Assessing Evolutionary Significant Units (ESU) of the Endangered Freshwater Pearl Mussel (Margaritifera margaritifera) in Southeast Norway on the Basis of Genetic Analysis
Abstract
1. Introduction
2. Materials and Methods
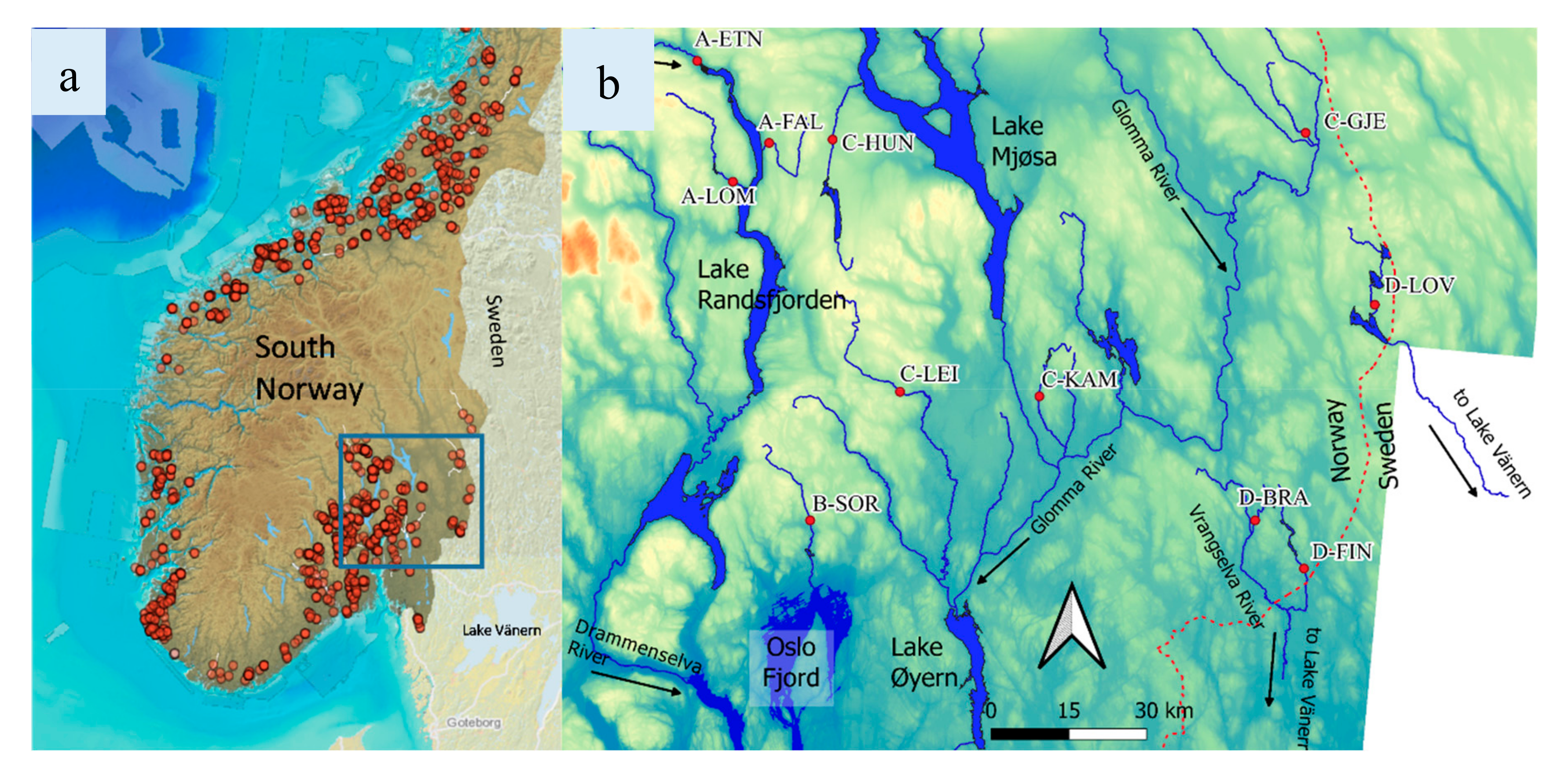
2.1. Study Area
2.2. Sampling and Analysis
2.3. Statistical Analysis
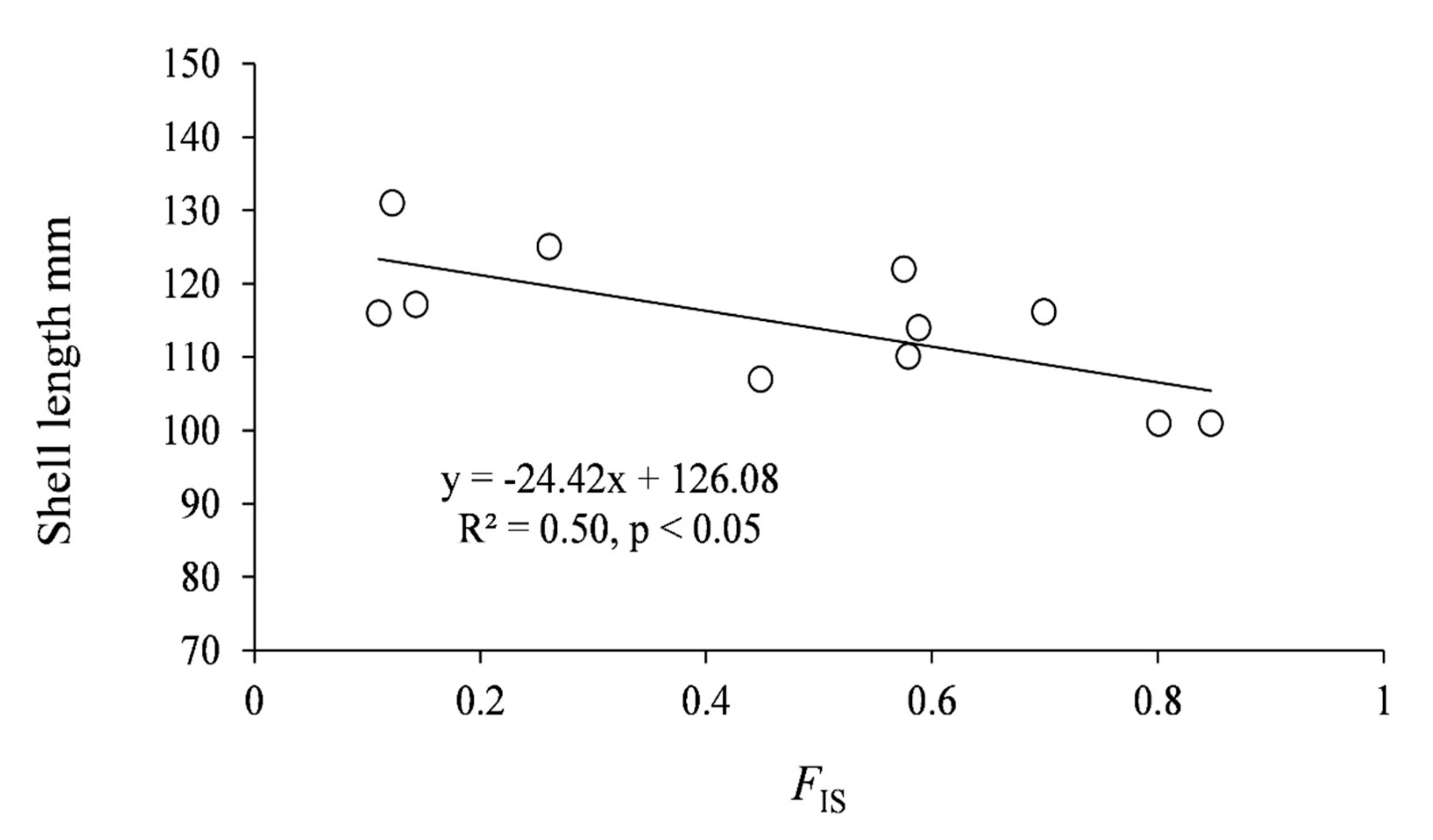
3. Results
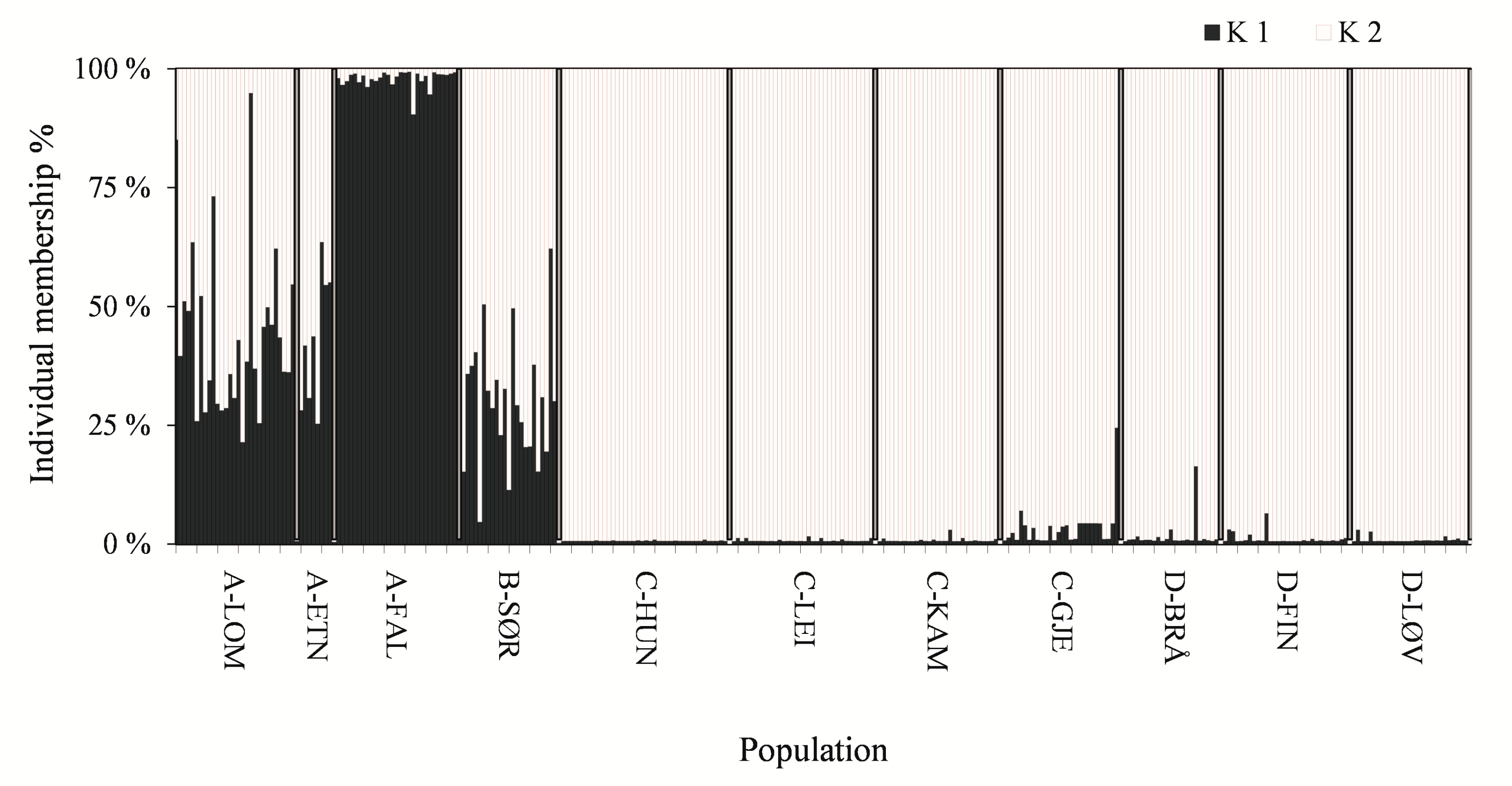
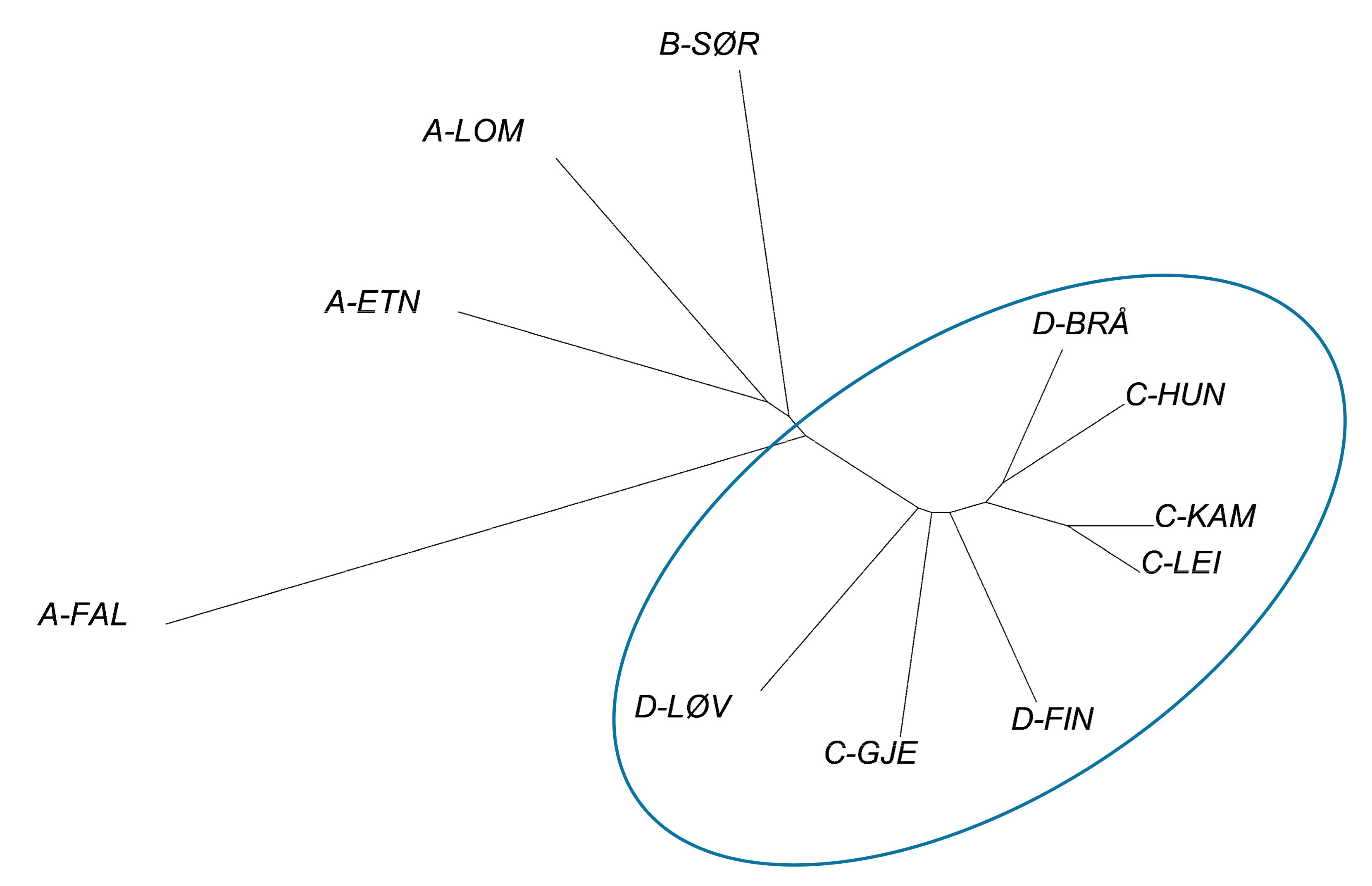
3.1. Genetic Structure
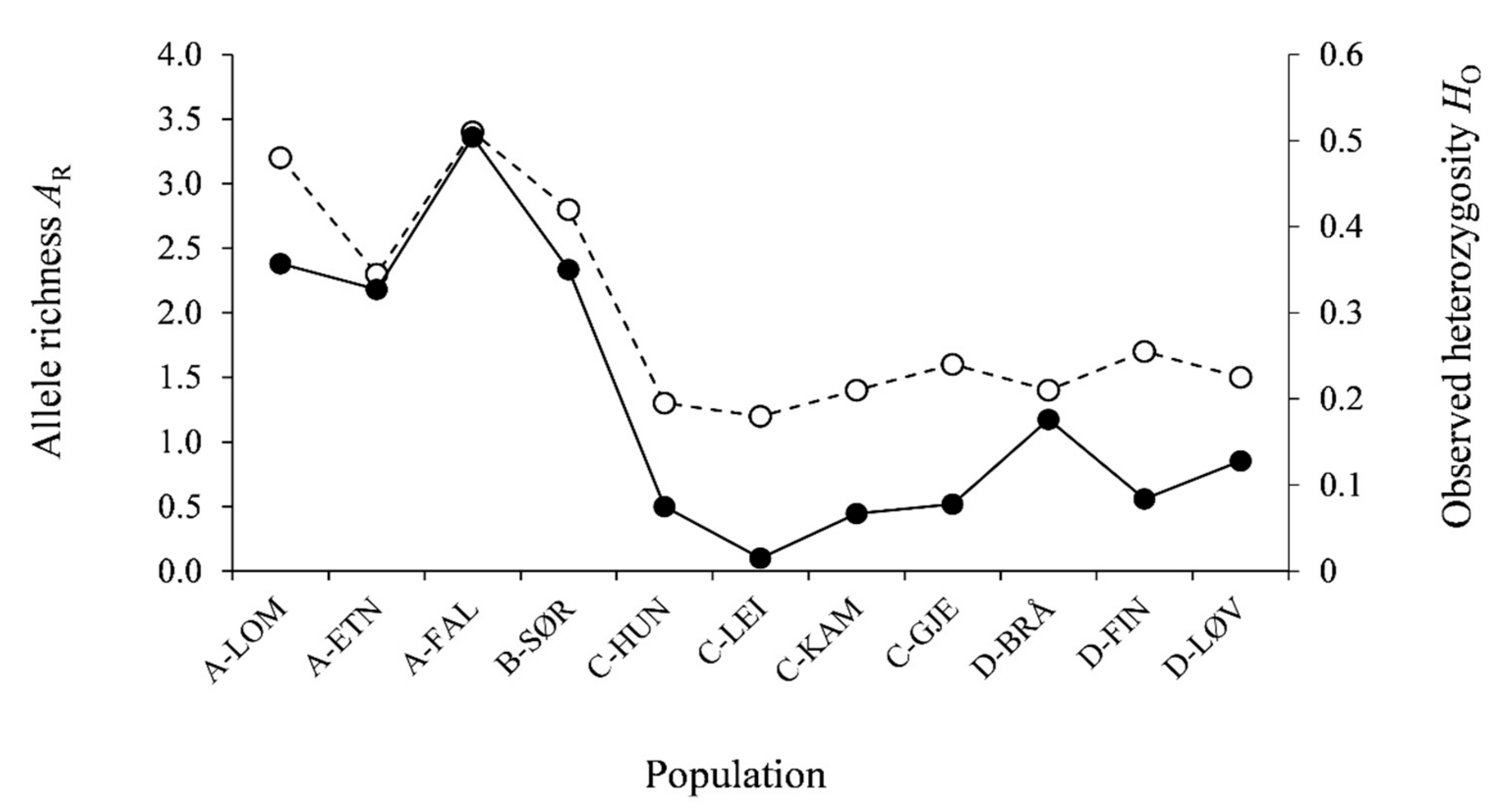
3.2. Genetic Diversity
4. Discussion
4.1. Genetic Structure and Probable Origin of the Study Populations
4.2. Genetic Diversity
5. Conclusions and Recommendations
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Geist, J.; Auerswald, K. Physicochemical stream bed characteristics and recruitment of freshwater pearl mussel (Margaritifera magaritifera). Freshw. Biol. 2007, 52, 2299–2316. [Google Scholar] [CrossRef]
- IUCN. International Union for Conservation of Nature Red List of Threatened Species. Available online: https://www.iucnredlist.org/search?query=margaritifera%20margaritifera&searchType=species (accessed on 7 July 2020).
- Lopes-Lima, M.; Sousa, R.; Teixeira, A.; Varandas, S.; Riccardi, N.; Aldridge, D.C.; Froufe, E. Newly developed microsatellite markers for the pan-European duck mussel, Anodonta anatina: Revisiting the main mitochondrial lineages. Aquat. Conserv. Mar. Freshw. Ecosyst. 2016, 26, 307–318. [Google Scholar] [CrossRef]
- Machordom, A.; Araujo, R.; Erpenbeck, D.; Ramos, M.A. Phylogeography and conservation genetics of endangered European Margaritiferidae (Bivalvia: Unionoidea). Biol. J. Linn. Soc. 2003, 78, 235–252. [Google Scholar] [CrossRef]
- Cuttelod, A.; Seddon, M.; Neubert, E. European Red List of Non-Marine Molluscs; Publication Office of the European Union: Luxembourg, 2011. [Google Scholar] [CrossRef]
- Young, M.R.; Cosgrove, P.J.; Hastie, L.C. The extent of, and causes for, the decline of highly threadened naiad: Margaritifera margaritifera. In Ecology and Evolution of the Freshwater Mussels Unionida; Bauer, G., Wachtler, K., Eds.; Springer: Berlin, Germany, 2001; pp. 337–358. [Google Scholar]
- Bauer, G.; Wächtler, K. Ecology and Evolution of the Freshwater Mussels Unionoida; Springer: Heidelberg, Germany, 2001; p. 145. [Google Scholar]
- Vaughn, C.C. Biodiversity Losses and Ecosystem Function in Freshwaters: Emerging Conclusions and Research Directions. Bioscience 2010, 60, 25–35. [Google Scholar] [CrossRef]
