BRCA1/2 Mutations in Vietnamese Patients with Hereditary Breast and Ovarian Cancer Syndrome
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Sample Collection
2.2. Genetic Testing
2.3. Statistical Analysis
2.3.1. Survival Analysis for Breast Cancer and Ovarian Cancer Patients
2.3.2. Breast and Ovarian Risk Prediction
2.3.3. Breast Cancer Familial Relative Risks
3. Results
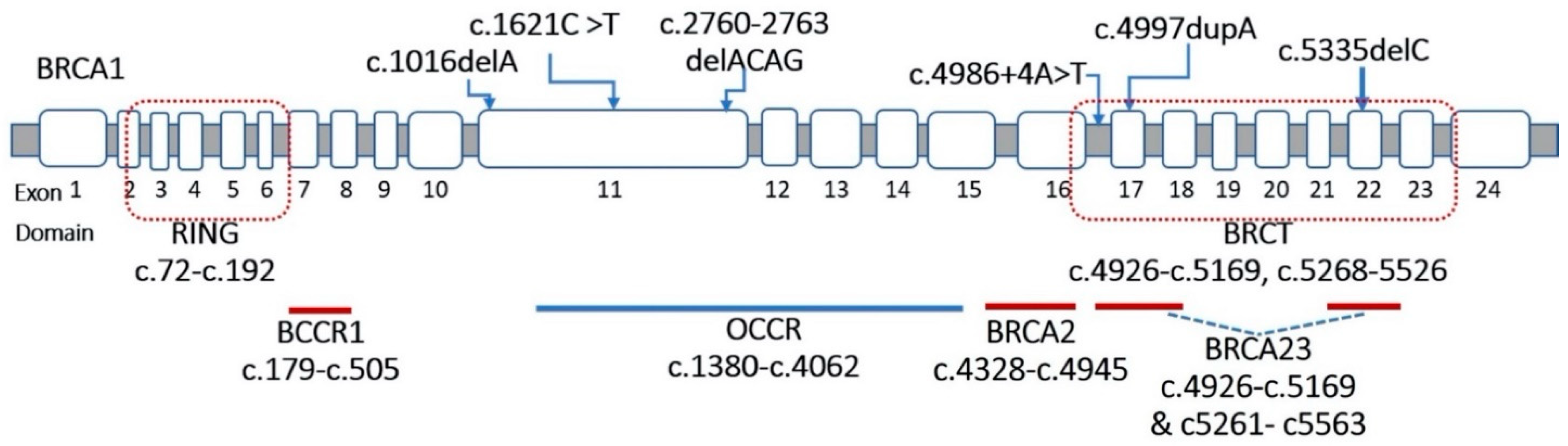
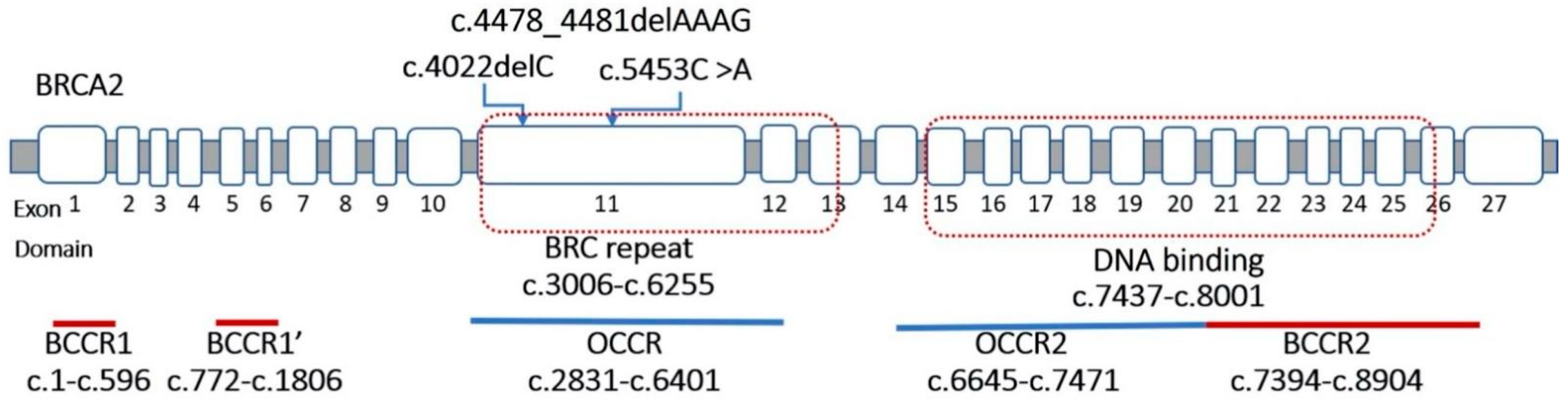
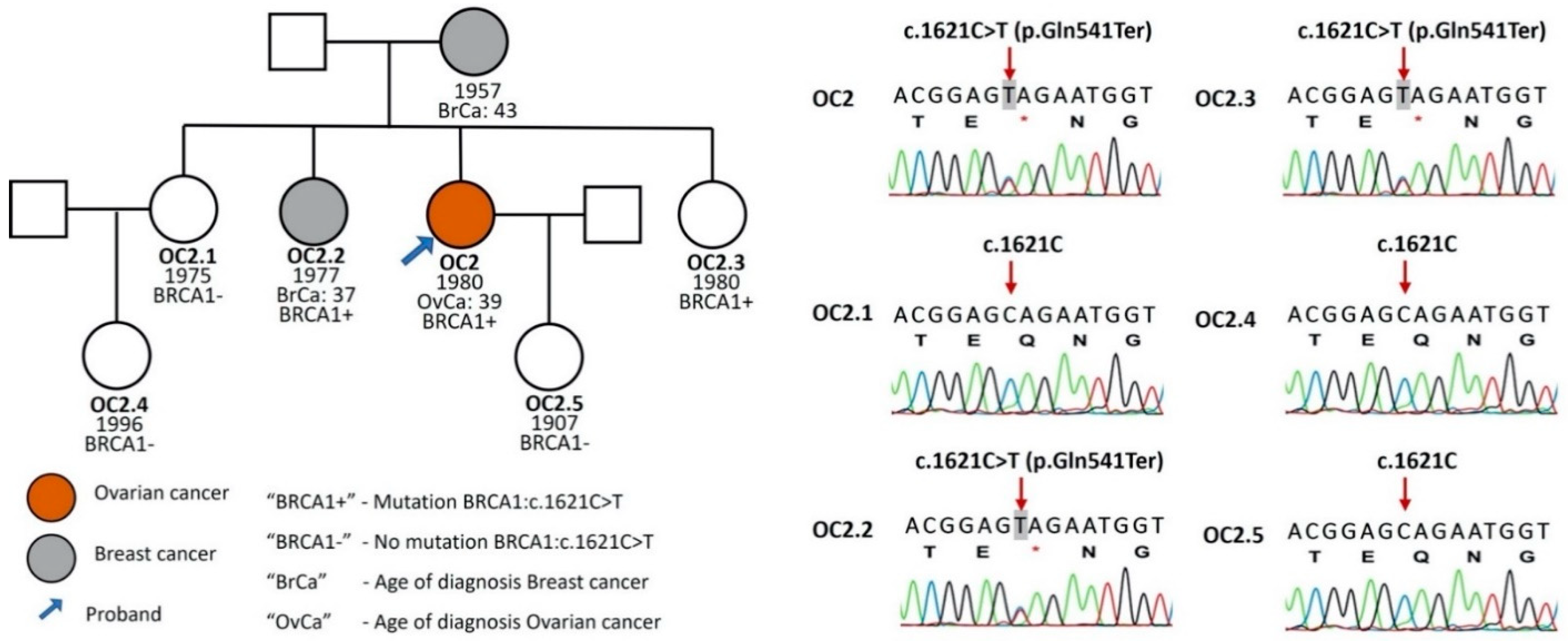
3.1. Detection of BRCA1 and BRCA2 Variants
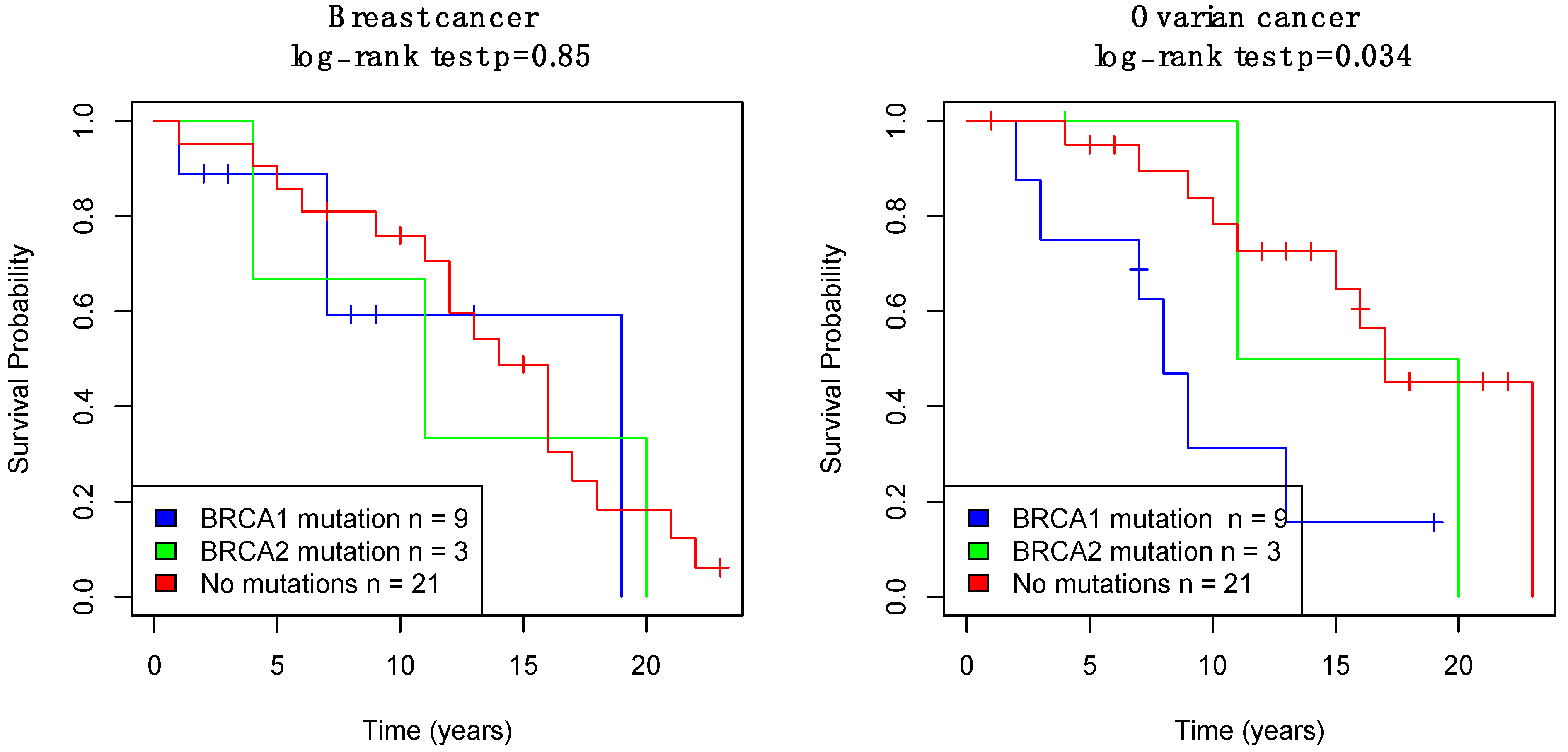
3.2. Survival Analysis for Breast Cancer and Ovarian Cancer Patients
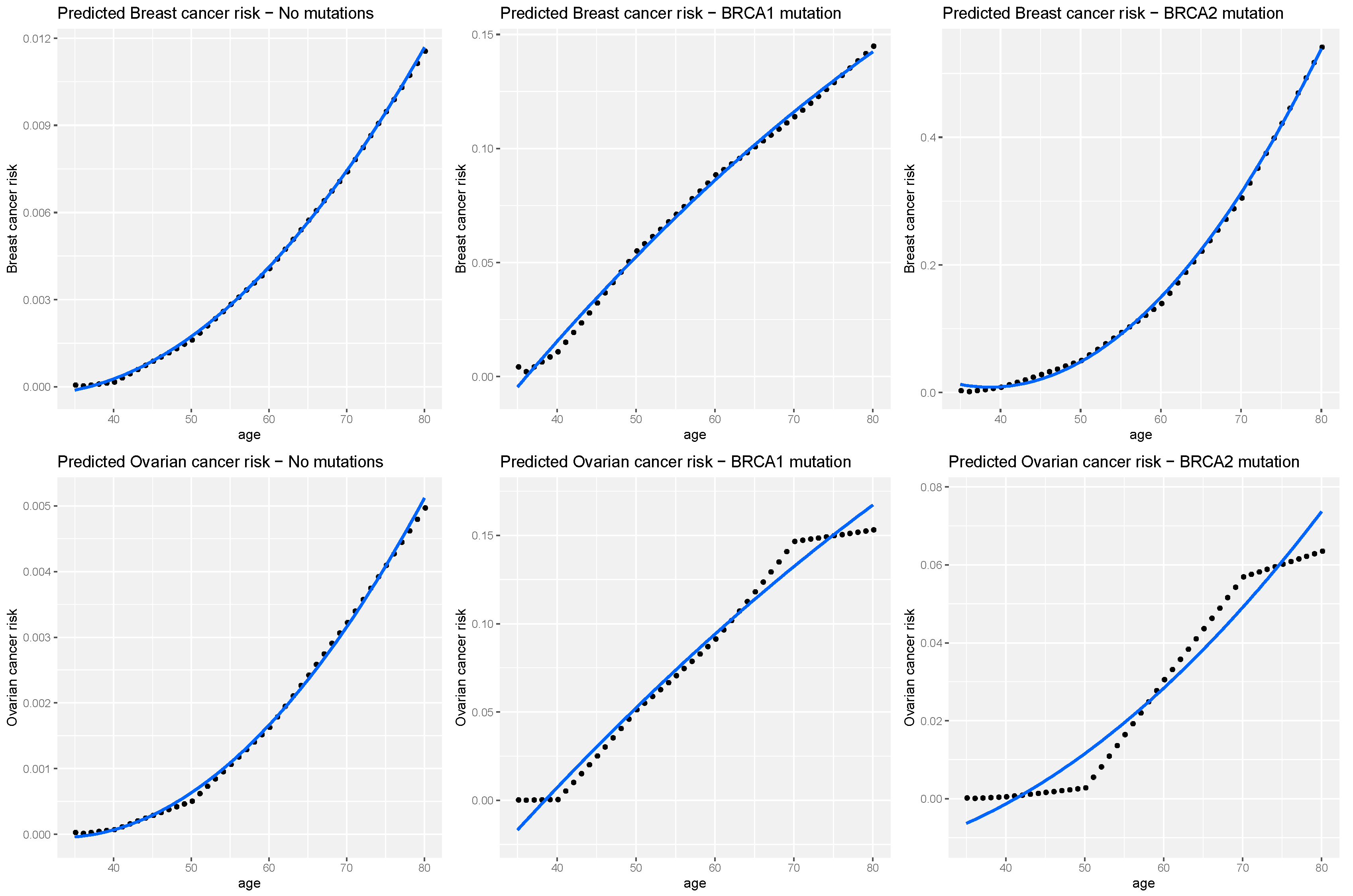
3.3. Cancer Risks
3.4. Breast Cancer Familial Relative Risks
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Neff, R.T.; Senter, L.; Salani, R. BRCA mutation in ovarian cancer: Testing, implications and treatment considerations. Ther. Adv. Med. Oncol. 2017, 9, 519–531. [Google Scholar] [CrossRef] [PubMed]
- Finch, A.P.; Lubinski, J.; Møller, P.; Singer, C.F.; Karlan, B.; Senter, L.; Rosen, B.; Maehle, L.; Ghadirian, P.; Cybulski, C.; et al. Impact of oophorectomy on cancer incidence and mortality in women with a BRCA1 or BRCA2 mutation. J. Clin. Oncol. 2014, 32, 1547–1553. [Google Scholar] [CrossRef] [PubMed]
