40 Years of RAS—A Historic Overview
Abstract
1. The Discovery of RAS Genes: From Viruses to Humans and Beyond
1.1. The Retroviral RAS Oncogenes
1.2. The Human RAS Oncogenes
1.3. The RAS Gene Products
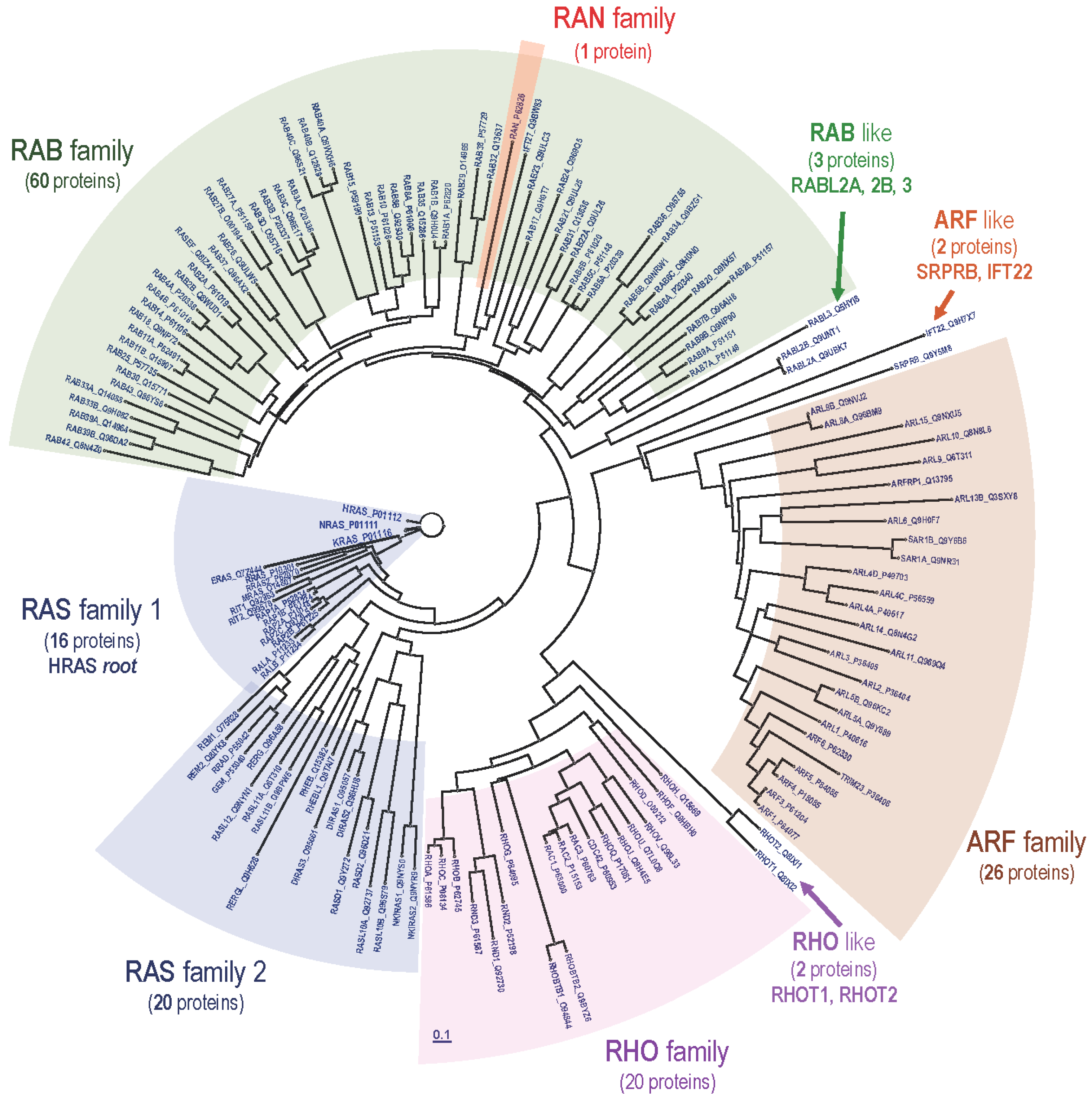
1.4. The Universal RAS Superfamily in Eukaryotes
2. The Progress of RAS Biology and Signaling during the 20th Century
2.1. Growth Factors and Their Receptors Activate Cellular RAS
2.2. RAS Activation. The Guanine Nucleotide Exchange Factors (GEFs)
2.3. RAS Inactivation. The GTPase Activating Proteins (GAPs)
2.4. The Link Between Surface Receptors and RAS Activation. The GRB2 Adaptor Proteins
2.5. RAS Functional Domains and Three-Dimensional Structure
2.6. RAS Downstream Effectors and Signaling Pathways
3. Further Advances on RAS Signaling during the 21st Century
3.1. RAS Regulation Through Covalent Modifications
3.2. RAS Inter-Molecular Interactions
3.3. Positive Feedback Loops of RAS Activation
3.4. RAS Signaling from Endomembranes
3.5. Regulation of RAS Expression by Micro RNAs
4. RAS in Physiology and Pathology
4.1. Central Role of RAS Signaling in Physiological Cellular Processes
4.2. Aberrant RAS Signaling in Cancer
4.3. Defective RAS Signaling in Developmental Syndromes
5. Animal Models for the Analysis of RAS Function
5.1. Transgenic Mice
5.2. Knockout Mice
5.3. Functional Specificity/Redundancy of RAS Isoforms
5.4. Mouse Models of RAS-Driven Cancer
5.5. Animal Models of Human RASopathies
6. Therapeutic Approaches to RAS-Driven Cancer
6.1. Blocking RAS Location at the Plasma Membrane
6.2. Inhibiting RAS Downstream Signaling
6.3. Direct RAS Inhibitors
6.4. Future Perspectives on RAS Therapy
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Harvey, J.J. An Unidentified Virus which causes the Rapid Production of Tumours in Mice. Nature 1964, 204, 1104–1105. [Google Scholar] [CrossRef]
- Kirsten, W.H.; Mayer, L.A. Morphologic Responses to a Murine Erythroblastosis Virus. J. Natl. Cancer Inst. 1967, 39, 311–335. [Google Scholar] [CrossRef]
- Ellis, R.W.; Defeo, D.; Shih, T.Y.; Gonda, M.A.; Young, H.A.; Tsuchida, N.; Lowy, D.R.; Scolnick, E.M. The p21 src genes of Harvey and Kirsten sarcoma viruses originate from divergent members of a family of normal vertebrate genes. Nature 1981, 292, 506–511. [Google Scholar] [CrossRef]
- Langbeheim, H.; Shih, T.Y.; Scolnick, E.M. Identification of a normal vertebrate cell protein related to the p21 src of harvey murine sarcoma virus. Virology 1980, 106, 292–300. [Google Scholar] [CrossRef]
- Shih, T.Y.; Papageorge, A.G.; Stokes, P.E.; Weeks, M.O.; Scolnick, E.M. Guanine nucleotide-binding and autophosphorylating activities associated with the p21src protein of Harvey murine sarcoma virus. Nature 1980, 287, 686–691. [Google Scholar] [CrossRef]
- Andersen, P.R.; Devare, S.G.; Tronick, S.R.; Ellis, R.W.; Aaronson, S.A.; Scolnick, E.M. Generation of BALB-MuSV and Ha-MuSV by type C virus transduction of homologous transforming genes from different species. Cell 1981, 26, 129–134. [Google Scholar] [CrossRef]
- Der, C.J.; Krontiris, T.G.; Cooper, G.M. Transforming genes of human bladder and lung carcinoma cell lines are homologous to the ras genes of Harvey and Kirsten sarcoma viruses. Proc. Natl. Acad. Sci. USA 1982, 79, 3637–3640. [Google Scholar] [CrossRef]
- Parada, L.F.; Tabin, C.J.; Shih, C.; Weinberg, R.A. Human EJ bladder carcinoma oncogene is homologue of Harvey sarcoma virus ras gene. Nature 1982, 297, 474–478. [Google Scholar] [CrossRef] [PubMed]
- Santos, E.; Tronick, S.R.; Aaronson, S.A.; Pulciani, S.; Barbacid, M. T24 human bladder carcinoma oncogene is an activated form of the normal human homologue of BALB- and Harvey-MSV transforming genes. Nature 1982, 298, 343–347. [Google Scholar] [CrossRef] [PubMed]
- Marshall, C.J.; Hall, A.; Weiss, R.A. A transforming gene present in human sarcoma cell lines. Nature 1982, 299, 171–173. [Google Scholar] [CrossRef]
- Shimizu, K.; Goldfarb, M.; Perucho, M.; Wigler, M. Isolation and preliminary characterization of the transforming gene of a human neuroblastoma cell line. Proc. Natl. Acad. Sci. USA 1983, 80, 383–387. [Google Scholar] [CrossRef]
- McBride, O.W.; Swan, D.C.; Santos, E.; Barbacid, M.; Tronick, S.R.; Aaronson, S.A. Localization of the normal allele of T24 human bladder carcinoma oncogene to chromosome 11. Nature 1982, 300, 773–774. [Google Scholar] [CrossRef]
- Sakaguchi, A.Y.; Naylor, S.L.; Shows, T.B.; Toole, J.J.; McCoy, M.; Weinberg, R.A. Human c-Ki-ras2 proto-oncogene on chromosome 12. Science 1983, 219, 1081–1083. [Google Scholar] [CrossRef] [PubMed]
- Hall, A.; Marshall, C.J.; Spurr, N.K.; Weiss, R.A. Identification of transforming gene in two human sarcoma cell lines as a new member of the ras gene family located on chromosome 1. Nature 1983, 303, 396–400. [Google Scholar] [CrossRef]
- Taparowsky, E.; Suard, Y.; Fasano, O.; Shimizu, K.; Goldfarb, M.; Wigler, M. Activation of the T24 bladder carcinoma transforming gene is linked to a single amino acid change. Nature 1982, 300, 762–765. [Google Scholar] [CrossRef] [PubMed]
- Reddy, E.P.; Reynolds, R.K.; Santos, E.; Barbacid, M. A point mutation is responsible for the acquisition of transforming properties by the T24 human bladder carcinoma oncogene. Nature 1982, 300, 149–152. [Google Scholar] [CrossRef]
- Tabin, C.J.; Bradley, S.M.; Bargmann, C.I.; Weinberg, R.A.; Papageorge, A.G.; Scolnick, E.M.; Dhar, R.; Lowy, D.R.; Chang, E.H. Mechanism of activation of a human oncogene. Nature 1982, 300, 143–149. [Google Scholar] [CrossRef]
- Bos, J.L. Ras oncogenes in human cancer: A review. Cancer Res. 1989, 49, 4682–4689. [Google Scholar]
- Santos, E.; Martin-Zanca, D.; Reddy, E.P.; Pierotti, M.A.; Della Porta, G.; Barbacid, M. Malignant activation of a K-ras oncogene in lung carcinoma but not in normal tissue of the same patient. Science 1984, 223, 661–664. [Google Scholar] [CrossRef] [PubMed]
- Feig, L.A.; Bast, R.C.; Knapp, R.C.; Cooper, G.M. Somatic activation of rasK gene in a human ovarian carcinoma. Science 1984, 223, 698–701. [Google Scholar] [CrossRef] [PubMed]
- Newbold, R.F.; Overell, R.W. Fibroblast immortality is a prerequisite for transformation by EJ c-Ha-ras oncogene. Nature 1983, 304, 648–651. [Google Scholar] [CrossRef] [PubMed]
