ADAMTS Proteases: Importance in Animal Reproduction
Abstract
1. Introduction
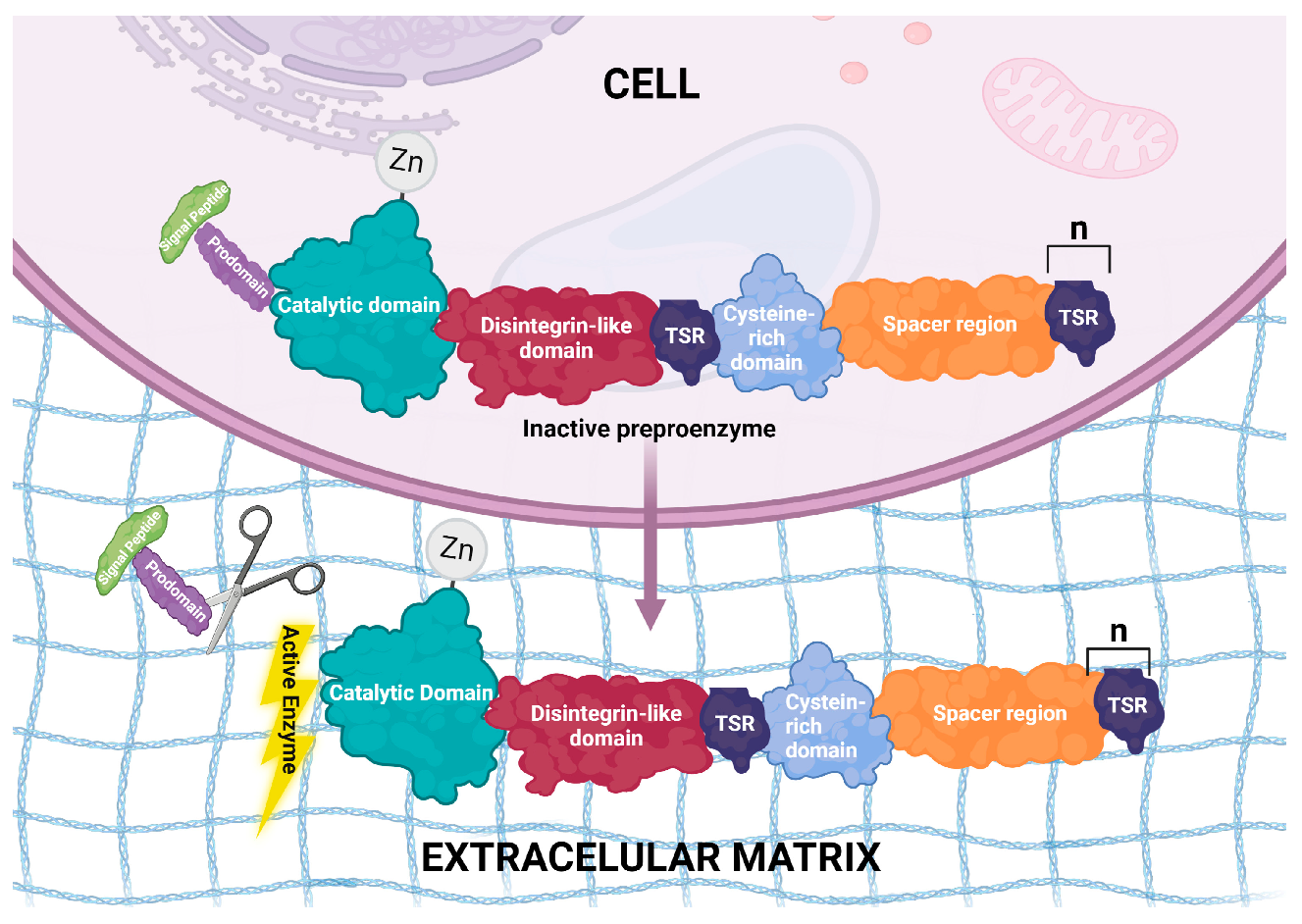
2. Structural Features of ADAMTS Genes
3. ADAMTS and Fertility in Females
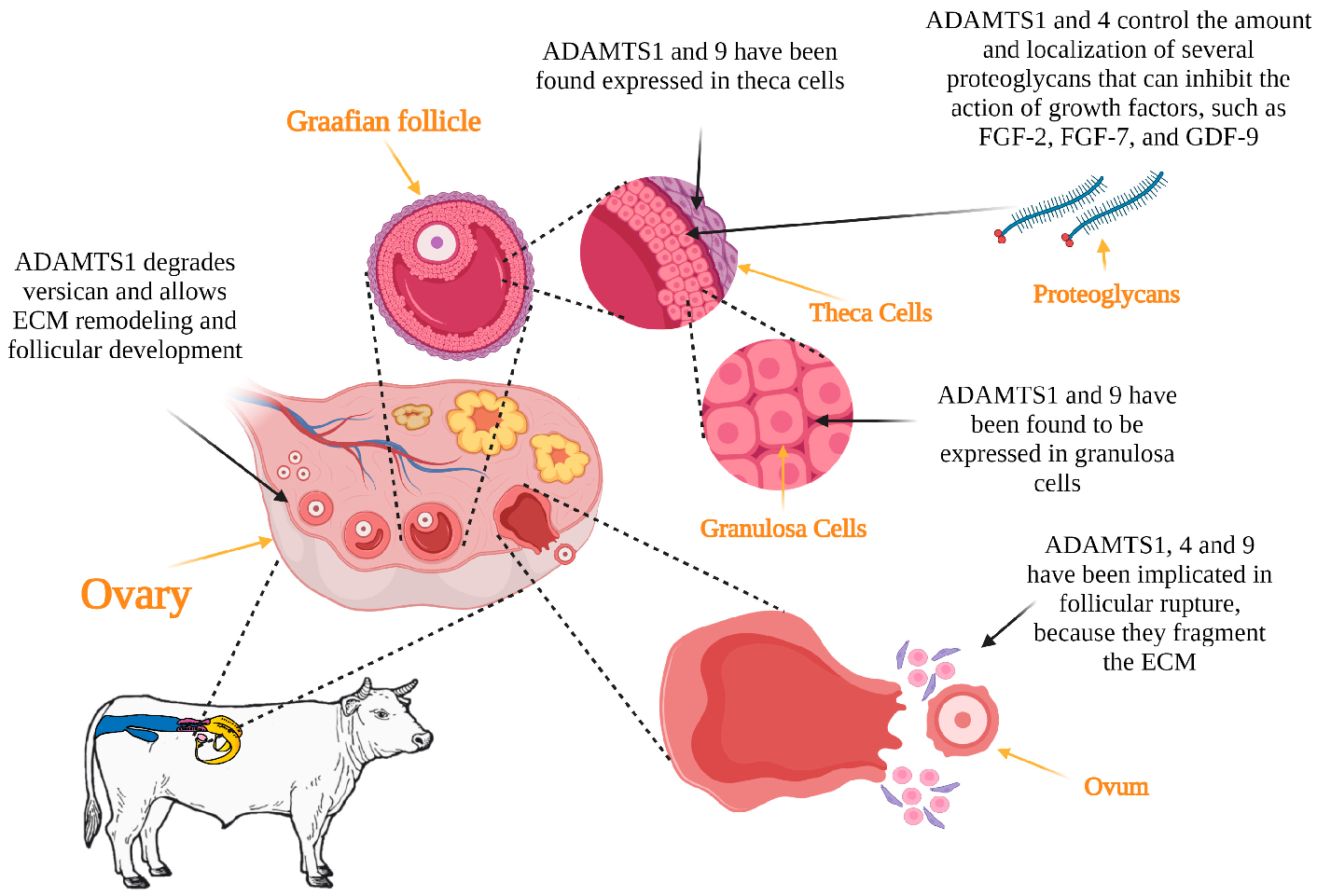
3.1. Folliculogenesis
3.2. Ovulation
3.3. Implantation, Placentation and Parturition
4. ADAMTS and Fertility in Males
4.1. Testicular Development
4.2. Spermatogenesis
4.3. Fertilization
5. Regulation of ADAMTS Genes Involved in Reproduction
6. ADAMTS in Omics Studies
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hill, W.G. Is continued genetic improvement of livestock sustainable? Genetics 2016, 202, 877–881. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.F.; Schnabel, R.D.; Sutovsky, P. Genomics of bull fertility. Animal 2018, 12, 172–183. [Google Scholar] [CrossRef] [PubMed]
- Notter, D. Genetic aspects of reproduction in sheep. Reprod. Domest. Anim. 2008, 43, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Albert, F.W.; Kruglyak, L. The role of regulatory variation in complex traits and disease. Nat. Rev. Genet. 2015, 16, 197–212. [Google Scholar] [CrossRef]
- Demircan, K.; Comertoglu, I.; Akyol, S.; Yigitoglu, B.N.; Sarikaya, E. A new biological marker candidate in female reproductive system diseases: Matrix metalloproteinase with thrombospondin motifs (ADAMTS). J. Turk. Ger. Gynecol. Assoc. 2014, 15, 250–255. [Google Scholar] [CrossRef]
- Russell, D.L.; Brown, H.M.; Dunning, K.R. ADAMTS proteases in fertility. Matrix Biol. 2015, 44–46, 54–63. [Google Scholar] [CrossRef]
- Copp, H.L.; Shortliffe, L.D. Undescended testes and testicular tumors. In Ashcraft’s Pediatric Surgery; Elsevier: Amsterdam, The Netherlands, 2010; pp. 676–686. [Google Scholar]
- Dun, M.D.; Anderson, A.L.; Bromfield, E.G.; Asquith, K.L.; Emmett, B.; McLaughlin, E.A.; Aitken, R.J.; Nixon, B. Investigation of the expression and functional significance of the novel mouse sperm protein, a disintegrin and metalloprotease with thrombospondin type 1 motifs number 10 (ADAMTS10). Int. J. Androl. 2012, 35, 572–589. [Google Scholar] [CrossRef]
- Fata, J.; Ho, A.-V.; Leco, K.; Moorehead, R.; Khokha, R. Cellular turnover and extracellular matrix remodeling in female reproductive tissues: Functions of metalloproteinases and their inhibitors. CMLS 2000, 57, 77–95. [Google Scholar] [CrossRef]
- Blobel, C.P.; Apte, S. ADAMs and ADAMTSs. In Encyclopedia of Respiratory Medicine; Elsevier: Amsterdam, The Netherlands, 2020; pp. 568–573. [Google Scholar]
- Sternlicht, M.D.; Werb, Z. How matrix metalloproteinases regulate cell behavior. Annu. Rev. Cell Dev. Biol. 2001, 17, 463–516. [Google Scholar] [CrossRef]
- Apte, S.S.; Parks, W.C. Metalloproteinases: A parade of functions in matrix biology and an outlook for the future. Matrix Biol. 2015, 44–46, 1–6. [Google Scholar] [CrossRef]
