Multisystem Proteinopathy Due to VCP Mutations: A Review of Clinical Heterogeneity and Genetic Diagnosis
Abstract
1. Introduction
2. Genetic Differential for VCP-MSP
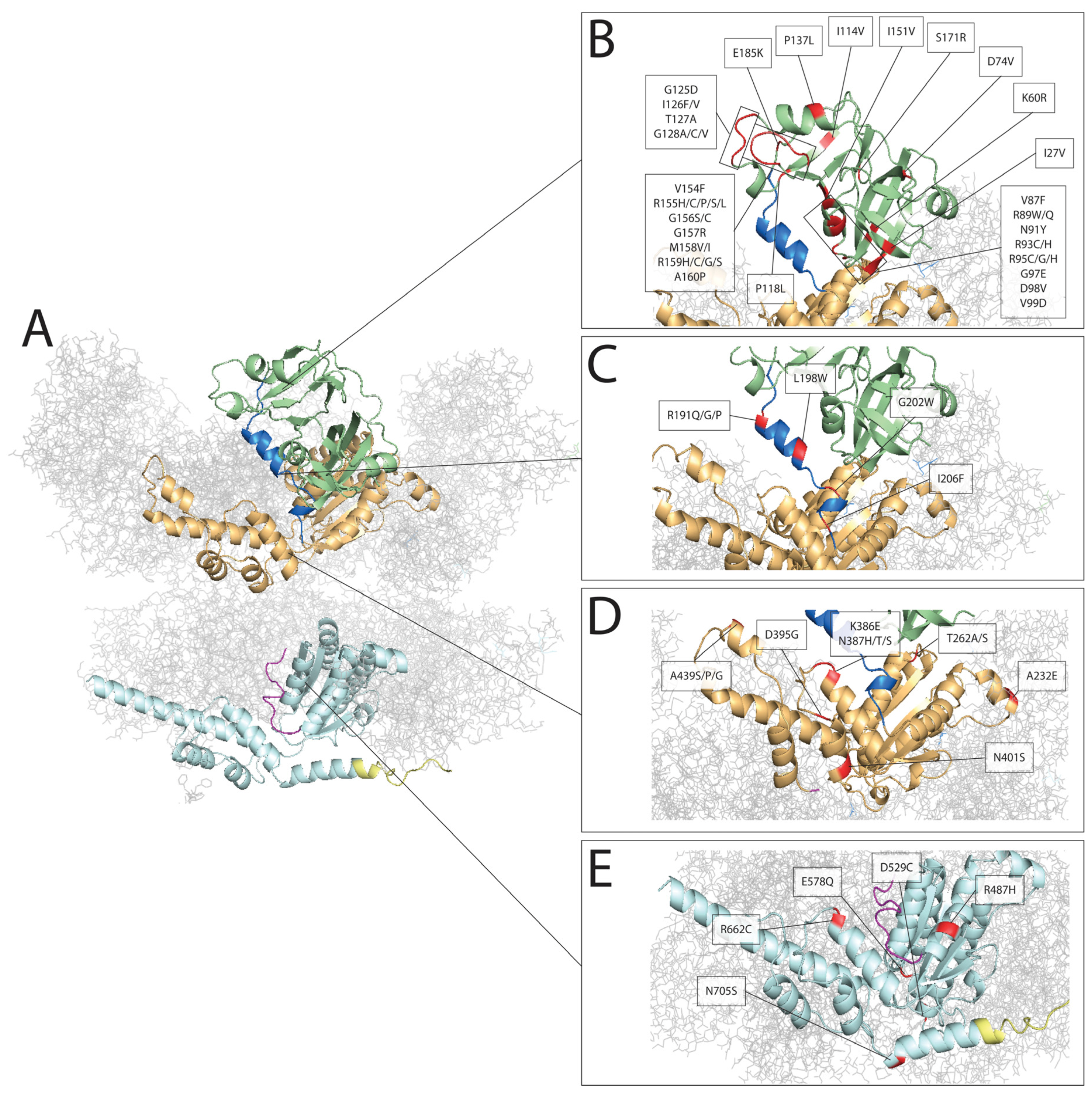
3. VCP and Genotype-Phenotype Correlations
| AA Substitution | Nucleotide Mutation | Domain | Phenotype or Disease | Inheritance | Reference |
|---|---|---|---|---|---|
| D6V | c.17A>T | N-Domain Cofactor and Substrate Binding | Myalgia, myopathy, arrhythmia | AD | [54] |
| I27V | c.79A>G | IBM, PDB, FTD | AD | [55,56] | |
| K60R | c.179A>G | ALS/FTD | SP | [57] | |
| D74V | c.221A>T | Chorea; FTD | AD | [58] | |
| V87F | c.259G>T | IBM | SP | [59] | |
| R89W | c.265C>T | IBM, FTD | SP | [60] | |
| R89Q | c.266G>A | ALS | SP | [61] | |
| N91Y | c.271A>T | IBM, ALS, FTD | AD | [62] | |
| R93C | c.277C>T | IBM, PDB, FTD | AD | [63] | |
| R93H | c.278G>A | HSP | SP | [64] | |
| R95C | c.283C>T | IBM | SP | [65] | |
| R95G | c.283C>G | IBM, PDB, FTD, ALS | AD | [10,42] | |
| R95H | c.284G>A | FTD | UN | [66] | |
| G97E | c.290G>A | IBM, PDB, FTD | AD | [51,67] | |
| D98V | c.293A>T | ALS, FTD | SP | [68] | |
| V99D | c.296T>A | PDB, FTD | AD | [69] | |
| I114V | c.340A>G | IBM (distal arm), ALS | SP | [70,71] | |
| P118L | c.353C>T | FTD | AD | [69] | |
| G125D | c.374G>A | IBM, PDB, FTD | AD | [69,72] | |
| I126F | c.376A>T | IBM | AD | [73] | |
| I126V | c.376A>G | IBM, FTD | SP | [59] | |
| T127A | c.379A>G | FTD | SP | [74] | |
| G128A | c.383G>C | IBM, PDB, FTD | AD | [42,75] | |
| G128C | c.382G>T | IBM | SP | [75] | |
| G128V | c.383G>T | IBM, PDB, FTD | AD | [76] | |
| P137L | c.410C>T | IBM (distal), PDB, FTD | AD | [77] | |
| I151V | c.451A>G | ALS | SP | [78] | |
| V154F | c.460G>T | FTD | AD | [79] | |
| R155H | c.464G>A | IBM, PDB, FTD, ALS | AD | [10] | |
| R155C | c.463C>T | IBM, PDB, FTD, ALS, PD | AD | [10,80] | |
| R155P | c.464G>C | IBM, PDB, FTD | AD | [10] | |
| R155S | c.463C>A | IBM, PDB, FTD | AD | [81] | |
| R155L | c.464G>T | IBM, PDB, FTD, ALS, SNH | AD | [82] | |
| G156S | c.466G>A | IBM, PDB, FTD | AD | [83,84] | |
| G156C | c.466G>T | ALS | AD | [85] | |
| G157R | c.469G>C | IBM, PDB, FTD, SNH | AD | [86] | |
| M158V | c.472A>G | PDB, ALS, FTD | AD | [87,88] | |
| M158I | c.474G>A | IBM, PDB, ALS | AD | [42,75] | |
| R159H | c.476G>A | IBM, PDB, FTD, ALS | AD AR | [70,72,89,90] | |
| R159C | c.475C>T | IBM, PDB, FTD, ALS, Parkinson disease, HSP | AD | [80,91,92,93] | |
| R159G | c.475C>G | FTD, ALS | AD | [94] | |
| R159S | c.475C>A | FTD | AD | [88] | |
| A160P | c.478G>C | IBM, PDB, FTD, ALS | AD | [42,75] | |
| E185K | c.553G>A | CMT2 | [44] | ||
| R191Q | c.572G>A | N-D1 Linker | IBM, PDB, FTD, ALS | AD | [10] |
| R191G | c.571C>G | IBM, ALS | AD | [80] | |
| R191P | c.572G>C | ALS | AD | [95] | |
| L198W | c.593T>G | IBM, PDB, FTD | AD | [96] | |
| G202W | c.604G>T | IBM, FTD | AD | [97] | |
| I206F | c.616A>T | IBM, PDB, FTD | AD | [98] | |
| A232E | c.695C>A | D1 Oligomerization Domain Heat Enhanced ATPase Domain | IBM, PDB, ALS | AD | [10] |
| T262A | c.784A>G | IBM, PDB, FTD, Parkinsonism | AD | [99] | |
| T262S | c.785C>G | FTD | AD | [88] | |
| K386E | c.1156A>G | Myopathy | NR | [100] | |
| N387H | c.1159A>C | IBM, FTD | AD | [96] | |
| N387T | c.1160A>C | ALS | SP | [101] | |
| N387S | c.1160G>A | IBM, PDB, FTD | AD | [102] | |
| D395G | c.1184A>G | FTD | AD | [53,79] | |
| D395A | c.1184A>C | FTD | AD | [52] | |
| N401S | c.1202A>G | Alzheimer Dementia | SP | [74] | |
| A439S | c.1315G>T | IBM, PDB | AD | [81] | |
| A439P | c.1315G>C | IBM, PDB, FTD | AD | [103] | |
| A439G | c.1316C>G | IBM, FTD | AD | [97] | |
| R487H | c.1460G>A | D2 Major ATPase Domain | FTD, ALS | AD | [104] |
| E578Q | IBM | UN | [105] | ||
| D592N | c.1774G>A | ALS | AD | [94] | |
| R662C | c.1984C>T | ALS | SP | [101] | |
| N750S | c.2249A>G | ALS | SP | [106] | |
| 9:35060456 del (5) Proposed fs at AA 515 | Autism | DN | [107] |
4. Genetic Diagnosis of VCP-MSP
4.1. Genetic Testing Methods
4.2. Access to Genetic Testing
4.3. Complementary Testing and Reverse Clinical Correlation
4.4. Interpreting Genetic Test Results
- (1)
- Positive: A positive result means that a variant in VCP known to be associated with VCP-MSP has been identified in the patient (i.e., a “pathogenic” or “likely pathogenic” variant). Thus far, all confirmed pathogenic variants in VCP are missense. A positive result may confirm the diagnosis in a person who meets clinical criteria or provide a pre-symptomatic diagnosis for an unaffected individual. If an unaffected individual has a positive result, there is a 90% chance that he or she will experience one or more of the features of VCP-MSP by the age of 45 [121]. Onset of symptoms is age-dependent and pre-symptomatic patients should be monitored by a healthcare professional on a regular basis.
- (2)
- Negative: A negative result means that no pathogenic variant has been identified in VCP. Depending on the test that was performed, and the purpose of testing, the implications of this result may vary:
- (a)
- Affected Individual: This means their condition is not caused by a mutation in VCP, but it remains possible that a mutation in another gene that was not tested could be responsible.
- (b)
- Unaffected individual in a family with a known pathogenic VCP mutation: In this situation, a negative result means that the person does not have the mutation and, therefore, will not develop VCP-MSP. The individual will not require follow-up care or monitoring for symptoms as they are confirmed to be unaffected and cannot pass it on to their children.
- (c)
