Bioinformatic and Molecular Analysis of Satellite Repeat Diversity in Vaccinium Genomes
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and DNA Extraction
2.2. Bioinformatic Identification of Satellite DNA Families
2.3. Satellite-Specific PCR Amplification in the Vaccinium Genotypes
2.4. Cloning, Sequencing, and Raw Sequence Analysis of Satellite Repeats
2.5. Analysis of Structural Variability and Diversity of Satellite Repeats
2.6. Analysis of Transposable Elements Similar to Satellite Repeats
2.7. Analysis of the Distribution of Satellite Repeats Monomer along the Pseudochromosomes of the Tetraploid V. corymbosum
3. Results
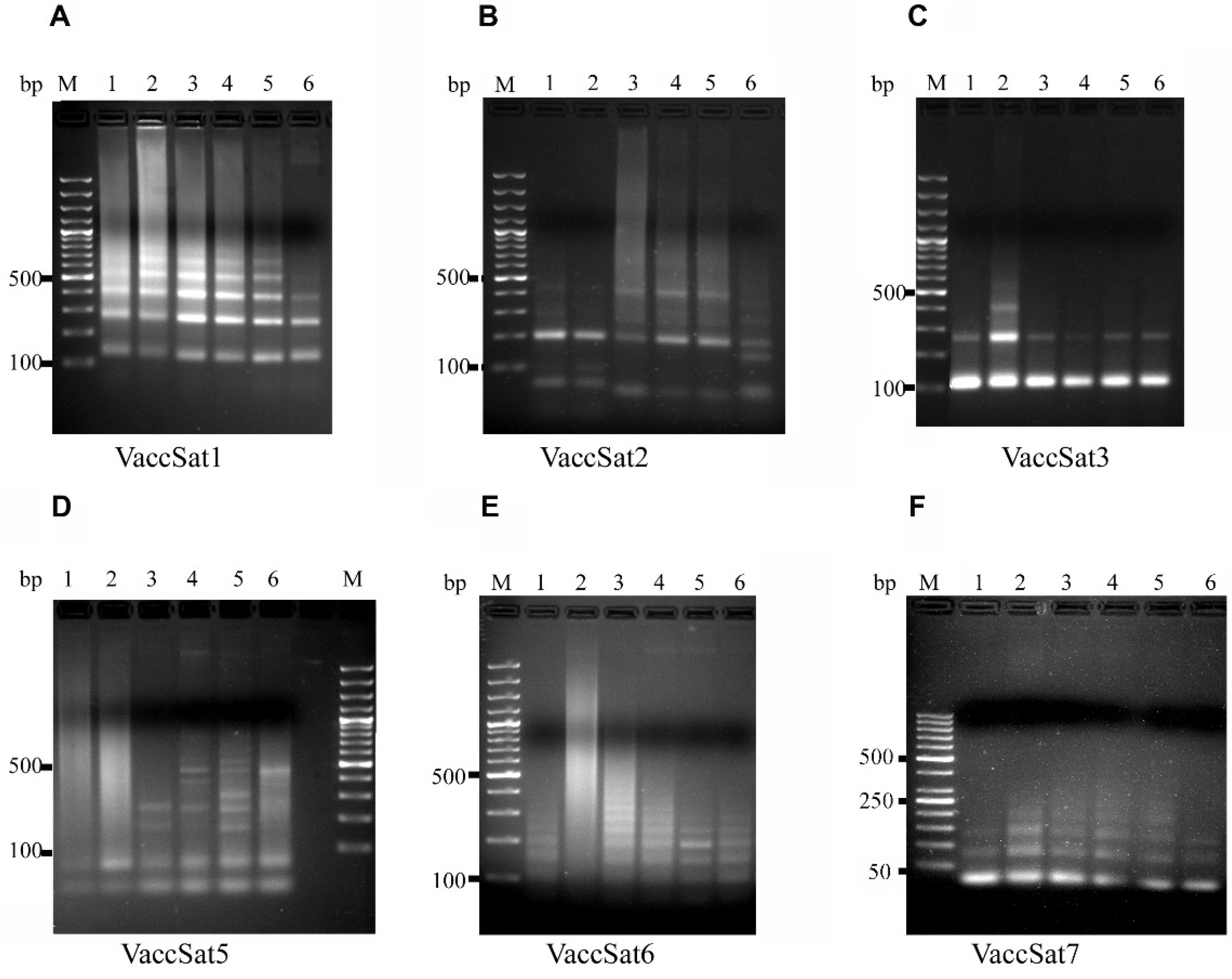
3.1. Analysis of VaccSat Satellite Repeats by PCR
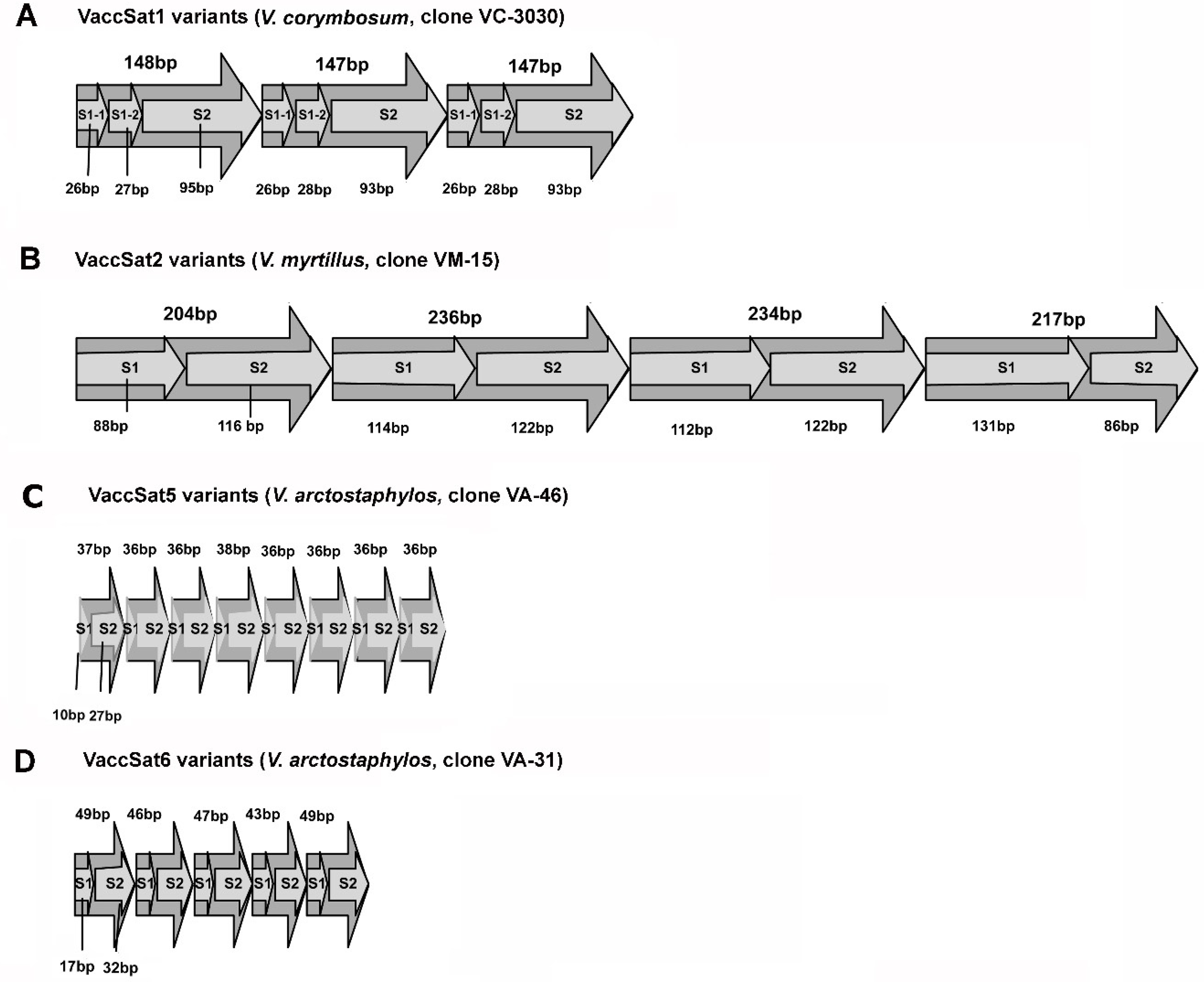
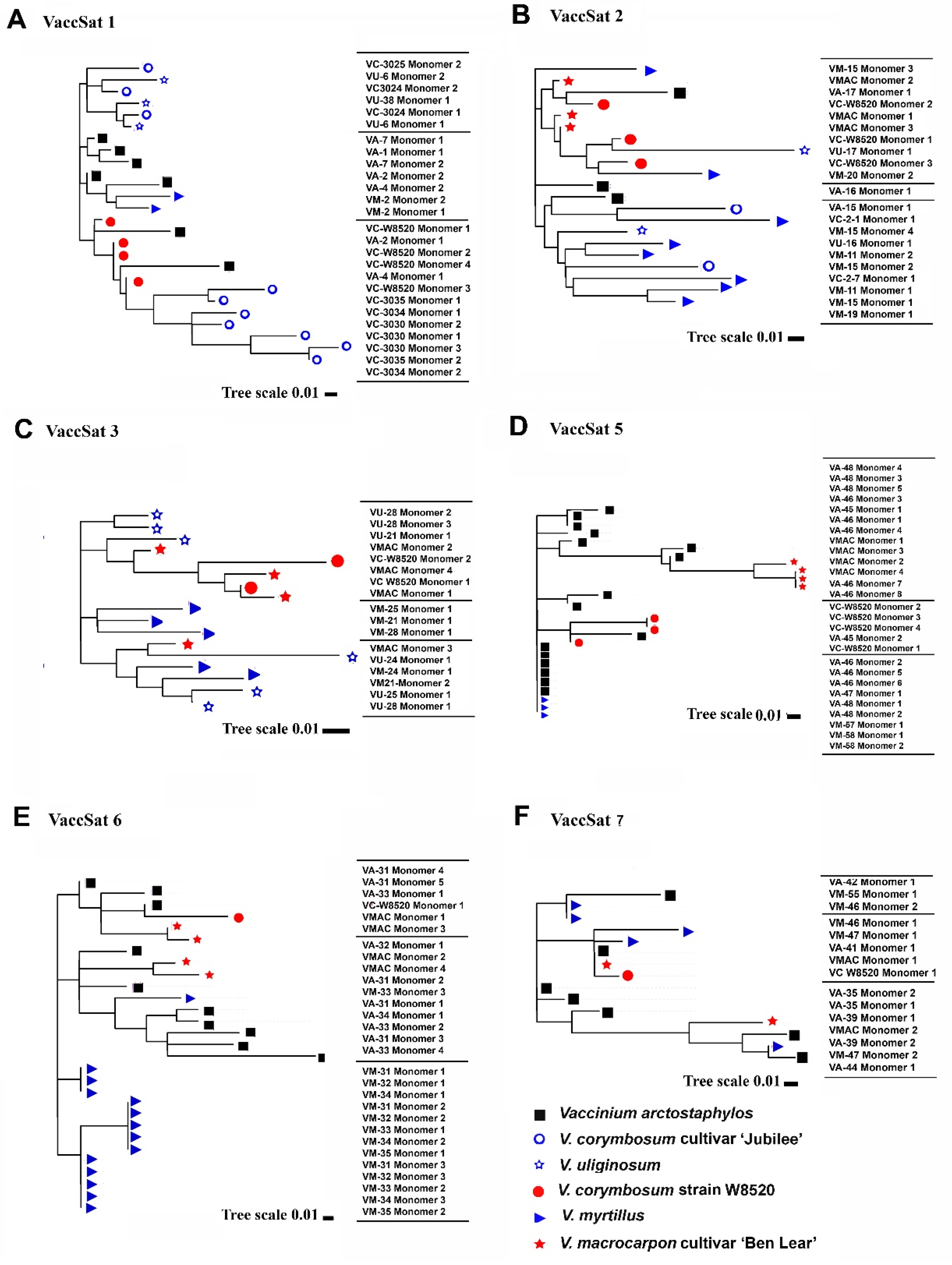
3.2. Satellite repeat Diversity and Organization across Different Vaccinium Species
3.2.1. VaccSat1
3.2.2. VaccSat2
3.2.3. VaccSat3
3.2.4. VaccSat5
3.2.5. VaccSat6
3.2.6. VaccSat7
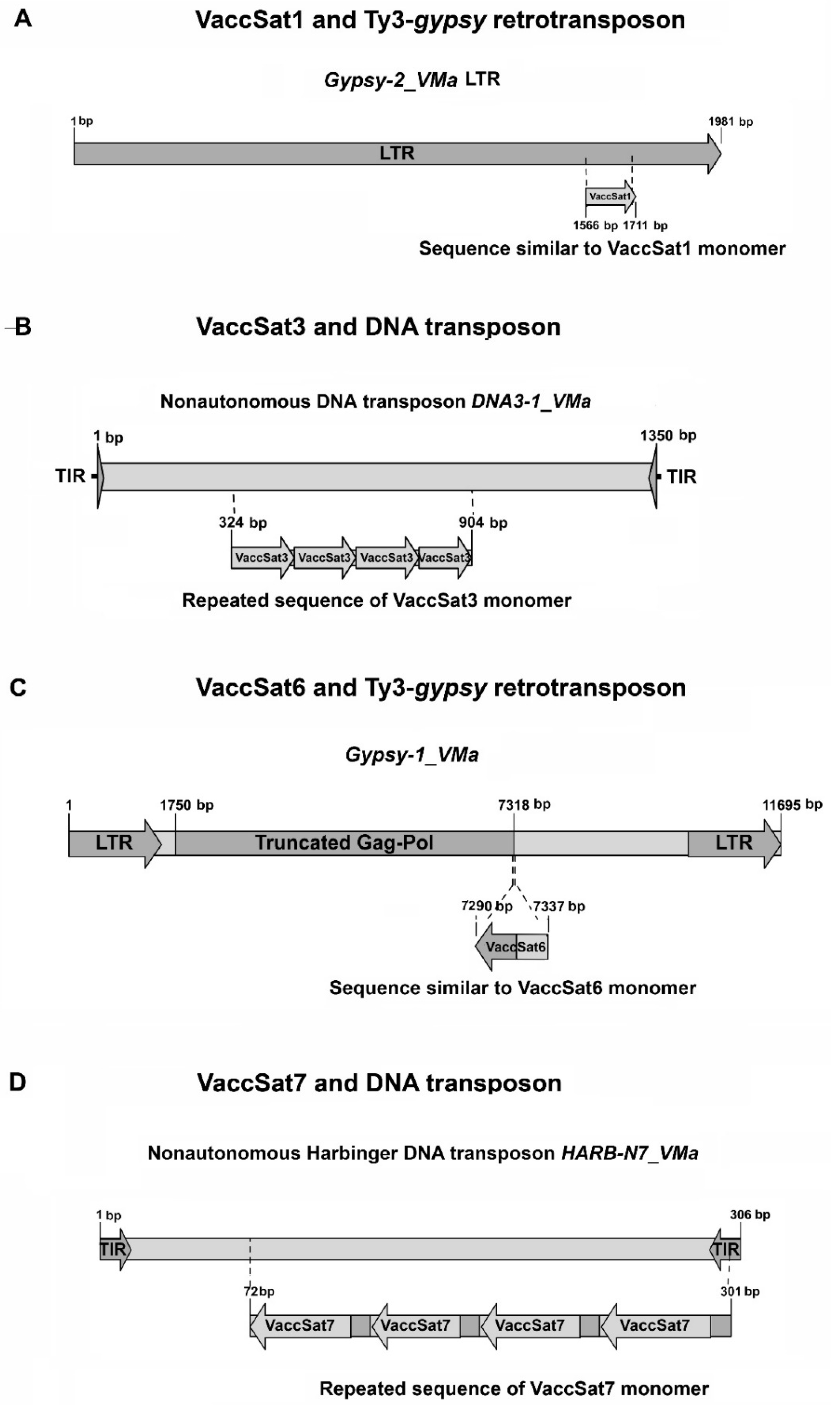
3.3. Similarities and Associations of Satellite Repeats with TEs
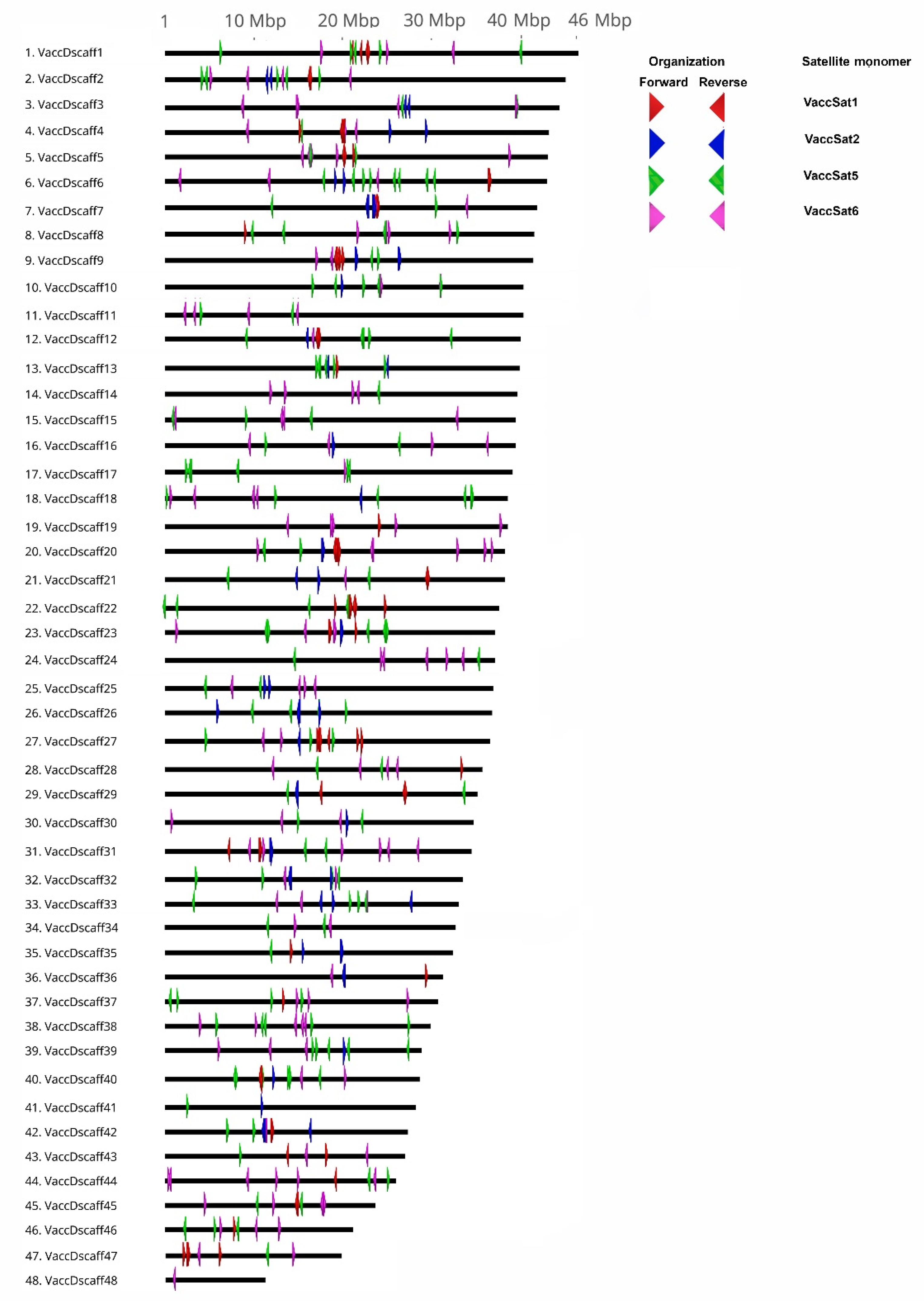
3.4. Pattern of Distribution and Number of Monomers of Each Satellite Repeat on the Pseudochromosomes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schmidt, T.; Heslop-Harrison, J.S. Genomes, genes and junk: The large-scale organization of plant chromosomes. Trends Plant Sci. 1998, 3, 195–199. [Google Scholar] [CrossRef]
- Heslop-Harrison, J.S.P.; Schwarzacher, T. Organisation of the plant genome in chromosomes. Plant J. 2011, 66, 18–33. [Google Scholar] [CrossRef] [PubMed]
- Kelly, L.J.; Renny-Byfield, S.; Pellicer, J.; Macas, J.; Novák, P.; Neumann, P.; Lysak, M.A.; Day, P.D.; Berger, M.; Fay, M.F.; et al. Analysis of the giant genomes of Fritillaria (Liliaceae) indicates that a lack of DNA removal characterizes extreme expansions in genome size. New Phytol. 2015, 208, 596–607. [Google Scholar] [CrossRef] [PubMed]
- Heitkam, T.; Petrasch, S.; Zakrzewski, F.; Kögler, A.; Wenke, T.; Wanke, S.; Schmidt, T. Next-generation sequencing reveals differentially amplified tandem repeats as a major genome component of Northern Europe’s oldest Camellia japonica. Chromosome Res. 2015, 23, 791–806. [Google Scholar] [CrossRef]
- Kirov, I.V.; Kiseleva, A.V.; Van Laere, K.; Van Roy, N.; Khrustaleva, L.I. Tandem repeats of Allium fistulosum associated with major chromosomal landmarks. Mol. Genet. Genom. 2017, 292, 453–464. [Google Scholar] [CrossRef]
- Garrido-Ramos, M. Satellite DNA: An evolving topic. Genes 2017, 8, 230. [Google Scholar] [CrossRef]
- Garrido-Ramos, M.A. Satellite DNA in plants: More than just rubbish. Cytogenet. Genome Res. 2015, 146, 153–170. [Google Scholar] [CrossRef]
- Ávila Robledillo, L.; Koblížková, A.; Novák, P.; Böttinger, K.; Vrbová, I.; Neumann, P.; Schubert, I.; Macas, J. Satellite DNA in Vicia faba is characterized by remarkable diversity in its sequence composition, association with centromeres, and replication timing. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Iwata, A.; Tek, A.L.; Richard, M.M.S.; Abernathy, B.; Fonsêca, A.; Schmutz, J.; Chen, N.W.G.; Thareau, V.; Magdelenat, G.; Li, Y.; et al. Identification and characterization of functional centromeres of the common bean. Plant J. 2013, 76, 47–60. [Google Scholar] [CrossRef]