- Vaughn, C.C.; Hakenkamp, C.C. The functional role of burrowing bivalves in freshwater ecosystems. Freshw. Biol. 2001, 46, 1431–1446. [Google Scholar] [CrossRef]
- Strayer, D.L.; Caraco, N.F.; Cole, J.J.; Findlay, S.; Pace, M.L. Transformation of Freshwater Ecosystems by Bivalves. A case study of zebra mussels in the Hudson River. BioScience 1999, 49, 19–27. [Google Scholar] [CrossRef]
- Stoeckl, K.; Denic, M.; Geist, J. Conservation status of two endangered freshwater mussel species in Bavaria, Germany: Habitat quality, threats, and implications for conservation management. Aquat. Conserv. Mar. Freshw. Ecosyst. 2020, 30, 647–661. [Google Scholar] [CrossRef]
- Bauer, G. Threats to the fresh-water pearl mussel Margaritifera margaritifera in Central-Europe. Biol. Conserv. 1988, 45, 239–253. [Google Scholar] [CrossRef]
- Jung, M.; Scheder, C.; Gumpinger, C.; Waringer, J. Habitat traits, population structure and host specificity of the freshwater pearl mussel Margaritifera margaritifera in the Waldaist River (Upper Austria). Biologia 2013, 68, 922–931. [Google Scholar] [CrossRef]
- Larsen, B.M.; Magerøy, J.H. Elvemuslinglokaliteter i Norge—En Beskrivelse av Status som Grunnlag for Arbeid med Kartlegging og Tiltak i Handlingsplanen for 2019–2028; Norwegian Institute of Nature Research: Trondheim, Norway, 2019. [Google Scholar]
- Geist, J. Strategies for the conservation of endangered freshwater pearl mussels (Margaritifera margaritifera L.): A synthesis of Conservation Genetics and Ecology. Hydrobiologia 2010, 644, 69–88. [Google Scholar] [CrossRef]
- Ryder, O.A. Species conservation and systematics: The dilemma of subspecies. Trends Ecol. Evol. 1986, 1, 9–10. [Google Scholar] [CrossRef]
- Fraser, D.J.; Bernatchez, L. Adaptive evolutionary conservation: Towards a unified concept for defining conservation units. Mol. Ecol. 2001, 10, 2741–2752. [Google Scholar] [CrossRef] [PubMed]
- Moritz, C. Defining ’Evolutionarily Significant Units’ for conservation. Trends Ecol. Evol. 1994, 9, 373–375. [Google Scholar] [CrossRef]
- Moritz, C. Strategies to protect biological diversity and the evolutionary processes that sustain it. Syst. Biol. 2002, 51, 238–254. [Google Scholar] [CrossRef]
- Moritz, C. Conservation units and translocations: Strategies for conserving evolutionary processes. Hereditas 1999, 130, 217–228. [Google Scholar] [CrossRef]
