Multiple Self-Healing Squamous Epithelioma (MSSE): A Digenic Trait Associated with Loss of Function Mutations in TGFBR1 and Variants at a Second Linked Locus on the Long Arm of Chromosome 9
Abstract
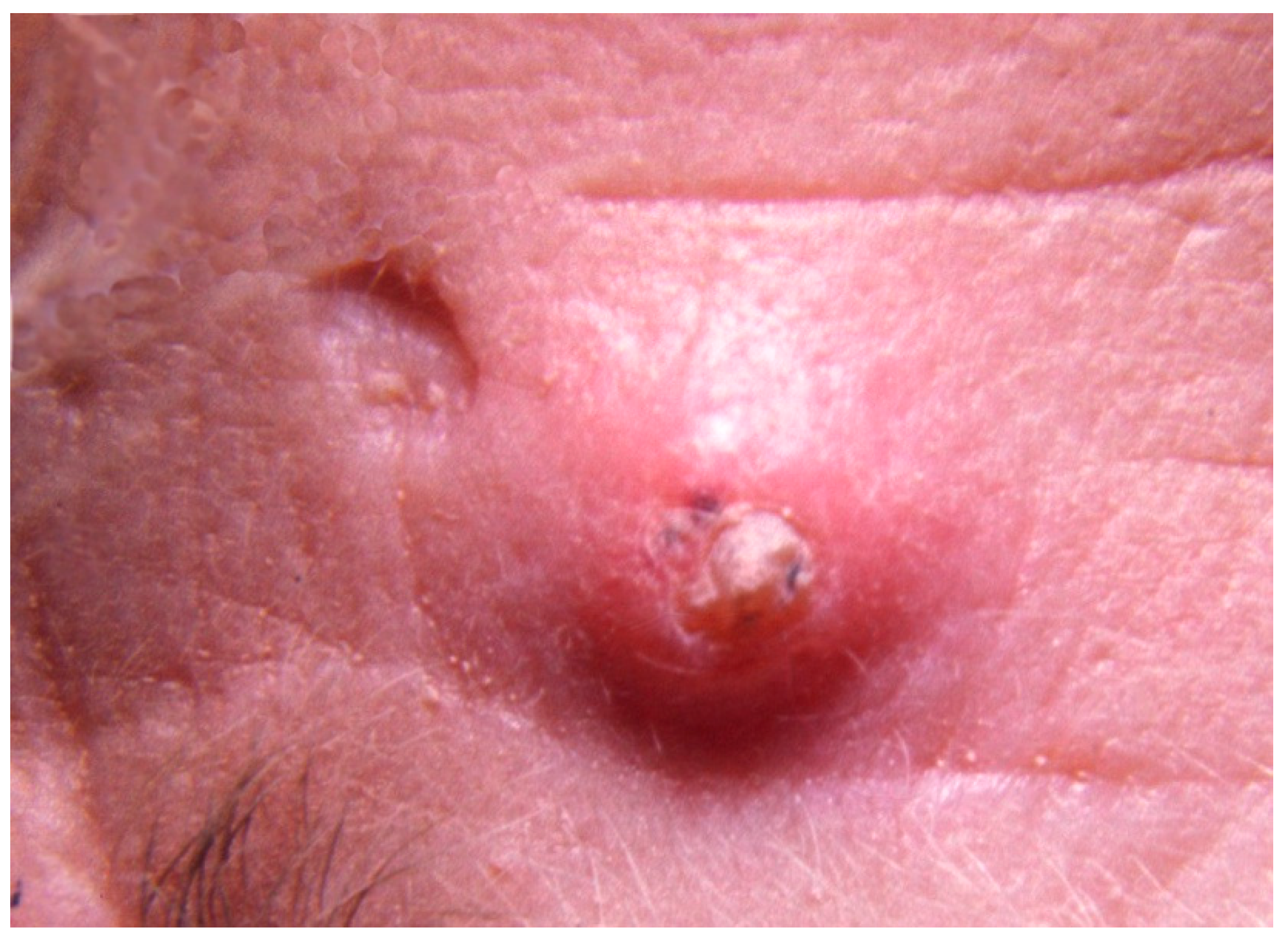
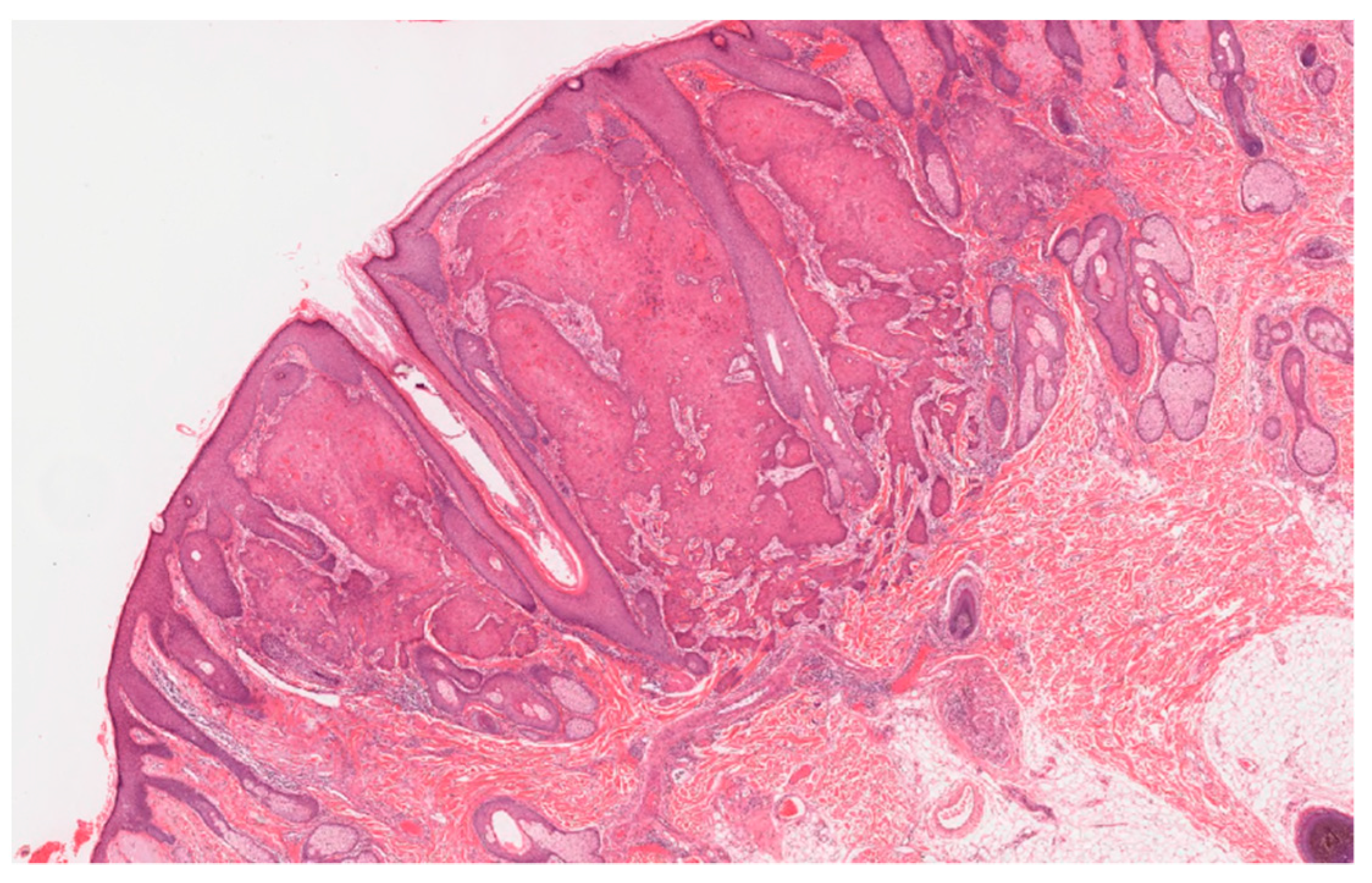
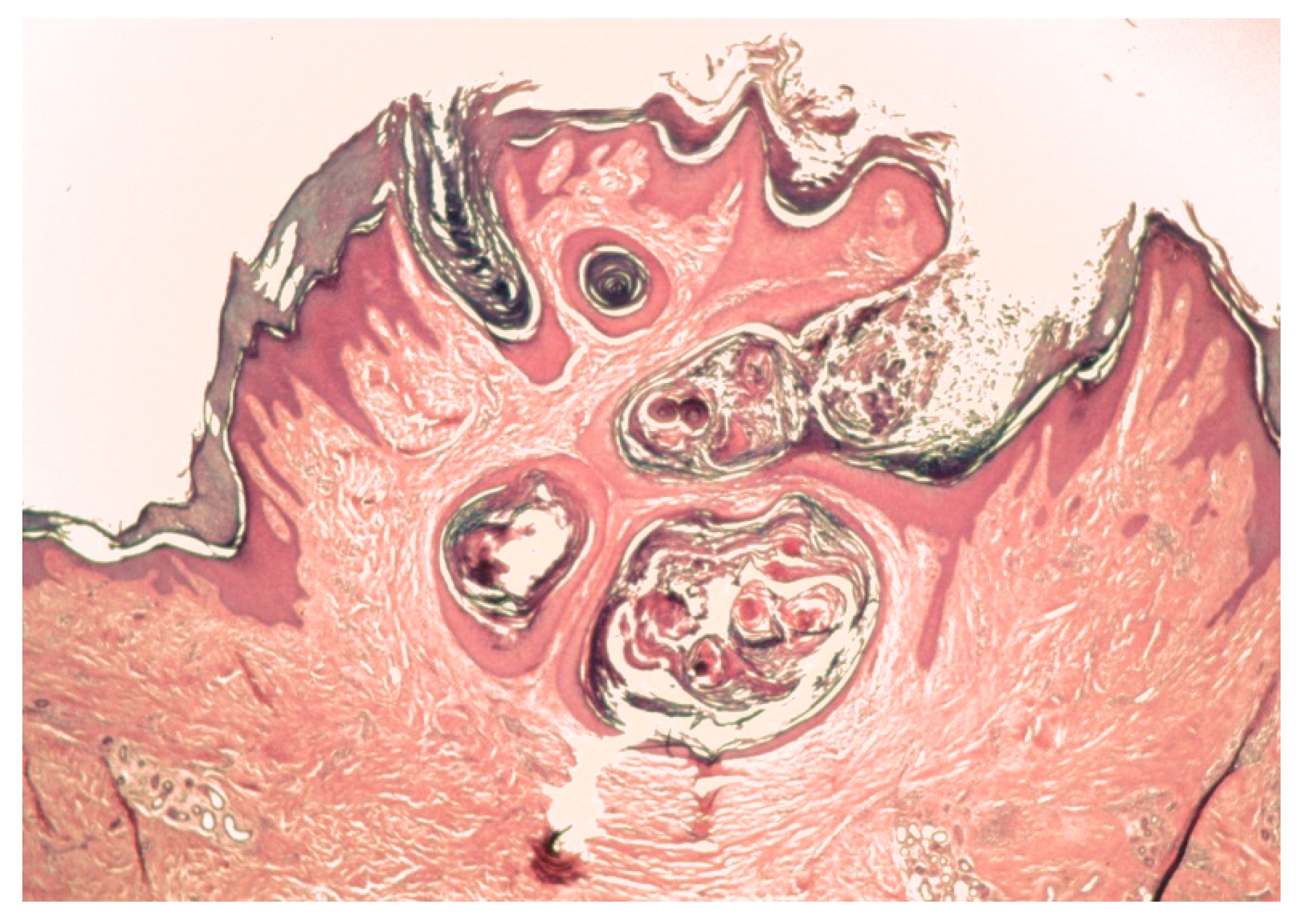
1. Clinical Features
2. Genetic Linkage
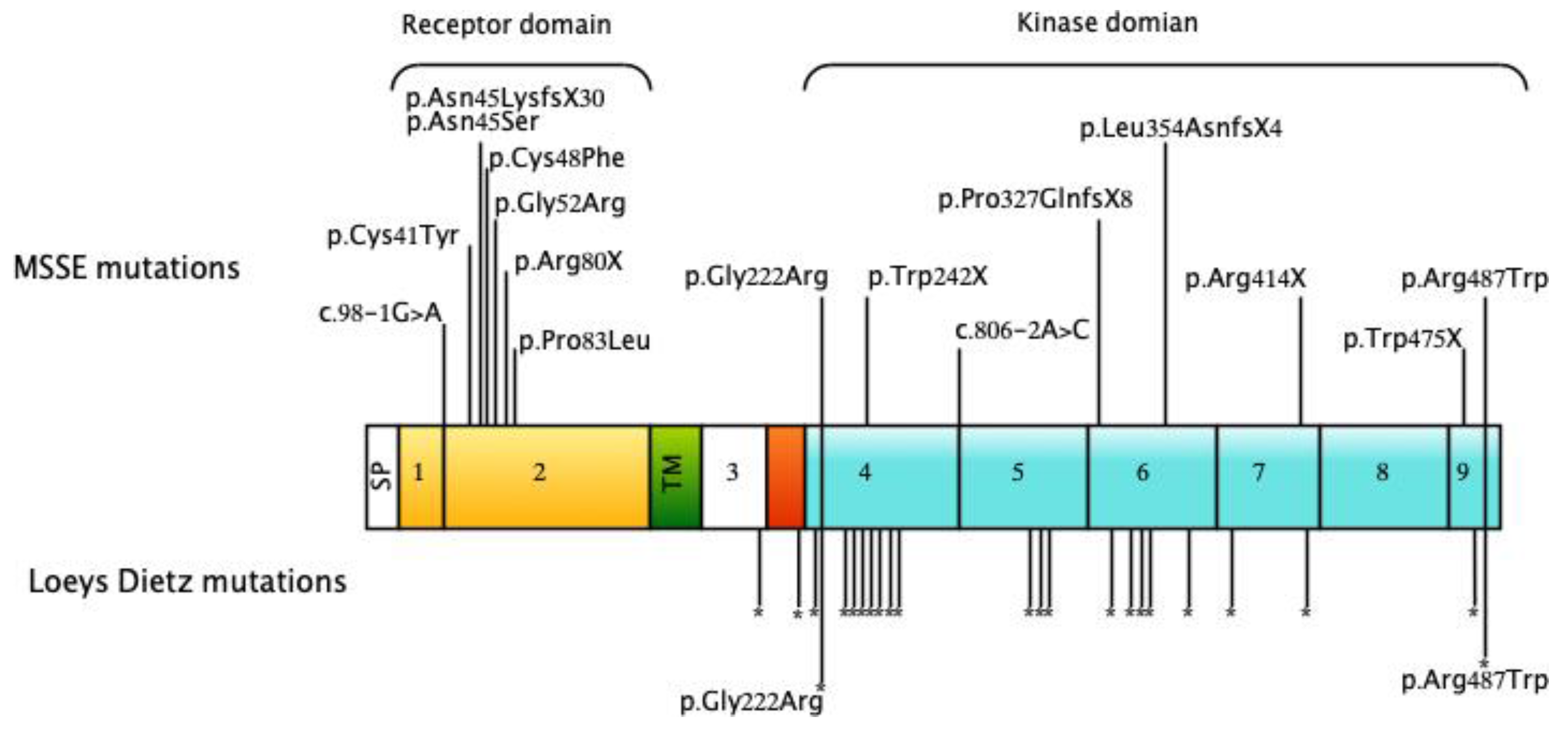
3. Pathogenic TGFBR1 Sequence Variants Cause MSSE
4. Digenic Inheritance: TGFBR1 and a Second Linked Locus on the Long Arm of Chromosome 9
5. MSSE Is Allelic to Loeys Dietz Syndrome Caused by TGFBR1 Mutations
6. TGFBR1 Variants in MSSE Patients with Aortic Aneurysms
7. The Second Locus and the Distinctive Spectrum of TGFBR1 Mutations in Loeys Diets Syndrome and MSSE
8. Keratoacanthomas/Squamous Cell Carcinomas in Patients Treated with an TGFβ Monoclonal Antibody
9. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Ferguson-Smith, J. A case of multiple primary squamous-celled carcinomata in a young man, with spontaneous healing. Br. J. Dermatol. 1934, 46, 267. [Google Scholar] [CrossRef]
- Ferguson-Smith, M.A.; Wallace, D.C.; James, Z.H.; Renwick, J.H. Multiple self-healing squamous epithelioma. Birth Defects Orig. Artic. Ser. 1971, 7, 157–163. [Google Scholar] [PubMed]
- Ferguson-Smith, M.A.; Goudie, D.R. Digenic/multilocus aetiology of multiple self-healing squamous epithelioma (Ferguson-Smith disease): TGFBR1 and a second linked locus. Int. J. Biochem. Cell Biol. 2014, 53, 520–525. [Google Scholar] [CrossRef] [PubMed]
- Wright, A.L.; Gawkrodger, D.J.; Branford, W.A.; McLaren, K.; Hunter, J.A. Self-healing epitheliomata of Ferguson-Smith: Cytogenetic and histological studies, and the therapeutic effect of etretinate. Dermatologica 1988, 176, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Robertson, S.J.; Bashir, S.J.; Pichert, G.; Robson, A.; Whittaker, S. Severe exacerbation of multiple self-healing squamous epithelioma (Ferguson-Smith disease) with radiotherapy, which was successfully treated with acitretin. Clin. Exp. Derm. 2009, 35, e100–e102. [Google Scholar] [CrossRef] [PubMed]