- Zhang, H.; Koblížková, A.; Wang, K.; Gong, Z.; Oliveira, L.; Torres, G.A.; Wu, Y.; Zhang, W.; Novák, P.; Buell, C.R.; et al. Boom-bust turnovers of megabase-sized centromeric DNA in Solanum species: Rapid evolution of DNA sequences associated with centromeres. Plant Cell 2014, 26, 1436–1447. [Google Scholar] [CrossRef]
- He, Q.; Cai, Z.; Hu, T.; Liu, H.; Bao, C.; Mao, W.; Jin, W. Repetitive sequence analysis and karyotyping reveals centromere-associated DNA sequences in radish (Raphanus sativus L.). BMC Plant Biol. 2015, 15, 105. [Google Scholar] [CrossRef]
- Jagannathan, M.; Cummings, R.; Yamashita, Y.M. A conserved function for pericentromeric satellite DNA. Elife 2018, 7, e34122. [Google Scholar] [CrossRef] [PubMed]
- Mizuno, H.; Wu, J.; Kanamori, H.; Fujisawa, M.; Namiki, N.; Saji, S.; Katagiri, S.; Katayose, Y.; Sasaki, T.; Matsumoto, T. Sequencing and characterization of telomere and subtelomere regions on rice chromosomes 1S, 2S, 2L, 6L, 7S, 7L and 8S. Plant J. 2006, 46, 206–217. [Google Scholar] [CrossRef] [PubMed]
- Churikov, D.; Price, C.M. Telomeric and Subtelomeric Repeat Sequences. In Encyclopedia of Life Sciences; John Wiley & Sons: Hoboken, NJ, USA, 2008; ISBN 0470016175. [Google Scholar]
- Torres, G.A.; Gong, Z.; Iovene, M.; Hirsch, C.D.; Buell, C.R.; Bryan, G.J.; Novák, P.; Macas, J.; Jiang, J. Organization and evolution of subtelomeric satellite repeats in the potato genome. G3 Genes Genomes Genetics 2011, 1, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, J.A.; von Sternberg, R. Why repetitive DNA is essential to genome function. Biol. Rev. 2005, 80, 227–250. [Google Scholar] [CrossRef] [PubMed]
- Ugarkovic, D. Functional elements residing within satellite DNAs. EMBO Rep. 2005, 6, 1035–1039. [Google Scholar] [CrossRef]
- Steflova, P.; Tokan, V.; Vogel, I.; Lexa, M.; Macas, J.; Novak, P.; Hobza, R.; Vyskot, B.; Kejnovsky, E. Contrasting patterns of transposable element and satellite distribution on sex chromosomes (XY1Y2) in the dioecious plant Rumex acetosa. Genome Biol. Evol. 2013, 5, 769–782. [Google Scholar] [CrossRef]
- Puterova, J.; Razumova, O.; Martinek, T.; Alexandrov, O.; Divashuk, M.; Kubat, Z.; Hobza, R.; Karlov, G.; Kejnovsky, E. Satellite DNA and transposable elements in seabuckthorn (Hippophae rhamnoides), a dioecious plant with small Y and large X chromosomes. Genome Biol. Evol. 2017, 9, 197–212. [Google Scholar] [CrossRef]
- Neumann, P.; Navrátilová, A.; Koblížková, A.; Kejnovský, E.; Hřibová, E.; Hobza, R.; Widmer, A.; Doležel, J.; Macas, J. Plant centromeric retrotransposons: A structural and cytogenetic perspective. Mob. DNA 2011, 2, 4. [Google Scholar] [CrossRef]
- Wendel, J.F.; Jackson, S.A.; Meyers, B.C.; Wing, R.A. Evolution of plant genome architecture. Genome Biol. 2016, 17, 37. [Google Scholar] [CrossRef]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory activities of transposable elements: From conflicts to benefits. Nat. Rev. Genet. 2016, 18, 71–86. [Google Scholar] [CrossRef] [PubMed]
- Pellicer, J.; Hidalgo, O.; Dodsworth, S.; Leitch, I.J. Genome size diversity and its impact on the evolution of land plants. Genes 2018, 9, 88. [Google Scholar] [CrossRef] [PubMed]
- Meštrović, N.; Mravinac, B.; Pavlek, M.; Vojvoda-Zeljko, T.; Šatović, E.; Plohl, M. Structural and functional liaisons between transposable elements and satellite DNAs. Chromosome Res. 2015, 23, 583–596. [Google Scholar] [CrossRef] [PubMed]