- Dubail, J.; Apte, S.S. Insights on ADAMTS proteases and ADAMTS-like proteins from mammalian genetics. Matrix Biol. 2015, 44, 24–37. [Google Scholar] [CrossRef]
- Menke, D.B.; Koubova, J.; Page, D.C. Sexual differentiation of germ cells in XX mouse gonads occurs in an anterior-to-posterior wave. J. Dev. Biol. 2003, 262, 303–312. [Google Scholar] [CrossRef]
- Doyle, K.M.; Russell, D.L.; Sriraman, V.; Richards, J.S. Coordinate transcription of the ADAMTS-1 gene by luteinizing hormone and progesterone receptor. Mol. Endocrinol. 2004, 18, 2463–2478. [Google Scholar] [CrossRef]
- Richards, J.S.; Hernandez-Gonzalez, I.; Gonzalez-Robayna, I.; Teuling, E.; Lo, Y.; Boerboom, D.; Falender, A.E.; Doyle, K.H.; LeBaron, R.G.; Thompson, V. Regulated expression of ADAMTS family members in follicles and cumulus oocyte complexes: Evidence for specific and redundant patterns during ovulation. Biol. Reprod. 2005, 72, 1241–1255. [Google Scholar] [CrossRef]
- Hurskainen, T.L.; Hirohata, S.; Seldin, M.F.; Apte, S.S. ADAM-TS5, ADAM-TS6, and ADAM-TS7, Novel Members of a New Family of Zinc Metalloproteases. J. Biol. Chem. 1999, 274, 25555–25563. [Google Scholar] [CrossRef]
- Kuno, K.; Kanada, N.; Nakashima, E.; Fujiki, F.; Ichimura, F.; Matsushima, K. Molecular cloning of a gene encoding a new type of metalloproteinase-disintegrin family protein with thrombospondin motifs as an inflammation associated gene. J. Biol. Chem. 1997, 272, 556–562. [Google Scholar] [CrossRef]
- Porter, S.; Clark, I.M.; Kevorkian, L.; Edwards, D.R. The ADAMTS metalloproteinases. Biochem. J. 2005, 386, 15–27. [Google Scholar] [CrossRef]
- Apte, S.S. A disintegrin-like and metalloprotease (reprolysin-type) with thrombospondin type 1 motif (ADAMTS) superfamily: Functions and mechanisms. J. Biol. Chem. 2009, 284, 31493–31497. [Google Scholar] [CrossRef]
- Rose, K.W.; Taye, N.; Karoulias, S.Z.; Hubmacher, D. Regulation of ADAMTS proteases. Front. Mol. Biosci. 2021, 8, 701959. [Google Scholar] [CrossRef]
- Huxley-Jones, J.; Apte, S.S.; Robertson, D.L.; Boot-Handford, R.P. The characterisation of six ADAMTS proteases in the basal chordate Ciona intestinalis provides new insights into the vertebrate ADAMTS family. Int. J. Biochem. Cell Biol. 2005, 37, 1838–1845. [Google Scholar] [CrossRef]
- Longpré, J.-M.; Leduc, R. Identification of prodomain determinants involved in ADAMTS-1 biosynthesis. J. Biol. Chem. 2004, 279, 33237–33245. [Google Scholar] [CrossRef] [PubMed]
- Kelwick, R.; Desanlis, I.; Wheeler, G.N.; Edwards, D.R. The ADAMTS (A Disintegrin and Metalloproteinase with Thrombospondin motifs) family. Genome Biol. 2015, 16, 113. [Google Scholar] [CrossRef] [PubMed]
- Gershon, E.; Dekel, N. Newly identified regulators of ovarian folliculogenesis and ovulation. Int. J. Mol. Sci. 2020, 21, 4565. [Google Scholar] [CrossRef] [PubMed]