- Land, H.; Parada, L.F.; Weinberg, R.A. Tumorigenic conversion of primary embryo fibroblasts requires at least two cooperating oncogenes. Nature 1983, 304, 596–602. [Google Scholar] [CrossRef] [PubMed]
- Ellis, R.W.; DeFeo, D.; Furth, M.E.; Scolnick, E.M. Mouse cells contain two distinct ras gene mRNA species that can be translated into a p21 onc protein. Mol. Cell. Biol. 1982, 2, 1339–1345. [Google Scholar] [CrossRef] [PubMed]
- McCoy, M.S.; Toole, J.J.; Cunningham, J.M.; Chang, E.H.; Lowy, D.R.; Weinberg, R.A. Characterization of a human co-lon/lung carcinoma oncogene. Nature 1983, 302, 79–81. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, J.B.; Sigal, I.S.; Poe, M.; Scolnick, E.M. Intrinsic GTPase activity distinguishes normal and oncogenic ras p21 molecules. Proc. Natl. Acad. Sci. USA 1984, 81, 5704–5708. [Google Scholar] [CrossRef] [PubMed]
- Sefton, B.M.; Trowbridge, I.S.; Cooper, J.A.; Scolnick, E.M. The transforming proteins of Rous sarcoma virus, Harvey sarcoma virus and Abelson virus contain tightly bound lipid. Cell 1982, 31, 465–474. [Google Scholar] [CrossRef]
- Willumsen, B.M.; Christensen, A.; Hubbert, N.L.; Papageorge, A.G.; Lowy, D.R. The p21 ras C-terminus is required for transformation and membrane association. Nature 1984, 310, 583–586. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.Q.; Ulsh, L.S.; Dubois, G.; Shih, T.Y. Posttranslational processing of p21 ras proteins involves palmitylation of the C-terminal tetrapeptide containing cysteine-186. J. Virol. 1985, 56. [Google Scholar] [CrossRef]
- Schafer, W.R.; Kim, R.; Sterne, R.; Thorner, J.; Kim, S.H.; Rine, J. Genetic and pharmacological suppression of oncogenic mutations in ras genes of yeast and humans. Science 1989, 245, 379–385. [Google Scholar] [CrossRef]
- Casey, P.J.; Solski, P.A.; Der, C.J.; Buss, J.E. p21ras is modified by a farnesyl isoprenoid. Proc. Natl. Acad. Sci. USA 1989, 86, 8323–8327. [Google Scholar] [CrossRef]
- Gibbs, J.B.; Kohl, N.E.; Koblan, K.S.; Omer, C.A.; Sepp-Lorenzino, L.; Rosen, N.; Anthony, N.J.; Conner, M.W.; Desolms, S.J.; Williams, T.M.; et al. Farnesyltransferase inhibitors and anti-Ras therapy. Breast Cancer Res. Treat. 1996, 38, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Goitre, L.; Trapani, E.; Trabalzini, L.; Retta, S.F. The ras superfamily of small GTPases: The unlocked secrets. Ras Signal. 2014, 1120, 1–18. [Google Scholar]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Bioinformatics 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Ellis, R.W.; Defeo, D.; Papageorge, A.; Scolnick, E.M. Expression of endogenous p21 ras genes. Prog. Clin. Boil. Res. 1983, 119, 67–77. [Google Scholar]
- Fukui, Y.; Kaziro, Y.; Yamamoto, M. Mating pheromone-like diffusible factor released by Schizosaccharomyces pombe. EMBO J. 1986, 5, 1991–1993. [Google Scholar] [CrossRef] [PubMed]
- Gallwitz, D.; Donath, C.; Sander, C. A yeast gene encoding a protein homologous to the human c-has/bas proto-oncogene product. Nature 1983, 306, 704–707. [Google Scholar] [CrossRef] [PubMed]
- Greenwald, I.; Broach, J.R. Cell fates in C. elegans: In medias ras. Cell 1990, 63, 1113–1116. [Google Scholar] [CrossRef]
- Van Dam, T.J.P.; Rehmann, H.; Bos, J.L.; Snel, B. Phylogeny of the CDC25 homology domain reveals rapid differentiation of Ras pathways between early animals and fungi. Cell. Signal. 2009, 21, 1579–1585. [Google Scholar] [CrossRef]
- Kamata, T.; Feramisco, J.R. Epidermal growth factor stimulates guanine nucleotide binding activity and phosphorylation of ras oncogene proteins. Nature 1984, 310, 147–150. [Google Scholar] [CrossRef]
- McKay, I.A.; Malone, P.; Marshall, C.J.; Hall, A. Malignant transformation of murine fibroblasts by a human c-Ha-ras-1 oncogene does not require a functional epidermal growth factor receptor. Mol. Cell. Biol. 1986, 6, 3382–3387. [Google Scholar] [CrossRef]
- Korn, L.J.; Siebel, C.W.; McCormick, F.; Roth, R.A. Ras p21 as a potential mediator of insulin action in Xenopus oocytes. Science 1987, 236, 840–843. [Google Scholar] [CrossRef] [PubMed]
- Nebreda, A.R.; Porras, A.; Santos, E. P21(ras)-induced meiotic maturation of Xenopus oocytes in the absence of protein syn-thesis: MPF activation is preceded by activation of MAP and S6 kinases. Oncogene 1993, 8, 467–477. [Google Scholar] [PubMed]
- Wakelam, M.J.O.; Davies, S.A.; Houslay, M.D.; McKay, I.; Marshall, C.J.; Hall, A. Normal p21N-ras couples bombesin and other growth factor receptors to inositol phosphate production. Nature 1986, 323, 173–176. [Google Scholar] [CrossRef] [PubMed]
- Alonso, T.; Morgan, R.O.; Marvizon, J.C.; Zarbl, H.; Santos, E. Malignant transformation by ras and other oncogenes produces common alterations in inositol phospholipid signaling pathways. Proc. Natl. Acad. Sci. USA 1988, 85, 4271–4275. [Google Scholar] [CrossRef] [PubMed]
- Broek, D.; Toda, T.; Michaeli, T.; Levin, L.; Birchmeier, C.; Zoller, M.; Powers, S.; Wigler, M. The S. cerevisiae CDC25 gene product regulates the RAS/adenylate cyclase pathway. Cell 1987, 48, 789–799. [Google Scholar] [CrossRef]
- West, M.; Kung, H.-F.; Kamata, T. A novel membrane factor stimulates guanine nucleotide exchange reaction of ras proteins. FEBS Lett. 1990, 259, 245–248. [Google Scholar] [CrossRef]
- Wolfman, A.; Macara, I.G. A cytosolic protein catalyzes the release of GDP from p21ras. Science 1990, 248, 67–69. [Google Scholar] [CrossRef] [PubMed]
- Simon, M.A.; Bowtell, D.D.L.; Dodson, G.S.; Laverty, T.R.; Rubin, G.M. Ras1 and a putative guanine nucleotide exchange factor perform crucial steps in signaling by the sevenless protein tyrosine kinase. Cell 1991, 67, 701–716. [Google Scholar] [CrossRef]
- Bowtell, D.; Fu, P.; Simon, M.; Senior, P. Identification of murine homologues of the Drosophila son of sevenless gene: Potential activators of ras. Proc. Natl. Acad. Sci. USA 1992, 89, 6511–6515. [Google Scholar] [CrossRef]
- Ebinu, J.O.; Bottorff, D.A.; Chan, E.Y.W.; Stang, S.L.; Dunn, R.J.; Stone, J.C. RasGRP, a Ras Guanyl Nucleotide- Releasing Protein with Calcium- and Diacylglycerol-Binding Motifs. Science 1998, 280, 1082–1086. [Google Scholar] [CrossRef]
- Ye, B.; Liao, D.; Zhang, X.; Zhang, P.; Dong, H.; Huganir, R.L. GRASP-1: A neuronal RasGEF associated with the AMPA receptor/GRIP complex. Neuron 2000, 26, 603–617. [Google Scholar] [CrossRef]
