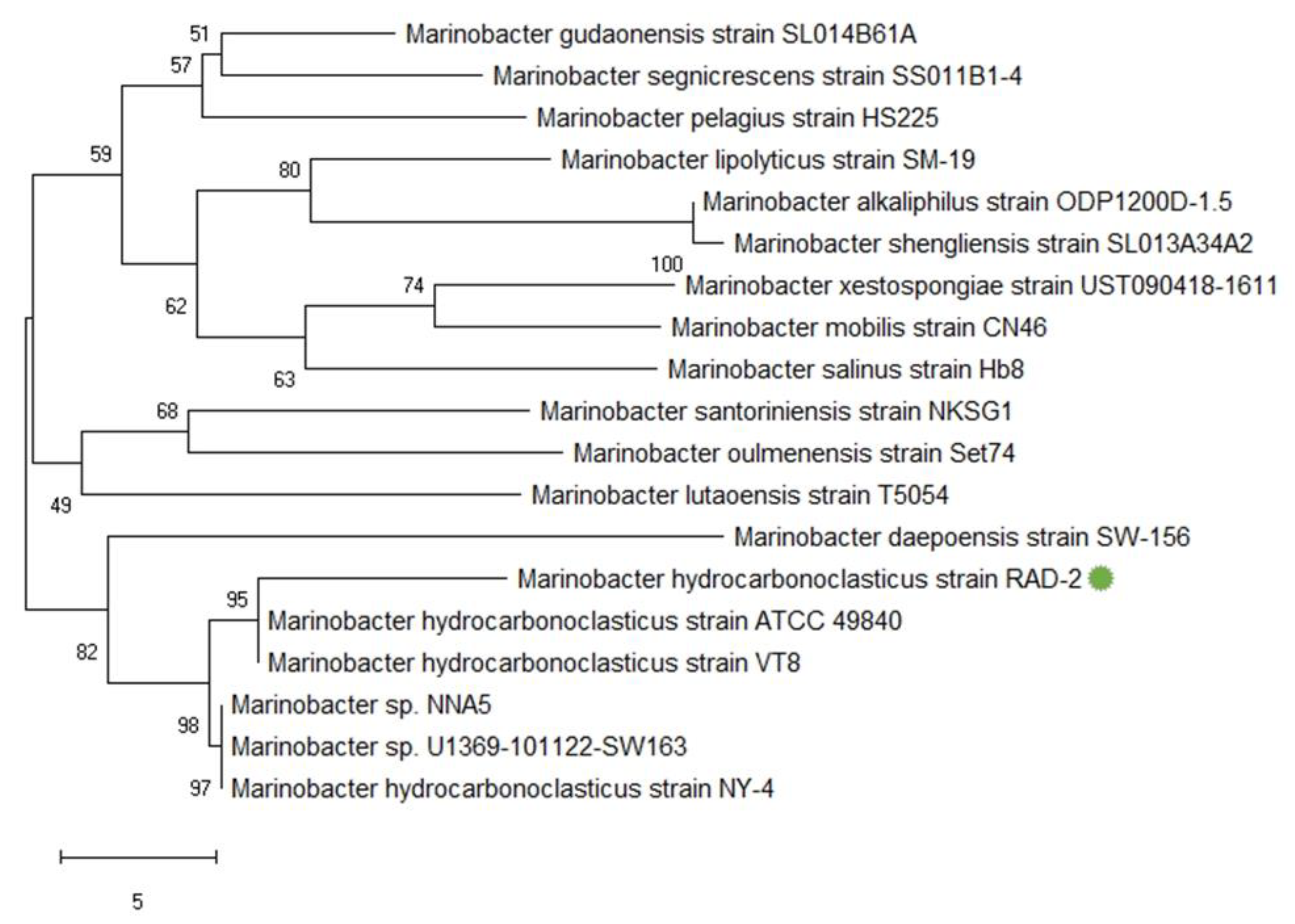
3.1. Characteristics and Identification
3.2. Nitrogen Removal Performance Evaluation
3.2.1. Aerobic Nitrogen-Removal Ability of Strain RAD-2
3.2.2. Single-Factor Experiments of Strain RAD-2
3.3. Expression of Denitrifying Genes by RT-qPCR Analysis
4. Discussion
4.1. Characteristics and Identification
4.2. Nitrogen-Removal Performance Evaluation
4.3. Aerobic Denitrification Pathways Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Martins, C.I.M.; Eding, E.H.; Verdegem, M.C.J.; Heinsbroek, L.T.N.; Schneider, O.; Blancheton, J.P.; d’Orbcastel, E.R.; Verreth, J.A.J. New developments in recirculating aquaculture systems in europe: A perspective on environmental sustainability. Aquac. Eng. 2010, 43, 83–93. [Google Scholar] [CrossRef]
- Gutierrez-Wing, M.T.; Malone, R.F. Biological filters in aquaculture: Trends and research directions for freshwater and marine applications. Aquac. Eng. 2006, 34, 163–171. [Google Scholar] [CrossRef]
- Van Bussel, C.G.J.; Schroeder, J.P.; Wuertz, S.; Schulz, C. The chronic effect of nitrate on production performance and health status of juvenile turbot (Psetta maxima). Aquaculture 2012, 326–329, 163–167. [Google Scholar] [CrossRef]
- Van Rijn, J.; Tal, Y.; Schreier, H.J. Denitrification in recirculating systems: Theory and applications. Aquac. Eng. 2006, 34, 364–376. [Google Scholar] [CrossRef]
- Kraft, B.; Tegetmeyer, H.E.; Sharma, R.; Klotz, M.G.; Ferdelman, T.G.; Hettich, R.L.; Geelhoed, J.S.; Strous, M. The environmental controls that govern the end product of bacterial nitrate respiration. Science 2014, 345, 676–679. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chu, L. Biological nitrate removal from water and wastewater by solid-phase denitrification process. Biotechnol. Adv. 2016, 34, 1103–1112. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Yang, L.; Wang, J. Denitrification using PBS as carbon source and biofilm support in a packed-bed bioreactor. Environ. Sci. Pollut. Res. 2013, 20, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Zhou, Y.; Wang, J. Comparison of denitrification performance and microbial diversity using starch/polylactic acid blends and ethanol as electron donor for nitrate removal. Bioresour. Technol. 2013, 131, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Chai, X. Effect of weight ratios of PHBV/PLA polymer blends on nitrate removal efficiency and microbial community during solid-phase denitrification. Int. Biodeterior. Biodegrad. 2017, 116, 175–183. [Google Scholar] [CrossRef]
- Zhu, S.M.; Deng, Y.L.; Ruan, Y.J.; Guo, X.S.; Shi, M.M.; Shen, J.Z. Biological denitrification using poly (butylene succinate) as carbon source and biofilm carrier for recirculating aquaculture system effluent treatment. Bioresour. Technol. 2015, 192, 603–610. [Google Scholar] [CrossRef] [PubMed]