- Affected individual in a family having a pathogenic VCP mutation: Individuals who are symptomatic but are non-carriers of the familial mutation may also have a “phenocopy” syndrome that should be considered in this situation [127]. Causes of phenocopy syndromes could be due to diagnostic confusion with acquired disease, a different genetic diagnosis, or non-organic symptoms.
4.5. Variant of Unknown Significance
5. Genetic Counselling
5.1. Risk to Other Family Members
5.2. Pre-Symptomatic Testing
5.3. Genetic Discrimination
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dalal, S.; Hanson, P.I. Membrane traffic: What drives the AAA motor? Cell 2001, 104, 5–8. [Google Scholar] [CrossRef]
- Meyer, H.; Weihl, C.C. The VCP/p97 system at a glance: Connecting cellular function to disease pathogenesis. J. Cell Sci. 2014, 127, 3877–3883. [Google Scholar] [CrossRef] [PubMed]
- Rabinovich, E.; Kerem, A.; Frohlich, K.U.; Diamant, N.; Bar-Nun, S. AAA-ATPase p97/Cdc48p, a cytosolic chaperone required for endoplasmic reticulum-associated protein degradation. Mol. Cell. Biol. 2002, 22, 626–634. [Google Scholar] [CrossRef] [PubMed]
- Shih, Y.T.; Hsueh, Y.P. VCP and ATL1 regulate endoplasmic reticulum and protein synthesis for dendritic spine formation. Nat. Commun. 2016, 7, 11020. [Google Scholar] [CrossRef]
- Rape, M.; Hoppe, T.; Gorr, I.; Kalocay, M.; Richly, H.; Jentsch, S. Mobilization of processed, membrane-tethered SPT23 transcription factor by CDC48(UFD1/NPL4), a ubiquitin-selective chaperone. Cell 2001, 107, 667–677. [Google Scholar] [CrossRef]
- Uchiyama, K.; Kondo, H. p97/p47-Mediated biogenesis of Golgi and ER. J. Biochem. 2005, 137, 115–119. [Google Scholar] [CrossRef]
- Frohlich, K.U.; Fries, H.W.; Rudiger, M.; Erdmann, R.; Botstein, D.; Mecke, D. Yeast cell cycle protein CDC48p shows full-length homology to the mammalian protein VCP and is a member of a protein family involved in secretion, peroxisome formation, and gene expression. J. Cell Biol. 1991, 114, 443–453. [Google Scholar] [CrossRef]
- Ju, J.S.; Fuentealba, R.A.; Miller, S.E.; Jackson, E.; Piwnica-Worms, D.; Baloh, R.H.; Weihl, C.C. Valosin-containing protein (VCP) is required for autophagy and is disrupted in VCP disease. J. Cell Biol. 2009, 187, 875–888. [Google Scholar] [CrossRef]
- Ferrari, V.; Cristofani, R.; Tedesco, B.; Crippa, V.; Chierichetti, M.; Casarotto, E.; Cozzi, M.; Mina, F.; Piccolella, M.; Galbiati, M.; et al. Valosin Containing Protein (VCP): A Multistep Regulator of Autophagy. Int. J. Mol. Sci. 2022, 23, 1939. [Google Scholar] [CrossRef]
- Watts, G.D.; Wymer, J.; Kovach, M.J.; Mehta, S.G.; Mumm, S.; Darvish, D.; Pestronk, A.; Whyte, M.P.; Kimonis, V.E. Inclusion body myopathy associated with Paget disease of bone and frontotemporal dementia is caused by mutant valosin-containing protein. Nat. Genet. 2004, 36, 377–381. [Google Scholar] [CrossRef]
- Korb, M.K.; Kimonis, V.E.; Mozaffar, T. Multisystem proteinopathy: Where myopathy and motor neuron disease converge. Muscle Nerve 2021, 63, 442–454. [Google Scholar] [CrossRef] [PubMed]
- Kimonis, V.E.; Mehta, S.G.; Fulchiero, E.C.; Thomasova, D.; Pasquali, M.; Boycott, K.; Neilan, E.G.; Kartashov, A.; Forman, M.S.; Tucker, S.; et al. Clinical studies in familial VCP myopathy associated with Paget disease of bone and frontotemporal dementia. Am. J. Med. Genet. A 2008, 146, 745–757. [Google Scholar] [CrossRef] [PubMed]
- Kimonis, V.E.; Fulchiero, E.; Vesa, J.; Watts, G. VCP disease associated with myopathy, Paget disease of bone and frontotemporal dementia: Review of a unique disorder. Biochim. Biophys. Acta 2008, 1782, 744–748. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.P. Multisystem proteinopathy: Intersecting genetics in muscle, bone, and brain degeneration. Neurology 2015, 85, 658–660. [Google Scholar] [CrossRef]
- Le Ber, I.; Van Bortel, I.; Nicolas, G.; Bouya-Ahmed, K.; Camuzat, A.; Wallon, D.; De Septenville, A.; Latouche, M.; Lattante, S.; Kabashi, E.; et al. hnRNPA2B1 and hnRNPA1 mutations are rare in patients with “multisystem proteinopathy” and frontotemporal lobar degeneration phenotypes. Neurobiol. Aging 2014, 35, 934.e5–934.e6. [Google Scholar] [CrossRef]
- Kim, H.J.; Kim, N.C.; Wang, Y.D.; Scarborough, E.A.; Moore, J.; Diaz, Z.; MacLea, K.S.; Freibaum, B.; Li, S.; Molliex, A.; et al. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 2013, 495, 467–473. [Google Scholar] [CrossRef]
- Liu, Y.; Shi, S.L. The roles of hnRNP A2/B1 in RNA biology and disease. Wiley Interdiscip. Rev. RNA 2021, 12, e1612. [Google Scholar] [CrossRef]
- Clarke, J.P.; Thibault, P.A.; Salapa, H.E.; Levin, M.C. A Comprehensive Analysis of the Role of hnRNP A1 Function and Dysfunction in the Pathogenesis of Neurodegenerative Disease. Front. Mol. Biosci. 2021, 8, 659610. [Google Scholar] [CrossRef]
- Liu, Q.; Shu, S.; Wang, R.R.; Liu, F.; Cui, B.; Guo, X.N.; Lu, C.X.; Li, X.G.; Liu, M.S.; Peng, B.; et al. Whole-exome sequencing identifies a missense mutation in hnRNPA1 in a family with flail arm ALS. Neurology 2016, 87, 1763–1769. [Google Scholar] [CrossRef]
- Kumar, A.V.; Mills, J.; Lapierre, L.R. Selective Autophagy Receptor p62/SQSTM1, a Pivotal Player in Stress and Aging. Front. Cell Dev. Biol. 2022, 10, 793328. [Google Scholar] [CrossRef]
- Bucelli, R.C.; Arhzaouy, K.; Pestronk, A.; Pittman, S.K.; Rojas, L.; Sue, C.M.; Evila, A.; Hackman, P.; Udd, B.; Harms, M.B.; et al. SQSTM1 splice site mutation in distal myopathy with rimmed vacuoles. Neurology 2015, 85, 665–674. [Google Scholar] [CrossRef] [PubMed]