- Huang, J.; Furuya, A.; Furuichi, T. Very-KIND, a KIND domain–containing RasGEF, controls dendrite growth by linking Ras small GTPases and MAP2. J. Cell Biol. 2007, 179, 539–552. [Google Scholar] [CrossRef] [PubMed]
- Osei-Sarfo, K.; Martello, L.; Ibrahim, S.; Pellicer, A. The human Rgr oncogene is overexpressed in T-cell malignancies and induces transformation by acting as a GEF for Ras and Ral. Oncogene 2011, 30, 3661–3671. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Henkels, K.M.; Mahankali, M.; Gomez-Cambronero, J. Increased cell growth due to a new lipase-GEF (Phospholipase D2) fastly acting on Ras. Cell. Signal. 2013, 25, 198–205. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Baltanás, F.C.; Zarich, N.; Rojas-Cabañeros, J.M.; Santos, E. SOS GEFs in health and disease. Biochim. Biophys. Acta-Rev. Cancer 2020, 1874, 188445. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Medarde, A.; Santos, E. The RasGrf family of mammalian guanine nucleotide exchange factors. Biochim. Biophys. Acta-Rev. Cancer 2011, 1815, 170–188. [Google Scholar] [CrossRef]
- Stone, J.C. Regulation and Function of the RasGRP Family of Ras Activators in Blood Cells. Genes Cancer 2011, 2, 320–334. [Google Scholar] [CrossRef]
- Trahey, M.; McCormick, F. A cytoplasmic protein stimulates normal N-ras p21 GTPase, but does not affect oncogenic mutants. Science 1987, 238, 542–545. [Google Scholar] [CrossRef]
- Calés, C.; Hancock, J.F.; Marshall, C.J.; Hall, A. The cytoplasmic protein GAP is implicated as the target for regulation by the ras gene product. Nature 1988, 332, 548–551. [Google Scholar] [CrossRef]
- Gibbs, J.B.; Schaber, M.D.; Allard, W.J.; Sigal, I.S.; Scolnick, E.M. Purification of ras GTPase activating protein from bovine brain. Proc. Natl. Acad. Sci. USA 1988, 85, 5026–5030. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Lin, B.; Tanaka, K.; Dunn, D.; Wood, D.; Gesteland, R.; White, R.; Weiss, R.; Tamanoi, F. The catalytic domain of the neurofibromatosis type 1 gene product stimulates ras GTPase and complements ira mutants of S. cerevisiae. Cell 1990, 63, 835–841. [Google Scholar] [CrossRef]
- Scheffzek, K.; Shivalingaiah, G. Ras-Specific GTPase-Activating Proteins—Structures, Mechanisms, and Interactions. Cold Spring Harb. Perspect. Med. 2019, 9, a031500. [Google Scholar] [CrossRef] [PubMed]
- Lowenstein, E.J.; Daly, R.J.; Batzer, A.G.; Li, W.; Margolis, B.; Lammers, R.; Ullrich, A.; Skolnik, E.Y.; Bar-Sagi, D.; Schlessinger, J. The SH2 and SH3 domain-containing protein GRB2 links receptor tyrosine kinases to ras signaling. Cell 1992, 70, 431–442. [Google Scholar] [CrossRef]
- Gale, N.W.; Kaplan, S.; Lowenstein, E.J.; Schlessinger, J.; Bar-Sagi, D. Grb2 mediates the EGF-dependent activation of guanine nucleotide exchange on Ras. Nature 1993, 363, 88–92. [Google Scholar] [CrossRef]
- Rozakis-Adcock, M.; Fernley, R.T.; Wade, J.D.; Pawson, T.; Bowtell, D.D.L. The SH2 and SH3 domains of mammalian Grb2 couple the EGF receptor to the Ras activator mSos1. Nature 1993, 363, 83–85. [Google Scholar] [CrossRef]
- Chardin, P.; Camonis, J.H.; Gale, N.W.; Van Aelst, L.; Schlessinger, J.; Wigler, M.H.; Bar-Sagi, D. Human Sos1: A guanine nucleotide exchange factor for Ras that binds to GRB2. Science 1993, 260, 1338–1343. [Google Scholar] [CrossRef]
- McCormick, F. Signal transduction. How receptors turn Ras on. Nature 1993, 363, 15–16. [Google Scholar] [CrossRef]
- Sigal, I.S.; Gibbs, J.B.; D’Alonzo, J.S.; Scolnick, E.M. Identification of effector residues and a neutralizing epitope of Ha-ras-encoded p21. Proc. Natl. Acad. Sci. USA 1986, 83, 4725–4729. [Google Scholar] [CrossRef]
- Adari, H.; Lowy, D.R.; Willumsen, B.M.; Der, C.J.; McCormick, F. Guanosine triphosphatase activating protein (GAP) interacts with the p21 ras effector binding domain. Science 1988, 240, 518–521. [Google Scholar] [CrossRef]
- Willumsen, B.M.; Papageorge, A.G.; Kung, H.F.; Bekesi, E.; Robins, T.; Johnsen, M.; Vass, W.C.; Lowy, D.R. Mutational analysis of a ras catalytic domain. Mol. Cell. Biol. 1986, 6, 2646–2654. [Google Scholar] [CrossRef]
- De Vos, A.M.; Tong, L.; Milburn, M.V.; Matias, P.M.; Jancarik, J.; Noguchi, S.; Nishimura, S.; Miura, K.; Ohtsuka, E.; Kim, S.H. Three-dimensional structure of an oncogene protein: Catalytic domain of human c-H-ras p21. Science 1988, 239, 888–893. [Google Scholar] [CrossRef] [PubMed]
- Brünger, A.T.; Milburn, M.V.; Tong, L.; DeVos, A.M.; Jancarik, J.; Yamaizumi, Z.; Nishimura, S.; Ohtsuka, E.; Kim, S.H. Crystal structure of an active form of RAS protein, a complex of a GTP analog and the HRAS p21 catalytic domain. Proc. Natl. Acad. Sci. USA 1990, 87, 4849–4853. [Google Scholar] [CrossRef] [PubMed]
- Mistou, M.Y.; Jacquet, E.; Poullet, P.; Rensland, H.; Gideon, P.; Schlichting, I.; Wittinghofer, A.; Parmeggiani, A. Mutations of Ha-ras p21 that define important regions for the molecular mechanism of the SDC25 C-domain, a guanine nucleotide dissociation stimulator. EMBO J. 1992, 11, 2391–2397. [Google Scholar] [CrossRef] [PubMed]
- Verrotti, A.C.; Crechet, J.B.; Di Blasi, F.; Seidita, G.; Mirisola, M.G.; Kavounis, C.; Nastopoulos, V.; Burderi, E.; De Vendittis, E.; Parmeggiani, A.; et al. RAS residues that are distant from the GDP binding site play a critical role in dissociation factor-stimulated release of GDP. EMBO J. 1992, 11, 2855–2862. [Google Scholar] [CrossRef] [PubMed]
- Segal, M.; Willumsen, B.M.; Levitzki, A. Residues crucial for Ras interaction with GDP-GTP exchangers. Proc. Natl. Acad. Sci. USA 1993, 90, 5564–5568. [Google Scholar] [CrossRef] [PubMed]
- Mosteller, R.D.; Han, J.; Broek, D. Identification of residues of the H-ras protein critical for functional interaction with guanine nucleotide exchange factors. Mol. Cell. Biol. 1994, 14, 1104–1112. [Google Scholar] [CrossRef] [PubMed]
- Quilliam, L.A.; Kato, K.; Rabun, K.M.; Hisaka, M.M.; Huff, S.Y.; Campbell-Burk, S.; Der, C.J. Identification of residues critical for Ras(17N) growth-inhibitory phenotype and for Ras interaction with guanine nucleotide exchange factors. Mol. Cell. Biol. 1994, 14, 1113–1121. [Google Scholar] [CrossRef]
