The Diversity and Evolution of Sex Chromosomes in Frogs
Abstract
1. Sex Chromosome Evolution
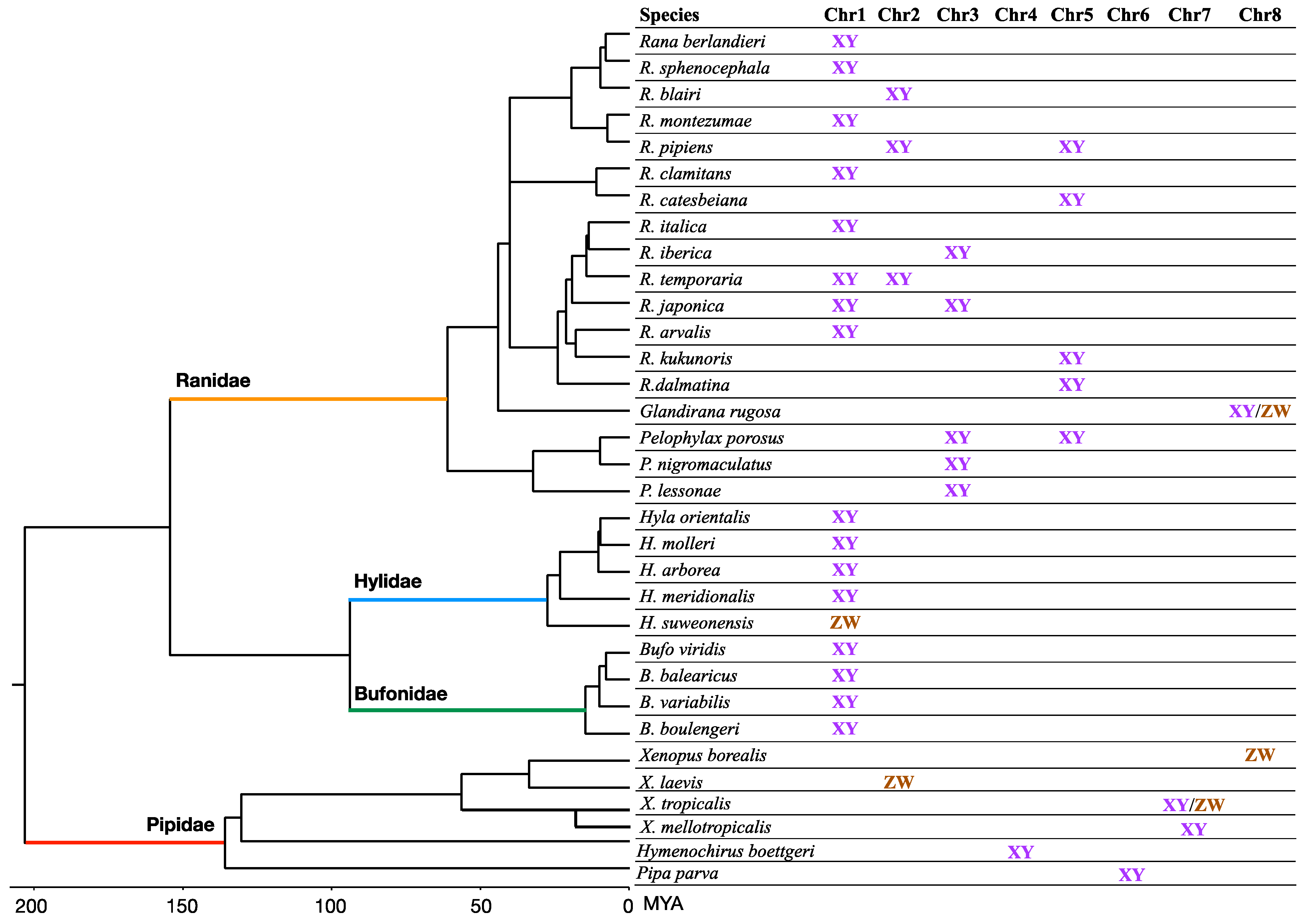
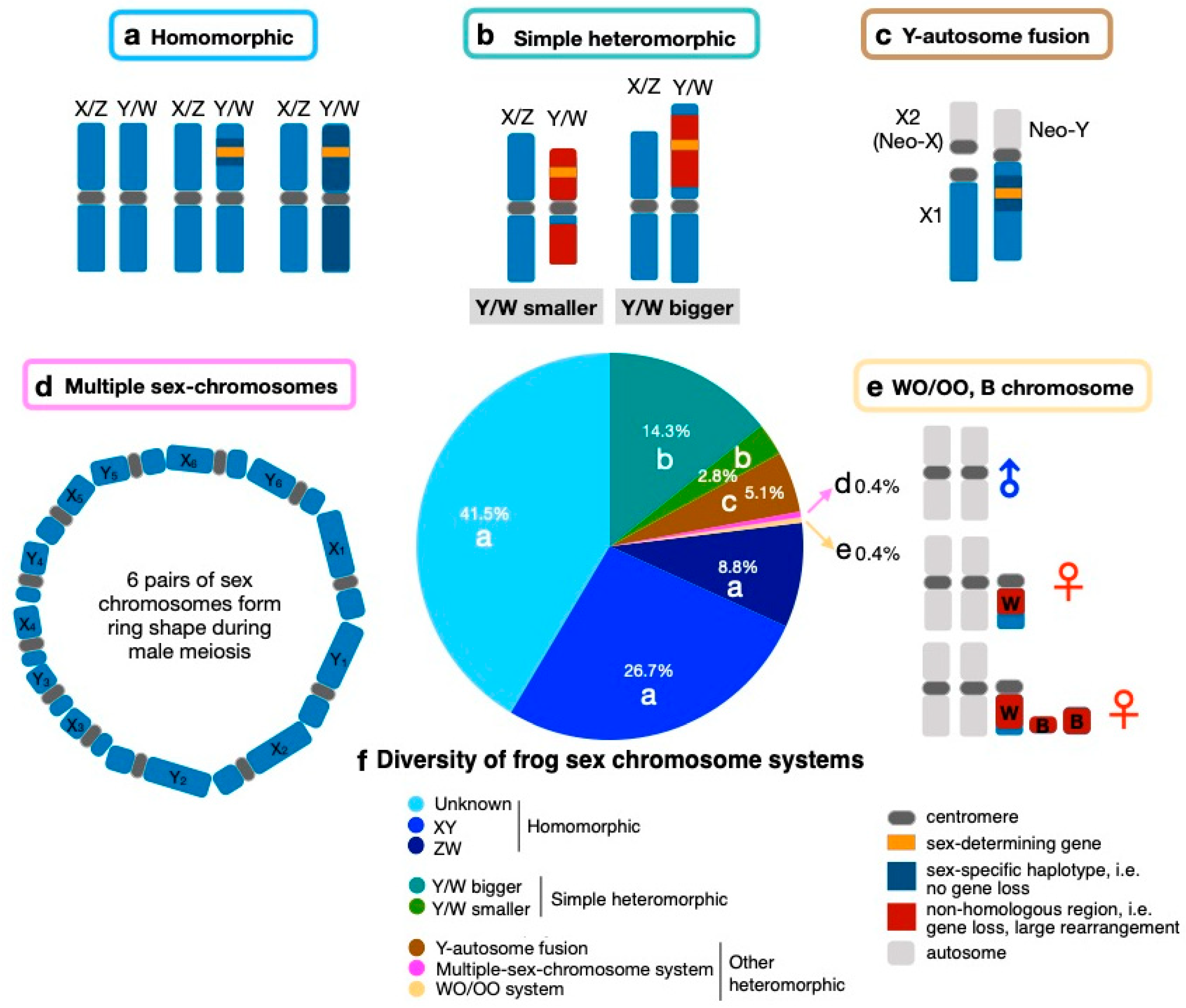
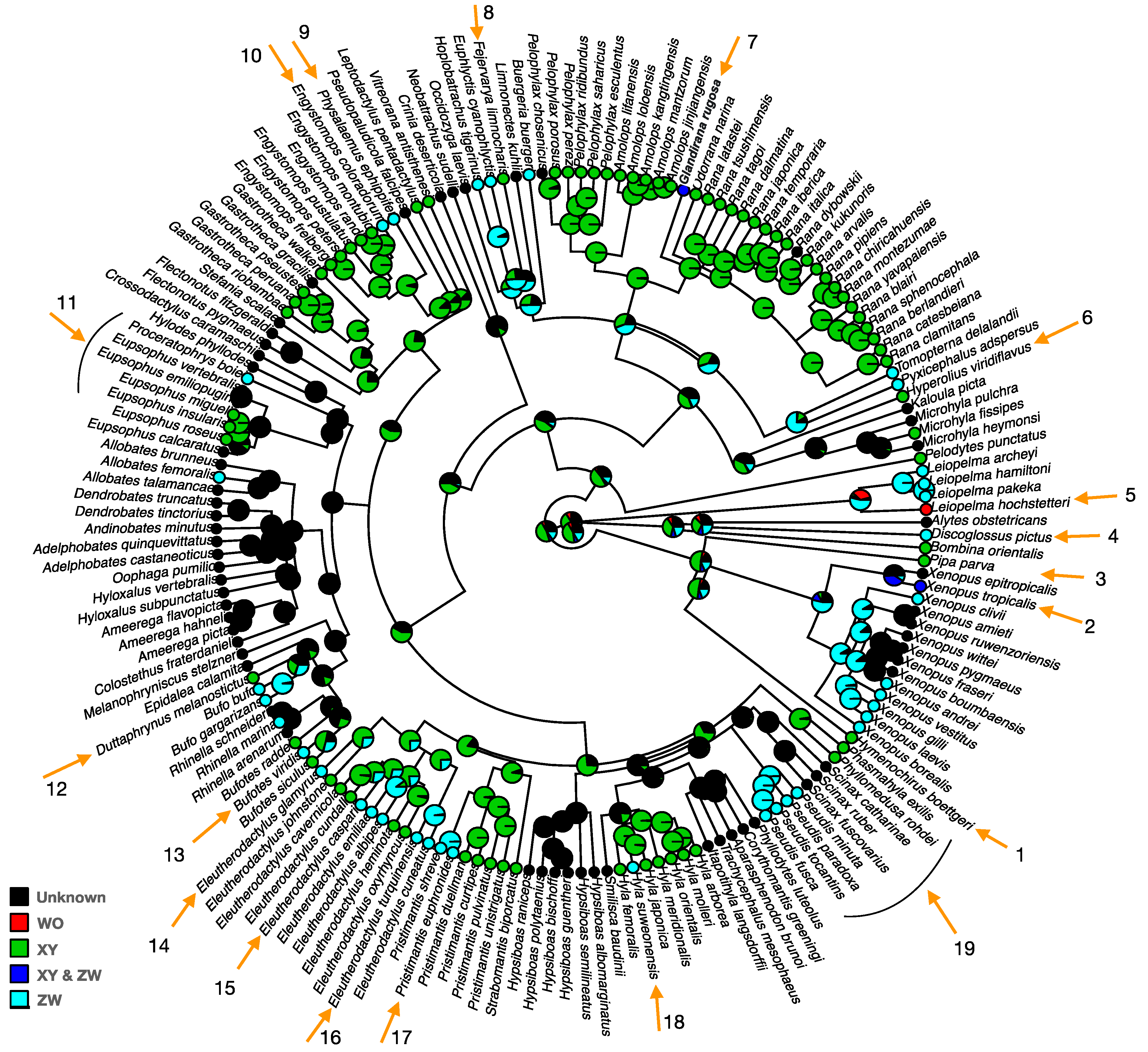
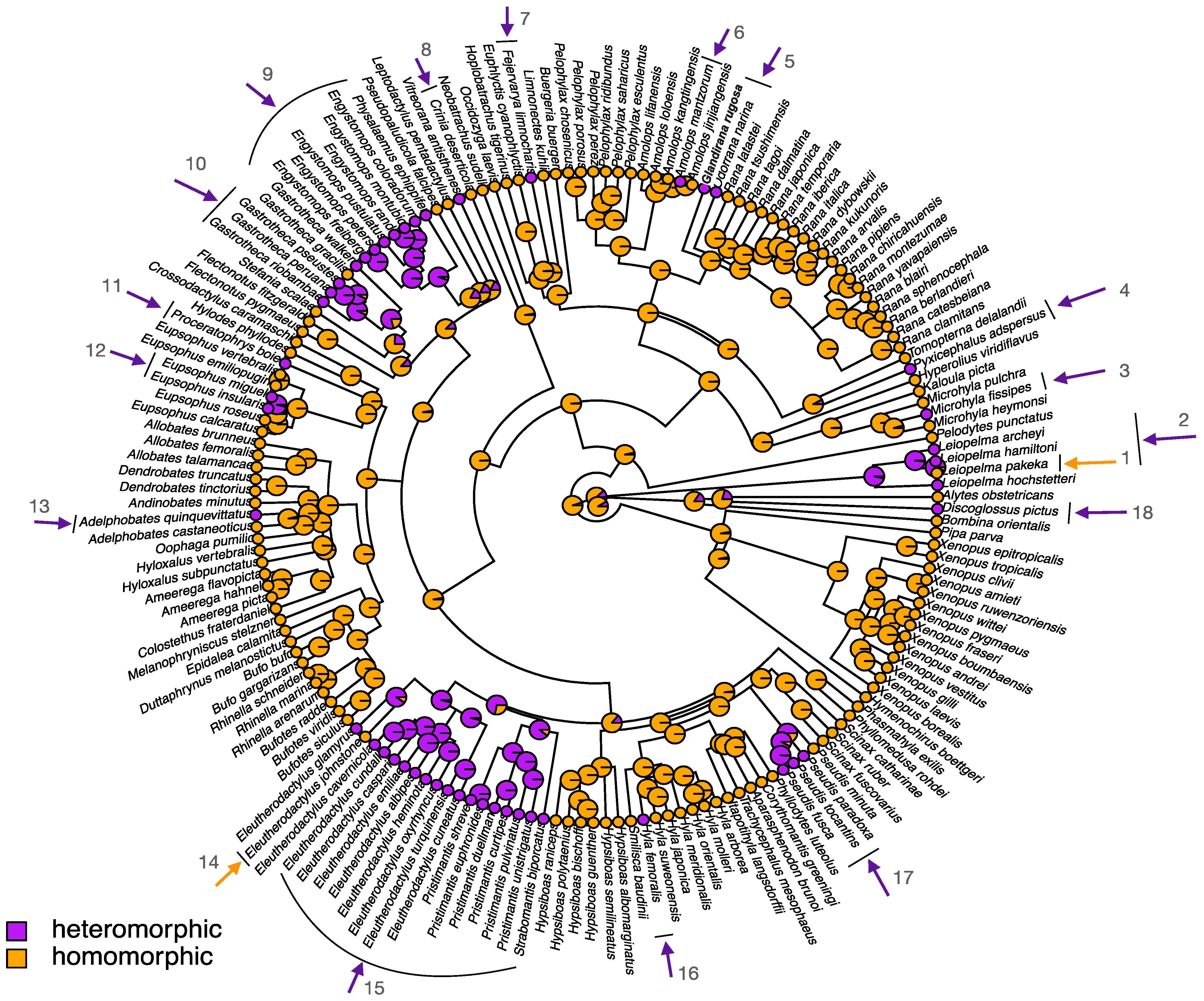
2. Sex Chromosome Diversity in Frogs
3. Sex Determination in Frogs
4. Homomorphic Sex Chromosomes in Frogs
4.1. Mechanism/Forces to Maintain Homomorphic Sex Chromosomes
5. Heteromorphic Sex Chromosomes in Frogs
5.1. What Forces Result in Heteromorphic Sex Chromosomes in Frog Lineages?
6. Sex Chromosome–Autosome Fusion
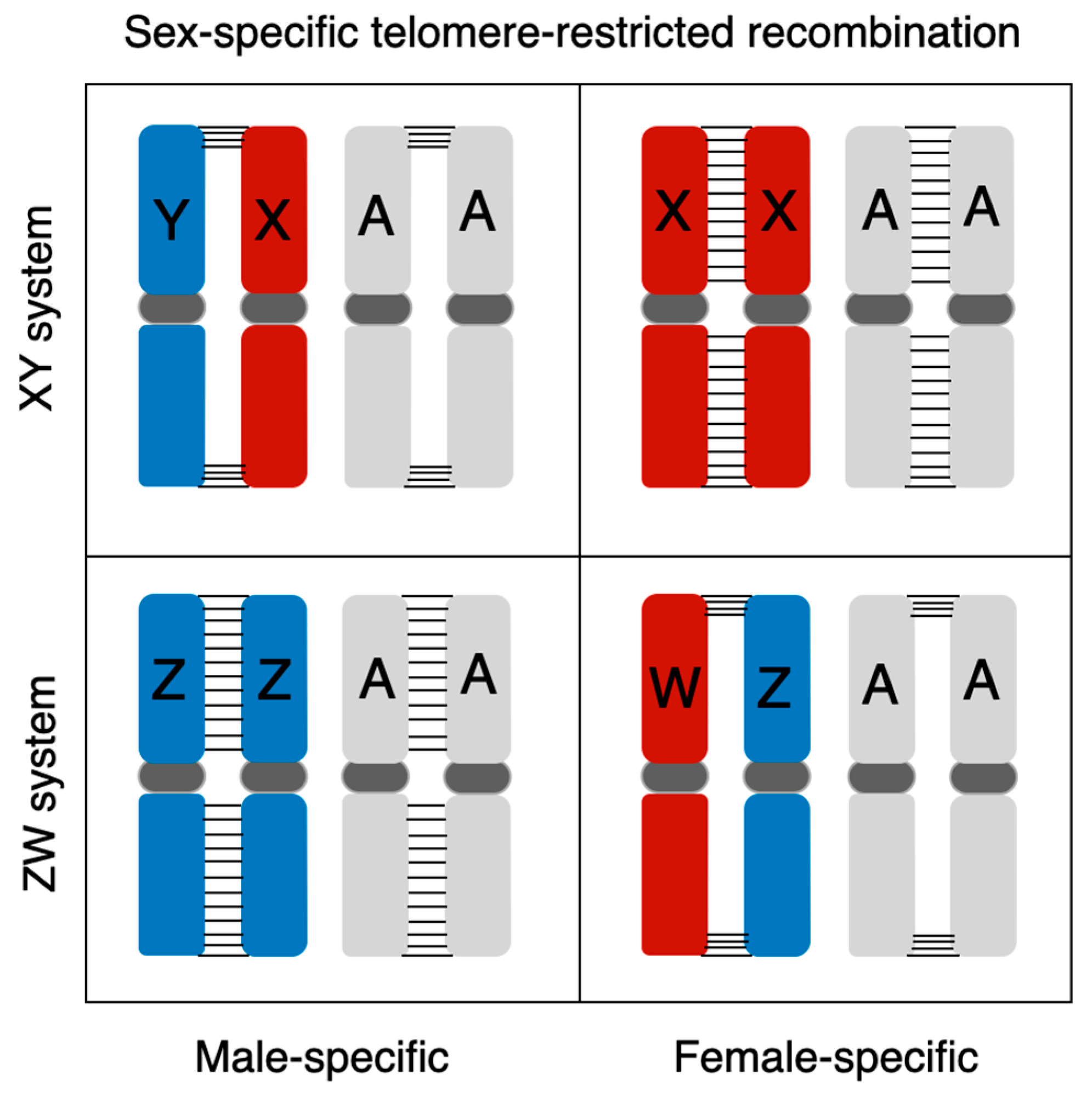
7. Extremely Sexual Dimorphic Recombination Pattern
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mullon, C.; Wright, A.E.; Reuter, M.; Pomiankowski, A.; Mank, J.E. Evolution of Dosage Compensation under Sexual Selection Differs between X and Z Chromosomes. Nat. Commun. 2015, 6, 7720. [Google Scholar] [CrossRef]
- Mank, J.E.; Hosken, D.J.; Wedell, N. Conflict on the Sex Chromosomes: Cause, Effect, and Complexity. Cold Spring Harb. Perspect. Biol. 2014, 6, 1–14. [Google Scholar] [CrossRef]
- Bachtrog, D. Y-chromosome Evolution: Emerging Insights into Processes of Y-chromosome Degeneration. Nat. Rev. Genet. 2013, 14, 113–124. [Google Scholar] [CrossRef]
- Wright, A.E.; Dean, R.; Zimmer, F.; Mank, J.E. How to Make a Sex Chromosome. Nat. Commun. 2016, 7, 12087. [Google Scholar] [CrossRef] [PubMed]
- Bull, J.J.J. Evolution of Sex Determining Mechanisms; Benjamin/Cummings Pub. Co.: Menlo Park, CA, USA, 1983. [Google Scholar]
- Bachtrog, D.; Kirkpatrick, M.; Mank, J.E.; McDaniel, S.F.; Pires, J.C.; Rice, W.R.; Valenzuela, N. Are All Sex Chromosomes Created Equal? Trends Genet. 2011, 27, 350–357. [Google Scholar] [CrossRef] [PubMed]
- Beukeboom, L.W.; Perrin, N. Evolution of Sex Determination; Oxford University Press: Oxford, UK, 2014. [Google Scholar]
- Bachtrog, D. The Temporal Dynamics of Processes Underlying Y Chromosome Degeneration. Genetics 2008, 179, 1513–1525. [Google Scholar] [CrossRef] [PubMed]
- Charlesworth, B.; Charlesworth, D. The Degeneration of Y Chromosomes. Philos. Trans. R. Soc. B Biol. Sci. 2000, 355, 1563–1572. [Google Scholar] [CrossRef] [PubMed]
- Bachtrog, D.; Mank, J.E.; Peichel, C.L.; Kirkpatrick, M.; Otto, S.P.; Ashman, T.L.; Hahn, M.W.; Kitano, J.; Mayrose, I.; Ming, R.; et al. Sex Determination: Why so Many Ways of Doing It? PLoS Biol. 2014, 12, e1001899. [Google Scholar] [CrossRef]
- Devlin, R.H.; Nagahama, Y. Sex Determination and Sex Differentiation in Fish: An Overview of Genetic, Physiological, and Environmental Influences. Aquaculture 2002, 208, 191–364. [Google Scholar] [CrossRef]
- Schmid, M.; Steinlein, C.; Bogart, J.P.; Feichtinger, W.; León, P.; La Marca, E.; Díaz, L.M.; Sanz, A.; Chen, S.H.; Hedges, S.B. The Chromosomes of Terraranan Frogs: Insights into Vertebrate Cytogenetics. Cytogenet. Genome Res. 2010, 130–131, 1–568. [Google Scholar] [CrossRef]
- Ming, R.; Bendahmane, A.; Renner, S.S. Sex Chromosomes in Land Plants. Annu. Rev. Plant Biol. 2011, 62, 485–514. [Google Scholar] [CrossRef]