- Rea, S.L.; Majcher, V.; Searle, M.S.; Layfield, R. SQSTM1 mutations--bridging Paget disease of bone and ALS/FTLD. Exp. Cell Res. 2014, 325, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Fecto, F.; Yan, J.; Vemula, S.P.; Liu, E.; Yang, Y.; Chen, W.; Zheng, J.G.; Shi, Y.; Siddique, N.; Arrat, H.; et al. SQSTM1 mutations in familial and sporadic amyotrophic lateral sclerosis. Arch. Neurol. 2011, 68, 1440–1446. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Jonson, P.H.; Sarparanta, J.; Palmio, J.; Sarkar, M.; Vihola, A.; Evila, A.; Suominen, T.; Penttila, S.; Savarese, M.; et al. TIA1 variant drives myodegeneration in multisystem proteinopathy with SQSTM1 mutations. J. Clin. Investig. 2018, 128, 1164–1177. [Google Scholar] [CrossRef] [PubMed]
- Niu, Z.; Pontifex, C.S.; Berini, S.; Hamilton, L.E.; Naddaf, E.; Wieben, E.; Aleff, R.A.; Martens, K.; Gruber, A.; Engel, A.G.; et al. Myopathy With SQSTM1 and TIA1 Variants: Clinical and Pathological Features. Front. Neurol. 2018, 9, 147. [Google Scholar] [CrossRef] [PubMed]
- Gilks, N.; Kedersha, N.; Ayodele, M.; Shen, L.; Stoecklin, G.; Dember, L.M.; Anderson, P. Stress granule assembly is mediated by prion-like aggregation of TIA-1. Mol. Biol. Cell 2004, 15, 5383–5398. [Google Scholar] [CrossRef]
- Malik, A.M.; Barmada, S.J. Matrin 3 in neuromuscular disease: Physiology and pathophysiology. JCI Insight 2021, 6, e143948. [Google Scholar] [CrossRef]
- Johnson, J.O.; Pioro, E.P.; Boehringer, A.; Chia, R.; Feit, H.; Renton, A.E.; Pliner, H.A.; Abramzon, Y.; Marangi, G.; Winborn, B.J.; et al. Mutations in the Matrin 3 gene cause familial amyotrophic lateral sclerosis. Nat. Neurosci. 2014, 17, 664–666. [Google Scholar] [CrossRef]
- Lin, K.P.; Tsai, P.C.; Liao, Y.C.; Chen, W.T.; Tsai, C.P.; Soong, B.W.; Lee, Y.C. Mutational analysis of MATR3 in Taiwanese patients with amyotrophic lateral sclerosis. Neurobiol. Aging 2015, 36, 2005.e1–2005.e4. [Google Scholar] [CrossRef]
- Leblond, C.S.; Gan-Or, Z.; Spiegelman, D.; Laurent, S.B.; Szuto, A.; Hodgkinson, A.; Dionne-Laporte, A.; Provencher, P.; de Carvalho, M.; Orru, S.; et al. Replication study of MATR3 in familial and sporadic amyotrophic lateral sclerosis. Neurobiol. Aging 2016, 37, 209.e17–209.e21. [Google Scholar] [CrossRef]
- Millecamps, S.; De Septenville, A.; Teyssou, E.; Daniau, M.; Camuzat, A.; Albert, M.; LeGuern, E.; Galimberti, D.; French research network on, F.T.D.; Ftd, A.L.S.; et al. Genetic analysis of matrin 3 gene in French amyotrophic lateral sclerosis patients and frontotemporal lobar degeneration with amyotrophic lateral sclerosis patients. Neurobiol. Aging 2014, 35, 2882.e13–2882.e15. [Google Scholar] [CrossRef] [PubMed]
- Castro, M.; Venkateswaran, N.; Peters, S.T.; Deyle, D.R.; Bower, M.; Koob, M.D.; Boeve, B.F.; Vossel, K. Case Report: Early-Onset Behavioral Variant Frontotemporal Dementia in Patient With Retrotransposed Full-Length Transcript of Matrin-3 Variant 5. Front. Neurol. 2020, 11, 600468. [Google Scholar] [CrossRef]
- Hackman, P.; Sarparanta, J.; Lehtinen, S.; Vihola, A.; Evila, A.; Jonson, P.H.; Luque, H.; Kere, J.; Screen, M.; Chinnery, P.F.; et al. Welander distal myopathy is caused by a mutation in the RNA-binding protein TIA1. Ann. Neurol. 2013, 73, 500–509. [Google Scholar] [CrossRef] [PubMed]
- Klar, J.; Sobol, M.; Melberg, A.; Mabert, K.; Ameur, A.; Johansson, A.C.; Feuk, L.; Entesarian, M.; Orlen, H.; Casar-Borota, O.; et al. Welander distal myopathy caused by an ancient founder mutation in TIA1 associated with perturbed splicing. Hum. Mutat. 2013, 34, 572–577. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, I.R.; Nicholson, A.M.; Sarkar, M.; Messing, J.; Purice, M.D.; Pottier, C.; Annu, K.; Baker, M.; Perkerson, R.B.; Kurti, A.; et al. TIA1 Mutations in Amyotrophic Lateral Sclerosis and Frontotemporal Dementia Promote Phase Separation and Alter Stress Granule Dynamics. Neuron 2017, 95, 808–816.e9. [Google Scholar] [CrossRef] [PubMed]
- van der Spek, R.A.; van Rheenen, W.; Pulit, S.L.; Kenna, K.P.; Ticozzi, N.; Kooyman, M.; McLaughlin, R.L.; Moisse, M.; van Eijk, K.R.; van Vugt, J.; et al. Reconsidering the causality of TIA1 mutations in ALS. Amyotroph. Lateral Scler. Frontotemporal. Degener. 2018, 19, 1–3. [Google Scholar] [CrossRef]
- Baradaran-Heravi, Y.; Dillen, L.; Nguyen, H.P.; Van Mossevelde, S.; Baets, J.; De Jonghe, P.; Engelborghs, S.; De Deyn, P.P.; Vandenbulcke, M.; Vandenberghe, R.; et al. No supportive evidence for TIA1 gene mutations in a European cohort of ALS-FTD spectrum patients. Neurobiol. Aging 2018, 69, 293.e9–293.e11. [Google Scholar] [CrossRef]
- Ryan, T.A.; Tumbarello, D.A. Optineurin: A Coordinator of Membrane-Associated Cargo Trafficking and Autophagy. Front. Immunol. 2018, 9, 1024. [Google Scholar] [CrossRef]
- Maruyama, H.; Morino, H.; Ito, H.; Izumi, Y.; Kato, H.; Watanabe, Y.; Kinoshita, Y.; Kamada, M.; Nodera, H.; Suzuki, H.; et al. Mutations of optineurin in amyotrophic lateral sclerosis. Nature 2010, 465, 223–226. [Google Scholar] [CrossRef]
- Albagha, O.M.; Visconti, M.R.; Alonso, N.; Langston, A.L.; Cundy, T.; Dargie, R.; Dunlop, M.G.; Fraser, W.D.; Hooper, M.J.; Isaia, G.; et al. Genome-wide association study identifies variants at CSF1, OPTN and TNFRSF11A as genetic risk factors for Paget’s disease of bone. Nat. Genet. 2010, 42, 520–524. [Google Scholar] [CrossRef]
- Pottier, C.; Bieniek, K.F.; Finch, N.; van de Vorst, M.; Baker, M.; Perkersen, R.; Brown, P.; Ravenscroft, T.; van Blitterswijk, M.; Nicholson, A.M.; et al. Whole-genome sequencing reveals important role for TBK1 and OPTN mutations in frontotemporal lobar degeneration without motor neuron disease. Acta Neuropathol. 2015, 130, 77–92. [Google Scholar] [CrossRef] [PubMed]