- Osunsade, L.; McEwan, C.N.; Lamont, D.; Yung, A. The importance of a multidisciplinary approach to the diagnosis and management of Ferguson-Smith disease: A patient’s 24-year journey to diagnosis. Eur. J. Plast. Surg. 2012, 35, 763–766. [Google Scholar] [CrossRef]
- Goudie, D.R.; Yuille, M.A.; Leversha, M.A.; Furlong, R.A.; Carter, N.P.; Lush, M.J.; Affara, N.A.; Ferguson-Smith, M.A. Multiple self-healing squamous epitheliomata (ESS1) mapped to chromosome 9q22-q31 in families with common ancestry. Nat. Genet. 1993, 3, 165–169. [Google Scholar] [CrossRef]
- Richards, F.M.; Goudie, D.R.; Cooper, W.N.; Jene, Q.; Barroso, I.; Wicking, C.; Wainwright, B.J.; Ferguson-Smith, M.A. Mapping the multiple self-healing squamous epithelioma (MSSE) gene and investigation of xeroderma pigmentosum group A (XPA) and PATCHED (PTCH) as candidate genes. Hum. Genet. 1997, 101, 317–322. [Google Scholar] [CrossRef]
- Goudie, D.R.; D’Alessandro, M.; Merriman, B.; Lee, H.; Szeverenyi, I.; Avery, S.; O’Connor, B.D.; Nelson, S.F.; Coats, S.E.; Stewart, A.; et al. Multiple self-healing squamous epithelioma is caused by a disease-specific spectrum of mutations in TGFBR1. Nat. Genet. 2011, 43, 365–369. [Google Scholar] [CrossRef]
- Bose, S.; Morgan, L.J.; Booth, D.R.; Goudie, D.R.; Ferguson-Smith, M.A.; Richards, F.M. The elusive multiple self-healing squamous epithelioma (MSSE) gene: Further mapping, analysis of candidates, and loss of heterozygosity. Oncogene 2006, 25, 806–812. [Google Scholar] [CrossRef]
- D’Alessandro, M.; Coats, S.E.; Morley, S.M.; Mackintosh, L.; Tessari, G.; Turco, A.; Gerdes, A.M.; Pichert, G.; Whittaker, S.; Brandrup, F.; et al. Multiple self-healing squamous epithelioma in different ethnic groups: More than a founder mutation disorder? J. Investig. Dermatol. 2007, 127, 2336–2344. [Google Scholar] [CrossRef] [PubMed]
- Debroy Kidambi, A.; Cook, J.; Messenger, A.G. A case of Ferguson–Smith disease. Clin. Exp. Dermatol. 2017, 42, 570–572. [Google Scholar] [CrossRef] [PubMed]