- Vu, H.A.; Phu, N.D.; Khuong, L.T.; Hoa, P.H.; Nhu, B.T.H.; Nhan, V.T.; Thanh, L.Q.; Sinh, N.D.; Chi, H.T.; Quan, N.D.; et al. Recurrent BRCA1 Mutation, but no BRCA2 Mutation, in Vietnamese Patients with Ovarian Carcinoma Detected with Next Generation Sequencing. Asian Pac. J. Cancer Prev. 2020, 21, 2331–2335. [Google Scholar] [CrossRef]
- Van Thuan, T.; Van Chu, N.; Khoa, P.H.; Quang, N.T.; Van Tu, D.; Tho, N.T.Q.; Huyen, P.T.; Ha, B.H.; Han, P.T.; Long, D.M.; et al. A Novel BRCA1 Gene Mutation Detected With Breast Cancer in a Vietnamese Family by Targeted Next-Generation Sequencing: A Case Report. Breast Cancer 2020, 14, 1178223420901555. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Peto, J.; Collins, N.; Barfoot, R.; Seal, S.; Warren, W.; Rahman, N.; Easton, D.F.; Evans, C.; Deacon, J.; Stratton, M.R. Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J. Natl. Cancer Inst. 1999, 91, 943–949. [Google Scholar] [CrossRef]
- Tilanus-Linthorst, M.M.; Lingsma, H.F.; Evans, D.G.; Thompson, D.; Kaas, R.; Manders, P.; van Asperen, C.J.; Adank, M.; Hooning, M.J.; Kwan Lim, G.E.; et al. Optimal age to start preventive measures in women with BRCA1/2 mutations or high familial breast cancer risk. Int. J. Cancer 2013, 133, 156–163. [Google Scholar] [CrossRef]
- van der Groep, P.; Bouter, A.; van der Zanden, R.; Siccama, I.; Menko, F.H.; Gille, J.J.P.; van Kalken, C.; van der Wall, E.; Verheijen, R.H.M.; van Diest, P.J. Distinction between hereditary and sporadic breast cancer on the basis of clinicopathological data. J. Clin. Pathol. 2006, 59, 611–617. [Google Scholar] [CrossRef]