- Waples, R.S.; Lindley, S.T. Genomics and conservation units: The genetic basis of adult migration timing in Pacific salmonids. Evol. Appl. 2018, 11, 1518–1526. [Google Scholar] [CrossRef]
- Frankel, O.H.; Soulé, M.E. Conservation and Evolution; Cambridge University Press: Cambridge, UK, 1981. [Google Scholar]
- Coates, D.J.; Byrne, M.; Moritz, C.C. Genetic Diversity and Conservation Units: Dealing With the Species-Population Continuum in the Age of Genomics. Front. Ecol. Evol. 2018, 6, 165. [Google Scholar] [CrossRef]
- Bauer, G. Variation in the life span and size of the freshwater pearl mussel. J. Anim. Ecol. 1992, 61, 425–436. [Google Scholar] [CrossRef]
- Chesney, H.C.G.; Oliver, P.G. Conservation issues for Margaritiferidae in British Isles and Western Europe. J. Conchol. Spec. Publ. 1998, 2, 231–242. [Google Scholar]
- Bauer, G. Reproductive strategy of the fresh-water pearl mussel Margaritifera margaritifera. J. Anim. Ecol. 1987, 56, 691–704. [Google Scholar] [CrossRef]
- Bouza, C.; Castro, J.; Martínez, P.; Amaro, R.; Fernández, C.; Ondina, P.; Outeiro, A.; San Miguel, E. Threatened freshwater pearl mussel Margaritifera margaritifera L. in NW Spain: Low and very structured genetic variation in southern peripheral populations assessed using microsatellite markers. Conserv. Genet. 2007, 8, 937–948. [Google Scholar] [CrossRef]
- Geist, J.; Kuehn, R. Genetic diversity and differentiation of central European freshwater pearl mussel (Margaritifera margaritifera L.) populations: Implications for conservation and management. Mol. Ecol. 2005, 14, 425–439. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, S.; Larsen, B.M.; Hindar, K. Host-dependent genetic variation in freshwater peral mussel (Margaritifera margaritifera L.). Hydrobiologia 2013, 735, 179–190. [Google Scholar] [CrossRef]
- Luck, G.W.; Daily, G.C.; Ehrlich, P.R. Population diversity and ecosystem services. Trends Ecol. Evol. 2003, 18, 331–336. [Google Scholar] [CrossRef]
- Zheng, H.; Li, L.; Zhang, G. Inbreeding depression for fitness-related traits and purging the genetic load in the hermaphroditic bay scallop Argopecten irradians irradians (Mollusca: Bivalvia). Aquaculture 2012, 366–367, 27–33. [Google Scholar] [CrossRef]
- Zheng, H.; Zhang, G.; Guo, X.; Liu, X. Inbreeding depression for various traits in two cultured populations of the American bay scallop, Argopecten irradians irradians Lamarck (1819) introduced into China. J. Exp. Mar. Biol. Ecol. 2008, 364, 42–47. [Google Scholar] [CrossRef]