- Müller, C.; Güvenc, B.; Henn, W.; Christmann, A.; Vogt, T.; Pföhler, C. Familiäre Keratoakanthome vom Typ Ferguson-Smith: Ein Fallbericht mit Nachweis einer bislang nicht beschriebenen Mutation im TGFBR-1-Gene. Aktuelle Dermatol. 2016, 42, 518–519. [Google Scholar] [CrossRef]
- Shukur, Z.; Chung, N.; Goudie, D.; Lacy, K.; Shaw, A. Novel phenotypic overlap between Ferguson-smith disease and Loeys-Dietz syn- drome. 11th BAD RCP medical dermatology meeting; London. Br. J. Dermatol. 2016, 174, 24–31. [Google Scholar]
- Sirisomboonwong, K.E.; Martindale, J.; Keefe, M.; Goudie, D.; Poke, G. Features of multiple self-healing squamous epithelioma and Loeys-Dietz syndrome in a patient with a novel TGFBR1 variant. Am. J. Med. Genet. A 2018, 176, 2892–2895. [Google Scholar] [CrossRef]
- Feldmeyer, L.; Szeverenyi, I.; Mandallaz, M.; Lane, E.B.; Hohl, D. Late-Onset Multiple Self-Healing Squamous Epithelioma Ferguson-Smith Recurrence Induced by Radiotherapy. Case Rep. Derm. 2016, 8, 344–349. [Google Scholar] [CrossRef]
- Loeys, B.L.; Schwarze, U.; Holm, T. Aneurysm syndromes caused by mutations in the TGF-beta receptor. N. Engl. J. Med. 2006, 355. [Google Scholar] [CrossRef]
- Loeys, B.L.; Chen, J.; Neptune, E.R. A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat. Genet. 2005, 37. [Google Scholar] [CrossRef]
- Mizuguchi, T.; Collod-Beroud, G.; Akiyama, T.; Abifadel, M.; Harada, N.; Morisaki, T.; Allard, D.; Varret, M.; Claustres, M.; Morisaki, H.; et al. Heterozygous TGFBR2 mutations in Marfan syndrome. Nat. Genet. 2004, 36, 855–860. [Google Scholar] [CrossRef]
- Pannu, H.; Fadulu, V.T.; Chang, J.; Lafont, A.; Hasham, S.N.; Sparks, E.; Giampietro, P.F.; Zaleski, C.; Estrera, A.L.; Safi, H.J.; et al. Mutations in transforming growth factor-beta receptor type II cause familial thoracic aortic aneurysms and dissections. Circulation 2005, 112, 513–520. [Google Scholar] [CrossRef]
