Heritability of Low ER Staining/HER2-Breast Tumors: Are We Missing an Opportunity for Germline Testing?
Abstract
1. Introduction
2. Materials and Methods
3. Results
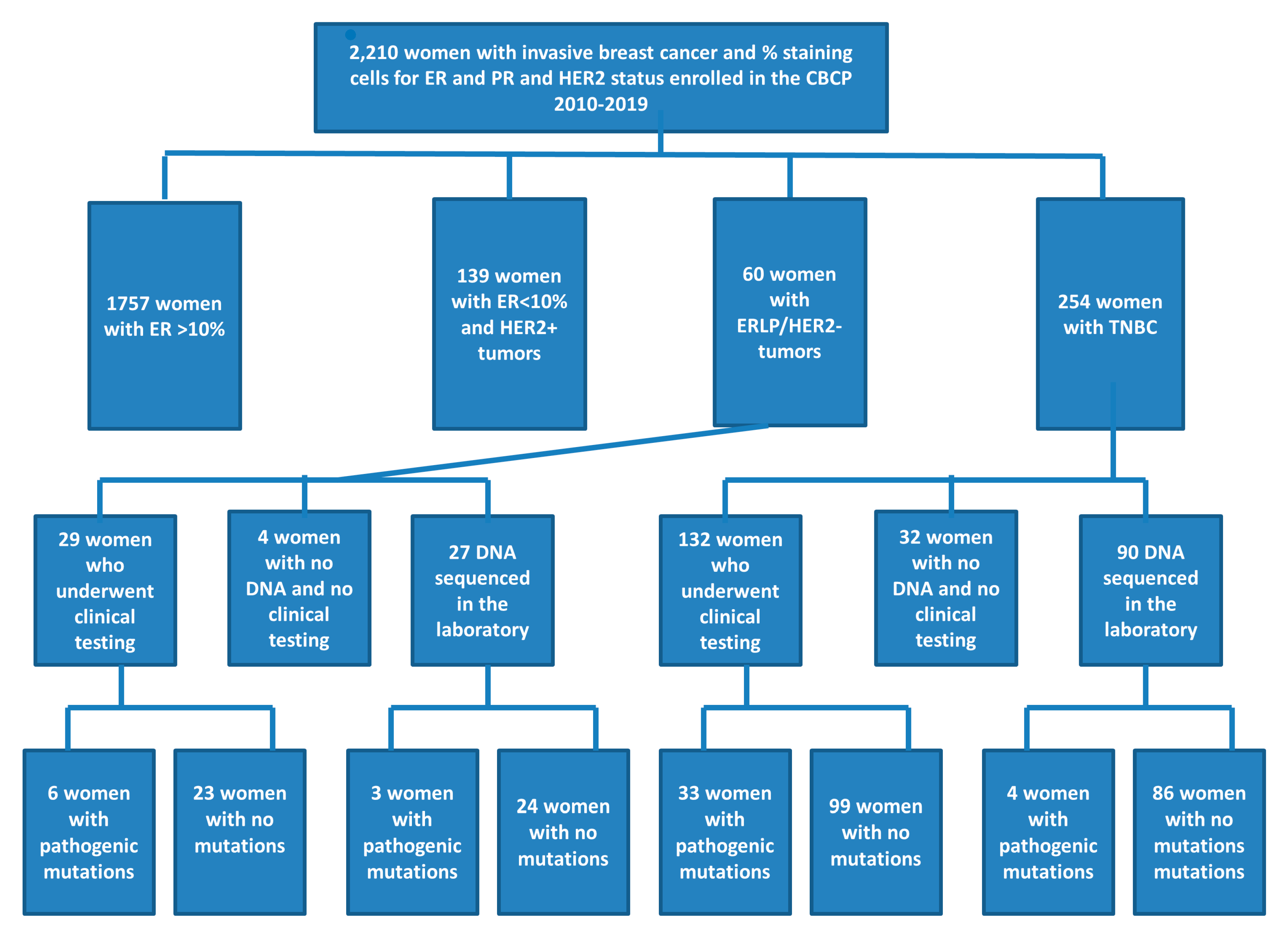
3.1. Patient Characteristics
3.2. Test Eligibility and Uptake
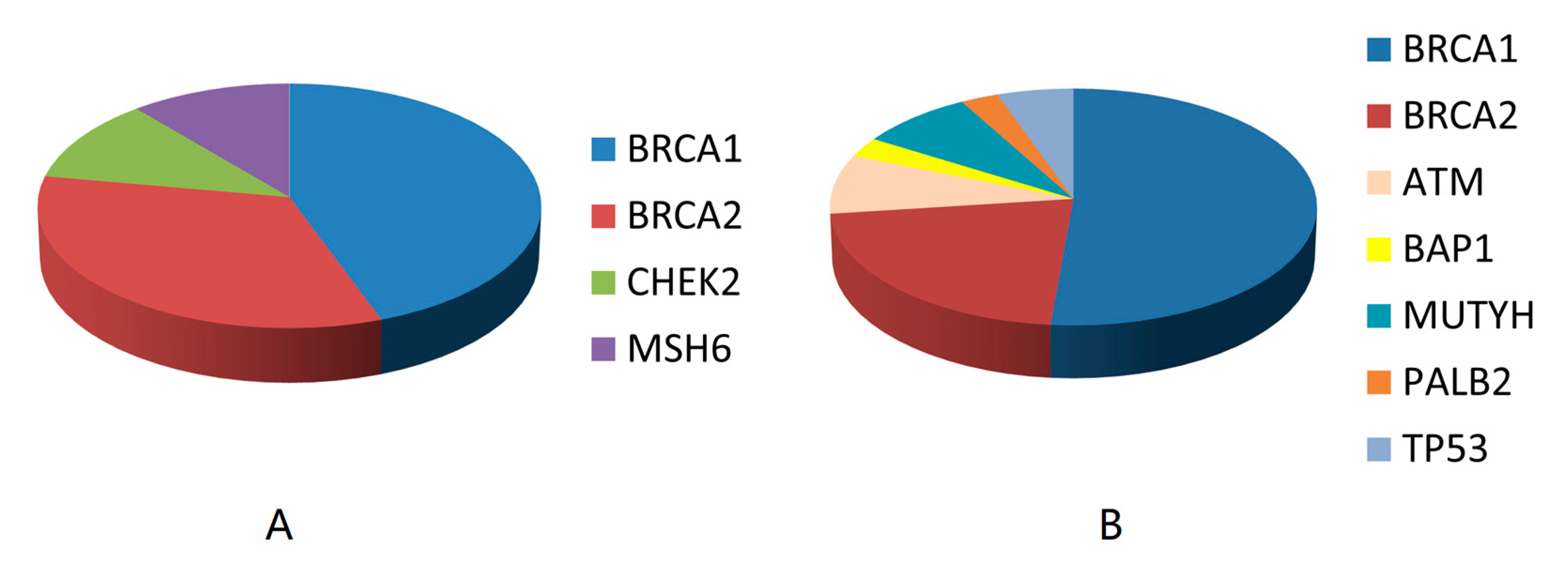
3.3. Genetic Results
3.4. Mutation Frequency in Women Not Eligible for Genetic Testing
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lin, N.U.; Claus, E.; Sohl, J.; Razzak, A.R.; Amaout, A.; Winer, E.P. Sites of distant recurrence and clinical outcomes in patients with metastatic triple-negative breast cancer: High incidence of central nervous system metastases. Cancer 2008, 113, 2638–2645. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Klemp, J.R.; Kimler, B.F.; Mahnken, J.D.; Geier, L.J.; Khan, Q.J.; Elia, M.; Connor, C.S.; McGinness, M.K.; Mammen, J.M.; et al. Germline BRCA mutation evaluation in a prospective triple-negative breast cancer registry: Implications for hereditary breast and/or ovarian cancer syndrome testing. Breast Cancer Res. Treat. 2014, 145, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Garutti, M.; Pelizzari, G.; Bartoletti, M.; Malfatti, M.C.; Gerratana, L.; Tell, G.; Puglisi, F. Platinum salts in patients with breast cancer: A focus on predictive factors. Int. J. Mol. Sci. 2019, 20, 3390. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, A.S.; Gillard, M.; Lipkowitz, S.; Lee, J.M. Update on PARP Inhibitors in breast cancer. Curr. Treat. Opt. Oncol. 2018, 19, 21. [Google Scholar] [CrossRef] [PubMed]
- National Comprehensive Cancer Network. Genetic/familial high-risk assessment: Breast and ovarian. JNCCN 2010, 8, 562–594. [Google Scholar]
- Hammond, M.E.H.; Hayes, D.F.; Dowsett, M.; Allred, D.C.; Hagerty, K.L.; Badve, S.; Fitzgibbons, P.L.; Francis, G.; Goldstein, N.S.; Hayes, M.; et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J. Clin. Oncol. 2010, 28, 2784–2795. [Google Scholar] [CrossRef] [PubMed]