- Antoniou, A.C.; Cunningham, A.P.; Peto, J.; Evans, D.G.; Lalloo, F.; Narod, S.A.; Risch, A.H.; Eyfjord, J.E.; Hopper, J.L.; Southey, M.C.; et al. The BOADICEA model of genetic susceptibility to breast and ovarian cancers: Updates and extensions. Br. J. Cancer 2008, 98, 1457–1466. [Google Scholar] [CrossRef]
- Lee, A.; Mavaddat, N.; Wilcox, A.N.; Cunningham, A.P.; Carver, T.; Hartley, S.; de Villiers, C.B.; Izquierdo, A.; Simard, J.; Schmidt, M.K.; et al. Correction: BOADICEA: A comprehensive breast cancer risk prediction model incorporating genetic and nongenetic risk factors. Genet. Med. 2019, 21, 1462. [Google Scholar] [CrossRef] [PubMed]
- Antoniou, A.C.; Pharoah, P.D.P.; McMullan, G.; Day, N.E.; Stratton, M.R.; Peto, J.; Ponder, B.J.; Easton, D.F. A comprehensive model for familial breast cancer incorporating BRCA1, BRCA2 and other genes. Br. J. Cancer 2002, 86, 76–83. [Google Scholar] [CrossRef] [PubMed]
- Antoniou, A.C.; Pharoah, P.P.D.; Smith, P.; Easton, D.F. The BOADICEA model of genetic susceptibility to breast and ovarian cancer. Br. J. Cancer 2004, 91, 1580–1590. [Google Scholar] [CrossRef] [PubMed]
- Hogervorst, F.B.; Cornelis, R.S.; Bout, M.; Van Vliet, M.; Oosterwijk, J.C.; Olmer, R.; Bakker, B.; Klijn, J.G.; Vasen, H.F.; Meijers-Heijboer, H.; et al. Rapid detection of BRCA1 mutations by the protein truncation test. Nat. Genet. 1995, 10, 208–212. [Google Scholar] [CrossRef] [PubMed]
