Recent Advances in the Enzymatic Synthesis of Polyester
Abstract
1. Introduction
2. Current Polyester Synthesis Method and a Possible Alternative
3. Enzyme-Catalyzed Synthesis of Polyesters
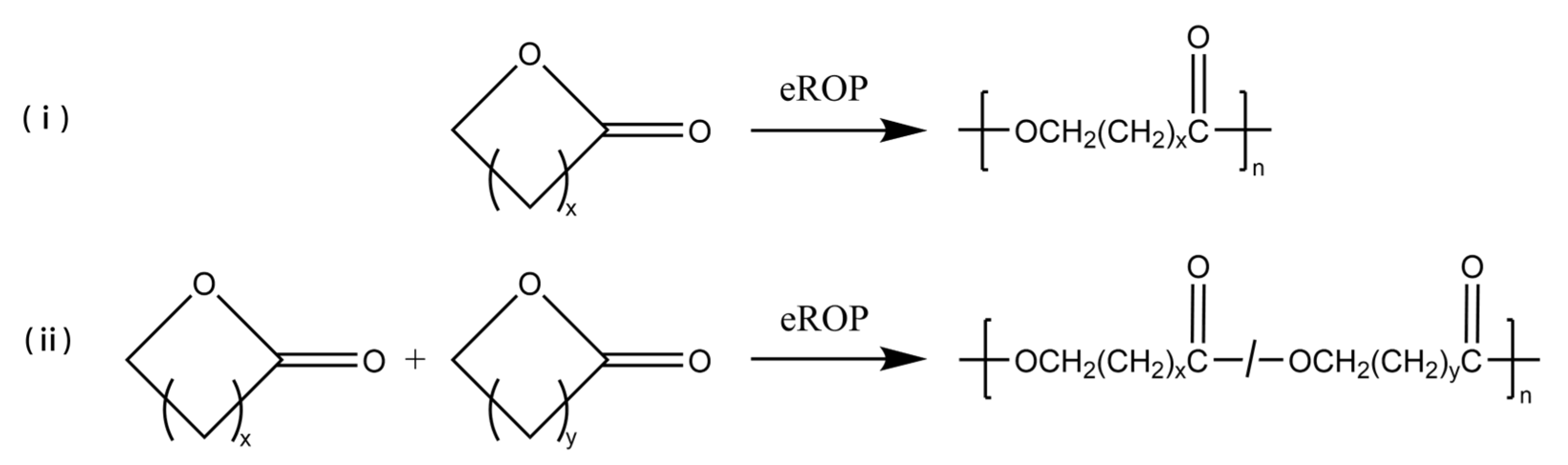
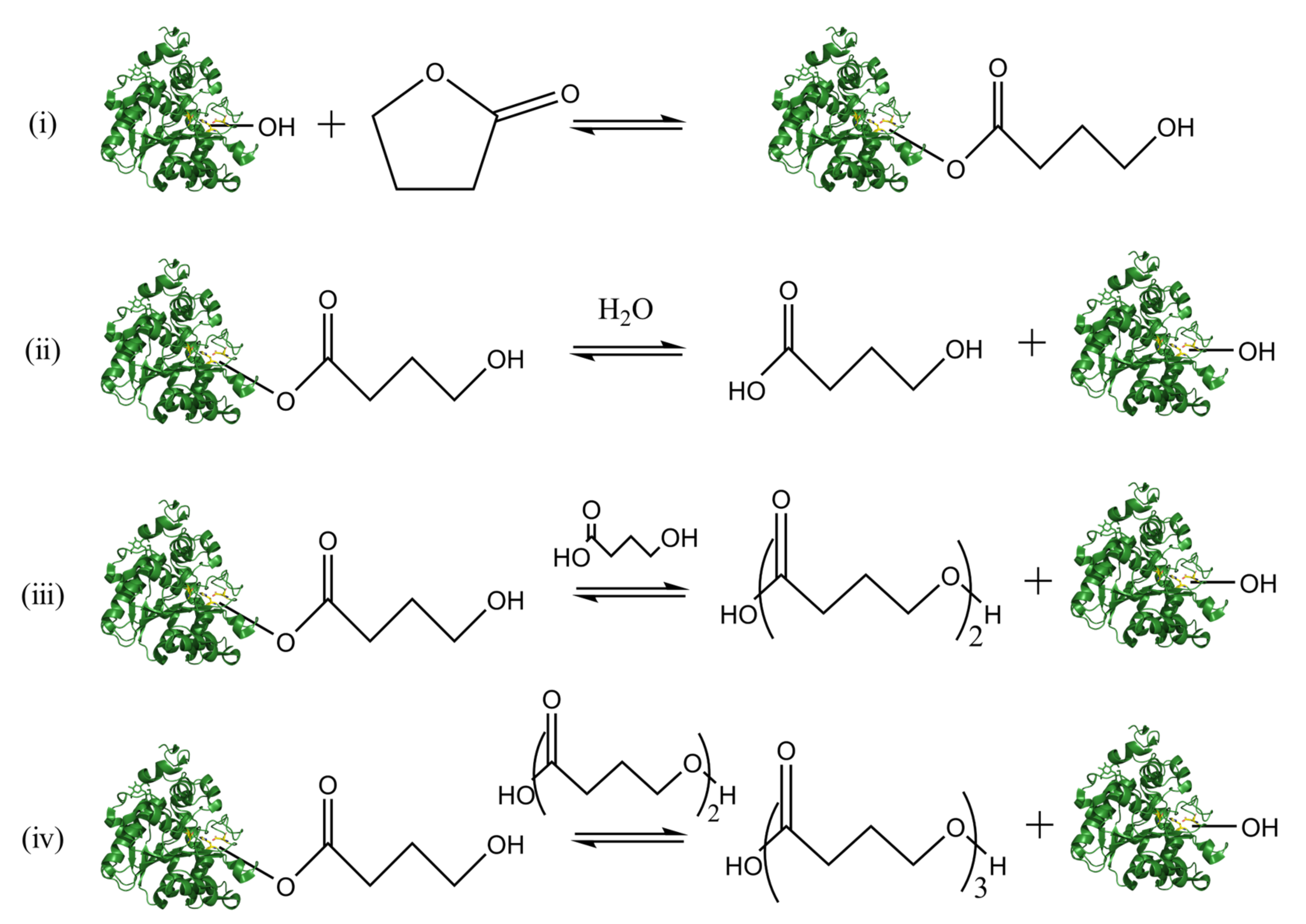
3.1. Enzyme-Catalyzed Ring-Opening Polymerization
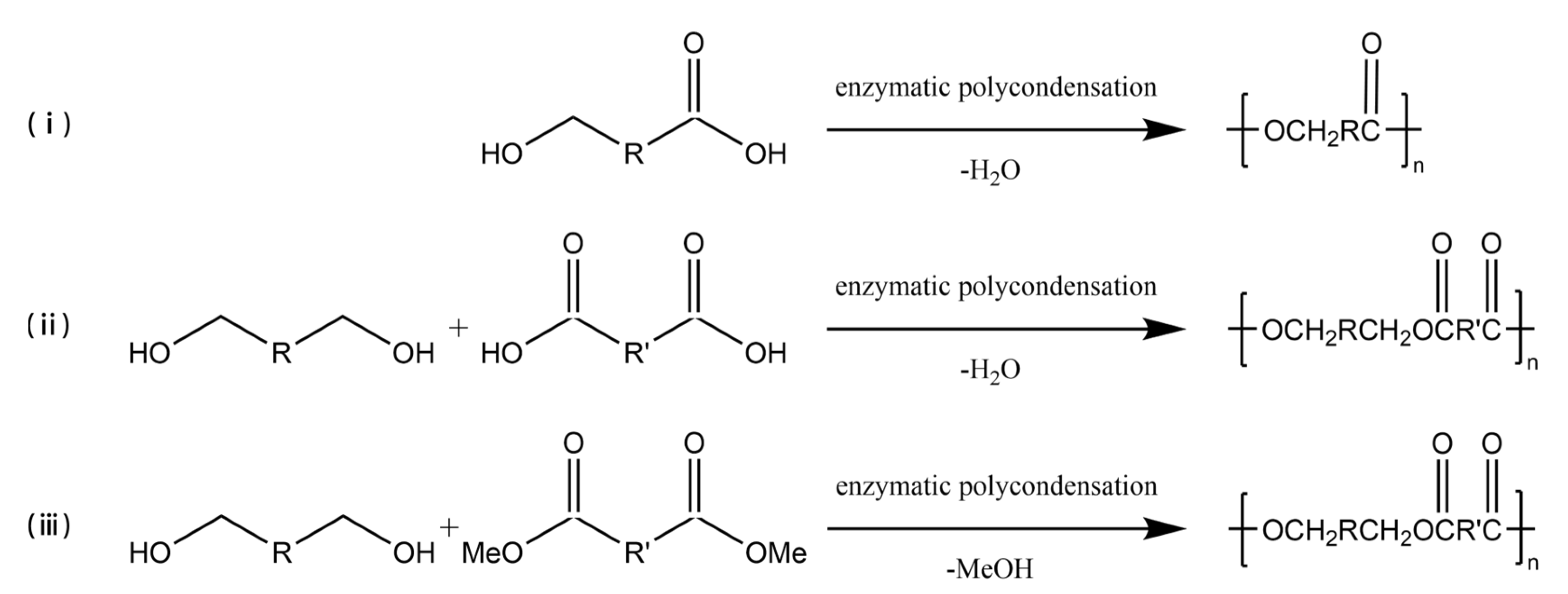
3.2. Enzyme-Catalyzed Polycondensation
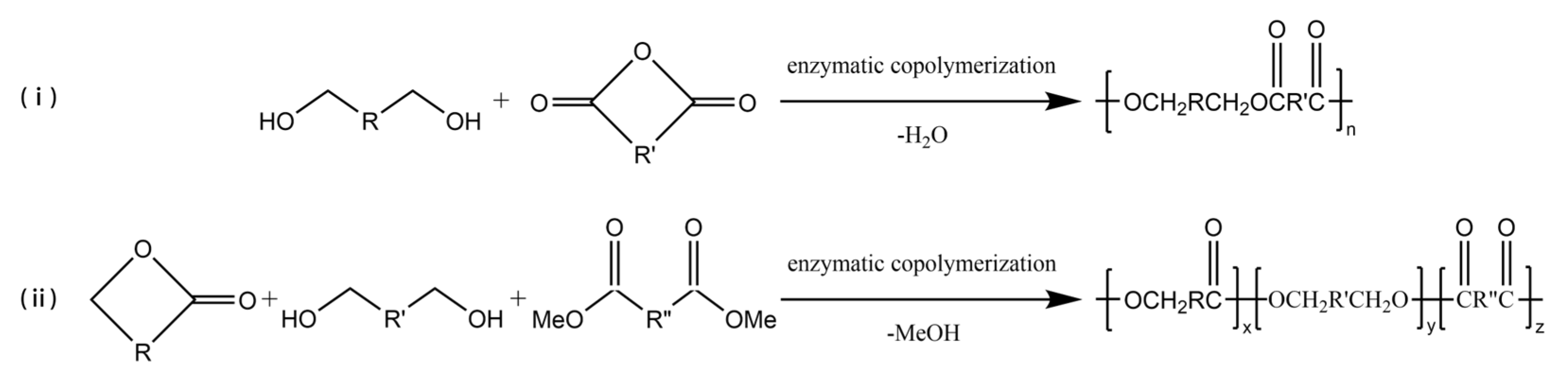
3.3. Enzyme-Catalyzed Copolymerization
4. Overview of the Biological Catalyst: Lipase
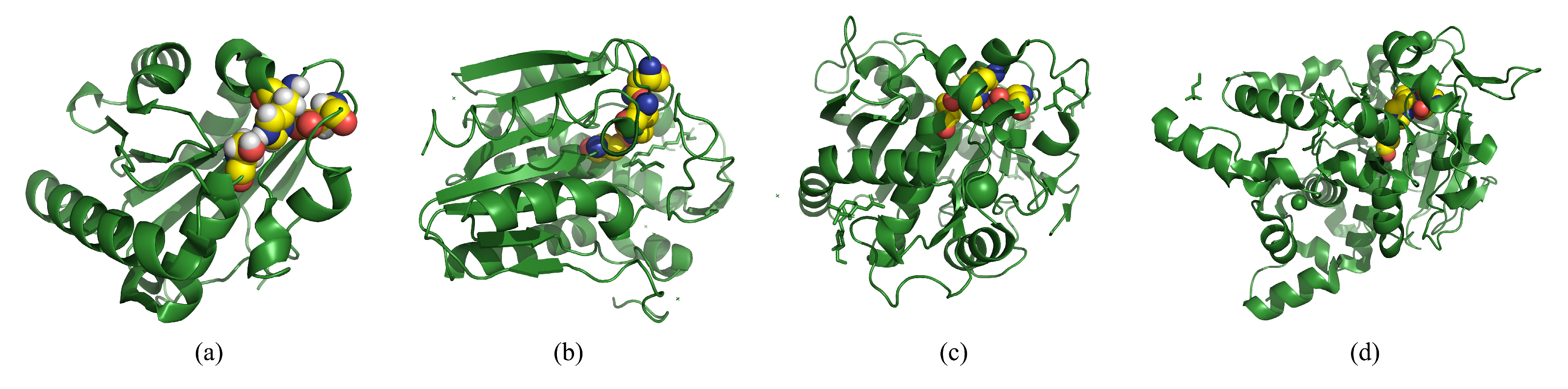
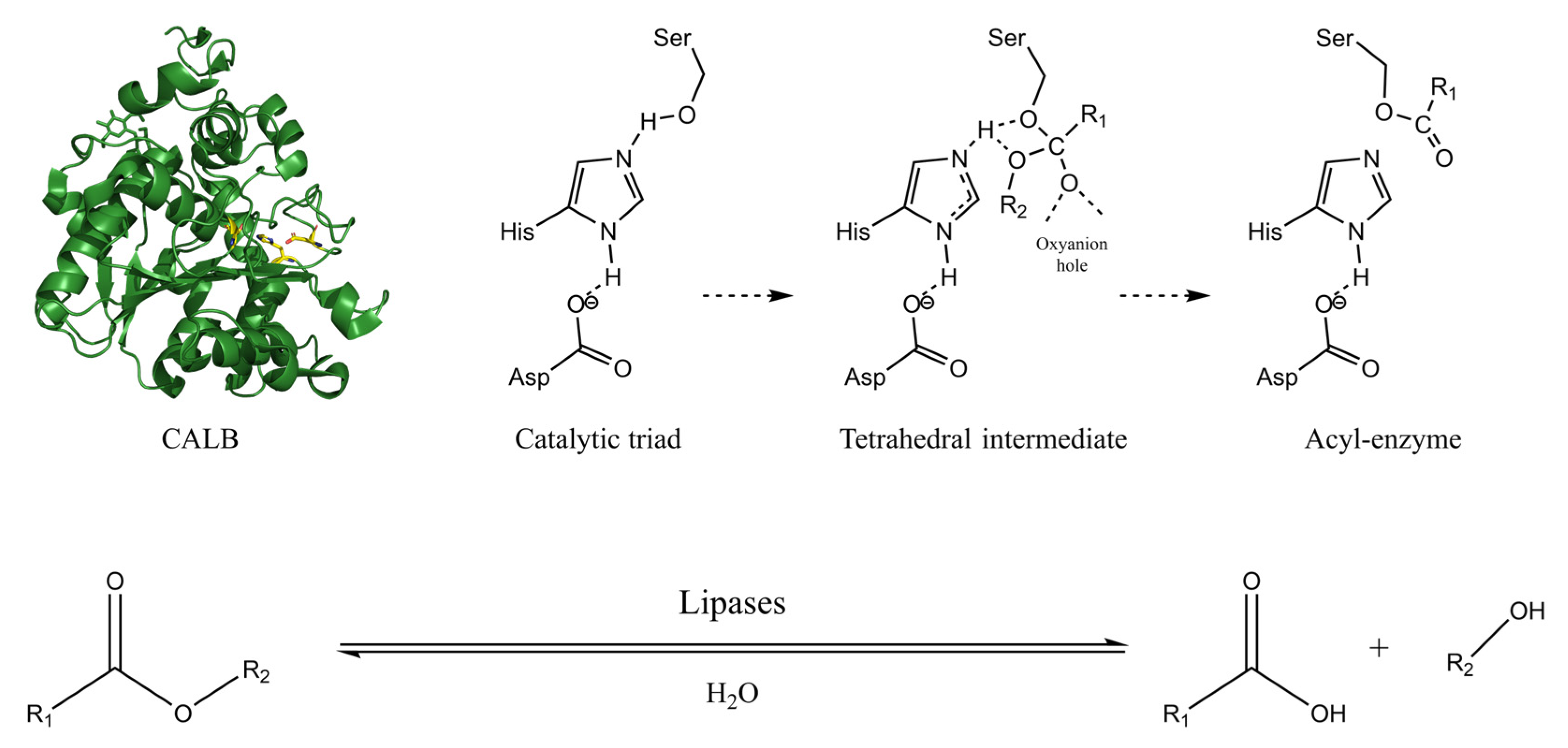
4.1. Lipase Catalyzed Hydrolysis: Structure and Mechanism
4.2. Mechanism of the Lipase-Catalyzed Polyester
4.3. Stability of Lipase in the Synthesis Reaction
5. Solvent System for Enzyme-Catalyzed Synthesis of Polyesters
5.1. Organic Solvent System
5.2. Solvent-Free System
5.3. Green Solvent System
6. The Challenges of Enzymatic Synthesis of Polyester
7. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Roumanet, P.J.; Laflèche, F.; Jarroux, N.; Raoul, Y.; Guégan, P. Novel aliphatic polyesters from an oleic acid based monomer. Synthesis, epoxidation, cross-linking and biodegradation. Eur. Polym. J. 2013, 49, 813–822. [Google Scholar] [CrossRef]
- Kolitz, M.; Cohen-Arazi, N.; Hagag, I.; Katzhendler, J.; Domb, A.J. Biodegradable Polyesters Derived from Amino Acids. Macromolecules 2009, 42, 4520–4530. [Google Scholar] [CrossRef]
- Gkountela, C.I.; Vouyiouka, S.N. Enzymatic polymerization as a green approach to synthesizing bio-based polyesters. Macromol 2022, 2, 30–57. [Google Scholar] [CrossRef]
- Jiang, Y.; Loos, K. Enzymatic synthesis of biobased polyesters and polyamides. Polymers 2016, 8, 243. [Google Scholar] [CrossRef] [PubMed]
- Darie-Niță, R.N.; Râpă, M.; Frąckowiak, S. Special Features of Polyester-Based Materials for Medical Applications. Polymers 2022, 14, 951. [Google Scholar] [CrossRef]
- Kai, D.; Zhang, K.; Jiang, L.; Wong, H.Z.; Li, Z.; Zhang, Z.; Loh, X.J. Sustainable and antioxidant lignin–polyester copolymers and nanofibers for potential healthcare applications. ACS Sustain. Chem. Eng. 2017, 5, 6016–6025. [Google Scholar] [CrossRef]