- Zheng, H.P.; Zhang, G.F.; Liu, X.A.; Guo, X.M. Sustained response to selection in an introduced population of the hermaphroditic bay scallop Argopecten irradians irradians Lamarck (1819). Aquaculture 2006, 255, 579–585. [Google Scholar] [CrossRef]
- Hollenbeck, C.M.; Johnston, I.A. Genomic Tools and Selective Breeding in Molluscs. Front. Genet. 2018, 9, 253. [Google Scholar] [CrossRef]
- Dolmen, D.; Kleiven, E. Elvemuslingen Margaritifera margaritifera i Norge. In Zoologisk Notat; Norges Teknisk-Naturvitenskapelig Universitet: Trondheim, Norway, 1997; p. 31. [Google Scholar]
- Kleiven, E.; Dolmen, D. Eldre opplysningar om elvemuslingen Margaritifera margaritifera etter Joh. O. Simonnæs. Del 1. Fauna 2010, 63, 60–67. [Google Scholar]
- Kleiven, E.; Dolmen, D. Eldre opplysningar om elvemuslingen Margaritifera margaritifera etter Joh. O. Simonnæs. Del 2. Fauna 2010, 63, 108–116. [Google Scholar]
- Dolmen, D.; Kleiven, E. Distribution, status and threats of the freshwater pearl mussel Margaritifera margaritifera (Linnaeus) (Bivalvia, Margaritiferidae) in Norway. Fauna Norwegica 2008, 26/27, 3–14. [Google Scholar]
- Araujo, R.; Ramos, A. Action Plan for Margaritifera margaritifera in Europe; Council of Europe: Madrid, Spain, 2000; p. 41. [Google Scholar]
- Taskinen, J.; Berg, P.; Saarinen-Valta, M.; Välilä, S.; Mäenpää, E.; Myllynen, K.; Pakkala, J. Effect of pH, iron and aluminum on survival of early life history stages of the endangered freshwater pearl mussel, Margaritifera margaritifera. Toxicol. Environ. Chem. 2011, 93, 1764–1777. [Google Scholar] [CrossRef]
- Huitfeldt-Kaas, H. Ferskvandsfiskenes Utbredelse og Innvandring i Norge med et Tillægg om Krebsen; Centraltrykkeriet: Oslo, Norway, 1918. [Google Scholar]
- Garmo, Ø.; Johnsen, S.I. Vannkjemisk Utvikling og Biologisk Tilstand etter Kalkslutt i Innsjøer Hedmark; Norwegian Institute of Water Reseach: Oslo, Norway, 2019; p. 119. [Google Scholar]
- NBIC. Norwegian Biodiversity Information Centre Database; Artsdatabanken: Trondheim, Norway, 2020. [Google Scholar]
- De Geer, G. Om Skandinaviens Geografiska Utveckling Efter Istiden; British Library, Historical Print Editions: London, UK, 1896; Available online: https://www.amazon.com/Skandinaviens-geografiska-utveckling-Istiden-Text-Kartor/dp/1241416575 (accessed on 7 July 2020).
- Blair, S.D.; Matheson, D.; He, Y.; Goss, G.G. Reduced salinity tolerance in the Arctic grayling (Thymallus arcticus) is associated with rapid development of a gill interlamellar cell mass: Implications of high-saline spills on native freshwater salmonids. Conservation Physiology 2016, 4, cow010. [Google Scholar] [CrossRef] [PubMed]
