Progress in Research on Key Factors Regulating Lactation Initiation in the Mammary Glands of Dairy Cows
Abstract
1. Introduction
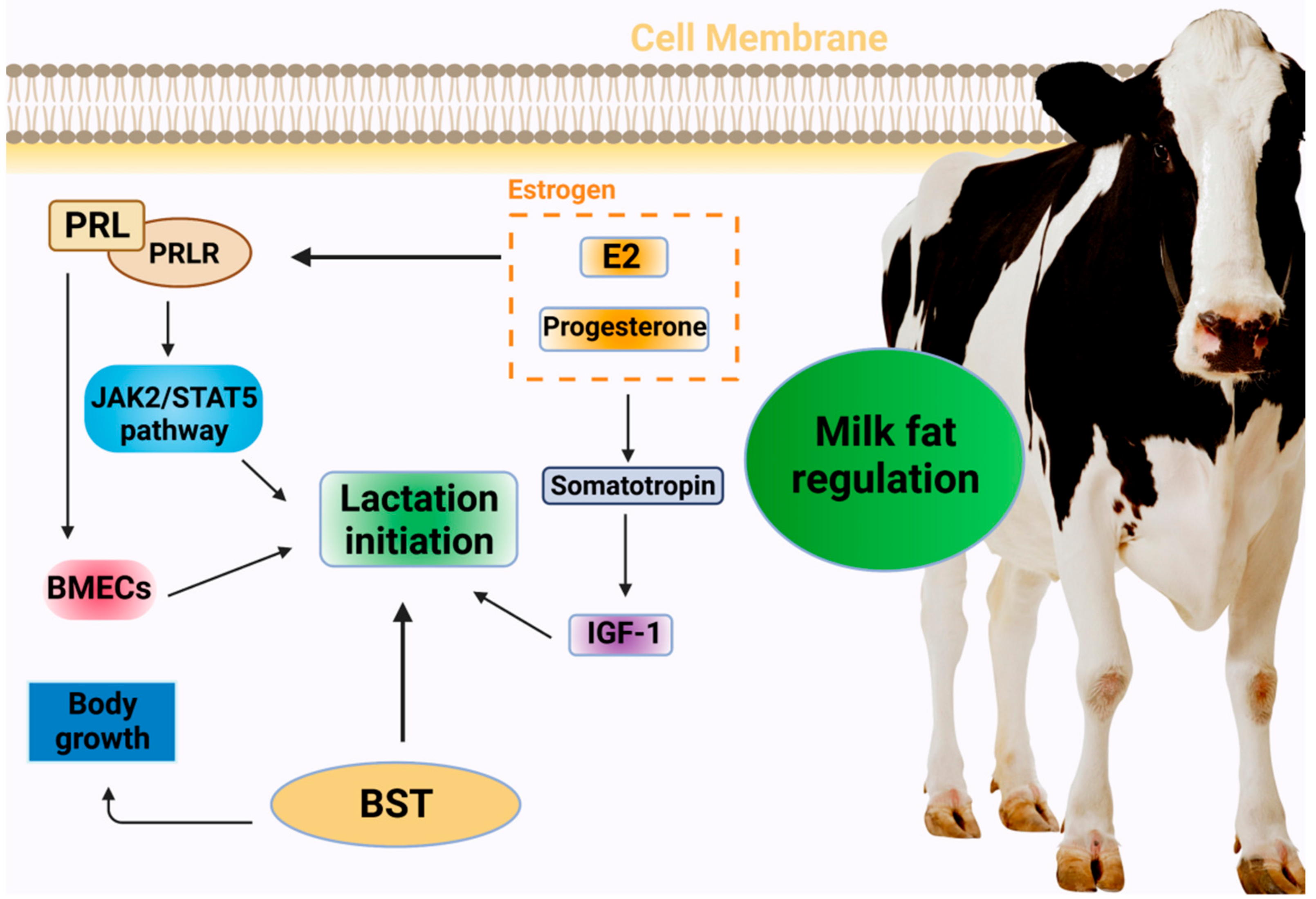
2. Hormonal Regulation of Lactation in Dairy Cows
2.1. PRL
2.2. Estrogen
2.3. Bovine Somatotropin (BST)
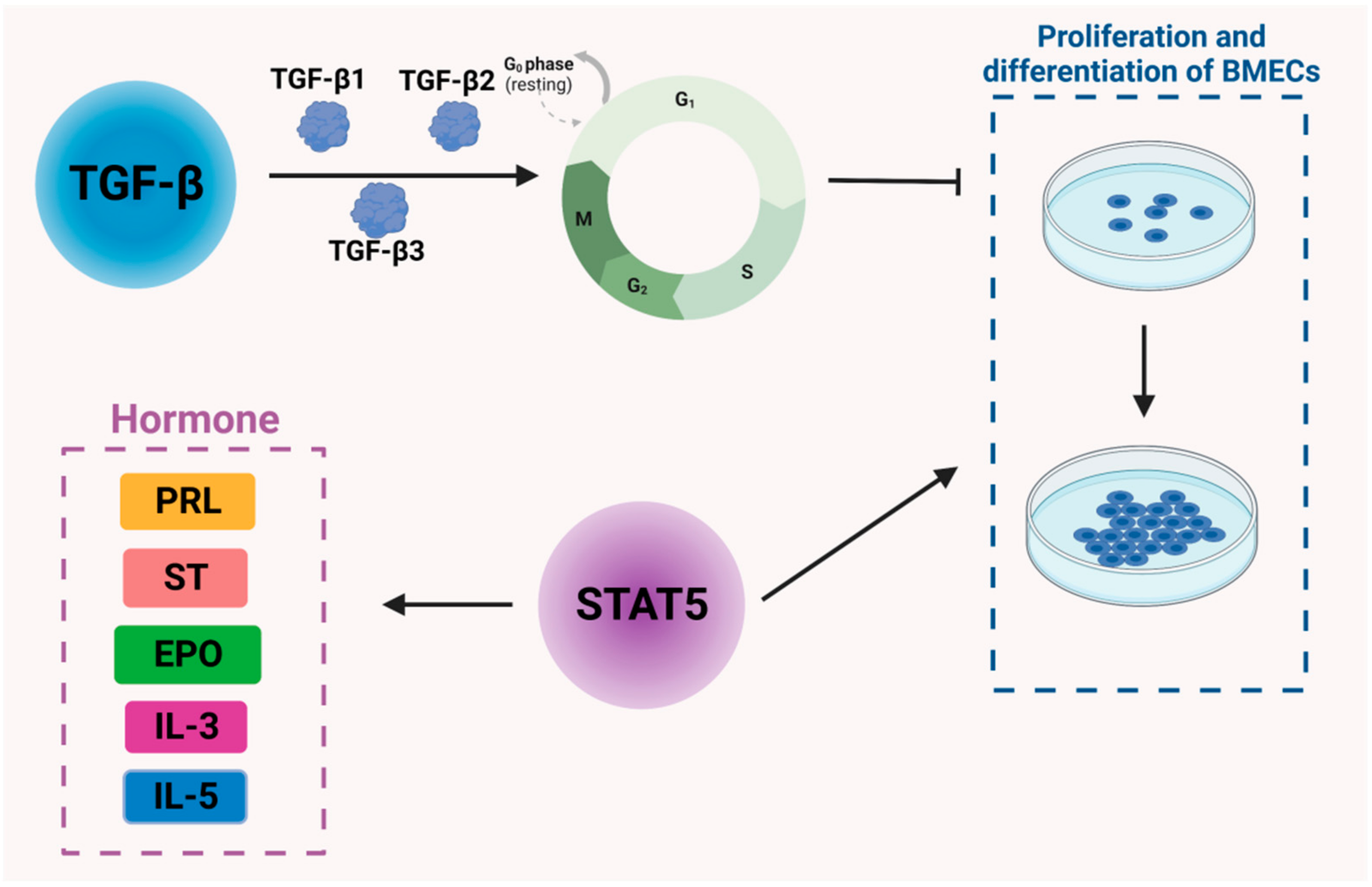
3. Regulation of Lactogenesis by Cytokines and Signaling Molecules in Dairy Cows
4. Regulation of Lactogenesis by Proteases in Dairy Cows
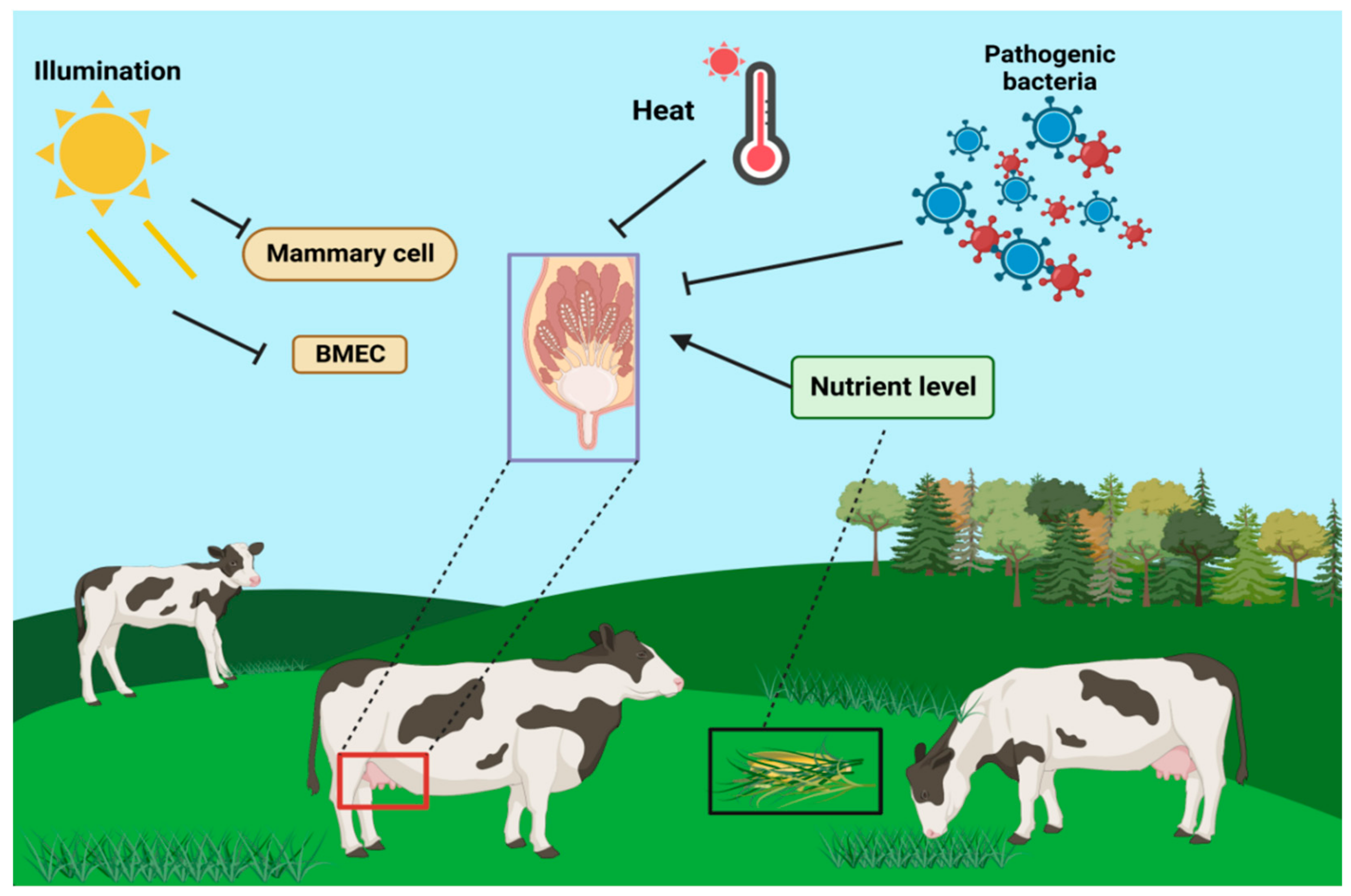
5. Regulation of Lactogenesis by Other Factors in Dairy Cows
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jaswal, S.; Jena, M.K.; Anand, V.; Jaswal, A.; Kancharla, S.; Kolli, P.; Mandadapu, G.; Kumar, S.; Mohanty, A.K. Critical Review on Physiological and Molecular Features during Bovine Mammary Gland Development: Recent Advances. Cells 2022, 11, 3325. [Google Scholar] [CrossRef] [PubMed]
- Bhat, S.A.; Ahmad, S.M.; Ibeagha-Awemu, E.M.; Bhat, B.A.; Dar, M.A.; Mumtaz, P.T.; Shah, R.A.; Ganai, N.A. Comparative transcriptome analysis of mammary epithelial cells at different stages of lactation reveals wide differences in gene expression and pathways regulating milk synthesis between Jersey and Kashmiri cattle. PLoS ONE 2019, 14, e0211773. [Google Scholar] [CrossRef] [PubMed]
- Roche, J.F.; Austin, E.J.; Ryan, M.; O‘Rourke, M.; Mihm, M.; Diskin, M.G. Regulation of follicle waves to maximize fertility in cattle. J. Reprod. Fertil. Suppl. 1999, 54, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Bova, T.L.; Chiavaccini, L.; Cline, G.F.; Hart, C.G.; Matheny, K.; Muth, A.M.; Voelz, B.E.; Kesler, D.; Memili, E. Environmental stressors influencing hormones and systems physiology in cattle. Reprod. Biol. Endocrinol. 2014, 12, 58. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Wei, Y.; Huang, Q.; Chen, Y.; Zeng, K.; Yang, W.; Chen, J.; Chen, J. Important Hormones Regulating Lipid Metabolism. Molecules 2022, 27, 7052. [Google Scholar] [CrossRef]