- Deyarmin, B.; Kane, J.L.; Valente, A.L.; van Laar, R.; Gallagher, C.; Shriver, C.D.; Ellsworth, R.E. Effect of ASCO/CAP guidelines for determining ER status on molecular subtype. Ann. Surg. Oncol. 2013, 20, 87–93. [Google Scholar] [CrossRef]
- Iwamoto, T.; Booser, D.; Valero, V.; Murray, J.L.; Koenig, K.; Esteva, F.J.; Ueno, N.T.; Zhang, J.; Shi, W.; Qi, Y.; et al. Estrogen receptor (ER) mRNA and ER-related gene expression in breast cancers that are 1% to 10% ER-positive by immunohistochemistry. J. Clin. Oncol. 2012, 30, 729–734. [Google Scholar] [CrossRef] [PubMed]
- Yi, M.; Huo, L.; Koenig, K.B.; Mittendorf, E.A.; Meric-Bernstam, F.; Kuerer, H.M.; Bedrosian, I.; Buzdar, A.U.; Symmans, W.F.; Crow, J.R.; et al. Which threshold for ER positivity? a retrospective study based on 9639 patients. Ann. Oncol. 2014, 25, 1004–1011. [Google Scholar] [CrossRef] [PubMed]
- Allison, K.H.; Hammond, M.E.H.; Dowsett, M.; McKernin, S.E.; Carey, L.A.; Fitzgibbons, P.L.; Hayes, D.F.; Lakhani, S.R.; Chavez-MacGregor, M.; Perlmutter, J.; et al. Estrogen and progesterone receptor testing in breast cancer: ASCO/CAP guideline update. J. Clin. Oncol. 2020, 38, 1346–1366. [Google Scholar] [CrossRef] [PubMed]
- Wolff, A.C.; Hammond, M.E.H.; Allison, K.H.; Harvey, B.E.; Mangu, P.B.; Bartlett, J.M.S.; Bilous, M.; Ellis, I.O.; Fitzgibbons, P.; Hanna, W.; et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline focused update. J. Clin. Oncol. 2018, 36, 2105–2122. [Google Scholar] [CrossRef] [PubMed]
- Rummel, S.K.; Lovejoy, L.A.; Turner, C.E.; Shriver, C.D.; Ellsworth, R.E. Should genetic testing for cancer predisposition be standard-of-care for women with invasive breast cancer? The Murtha Cancer Center experience. Cancers 2020, 12, 234. [Google Scholar] [CrossRef] [PubMed]
- Miki, Y.; Swensen, J.; Shattuck-Eidens, D.; Futreal, P.A.; Harshman, K.; Tavtigian, S.; Liu, Q.; Cochran, C.; Bennett, L.M.; Ding, W.; et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA. Science 1994, 266, 66–71. [Google Scholar] [CrossRef] [PubMed]