- Macas, J.; Koblížková, A.; Navrátilová, A.; Neumann, P. Hypervariable 3′ UTR region of plant LTR-retrotransposons as a source of novel satellite repeats. Gene 2009, 448, 198–206. [Google Scholar] [CrossRef]
- Stupar, R.M.; Song, J.; Tek, A.L.; Cheng, Z.; Dong, F.; Jiang, J. Highly condensed potato pericentromeric heterochromatin contains rDNA-related tandem repeats. Genetics 2002, 162, 1435–1444. [Google Scholar]
- Mehrotra, S.; Goyal, V. Repetitive sequences in plant nuclear DNA: Types, distribution, evolution and function. Genom. Proteom. Bioinform. 2014, 12, 164–171. [Google Scholar] [CrossRef]
- Del Bosque, M.E.Q.; López-Flores, I.; Suárez-Santiago, V.N.; Garrido-Ramos, M.A. Satellite-DNA diversification and the evolution of major lineages in Cardueae (Carduoideae Asteraceae). J. Plant Res. 2014, 127, 575–583. [Google Scholar] [CrossRef]
- Plohl, M.; Meštrovic, N.; Mravinac, B. Satellite DNA Evolution. In Repetitive DNA Genome Dynamic; Garrido-Ramos, M.A., Ed.; Karger: Basel, Switzerland, 2012; Volume 7, pp. 126–152. [Google Scholar]
- Yang, X.; Zhao, H.; Zhang, T.; Zeng, Z.; Zhang, P.; Zhu, B.; Han, Y.; Braz, G.T.; Casler, M.D.; Schmutz, J.; et al. Amplification and adaptation of centromeric repeats in polyploid switchgrass species. New Phytol. 2018, 218, 1645–1657. [Google Scholar] [CrossRef]
- Menzel, G.; Dechyeva, D.; Wenke, T.; Holtgräwe, D.; Weisshaar, B.; Schmidt, T. Diversity of a complex centromeric satellite and molecular characterization of dispersed sequence families in sugar beet (Beta vulgaris). Ann. Bot. 2008, 102, 521–530. [Google Scholar] [CrossRef]
- Cechova, M.; Harris, R.S.; Tomaszkiewicz, M.; Arbeithuber, B.; Chiaromonte, F.; Makova, K.D. High satellite repeat turnover in great apes studied with short- and long-read technologies. Mol. Biol. Evol. 2019, 36, 2415–2431. [Google Scholar] [CrossRef]
- Lunerová, J.; Herklotz, V.; Laudien, M.; Vozárová, R.; Groth, M.; Kovařík, A.; Ritz, C.M. Asymmetrical canina meiosis is accompanied by the expansion of a pericentromeric satellite in non-recombining univalent chromosomes in the genus Rosa. Ann. Bot. 2020. [Google Scholar] [CrossRef]
- Heitkam, T.; Weber, B.; Walter, I.; Liedtke, S.; Ost, C.; Schmidt, T. Satellite DNA landscapes after allotetraploidisation of quinoa (Chenopodium quinoa) reveal unique A and B subgenomes. Plant J. 2020. [Google Scholar] [CrossRef]
- Belyayev, A.; Josefiová, J.; Jandová, M.; Kalendar, R.; Krak, K.; Mandák, B. Natural history of a satellite DNA family: From the ancestral genome component to species-specific sequences, concerted and non-concerted evolution. Int. J. Mol. Sci. 2019, 20, 1201. [Google Scholar] [CrossRef] [PubMed]
- Trehane, J. Blueberries, Cranberries, and other Vacciniums. In Plant Collector Guide; Royal Horticultral Society: London, UK, 2004; Volume 19. [Google Scholar]
- Hancock, J.F.; Lyrene, P.; Finn, C.E.; Vorsa, N.; Lobos, G.A. Blueberries and Cranberries. In Temperate Fruit Crop Breeding: Germplasm to Genomics; Hancock, J.F., Ed.; Springer: Dordrecht, The Netherlands, 2008; pp. 115–150. ISBN 978-1-4020-6907-9. [Google Scholar]
- Vander Kloet, S.P. The genus Vaccinium in North America; Agriculture Canada: Ottawa, ON, Canada, 1988. [Google Scholar]
- Ehlenfeldt, M.K.; Ballington, J.R. Vaccinium species of section Hemimyrtillus: Their value to cultivated blueberry and approaches to utilization. Botany 2012, 90, 243–252. [Google Scholar] [CrossRef]