- Idrees, M.; Oh, S.H.; Muhammad, T.; El-Sheikh, M.; Song, S.H.; Lee, K.L.; Kong, I.K. Growth Factors, and Cytokines; Understanding the Role of Tyrosine Phosphatase SHP2 in Gametogenesis and Early Embryo Development. Cells 2020, 9, 1798. [Google Scholar] [CrossRef]
- Ivanova, E.; Le Guillou, S.; Hue-Beauvais, C.; Le Provost, F. Epigenetics: New Insights into Mammary Gland Biology. Genes 2021, 12, 231. [Google Scholar] [CrossRef]
- Arendt, L.M.; Kuperwasser, C. Form and function: How estrogen and progesterone regulate the mammary epithelial hierarchy. J. Mammary Gland Biol. Neoplasia 2015, 20, 9–25. [Google Scholar] [CrossRef]
- Perruchot, M.H.; Arévalo-Turrubiarte, M.; Dufreneix, F.; Finot, L.; Lollivier, V.; Chanat, E.; Mayeur, F.; Dessauge, F. Mammary Epithelial Cell Hierarchy in the Dairy Cow Throughout Lactation. Stem Cells Dev. 2016, 25, 1407–1418. [Google Scholar] [CrossRef]
- Fan, Y.; Han, Z.; Lu, X.; Arbab, A.A.I.; Nazar, M.; Yang, Y.; Yang, Z. Short Time-Series Expression Transcriptome Data Reveal the Gene Expression Patterns of Dairy Cow Mammary Gland as Milk Yield Decreased Process. Genes 2021, 12, 336. [Google Scholar] [CrossRef]
- Slepicka, P.F.; Somasundara, A.V.H.; Dos Santos, C.O. The molecular basis of mammary gland development and epithelial differentiation. Semin. Cell Dev. Biol. 2021, 114, 93–112. [Google Scholar] [CrossRef]
- Collier, R.J.; Bauman, D.E. Triennial Lactation Symposium/Bolfa: Historical perspectives of lactation biology in the late 20th and early 21st centuries. J. Anim. Sci. 2017, 95, 5639–5652. [Google Scholar] [CrossRef]
- Raven, L.A.; Cocks, B.G.; Goddard, M.E.; Pryce, J.E.; Hayes, B.J. Genetic variants in mammary development, prolactin signalling and involution pathways explain considerable variation in bovine milk production and milk composition. Genet. Sel. Evol. 2014, 46, 29. [Google Scholar] [CrossRef]