- Wooster, R.; Bignell, G.; Lancaseter, J.; Swift, S.; Seal, S.; Mangion, J.; Collins, N.; Gregory, S.; Gumbs, C.; Micklem, G.; et al. Identification of the breast cancer susceptibility gene BRCA. Nature 1995, 378, 789–792. [Google Scholar] [CrossRef] [PubMed]
- Song, C.V.; Teo, S.H.; Taib, N.A.; Yip, C.H. Surgery for BRCA, TP53 and PALB2: A literature review. Ecancermedicalscience 2018, 12, 863. [Google Scholar] [CrossRef] [PubMed]
- Nicolas, E.; Bertucci, F.; Sabatier, R.; Goncalves, A. Targeting BRCA deficiency in breast cancer: What are the clinical evidences and the next perspectives? Cancers 2018, 10, 506. [Google Scholar] [CrossRef] [PubMed]
- Piombino, C.; Cortesi, L.; Lambertini, M.; Punie, K.; Grandi, G.; Toss, A. Secondary prevention in hereditary breast and/or ovarian cancer syndromes other than BRCA. J. Oncol. 2020, 2020, 6384190. [Google Scholar] [CrossRef] [PubMed]
- Sanford, R.A.; Song, J.; Gutierrez-Barrera, A.M.; Profato, J.; Woodson, A.; Litton, J.K.; Bedrosian, I.; Albarracin, C.T.; Valero, V.; Arun, B. High incidence of germline BRCA mutation in patients with ER low-positive/PR low-positive/HER-2 neu negative tumors. Cancer 2015, 121, 3422–3427. [Google Scholar] [CrossRef] [PubMed]
- Buys, S.S.; Sandbach, J.F.; Gammon, A.; Patel, G.; Kidd, J.; Brown, K.L.; Sharma, L.; Saam, J.; Lancaster, J.; Daly, M.B. A study of over 35,000 women with breast cancer tested with a 25-gene panel of hereditary cancer genes. Cancer 2017, 123, 1721–1730. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | ERLPHN (n = 60) | TNBC (n = 254) | p-Value |
|---|---|---|---|
| Age | 0.443 | ||
| ≤45 years | 11 (18.3%) | 55 (21.7%) | |
| 46–60 years | 30 (50.0%) | 104 (40.9%) | |
| >60 years | 19 (31.7%) | 95 (37.4%) | |
| Ethnicity | 0.178 | ||
| African American | 13 (21.6%) | 85 (33.4) | |
| Asian | 2 (3.3%) | 5 (2.0%) | |