- Madan, P.; Bridges, P.J.; Komar, C.M.; Beristain, A.G.; Rajamahendran, R.; Fortune, J.E.; MacCalman, C.D. Expression of messenger RNA for ADAMTS subtypes changes in the periovulatory follicle after the gonadotropin surge and during luteal development and regression in cattle. Biol. Reprod. 2003, 69, 1506–1514. [Google Scholar] [CrossRef]
- Boerboom, D.; Russell, D.L.; Richards, J.S.; Sirois, J. Regulation of transcripts encoding ADAMTS-1 (a disintegrin and metalloproteinase with thrombospondin-like motifs-1) and progesterone receptor by human chorionic gonadotropin in equine preovulatory follicles. J. Mol. Endocrinol. 2003, 31, 473–485. [Google Scholar] [CrossRef]
- Shimada, M.; Nishibori, M.; Yamashita, Y.; Ito, J.; Mori, T.; Richards, J.S. Down-regulated expression of A disintegrin and metalloproteinase with thrombospondin-like repeats-1 by progesterone receptor antagonist is associated with impaired expansion of porcine cumulus-oocyte complexes. Endocrinology 2004, 145, 4603–4614. [Google Scholar] [CrossRef]
- Brown, H.M.; Dunning, K.R.; Robker, R.L.; Pritchard, M.; Russell, D.L. Requirement for ADAMTS-1 in extracellular matrix remodeling during ovarian folliculogenesis and lymphangiogenesis. Dev. Biol. 2006, 300, 699–709. [Google Scholar] [CrossRef]
- Park, P.W.; Reizes, O.; Bernfield, M. Cell surface heparan sulfate proteoglycans: Selective regulators of ligand-receptor encounters. J. Biol. Chem. 2000, 275, 29923–29926. [Google Scholar] [CrossRef]
- Berisha, B.; Sinowatz, F.; Schams, D. Expression and localization of fibroblast growth factor (FGF) family members during the final growth of bovine ovarian follicles. Mol. Reprod. Dev. 2004, 67, 162–171. [Google Scholar] [CrossRef]
- Mishra, S.R.; Thakur, N.; Somal, A.; Parmar, M.S.; Reshma, R.; Rajesh, G.; Yadav, V.P.; Bharti, M.K.; Bharati, J.; Paul, A.; et al. Expression and localization of fibroblast growth factor (FGF) family in buffalo ovarian follicle during different stages of development and modulatory role of FGF2 on steroidogenesis and survival of cultured buffalo granulosa cells. Res. Vet. Sci. 2016, 108, 98–111. [Google Scholar] [CrossRef]
- Matos, M.H.; Lima-Verde, I.B.; Bruno, J.B.; Lopes, C.A.; Martins, F.S.; Santos, K.D.; Rocha, R.M.; Silva, J.R.; Bao, S.N.; Figueiredo, J.R. Follicle stimulating hormone and fibroblast growth factor-2 interact and promote goat primordial follicle development in vitro. Reprod. Fertil. Dev. 2007, 19, 677–684. [Google Scholar] [CrossRef]
- Santos, J.M.; Menezes, V.G.; Barberino, R.S.; Macedo, T.J.; Lins, T.L.; Gouveia, B.B.; Barros, V.R.; Santos, L.P.; Goncalves, R.J.; Matos, M.H. Immunohistochemical localization of fibroblast growth factor-2 in the sheep ovary and its effects on pre-antral follicle apoptosis and development in vitro. Reprod. Domest. Anim. 2014, 49, 522–528. [Google Scholar] [CrossRef]