- Al-Obeidi, E.; Al-Tahan, S.; Surampalli, A.; Goyal, N.; Wang, A.K.; Hermann, A.; Omizo, M.; Smith, C.; Mozaffar, T.; Kimonis, V. Genotype-phenotype study in patients with valosin-containing protein mutations associated with multisystem proteinopathy. Clin. Genet. 2018, 93, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Farpour, F.; Tehranzadeh, J.; Donkervoort, S.; Smith, C.; Martin, B.; Vanjara, P.; Osann, K.; Kimonis, V.E. Radiological features of Paget disease of bone associated with VCP myopathy. Skeletal. Radiol. 2012, 41, 329–337. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, M.A.; Feely, S.M.; Speziani, F.; Strickland, A.V.; Danzi, M.; Bacon, C.; Lee, Y.; Chou, T.F.; Blanton, S.H.; Weihl, C.C.; et al. A novel mutation in VCP causes Charcot-Marie-Tooth Type 2 disease. Brain 2014, 137, 2897–2902. [Google Scholar] [CrossRef] [PubMed]
- Gite, J.; Milko, E.; Brady, L.; Baker, S.K. Phenotypic convergence in Charcot-Marie-Tooth 2Y with novel VCP mutation. Neuromuscul. Disord. 2020, 30, 232–235. [Google Scholar] [CrossRef]
- van de Warrenburg, B.P.; Schouten, M.I.; de Bot, S.T.; Vermeer, S.; Meijer, R.; Pennings, M.; Gilissen, C.; Willemsen, M.A.; Scheffer, H.; Kamsteeg, E.J. Clinical exome sequencing for cerebellar ataxia and spastic paraplegia uncovers novel gene-disease associations and unanticipated rare disorders. Eur. J. Hum. Genet. 2017, 25, 393. [Google Scholar] [CrossRef]
- Kimonis, V.E.; Watts, G.D. Autosomal dominant inclusion body myopathy, Paget disease of bone, and frontotemporal dementia. Alzheimer Dis. Assoc. Disord. 2005, 19 (Suppl. S1), S44–S47. [Google Scholar] [CrossRef]
- Miller, T.D.; Jackson, A.P.; Barresi, R.; Smart, C.M.; Eugenicos, M.; Summers, D.; Clegg, S.; Straub, V.; Stone, J. Inclusion body myopathy with Paget disease and frontotemporal dementia (IBMPFD): Clinical features including sphincter disturbance in a large pedigree. J. Neurol. Neurosurg. Psychiatry 2009, 80, 583–584. [Google Scholar] [CrossRef]
- Pfeffer, G.; Povitz, M. Respiratory management of patients with neuromuscular disease: Current perspectives. Degener Neurol Neuromuscul. Dis. 2016, 6, 111–118. [Google Scholar] [CrossRef][Green Version]
- Korb, M.; Peck, A.; Alfano, L.N.; Berger, K.I.; James, M.K.; Ghoshal, N.; Healzer, E.; Henchcliffe, C.; Khan, S.; Mammen, P.P.A.; et al. Development of a standard of care for patients with valosin-containing protein associated multisystem proteinopathy. Orphanet J. Rare Dis. 2022, 17, 23. [Google Scholar] [CrossRef]
- Jerath, N.U.; Crockett, C.D.; Moore, S.A.; Shy, M.E.; Weihl, C.C.; Chou, T.F.; Grider, T.; Gonzalez, M.A.; Zuchner, S.; Swenson, A. Rare Manifestation of a c.290 C>T, p.Gly97Glu VCP Mutation. Case Rep. Genet. 2015, 2015, 239167. [Google Scholar] [CrossRef] [PubMed]
- Bruno, F.; Conidi, M.E.; Puccio, G.; Frangipane, F.; Lagana, V.; Bernardi, L.; Smirne, N.; Mirabelli, M.; Colao, R.; Curcio, S.; et al. A Novel Mutation (D395A) in Valosin-Containing Protein Gene Is Associated With Early Onset Frontotemporal Dementia in an Italian Family. Front. Genet. 2021, 12, 795029. [Google Scholar] [CrossRef] [PubMed]
- Darwich, N.F.; Phan, J.M.; Kim, B.; Suh, E.; Papatriantafyllou, J.D.; Changolkar, L.; Nguyen, A.T.; O’Rourke, C.M.; He, Z.; Porta, S.; et al. Autosomal dominant VCP hypomorph mutation impairs disaggregation of PHF-tau. Science 2020, 370, eaay8826. [Google Scholar] [CrossRef] [PubMed]
- Meinke, P.; Kerr, A.R.W.; Czapiewski, R.; de Las Heras, J.I.; Dixon, C.R.; Harris, E.; Kolbel, H.; Muntoni, F.; Schara, U.; Straub, V.; et al. A multistage sequencing strategy pinpoints novel candidate alleles for Emery-Dreifuss muscular dystrophy and supports gene misregulation as its pathomechanism. EBioMedicine 2019, 53, 102587. [Google Scholar] [CrossRef] [PubMed]
- Majounie, E.; Traynor, B.J.; Chio, A.; Restagno, G.; Mandrioli, J.; Benatar, M.; Taylor, J.P.; Singleton, A.B. Mutational analysis of the VCP gene in Parkinson’s disease. Neurobiol. Aging 2012, 33, 209.e1–209.e2. [Google Scholar] [CrossRef] [PubMed]
- Rohrer, J.D.; Warren, J.D.; Reiman, D.; Uphill, J.; Beck, J.; Collinge, J.; Rossor, M.N.; Isaacs, A.M.; Mead, S. A novel exon 2 I27V VCP variant is associated with dissimilar clinical syndromes. J. Neurol. 2011, 258, 1494–1496. [Google Scholar] [CrossRef][Green Version]
- Pensato, V.; Magri, S.; Bella, E.D.; Tannorella, P.; Bersano, E.; Soraru, G.; Gatti, M.; Ticozzi, N.; Taroni, F.; Lauria, G.; et al. Sorting Rare ALS Genetic Variants by Targeted Re-Sequencing Panel in Italian Patients: OPTN, VCP, and SQSTM1 Variants Account for 3% of Rare Genetic Forms. J. Clin. Med. 2020, 9, 412. [Google Scholar] [CrossRef]
- Mariani, L.L.; Tesson, C.; Charles, P.; Cazeneuve, C.; Hahn, V.; Youssov, K.; Freeman, L.; Grabli, D.; Roze, E.; Noel, S.; et al. Expanding the Spectrum of Genes Involved in Huntington Disease Using a Combined Clinical and Genetic Approach. JAMA Neurol. 2016, 73, 1105–1114. [Google Scholar] [CrossRef]
- Inoue, M.; Iida, A.; Hayashi, S.; Mori-Yoshimura, M.; Nagaoka, A.; Yoshimura, S.; Shiraishi, H.; Tsujino, A.; Takahashi, Y.; Nonaka, I.; et al. Two novel VCP missense variants identified in Japanese patients with multisystem proteinopathy. Hum. Genome Var. 2018, 5, 9. [Google Scholar] [CrossRef]
- Falcao de Campos, C.; de Carvalho, M. Distal myopathy and rapidly progressive dementia associated with a novel mutation in the VCP gene: Expanding inclusion body myopathy with early-onset Paget disease and frontotemporal dementia spectrum. J. Clin. Neurosci. 2019, 64, 8–10. [Google Scholar] [CrossRef]
- Deng, J.; Wu, W.; Xie, Z.; Gang, Q.; Yu, M.; Liu, J.; Wang, Q.; Lv, H.; Zhang, W.; Huang, Y.; et al. Novel and Recurrent Mutations in a Cohort of Chinese Patients With Young-Onset Amyotrophic Lateral Sclerosis. Front. Neurosci. 2019, 13, 1289. [Google Scholar] [CrossRef] [PubMed]
- Abrahao, A.; Abath Neto, O.; Kok, F.; Zanoteli, E.; Santos, B.; Pinto, W.B.; Barsottini, O.G.; Oliveira, A.S.; Pedroso, J.L. One family, one gene and three phenotypes: A novel VCP (valosin-containing protein) mutation associated with myopathy with rimmed vacuoles, amyotrophic lateral sclerosis and frontotemporal dementia. J. Neurol. Sci. 2016, 368, 352–358. [Google Scholar] [CrossRef] [PubMed]