- Maruoka, M.; Kedashiro, S.; Ueda, Y.; Mizutani, K.; Takai, Y. Nectin-4 co-stimulates the prolactin receptor by interacting with SOCS1 and inhibiting its activity on the JAK2-STAT5a signaling pathway. J. Biol. Chem. 2017, 292, 6895–6909. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, M.; Li, D.; Li, M.; Kong, L.; Cao, M.; Wang, Y.; Song, C.; Fang, X.; Chen, H.; et al. Prolactin-Responsive Circular RNA circHIPK3 Promotes Proliferation of Mammary Epithelial Cells from Dairy Cow. Genes 2020, 11, 336. [Google Scholar] [CrossRef]
- Zhao, D.; Ma, G.; Zhang, X.; He, Y.; Li, M.; Han, X.; Fu, L.; Dong, X.Y.; Nagy, T.; Zhao, Q.; et al. Zinc Finger Homeodomain Factor Zfhx3 Is Essential for Mammary Lactogenic Differentiation by Maintaining Prolactin Signaling Activity. J. Biol. Chem. 2016, 291, 12809–12820. [Google Scholar] [CrossRef]
- De Souza, L.B.; Dupras, R.; Mills, L.; Chorfi, Y.; Price, C.A. Effect of synchronization of follicle-wave emergence with estradiol and progesterone and superstimulation with follicle-stimulating hormone on milk estrogen concentrations in dairy cattle. Can. J. Vet. Res. Rev. Can. Rech. Vet. 2013, 77, 75–78. [Google Scholar]
- Wall, E.H.; Hewitt, S.C.; Case, L.K.; Lin, C.Y.; Korach, K.S.; Teuscher, C. The role of genetics in estrogen responses: A critical piece of an intricate puzzle. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2014, 28, 5042–5054. [Google Scholar] [CrossRef]
- Hewitt, S.C.; Lierz, S.L.; Garcia, M.; Hamilton, K.J.; Gruzdev, A.; Grimm, S.A.; Lydon, J.P.; Demayo, F.J.; Korach, K.S. A distal super enhancer mediates estrogen-dependent mouse uterine-specific gene transcription of Igf1 (insulin-like growth factor 1). J. Biol. Chem. 2019, 294, 9746–9759. [Google Scholar] [CrossRef]