- Larsen, B.M.; Saksgård, R. Overvåking av Elvemusling i Norge; Norwegien Institute of Nature Research: Trondheim, Norway, 2010; Report 718. [Google Scholar]
- Sandaas, K.; Enerud, J. Elvemusling (Margaritifera margaritifera) i Kampåa, Nes kommune i Akershus 2008–2010; Fylkesmannen i Oslo og Akershus. Miljøvernavdelingen: Oslo, Norway, 2011.
- Sandaas, K.; Enerud, J. Elvemusling Margaritifera margaritifera i Bråtaåa 2000–2012, Eidskog Kommune, Hedmark 2012; Naturfaglige Konsulenttjenester & Fisk-og Miljøundersøkelser: Oslo, Norway, 2012; p. 21. [Google Scholar]
- Sandaas, K.; Enerud, J. Elvemusling Margaritifera margaritifera i Trøftåa, Løvhaugsåa og Gjerda, Hedmark fylke 2014; Naturfaglige Konsulenttjenester & Fisk-og Miljøundersøkelser: Oslo, Norway, 2014; p. 13. [Google Scholar]
- Walsh, P.S.; Metzger, D.A.; Higuchi, R. Chelex 100 as a Medium for Simple Extraction of DNA for PCR-Based Typing from Forensic Material. BioTechniques 2013, 54, 134–139. [Google Scholar] [CrossRef] [PubMed]
- Garlie, S. Utvikling av Mikrosatellitt Multipleks PCR for Genetiske Studier av Margaritifera margaritifera. Master’s Thesis, Hedmark College, Hamar, Norway, 2010. [Google Scholar]
- Geist, J.; Rottmann, O.; Schröder, W.; Kühn, R. Development of microsatellite markers for the endangered freshwater pearl mussel Margaritifera margaritifera L. (Bivalvia: Unionoidea). Mol. Ecol. Notes 2003, 3, 444–446. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Earl, D.A.; von Holdt, B.M. Structure harvester: A website and program for visualizing structure output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Jakobsson, M.; Rosenberg, N.A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 2007, 23, 1801–1806. [Google Scholar] [CrossRef]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform. 2005, 1, 47–50. [Google Scholar] [CrossRef]
- Kalinowski, S.T. HP-Rare: A computer program performing rarefaction on measures of allelic diversity. Mol. Ecol. Notes 2005, 5, 187–189. [Google Scholar] [CrossRef]
- Kalinowski, S.T.; Wagner, A.P.; Taper, M.L. ML-Relate: A computer program for maximum likelihood estimation of relatedness and relationship. Mol. Ecol. Notes 2006, 6, 576–579. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GENALEX 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Weir, B.S.; Cockerham, C.C. Estimating F-statistics for the analysis of population-structure. Evolution 1984, 38, 1358–1370. [Google Scholar] [CrossRef] [PubMed]