- Hubbers, C.U.; Clemen, C.S.; Kesper, K.; Boddrich, A.; Hofmann, A.; Kamarainen, O.; Tolksdorf, K.; Stumpf, M.; Reichelt, J.; Roth, U.; et al. Pathological consequences of VCP mutations on human striated muscle. Brain 2007, 130, 381–393. [Google Scholar] [CrossRef] [PubMed]
- Neveling, K.; Feenstra, I.; Gilissen, C.; Hoefsloot, L.H.; Kamsteeg, E.J.; Mensenkamp, A.R.; Rodenburg, R.J.; Yntema, H.G.; Spruijt, L.; Vermeer, S.; et al. A post-hoc comparison of the utility of sanger sequencing and exome sequencing for the diagnosis of heterogeneous diseases. Hum. Mutat. 2013, 34, 1721–1726. [Google Scholar] [CrossRef] [PubMed]
- Crisp, M.J.; Mawuenyega, K.G.; Patterson, B.W.; Reddy, N.C.; Chott, R.; Self, W.K.; Weihl, C.C.; Jockel-Balsarotti, J.; Varadhachary, A.S.; Bucelli, R.C.; et al. In vivo kinetic approach reveals slow SOD1 turnover in the CNS. J. Clin. Investig. 2015, 125, 2772–2780. [Google Scholar] [CrossRef] [PubMed]
- Kaleem, M.; Zhao, A.; Hamshere, M.; Myers, A.J. Identification of a novel valosin-containing protein polymorphism in late-onset Alzheimer’s disease. Neurodegener. Dis. 2007, 4, 376–381. [Google Scholar] [CrossRef]
- Gu, J.M.; Ke, Y.H.; Yue, H.; Liu, Y.J.; Zhang, Z.; Zhang, H.; Hu, W.W.; Wang, C.; He, J.W.; Hu, Y.Q.; et al. A novel VCP mutation as the cause of atypical IBMPFD in a Chinese family. Bone 2013, 52, 9–16. [Google Scholar] [CrossRef]
- Ando, T.; Nakamura, R.; Kuru, S.; Yokoi, D.; Atsuta, N.; Koike, H.; Suzuki, M.; Hara, K.; Iguchi, Y.; Harada, Y.; et al. The wide-ranging clinical and genetic features in Japanese families with valosin-containing protein proteinopathy. Neurobiol. Aging 2020, 100, 120.e1–120.e6. [Google Scholar] [CrossRef]
- Saracino, D.; Clot, F.; Camuzat, A.; Anquetil, V.; Hannequin, D.; Guyant-Marechal, L.; Didic, M.; Guillot-Noel, L.; Rinaldi, D.; Latouche, M.; et al. Novel VCP mutations expand the mutational spectrum of frontotemporal dementia. Neurobiol. Aging 2018, 72, 187.e11–187.e14. [Google Scholar] [CrossRef]
- Koppers, M.; van Blitterswijk, M.M.; Vlam, L.; Rowicka, P.A.; van Vught, P.W.; Groen, E.J.; Spliet, W.G.; Engelen-Lee, J.; Schelhaas, H.J.; de Visser, M.; et al. VCP mutations in familial and sporadic amyotrophic lateral sclerosis. Neurobiol. Aging 2012, 33, 837.e7–837.e13. [Google Scholar] [CrossRef]
- Boland-Freitas, R.; Graham, J.; Davis, M.; Geevasinga, N.; Vucic, S.; Ng, K. Late onset distal myopathy of the upper limbs due to p.Ile151Val mutation in VCP. Muscle Nerve 2016, 54, 165–166. [Google Scholar] [CrossRef] [PubMed]
- De Ridder, W.; Azmi, A.; Clemen, C.S.; Eichinger, L.; Hofmann, A.; Schroder, R.; Johnson, K.; Topf, A.; Straub, V.; De Jonghe, P.; et al. Multisystem proteinopathy due to a homozygous p.Arg159His VCP mutation: A tale of the unexpected. Neurology 2019, 94, e785–e796. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, S.; Shimizu, T.; Komori, T.; Mori-Yoshimura, M.; Minami, N.; Hayashi, Y.K. Nuclear inclusions mimicking poly(A)-binding protein nuclear 1 inclusions in a case of inclusion body myopathy associated with Paget disease of bone and frontotemporal dementia with a novel mutation in the valosin-containing protein gene. Neuromuscul. Disord. 2016, 26, 436–440. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.; Liu, S.; Xiang, L.; Wang, Y.; Liu, M.; Liu, S.; Han, T.; Zhou, Y.; Wang, J.; Cai, L.; et al. Frontotemporal dementia-related gene mutations in clinical dementia patients from a Chinese population. J. Hum. Genet. 2016, 61, 1003–1008. [Google Scholar] [CrossRef]
- Al-Tahan, S.; Al-Obeidi, E.; Yoshioka, H.; Lakatos, A.; Weiss, L.; Grafe, M.; Palmio, J.; Wicklund, M.; Harati, Y.; Omizo, M.; et al. Novel valosin-containing protein mutations associated with multisystem proteinopathy. Neuromuscul. Disord. 2018, 28, 491–501. [Google Scholar] [CrossRef]
- Lucente, G.; Almendrote, M.; Ramos-Fransi, A.; Martinez-Pineiro, A.; Camano, P.; Ballester-Lopez, A.; Lucia, A.; Carrato, C.; Nogales-Gadea, G.; Coll-Canti, J. A novel mutation in the valosin-containing-protein gene found in a Spanish family. J. Neurol. Sci. 2018, 391, 112–113. [Google Scholar] [CrossRef]
- Palmio, J.; Sandell, S.; Suominen, T.; Penttila, S.; Raheem, O.; Hackman, P.; Huovinen, S.; Haapasalo, H.; Udd, B. Distinct distal myopathy phenotype caused by VCP gene mutation in a Finnish family. Neuromuscul. Disord. 2011, 21, 551–555. [Google Scholar] [CrossRef]
- DeJesus-Hernandez, M.; Desaro, P.; Johnston, A.; Ross, O.A.; Wszolek, Z.K.; Ertekin-Taner, N.; Graff-Radford, N.R.; Rademakers, R.; Boylan, K. Novel p.Ile151Val mutation in VCP in a patient of African American descent with sporadic ALS. Neurology 2011, 77, 1102–1103. [Google Scholar] [CrossRef]
- Ramos, E.M.; Dokuru, D.R.; Van Berlo, V.; Wojta, K.; Wang, Q.; Huang, A.Y.; Deverasetty, S.; Qin, Y.; van Blitterswijk, M.; Jackson, J.; et al. Genetic screening of a large series of North American sporadic and familial frontotemporal dementia cases. Alzheimers Dement. 2020, 16, 118–130. [Google Scholar] [CrossRef]
- Gonzalez-Perez, P.; Cirulli, E.T.; Drory, V.E.; Dabby, R.; Nisipeanu, P.; Carasso, R.L.; Sadeh, M.; Fox, A.; Festoff, B.W.; Sapp, P.C.; et al. Novel mutation in VCP gene causes atypical amyotrophic lateral sclerosis. Neurology 2012, 79, 2201–2208. [Google Scholar] [CrossRef]
- Stojkovic, T.; Hammouda el, H.; Richard, P.; Lopez de Munain, A.; Ruiz-Martinez, J.; Camano, P.; Laforet, P.; Penisson-Besnier, I.; Ferrer, X.; Lacour, A.; et al. Clinical outcome in 19 French and Spanish patients with valosin-containing protein myopathy associated with Paget’s disease of bone and frontotemporal dementia. Neuromuscul. Disord. 2009, 19, 316–323. [Google Scholar] [CrossRef] [PubMed]