- Fisher, R.A. The Evolution of Dominance. Biol. Rev. Biol. Proc. Camb. Philos. Soc. 1931, 6, 345–368. [Google Scholar] [CrossRef]
- Charlesworth, B.; Crow, J.F. Model for Evolution of Y Chromosomes and Dosage Compensation. Proc. Natl. Acad. Sci. USA 1978, 75, 5618–5622. [Google Scholar] [CrossRef] [PubMed]
- Charlesworth, D.; Charlesworth, B.; Marais, G. Steps in the Evolution of Heteromorphic Sex Chromosomes. Heredity 2005, 95, 118–128. [Google Scholar] [CrossRef]
- Rice, W.R. The Accumulation of Sexually Antagonistic Genes as a Selective Agent Promoting the Evolution of Reduced Recombination between Primitive Sex Chromosomes. Evolution 1987, 41, 911–914. [Google Scholar] [CrossRef] [PubMed]
- Bachtrog, D. A Dynamic View of Sex Chromosome Evolution. Curr. Opin. Genet. Dev. 2006, 16, 578–585. [Google Scholar] [CrossRef]
- Furman, B.L.S.; Metzger, D.C.H.; Darolti, I.; Wright, A.E.; Sandkam, B.A.; Almeida, P.; Shu, J.J.; Mank, J.E. Sex Chromosome Evolution: So Many Exceptions to the Rules. Genome Biol. Evol. 2020, 12, 750–763. [Google Scholar] [CrossRef]
- Stöck, M.; Horn, A.; Grossen, C.; Lindtke, D.; Sermier, R.; Betto-Colliard, C.; Dufresnes, C.; Bonjour, E.; Dumas, Z.; Luquet, E.; et al. Ever-young Sex Chromosomes in European Tree Frogs. PLoS Biol. 2011, 9, e1001062. [Google Scholar] [CrossRef]
- Guerrero, R.F.; Kirkpatrick, M.; Perrin, N. Cryptic Recombination in the Ever-young Sex Chromosomes of Hylid Frogs. J. Evol. Biol. 2012, 25, 1947–1954. [Google Scholar] [CrossRef]
- Stöck, M.; Savary, R.; Betto-Colliard, C.; Biollay, S.; Jourdan-Pineau, H.; Perrin, N. Low Rates of X-Y Recombination, not Turnovers, Account for Homomorphic Sex Chromosomes in Several Diploid Species of Palearctic Green Toads (Bufo viridis Subgroup). J. Evol. Biol. 2013, 26, 674–682. [Google Scholar] [CrossRef]
- Yazdi, H.P.; Silva, W.T.A.F.; Suh, A. Why do Some Sex Chromosomes Degenerate More Slowly Than Others? The Odd Case of Ratite Sex Chromosomes. Genes 2020, 11, 1153. [Google Scholar] [CrossRef]
- Yazdi, H.P.; Ellegren, H. Old but Not (So) Degenerated-slow Evolution of Largely Homomorphic Sex Chromosomes in Ratites. Mol. Biol. Evol. 2014, 31, 1444–1453. [Google Scholar] [CrossRef]
- Furman, B.L.S.; Evans, B.J. Divergent Evolutionary Trajectories of Two Young, Homomorphic, and Closely Related Sex Chromosome Systems. Genome Biol. Evol. 2018, 10, 742–755. [Google Scholar] [CrossRef]
- Perrin, N. Sex-chromosome Evolution in Frogs: What Role for Sex-antagonistic Genes? Phil. Trans. R. Soc. B 2020. [Google Scholar] [CrossRef]
- Ma, W.-J.; Veltsos, P.; Toups, M.A.; Rodrigues, N.; Sermier, R.; Jeffries, D.L.; Perrin, N. Tissue Specificity and Dynamics of Sex-biased Gene Expression in a Common Frog Population with Differentiated, yet Homomorphic, Sex Chromosomes. Genes 2018, 9, 294. [Google Scholar] [CrossRef]
- Ma, W.-J.; Veltsos, P.; Sermier, R.; Parker, D.J.; Perrin, N. Evolutionary and Developmental Dynamics of Sex-biased Gene Expression in Common Frogs with Proto-Y chromosomes. Genome Biol. 2018, 19, 156. [Google Scholar] [CrossRef] [PubMed]