- Leonardsen, L.; DeClue, J.E.; Lybæk, H.; Lowy, D.R.; Willumsen, B.M. Rasp21 sequences opposite the nucleotide binding pocket are required for GRF-mediated nucleotide release. Oncogene 1996, 13, 2177–2187. [Google Scholar]
- Boriack-Sjodin, P.A.; Margarit, S.M.; Bar-Sagi, D.; Kuriyan, J. The structural basis of the activation of Ras by Sos. Nature 1998, 394, 337–343. [Google Scholar] [CrossRef]
- McCormick, F. Working hypothesis. GAP as ras effector or negative regulator? Mol. Carcinog. 1990, 3, 185–187. [Google Scholar] [CrossRef]
- Pamonsinlapatham, P.; Hadj-Slimane, R.; Lepelletier, Y.; Allain, B.; Toccafondi, M.; Garbay, C.; Raynaud, F. P120-Ras GTPase activating protein (RasGAP): A multi-interacting protein in downstream signaling. Biochimie 2009, 91, 320–328. [Google Scholar] [CrossRef] [PubMed]
- Moodie, S.; Willumsen, B.; Weber, M.; Wolfman, A. Complexes of Ras.GTP with Raf-1 and mitogen-activated protein kinase kinase. Science 1993, 260, 1658–1661. [Google Scholar] [CrossRef]
- Rodriguez-Viciana, P.; Warne, P.H.; Dhand, R.; Vanhaesebroeck, B.; Gout, I.; Fry, M.J.; Waterfield, M.D.; Downward, J. Phosphatidylinositol-3-OH kinase direct target of Ras. Nature 1994, 370, 527–532. [Google Scholar] [CrossRef] [PubMed]
- Hofer, F.; Fields, S.; Schneider, C.; Martin, G.S. Activated Ras interacts with the Ral guanine nucleotide dissociation stimulator. Proc. Natl. Acad. Sci. USA 1994, 91, 11089–11093. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Meco, M.T.; Lozano, J.; Municio, M.M.; Berra, E.; Frutos, S.; Sanz, L.; Moscat, J. Evidence for the in vitro and in vivo interaction of Ras with protein kinase C ζ. J. Biol. Chem. 1994, 269, 31706–31710. [Google Scholar] [CrossRef]
- Joneson, T.; Bar-Sagi, D. Ras effectors and their role in mitogenesis and oncogenesis. J. Mol. Med. 1997, 75, 587–593. [Google Scholar] [CrossRef]
- Kyriakis, J.M.; App, H.; Zhang, X.-F.; Banerjee, P.; Brautigan, D.L.; Rapp, U.R.; Avruch, J. Raf-1 activates MAP kinase-kinase. Nature 1992, 358, 417–421. [Google Scholar] [CrossRef]
- Gómez, N.; Cohen, P. Dissection of the protein kinase cascade by which nerve growth factor activates MAP kinases. Nature 1991, 353, 170–173. [Google Scholar] [CrossRef]
- Brunet, A.; Pagès, G.; Pouysségur, J. Constitutively active mutants of MAP kinase kinase (MEK1) induce growth fac-tor-relaxation and oncogenicity when expressed in fibroblasts. Oncogene 1994, 9, 3379–3387. [Google Scholar]
- Pages, G.; Lenormand, P.; L’Allemain, G.; Chambard, J.C.; Meloche, S.; Pouyssegur, J. Mitogen-activated protein kinases p42mapk and p44mapk are required for fibroblast proliferation. Proc. Natl. Acad. Sci. USA 1993, 90, 8319–8323. [Google Scholar] [CrossRef]
- Burgering, B.M.T.; Coffer, P.J. Protein kinase B (c-Akt) in phosphatidylinositol-3-OH kinase signal transduction. Nature 1995, 376, 599–602. [Google Scholar] [CrossRef] [PubMed]
- Myers, M.G.; Grammer, T.C.; Wang, L.M.; Sun, X.J.; Pierce, J.H.; Blenis, J.; White, M.F. Insulin receptor substrate-1 mediates phosphatidylinositol 3’-kinase and p70S6k signaling during insulin, insulin-like growth factor-1, and interleukin-4 stimulation. J. Biol. Chem. 1994, 269, 28783–28789. [Google Scholar] [CrossRef]
- Tsakiridis, T.; Mc dowell, H.E.; Walker, T.; Downes, C.P.; Hundals, H.S.; Vranic, M.; Klip, A. Multiple roles of phosphatidylinositol 3-kinase in regulation of glucose transport, amino acid transport, and glucose transporters in L6 skeletal muscle cells. Endocrinology 1995, 136, 4315–4322. [Google Scholar] [CrossRef]
- Mèndez, R.; Myers, M.G.; White, M.F.; Rhoads, R.E. Stimulation of protein synthesis, eukaryotic translation initiation factor 4E phosphorylation, and PHAS-I phosphorylation by insulin requires insulin receptor substrate 1 and phosphatidylinositol 3-kinase. Mol. Cell. Biol. 1996, 16, 2857–2864. [Google Scholar] [CrossRef] [PubMed]
- Cross, D.A.E.; Alessi, D.R.; Vandenheede, J.R.; McDowell, H.E.; Hundal, H.S.; Cohen, P. The inhibition of glycogen synthase kinase-3 by insulin or insulin-like growth factor 1 in the rat skeletal muscle cell line L6 is blocked by wortmannin, but not by rapamycin: Evidence that wortmannin blocks activation of the mitogen-activated protein kinase pathway in L6 cells between Ras and Raf. Biochem. J. 1994, 303, 21–26. [Google Scholar] [CrossRef]
- Downward, J. Targeting RAS signalling pathways in cancer therapy. Nat. Rev. Cancer 2003, 3, 11–22. [Google Scholar] [CrossRef]
- Ballester, R.; Furth, M.E.; Rosen, O.M. Phorbol ester- and protein kinase C-mediated phosphorylation of the cellular Kirsten ras gene product. J. Biol. Chem. 1987, 262, 2688–2695. [Google Scholar] [CrossRef]
- Bivona, T.G.; Quatela, S.E.; Bodemann, B.O.; Ahearn, I.M.; Soskis, M.J.; Mor, A.; Miura, J.; Wiener, H.H.; Wright, L.; Saba, S.G.; et al. PKC Regulates a Farnesyl-Electrostatic Switch on K-Ras that Promotes its Association with Bcl-Xl on Mitochondria and Induces Apoptosis. Mol. Cell 2006, 21, 481–493. [Google Scholar] [CrossRef]
- Barceló, C.; Paco, N.; Morell, M.; Alvarez-Moya, B.; Bota-Rabassedas, N.; Jaumot, M.; Vilardell, F.; Capella, G.; Agell, N. Phosphorylation at Ser-181 of Oncogenic KRAS Is Required for Tumor Growth. Cancer Res. 2014, 74, 1190–1199. [Google Scholar] [CrossRef]
- Jura, N.; Scotto-Lavino, E.; Sobczyk, A.; Bar-Sagi, D. Differential Modification of Ras Proteins by Ubiquitination. Mol. Cell 2006, 21, 679–687. [Google Scholar] [CrossRef]
- Baker, R.; Lewis, S.M.; Sasaki, A.T.; Wilkerson, E.M.; Locasale, J.W.; Cantley, L.C.; Kuhlman, B.; Dohlman, H.G.; Campbell, S.L. Site-specific monoubiquitination activates Ras by impeding GTPase-activating protein function. Nat. Struct. Mol. Biol. 2012, 20, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Agudo-Ibáñez, L.; Herrero, A.; Barbacid, M.; Crespo, P. H-Ras Distribution and Signaling in Plasma Membrane Microdomains Are Regulated by Acylation and Deacylation Events. Mol. Cell. Biol. 2015, 35, 1898–1914. [Google Scholar] [CrossRef]
- Jeong, W.J.; Yoon, J.; Park, J.C.; Lee, S.H.; Lee, S.H.; Kaduwal, S.; Kim, H.; Yoon, J.B.; Choi, K.Y. Ras Stabilization Through Aberrant Activation of Wnt/ -Catenin Signaling Promotes Intestinal Tumorigenesis. Sci. Signal. 2012, 5, 30. [Google Scholar] [CrossRef] [PubMed]