- Vander Kloet, S.P.; Dickinson, T.A. A subgeneric classification of the genus Vaccinium and the metamorphosis of V. section Bracteata Nakai: More terrestrial and less epiphytic in habit, more continental and less insular in distribution. J. Plant Res. 2009, 122, 253–268. [Google Scholar] [CrossRef] [PubMed]
- Mudd, A.B.; White, E.J.; Bolloskis, M.P.; Kapur, N.P.; Everhart, K.W.; Lin, Y.C.; Bussler, W.W.; Reid, R.W.; Brown, R.H. Students’ perspective on genomics: From sample to sequence using the case study of blueberry. Front. Genet. 2013, 4, 245. [Google Scholar] [CrossRef]
- Ballington, J.R. Collection, utilization, and preservation of genetic resources in Vaccinium. HortScience 2001, 36, 213–220. [Google Scholar] [CrossRef]
- Vicient, C.M.; Casacuberta, J.M. Impact of transposable elements on polyploid plant genomes. Ann. Bot. 2017, 120, 195–207. [Google Scholar] [CrossRef]
- Powell, E.A.; Kron, K.A. Hawaiian blueberries and their relatives—A phylogenetic analysis of Vaccinium sections Macropelma, Myrtillus, and Hemimyrtillus (Ericaceae). Syst. Bot. 2002, 27, 768–779. [Google Scholar]
- Rowland, L.J.; Bell, D.J.; Alkharouf, N.; Bassil, N.V.; Drummond, F.A.; Beers, L.; Buck, E.J.; Finn, C.E.; Graham, J.; McCallum, S.; et al. Generating genomic tools for blueberry improvement. Int. J. Fruit Sci. 2012, 12, 276–287. [Google Scholar] [CrossRef]
- Zdepski, A.; Debnath, S.C.; Howell, A.; Polashock, J.; Oudemans, P.; Vorsa, N.; Michael, T.P. Cranberry. In Genetics, Genomics and Breeding of Berries; Folta, K., Kole, C., Eds.; CRC Press: Boca Raton, FL, USA, 2011; p. 200. ISBN 9781578087075. [Google Scholar]
- Costich, D.E.; Ortiz, R.; Meagher, T.R.; Bruederle, L.P.; Vorsa, N. Determination of ploidy level and nuclear DNA content in blueberry by flow cytometry. Theor. Appl. Genet. 1993, 86, 1001–1006. [Google Scholar] [CrossRef] [PubMed]
- Colle, M.; Leisner, C.P.; Wai, C.M.; Ou, S.; Bird, K.A.; Wang, J.; Wisecaver, J.H.; Yocca, A.E.; Alger, E.I.; Tang, H.; et al. Haplotype-phased genome and evolution of phytonutrient pathways of tetraploid blueberry. Gigascience 2019, 8, giz012. [Google Scholar] [CrossRef] [PubMed]
- Hummer, K.E.; Bassil, N.V.; Rodríquez Armenta, H.P.; Olmstead, J.W. Vaccinium species ploidy assessment. Acta Hortic. 2015, IV 1101, 199–204. [Google Scholar] [CrossRef]
- Sultana, N.; Pascual-Díaz, J.P.; Gers, A.; Ilga, K.; Serçe, S.; Vitales, D.; Garcia, S. Contribution to the knowledge of genome size evolution in edible blueberries (genus Vaccinium). J. Berry Res. 2019, 1–15. [Google Scholar] [CrossRef]
- Sultana, N.; Menzel, G.; Heitkam, T.; Schmidt, T.; Serce, S. Comparative analysis of repetitive sequences reveals genome differences between two common cultivated Vaccinium Species (V. corymbosum and V. macrocarpon). J. Mol. Biol. Biotech. 2017, 1, 7–15. [Google Scholar]
- Fajardo, D.; Senalik, D.; Ames, M.; Zhu, H.; Steffan, S.A.; Harbut, R.; Polashock, J.; Vorsa, N.; Gillespie, E.; Kron, K.; et al. Complete plastid genome sequence of Vaccinium macrocarpon: Structure, gene content, and rearrangements revealed by next generation sequencing. Tree Genet. Genomes 2012, 9, 489–498. [Google Scholar] [CrossRef]
- Polashock, J.; Zelzion, E.; Fajardo, D.; Zalapa, J.; Georgi, L.; Bhattacharya, D.; Vorsa, N. The American cranberry: First insights into the whole genome of a species adapted to bog habitat. BMC Plant Biol. 2014, 14, 165. [Google Scholar] [CrossRef]