- Kumar, K.R.; Needham, M.; Mina, K.; Davis, M.; Brewer, J.; Staples, C.; Ng, K.; Sue, C.M.; Mastaglia, F.L. Two Australian families with inclusion-body myopathy, Paget’s disease of bone and frontotemporal dementia: Novel clinical and genetic findings. Neuromuscul. Disord. 2010, 20, 330–334. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, J.; Iwasa, K.; Yanase, D.; Yamada, M. Inclusion body myopathy with Paget disease of the bone and frontotemporal dementia associated with a novel G156S mutation in the VCP gene. Muscle Nerve 2013. [Google Scholar] [CrossRef] [PubMed]
- Pellerin, D.; Ellezam, B.; Korathanakhun, P.; Renaud, M.; Dicaire, M.J.; Pilote, L.; Levy, J.P.; Karamchandani, J.; Ducharme, S.; Massie, R.; et al. Multi-system proteinopathy associated with a VCP G156S mutation in a French Canadian family. Can. J. Neurol. Sci. 2020, 47, 412–415. [Google Scholar] [CrossRef] [PubMed]
- Segawa, M.; Hoshi, A.; Naruse, H.; Kuroda, M.; Bujo, H.; Ugawa, Y. A patient with familial amyotrophic lateral sclerosis associated with a new valosin-containing protein (VCP) gene mutation. Rinsho Shinkeigaku 2015, 55, 914–920. [Google Scholar] [CrossRef] [PubMed]
- Djamshidian, A.; Schaefer, J.; Haubenberger, D.; Stogmann, E.; Zimprich, F.; Auff, E.; Zimprich, A. A novel mutation in the VCP gene (G157R) in a German family with inclusion-body myopathy with Paget disease of bone and frontotemporal dementia. Muscle Nerve 2009, 39, 389–391. [Google Scholar] [CrossRef] [PubMed]
- Ayaki, T.; Ito, H.; Fukushima, H.; Inoue, T.; Kondo, T.; Ikemoto, A.; Asano, T.; Shodai, A.; Fujita, T.; Fukui, S.; et al. Immunoreactivity of valosin-containing protein in sporadic amyotrophic lateral sclerosis and in a case of its novel mutant. Acta Neuropathol. Commun. 2014, 2, 172. [Google Scholar] [CrossRef]
- Hang Wong, T.; Pottier, C.; Hondius, D.; Meeter, L.H.H.; van Rooij, J.G.J.; Melhem, S.; Brain Bank, T.N.; van Minkelen, R.; van Duijn, C.M.; Rozemuller, A.J.M.; et al. Three VCP Mutations in Patients with Frontotemporal Dementia. J. Alzheimers Dis. 2018, 65, 1139–1146. [Google Scholar] [CrossRef]
- Haubenberger, D.; Bittner, R.E.; Rauch-Shorny, S.; Zimprich, F.; Mannhalter, C.; Wagner, L.; Mineva, I.; Vass, K.; Auff, E.; Zimprich, A. Inclusion body myopathy and Paget disease is linked to a novel mutation in the VCP gene. Neurology 2005, 65, 1304–1305. [Google Scholar] [CrossRef]
- van der Zee, J.; Pirici, D.; Van Langenhove, T.; Engelborghs, S.; Vandenberghe, R.; Hoffmann, M.; Pusswald, G.; Van den Broeck, M.; Peeters, K.; Mattheijssens, M.; et al. Clinical heterogeneity in 3 unrelated families linked to VCP p.Arg159His. Neurology 2009, 73, 626–632. [Google Scholar] [CrossRef]
- Bersano, A.; Del Bo, R.; Lamperti, C.; Ghezzi, S.; Fagiolari, G.; Fortunato, F.; Ballabio, E.; Moggio, M.; Candelise, L.; Galimberti, D.; et al. Inclusion body myopathy and frontotemporal dementia caused by a novel VCP mutation. Neurobiol. Aging 2009, 30, 752–758. [Google Scholar] [CrossRef] [PubMed]
- Chan, N.; Le, C.; Shieh, P.; Mozaffar, T.; Khare, M.; Bronstein, J.; Kimonis, V. Valosin-containing protein mutation and Parkinson’s disease. Parkinsonism Relat. Disord. 2012, 18, 107–109. [Google Scholar] [CrossRef] [PubMed]
- de Bot, S.T.; Schelhaas, H.J.; Kamsteeg, E.J.; van de Warrenburg, B.P. Hereditary spastic paraplegia caused by a mutation in the VCP gene. Brain 2012, 135, e223. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.O.; Mandrioli, J.; Benatar, M.; Abramzon, Y.; Van Deerlin, V.M.; Trojanowski, J.Q.; Gibbs, J.R.; Brunetti, M.; Gronka, S.; Wuu, J.; et al. Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 2010, 68, 857–864. [Google Scholar] [CrossRef]
- Kotan, D.; Ozozen Ayas, Z.; Tunca, C.; Gungen, B.D.; Akcimen, F.; Basak, A.N. Phenotypic and genotypic features of patients diagnosed with ALS in the city of Sakarya, Turkey. Acta Neurol. Belg. 2020, 120, 1411–1418. [Google Scholar] [CrossRef]
- Watts, G.D.; Thomasova, D.; Ramdeen, S.K.; Fulchiero, E.C.; Mehta, S.G.; Drachman, D.A.; Weihl, C.C.; Jamrozik, Z.; Kwiecinski, H.; Kaminska, A.; et al. Novel VCP mutations in inclusion body myopathy associated with Paget disease of bone and frontotemporal dementia. Clin. Genet. 2007, 72, 420–426. [Google Scholar] [CrossRef]
- Figueroa-Bonaparte, S.; Hudson, J.; Barresi, R.; Polvikoski, T.; Williams, T.; Topf, A.; Harris, E.; Hilton-Jones, D.; Petty, R.; Willis, T.A.; et al. Mutational spectrum and phenotypic variability of VCP-related neurological disease in the UK. J. Neurol. Neurosurg. Psychiatry 2015, 87, 680–681. [Google Scholar] [CrossRef]
- Peyer, A.K.; Kinter, J.; Hench, J.; Frank, S.; Fuhr, P.; Thomann, S.; Fischmann, A.; Kneifel, S.; Camano, P.; Lopez de Munain, A.; et al. Novel valosin containing protein mutation in a Swiss family with hereditary inclusion body myopathy and dementia. Neuromuscul. Disord. 2013, 23, 149–154. [Google Scholar] [CrossRef]
- Spina, S.; Van Laar, A.D.; Murrell, J.R.; Hamilton, R.L.; Kofler, J.K.; Epperson, F.; Farlow, M.R.; Lopez, O.L.; Quinlan, J.; DeKosky, S.T.; et al. Phenotypic variability in three families with valosin-containing protein mutation. Eur. J. Neurol. 2013, 20, 251–258. [Google Scholar] [CrossRef]
- Levesque, S.; Auray-Blais, C.; Gravel, E.; Boutin, M.; Dempsey-Nunez, L.; Jacques, P.E.; Chenier, S.; Larue, S.; Rioux, M.F.; Al-Hertani, W.; et al. Diagnosis of late-onset Pompe disease and other muscle disorders by next-generation sequencing. Orphanet J. Rare Dis. 2016, 11, 8. [Google Scholar] [CrossRef]