- Kasuya, E. Secretory pattern and regulatory mechanism of growth hormone in cattle. Anim. Sci. J. Nihon Chikusan Gakkaiho 2016, 87, 178–182. [Google Scholar] [CrossRef]
- Sciascia, Q.L.; Pacheco, D.; McCoard, S.A. Administration of Exogenous Growth Hormone Is Associated with Changes in Plasma and Intracellular Mammary Amino Acid Profiles and Abundance of the Mammary Gland Amino Acid Transporter SLC3A2 in Mid-Lactation Dairy Cows. PLoS ONE 2015, 10, e0134323. [Google Scholar] [CrossRef] [PubMed]
- Gurevich, E.; Segev, Y.; Landau, D. Growth Hormone and IGF1 Actions in Kidney Development and Function. Cells 2021, 10, 3371. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.V.; Li Calzi, S.; Shaw, L.C.; Kielczewski, J.L.; Korah, H.E.; Grant, M.B. An ocular view of the IGF-IGFBP system. Growth Horm. IGF Res. Off. J. Growth Horm. Res. Soc. Int. IGF Res. Soc. 2013, 23, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Osher, E.; Macaulay, V.M. Therapeutic Targeting of the IGF Axis. Cells 2019, 8, 895. [Google Scholar] [CrossRef] [PubMed]
- Derynck, R.; Budi, E.H. Specificity, versatility, and control of TGF-β family signaling. Sci. Signal. 2019, 12. [Google Scholar] [CrossRef]
- Kahata, K.; Maturi, V.; Moustakas, A. TGF-β Family Signaling in Ductal Differentiation and Branching Morphogenesis. Cold Spring Harb. Perspect. Biol. 2018, 10, a031997. [Google Scholar] [CrossRef]
- Biswas, S.K.; Banerjee, S.; Baker, G.W.; Kuo, C.Y.; Chowdhury, I. The Mammary Gland: Basic Structure and Molecular Signaling during Development. Int. J. Mol. Sci. 2022, 23, 3883. [Google Scholar] [CrossRef]
- Guo, Q.; Betts, C.; Pennock, N.; Mitchell, E.; Schedin, P. Mammary Gland Involution Provides a Unique Model to Study the TGF-β Cancer Paradox. J. Clin. Med. 2017, 6, 10. [Google Scholar] [CrossRef]