- Veltsos, P.; Rodrigues, N.; Studer, T.; Ma, W.-J.; Sermier, R.; Leuenberger, J.; Perrin, N. No Evidence That Y-chromosome Differentiation Affects Male Fitness in a Swiss Population of Common Frogs. J. Evol. Biol. 2020, 33, 401–409. [Google Scholar] [CrossRef]
- Schmid, M.; Nanda, I.; Steinlein, C.; Kausch, K.; Haaf, T.; Epplen, J.T. Sex-Determining Mechanisms and Sex Chromosomes in Amphibia. In Amphibian Cytogenetics and Evolution; Green, D.M., Sessions, S.K., Eds.; Academic Press: San Diego, CA, USA, 1991. [Google Scholar]
- Malcom, J.W.; Kudra, R.S.; Malone, J.H. The Sex Chromosomes of Frogs: Variability and Tolerance Offer Clues to Genome Evolution and Function. J. Genom. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, M. Sex Determination in Amphibians. Semin. Cell Dev. Biol. 2009, 20, 271–282. [Google Scholar] [CrossRef]
- Ggert, C.E. Sex Determination: The Amphibian Models. Reprod. Nutr. Dev. 2004, 4, 539–549. [Google Scholar] [CrossRef]
- Gatto, K.P.; Busin, C.S.; Lourenço, L.B. Unraveling the Sex Chromosome Heteromorphism of the Paradoxical Frog Pseudis tocantins. PLoS ONE 2016, 11, e0156176. [Google Scholar] [CrossRef] [PubMed]
- Schmid, M.; Steinlein, C. Chromosome Banding in Amphibia. XXXVI. Multimorphic Sex Chromosomes and an Enigmatic Sex Determination in Eleutherodactylus johnstonei (Anura, Eleutherodactylidae). Cytogenet. Genome Res. 2018, 154, 86–98. [Google Scholar] [CrossRef]
- Schmid, M.; Feichtinger, W.; Steinlein, C.; Rupprecht, A.; Haaf, T.; Kaiser, H. Chromosome Banding in Amphibia: XXIII. Giant W Sex Chromosomes and Extremely Small Genomes in Eleutherodactylus euphronides and Eleutherodactylus shrevei (Anura, Leptodactylidae). Cytogenet. Genome Res. 2002, 97, 81–94. [Google Scholar] [CrossRef]
- Schartl, M.; Schmid, M.; Nanda, I. Dynamics of Vertebrate Sex Chromosome Evolution: From Equal Size to Giants and Dwarfs. Chromosoma 2016, 125, 553–571. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.-J.; Rodrigues, N.; Sermier, R.; Brelsford, A.; Perrin, N. Dmrt1 Polymorphism Covaries with Sex-determination Patterns in Rana temporaria. Ecol. Evol. 2016, 6, 5017–5117. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, N.; Studer, T.; Dufresnes, C.; Ma, W.; Veltsos, P.; Perrin, N. Dmrt1 Polymorphism and Sex-chromosome Differentiation in Rana temporaria. Mol. Ecol. 2017, 26, 4897–4905. [Google Scholar] [CrossRef] [PubMed]
- Gazoni, T.; Haddad, C.F.B.; Narimatsu, H.; Cabral-de-Mello, D.C.; Lyra, M.L.; Parise-Maltempi, P.P. More Sex Chromosomes than Autosomes in the Amazonian Frog Leptodactylus pentadactylus. Chromosoma 2018, 127, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Green, D.M. Structure and Evolution of B Chromosomes in Amphibians. Cytogenet. Genome Res. 2004, 106, 235–242. [Google Scholar] [CrossRef]
- Green, D.M.; Zeyl, C.W.; Sharbel, T.F. The Evolution of Hypervariable Sex and Supernumerary (B) Chromosomes in the Relict New Zealand Frog, Leiopelma hochstetteri. J. Evol. Biol. 1993, 6, 417–441. [Google Scholar] [CrossRef]