- Ades, L.C.; Sullivan, K.; Biggin, A.; Haan, E.A.; Brett, M.; Holman, K.J.; Dixon, J.; Robertson, S.; Holmes, A.D.; Rogers, J.; et al. FBN1, TGFBR1, and the Marfan-craniosynostosis/mental retardation disorders revisited. Am. J. Med. Genet. A 2006, 140, 1047–1058. [Google Scholar] [CrossRef] [PubMed]
- Akutsu, K.; Morisaki, H.; Takeshita, S.; Sakamoto, S.; Tamori, Y.; Yoshimuta, T.; Yokoyama, N.; Nonogi, H.; Ogino, H.; Morisaki, T. Phenotypic heterogeneity of Marfan-like connective tissue disorders associated with mutations in the transforming growth factor-beta receptor genes. Circ. J. 2007, 71, 1305–1309. [Google Scholar] [CrossRef] [PubMed]
- Drera, B.; Tadini, G.; Barlati, S.; Colombi, M. Identification of a novel TGFBR1 mutation in a Loeys-Dietz syndrome type II patient with vascular Ehlers-Danlos syndrome phenotype. Clin. Genet. 2008, 73, 290–293. [Google Scholar] [CrossRef] [PubMed]
- Drera, B.; Ritelli, M.; Zoppi, N.; Wischmeijer, A.; Gnoli, M.; Fattori, R.; Calzavara-Pinton, P.G.; Barlati, S.; Colombi, M. Loeys-Dietz syndrome type I and type II: Clinical findings and novel mutations in two Italian patients. Orphanet J. Rare Dis. 2009, 4, 24. [Google Scholar] [CrossRef]
- Matyas, G.; Arnold, E.; Carrel, T.; Baumgartner, D.; Boileau, C.; Berger, W.; Steinmann, B. Identification and in silico analyses of novel TGFBR1 and TGFBR2 mutations in Marfan syndrome-related disorders. Hum. Mutat. 2006, 27, 760–769. [Google Scholar] [CrossRef]
- Sakai, H.; Visser, R.; Ikegawa, S.; Ito, E.; Numabe, H.; Watanabe, Y.; Mikami, H.; Kondoh, T.; Kitoh, H.; Sugiyama, R.; et al. Comprehensive genetic analysis of relevant four genes in 49 patients with Marfan syndrome or Marfan-related phenotypes. Am. J. Med. Genet. A 2006, 140, 1719–1725. [Google Scholar] [CrossRef]
- Singh, K.K.; Rommel, K.; Mishra, A.; Karck, M.; Haverich, A.; Schmidtke, J.; Arslan-Kirchner, M. TGFBR1 and TGFBR2 mutations in patients with features of Marfan syndrome and Loeys-Dietz syndrome. Hum. Mutat. 2006, 27, 770–777. [Google Scholar] [CrossRef]