- Clavreul, N.; Adachi, T.; Pimental, D.R.; Ido, Y.; Schöneich, C.; Cohen, R.A. S-glutathiolation by peroxynitrite of p21ras at cysteine-118 mediates its direct activation and downstream signaling in endothelial cells. FASEB J. 2006, 20, 518–520. [Google Scholar] [CrossRef] [PubMed]
- Batista, W.L.; Ogata, F.T.; Curcio, M.F.; Miguel, R.B.; Arai, R.J.; Matsuo, A.L.; Moraes, M.S.; Stern, A.; Monteiro, H.P. S-Nitrosoglutathione and Endothelial Nitric Oxide Synthase-Derived Nitric Oxide Regulate Compartmentalized RasS-Nitrosylation and Stimulate Cell Proliferation. Antioxidants Redox Signal. 2013, 18, 221–238. [Google Scholar] [CrossRef]
- Santos, A.I.; Carreira, B.P.; Izquierdo-Álvarez, A.; Ramos, E.; Lourenço, A.S.; Santos, D.F.; Morte, M.I.; Ribeiro, L.F.; Marreiros, A.; Sánchez-López, N.; et al. S-Nitrosylation of Ras Mediates Nitric Oxide-Dependent Post-Injury Neurogenesis in a Seizure Model. Antioxidants Redox Signal. 2018, 28, 15–30. [Google Scholar] [CrossRef]
- Yang, M.H.; Nickerson, S.; Kim, E.T.; Liot, C.; Laurent, G.; Spang, R.; Philips, M.R.; Shan, Y.; Shaw, D.E.; Bar-Sagi, D.; et al. Regulation of RAS oncogenicity by acetylation. Proc. Natl. Acad. Sci. USA 2012, 109, 10843–10848. [Google Scholar] [CrossRef]
- Santos, E.; Nebreda, A.R.; Bryan, T.; Kempner, E.S. Oligomeric structure of p21 ras proteins as determined by radiation inactivation. J. Biol. Chem. 1988, 263, 9853–9858. [Google Scholar] [CrossRef]
- Prior, I.A.; Muncke, C.; Parton, R.G.; Hancock, J.F. Direct visualization of Ras proteins in spatially distinct cell surface microdomains. J. Cell Biol. 2003, 160, 165–170. [Google Scholar] [CrossRef]
- Nicolini, C.; Baranski, J.; Schlummer, S.; Palomo, J.; Lumbierres-Burgues, M.; Kahms, M.; Kuhlmann, J.; Sánchez, S.; Gratton, E.; Waldmann, H.; et al. Visualizing Association of N-Ras in Lipid Microdomains: Influence of Domain Structure and Interfacial Adsorption. J. Am. Chem. Soc. 2006, 128, 192–201. [Google Scholar] [CrossRef]
- Shalom-Feuerstein, R.; Plowman, S.J.; Rotblat, B.; Ariotti, N.; Tian, T.; Hancock, J.F.; Kloog, Y. K-Ras Nanoclustering Is Subverted by Overexpression of the Scaffold Protein Galectin-3. Cancer Res. 2008, 68, 6608–6616. [Google Scholar] [CrossRef]
- Michael, J.V.; Wurtzel, J.G.T.; Goldfinger, L.E. Regulation of H-Ras-driven MAPK signaling, transformation and tumorigenesis, but not PI3K signaling and tumor progression, by plasma membrane microdomains. Oncogenesis 2016, 5, e228. [Google Scholar] [CrossRef] [PubMed]
- Santos, E. Dimerization Opens New Avenues into Ras Signaling Research. Sci. Signal. 2014, 7, 12. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nan, X.; Tamgüney, T.M.; Collisson, E.A.; Lin, L.-J.; Pitt, C.; Galeas, J.; Lewis, S.; Gray, J.W.; McCormick, F.; Chu, S. Ras-GTP dimers activate the Mitogen-Activated Protein Kinase (MAPK) pathway. Proc. Natl. Acad. Sci. USA 2015, 112, 7996–8001. [Google Scholar] [CrossRef]
- Santos, E.; Crespo, P. The RAS-ERK pathway: A route for couples. Sci. Signal. 2018, 11, eaav0917. [Google Scholar] [CrossRef] [PubMed]
- Muratcioglu, S.; Chavan, T.S.; Freed, B.C.; Jang, H.; Khavrutskii, L.; Freed, R.N.; Dyba, M.A.; Stefanisko, K.; Tarasov, S.G.; Gursoy, A.; et al. GTP-Dependent K-Ras Dimerization. Structure 2015, 23, 1325–1335. [Google Scholar] [CrossRef]
- Ambrogio, C.; Köhler, J.; Zhou, Z.-W.; Wang, H.; Paranal, R.; Li, J.; Capelletti, M.; Caffarra, C.; Li, S.; Lv, Q.; et al. KRAS Dimerization Impacts MEK Inhibitor Sensitivity and Oncogenic Activity of Mutant KRAS. Cell 2018, 172, 857–868.e15. [Google Scholar] [CrossRef]
- Margarit, S.M.; Sondermann, H.; Hall, B.E.; Nagar, B.; Hoelz, A.; Pirruccello, M.; Bar-Sagi, D.; Kuriyan, J. Structural Evidence for Feedback Activation by Ras·GTP of the Ras-Specific Nucleotide Exchange Factor SOS. Cell 2003, 112, 685–695. [Google Scholar] [CrossRef]
- Jeng, H.-H.; Taylor, L.J.; Bar-Sagi, D. Sos-mediated cross-activation of wild-type Ras by oncogenic Ras is essential for tumorigenesis. Nat. Commun. 2012, 3, 1–8. [Google Scholar] [CrossRef]
- Bentley, C.; Jurinka, S.S.; Kljavin, N.M.; Vartanian, S.; Ramani, S.R.; Gonzalez, L.C.; Yu, K.; Modrusan, Z.; Du, P.; Bourgon, R.; et al. A requirement for wild-type Ras isoforms in mutant KRas-driven signalling and transformation. Biochem. J. 2013, 452, 313–320. [Google Scholar] [CrossRef]
- Chiu, V.K.; Bivona, T.; Hach, A.; Sajous, J.B.; Silletti, J.; Wiener, H.; Ii, R.L.J.; Cox, A.D.; Philips, M.R. Ras signalling on the endoplasmic reticulum and the Golgi. Nature 2002, 4, 343–350. [Google Scholar] [CrossRef]
- Matallanas, D.; Sanz-Moreno, V.; Arozarena, I.; Calvo, F.; Agudo-Ibáñez, L.; Santos, E.; Berciano, M.T.; Crespo, P. Distinct Utilization of Effectors and Biological Outcomes Resulting from Site-Specific Ras Activation: Ras Functions in Lipid Rafts and Golgi Complex Are Dispensable for Proliferation and Transformation. Mol. Cell. Biol. 2006, 26, 100–116. [Google Scholar] [CrossRef]
- Calvo, F.; Agudo-Ibáñez, L.; Crespo, P. The Ras-ERK pathway: Understanding site-specific signaling provides hope of new anti-tumor therapies. BioEssays 2010, 32, 412–421. [Google Scholar] [CrossRef]
- Johnson, S.M.; Grosshans, H.; Shingara, J.; Byrom, M.; Jarvis, R.; Cheng, A.; Labourier, E.; Reinert, K.L.; Brown, D.; Slack, F.J. RAS Is Regulated by the let-7 MicroRNA Family. Cell 2005, 120, 635–647. [Google Scholar] [CrossRef] [PubMed]
- Karmakar, S.; Kaushik, G.; Nimmakayala, R.; Rachagani, S.; Ponnusamy, M.P.; Batra, S.K. MicroRNA regulation of K-Ras in pancreatic cancer and opportunities for therapeutic intervention. Semin. Cancer Biol. 2019, 54, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Mulcahy, L.S.; Smith, M.R.; Stacey, D.W. Requirement for ras proto-oncogene function during serum-stimulated growth of NIH 3T3 cells. Nature 1985, 313, 241–243. [Google Scholar] [CrossRef]
- Yuspa, S.H.; Vass, W.; Scolnick, E. Altered growth and differentiation of cultured mouse epidermal cells infected with onco-genic retrovirus: Contrasting effects of viruses and chemicals. Cancer Res. 1983, 43, 6021–6030. [Google Scholar]
- Bar-Sagi, D.; Feramisco, J.R. Microinjection of the ras oncogene protein into PC12 cells induces morphological differentiation. Cell 1985, 42, 841–848. [Google Scholar] [CrossRef]