- Jeffries, D.L.; Lavanchy, G.; Sermier, R.; Sredl, M.J.; Miura, I.; Borzée, A.; Barrow, L.N.; Canestrelli, D.; Crochet, P.A.; Dufresnes, C.; et al. A Rapid Rate of Sex-chromosome Turnover and Non-random Transitions in True Frogs. Nat. Commun. 2018, 9, 4088. [Google Scholar] [CrossRef]
- Gamble, T.; Coryell, J.; Ezaz, T.; Lynch, J.; Scantlebury, D.P.; Zarkower, D. Restriction site-associated DNA Sequencing (RAD-seq) Reveals an Extraordinary Number of Transitions among Gecko Sex-determining Systems. Mol. Biol. Evol. 2015, 32, 1296–1309. [Google Scholar] [CrossRef]
- Ross, J.A.; Urton, J.R.; Boland, J.; Shapiro, M.D.; Peichel, C.L. Turnover of Sex Chromosomes in the Stickleback Fishes (Gasterosteidae). PLoS Genet. 2009, 5, e1000391. [Google Scholar] [CrossRef]
- Phillips, R.B. Evolution of the Sex Chromosomes in Salmonid Fishes. Cytogenet. Genome Res. 2013, 141, 177–185. [Google Scholar] [CrossRef]
- Kikuchi, K.; Hamaguchi, S. Novel Sex-determining Genes in Fish and Sex Chromosome Evolution. Dev. Dyn. 2013, 242, 339–353. [Google Scholar] [CrossRef] [PubMed]
- Revell, M.L.J. Phytools: An R Package for Phylogenetic Comparative Biology (and Other Things). Methods Ecol. Evol. 2012, 3, 217–223. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef] [PubMed]
- Brelsford, A.; Rodrigues, N.; Perrin, N. High-density Linkage Maps Fail to Detect Any Genetic Component to Sex Determination in a Rana temporaria Family. J. Evol. Biol. 2016, 29, 220–225. [Google Scholar] [CrossRef]
- Miura, I. Sex Determination and Sex Chromosomes in Amphibia. Sex. Dev. 2017, 11, 298–306. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, S.; Okada, E.; Umemoto, H.; Tamura, K.; Uno, Y.; Nishida-umehara, C.; Matsuda, Y.; Takamatsu, N.; Shiba, T.; Ito, M. A W-linked DM-domain Gene, DM-W, Participates in Primary Ovary Development in Xenopus laevis. Proc. Natl. Acad. Sci. USA 2008, 105, 2469–2474. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, S.; Ikeda, N.; Izutsu, Y.; Shiba, T.; Takamatsu, N.; Ito, M. Opposite Roles of DMRT1 and Its W-linked Paralogue, DM-W, in Sexual Dimorphism of Xenopus laevis: Implications of a ZZ/ZW-type Sex-determining System. Development 2010, 137, 2519–2526. [Google Scholar] [CrossRef]
- Mawaribuchi, S.; Yoshimoto, S.; Ohashi, S.; Takamatsu, N.; Ito, M. Molecular Evolution of Vertebrate Sex-determining Genes. Chromosom. Res. 2012, 20, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Cauret, C.M.S.; Gansauge, M.T.; Tupper, A.S.; Furman, B.L.S.; Knytl, M.; Song, X.Y.; Greenbaum, E.; Meyer, M.; Evans, B.J. Developmental Systems Drift and the Drivers of Sex Chromosome Evolution. Mol. Biol. Evol. 2020, 37, 799–810. [Google Scholar] [CrossRef]
- Sun, Y.; Xiong, Z.; Xiang, X.; Liu, S.; Zhou, W.; Tu, X.; Zhong, L. Whole-genome Sequence of the Tibetan Frog Nanorana parkeri and the Comparative Evolution of Tetrapod Genomes. Proc. Natl. Acad. Sci. USA 2015, 112, E1257–E1262. [Google Scholar] [CrossRef] [PubMed]
- Brelsford, A.; Stöck, M.; Betto-Colliard, C.; Dubey, S.; Dufresnes, C.; Jourdan-Pineau, H.; Rodrigues, N.; Savary, R.; Sermier, R.; Perrin, N. Homologous Sex Chromosomes in Three Deeply Divergent Anuran Species. Evolution 2013, 67, 2434–2440. [Google Scholar] [CrossRef]