- Gupta, V.; Estrada, A.D.; Blakley, I.; Reid, R.; Patel, K.; Meyer, M.D.; Andersen, S.U.; Brown, A.F.; Lila, M.A.; Loraine, A.E. RNA-Seq analysis and annotation of a draft blueberry genome assembly identifies candidate genes involved in fruit ripening, biosynthesis of bioactive compounds, and stage-specific alternative splicing. Gigascience 2015, 4, 5. [Google Scholar] [CrossRef]
- Bian, Y.; Ballington, J.; Raja, A.; Brouwer, C.; Reid, R.; Burke, M.; Wang, X.; Rowland, L.J.; Bassil, N.; Brown, A. Patterns of simple sequence repeats in cultivated blueberries (Vaccinium section Cyanococcus spp.) and their use in revealing genetic diversity and population structure. Mol. Breed. 2014, 34, 675–689. [Google Scholar] [CrossRef]
- Li, L.; Zhang, H.; Liu, Z.; Cui, X.; Zhang, T.; Li, Y.; Zhang, L. Comparative transcriptome sequencing and de novo analysis of Vaccinium corymbosum during fruit and color development. BMC Plant Biol. 2016, 16, 223. [Google Scholar] [CrossRef]
- Novak, P.; Neumann, P.; Pech, J.; Steinhaisl, J.; Macas, J. RepeatExplorer: A Galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 2013, 29, 792–793. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 1999, 27, 573–580. [Google Scholar] [CrossRef]
- Rice, P.; Longden, I.; Bleasby, A. EMBOSS: The the European molecular biology open software suite. Trends Genet. 2000, 16, 276–277. [Google Scholar] [CrossRef]
- Seibt, K.M.; Schmidt, T.; Heitkam, T. FlexiDot: Highly customizable, ambiguity-aware dotplots for visual sequence analyses. Bioinformatics 2018, 34, 3575–3577. [Google Scholar] [CrossRef]
- Kohany, O.; Gentles, A.J.; Hankus, L.; Jurka, J. Annotation, submission and screening of repetitive elements in Repbase: Repbase Submitter and Censor. BMC Bioinform. 2006, 25, 474. [Google Scholar]
- Blastclust. Available online: Ftp://ftp.ncbi.nih.gov/blast/documents/blastclust.html/ (accessed on 15 August 2013).
- Kapitonov, V.V.; Jurka, J. A universal classification of eukaryotic transposable elements implemented in Repbase. Nat. Rev. Gen. 2008, 9, 411. [Google Scholar] [CrossRef]
- Bao, W.; Kojima, K.K.; Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 2015, 6, 11. [Google Scholar] [CrossRef]
- Begum, R.; Alam, S.S.; Menzel, G.; Schmidt, T. Comparative molecular cytogenetics of major repetitive sequence families of three Dendrobium species (Orchidaceae) from Bangladesh. Ann. Bot. 2009, 104, 863–872. [Google Scholar] [CrossRef] [PubMed]
- Macas, J.; Novák, P.; Pellicer, J.; Čížková, J.; Koblížková, A.; Neumann, P.; Fuková, I.; Doležel, J.; Kelly, L.J.; Leitch, I.J. In depth characterization of repetitive DNA in 23 plant genomes reveals sources of genome size variation in the legume tribe Fabeae. PLoS ONE 2015, 10, e0143424. [Google Scholar] [CrossRef] [PubMed]
- Bolsheva, N.L.; Melnikova, N.V.; Kirov, I.V.; Dmitriev, A.A.; Krasnov, G.S.; Amosova, A.V.; Samatadze, T.E.; Yurkevich, O.Y.; Zoshchuk, S.A.; Kudryavtseva, A.V.; et al. Characterization of repeated DNA sequences in genomes of blue-flowered flax. BMC Evol. Biol. 2019, 19, 49. [Google Scholar] [CrossRef] [PubMed]
- Tek, A.L.; Song, J.; Macas, J.; Jiang, J. Sobo, a recently amplified satellite repeat of potato, and its implications for the origin of tandemly repeated sequences. Genetics 2005, 170, 1231–1238. [Google Scholar] [CrossRef]