- Nei, M. Genetic Distance between Populations. Am. Nat. 1972, 106, 283–292. [Google Scholar] [CrossRef]
- Paradis, E.; Blomberg, S.; Bolker, B.; Brown, J.; Claude, J.; Cuong, H.S.; Desper, R.; Didier, G.; Durand, B.; Dutheil, J.; et al. Analyses of Phylogenetics and Evolution (Ape). In R Package Ape, 5.4 ed.; APE: Marseille, France, 2020; p. 293. [Google Scholar]
- Cornuet, J.M.; Luikart, G. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 1996, 144, 2001–2014. [Google Scholar]
- Luikart, G.; Cornuet, J.M. Empirical evaluation of a test identifying recently bottlenecked populations from allele frequency data. Conserv. Biol. 1998, 12, 228–237. [Google Scholar] [CrossRef]
- Peery, M.Z.; Kirby, R.; Reid, B.N.; Stoelting, R.; Doucet-Bëer, E.; Robinson, S.; Vásquez-Carrillo, C.; Pauli, J.N.; Palsbøli, P.J. Reliability of genetic bottleneck tests for detecting recent population declines. Mol. Ecol. 2012, 21, 3403–3418. [Google Scholar] [CrossRef]
- Garza, J.C.; Williamson, E.G. Detection of reduction in population size using data from microsatellite loci. Mol. Ecol. 2001, 10, 305–318. [Google Scholar] [CrossRef]
- Karlsson, S.; Larsen, B.M. Genetiske Analyser av Elvemusling Margaritifera margaritifera (L.)—Et Nødvendig Verktøy for Riktig Forvaltning av Arten; Norwegian Institue of Nature Research: Trondheim, Norway, 2013; p. 44. [Google Scholar]
- Geist, J.; Söderberg, H.; Karlberg, A.; Kuehn, R. Drainage-independent genetic structure and high genetic diversity of endangered freshwater pearl mussels (Margaritifera margaritifera) in northern Europe. Conserv. Genet. 2010, 11, 1339–1350. [Google Scholar] [CrossRef]
- Larsen, B.M. Kalking har positiv effekt på elvemuslingen i Ogna. pH Status 2012, 4, 7–9. [Google Scholar]
- Haasl, R.J.; Payseur, B.A. Microsatellites as targets of natural selection. Mol. Biol. Evol. 2013, 30, 285–298. [Google Scholar] [CrossRef] [PubMed]
- Leberg, P.L.; Firmin, B.D. Role of inbreeding depression and purging in captive breeding and restoration programmes. Mol. Ecol. 2008, 17, 334–343. [Google Scholar] [CrossRef] [PubMed]
- Österling, M.E.; Larsen, B.M. Impact of origin and condition of host fish (Salmo trutta) on parasitic larvae of Margaritifera margaritifera. Aquat. Conserv. Mar. Freshw. Ecosyst. 2013, 23, 564–570. [Google Scholar] [CrossRef]