- Palomar, G.; Ahmad, F.; Vasemägi, A.; Matsuba, C.; Nicieza, A.G.; Cano, J.M. Comparative High-density Linkage Mapping Reveals Conserved Genome Structure but Variation in Levels of Heterochiasmy and Location of Recombination Cold Spots in the Common Frog. G3 Genes Genomes Genet. 2017, 7, 637–645. [Google Scholar] [CrossRef]
- Dufresnes, C.; Borzee, A.; Horn, A.; Stock, M.; Ostini, M.; Sermier, R.; Wassef, J.; Litvinchuck, S.N.; Kosch, T.A.; Waldman, B.; et al. Sex-chromosome Homomorphy in Palearctic Tree Frogs Results from Both Turnovers and X-Y Recombination. Mol. Biol. Evol. 2015, 32, 2328–2337. [Google Scholar] [CrossRef]
- Stöck, M.; Croll, D.; Dumas, Z.; Biollay, S.; Wang, J.; Perrin, N. A Cryptic Heterogametic Transition Revealed by Sex-linked DNA Markers in Palearctic Green Toads. J. Evol. Biol. 2011, 24, 1064–1070. [Google Scholar] [CrossRef]
- Miura, I. An Evolutionary Witness: The Frog Rana rugosa Underwent Change of Heterogametic Sex from XY Male to ZW Female. Sex. Dev. 2008, 1, 323–331. [Google Scholar] [CrossRef]
- Furman, B.L.S.; Cauret, C.M.S.; Knytl, M.; Song, X.Y.; Premachandra, T.; Ofori-Boateng, C.; Jordan, D.C.; Horb, M.E.; Evans, B.J. A Frog with Three Sex Chromosomes That Co-mingle Together in Nature: Xenopus tropicalis Has a Degenerate W and a Y That Evolved from a Z Chromosome. PLoS Genet. 2020, 16, e1009121. [Google Scholar] [CrossRef] [PubMed]
- Roco, Á.S.; Olmstead, A.W.; Degitz, S.J.; Amano, T.; Zimmerman, L.B.; Bullejos, M. Coexistence of Y, W, and Z Sex Chromosomes in Xenopus tropicalis. Proc. Natl. Acad. Sci. USA 2015, 112, E4752–E4761. [Google Scholar] [CrossRef] [PubMed]
- Bewick, A.J.; Anderson, D.W.; Evans, B.J. Evolution of the Closely Related, Sex-related Genes Dm-w and Dmrt1 in African Clawed Frogs (Xenopus). Evolution 2011, 65, 698–712. [Google Scholar] [CrossRef]
- Uno, Y.; Nishida, C.; Takagi, C.; Igawa, T.; Ueno, N.; Sumida, M.; Matsuda, Y. Extraordinary Diversity in the Origins of Sex Chromosomes in Anurans Inferred from Comparative Gene Mapping. Cytogenet. Genome Res. 2015, 145, 218–229. [Google Scholar] [CrossRef]
- Rodrigues, N.; Vuille, Y.; Brelsford, A.; Merilä, J.; Perrin, N. The Genetic Contribution to Sex Determination and Number of Sex Chromosomes Vary among Populations of Common Frogs (Rana temporaria). Heredity 2016, 117, 25–32. [Google Scholar] [CrossRef]
- Toups, M.A.; Rodrigues, N.; Perrin, N.; Kirkpatrick, M. A Reciprocal Translocation Radically Reshapes Sex-linked Inheritance in the Common Frog. Mol. Ecol. 2019, 28, 1877–1889. [Google Scholar] [CrossRef] [PubMed]
- Witschi, E. Studies on Sex Differentiation and Sex Determination in Amphibians. IV. The Geographical Distribution of the Sex Races of the European Grass Frog (Rana temporaria, L.). A Contribution to the Problem of the Evolution of Sex. J. Exp. Zool. 1930, 56, 149–165. [Google Scholar] [CrossRef]
- Teacher, A.G.F.; Garner, T.W.J.; Nichols, R.A. European Phylogeography of the Common Frog (Rana temporaria): Routes of Postglacial Colonization into the British Isles, and Evidence for an Irish Glacial Refugium. Heredity 2009, 102, 490–496. [Google Scholar] [CrossRef]