- Debatin, I.; Tonin, P.; Royer-Pokora, B.; Dong, J.; Chang-Claude, J.; Wu, Y.; Schumacher, V. A high proportion of mutations in the BRCA1 gene in German breast/ovarian cancer families with clustering of mutations in the 3′ third of the gene. Hum. Genet. 1998, 103, 154–161. [Google Scholar] [CrossRef]
- van der Hout, A.H.; van den Ouweland, A.M.; van der Luijt, R.B.; Gille, H.J.; Bodmer, D.; Brüggenwirth, H.; Mulder, I.M.; van der Vlies, P.; Elfferich, P.; Huisman, M.T.; et al. A DGGE system for comprehensive mutation screening of BRCA1 and BRCA2: Application in a Dutch cancer clinic setting. Hum. Mutat. 2006, 27, 654–666. [Google Scholar] [CrossRef] [PubMed]
- Konecny, M.; Milly, M.; Zavodna, K.; Weismanova, E.; Gregorova, J.; Mlkva, I.; Ilencikova, D.; Kausitz, J.; Bartosova, Z. Comprehensive genetic characterization of hereditary breast/ovarian cancer families from Slovakia. Breast Cancer Res. Treat. 2011, 126, 119–130. [Google Scholar] [CrossRef][Green Version]
- Ang, P.; Lim, I.H.; Lee, T.-C.; Luo, J.-T.; Ong, D.C.; Tan, P.H.; Lee, A.S. BRCA1 and BRCA2 mutations in an Asian clinic-based population detected using a comprehensive strategy. Cancer Epidemiol. Biomarkers Prev. 2007, 16, 2276–2284. [Google Scholar] [CrossRef]
- Vehmanen, P.; Friedman, L.S.; Eerola, H.; McClure, M.; Ward, B.; Sarantaus, L.; Kainu, T.; Syrjäkoski, K.; Pyrhönen, S.; Kallioniemi, O.-P.; et al. Low proportion of BRCA1 and BRCA2 mutations in Finnish breast cancer families: Evidence for additional susceptibility genes. Hum. Mol. Genet. 1997, 6, 2309–2315. [Google Scholar] [CrossRef][Green Version]