- Soylen, B.; Singh, K.K.; Abuzainin, A.; Rommel, K.; Becker, H.; Arslan-Kirchner, M.; Schmidtke, J. Prevalence of dural ectasia in 63 gene-mutation-positive patients with features of Marfan syndrome type 1 and Loeys-Dietz syndrome and report of 22 novel FBN1 mutations. Clin. Genet. 2009, 75, 265–270. [Google Scholar] [CrossRef]
- Tran-Fadulu, V.; Pannu, H.; Kim, D.H.; Vick, G.W., 3rd; Lonsford, C.M.; Lafont, A.L.; Boccalandro, C.; Smart, S.; Peterson, K.L.; Hain, J.Z.; et al. Analysis of multigenerational families with thoracic aortic aneurysms and dissections due to TGFBR1 or TGFBR2 mutations. J. Med. Genet. 2009, 46, 607–613. [Google Scholar] [CrossRef]
- Renard, M.; Trachet, B.; Casteleyn, C.; Campens, L.; Cornillie, P.; Callewaert, B.; Deleye, S.; Vandeghinste, B.; van Heijningen, P.M.; Dietz, H.; et al. Absence of cardiovascular manifestations in a haploinsufficient Tgfbr1 mouse model. PLoS ONE 2014, 9, e89749. [Google Scholar] [CrossRef]
- Chung Kang, H.; Quigley, D.A.; Kim, I.J.; Wakabayashi, Y.; Ferguson-Smith, M.A.; D’Alessandro, M.; Birgitte Lane, E.; Akhurst, R.J.; Goudie, D.R.; Balmain, A. Multiple Self-Healing Squamous Epithelioma (MSSE): Rare Variants in an Adjacent Region of Chromosome 9q22.3 to Known TGFBR1 Mutations Suggest a Digenic or Multilocus Etiology. J. Investig. Dermatol. 2013. [Google Scholar] [CrossRef] [PubMed]
- van de Laar, I.M.B.H.; Oldenburg, R.A.; Pals, G.; Roos-Hesselink, J.W.; de Graaf, B.M.; Verhagen, J.M.A.; Hoedemaekers, Y.M.; Willemsen, R.; Severijnen, L.-A.; Venselaar, H.; et al. Mutations in SMAD3 cause a syndromic form of aortic aneurysms and dissections with early-onset osteoarthritis. Nat. Genet. Adv. Online Publ. 2011, 43, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Bertoli-Avella, A.M.; Gillis, E.; Morisaki, H.; Verhagen, J.M.A.; de Graaf, B.M.; van de Beek, G.; Gallo, E.; Kruithof, B.P.T.; Venselaar, H.; Myers, L.A.; et al. Mutations in a TGF-β Ligand, TGFB3, Cause Syndromic Aortic Aneurysms and Dissections. J. Am. Coll. Cardiol. 2015, 65, 1324–1336. [Google Scholar] [CrossRef] [PubMed]