- Daniel, C.W.; Robinson, S.D. Regulation of mammary growth and function by TGF-beta. Mol. Reprod. Dev. 1992, 32, 145–151. [Google Scholar] [CrossRef]
- Ginjala, V.; Pakkanen, R. Determination of transforming growth factor-beta 1 (TGF-beta 1) and insulin-like growth factor (IGF-1) in bovine colostrum samples. J. Immunoass. 1998, 19, 195–207. [Google Scholar] [CrossRef]
- Flanders, K.C.; Yang, Y.A.; Herrmann, M.; Chen, J.; Mendoza, N.; Mirza, A.M.; Wakefield, L.M. Quantitation of TGF-β proteins in mouse tissues shows reciprocal changes in TGF-β1 and TGF-β3 in normal vs neoplastic mammary epithelium. Oncotarget 2016, 7, 38164–38179. [Google Scholar] [CrossRef] [PubMed]
- Schauwecker, S.M.; Kim, J.J.; Licht, J.D.; Clevenger, C.V. Histone H1 and Chromosomal Protein HMGN2 Regulate Prolactin-induced STAT5 Transcription Factor Recruitment and Function in Breast Cancer Cells. J. Biol. Chem. 2017, 292, 2237–2254. [Google Scholar] [CrossRef] [PubMed]
- Metser, G.; Shin, H.Y.; Wang, C.; Yoo, K.H.; Oh, S.; Villarino, A.V.; O‘Shea, J.J.; Kang, K.; Hennighausen, L. An autoregulatory enhancer controls mammary-specific STAT5 functions. Nucleic Acids Res. 2016, 44, 1052–1063. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.Z.; Khan, A.; Xiao, J.; Ma, Y.; Ma, J.; Gao, J.; Cao, Z. Role of the JAK-STAT Pathway in Bovine Mastitis and Milk Production. Animals 2020, 10, 101. [Google Scholar] [CrossRef] [PubMed]
- Grinman, D.Y.; Boras-Granic, K.; Takyar, F.M.; Dann, P.; Hens, J.R.; Marmol, C.; Lee, J.; Choi, J.; Chodosh, L.A.; Sola, M.E.G.; et al. PTHrP induces STAT5 activation, secretory differentiation and accelerates mammary tumor development. Breast Cancer Res. 2022, 24, 30. [Google Scholar] [CrossRef]
- Willi, M.; Yoo, K.H.; Wang, C.; Trajanoski, Z.; Hennighausen, L. Differential cytokine sensitivities of STAT5-dependent enhancers rely on Stat5 autoregulation. Nucleic Acids Res. 2016, 44, 10277–10291. [Google Scholar] [CrossRef]