- Matsuda, M.L.D.L.; Liede, A.; Kwan, E.; Mapua, C.A.; Cutiongco, E.M.C.; Tan, A.; Narod, S.A. BRCA1 and BRCA2 mutations among breast cancer patients from the Philippines. Int. J. Cancer 2002, 98, 596–603. [Google Scholar] [CrossRef]
- Tavtigian, S.V.; Simard, J.; Rommens, J.M.; Couch, F.J.; Shattuck-Eidens, D.; Neuhausen, S.L.; Merajver, S.D.; Thorlacius, S.; Offit, K.; Stoppalyonnet, D.; et al. The complete BRCA2 gene and mutations in chromosome 13q-linked kindreds. Nat. Genet. 1996, 12, 333–337. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Yang, J.; Li, L.; Cao, D.; Yu, M.; Shen, K. Germline and somatic mutations in homologous recombination genes among Chinese ovarian cancer patients detected using next-generation sequencing. J Gynecol. Oncol. 2017, 28, e39. [Google Scholar] [CrossRef] [PubMed]
- Ginsburg, O.; Dinh, N.; To, T.; Quang, L.; Linh, N.; Duong, B.; Royer, R.; Llacuachaqui, M.; Tulman, A.; Vichodez, G.; et al. Family history, BRCA mutations and breast cancer in Vietnamese women. Clin. Genet. 2011, 80, 89–92. [Google Scholar] [CrossRef] [PubMed]
- Frank, T.S.; Deffenbaugh, A.M.; Reid, J.E.; Hulick, M.; Ward, B.E.; Lingenfelter, B.; Gumpper, K.L.; Scholl, T.; Tavtigian, S.V.; Pruss, D.R.; et al. Clinical characteristics of individuals with germline mutations in BRCA1 and BRCA2: Analysis of 10,000 individuals. J. Clin. Oncol. 2002, 20, 1480–1490. [Google Scholar] [CrossRef]