- Lindsay, M.E.; Schepers, D.; Bolar, N.A.; Doyle, J.J.; Gallo, E.; Fert-Bober, J.; Kempers, M.J.E.; Fishman, E.K.; Chen, Y.; Myers, L.; et al. Loss-of-function mutations in TGFB2 cause a syndromic presentation of thoracic aortic aneurysm. Nat. Genet. 2012, 44, 922–927. [Google Scholar] [CrossRef] [PubMed]
- Rienhoff, H.Y., Jr.; Yeo, C.-Y.; Morissette, R.; Khrebtukova, I.; Melnick, J.; Luo, S.; Leng, N.; Kim, Y.-J.; Schroth, G.; Westwick, J.; et al. A mutation in TGFB3 associated with a syndrome of low muscle mass, growth retardation, distal arthrogryposis and clinical features overlapping with marfan and loeys–dietz syndrome. Am. J. Med. Genet. Part. A 2013, 161, 2040–2046. [Google Scholar] [CrossRef] [PubMed]
- Matyas, G.; Naef, P.; Tollens, M.; Oexle, K. De novo mutation of the latency-associated peptide domain of TGFB3 in a patient with overgrowth and Loeys–Dietz syndrome features. Am. J. Med. Genet. Part. A 2014, 164, 2141–2143. [Google Scholar] [CrossRef]
- Makino, K.; Jinnin, M.; Hirano, A.; Yamane, K.; Eto, M.; Kusano, T.; Honda, N.; Kajihara, I.; Makino, T.; Sakai, K.; et al. The downregulation of microRNA let-7a contributes to the excessive expression of type I collagen in systemic and localized scleroderma. J. Immunol. 2013, 190, 3905–3915. [Google Scholar] [CrossRef]
- Fujiwara, T.; Takeda, N.; Hara, H.; Morita, H.; Kishihara, J.; Inuzuka, R.; Yagi, H.; Maemura, S.; Toko, H.; Harada, M.; et al. Distinct variants affecting differential splicing of TGFBR1 exon 5 cause either Loeys–Dietz syndrome or multiple self-healing squamous epithelioma. Eur. J. Hum. Genet. 2018, 26, 1151–1158. [Google Scholar] [CrossRef]
- Gallo, E.M.; Loch, D.C.; Habashi, J.P.; Calderon, J.F.; Chen, Y.; Bedja, D.; van Erp, C.; Gerber, E.E.; Parker, S.J.; Sauls, K.; et al. Angiotensin II-dependent TGF-beta signaling contributes to Loeys-Dietz syndrome vascular pathogenesis. J. Clin. Investig. 2014, 124, 448–460. [Google Scholar] [CrossRef]