- Harvatine, K.J.; Boisclair, Y.R.; Bauman, D.E. Time-dependent effect of trans-10,cis-12 conjugated linoleic acid on gene expression of lipogenic enzymes and regulators in mammary tissue of dairy cows. J. Dairy Sci. 2018, 101, 7585–7592. [Google Scholar] [CrossRef]
- Shu, G.; Huang, J.; Bao, C.; Meng, J.; Chen, H.; Cao, J. Effect of Different Proteases on the Degree of Hydrolysis and Angiotensin I-Converting Enzyme-Inhibitory Activity in Goat and Cow Milk. Biomolecules 2018, 8, 101. [Google Scholar] [CrossRef]
- Liberg, P. The fibrinogen concentration in blood of dairy cows and its influence on the interpretation of the glutaraldehyde and formol-gel test reactions. Acta Vet. Scand. 1978, 19, 413–421. [Google Scholar] [CrossRef]
- Mahmood, N.; Mihalcioiu, C.; Rabbani, S.A. Multifaceted Role of the Urokinase-Type Plasminogen Activator (uPA) and Its Receptor (uPAR): Diagnostic, Prognostic, and Therapeutic Applications. Front. Oncol. 2018, 8, 24. [Google Scholar] [CrossRef]
- Dow, M.P.; Bakke, L.J.; Cassar, C.A.; Peters, M.W.; Pursley, J.R.; Smith, G.W. Gonadotropin surge-induced up-regulation of the plasminogen activators (tissue plasminogen activator and urokinase plasminogen activator) and the urokinase plasminogen activator receptor within bovine periovulatory follicular and luteal tissue. Biol. Reprod. 2002, 66, 1413–1421. [Google Scholar] [CrossRef]
- Uhl, B.; Mittman, L.A.; Dominik, J.; Hennel, R.; Smiljanov, B.; Haring, F.; Schaubächer, J.B.; Braun, C.; Padovan, L.; Pick, R.; et al. uPA-PAI-1 heteromerization promotes breast cancer progression by attracting tumorigenic neutrophils. EMBO Mol. Med. 2021, 13, e13110. [Google Scholar] [CrossRef]