- Richards, J.S.; Russell, D.L.; Ochsner, S.; Espey, L.L. Ovulation: New dimensions and new regulators of the inflammatory-like response. Annu. Rev. Physiol. 2002, 64, 69–92. [Google Scholar] [CrossRef]
- Shozu, M.; Minami, N.; Yokoyama, H.; Inoue, M.; Kurihara, H.; Matsushima, K.; Kuno, K. ADAMTS-1 is involved in normal follicular development, ovulatory process and organization of the medullary vascular network in the ovary. J. Mol. Endocrinol. 2005, 35, 343–355. [Google Scholar] [CrossRef]
- Russell, D.L.; Ochsner, S.A.; Hsieh, M.; Mulders, S.; Richards, J.S. Hormone-regulated expression and localization of versican in the rodent ovary. Endocrinology 2003, 144, 1020–1031. [Google Scholar] [CrossRef]
- McArthur, M.E.; Irving-Rodgers, H.F.; Byers, S.; Rodgers, R.J. Identification and immunolocalization of decorin, versican, perlecan, nidogen, and chondroitin sulfate proteoglycans in bovine small-antral ovarian follicles. Biol. Reprod. 2000, 63, 913–924. [Google Scholar] [CrossRef] [PubMed]
- Rodgers, R.J.; Irving-Rodgers, H.F. Formation of the ovarian follicular antrum and follicular fluid. Biol. Reprod. 2010, 82, 1021–1029. [Google Scholar] [CrossRef] [PubMed]
- Richards, J.S.; Russell, D.L.; Robker, R.L.; Dajee, M.; Alliston, T.N. Molecular mechanisms of ovulation and luteinization. Mol. Cell. Endocrinol. 1998, 145, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Russell, D.L.; Doyle, K.M.; Ochsner, S.A.; Sandy, J.D.; Richards, J.S. Processing and localization of ADAMTS-1 and proteolytic cleavage of versican during cumulus matrix expansion and ovulation. J. Biol. Chem. 2003, 278, 42330–42339. [Google Scholar] [CrossRef]
- Curry, T.E.; Smith, M.F. Impact of extracellular matrix remodeling on ovulation and the folliculo-luteal transition. Semin. Reprod. Med. 2006, 24, 228–241. [Google Scholar] [CrossRef]
- Mittaz, L.; Russell, D.; Wilson, T.; Brasted, M.; Tkalcevic, J.; Salamonsen, L.A.; Hertzog, P.J.; Pritchard, M.A. Adamts-1 is essential for the development and function of the urogenital system. Biol. Reprod. 2004, 70, 1096–1105. [Google Scholar] [CrossRef]
- Brown, H.M.; Dunning, K.R.; Robker, R.L.; Boerboom, D.; Pritchard, M.; Lane, M.; Russell, D.L. ADAMTS1 cleavage of versican mediates essential structural remodeling of the ovarian follicle and cumulus-oocyte matrix during ovulation in mice. Biol. Reprod. 2010, 83, 549–557. [Google Scholar] [CrossRef]
- Lussier, J.G.; Diouf, M.N.; Lévesque, V.; Sirois, J.; Ndiaye, K. Gene expression profiling of upregulated mRNAs in granulosa cells of bovine ovulatory follicles following stimulation with hCG. Reprod. Biol. Endocrinol. 2017, 15, 88. [Google Scholar] [CrossRef]