- Taeubert, J.E.; Denic, M.; Gum, B.; Lange, M.; Geist, J. Suitability of different salmonid strains as hosts for the endangered freshwater pearl mussel (Margaritifera margaritifera L.). Aquat. Conserv. Mar. Freshw. Ecosyst. 2010, 20, 728–734. [Google Scholar] [CrossRef]
- Söderberg, H. Undersökningtyp: Övervakning av flodpärlmussla. In Flodpärlmusslan i Sverige; Eriksson, M.O.G., Henrikson, L., Söderberg, H., Eds.; Naturvårdsverket: Stockholm, Sweden, 1998. [Google Scholar]

| Stream | Abbreviation | Alt. | pH | Conductivity | Color | Ca | L mm | Status |
|---|---|---|---|---|---|---|---|---|
| m a.s.l. | mS m−1 | mg Pt L−1 | mg L−1 | Min–Max | Score 1 | |||
| Lomsdalselva | A-LOM | 170 | 5.5 2–6.4 | 1.6 | 70 | 1.7 | 79–131 | 8 |
| Etna | A-ETN | 143 | 6.8 | 3.1 | 20 | 5.6 | 44–125 | 6 |
| Fallselva | A-FAL | 304 | 6.5 | 2.4 | 52 | 3.5 | 55–115 | 9 |
| Sørkedalselva | B-SØR | 150 | 7.0 | 2.8 | 43 | 5.6 | 84–116 | 22 |
| Hunnselva | C-HUN | 300 | 7.6 | 11.5 | 36 | 16 | 80–115 | 7 |
| Leira | C-LEI | 240 | 5.3 2–6.4 | 1.7 | 22 | 1.9 | 63–101 | 18 |
| Kampåa | C-KAM | 185 | 5.3 2–6.8 | 2.7 | 68 | 3.2 | 62–107 | 15 |
| Gjerda | C-GJE | 267 | 6.0 3–6.1 | 2.8 | 68 | 1.8 | 78–101 | 6 |
| Bråtåa | D-BRÅ | 160 | 5.7 3–6.4 | 3.5 | 72 | 3.3 | 65–114 | 8 |
| Finnsrudåa | D-FIN | 150 | 6.1 | 4.1 | 40 | 4.2 | 61–118 | 20 |
| Løvhaugsåa | D-LØV | 293 | 5.6 2–6.0 | 2.1 | 78 | 1.4 | 86–116 | 6 |
| A-LOM | A-ETN | A-FAL | B-SOR | C-HUN | C-LEI | C-KAM | C-GJE | D-BRA | D-FIN | D-LOV | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| A-LOM | - | 0.149 | 0.638 | 0.181 | 0.198 | 0.136 | 0.135 | 0.206 | 0.125 | 0.195 | 0.189 |
| A-ETN | 0.099 | - | 0.624 | 0.170 | 0.222 | 0.139 | 0.144 | 0.191 | 0.161 | 0.215 | 0.225 |
| A-FAL | 0.199 | 0.209 | - | 0.770 | 0.819 | 0.891 | 0.889 | 0.673 | 0.816 | 0.834 | 0.788 |
| B-SOR | 0.120 | 0.113 | 0.266 | - | 0.178 | 0.109 | 0.094 | 0.189 | 0.142 | 0.168 | 0.153 |
| C-HUN | 0.172 | 0.188 | 0.370 | 0.155 | - | 0.077 | 0.080 | 0.104 | 0.098 | 0.038 | 0.163 |
| C-LEI | 0.154 | 0.157 | 0.391 | 0.137 | 0.134 | - | 0.001 | 0.179 | 0.090 | 0.065 | 0.068 |
| C-KAM | 0.155 | 0.159 | 0.390 | 0.115 | 0.149 | 0.013 | - | 0.168 | 0.084 | 0.068 | 0.063 |
| C-GJE | 0.157 | 0.153 | 0.307 | 0.173 | 0.170 | 0.317 | 0.247 | - | 0.099 | 0.164 | 0.208 |
| D-BRA | 0.121 | 0.142 | 0.332 | 0.148 | 0.198 | 0.256 | 0.197 | 0.160 | - | 0.078 | 0.123 |
| D-FIN | 0.162 | 0.175 | 0.346 | 0.136 | 0.053 | 0.094 | 0.101 | 0.172 | 0.127 | - | 0.122 |
| D-LOV | 0.170 | 0.183 | 0.331 | 0.148 | 0.267 | 0.150 | 0.139 | 0.285 | 0.239 | 0.166 | - |