- Veith, M.; Baumgart, A.; Dubois, A.; Ohler, A.; Galán, P.; Vieites, D.R.; Nieto-Román, S.; Vences, M. Discordant Patterns of Nuclear and Mitochondrial Introgression in Iberian Populations of the European Common Frog (Rana temporaria). J. Hered. 2012, 103, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Spasić-Bošković, O.; Tanić, N.; Blagojević, J.; Vujošević, M. Comparative Cytogenetic Analysis of European Brown Frogs: Rana temporaria, R. dalmatina and R. graeca. Caryologia 1997, 50, 139–149. [Google Scholar] [CrossRef]
- Rodrigues, N.; Betto-Colliard, C.; Jourdan-Pineau, H.; Perrin, N. Within-population Polymorphism of Sex-determination Systems in the Common Frog (Rana temporaria). J. Evol. Biol. 2013, 26, 1569–1577. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, N.; Vuille, Y.; Loman, J.; Perrin, N.; Konsult, R.; Rodrigues, N. Sex-chromosome Differentiation and ‘Sex Races’ in the Common Frog (Rana temporaria). Proc. R. Soc. B Biol. Sci. 2015, 282, 20142726. [Google Scholar] [CrossRef] [PubMed]
- Phillips, B.C.; Rodrigues, N.; van Rensburg, A.J.; Perrin, N. Phylogeography, More Than Elevation, Accounts for Sex-chromosome Differentiation in Swiss Populations of the Common Frog. Evolution 2020, 644–654. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, F.E.; Ma, W.-J. Digest: Climate Plays Marginal Role for Homomorphic Sex Chromosome Differentiation in Common Frogs†. Evolution 2020, 74, 690–693. [Google Scholar] [CrossRef] [PubMed]
- Nagai, Y.; Doi, T.; Ito, K.; Yuasa, Y.; Fujitani, T.; Naito, J.I.; Ogata, M.; Miura, I. The Distributions and Boundary of Two Distinct, Local Forms of Japanese Pond Frog, Pelophylax porosus Brevipodus, Inferred from Sequences of Mitochondrial DNA. Front. Genet. 2018, 9, 5–10. [Google Scholar] [CrossRef]
- Wright, D.A.; Richards, C.M. Two Sex-linked Loci in the Leopard Frog, Rana pipiens. Genetics 1983, 103, 249–261. [Google Scholar] [CrossRef]
- Sumida, M.; Nishioka, M. Geographic Variability of Sex-linked Loci in the Japanese Brown Frog, Rana japonica. Sci. Rep. Lab. Amphib. Biol. Hiroshima Univ. 1994, 13, 173–195. [Google Scholar]
- Perrin, N. Sex Reversal: A Fountain of Youth for Sex Chromosomes? Evolution 2009, 63, 3043–3049. [Google Scholar] [CrossRef]
- Brelsford, A.; Dufresnes, C.; Perrin, N. Trans-species Variation in Dmrt1 is Associated with Sex Determination in Four European Tree-frog Species. Evolution 2016, 70, 840–847. [Google Scholar] [CrossRef]
- Wallace, H.; Wallace, B.M.N.; Badawy, G.M.I. Lampbrush Chromosomes and Chiasmata of Sex-reversed Crested Newts. Chromosoma 1997, 106, 526–533. [Google Scholar] [CrossRef]
- Kondo, M.; Nagao, E.; Mitani, H.; Shima, A. Differences in Recombination Frequencies during Female and Male Meioses of the Sex Chromosomes of the Medaka, Oryzias latipes. Genet. Res. 2001, 78, 23–30. [Google Scholar] [CrossRef]
- Inoue, H.; Fukumori, Y.; Hiboyoshi, T. Mapping of Autosomal Male-determining Factors of the Housefly, Musca domestica L., by Means of Sex-reversal. Jpn. J. Genet. 1983, 58, 451–461. [Google Scholar] [CrossRef]