- Cardoso, S.; Robertson, S.P.; Daniel, P.B. TGFBR1 mutations associated with Loeys-Dietz syndrome are inactivating. J. Recept. Signal. Transduct. Res. 2012, 32, 150–155. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alfoldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.C.; Tan, A.R.; Olencki, T.E.; Shapiro, G.I.; Dezube, B.J.; Reiss, M.; Hsu, F.J.; Berzofsky, J.A.; Lawrence, D.P. Phase I Study of GC1008 (Fresolimumab): A Human Anti-Transforming Growth Factor-beta (TGFβ) Monoclonal Antibody in Patients with Advanced Malignant Melanoma or Renal Cell Carcinoma. PLoS ONE 2014, 9, e90353. [Google Scholar] [CrossRef] [PubMed]
- Derynck, R.; Turley, S.J.; Akhurst, R.J. TGFbeta biology in cancer progression and immunotherapy. Nat. Rev. Clin. Oncol. 2020. [Google Scholar] [CrossRef]
- Cui, W.; Fowlis, D.J.; Bryson, S.; Duffie, E.; Ireland, H.; Balmain, A.; Akhurst, R.J. TGFbeta1 inhibits the formation of benign skin tumors, but enhances progression to invasive spindle carcinomas in transgenic mice. Cell 1996, 86, 531–542. [Google Scholar] [CrossRef]
- Cammareri, P.; Rose, A.M.; Vincent, D.F.; Wang, J.; Nagano, A.; Libertini, S.; Ridgway, R.A.; Athineos, D.; Coates, P.J.; McHugh, A.; et al. Inactivation of TGFbeta receptors in stem cells drives cutaneous squamous cell carcinoma. Nat. Commun. 2016, 7, 12493. [Google Scholar] [CrossRef]
- Ji, A.L.; Rubin, A.J.; Thrane, K.; Jiang, S.; Reynolds, D.L.; Meyers, R.M.; Guo, M.G.; George, B.M.; Mollbrink, A.; Bergenstrahle, J.; et al. Multimodal Analysis of Composition and Spatial Architecture in Human Squamous Cell Carcinoma. Cell 2020, 182, 497–514.e422. [Google Scholar] [CrossRef]
| Marker loci | Position | Families | ||
|---|---|---|---|---|
| 4–10 | 2 | 18 | ||
| Assembly GRCh37/hg19 | Alleles | |||
| D9S119 | 95867389-95867529 | 1 | 1 | 1 |
| D9S197 | 95875618-95875968 | 2 | 2/4 * | 2 |
| D9S196 | 95906010-95906446 | 5 | 4/5 * | 5 |
| ZNF169CA | 96264697-96265065 | 8 | 8 | 8 |
| FBP1 (GA) | 96474007-96474379 | 2 | 2 | 1/2 * |
| AFM070xb11 | 97021578-97065291 | 1 | 1 | 1 |
| AFM086yf1 | 97365423-97402531 | 1 | 1 | 1 |
| D9S280 | 97365423-97344962 | 2 | 2 | 2 |
| FANCC CA Intr2 | 97912205-98079991 | 2/3 * | 2 | 2/4 * |
| FANCC EcoRI | 97912205-98079991 | 2 | 1/2 * | 2 |