- Powell, C.B.; Kenley, E.; Chen, L.-M.; Crawford, B.; McLennan, J.; Zaloudek, C.; Komaromy, M.; Beattie, M.; Ziegler, J. Risk-reducing salpingo-oophorectomy in BRCA mutation carriers: Role of serial sectioning in the detection of occult malignancy. J. Clin. Oncol. 2005, 23, 127–132. [Google Scholar] [CrossRef]
- Findlay, G.M.; Daza, R.M.; Martin, B.; Zhang, M.D.; Leith, A.P.; Gasperini, M.; Janizek, J.D.; Huang, X.; Starita, L.M.; Shendure, J. Accurate classification of BRCA1 variants with saturation genome editing. Nature 2018, 562, 217–222. [Google Scholar] [CrossRef]
- Rebbeck, T.R.; Mitra, N.; Wan, F.; Sinilnikova, O.M.; Healey, S.; McGuffog, L.; Mazoyer, S.; Chenevix-Trench, G.; Easton, D.F.; Antoniou, A.C.; et al. Association of type and location of BRCA1 and BRCA2 mutations with risk of breast and ovarian cancer. JAMA 2015, 313, 1347–1361. [Google Scholar] [CrossRef]
- Kurian, A.W.; Gong, G.D.; Chun, N.M.; Mills, M.A.; Staton, A.D.; Kingham, K.E.; Crawford, B.B.; Lee, R.; Chan, S.; Donlon, S.S.; et al. Performance of BRCA1/2 mutation prediction models in Asian Americans. J. Clin. Oncol. 2008, 26, 4752–4758. [Google Scholar] [CrossRef]
- NCCN. NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) Genetic/Familial High-Risk Assecessment: Breast and Ovarian Version 2; Cold Spring Publishing LLC: Long Island, NY, USA, 2017; pp. 1–77. [Google Scholar]