- Willis, E.L.; Bridges, P.J.; Fortune, J.E. Progesterone receptor and prostaglandins mediate luteinizing hormone-induced changes in messenger RNAs for ADAMTS proteases in theca cells of bovine periovulatory follicles. Mol. Reprod. Dev. 2017, 84, 55–66. [Google Scholar] [CrossRef]
- Fortune, J.E.; Willis, E.L.; Bridges, P.J.; Yang, C.S. The periovulatory period in cattle: Progesterone, prostaglandins, oxytocin and ADAMTS proteases. Anim. Reprod. 2009, 6, 60–71. [Google Scholar]
- Hernández-Montiel, W.; Collí-Dula, R.C.; Ramón-Ugalde, J.P.; Martínez-Núñez, M.A.; Zamora-Bustillos, R. RNA-seq transcriptome analysis in ovarian tissue of pelibuey breed to explore the regulation of prolificacy. Genes 2019, 10, 358. [Google Scholar] [CrossRef]
- Hu, W.; Tang, J.; Zhang, Z.; Tang, Q.; Yan, Y.; Wang, P.; Wang, X.; Liu, Q.; Guo, X.; Jin, M.; et al. Polymorphisms in the ASMT and ADAMTS1 gene may increase litter size in goats. Vet. Med. Sci. 2020, 6, 775–787. [Google Scholar] [CrossRef]
- Guillomot, M. Changes in extracellular matrix components and cytokeratins in the endometrium during goat implantation. Placenta 1999, 20, 339–345. [Google Scholar] [CrossRef]
- Nardo, L.G.; Li, T.C.; Edwards, R.G. Introduction: Human embryo implantation failure and recurrent miscarriage: Basic science and clinical practice. Reprod. Biomed. Online 2006, 13, 11–12. [Google Scholar] [CrossRef]
- Guzeloglu-Kayisli, O.; Basar, M.; Arici, A. Basic aspects of implantation. Reprod. Biomed. Online 2007, 15, 728–739. [Google Scholar] [CrossRef]
- Das, S.K.; Yano, S.; Wang, J.; Edwards, D.R.; Nagase, H.; Dey, S.K. Expression of matrix metalloproteinases and tissue inhibitors of metalloproteinases in the mouse uterus during the peri-implantation period. Dev. Genet. 1997, 21, 44–54. [Google Scholar] [CrossRef]
- Smith, S.E.; Cullen, W.C.; Godkin, J.D. Ultrastructural morphometric analysis of the uterine epithelium during early pregnancy in the sheep. J. Reprod. Fertil. 1990, 89, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, L.P.; Redmer, D.A. Growth and microvascular development of the uterus during early pregnancy in ewes. Biol. Reprod. 1992, 47, 698–708. [Google Scholar] [CrossRef] [PubMed]
- Shindo, T.; Kurihara, H.; Kuno, K.; Yokoyama, H.; Wada, T.; Kurihara, Y.; Imai, T.; Wang, Y.; Ogata, M.; Nishimatsu, H.; et al. ADAMTS-1: A metalloproteinase-disintegrin essential for normal growth, fertility, and organ morphology and function. J. Clin. Investig. 2000, 105, 1345–1352. [Google Scholar] [CrossRef]
- Mishra, B.; Koshi, K.; Kizaki, K.; Ushizawa, K.; Takahashi, T.; Hosoe, M.; Sato, T.; Ito, A.; Hashizume, K. Expression of ADAMTS1 mRNA in bovine endometrium and placenta during gestation. Domest. Anim. Endocrinol. 2013, 45, 43–48. [Google Scholar] [CrossRef]
- Riemer, R.K.; Heymann, M.A. Regulation of uterine smooth muscle function during gestation. Pediatr. Res. 1998, 44, 615–627. [Google Scholar] [CrossRef]