| FANCC CA Intr1 | 97912205-98079991 | 1 | 1 | 1 |
| AFM203WH8 * | 98073744-98074099 | 1 | 1 | 1 |
| PTC INT5 | 98205266-98270831 | T | T | T |
| PTC INT9 | 98205266-98270831 | C | C | C |
| PTC INT10 | 98205266-98270831 | G | G | G |
| D9S1816 | 98276797-98277080 | 5 | 5 | 5/8 * |
| D9S287 | 98466090-98466414 | 3 | 3 | 3 |
| AFMa350xgi | 98544811-98545188 | 2 | 2 | 2 |
| D9S1809 | 98558557-98558805 | 5 | 5 | 5 |
| AFM023xh8 | 99032436-99032712 | 1 | 1/2 * | 1 |
| D9S1851 | 99570721-99571084 | 2 | 2 | 2 |
| XPA (MspI) | 100437192-100459691 | A | G | G |
| D9S180 | 100649474-100649706 | 1 | 2 | 2 |
| D9S6 | 100931590-100931609 | A2 | A1 | A1 |
| D9S173 | 101890198-101916471 | 2 | 4 | 2/4 * |
| D9S176 | 102058162-102058520 | 7 | 3/5 * | 5 |
| TGFBR1 | 103142351-103142640 | p.(G52R) | p.(N45S) | p.(R414X) |
| ALDOB | 104182842-104198062 | 1 | 1 | 1/2 * |
| D9S109 | 105566169-10556648 | 2 | 3 | 6 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goudie, D. Multiple Self-Healing Squamous Epithelioma (MSSE): A Digenic Trait Associated with Loss of Function Mutations in TGFBR1 and Variants at a Second Linked Locus on the Long Arm of Chromosome 9. Genes 2020, 11, 1410. https://doi.org/10.3390/genes11121410
Goudie D. Multiple Self-Healing Squamous Epithelioma (MSSE): A Digenic Trait Associated with Loss of Function Mutations in TGFBR1 and Variants at a Second Linked Locus on the Long Arm of Chromosome 9. Genes. 2020; 11(12):1410. https://doi.org/10.3390/genes11121410
Chicago/Turabian StyleGoudie, David. 2020. "Multiple Self-Healing Squamous Epithelioma (MSSE): A Digenic Trait Associated with Loss of Function Mutations in TGFBR1 and Variants at a Second Linked Locus on the Long Arm of Chromosome 9" Genes 11, no. 12: 1410. https://doi.org/10.3390/genes11121410
APA StyleGoudie, D. (2020). Multiple Self-Healing Squamous Epithelioma (MSSE): A Digenic Trait Associated with Loss of Function Mutations in TGFBR1 and Variants at a Second Linked Locus on the Long Arm of Chromosome 9. Genes, 11(12), 1410. https://doi.org/10.3390/genes11121410