| Characteristics | Patients with BRCA1/2 Mutations | Non-Mutation Patient | Total | ||
|---|---|---|---|---|---|
| BRCA1 Mutation Patient | BRCA2 Mutation Patient | Total | |||
| N = 9 (27.3%) | N = 3 (9.1%) | N = 12 (36.4%) | N = 21 (64.6%) | N = 33 (100%) | |
| Mean of diagnosed age (years) | 48.0 | 61.0 | 51.25 | 54.86 | 53.55 |
| HBOC | |||||
| FH (OC and/or BC) only | 8 | 1 | 9 | 16 | 25 |
| Duplex cancer only | 0 | 2 | 2 | 4 | 6 |
| Both (FH & Duplex cancer) | 1 | 0 | 1 | 1 | 2 |
| No. | HGVS cDNA | HGVS Protein | Local | BCCR/ OCCR | Domain /Binding | Type | Sign. | Ref. |
|---|---|---|---|---|---|---|---|---|
| 01 | c.1016delA | p.Lys339ArgfsTer2 | BRCA1-Exon 11 | − | − | F | P | [13] |
| 02 | c.1621C > T | p.Gln541Ter | BRCA1-Exon 11 | OCCR1 | − | N | P | [14] |
| 03 | c.2760_2763delACAG | p.Thr922LeufsTer77 | BRCA1-Exon 11 | OCCR3 | − | F | P | [15] |
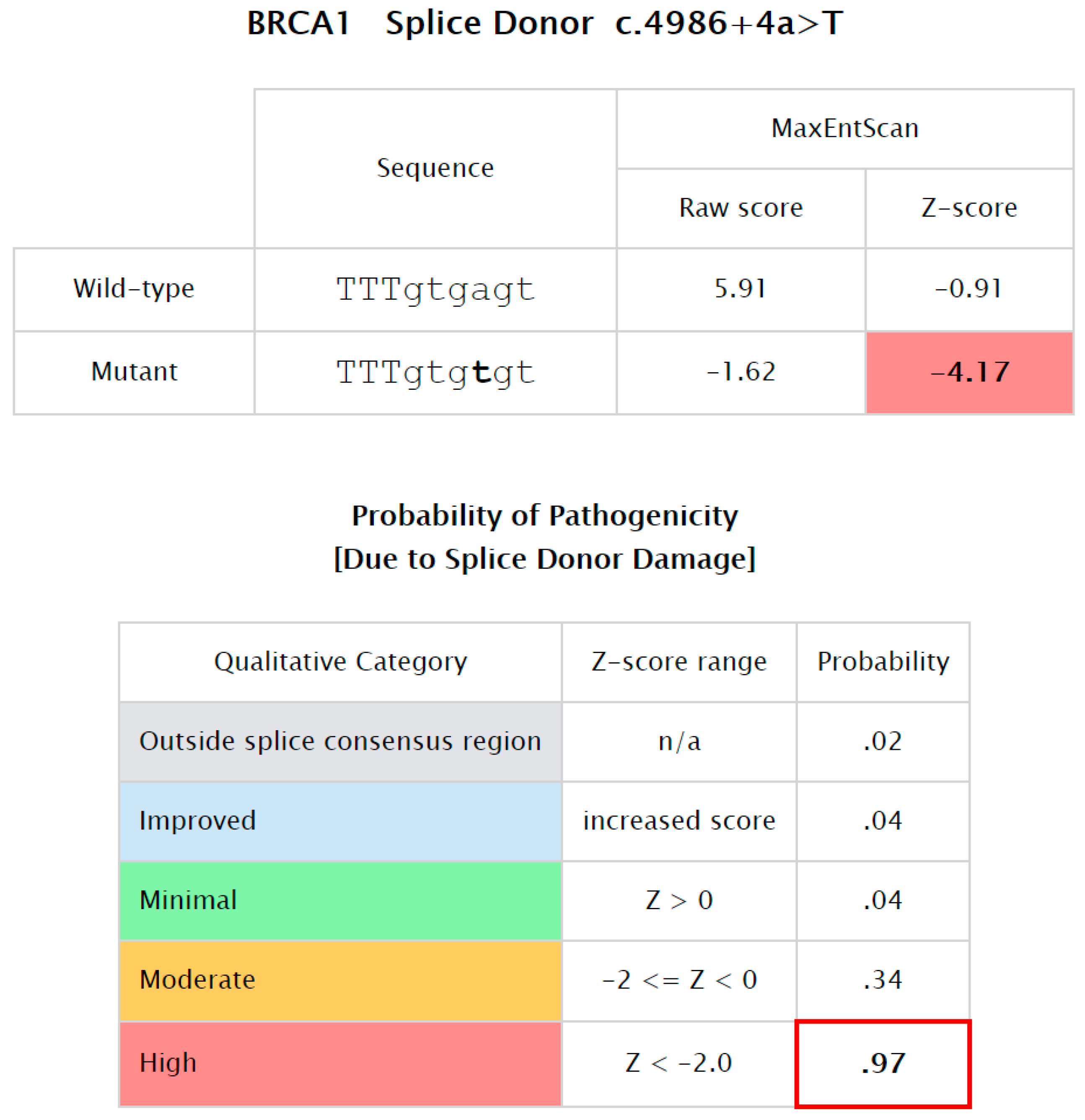
| 04 | c.4986 + 4A > T | IVS16+4A > T | BRCA1-Intron 16 | BCCR23 | BRCT/BACH1 | Ss | P | [16] |
| 05 | c.4997dupA | p.Tyr1666Ter | BRCA1-Exon 17 | BCCR23 | BRCT/BACH1 | N | P | CS |
| 06 | c.5068A > C | p.Lys1690Gln | BRCA1-Exon 17 | BCCR23 | BRCT/BACH1 | M | VUS | [17] |
| 07 | c.5251C > T | p.Arg1751Ter | BRCA1-Exon 20 | − | − | N | P | [18] |
| 08 | c.5335delC | p.Gln1779AsnfsTer14 | BRCA1-Exon 22 | BCCR23 | BRCT/BACH1 | F | P | [19] |
| 09 | c.4022delC | p.Ser1341Ter | BRCA2-Exon 11 | OCCR1 | − | N | P | CS |