- Gross, R.A.; Ganesh, M.; Lu, W. Enzyme-catalysis breathes new life into polyester condensation polymerizations. Trends Biotechnol. 2010, 28, 435–443. [Google Scholar] [CrossRef]
- Fliedel, C.; Mameri, S.; Dagorne, S.; Aviles, T. Controlled ring-opening polymerization of trimethylene carbonate and access to PTMC-PLA block copolymers mediated by well-defined N-heterocyclic carbene zinc alkoxides. Appl. Organomet. Chem. 2014, 28, 504–511. [Google Scholar] [CrossRef]
- You, L.; Shen, Z.; Kong, J.; Ling, J. A novel approach to RE–OR bond from in situ reaction of rare earth triflates and sodium alkoxides: A versatile catalyst for living ring-opening polymerization of ε-caprolactone. Polymer 2014, 55, 2404–2410. [Google Scholar] [CrossRef]
- Moon, S.I.; Chan, W.L.; Miyamoto, M.; Kimura, Y. Melt polycondensation of L-lactic acid with Sn(II) catalysts activated by various proton acids: A direct manufacturing route to high molecular weight poly(L-lactic acid). J Polym Sci A Polym Chem. J. Polym. Sci. Part A Polym. Chem. 2015, 38, 1673–1679. [Google Scholar] [CrossRef]