- Abramzon, Y.; Johnson, J.O.; Scholz, S.W.; Taylor, J.P.; Brunetti, M.; Calvo, A.; Mandrioli, J.; Benatar, M.; Mora, G.; Restagno, G.; et al. Valosin-containing protein (VCP) mutations in sporadic amyotrophic lateral sclerosis. Neurobiol. Aging 2012, 33, 2231.e1–2231.e6. [Google Scholar] [CrossRef] [PubMed]
- Liewluck, T.; Milone, M.; Mauermann, M.L.; Castro-Couch, M.; Cerhan, J.H.; Murthy, N.S. A novel VCP mutation underlies scapuloperoneal muscular dystrophy and dropped head syndrome featuring lobulated fibers. Muscle Nerve 2014, 50, 295–299. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.; Hayashi, Y.K.; Mitsuhashi, S.; Goto, K.; Kaneda, D.; Choi, Y.C.; Toyoda, C.; Hieda, S.; Kamiyama, T.; Sato, H.; et al. Characterization of the Asian myopathy patients with VCP mutations. Eur. J. Neurol. 2012, 19, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Hirano, M.; Nakamura, Y.; Saigoh, K.; Sakamoto, H.; Ueno, S.; Isono, C.; Mitsui, Y.; Kusunoki, S. VCP gene analyses in Japanese patients with sporadic amyotrophic lateral sclerosis identify a new mutation. Neurobiol. Aging 2015, 36, 1604.e1–1604.e6. [Google Scholar] [CrossRef]
- Papadopoulos, C.; Kirchner, P.; Bug, M.; Grum, D.; Koerver, L.; Schulze, N.; Poehler, R.; Dressler, A.; Fengler, S.; Arhzaouy, K.; et al. VCP/p97 cooperates with YOD1, UBXD1 and PLAA to drive clearance of ruptured lysosomes by autophagy. EMBO J. 2017, 36, 135–150. [Google Scholar] [CrossRef]
- Kenna, K.P.; McLaughlin, R.L.; Byrne, S.; Elamin, M.; Heverin, M.; Kenny, E.M.; Cormican, P.; Morris, D.W.; Donaghy, C.G.; Bradley, D.G.; et al. Delineating the genetic heterogeneity of ALS using targeted high-throughput sequencing. J. Med. Genet. 2013, 50, 776–783. [Google Scholar] [CrossRef]
- Iossifov, I.; Ronemus, M.; Levy, D.; Wang, Z.; Hakker, I.; Rosenbaum, J.; Yamrom, B.; Lee, Y.H.; Narzisi, G.; Leotta, A.; et al. De novo gene disruptions in children on the autistic spectrum. Neuron 2012, 74, 285–299. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Schrodinger, L.; DeLano, W. PyMOL. Available online: http://www.pymol.org/pymol (accessed on 24 April 2022).
- Pan, M.; Yu, Y.; Ai, H.; Zheng, Q.; Xie, Y.; Liu, L.; Zhao, M. Mechanistic insight into substrate processing and allosteric inhibition of human p97. Nat. Struct. Mol. Biol. 2021, 28, 614–625. [Google Scholar] [CrossRef]
- Giordano, L.; Deceglie, S.; d’Adamo, P.; Valentino, M.L.; La Morgia, C.; Fracasso, F.; Roberti, M.; Cappellari, M.; Petrosillo, G.; Ciaravolo, S.; et al. Cigarette toxicity triggers Leber’s hereditary optic neuropathy by affecting mtDNA copy number, oxidative phosphorylation and ROS detoxification pathways. Cell Death Dis. 2015, 6, e2021. [Google Scholar] [CrossRef]
- Pfeffer, G.; Cote, H.C.; Montaner, J.S.; Li, C.C.; Jitratkosol, M.; Mezei, M.M. Ophthalmoplegia and ptosis: Mitochondrial toxicity in patients receiving HIV therapy. Neurology 2009, 73, 71–72. [Google Scholar] [CrossRef] [PubMed]
- Pfeffer, G.; Sirrs, S.; Wade, N.K.; Mezei, M.M. Multisystem disorder in late-onset chronic progressive external ophthalmoplegia. Can. J. Neurol. Sci. 2011, 38, 119–123. [Google Scholar] [CrossRef] [PubMed]
- Blacher, E.; Bashiardes, S.; Shapiro, H.; Rothschild, D.; Mor, U.; Dori-Bachash, M.; Kleimeyer, C.; Moresi, C.; Harnik, Y.; Zur, M.; et al. Potential roles of gut microbiome and metabolites in modulating ALS in mice. Nature 2019, 572, 474–480. [Google Scholar] [CrossRef] [PubMed]
- Mehta, S.G.; Watts, G.D.; Adamson, J.L.; Hutton, M.; Umberger, G.; Xiong, S.; Ramdeen, S.; Lovell, M.A.; Kimonis, V.E.; Smith, C.D. APOE is a potential modifier gene in an autosomal dominant form of frontotemporal dementia (IBMPFD). Genet. Med. 2007, 9, 9–13. [Google Scholar] [CrossRef]
- Kirby, A.E.; Kimonis, V.; Kompoliti, K. Ataxia and Parkinsonism in a Woman With a VCP Variant and Long-Normal Repeats in the SCA2 Allele. Neurol. Genet. 2021, 7, e595. [Google Scholar] [CrossRef]
- Chalmers, M.R.; Kim, J.; Kim, N.C. Eip74EF is a dominant modifier for ALS-FTD linked mutant VCP phenotypes in Drosophila, but not miR-34. Biorxiv 2020. [Google Scholar] [CrossRef]
- Ikenaga, C.; Findlay, A.R.; Goyal, N.A.; Robinson, S.; Cauchi, J.; Hussain, Y.; Wang, L.H.; Kershen, J.C.; Beson, B.A.; Wallendorf, M.; et al. Clinical utility of anti-cytosolic 5’-nucleotidase 1A antibody in idiopathic inflammatory myopathies. Ann. Clin. Transl. Neurol. 2021, 8, 571–578. [Google Scholar] [CrossRef]
- Posey, J.E.; Harel, T.; Liu, P.; Rosenfeld, J.A.; James, R.A.; Coban Akdemir, Z.H.; Walkiewicz, M.; Bi, W.; Xiao, R.; Ding, Y.; et al. Resolution of Disease Phenotypes Resulting from Multilocus Genomic Variation. N. Engl. J. Med. 2017, 376, 21–31. [Google Scholar] [CrossRef]
- Estiar, M.A.; Yu, E.; Haj Salem, I.; Ross, J.P.; Mufti, K.; Akcimen, F.; Leveille, E.; Spiegelman, D.; Ruskey, J.A.; Asayesh, F.; et al. Evidence for Non-Mendelian Inheritance in Spastic Paraplegia 7. Mov. Disord. 2021, 36, 1664–1675. [Google Scholar] [CrossRef]
- Weihl, C.C.; Pestronk, A.; Kimonis, V.E. Valosin-containing protein disease: Inclusion body myopathy with Paget’s disease of the bone and fronto-temporal dementia. Neuromuscul. Disord. 2009, 19, 308–315. [Google Scholar] [CrossRef]
- Bugiardini, E.; Morrow, J.M.; Shah, S.; Wood, C.L.; Lynch, D.S.; Pitmann, A.M.; Reilly, M.M.; Houlden, H.; Matthews, E.; Parton, M.; et al. The Diagnostic Value of MRI Pattern Recognition in Distal Myopathies. Front. Neurol. 2018, 9, 456. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Manera, J.; Llauger, J.; Gallardo, E.; Illa, I. Muscle MRI in muscular dystrophies. Acta Myol. 2015, 34, 95–108. [Google Scholar] [PubMed]
- Papadimas, G.K.; Paraskevas, G.P.; Zambelis, T.; Karagiaouris, C.; Bourbouli, M.; Bougea, A.; Walter, M.C.; Schumacher, N.U.; Krause, S.; Kapaki, E. The multifaceted clinical presentation of VCP-proteinopathy in a Greek family. Acta Myol. 2017, 36, 203–206. [Google Scholar] [PubMed]