| 10 | c.4478_4481delAAAG | p.Glu1493ValfsTer10 | BRCA2-Exon 11 | OCCR1 | − | F | P | [20] |
| 11 | c.5453C > A | p.Ser1818Ter | BRCA2-Exon 11 | OCCR1 | − | N | P | [21] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Le, T.-N.N.; Tran, V.-K.; Nguyen, T.-T.; Vo, N.S.; Hoang, T.H.; Vo, H.-L.; Nguyen, T.-H.T.; Nguyen, P.-D.; Nguyen, V.-T.; Ta, T.-V.; et al. BRCA1/2 Mutations in Vietnamese Patients with Hereditary Breast and Ovarian Cancer Syndrome. Genes 2022, 13, 268. https://doi.org/10.3390/genes13020268
Le T-NN, Tran V-K, Nguyen T-T, Vo NS, Hoang TH, Vo H-L, Nguyen T-HT, Nguyen P-D, Nguyen V-T, Ta T-V, et al. BRCA1/2 Mutations in Vietnamese Patients with Hereditary Breast and Ovarian Cancer Syndrome. Genes. 2022; 13(2):268. https://doi.org/10.3390/genes13020268
Chicago/Turabian StyleLe, Trong-Nhan N., Van-Khanh Tran, Thu-Thuy Nguyen, Nam S. Vo, Tham H. Hoang, Hoang-Long Vo, Thanh-Hai T. Nguyen, Phuoc-Dung Nguyen, Viet-Tien Nguyen, Thanh-Van Ta, and et al. 2022. "BRCA1/2 Mutations in Vietnamese Patients with Hereditary Breast and Ovarian Cancer Syndrome" Genes 13, no. 2: 268. https://doi.org/10.3390/genes13020268
APA StyleLe, T.-N. N., Tran, V.-K., Nguyen, T.-T., Vo, N. S., Hoang, T. H., Vo, H.-L., Nguyen, T.-H. T., Nguyen, P.-D., Nguyen, V.-T., Ta, T.-V., & Tran, H.-T. (2022). BRCA1/2 Mutations in Vietnamese Patients with Hereditary Breast and Ovarian Cancer Syndrome. Genes, 13(2), 268. https://doi.org/10.3390/genes13020268