- Qiu, T.; Liu, L.; Gao, M.; Zhang, L.; Tursun, H.; Wang, X. Effects of solid-phase denitrification on the nitrate removal and bacterial community structure in recirculating aquaculture system. Biodegradation 2016, 27, 165–178. [Google Scholar] [CrossRef] [PubMed]
- Feng, L.; Chen, K.; Han, D.; Zhao, J.; Lu, Y.; Yang, G.; Mu, J.; Zhao, X. Comparison of nitrogen removal and microbial properties in solid-phase denitrification systems for water purification with various pretreated lignocellulosic carriers. Bioresour. Technol. 2017, 224, 236–245. [Google Scholar] [CrossRef] [PubMed]
- Zumft, W.G. Cell biology and molecular basis of denitrification. Microbiol. Mol. Biol. Rev. 1997, 61, 533–616. [Google Scholar] [PubMed]
- Körner, H.; Zumft, W.G. Expression of denitrification enzymes in response to the dissolved oxygen level and respiratory substrate in continuous culture of Pseudomonas stutzeri. Appl. Environ. Microbiol. 1989, 55, 1670–1676. [Google Scholar] [PubMed]
- Giblin, A.E.; Tobias, C.R.; Song, B.; Weston, N.; Banta, G.T.; Rivera-Monroy, V.H. The importance of dissimilatory nitrate reduction to ammonium (DNRA) in the nitrogen cycle of coastal ecosystems. Oceanography 2013, 26, 124–131. [Google Scholar] [CrossRef]
- Chu, L.; Wang, J. Denitrification performance and biofilm characteristics using biodegradable polymers PCL as carriers and carbon source. Chemosphere 2013, 91, 1310–1316. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Wang, J. Biological denitrification using cross-linked starch/PCL blends as solid carbon source and biofilm carrier. Bioresour. Technol. 2011, 102, 8835–8838. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Y.J.; Deng, Y.L.; Guo, X.S.; Timmons, M.B.; Lu, H.F.; Han, Z.Y.; Ye, Z.Y.; Shi, M.M.; Zhu, S.M. Simultaneous ammonia and nitrate removal in an airlift reactor using poly (butylene succinate) as carbon source and biofilm carrier. Bioresour. Technol. 2016, 216, 1004–1013. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.L.; Ruan, Y.J.; Zhu, S.M.; Guo, X.S.; Han, Z.Y.; Ye, Z.Y.; Liu, G.; Shi, M.M. The impact of DO and salinity on microbial community in poly (butylene succinate) denitrification reactors for recirculating aquaculture system wastewater treatment. AMB Express 2017, 7, 113. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ai, G.M.; Miao, L.L.; Liu, Z.P. Marinobacter strain NNA5, a newly isolated and highly efficient aerobic denitrifier with zero N2O emission. Bioresour. Technol. 2016, 206, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Sun, X.; Fan, Y.; Qiu, T.; Gao, M.; Wang, X. Heterotrophic nitrification and aerobic denitrification by diaphorobacter polyhydroxybutyrativorans SL-205 using poly (3-hydroxybutyrate-co-3-hydroxyvalerate) as the sole carbon source. Bioresour. Technol. 