- Paul Tuck, S.; Layfield, R.; Walker, J.; Mekkayil, B.; Francis, R. Adult Paget’s disease of bone: A review. Rheumatology 2017, 56, 2050–2059. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Grimes, D.A.; Barclay, C.L.; Duff, J.; Furukawa, Y.; Lang, A.E. Phenocopies in a large GCH1 mutation positive family with dopa responsive dystonia: Confusing the picture? J. Neurol. Neurosurg. Psychiatry 2002, 72, 801–804. [Google Scholar] [CrossRef] [PubMed]
- Garrison, N.A.; Brothers, K.B.; Goldenberg, A.J.; Lynch, J.A. Genomic Contextualism: Shifting the Rhetoric of Genetic Exceptionalism. Am. J. Bioeth. 2019, 19, 51–63. [Google Scholar] [CrossRef]
- Laberge, A.M.; Richer, J.; Ravitsky, V. Toward Broader Genetic Contextualism: Genetic Testing Enters the Age of Evidence-Based Medicine. Am. J. Bioeth. 2019, 19, 77–79. [Google Scholar] [CrossRef]
- Slade, I.; Riddell, D.; Turnbull, C.; Hanson, H.; Rahman, N.; programme, M.C.G. Development of cancer genetic services in the UK: A national consultation. Genome Med. 2015, 7, 18. [Google Scholar] [CrossRef]
- Cohen, S.A.; Bradbury, A.; Henderson, V.; Hoskins, K.; Bednar, E.; Arun, B.K. Genetic Counseling and Testing in a Community Setting: Quality, Access, and Efficiency. Am. Soc. Clin. Oncol. Educ. Book 2019, 39, e33–e44. [Google Scholar] [CrossRef]
- Yuen, J.; Cousens, N.; Barlow-Stewart, K.; O’Shea, R.; Andrews, L. Online BRCA1/2 screening in the Australian Jewish community: A qualitative study. J. Community Genet. 2020, 11, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Scuffham, T.M.; MacMillan, J.C. Huntington disease: Who seeks presymptomatic genetic testing, why and what are the outcomes? J. Genet. Couns. 2014, 23, 754–761. [Google Scholar] [CrossRef] [PubMed]
- Godino, L.; Turchetti, D.; Jackson, L.; Hennessy, C.; Skirton, H. Impact of presymptomatic genetic testing on young adults: A systematic review. Eur. J. Hum. Genet. 2016, 24, 496–503. [Google Scholar] [CrossRef] [PubMed]
- Tilleras, K.H.; Kjoelaas, S.H.; Dramstad, E.; Feragen, K.B.; von der Lippe, C. Psychological reactions to predictive genetic testing for Huntington’s disease: A qualitative study. J. Genet. Couns. 2020, 29, 1093–1105. [Google Scholar] [CrossRef]
- Surampalli, A.; Khare, M.; Kubrussi, G.; Wencel, M.; Tanaja, J.; Donkervoort, S.; Osann, K.; Simon, M.; Wallace, D.; Smith, C.; et al. Psychological Impact of Predictive Genetic Testing in VCP Inclusion Body Myopathy, Paget Disease of Bone and Frontotemporal Dementia. J. Genet. Couns. 2015, 24, 842–850. [Google Scholar] [CrossRef]
- Clement, S.; Gargiulo, M.; Feingold, J.; Durr, A. Guidelines for presymptomatic testing for Huntington’s disease: Past, present and future in France. Rev. Neurol. 2015, 171, 572–580. [Google Scholar] [CrossRef]
- Dalpe, G.; Pinkesz, M.; Oliviero, E.; Tolymbek, M.; Joly, Y. Genetic discrimination views in online discussion forums: Perspectives from Canadian forumites. J. Genet. Couns. 2021, 30, 1613–1628. [Google Scholar] [CrossRef]
- Burnett-Hartman, A.N.; Blum-Barnett, E.; Carroll, N.M.; Madrid, S.D.; Jonas, C.; Janes, K.; Alvarado, M.; Bedoy, R.; Paolino, V.; Aziz, N.; et al. Return of Research-Related Genetic Test Results and Genetic Discrimination Concerns: Facilitators and Barriers of Genetic Research Participation in Diverse Groups. Public Health Genomics. 2020, 23, 59–68. [Google Scholar] [CrossRef]
- Joly, Y.; Dalpe, G.; Dupras, C.; Beviere-Boyer, B.; de Paor, A.; Dove, E.S.; Granados Moreno, P.; Ho, C.W.L.; Ho, C.H.; K, O.C.; et al. Establishing the International Genetic Discrimination Observatory. Nat. Genet. 2020, 52, 466–468. [Google Scholar] [CrossRef]
- de Paor, A. Genetic Discrimination: A Case for a European Legislative Response? Eur. J. Health Law 2017, 24, 135–159. [Google Scholar] [CrossRef]
- Clifton, J.M.; VanBeuge, S.S.; Mladenka, C.; Wosnik, K.K. The Genetic Information Nondiscrimination Act 2008: What clinicians should understand. J. Am. Acad. Nurse Pract. 2010, 22, 246–249. [Google Scholar] [CrossRef] [PubMed]
| Localization | Phenotype or Disease | Frequency in VCP-MSP |
|---|---|---|
| Central nervous system | Frontotemporal dementia | 30% |
| Alzheimer disease | 2% | |
| Parkinson disease | 4% | |
| Spastic paraplegia | Case reports | |
| Peripheral nervous system | Inclusion body myopathy | 90% |
| Amyotrophic lateral sclerosis | 10% | |
| Charcot-Marie-Tooth disease | Case reports | |
| Sensory polyneuropathy | Case reports | |
| Non-neurological | Paget disease of bone | 40% |
| Cardiomyopathy | Uncertain |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pfeffer, G.; Lee, G.; Pontifex, C.S.; Fanganiello, R.D.; Peck, A.; Weihl, C.C.; Kimonis, V. Multisystem Proteinopathy Due to VCP Mutations: A Review of Clinical Heterogeneity and Genetic Diagnosis. Genes 2022, 13, 963. https://doi.org/10.3390/genes13060963
Pfeffer G, Lee G, Pontifex CS, Fanganiello RD, Peck A, Weihl CC, Kimonis V. Multisystem Proteinopathy Due to VCP Mutations: A Review of Clinical Heterogeneity and Genetic Diagnosis. Genes. 2022; 13(6):963. https://doi.org/10.3390/genes13060963
Chicago/Turabian StylePfeffer, Gerald, Grace Lee, Carly S. Pontifex, Roberto D. Fanganiello, Allison Peck, Conrad C. Weihl, and Virginia Kimonis. 2022. "Multisystem Proteinopathy Due to VCP Mutations: A Review of Clinical Heterogeneity and Genetic Diagnosis" Genes 13, no. 6: 963. https://doi.org/10.3390/genes13060963
APA StylePfeffer, G., Lee, G., Pontifex, C. S., Fanganiello, R. D., Peck, A., Weihl, C. C., & Kimonis, V. (2022). Multisystem Proteinopathy Due to VCP Mutations: A Review of Clinical Heterogeneity and Genetic Diagnosis. Genes, 13(6), 963. https://doi.org/10.3390/genes13060963