- Rodrigues, N.; Studer, T.; Dufresnes, C.; Perrin, N. Sex-chromosome Recombination in Common Frogs Brings Water to the Fountain-of-youth. Mol. Biol. Evol. 2018, 35, 942–948. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M. Sex Chromosome Evolution in Non-mammalian Vertebrates. Curr. Opin. Genet. Dev. 2004, 14, 634–641. [Google Scholar] [CrossRef]
- Volff, J.N.; Nanda, I.; Schmid, M.; Schartl, M. Governing Sex Determination in Fish: Regulatory Putsches and Ephemeral Dictators. Sex. Dev. 2007, 1, 85–99. [Google Scholar] [CrossRef] [PubMed]
- Van Doorn, G.S.; Kirkpatrick, M. Turnover of Sex Chromosomes Induced by Sexual Conflict. Nature 2007, 449, 909–912. [Google Scholar] [CrossRef]
- Van Doorn, G.S.; Kirkpatrick, M. Transitions between Male and Female Heterogamety Caused by Sex-antagonistic Selection. Genetics 2010, 186, 629–645. [Google Scholar] [CrossRef]
- Gammerdinger, W.J.; Conte, M.A.; Sandkam, B.A.; Ziegelbecker, A.; Koblmüller, S.; Kocher, T.D. Novel Sex Chromosomes in 3 Cichlid Fishes from Lake Tanganyika. J. Hered. 2018, 109, 489–500. [Google Scholar] [CrossRef]
- Blaser, O.; Grossen, C.; Neuenschwander, S.; Perrin, N. Sex-chromosome Turnovers Induced by Deleterious Mutation Load. Evolution. 2013, 67, 635–645. [Google Scholar] [CrossRef] [PubMed]
- Blaser, O.; Neuenschwander, S.; Perrin, N. Sex-chromosome Turnovers: The Hot-Potato Model. Am. Nat. 2014, 183, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Gunski, R.J.; Kretschmer, R.; Santos De Souza, M.; De Oliveira Furo, I.; Barcellos, S.A.; Costa, A.L.; Cioffi, M.B.; De Oliveira, E.H.C.; Del Valle Garnero, A. Evolution of Bird Sex Chromosomes Narrated by Repetitive Sequences: Unusual W Chromosome Enlargement in Gallinula melanops (Aves: Gruiformes: Rallidae). Cytogenet. Genome Res. 2019. [Google Scholar] [CrossRef]
- Chalopin, D.; Volff, J.; Galiana, D.; Anderson, J.L.; Schartl, M. Transposable Elements and Early Evolution of Sex Chromosomes in Fish. Chromosom. Res. 2015, 23, 545–560. [Google Scholar] [CrossRef]
- Xu, L.; Wa Sin, S.Y.; Grayson, P.; Edwards, S.V.; Sackton, T.B.; Mank, J. Evolutionary Dynamics of Sex Chromosomes of Paleognathous Birds. Genome Biol. Evol. 2019, 11, 2376–2390. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhang, J.; Xu, X.; Witt, C.; Deng, Y.; Chen, G.; Meng, G.; Feng, S.; Szekely, T.; Zhang, G.; et al. Phylogeny, Transposable Element and Sex Chromosome Evolution of the Basal Lineage of Birds. bioRxiv 2019. [Google Scholar] [CrossRef]
- Wang, Z.-J.; Chen, G.-J.; Zhang, G.-J.; Zhou, Q. Dynamic Evolution of Transposable Elements, Demographic History, and Gene Content of Paleognathous Birds. Zool. Res. 2021, 42, 51–61. [Google Scholar] [CrossRef]
- Townsend, D.S.; Stewart, M.M. Direct Development in Eleutherodactylus coqui (Anura: Leptodactylidae): A Staging Table. Copeia 1985, 2, 423–436. [Google Scholar] [CrossRef]