- Benito, M.; Porras, A.; Nebreda, A.R.; Santos, E. Differentiation of 3T3-L1 fibroblasts to adipocytes induced by transfection of ras oncogenes. Science 1991, 253, 565–568. [Google Scholar] [CrossRef]
- Liotta, L.A.; Mandler, R.; Murano, G.; Katz, D.A.; Gordon, R.K.; Chiang, P.K.; Schiffmann, E. Tumor cell autocrine motility factor. Proc. Natl. Acad. Sci. USA 1986, 83, 3302–3306. [Google Scholar] [CrossRef]
- Clark, G.J.; Westwick, J.K.; Der, C.J. p120 GAP Modulates Ras Activation of Jun Kinases and Transformation. J. Biol. Chem. 1997, 272, 1677–1681. [Google Scholar] [CrossRef]
- Yamauchi, J.; Miyamoto, Y.; Tanoue, A.; Shooter, E.M.; Chan, J.R. Ras activation of a Rac1 exchange factor, Tiam1, mediates neurotrophin-3-induced Schwann cell migration. Proc. Natl. Acad. Sci. USA 2005, 102, 14889–14894. [Google Scholar] [CrossRef] [PubMed]
- Gallego, C.; Gupta, S.K.; Heasley, L.E.; Qian, N.X.; Johnson, G.L. Mitogen-activated protein kinase activation resulting from selective oncogene expression in NIH 3T3 and rat 1a cells. Proc. Natl. Acad. Sci. USA 1992, 89, 7355–7359. [Google Scholar] [CrossRef] [PubMed]
- Rapp, U.R.; Goldsborough, M.D.; Mark, G.E.; Bonner, T.I.; Groffen, J.; Reynolds, F.H.; Stephenson, J.R. Structure and biological activity of v-raf, a unique oncogene transduced by a retrovirus. Proc. Natl. Acad. Sci. USA 1983, 80, 4218–4222. [Google Scholar] [CrossRef] [PubMed]
- Davies, H.; Bignell, G.R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Teague, J.; Woffendin, H.; Garnett, M.J.; Bottomley, W.; et al. Mutations of the BRAF gene in human cancer. Nature 2002, 417, 949–954. [Google Scholar] [CrossRef]
- Neuzillet, C.; Tijeras-Raballand, A.; De Mestier, L.; Cros, J.; Faivre, S.; Raymond, E. MEK in cancer and cancer therapy. Pharmacol. Ther. 2014, 141, 160–171. [Google Scholar] [CrossRef]
- Torii, S.; Yamamoto, T.; Tsuchiya, Y.; Nishida, E. ERK MAP kinase in G1 cell cycle progression and cancer. Cancer Sci. 2006, 97, 697–702. [Google Scholar] [CrossRef]
- Samuels, Y.; Wang, Z.; Bardelli, A.; Silliman, N.; Ptak, J.; Szabo, S.; Yan, H.; Gazdar, A.; Powell, S.M.; Riggins, G.J.; et al. High Frequency of Mutations of the PIK3CA Gene in Human Cancers. Science 2004, 304, 554. [Google Scholar] [CrossRef]
- Li, J.; Yen, C.; Liaw, D.; Podsypanina, K.; Bose, S.; Wang, S.I.; Puc, J.; Miliaresis, C.; Rodgers, L.; McCombie, R.; et al. PTEN, a Putative Protein Tyrosine Phosphatase Gene Mutated in Human Brain, Breast, and Prostate Cancer. Science 1997, 275, 1943–1947. [Google Scholar] [CrossRef]
- Maehama, T.; Dixon, J.E. The Tumor Suppressor, PTEN/MMAC1, Dephosphorylates the Lipid Second Messenger, Phosphatidylinositol 3,4,5-Trisphosphate. J. Biol. Chem. 1998, 273, 13375–13378. [Google Scholar] [CrossRef]
- Garnett, M.J.; Marais, R. Guilty as Charged: B-RAF is a Human Oncogene. Cancer Cell 2004, 6, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Cisowski, J.; Sayin, V.I.; Liu, M.; Karlsson, C.; Bergo, M.O. Oncogene-induced senescence underlies the mutual exclusive nature of oncogenic KRAS and BRAF. Oncogene 2016, 35, 1328–1333. [Google Scholar] [CrossRef] [PubMed]
- Cisowski, J.; Bergo, M.O. What makes oncogenes mutually exclusive? Small GTPases 2016, 8, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Janku, F.; Lee, J.J.; Tsimberidou, A.M.; Hong, D.S.; Naing, A.; Falchook, G.S.; Fu, S.; Luthra, R.; Garrido-Laguna, I.; Kurzrock, R. PIK3CA Mutations Frequently Coexist with RAS and BRAF Mutations in Patients with Advanced Cancers. PLoS ONE 2011, 6, e22769. [Google Scholar] [CrossRef]
- Asati, V.; Mahapatra, D.K.; Bharti, S.K. PI3K/Akt/mTOR and Ras/Raf/MEK/ERK signaling pathways inhibitors as anticancer agents: Structural and pharmacological perspectives. Eur. J. Med. Chem. 2016, 109, 314–341. [Google Scholar] [CrossRef]
- Aoki, Y.; Niihori, T.; Kawame, H.; Kurosawa, K.; Ohashi, H.; Tanaka, Y.; Filocamo, M.; Kato, K.; Suzuki, Y.; Kure, S.; et al. Germline mutations in HRAS proto-oncogene cause Costello syndrome. Nat. Genet. 2005, 37, 1038–1040. [Google Scholar] [CrossRef]
- Tidyman, W.E.; Rauen, K.A. The RASopathies: Developmental syndromes of Ras/MAPK pathway dysregulation. Curr. Opin. Genet. Dev. 2009, 19, 230–236. [Google Scholar] [CrossRef]
- Roberts, A.E.; Araki, T.; Swanson, K.D.; Montgomery, K.T.; Schiripo, T.A.; Joshi, V.A.; Li, L.; Yassin, Y.; Tamburino, A.M.; Neel, B.G.; et al. Germline gain-of-function mutations in SOS1 cause Noonan syndrome. Nat. Genet. 2007, 39, 70–74. [Google Scholar] [CrossRef]
- Aoki, Y.; Niihori, T.; Inoue, S.-I.; Matsubara, Y. Recent advances in RASopathies. J. Hum. Genet. 2016, 61, 33–39. [Google Scholar] [CrossRef]
- Zhou, H.; Yao, C.; Bian, A.; Qian, J.; Zhao, X.; Zhao, Y.; Wang, W.; Xing, C. The Ras GTPase-activating-like protein IQGAP1 is downregulated in human diabetic nephropathy and associated with ERK1/2 pathway activation. Mol. Cell. Biochem. 2014, 391, 21–25. [Google Scholar] [CrossRef]
- Gamache, T.R.; Araki, Y.; Huganir, R.L. Twenty Years of SynGAP Research: From Synapses to Cognition. J. Neurosci. 2020, 40, 1596–1605. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Shi, Z.; Zhang, Y.; Yan, J.; Yu, W.; Chen, L. Oligomer β-amyloid Induces Hyperactivation of Ras to Impede NMDA Receptor-Dependent Long-Term Potentiation in Hippocampal CA1 of Mice. Front. Pharmacol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Katsuki, M.; Kimura, M.; Yokoyama, M.; Sato, M.; Kimura, S.; Hata, J.; Sekiya, T.; Izawa, M.; Nishimura, S. Teratocarcinoma derived from mouse zygotes after the introduction of activated human c-Ha-ras DNA. Princess Takamatsu Symp. 1986, 17, 269–276. [Google Scholar] [PubMed]
- Mitsumori, K. Possible mechanism on enhanced carcinogenesis of genotoxic carcinogens and unsolved mechanisms on lesser carcinogenic susceptibility to some carcinogens in rash2 mice. J. Toxicol. Sci. 2003, 28, 371–383. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Storer, R.D.; Sistare, F.D.; Reddy, M.V.; DeGeorge, J.J. An Industry Perspective on the Utility of Short-Term Carcinogenicity Testing in Transgenic Mice in Pharmaceutical Development. Toxicol. Pathol. 2009, 38, 51–61. [Google Scholar] [CrossRef] [PubMed]
- Chin, L.; Tam, A.; Pomerantz, J.; Wong, M.; Holash, J.; Bardeesy, N.; Shen, Q.; O’Hagan, R.; Pantginis, J.; Zhou, H.; et al. Essential role for oncogenic Ras in tumour maintenance. Nature 1999, 400, 468–472. [Google Scholar] [CrossRef]