| Population | n | AL | AP | PL | FIS | HE |
|---|---|---|---|---|---|---|
| A-LOM | 30 | 3.6 | 11 | 11 | 0.122 * | 0.405 |
| A-ETN | 8 | 2.8 | 1 | 11 | 0.261 * | 0.404 |
| A-FAL | 30 | 4.1 | 9 | 11 | 0.143 * | 0.586 |
| B-SØR | 24 | 3. | 5 | 10 | 0.110 | 0.392 |
| C-HUN | 41 | 1.4 | 1 | 4 | 0.579 * | 0.176 |
| C-LEI | 32 | 1.4 | 0 | 4 | 0.847 * | 0.099 |
| C-KAM | 28 | 1.5 | 3 | 4 | 0.448 * | 0.120 |
| C-GJE | 29 | 1.8 | 2 | 4 | 0.801 * | 0.385 |
| D-BRÅ | 22 | 1.5 | 0 | 4 | 0.588 * | 0.422 |
| D-FIN | 30 | 2.1 | 3 | 7 | 0.575 * | 0.195 |
| D-LØV | 29 | 1.6 | 2 | 4 | 0.699 * | 0.420 |
| Population | FullSib | % FullSib | Max. Family | M | |
|---|---|---|---|---|---|
| Group Size | Mean | Min–Max | |||
| A-LOM | 23 | 5.3 | 6 | 0.23 | 0.04–0.80 |
| A-ETN | 0 | 0 | 0 | 0.16 | 0.04–0.50 |
| A-FAL | 2 | 3.5 | 3 | 0.20 | 0.03–0.70 |
| B-SØR | 12 | 4.4 | 5 | 0.18 | 0.02–0.60 |
| C-HUN | 233 | 28.4 | 22 | 0.07 | 0.02–0.25 |
| C-LEI | 416 | 83.8 | 29 | 0.07 | 0.02–0.25 |
| C-KAM | 220 | 58.2 | 28 | 0.09 | 0.02–0.40 |
| C-GJE | 23 | 15.3 | 11 | 0.11 | 0.02–0.40 |
| D-LØV | 17 | 4.2 | 4 | 0.08 | 0.02–0.30 |
| D-BRÅ | 35 | 15.2 | 6 | 0.10 | 0.02–0.25 |
| D-FIN | 221 | 50.8 | 27 | 0.08 | 0.02–0.25 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Linløkken, A.N.; Garlie, S.; Johansen, W.; Wilson, R.C. Assessing Evolutionary Significant Units (ESU) of the Endangered Freshwater Pearl Mussel (Margaritifera margaritifera) in Southeast Norway on the Basis of Genetic Analysis. Genes 2020, 11, 1061. https://doi.org/10.3390/genes11091061
Linløkken AN, Garlie S, Johansen W, Wilson RC. Assessing Evolutionary Significant Units (ESU) of the Endangered Freshwater Pearl Mussel (Margaritifera margaritifera) in Southeast Norway on the Basis of Genetic Analysis. Genes. 2020; 11(9):1061. https://doi.org/10.3390/genes11091061
Chicago/Turabian StyleLinløkken, Arne N., Silje Garlie, Wenche Johansen, and Robert C. Wilson. 2020. "Assessing Evolutionary Significant Units (ESU) of the Endangered Freshwater Pearl Mussel (Margaritifera margaritifera) in Southeast Norway on the Basis of Genetic Analysis" Genes 11, no. 9: 1061. https://doi.org/10.3390/genes11091061
APA StyleLinløkken, A. N., Garlie, S., Johansen, W., & Wilson, R. C. (2020). Assessing Evolutionary Significant Units (ESU) of the Endangered Freshwater Pearl Mussel (Margaritifera margaritifera) in Southeast Norway on the Basis of Genetic Analysis. Genes, 11(9), 1061. https://doi.org/10.3390/genes11091061