- García-R, J.C.; Crawford, A.J.; Mendoza, Á.M.; Ospina, O.; Cardenas, H.; Castro, F. Comparative Phylogeography of Direct-developing Frogs (Anura: Craugastoridae: Pristimantis) in the Southern Andes of Colombia. PLoS ONE 2012, 7, e0046077. [Google Scholar] [CrossRef]
- Callery, E.M.; Fang, H.; Elinson, R.P. Frogs without Polliwogs: Evolution of Anuran Direct Development. BioEssays 2001, 23, 233–241. [Google Scholar] [CrossRef]
- Elinson, R.P.; del Pino, E.M. Developmental Diversity of Amphibians. Wiley Interdiscip. Rev. Dev. Biol. 2012, 1, 345–369. [Google Scholar] [CrossRef] [PubMed]
- Ohta, S. Sex Determination Mechanism in Buergeria buergeri (Schlegel) I. Heterozygosity of Chromosome Pair No.7 in the Female. Sci. Rep. Lab. Amphib. Biol. Hiroshima Univ. 1986, 8, 29–43. [Google Scholar]
- Pennell, M.W.; Kirkpatrick, M.; Otto, S.P.; Vamosi, J.C. Y Fuse? Sex Chromosome Fusions in Fishes and Reptiles. PLoS Genet 2015, 11, e1005237. [Google Scholar] [CrossRef] [PubMed]
- Burt, A.; Bell, G.; Harvey, P.H. Sex Differences in Recombination. J. Evol. Biol. 1991, 4, 259–277. [Google Scholar] [CrossRef]
- Lenormand, T.; Dutheil, J. Recombination Difference between Sexes: A Role for Haploid Selection. PLoS Biol. 2005, 3, e63. [Google Scholar] [CrossRef] [PubMed]
- Lenormand, T.; Engelstädter, J.; Johnston, S.E.; Wijnker, E.; Haag, C.R. Evolutionary Mysteries in Meiosis. Philos. Trans. R. Soc. B Biol. Sci. 2016, 371. [Google Scholar] [CrossRef] [PubMed]
- Sardell, J.M.; Kirkpatrick, M. Sex Differences in the Recombination Landscape. Am. Nat. 2020, 195, 361–379. [Google Scholar] [CrossRef] [PubMed]
- Haldane, J.B.S. Sex Ratio and Unisexual Sterility in Hybrid Animals. J. Genet. 1922, 12, 101–109. [Google Scholar] [CrossRef]
- Huxley, J.S. Sexual Difference of Linkage in Gammarus chevreuyxi. J. Genet. 1928, 20, 145–156. [Google Scholar] [CrossRef]
- Brelsford, A.; Dufresnes, C.; Perrin, N. High-density Sex-specific Linkage Maps of a European Tree Frog (Hyla arborea) Identify the Sex Chromosome without Information on Offspring Sex. Heredity 2016, 116, 177–181. [Google Scholar] [CrossRef]
- Bergero, R.; Gardner, J.; Bader, B.; Yong, L.; Charlesworth, D. Exaggerated Heterochiasmy in a Fish with Sex-linked Male Coloration Polymorphisms. Proc. Natl. Acad. Sci. USA 2019, 116, 6924–6931. [Google Scholar] [CrossRef] [PubMed]
- Dufresnes, C.; Brelsford, A.; Baier, F.; Perrin, N. When Sex Chromosomes Recombine only in the Heterogametic Sex: Heterochiasmy and Heterogamety in Hyla Tree Frogs. Mol. Biol. Evol. 2021, 38, 192–200. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, W.-J.; Veltsos, P. The Diversity and Evolution of Sex Chromosomes in Frogs. Genes 2021, 12, 483. https://doi.org/10.3390/genes12040483
Ma W-J, Veltsos P. The Diversity and Evolution of Sex Chromosomes in Frogs. Genes. 2021; 12(4):483. https://doi.org/10.3390/genes12040483
Chicago/Turabian StyleMa, Wen-Juan, and Paris Veltsos. 2021. "The Diversity and Evolution of Sex Chromosomes in Frogs" Genes 12, no. 4: 483. https://doi.org/10.3390/genes12040483
APA StyleMa, W.-J., & Veltsos, P. (2021). The Diversity and Evolution of Sex Chromosomes in Frogs. Genes, 12(4), 483. https://doi.org/10.3390/genes12040483