2017, 241, 500–507. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Strous, M. Denitrification and aerobic respiration, hybrid electron transport chains and co-evolution. Biochim. Biophys. Acta Bioenergy 2013, 1827, 136–144. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; He, D.; Ma, T.; Chen, Q.; Liu, S.; Ahmad, M.; Gui, M.; Ni, J. Reducing NO and N2O emission during aerobic denitrification by newly isolated Pseudomonas Stutzeri PCN-1. Bioresour. Technol. 2014, 162, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Robertson, L.A.; Kuenen, J.G. Aerobic denitrification: A controversy revived. Arch. Microbiol. 1984, 139, 351–354. [Google Scholar] [CrossRef]
- Zheng, H.Y.; Liu, Y.; Gao, X.Y.; Ai, G.M.; Miao, L.L.; Liu, Z.P. Characterization of a marine origin aerobic nitrifying-denitrifying bacterium. J. Biosci. Bioeng. 2012, 114, 33–37. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.; Wang, Y.; Liu, H.; Xi, C.; Song, L. A novel heterotrophic nitrifying and aerobic denitrifying bacterium, Zobellella taiwanensis DN-7, can remove high-strength ammonium. Appl. Microbiol. Biotechnol. 2016, 100, 4219–4229. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.; Zhang, Y.; Zhou, J.; Chen, M.; Wang, X. Biological removal of nitrate and ammonium under aerobic atmosphere by Paracoccus versutus LYM. Bioresour. Technol. 2013, 148, 144–148. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Sun, X.; Wang, X.; Qiu, T.; Gao, M.; Sun, Y.; Cheng, S.; Zhang, Q. Bioaugmentation with Diaphorobacter polyhydroxybutyrativorans to enhance nitrate removal in a poly (3-hydroxybutyrate-co-3-hydroxyvalerate)-supported denitrification reactor. Bioresour. Technol. 2018, 263, 499–507. [Google Scholar] [CrossRef] [PubMed]
- Qiu, T.; Zuo, Z.; Gao, J.; Gao, M.; Han, M.; Sun, L.; Zhang, L.; Wang, X. Diaphorobacter polyhydroxybutyrativorans sp. Nov., a novel poly (3-hydroxybutyrate-co-3-hydroxyvalerate)-degrading bacterium isolated from biofilms. Int. J. Syst. Evol. Microbiol. 2015, 65, 2913–2918. [Google Scholar] [CrossRef] [PubMed]
- Gui, M.; Chen, Q.; Ni, J. Effect of sulfamethoxazole on aerobic denitrification by strain Pseudomonas stutzeri PCN-1. Bioresour. Technol. 2017, 235, 325. [Google Scholar] [CrossRef] [PubMed]
- Clesceri, L.S. Standard Methods for the Examination of Water and Wastewater, 20th ed.; American Public Health Association: Washington, DC, USA, 1998. [Google Scholar]
- Gauthier, M.J.; Lafay, B.; Christen, R.; Fernandez, L.; Acquaviva, M.; Bonin, P.; Bertrand, J.C. Marinobacter hydrocarbonoclasticus gen. nov., sp. nov., a new, extremely halotolerant, hydrocarbon-degrading marine bacterium. Int. J. Syst. Bacteriol. 1992, 42, 568. [Google Scholar] [CrossRef] [PubMed]