- Simoni, R.D.; Hill, R.L.; Vaughan, M. Urease, the first crystalline enzyme and the proof that enzymes are proteins: The work of James B. Sumner. J. Biol. Chem. 2002, 277, e1–e2. [Google Scholar] [CrossRef]
- Kobayashi, S. Recent developments in lipase-catalyzed synthesis of polyesters. Macromol. Rapid Commun. 2009, 30, 237–266. [Google Scholar] [CrossRef]
- Jiang, Z. Lipase-catalyzed synthesis of aliphatic polyesters via copolymerization of lactide with diesters and diols. Polymer 2013, 54, 6105–6113. [Google Scholar] [CrossRef]
- Liu, C.; Wang, Y.; Liu, J.; Xu, J.; Zhang, R.; Wang, F.; Nie, K.; Deng, L. One-Step Synthesis of 4-Octyl Itaconate through the Structure Control of Lipase. J. Org. Chem. 2021, 86, 7895–7903. [Google Scholar] [CrossRef] [PubMed]
- Albertsson, A.C.; Srivastava, R.K. Recent developments in enzyme-catalyzed ring-opening polymerization. Adv. Drug Deliv. Rev. 2008, 60, 1077–1093. [Google Scholar] [CrossRef]
- Uyama, H.; Kobayashi, S. Enzymatic ring-opening polymerization of lactones catalyzed by lipase. Chem. Lett. 1993, 22, 1149–1150. [Google Scholar] [CrossRef]