- Weigand, A.; Boos, A.M.; Tasbihi, K.; Beier, J.P.; Dalton, P.D.; Schrauder, M.; Horch, R.E.; Beckmann, M.W.; Strissel, P.L.; Strick, R. Selective isolation and characterization of primary cells from normal breast and tumors reveal plasticity of adipose derived stem cells. Breast Cancer Res. 2016, 18, 32. [Google Scholar] [CrossRef]
- Cui, N.; Hu, M.; Khalil, R.A. Biochemical and Biological Attributes of Matrix Metalloproteinases. Prog. Mol. Biol. Transl. Sci. 2017, 147, 1–73. [Google Scholar] [CrossRef]
- Wilson, A.M.; Wright, T.C.; Cant, J.P.; Osborne, V.R. Preferences of Dairy Cattle for Supplemental Light-Emitting Diode Lighting in the Resting Area. Animals 2022, 12, 1894. [Google Scholar] [CrossRef]
- Petrovski, K.R.; Cusack, P.; Malmo, J.; Cockcroft, P. The Value of ‘Cow Signs‘ in the Assessment of the Quality of Nutrition on Dairy Farms. Animals 2022, 12, 1352. [Google Scholar] [CrossRef]
- Kim, S.H.; Ramos, S.C.; Valencia, R.A.; Cho, Y.I.; Lee, S.S. Heat Stress: Effects on Rumen Microbes and Host Physiology, and Strategies to Alleviate the Negative Impacts on Lactating Dairy Cows. Front. Microbiol. 2022, 13, 804562. [Google Scholar] [CrossRef]
- Shandilya, U.K.; Sharma, A.; Sodhi, M.; Mukesh, M. Heat stress modulates differential response in skin fibroblast cells of native cattle (Bos indicus) and riverine buffaloes (Bubalus bubalis). Biosci. Rep. 2020, 40, BSR20191544. [Google Scholar] [CrossRef]
- Hughes, K.; Watson, C.J. The Mammary Microenvironment in Mastitis in Humans, Dairy Ruminants, Rabbits and Rodents: A One Health Focus. J. Mammary Gland Biol. Neoplasia 2018, 23, 27–41. [Google Scholar] [CrossRef]
- Edwards-Callaway, L.N.; Cramer, M.C.; Cadaret, C.N.; Bigler, E.J.; Engle, T.E.; Wagner, J.J.; Clark, D.L. Impacts of shade on cattle well-being in the beef supply chain. J. Anim. Sci. 2021, 99, skaa375. [Google Scholar] [CrossRef]
- Galán, E.; Llonch, P.; Villagrá, A.; Levit, H.; Pinto, S.; Del Prado, A. A systematic review of non-productivity-related animal-based indicators of heat stress resilience in dairy cattle. PLoS ONE 2018, 13, e0206520. [Google Scholar] [CrossRef] [PubMed]