- Zenati, B.; Chebbi, A.; Badis, A.; Eddouaouda, K.; Boutoumi, H.; El Hattab, M.; Hentati, D.; Chelbi, M.; Sayadi, S.; Chamkha, M.; et al. A non-toxic microbial surfactant from Marinobacter hydrocarbonoclasticus sdk644 for crude oil solubilization enhancement. Ecotoxicol. Environ. Saf. 2018, 154, 100–107. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Zi, X.; Wang, X.; Zhang, X.; Gao, H.; Hu, N. Marinobacter hydrocarbonoclasticus NY-4, a novel denitrifying, moderately halophilic marine bacterium. SpringerPlus 2013, 2, 346. [Google Scholar] [CrossRef] [PubMed]
- Wan, C.; Yang, X.; Lee, D.-J.; Du, M.; Wan, F.; Chen, C. Aerobic denitrification by novel isolated strain using NO2−-N as nitrogen source. Bioresour. Technol. 2011, 102, 7244–7248. [Google Scholar] [CrossRef] [PubMed]
- Qu, D.; Wang, C.; Wang, Y.; Zhou, R.; Ren, H. Heterotrophic nitrification and aerobic denitrification by a novel groundwater origin cold-adapted bacterium at low temperatures. RSC Adv. 2015, 5, 5149–5157. [Google Scholar] [CrossRef]
- Ye, Q.; Li, K.; Li, Z.; Xu, Y.; He, T.; Tang, W.; Xiang, S. Heterotrophic nitrification-aerobic denitrification performance of strain Y-12 under low temperature and high concentration of inorganic nitrogen conditions. Water 2017, 9, 835. [Google Scholar] [CrossRef]
- Xin, X.; Huang, G.; Liu, X.; An, C.; Yao, Y.; Weger, H.; Zhang, P.; Chen, X. Molecular toxicity of triclosan and carbamazepine to green algae Chlorococcum sp.: A single cell view using synchrotron-based Fourier transform infrared spectromicroscopy. Environ. Pollut. 2017, 226, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Xin, X.; Huang, G.; An, C.; Huang, C.; Weger, H.; Zhao, S.; Zhou, Y.; Rosendahl, S. Insights into the Toxicity of Triclosan to Green Microalga Chlorococcum sp. Using Synchrotron-Based Fourier Transform Infrared Spectromicroscopy: Biophysiological Analyses and Roles of Environmental Factors. Environ. Sci. Technol. 2018, 52, 2295–2306. [Google Scholar] [CrossRef] [PubMed]
- Caniani, D.; Esposito, G.; Gori, R.; Mannina, G. Towards a new decision support system for design, management and operation of wastewater treatment plants for the reduction of greenhouse gases emission. Water 2015, 7, 5599–5616. [Google Scholar] [CrossRef] [Green Version]
- Hu, Z.; Lee, J.W.; Chandran, K.; Kim, S.; Khanal, S.K. Nitrous oxide (N2O) emission from aquaculture: A review. Environ. Sci. Technol. 2012, 46, 6470–6480. [Google Scholar] [CrossRef] [PubMed]



| API 20 NE Results | Strain RAD-2 |
|---|---|
| Oxidase test | + |
| Nitrate reduction | + |
| Arginine dihydrolase | - |
| Urease | - |
| β-glucosidase | - |