- Knani, D.; Gutman, A.L.; Kohn, D.H. Enzymatic polyesterification in organic media. Enzyme-catalyzed synthesis of linear polyesters. I. Condensation polymerization of linear hydroxyesters. II. Ring-opening polymerization of ϵ-caprolactone. J. Polym. Sci. Part A: Polym. Chem. 1993, 31, 1221–1232. [Google Scholar] [CrossRef]
- Zhu, J.; An, S.; Lu, D.; Liu, Z. Development and prospects of enzymatic polymerization. Huagong Xuebao/CIESC J. 2013, 64, 407–414. [Google Scholar] [CrossRef]
- Matsumura, S.; Beppu, H.; Tsukada, K.; Toshima, K. Enzyme-catalyzed ring-opening polymerization of β-propiolactone. Biotechnol. Lett. 1996, 18, 1041–1046. [Google Scholar] [CrossRef]
- Champagne, E.; Strandman, S.; Zhu, X.X. Recent developments and optimization of lipase-catalyzed lactone formation and ring-opening polymerization. Macromol. Rapid Commun. 2016, 37, 1986–2004. [Google Scholar] [CrossRef]
- Yang, Y.; Yu, Y.; Zhang, Y.; Liu, C.; Shi, W.; Li, Q. Lipase/esterase-catalyzed ring-opening polymerization: A green polyester synthesis technique. Process Biochem. 2011, 46, 1900–1908. [Google Scholar] [CrossRef]