| Hispanic | 1 (1.7%) | 6 (2.4%) | |
| Other | 1 (1.7%) | 3 (1.2%) | |
| European American | 42 (70.0%) | 155 (61.0%) | |
| Unknown | 1 (1.7%) | 0 (0.0%) | |
| Family history | 0.197 | ||
| 0 | 18 (30.0%) | 108 (42.5%) | |
| 1 | 25 (41.7%) | 78 (30.7%) | |
| 2 | 9 (15.0%) | 46 (18.1%) | |
| ≥3 | 8 (13.3%) | 20 (7.9%) | |
| Unknown | 0 (0.0%) | 2 (0.8%) | |
| Tumor size | 0.189 | ||
| T1 | 22 (36.7%) | 131 (51.6%) | |
| T2 | 30 (50.0%) | 97 (38.2%) | |
| T3/4 | 6 (10.0%) | 22 (8.6%) | |
| Unknown | 2 (3.3%) | 4 (1.6%) | |
| Tumor grade | 0.488 | ||
| Well-differentiated | 2 (3.3%) | 4 (1.6%) | |
| Moderately-differentiated | 12 (20.0%) | 35 (13.8%) | |
| Poorly-differentiated | 45 (75.0%) | 211 (83.0%) | |
| Unknown | 1 (1.7%) | 4 (1.6%) | |
| Tumor stage | 0.092 | ||
| I | 18 (30.0%) | 109 (42.9%) | |
| II | 30 (50.0%) | 111 (43.7%) | |
| III | 9 (15.0%) | 31 (12.2%) | |
| IV | 3 (5.0%) | 3 (1.2%) |
| Patient | Gene | Mutation | ≤45 Years at Diagnosis | <60 Years at Diagnosis | Family History a |
|---|---|---|---|---|---|
| ERLP/HER2- | |||||
| 38 | BRCA1 | Glu1250Ter | √ | √ | √ |
| 185 | BRCA1 | Mutation not provided b | √ | √ | |
| 191 | BRCA1 | Glu1250Ter | √ | √ | √ |
| 226 | BRCA1 | Gln1756Profs | √ | √ | √ |
| 78 | BRCA2 | Leu1466Phefs | √ | ||
| 117 | BRCA2 | Leu2510Pro | √ | ||
| 125 | BRCA2 | Ile1859Lysfs | √ | √ | |
| 171 | CHEK2 | Thr367Metfs | √ | ||
| 213 | MSH6 | Phe1088Leufs | √ | √ | |
| TNBC | |||||
| 280 | ATM | Glu2236Ter | |||
| 230 | ATM | 901 + 1G > A | √ | ||
| 289 | ATM | Tyr1124Ter | √ | ||
| 303 | BAP1 | Leu573fs | √ | ||
| 5 | BRCA1 | 5193 + 2del | √ | ||
| 48 | BRCA1 | Gln1408Ter | √ | √ | |
| 54 | BRCA1 | Met1775Arg | √ | √ | |
| 73 | BRCA1 | 5467 + 1G > A | √ | √ | |
| 80 | BRCA1 | Glu908Ter | √ | ||
| 110 | BRCA1 | Gln1756Profs | √ | ||
| 112 | BRCA1 | Ser1655fs | √ | √ | √ |
| 123 | BRCA1 | 4986 + 6T > G | √ | √ | √ |
| 150 | BRCA1 | Val1234fs | √ | √ | √ |
| 159 | BRCA1 | Val1688del | √ | √ | |