- Fisher, G.H.; Wellen, S.L.; Klimstra, D.; Lenczowski, J.M.; Tichelaar, J.W.; Lizak, M.J.; Whitsett, J.A.; Koretsky, A.; Varmus, H.E. Induction and apoptotic regression of lung adenocarcinomas by regulation of a K-Ras transgene in the presence and absence of tumor suppressor genes. Genes Dev. 2001, 15, 3249–3262. [Google Scholar] [CrossRef]
- Hingorani, S.R.; Wang, L.; Multani, A.S.; Combs, C.; Deramaudt, T.B.; Hruban, R.H.; Rustgi, A.K.; Chang, S.; Tuveson, D.A. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 2005, 7, 469–483. [Google Scholar] [CrossRef]
- Drosten, M.; Simón-Carrasco, L.; Hernández-Porras, I.; Lechuga, C.G.; Blasco, M.T.; Jacob, H.K.; Fabbiano, S.; Potenza, N.; Bustelo, X.R.; Guerra, C.; et al. H-Ras and K-Ras Oncoproteins Induce Different Tumor Spectra When Driven by the Same Regulatory Sequences. Cancer Res. 2016, 77, 707–718. [Google Scholar] [CrossRef]
- Johnson, L.; Greenbaum, D.; Cichowski, K.; Mercer, K.; Murphy, E.; Schmitt, E.; Bronson, R.T.; Umanoff, H.; Edelmann, W.; Kucherlapati, R.; et al. K-ras is an essential gene in the mouse with partial functional overlap with N-ras. Genes Dev. 1997, 11, 2468–2481. [Google Scholar] [CrossRef]
- Esteban, L.M.; Vicario-Abejón, C.; Fernández-Salguero, P.; Fernández-Medarde, A.; Swaminathan, N.; Yienger, K.; Lopez, E.; Malumbres, M.; McKay, R.; Ward, J.M.; et al. Targeted Genomic Disruption of H-ras and N-ras, Individually or in Combination, Reveals the Dispensability of Both Loci for Mouse Growth and Development. Mol. Cell. Biol. 2001, 21, 1444–1452. [Google Scholar] [CrossRef] [PubMed]
- Fuentes-Mateos, R.; Jimeno, D.; Gómez, C.; Calzada, N.; Fernández-Medarde, A.; Santos, E. Concomitant deletion of HRAS and NRAS leads to pulmonary immaturity, respiratory failure and neonatal death in mice. Cell Death Dis. 2019, 10, 1–16. [Google Scholar] [CrossRef]
- Potenza, N.; Vecchione, C.; Notte, A.; De Rienzo, A.; Rosica, A.; Bauer, L.; Affuso, A.; De Felice, M.; Russo, T.; Poulet, R.; et al. Replacement of K-Ras with H-Ras supports normal embryonic development despite inducing cardiovascular pathology in adult mice. EMBO Rep. 2005, 6, 432–437. [Google Scholar] [CrossRef]
- Nakamura, K.; Ichise, H.; Nakao, K.; Hatta, T.; Otani, H.; Sakagami, H.; Kondo, H.; Katsuki, M. Partial functional overlap of the three ras genes in mouse embryonic development. Oncogene 2007, 27, 2961–2968. [Google Scholar] [CrossRef][Green Version]
- Castellano, E.; Santos, E. Functional Specificity of Ras Isoforms: So Similar but So Different. Genes Cancer 2011, 2, 216–231. [Google Scholar] [CrossRef] [PubMed]
- Jackson, E.L.; Willis, N.; Mercer, K.; Bronson, R.T.; Crowley, D.; Montoya, R.; Jacks, T.; Tuveson, D.A. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 2001, 15, 3243–3248. [Google Scholar] [CrossRef] [PubMed]
- Guerra, C.; Mijimolle, N.; Dhawahir, A.; Dubus, P.; Barradas, M.; Serrano, M.; Campuzano, V.; Barbacid, M. Tumor induction by an endogenous K-ras oncogene is highly dependent on cellular context. Cancer Cell 2003, 4, 111–120. [Google Scholar] [CrossRef]
- Johnson, L.; Mercer, K.; Greenbaum, D.; Bronson, R.T.; Crowley, D.; Tuveson, D.A.; Jacks, T. Somatic activation of the K-ras oncogene causes early onset lung cancer in mice. Nature 2001, 410, 1111–1116. [Google Scholar] [CrossRef] [PubMed]
- Hingorani, S.R.; Petricoin, E.F.; Maitra, A.; Rajapakse, V.; King, C.; Jacobetz, M.A.; Ross, S.; Conrads, T.P.; Veenstra, T.D.; Hitt, B.A.; et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 2003, 4, 437–450. [Google Scholar] [CrossRef]
- Guerra, C.; Schuhmacher, A.J.; Cañamero, M.; Grippo, P.J.; Verdaguer, L.; Pérez-Gallego, L.; Dubus, P.; Sandgren, E.P.; Barbacid, M. Chronic Pancreatitis Is Essential for Induction of Pancreatic Ductal Adenocarcinoma by K-Ras Oncogenes in Adult Mice. Cancer Cell 2007, 11, 291–302. [Google Scholar] [CrossRef]
- Guerra, C.; Collado, M.; Navas, C.; Schuhmacher, A.J.; Hernández-Porras, I.; Cañamero, M.; Rodriguez-Justo, M.; Serrano, M.; Barbacid, M. Pancreatitis-Induced Inflammation Contributes to Pancreatic Cancer by Inhibiting Oncogene-Induced Senescence. Cancer Cell 2011, 19, 728–739. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, J.; Frutschi, M.; Kaloulis, K.; McKee, T.; Trumpp, A.; Beermann, F. Metastasizing Melanoma Formation Caused by Expression of Activated N-RasQ61K on an INK4a-Deficient Background. Cancer Res. 2005, 65, 4005–4011. [Google Scholar] [CrossRef] [PubMed]
- Calcagno, S.R.; Li, S.; Colon, M.; Kreinest, P.A.; Thompson, E.A.; Fields, A.P.; Murray, N.R. OncogenicK-ras promotes early carcinogenesis in the mouse proximal colon. Int. J. Cancer 2008, 122, 2462–2470. [Google Scholar] [CrossRef]
- Maitra, R.; Thavornwatanayong, T.; Venkatesh, M.K.; Chandy, C.; Vachss, D.; Augustine, T.; Guzik, H.; Koba, W.; Liu, Q.; Goel, S. Development and Characterization of a Genetic Mouse Model of KRAS Mutated Colorectal Cancer. Int. J. Mol. Sci. 2019, 20, 5677. [Google Scholar] [CrossRef]
- Drosten, M.; Guerra, C.; Barbacid, M. Genetically Engineered Mouse Models of K-Ras-Driven Lung and Pancreatic Tumors: Validation of Therapeutic Targets. Cold Spring Harb. Perspect. Med. 2017, 8, a031542. [Google Scholar] [CrossRef] [PubMed]
- Brannan, C.I.; Perkins, A.S.; Vogel, K.S.; Ratner, N.; Nordlund, M.L.; Reid, S.W.; Buchberg, A.M.; Jenkins, N.A.; Parada, L.F.; Copeland, N.G. Targeted disruption of the neurofibromatosis type-1 gene leads to developmental abnormalities in heart and various neural crest-derived tissues. Genes Dev. 1994, 8, 1019–1029. [Google Scholar] [CrossRef]
- Araki, T.; Mohi, M.G.; Ismat, F.A.; Bronson, R.T.; Williams, I.R.; Kutok, J.L.; Yang, W.; Pao, L.I.; Gilliland, D.G.; Epstein, J.A.; et al. Mouse model of Noonan syndrome reveals cell type– and gene dosage–dependent effects of Ptpn11 mutation. Nat. Med. 2004, 10, 849–857. [Google Scholar] [CrossRef]
- Schuhmacher, A.J.; Guerra, C.; Sauzeau, V.; Cañamero, M.; Bustelo, X.R.; Barbacid, M. A mouse model for Costello syndrome reveals an Ang II–mediated hypertensive condition. J. Clin. Investig. 2008, 118, 2169–2179. [Google Scholar] [CrossRef]