- Gross, J.J. Limiting factors for milk production in dairy cows: Perspectives from physiology and nutrition. J. Anim. Sci. 2022, 100, skac044. [Google Scholar] [CrossRef] [PubMed]
- Herforth, A.; Arimond, M.; Álvarez-Sánchez, C.; Coates, J.; Christianson, K.; Muehlhoff, E. A Global Review of Food-Based Dietary Guidelines. Adv. Nutr. 2019, 10, 590–605. [Google Scholar] [CrossRef] [PubMed]
- Vlasova, A.N.; Saif, L.J. Bovine Immunology: Implications for Dairy Cattle. Front. Immunol. 2021, 12, 643206. [Google Scholar] [CrossRef]
- Hannan, F.M.; Elajnaf, T.; Vandenberg, L.N.; Kennedy, S.H.; Thakker, R.V. Hormonal regulation of mammary gland development and lactation. Nat. Rev. Endocrinol. 2023, 19, 46–61. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, H.; Li, J.; Wang, Y.; Cao, X.; Lv, X.; Yang, Z.; Chen, Z. Progress in Research on Key Factors Regulating Lactation Initiation in the Mammary Glands of Dairy Cows. Genes 2023, 14, 1163. https://doi.org/10.3390/genes14061163
Guo H, Li J, Wang Y, Cao X, Lv X, Yang Z, Chen Z. Progress in Research on Key Factors Regulating Lactation Initiation in the Mammary Glands of Dairy Cows. Genes. 2023; 14(6):1163. https://doi.org/10.3390/genes14061163
Chicago/Turabian StyleGuo, Haoyue, Jianyuan Li, Yuhao Wang, Xiang Cao, Xiaoyang Lv, Zhangping Yang, and Zhi Chen. 2023. "Progress in Research on Key Factors Regulating Lactation Initiation in the Mammary Glands of Dairy Cows" Genes 14, no. 6: 1163. https://doi.org/10.3390/genes14061163
APA StyleGuo, H., Li, J., Wang, Y., Cao, X., Lv, X., Yang, Z., & Chen, Z. (2023). Progress in Research on Key Factors Regulating Lactation Initiation in the Mammary Glands of Dairy Cows. Genes, 14(6), 1163. https://doi.org/10.3390/genes14061163