- Zakrzewski, F.; Weber, B.; Schmidt, T. A Molecular Cytogenetic Analysis of the Structure, Evolution, and Epigenetic Modifications of Major DNA Sequences in Centromeres of Beta Species. In Plant Centromere Biology; Jiang, J., Birchler, J.A., Eds.; Wiley-Blackwell: Hoboken, NJ, USA, 2013; pp. 39–55. [Google Scholar]
- Macas, J.; Navrátilová, A.; Koblízková, A. Sequence homogenization and chromosomal localization of VicTR-B satellites differ between closely related Vicia species. Chromosoma 2006, 115, 437–447. [Google Scholar] [CrossRef]
- Sharma, S.; Raina, S.N. Organization and evolution of highly repeated satellite DNA sequences in plant chromosomes. Cytogenet. Genome Res. 2005, 109, 15–26. [Google Scholar] [CrossRef]
- Melters, D.P.; Bradnam, K.R.; Young, H.A.; Telis, N.; May, M.R.; Ruby, J.; Sebra, R.; Peluso, P.; Eid, J.; Rank, D.; et al. Comparative analysis of tandem repeats from hundreds of species reveals unique insights into centromere evolution. Genome Biol. 2013, 14, R10. [Google Scholar] [CrossRef]
- Dover, G. Molecular drive: A cohesive mode of species evolution. Nature 1982, 299, 111–117. [Google Scholar] [CrossRef]
- Macas, J.; Mészáros, T.; Nouzová, M. PlantSat: A specialized database for plant satellite repeats. Bioinformatics 2002, 18, 28–35. [Google Scholar] [CrossRef]
- Cohen, S.; Houben, A.; Segal, D. Extrachromosomal circular DNA derived from tandemly repeated genomic sequences in plants. Plant J. 2008, 53, 1027–1034. [Google Scholar] [CrossRef]
- Heslop-Harrison, J.S.P.; Schwarzacher, T. Nucleosomes and centromeric DNA packaging. Proc. Natl. Acad. Sci. USA 2013, 110, 19974–19975. [Google Scholar] [CrossRef]
- Qu, L.; Hancock, J.F.; Whallon, J.H. Evolution in an autopolyploid group displaying predominantly bivalent pairing at meiosis: Genomic similarity of diploid Vaccinium darrowi and autotetraploid V. corymbosum (Ericaceae). Am. J. Bot. 1998, 85, 698–703. [Google Scholar] [CrossRef]
- Jain, M.; Olsen, H.E.; Turner, D.J.; Stoddart, D.; Bulazel, K.V.; Paten, B.; Haussler, D.; Willard, H.F.; Akeson, M.; Miga, K.H. Linear assembly of a human centromere on the Y chromosome. Nat. Biotechnol. 2018, 36, 321. [Google Scholar] [CrossRef]
- Miga, K. Completing the human genome: The progress and challenge of satellite DNA assembly. Chromosome Res. 2015, 23, 421–426. [Google Scholar] [CrossRef]
- Peona, V.; Weissensteiner, M.H.; Suh, A. How complete are “complete” genome assemblies?—An avian perspective. Mol. Ecol. Resour. 2018, 18, 1188–1195. [Google Scholar] [CrossRef]
- Mlinarec, J.; Chester, M.; Siljak-Yakovlev, S.; Papes, D.; Leitch, A.R.; Besendorfer, V. Molecular structure and chromosome distribution of three repetitive DNA families in Anemone hortensis L. (Ranunculaceae). Chromosome Res. 2009, 17, 331–346. [Google Scholar] [CrossRef]
- Zakrzewski, F.; Wenke, T.; Weisshaar, B.; Schmidt, T.; Holtgräwe, D. Analysis of a c0t-1 library enables the targeted identification of minisatellite and satellite families in Beta vulgaris. BMC Plant Biol. 2010, 10, 8. [Google Scholar] [CrossRef]
- Schmidt, T.; Heitkam, T.; Liedtke, S.; Schubert, V.; Menzel, G. Adding color to a century-old enigma: Multi-color chromosome identification unravels the autotriploid nature of saffron (Crocus sativus) as a hybrid of wild Crocus cartwrightianus cytotypes. New Phytol. 2019, 222, 1965–1980. [Google Scholar] [CrossRef]
- Plohl, M. Those mysterious sequences of satellite DNAs. Period. Biol. 2010, 112, 403–410. [Google Scholar]