- Mead, T.J.; Du, Y.; Nelson, C.M.; Gueye, N.A.; Drazba, J.; Dancevic, C.M.; Vankemmelbeke, M.; Buttle, D.J.; Apte, S.S. ADAMTS9-Regulated Pericellular Matrix Dynamics Governs Focal Adhesion-Dependent Smooth Muscle Differentiation. Cell Rep. 2018, 23, 485–498. [Google Scholar] [CrossRef]
- Abdul-Majeed, S.; Mell, B.; Nauli, S.M.; Joe, B. Cryptorchidism and infertility in rats with targeted disruption of the Adamts16 locus. PLoS ONE 2014, 9, e100967. [Google Scholar] [CrossRef]
- Jacobi, C.L.; Rudigier, L.J.; Scholz, H.; Kirschner, K.M. Transcriptional regulation by the Wilms tumor protein, Wt1, suggests a role of the metalloproteinase Adamts16 in murine genitourinary development. J. Biol. Chem. 2013, 288, 18811–18824. [Google Scholar] [CrossRef]
- Sarila, G.; Bao, T.; Abeydeera, S.A.; Li, R.; Mell, B.; Joe, B.; Catubig, A.; Hutson, J. Interplay between collagenase and undescended testes in Adamts16 knockout rats. J. Pediatr. Surg. 2020, 55, 1952–1958. [Google Scholar] [CrossRef]
- Livermore, C.; Warr, N.; Chalon, N.; Siggers, P.; Mianne, J.; Codner, G.; Teboul, L.; Wells, S.; Greenfield, A. Male mice lacking ADAMTS-16 are fertile but exhibit testes of reduced weight. Sci. Rep. 2019, 9, 17195. [Google Scholar] [CrossRef]
- Carré, G.-A.; Couty, I.; Hennequet-Antier, C.; Govoroun, M.S. Gene expression profiling reveals new potential players of gonad differentiation in the chicken embryo. PLoS ONE 2011, 6, e23959. [Google Scholar] [CrossRef]
- Staub, C.; Johnson, L. Review: Spermatogenesis in the bull. Animal 2018, 12, 27–35. [Google Scholar] [CrossRef]
- Gurupriya, V.S.; Roy, S.C.; Javvaji, P.K.; Dhali, A.; Badami, S.; Rahim, F.; Divyashree, B.C.; Panda, A.P. Expression of ADAMTS10 in male reproductive tract of buffaloes (Bubalus bubalis). J. Exp. Biol. Agric. Sci. 2018, 6, 800–807. [Google Scholar] [CrossRef]
- Cornwall, G.A. New insights into epididymal biology and function. Hum. Reprod. Update 2009, 15, 213–227. [Google Scholar] [CrossRef]
- Li, S.W.; Arita, M.; Fertala, A.; Bao, Y.; Kopen, G.C.; Langsjo, T.K.; Hyttinen, M.M.; Helminen, H.J.; Prockop, D.J. Transgenic mice with inactive alleles for procollagen N-proteinase (ADAMTS-2) develop fragile skin and male sterility. Biochem. J. 2001, 355, 271–278. [Google Scholar] [CrossRef]
- González-Barrio, D.; Diezma-Díaz, C.; Gutiérrez-Expósito, D.; Tabanera, E.; Jiménez-Meléndez, A.; Pizarro, M.; González-Huecas, M.; Ferre, I.; Ortega-Mora, L.M.; Álvarez-García, G. Identification of molecular biomarkers associated with disease progression in the testis of bulls infected with Besnoitia besnoiti. Vet. Res. 2021, 52, 106. [Google Scholar] [CrossRef]
- Olias, P.; Schade, B.; Mehlhorn, H. Molecular pathology, taxonomy and epidemiology of Besnoitia species (Protozoa: Sarcocystidae). Infect. Genet. Evol. 2011, 11, 1564–1576. [Google Scholar] [CrossRef]