| Protease | - |
| β-galactosidase | - |
| Assimilation of Glucose | + |
| Arabinose | - |
| Mannose | - |
| Mannitol | - |
| N-acetyl-glucosamine | - |
| Maltose | - |
| Gluconate | + |
| Capric acid | - |
| Adipic acid | - |
| Malic acid | - |
| Citric acid | - |
| Phenylacetic acid | - |
| Factor | Variations | Growth (OD600) | Initial Nitrate Concentration (mg/L) | Final Nitrate Concentration (mg/L) | Removal Efficiency (%) |
|---|---|---|---|---|---|
| C/N Ratio | 2 | 0.32 ± 0.03 | 306.33 ± 0.95 | 203.44 ± 13.05 | 33.59 ± 4.17 |
| 5 | 0.79 ± 0.13 | 305.25 ± 0.65 | 4.06 ± 0.58 | 98.67 ± 0.19 | |
| 10 | 1.34 ± 0.05 | 305.21 ± 0.31 | 6.48 ± 1.93 | 97.88 ± 0.63 | |
| 15 | 0.94 ± 0.18 | 304.02 ± 0.26 | 40.72 ± 5.77 | 86.61 ± 1.90 | |
| 20 | 0.42 ± 0.05 | 304.67 ± 0.45 | 213.90 ± 12.26 | 29.80 ± 3.92 | |
| Temperature (°C) | 5 | 0.012 ± 0.06 | 306.37 ± 0.36 | 305.74 ± 0.40 | 0.21 ± 0.25 |
| 10 | 0.11 ± 0.03 | 305.38 ± 0.35 | 298.75 ± 2.59 | 2.17 ± 0.85 | |
| 15 | 0.24 ± 0.06 | 304.01 ± 0.38 | 274.22 ± 6.48 | 9.79 ± 2.25 | |
| 25 | 1.12 ± 0.12 | 303.29 ± 0.17 | 6.23 ± 3.61 | 97.95 ± 1.19 | |
| 40 | 1.02 ± 0.19 | 304.52 ± 0.37 | 10.18 ± 1.18 | 96.66 ± 0.39 | |
| Rotation Speed (rpm) | 0 | 0.52 ± 0.11 | 304.86 ± 0.50 | 138.99 ± 17.06 | 54.41 ± 5.53 |
| 50 | 0.67 ± 0.05 | 303.62 ± 0.30 | 92.14 ± 11.05 | 69.65 ± 3.61 | |
| 100 | 0.95 ± 0.13 | 305.34 ± 0.21 | 24.38 ± 6.33 | 92.02 ± 2.08 | |
| 150 | 1.25 ± 0.11 | 305.38 ± 0.24 | 7.02 ± 1.51 | 97.70 ± 0.49 | |
| 200 | 1.03 ± 0.07 | 304.43 ± 0.27 | 30.45 ± 8.37 | 89.90 ± 2.75 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kong, D.; Li, W.; Deng, Y.; Ruan, Y.; Chen, G.; Yu, J.; Lin, F. Denitrification-Potential Evaluation and Nitrate-Removal-Pathway Analysis of Aerobic Denitrifier Strain Marinobacter hydrocarbonoclasticus RAD-2. Water 2018, 10, 1298. https://doi.org/10.3390/w10101298
Kong D, Li W, Deng Y, Ruan Y, Chen G, Yu J, Lin F. Denitrification-Potential Evaluation and Nitrate-Removal-Pathway Analysis of Aerobic Denitrifier Strain Marinobacter hydrocarbonoclasticus RAD-2. Water. 2018; 10(10):1298. https://doi.org/10.3390/w10101298
Chicago/Turabian StyleKong, Dedong, Wenbing Li, Yale Deng, Yunjie Ruan, Guangsuo Chen, Jianhai Yu, and Fucheng Lin. 2018. "Denitrification-Potential Evaluation and Nitrate-Removal-Pathway Analysis of Aerobic Denitrifier Strain Marinobacter hydrocarbonoclasticus RAD-2" Water 10, no. 10: 1298. https://doi.org/10.3390/w10101298
APA StyleKong, D., Li, W., Deng, Y., Ruan, Y., Chen, G., Yu, J., & Lin, F. (2018). Denitrification-Potential Evaluation and Nitrate-Removal-Pathway Analysis of Aerobic Denitrifier Strain Marinobacter hydrocarbonoclasticus RAD-2. Water, 10(10), 1298. https://doi.org/10.3390/w10101298