| 172 | BRCA1 | 4986 + 3G > C | √ | √ | |
| 182 | BRCA1 | Glu23fs | √ | √ | |
| 201 | BRCA1 | Mutation not provided b | √ | √ | √ |
| 219 | BRCA1 | Val1838Glu | √ | √ | √ |
| 237 | BRCA1 | Glu23fs | √ | √ | |
| 239 | BRCA1 | Mutation not provided b | √ | ||
| 280 | BRCA1 | Met1775Arg | |||
| 283 | BRCA1 | 135-1G > T | √ | √ | √ |
| 288 | BRCA1 | Thr276fs | √ | √ | |
| 310 | BRCA1 | Lys894fs | √ | √ | √ |
| 62 | BRCA2 | Asp1199_Cys1200insTer | |||
| 109 | BRCA2 | Tyr1894Terfs | √ | √ | |
| 139 | BRCA2 | Asn1784fs | √ | ||
| 236 | BRCA2 | Ile2315fs | |||
| 238 | BRCA2 | Asn1626Serfs | |||
| 268 | BRCA2 | Asn986fs | √ | ||
| 277 | BRCA2 | Leu2805fs | √ | √ | |
| 308 | BRCA2 | Cys1200Terfs | √ | ||
| 133 | MUTYH | Arg233Ter | √ | ||
| 221 | MUTYH | 850-2A > G | √ | √ | |
| 241 | PALB2 | Tyr1183Ter | √ | √ | |
| 220 | TP53 | Arg175Gly | √ | √ | |
| 300 | TP53 | Arg273Cys | √ | √ | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lovejoy, L.A.; Turner, C.E.; Wells, J.M.; Shriver, C.D.; Ellsworth, R.E. Heritability of Low ER Staining/HER2-Breast Tumors: Are We Missing an Opportunity for Germline Testing? Genes 2020, 11, 1469. https://doi.org/10.3390/genes11121469
Lovejoy LA, Turner CE, Wells JM, Shriver CD, Ellsworth RE. Heritability of Low ER Staining/HER2-Breast Tumors: Are We Missing an Opportunity for Germline Testing? Genes. 2020; 11(12):1469. https://doi.org/10.3390/genes11121469
Chicago/Turabian StyleLovejoy, Leann A., Clesson E. Turner, Justin M. Wells, Craig D. Shriver, and Rachel E. Ellsworth. 2020. "Heritability of Low ER Staining/HER2-Breast Tumors: Are We Missing an Opportunity for Germline Testing?" Genes 11, no. 12: 1469. https://doi.org/10.3390/genes11121469
APA StyleLovejoy, L. A., Turner, C. E., Wells, J. M., Shriver, C. D., & Ellsworth, R. E. (2020). Heritability of Low ER Staining/HER2-Breast Tumors: Are We Missing an Opportunity for Germline Testing? Genes, 11(12), 1469. https://doi.org/10.3390/genes11121469