- Wu, S.; Mipam, T.; Xu, C.; Zhao, W.; Shah, M.A.; Yi, C.; Luo, H.; Cai, X.; Zhong, J. Testis transcriptome profiling identified genes involved in spermatogenic arrest of cattleyak. PLoS ONE 2020, 15, e0229503. [Google Scholar] [CrossRef]
- Le Goff, C.; Cormier-Daire, V. The ADAMTS(L) family and human genetic disorders. Hum. Mol. Genet. 2011, 20, R163–R167. [Google Scholar] [CrossRef]
- Sayasith, K.; Lussier, J.; Sirois, J. Molecular characterization and transcriptional regulation of a disintegrin and metalloproteinase with thrombospondin motif 1 (ADAMTS1) in bovine preovulatory follicles. Endocrinology 2013, 154, 2857–2869. [Google Scholar] [CrossRef] [PubMed]
- Binelli, M.; Scolari, S.C.; Pugliesi, G.; Van Hoeck, V.; Gonella-Diaza, A.M.; Andrade, S.C.; Gasparin, G.R.; Coutinho, L.L. The transcriptome signature of the receptive bovine uterus determined at early gestation. PLoS ONE 2015, 10, e0122874. [Google Scholar] [CrossRef] [PubMed]
- Sammad, A.; Luo, H.; Hu, L.; Zhu, H.; Wang, Y. Transcriptome reveals granulosa cells coping through redox, inflammatory and metabolic mechanisms under acute heat stress. Cells 2022, 11, 1443. [Google Scholar] [CrossRef] [PubMed]
- Gootwine, E. Invited review: Opportunities for genetic improvement toward higher prolificacy in sheep. Small Rumin. Res. 2020, 186, 106090. [Google Scholar] [CrossRef]

| Gene | Species | Chr | Exon No | Size (bp) | AA No | Gene | Species | Chr | Exon No | Size (bp) | AA No |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ADAMTS1 | Mus musculus | 16 | 9 | 9287 | 968 | ADAMTS9 | M. musculus | 6 | 40 | 171,752 | 1931 |
| Bos taurus | 1 | 9 | 8698 | 970 | B. taurus | 22 | 41 | 163,000 | 1937 | ||
| Ovis aries | 1 | 9 | 9452 | 968 | O. aries | 19 | 41 | 162,411 | 1935 | ||
| Equus caballus | 26 | 9 | 9002 | 949 | E. caballus | 16 | 40 | 166,329 | 1936 | ||
| Sus scrofa | 13 | 9 | 8783 | 947 | S. scrofa | 13 | 40 | 175,486 | 2028 | ||
| Gallus gallus | 1 | 9 | 6915 | 923 | G. gallus | 12 | 41 | 67,065 | 1928 | ||
| ADAMTS2 | M. musculus | 11 | 22 | 205,489 | 1213 | ADAMTS10 | M. musculus | 17 | 26 | 30,137 | 1104 |
| B. taurus | 7 | 22 | 245,248 | 1205 | B. taurus | 7 | 26 | 21,940 | 1103 | ||
| O. aries | 5 | 23 | 255,023 | 1206 | O. aries | 5 | 27 | 21,873 | 1103 | ||
| E. caballus | 14 | 23 | 215,470 | 1211 | E. caballus | 7 | 27 | 21,061 | 1191 | ||
| S. scrofa | 2 | 23 | 223,979 | 1208 | S. scrofa | 2 | 24 | 16,972 | 1103 | ||
| G. gallus | 13 | 27 | 236,034 | 1187 | G. gallus | 28 | 25 | 47,963 | 1109 | ||
| ADAMTS4 | M. musculus | 1 | 9 | 12,139 | 833 | ADAMTS12 | M. musculus | 15 | 24 | 284,541 | 1600 |
| B. taurus | 3 | 9 | 8057 | 839 | B. taurus | 20 | 25 | 394,653 | 1607 | ||
| O. aries | 1 | 9 | 8091 | 872 | O. aries | 16 | 24 | 402,782 | 1605 | ||