- Skaletsky, H.; Kuroda-Kawaguchi, T.; Minx, P.J.; Cordum, H.S.; Hillier, L.; Brown, L.G.; Repping, S.; Pyntikova, T.; Ali, J.; Bieri, T.; et al. The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 2003, 423, 825–837. [Google Scholar] [CrossRef]
- Mariotti, B.; Manzano, S.; Kejnovský, E.; Vyskot, B.; Jamilena, M. Accumulation of Y-specific satellite DNAs during the evolution of Rumex acetosa sex chromosomes. Mol. Genet. Genom. 2009, 281, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Richard, M.M.S.; Chen, N.W.G.; Thareau, V.; Pflieger, S.; Blanchet, S.; Pedrosa-Harand, A.; Iwata, A.; Chavarro, C.; Jackson, S.A.; Geffroy, V.T. The subtelomeric khipu satellite repeat from Phaseolus vulgaris: Lessons learned from the genome analysis of the Andean genotype G19833. Front. Plant Sci. 2013, 4, 109. [Google Scholar] [CrossRef] [PubMed]
- Koukalova, B.; Moraes, A.P.; Renny-Byfield, S.; Matyasek, R.; Leitch, A.R.; Kovarik, A. Fall and rise of satellite repeats in allopolyploids of Nicotiana over c. 5 million years. New Phytol. 2010, 186, 148–160. [Google Scholar] [CrossRef] [PubMed]
- Dover, G. Molecular drive. Trends Genet. 2002, 18, 587–589. [Google Scholar] [CrossRef]
- Fry, K.; Salser, W. Nucleotide sequences of HS-alpha satellite DNA from kangaroo rat Dipodomys ordii and characterization of similar sequences in other rodents. Cell 1977, 12, 1069–1084. [Google Scholar] [CrossRef]
- Kejnovsky, E.; Kubat, Z.; Macas, J.; Hobza, R.; Mracek, J.; Vyskot, B. Retand: A novel family of gypsy-like retrotransposons harboring an amplified tandem repeat. Mol. Genet. Genom. 2006, 276, 254–263. [Google Scholar] [CrossRef] [PubMed]
- Vondrak, T.; Ávila Robledillo, L.; Novák, P.; Koblížková, A.; Neumann, P.; Macas, J. Characterization of repeat arrays in ultra-long nanopore reads reveals frequent origin of satellite DNA from retrotransposon-derived tandem repeats. Plant J. 2020, 101, 484–500. [Google Scholar] [CrossRef]
- McGurk, M.P.; Barbash, D.A. Double insertion of transposable elements provides a substrate for the evolution of satellite DNA. Genome Res. 2018, 28, 714–725. [Google Scholar] [CrossRef]
- Dodsworth, S.; Chase, M.W.; Kelly, L.J.; Leitch, I.J.; Macas, J.; Novak, P.; Piednoel, M.; Weiss-Schneeweiss, H.; Leitch, A.R. Genomic repeat abundances contain phylogenetic signal. Syst. Biol. 2014, 64, 112–126. [Google Scholar] [CrossRef]
| Species | Section * | General Information of Specimens | Reference | ||||
|---|---|---|---|---|---|---|---|
| Place of Collection | Latitude | Longitude | Elevation [m above Sea Level] | Chromosome Number | |||
| Vaccinium corymbosum L. (cultivar ‘Jubilee’) | Cyanococcus | Turkey (NOHU) | 37.563200 | 34.372500 | 1,300 | 48 | Trehane, 2004 [36] |
| V. corymbosum L. (cultivar ‘Misty’) | Cyanococcus | Turkey (NOHU) | 37.940462 | 37.940462 | 1,300 | 48 | Trehane, 2004 [36] |
| Vaccinium corymbosum strain W8520 | Cyanococcus | - | - | - | - | 24 | Sultana et al., 2017 [51] |
| V. corymbosum L. (cultivar ‘Draper’) | Cyanococcus | - | - | - | - | 48 | Colle et al., 2019 [48] |
| Vaccinium arctostaphylos L. (round shaped fruit) | Hemimyrtillus | Turkey (KM) | 40.840298 | 41.102927 | 2641 | 48 | Sultana et al., 2019 [50] |
| V. arctostaphylos L. (elongated shaped fruit) | Hemimyrtillus | Turkey (KM) | 40.843791 | 41.081443 | 2779 | 48 | Sultana et al., 2019 [50] |
| Vaccinium myrtillus L. | Myrtillus | Turkey (KM) | 40.952462 | 41.101182 | 1251 | 24 | Sultana et al., 2019 [50] |