- Hernández-Porras, I.; Guerra, C. Modeling RASopathies with Genetically Modified Mouse Models. In Methods in Molecular Biology; Springer Nature: Cham, Switzerland, 2017. [Google Scholar]
- Pai, E.F.; Kabsch, W.; Krengel, U.; Holmes, K.C.; John, J.; Wittinghofer, A. Structure of the guanine-nucleotide-binding domain of the Ha-ras oncogene product p21 in the triphosphate conformation. Nature 1989, 341, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Brunner, T.B.; Hahn, S.M.; Gupta, A.K.; Muschel, R.J.; McKenna, W.G.; Bernhard, E.J. Farnesyltransferase inhibitors: An overview of the results of preclinical and clinical investigations. Cancer Res. 2003, 63, 5656–5668. [Google Scholar]
- Zhang, F.L.; Kirschmeier, P.; Carr, D.; James, L.; Bond, R.W.; Wang, L.; Patton, R.; Windsor, W.T.; Syto, R.; Zhang, R.; et al. Characterization of Ha-Ras, N-Ras, Ki-Ras4A, and Ki-Ras4B as in Vitro Substrates for Farnesyl Protein Transferase and Geranylgeranyl Protein Transferase Type I. J. Biol. Chem. 1997, 272, 10232–10239. [Google Scholar] [CrossRef] [PubMed]
- Tamanoi, F.; Lu, J. Recent Progress in Developing Small Molecule Inhibitors Designed to Interfere with Ras Membrane Association: Toward Inhibiting K-Ras and N-Ras Functions, 1st ed.; Elsevier Inc: Amsterdam, The Netherlands, 2013; Volume 34, ISBN 9780124201460. [Google Scholar]
- Zimmermann, G.; Papke, B.; Ismail, S.; Vartak, N.; Chandra, A.; Hoffmann, M.; Hahn, S.A.; Triola, G.; Wittinghofer, A.; Bastiaens, P.I.H.; et al. Small molecule inhibition of the KRAS–PDEδ interaction impairs oncogenic KRAS signalling. Nature 2013, 497, 638–642. [Google Scholar] [CrossRef] [PubMed]
- Novotny, C.J.; Hamilton, G.L.; McCormick, F.; Shokat, K.M. Farnesyltransferase-Mediated Delivery of a Covalent Inhibitor Overcomes Alternative Prenylation to Mislocalize K-Ras. ACS Chem. Biol. 2017, 12, 1956–1962. [Google Scholar] [CrossRef] [PubMed]
- Cox, A.D.; Fesik, S.W.; Kimmelman, A.C.; Luo, J.; Der, C.J. Drugging the undruggable RAS: Mission Possible? Nat. Rev. Drug Discov. 2014, 13, 828–851. [Google Scholar] [CrossRef] [PubMed]
- Poulikakos, P.I.; Zhang, C.; Bollag, G.; Shokat, K.M.; Rosen, N. RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature 2010, 464, 427–430. [Google Scholar] [CrossRef] [PubMed]
- Heidorn, S.J.; Milagre, C.; Whittaker, S.; Nourry, A.; Niculescu-Duvas, I.; Dhomen, N.; Hussain, J.; Reis-Filho, J.S.; Springer, C.J.; Pritchard, C.; et al. Kinase-Dead BRAF and Oncogenic RAS Cooperate to Drive Tumor Progression through CRAF. Cell 2010, 140, 209–221. [Google Scholar] [CrossRef]
- Karoulia, Z.; Gavathiotis, E.; Poulikakos, P.I. New perspectives for targeting RAF kinase in human cancer. Nat. Rev. Cancer 2017, 17, 676–691. [Google Scholar] [CrossRef]
- Degirmenci, U.; Wang, M.; Hu, J. Targeting Aberrant RAS/RAF/MEK/ERK Signaling for Cancer Therapy. Cells 2020, 9, 198. [Google Scholar] [CrossRef]
- Ye, M.; Shima, F.; Muraoka, S.; Liao, J.; Okamoto, H.; Yamamoto, M.; Tamura, A.; Yagi, N.; Ueki, T.; Kataoka, T. Crystal Structure of M-Ras Reveals a GTP-bound “Off” State Conformation of Ras Family Small GTPases. J. Biol. Chem. 2005, 280, 31267–31275. [Google Scholar] [CrossRef]
- Orgován, Z.; Keserű, G.M. Small molecule inhibitors of RAS proteins with oncogenic mutations. Cancer Metastasis Rev. 2020, 39, 1107–1126. [Google Scholar] [CrossRef]
- Ostrem, J.M.; Peters, U.; Sos, M.L.; Wells, J.A.; Shokat, K.M. K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions. Nature 2013, 503, 548–551. [Google Scholar] [CrossRef] [PubMed]
- Ni, D.; Li, X.; He, X.; Zhang, H.; Zhang, J.; Lu, S. Drugging K-RasG12C through covalent inhibitors: Mission possible? Pharmacol. Ther. 2019, 202, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Canon, J.; Rex, K.; Saiki, A.Y.; Mohr, C.; Cooke, K.; Bagal, D.; Gaida, K.; Holt, T.; Knutson, C.G.; Koppada, N.; et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 2019, 575, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Hallin, J.; Engstrom, L.D.; Hargis, L.; Calinisan, A.; Aranda, R.; Briere, D.M.; Sudhakar, N.; Bowcut, V.; Baer, B.R.; Ballard, J.A.; et al. The KRASG12C Inhibitor MRTX849 Provides Insight toward Therapeutic Susceptibility of KRAS-Mutant Cancers in Mouse Models and Patients. Cancer Discov. 2020, 10, 54–71. [Google Scholar] [CrossRef] [PubMed]
- Akhave, N.S.; Biter, A.B.; Hong, D.S. Mechanisms of Resistance to KRASG12C-Targeted Therapy. Cancer Discov. 2021. [Google Scholar] [CrossRef]
- Sanclemente, M.; Francoz, S.; Esteban-Burgos, L.; Bousquet-Mur, E.; Djurec, M.; Lopez-Casas, P.P.; Hidalgo, M.; Guerra, C.; Drosten, M.; Musteanu, M.; et al. c-RAF Ablation Induces Regression of Advanced Kras/Trp53 Mutant Lung Adenocarcinomas by a Mechanism Independent of MAPK Signaling. Cancer Cell 2018, 33, 217–228.e4. [Google Scholar] [CrossRef]
- Jinesh, G.G.; Sambandam, V.; Vijayaraghavan, S.; Balaji, K.; Mukherjee, S. Molecular genetics and cellular events of K-Ras-driven tumorigenesis. Oncogene 2018, 37, 839–846. [Google Scholar] [CrossRef]
- Heng, H.H.; Bremer, S.W.; Stevens, J.B.; Horne, S.D.; Liu, G.; Abdallah, B.Y.; Ye, K.J.; Ye, C.J. Chromosomal instability (CIN): What it is and why it is crucial to cancer evolution. Cancer Metastasis Rev. 2013, 32, 325–340. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Pérez-Herrero, E.; Fernández-Medarde, A. Advanced targeted therapies in cancer: Drug nanocarriers, the future of chemotherapy. Eur. J. Pharm. Biopharm. 2015, 93, 52–79. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations All authors have read and agreed to the published version of the manuscript.. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernández-Medarde, A.; De Las Rivas, J.; Santos, E. 40 Years of RAS—A Historic Overview. Genes 2021, 12, 681. https://doi.org/10.3390/genes12050681
Fernández-Medarde A, De Las Rivas J, Santos E. 40 Years of RAS—A Historic Overview. Genes. 2021; 12(5):681. https://doi.org/10.3390/genes12050681
Chicago/Turabian StyleFernández-Medarde, Alberto, Javier De Las Rivas, and Eugenio Santos. 2021. "40 Years of RAS—A Historic Overview" Genes 12, no. 5: 681. https://doi.org/10.3390/genes12050681
APA StyleFernández-Medarde, A., De Las Rivas, J., & Santos, E. (2021). 40 Years of RAS—A Historic Overview. Genes, 12(5), 681. https://doi.org/10.3390/genes12050681