| E. caballus | 5 | 9 | 7009 | 837 | E. caballus | 21 | 24 | 314,874 | 1598 | ||
| S. scrofa | 4 | 9 | 8885 | 872 | S. scrofa | 16 | 24 | 373,778 | 1599 | ||
| G. gallus | 25 | 9 | 7976 | 662 | G. gallus | Z | 23 | 170,964 | 1580 | ||
| ADAMTS5 | M. musculus | 16 | 9 | 42,969 | 930 | ADAMTS16 | M. musculus | 13 | 24 | 114,040 | 1222 |
| B. taurus | 1 | 9 | 50,445 | 934 | B. taurus | 20 | 23 | 193,779 | 1246 | ||
| O. aries | 1 | 8 | 57,118 | 934 | O. aries | 16 | 23 | 183,599 | 1226 | ||
| E. caballus | 26 | 9 | 49,839 | 929 | E. caballus | 21 | 23 | 154,339 | 1249 | ||
| S. scrofa | 13 | 10 | 60,092 | 929 | S. scrofa | 16 | 24 | 164,333 | 1238 | ||
| G. gallus | 1 | 8 | 46,807 | 877 | G. gallus | - | - | - | - | ||
| ADAMTS6 | M. musculus | 13 | 28 | 210,178 | 1117 | ADAMTS18 | M. musculus | 8 | 24 | 152,221 | 1219 |
| B. taurus | 20 | 26 | 286,175 | 1117 | B. taurus | 18 | 24 | 145,453 | 1223 | ||
| O. aries | 16 | 28 | 289,952 | 1117 | O. aries | 14 | 21 | 147,254 | 1050 | ||
| E. caballus | 21 | 25 | 358,367 | 1117 | E. caballus | 3 | 23 | 125,706 | 1199 | ||
| S. scrofa | 16 | 24 | 308,302 | 1117 | S. scrofa | 6 | 23 | 164,005 | 1224 | ||
| G. gallus | Z | 26 | 159,448 | 1119 | G. gallus | 11 | 23 | 69,793 | 1228 |
| Species | ADAMTS | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 4 | 5 | 6 | 9 | 10 | 12 | 16 | 18 | |||||||||||
| I | S | I | S | I | S | I | S | I | S | I | S | I | S | I | S | I | S | I | S | |
| B. taurus | 81 | 87 | 89 | 93 | 88 | 92 | 88 | 90 | 96 | 97 | 88 | 93 | 95 | 96 | 78 | 85 | 78 | 85 | 86 | 91 |
| O. aries | 80 | 87 | 89 | 93 | 88 | 92 | 88 | 90 | 95 | 97 | 88 | 93 | 95 | 96 | 79 | 85 | 78 | 86 | 87 | 93 |
| E. caballus | 86 | 91 | 87 | 91 | 89 | 92 | 91 | 93 | 96 | 97 | 90 | 94 | 96 | 97 | 79 | 86 | 80 | 86 | 89 | 93 |
| S. scrofa | 86 | 91 | 88 | 92 | 89 | 92 | 91 | 93 | 95 | 97 | 89 | 94 | 95 | 96 | 80 | 86 | 79 | 86 | 87 | 91 |
| G. gallus | 74 | 81 | 74 | 84 | 60 | 71 | 79 | 86 | 91 | 95 | 74 | 85 | 73 | 83 | 57 | 70 | - | - | 75 | 84 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hernández-Delgado, P.; Felix-Portillo, M.; Martínez-Quintana, J.A. ADAMTS Proteases: Importance in Animal Reproduction. Genes 2023, 14, 1181. https://doi.org/10.3390/genes14061181
Hernández-Delgado P, Felix-Portillo M, Martínez-Quintana JA. ADAMTS Proteases: Importance in Animal Reproduction. Genes. 2023; 14(6):1181. https://doi.org/10.3390/genes14061181
Chicago/Turabian StyleHernández-Delgado, Pamela, Monserrath Felix-Portillo, and José A. Martínez-Quintana. 2023. "ADAMTS Proteases: Importance in Animal Reproduction" Genes 14, no. 6: 1181. https://doi.org/10.3390/genes14061181
APA StyleHernández-Delgado, P., Felix-Portillo, M., & Martínez-Quintana, J. A. (2023). ADAMTS Proteases: Importance in Animal Reproduction. Genes, 14(6), 1181. https://doi.org/10.3390/genes14061181