| Vaccinium uliginosum L. | Vaccinium | Turkey (KM) | 40.952461 | 41.101182 | 1251 | 24 | Sultana et al., 2019 [50] |
| Vaccinium macrocarpon cultivar ‘Ben Lear’ | Oxycococus | - | - | - | - | 24 | Sultana et al., 2017 [51] |
| Satellite Repeat | Forward Primer (5′–3′) | Reverse Primer (5′–3′) | Product Size (bp) | Annealing Temperature Used (°C) |
|---|---|---|---|---|
| VaccSat1/VaccSat4 | ATTTAAAATGATTTTGTCGC | GCAAATAATAATGGTATTTAGC | 119 | 48.9 |
| VaccSat2 | GTACGGGCTACTGACCAC | TATCGCTCAAACAACAAGTGG | 198 | 56.8 |
| VaccSat3 | ATTTGACATTGTTGGCTTGC | GATCTCAATTAGTAGTTTAATTTGGTG | 111 | 55.2 |
| VaccSat5 | ATTAAATCCATTTAAATCATTTTCTG | GATTTAAATGGATTTAATTAAAAATCC | 55 | 51.3 |
| VaccSat6 | CTGACGGATTTTAAAAACGATG | TCCGTCAGGTATTATTATGATTTTC | 54 | 54.4 |
| VaccSat7 | CAAGTTAGTTTTTTTGCAAAAC | GTTTTGCAAAAAAACTAACTTG | 49–70 | 51.9 |
| Satellite Repeat | Monomer Length (bp) | G/C Content (%) | Genome Proportion (%) a | Copy Number b |
|---|---|---|---|---|
| VaccSat1 | 146–147 | 18.4 | 0.999 | 31,439 |
| VaccSat2 | 238 | 40.9 | 0.124 | 2436 |
| VaccSat3 | 154 | 21.1 | 0.36 | 10,987 |
| VaccSat4 | 101 | 17.4 | 0.050 | 2326 |
| VaccSat5 | 36–38 | 19.9 | 0.015 | 1905 |
| VaccSat6 | 49 | 22.4 | 0.015 | 1438 |
| VaccSat7 | 49–70 | 31.4 | 0.069 | 4632 to 6618 |
| Satellite Family | Species | Number of Clones | Number of Monomers | Identity to Monomer Consensus [%] |
|---|---|---|---|---|
| VaccSat1/VaccSat4 | V. corymbosum | 8 | 16 | 78–92 |
| V. arctostaphylos | 4 | 11 | 68–92 | |
| V. myrtillus | 2 | 4 | 67–86 | |
| V. uliginosum | 7 | 10 | 83–97 | |
| VaccSat2 | V. corymbosum | 2 | 4 | 86–88 |
| V. arctostaphylos | 4 | 7 | 82–90 | |
| V. myrtillus | 4 | 11 | 83–90 | |
| V. uliginosum | 4 | 6 | 71–79 | |
| VaccSat3 | V. corymbosum | 3 | 3 | 88–90 |
| V. arctostaphylos | 4 | 4 | 84–90 | |
| V. myrtillus | 4 | 11 | 65–91 | |
| V. uliginosum | 4 | 10 | 65–89 | |
| VaccSat5 | V. corymbosum | 1 | 1 | - |
| V. arctostaphylos | 4 | 20 | 88–96 | |
| V. myrtillus | 2 | 5 | 96–97 | |
| V. uliginosum | - | - | - | |
| VaccSat6 | V. corymbosum | 1 | 1 | - |
| V. arctostaphylos | 4 | 14 | 89–90 | |
| V. myrtillus | 5 | 19 | 81–88 | |
| V. uliginosum | - | - | - | |
| VaccSat7 | V. corymbosum | - | - | - |
| V. arctostaphylos | 5 | 13 | 66–75 | |
| V. myrtillus | 4 | 10 | 69–78 | |
| V. uliginosum | - | - | - | |
| Total | 76 | 180 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sultana, N.; Menzel, G.; Heitkam, T.; Kojima, K.K.; Bao, W.; Serçe, S. Bioinformatic and Molecular Analysis of Satellite Repeat Diversity in Vaccinium Genomes. Genes 2020, 11, 527. https://doi.org/10.3390/genes11050527
Sultana N, Menzel G, Heitkam T, Kojima KK, Bao W, Serçe S. Bioinformatic and Molecular Analysis of Satellite Repeat Diversity in Vaccinium Genomes. Genes. 2020; 11(5):527. https://doi.org/10.3390/genes11050527
Chicago/Turabian StyleSultana, Nusrat, Gerhard Menzel, Tony Heitkam, Kenji K. Kojima, Weidong Bao, and Sedat Serçe. 2020. "Bioinformatic and Molecular Analysis of Satellite Repeat Diversity in Vaccinium Genomes" Genes 11, no. 5: 527. https://doi.org/10.3390/genes11050527
APA StyleSultana, N., Menzel, G., Heitkam, T., Kojima, K. K., Bao, W., & Serçe, S. (2020). Bioinformatic and Molecular Analysis of Satellite Repeat Diversity in Vaccinium Genomes. Genes, 11(5), 527. https://doi.org/10.3390/genes11050527