- van der Mee, L.; Helmich, F.; de Bruijn, R.; Vekemans, J.A.; Palmans, A.R.; Meijer, E. Investigation of lipase-catalyzed ring-opening polymerizations of lactones with various ring sizes: Kinetic evaluation. Macromolecules 2006, 39, 5021–5027. [Google Scholar] [CrossRef]
- Albertsson, A.-C.; Varma, I.K.; Srivastava, R.K. Polyesters from large lactones. In Handbook of Ring-Opening Polymerization; Wiley-VCH: Weinheim, Germany, 2009; pp. 287–306. [Google Scholar] [CrossRef]
- Manzini, B.; Hodge, P.; Ben-Haida, A. Entropically-driven ring-opening polymerization of macrocyclic esters with up to 84-membered rings catalysed by polymer-supported Candida antarctica lipase B. Polym. Chem. 2010, 1, 339–346. [Google Scholar] [CrossRef]
- Srivastava, R.K.; Albertsson, A.C. Porous scaffolds from high molecular weight polyesters synthesized via enzyme-catalyzed ring-opening polymerization. Biomacromolecules 2006, 7, 2531–2538. [Google Scholar] [CrossRef] [PubMed]
- Piotrowska, U.; Sobczak, M. Enzymatic Polymerization of Cyclic Monomers in Ionic Liquids as a Prospective Synthesis Method for Polyesters Used in Drug Delivery Systems. Molecules 2015, 20, 1–23. [Google Scholar] [CrossRef]
- Seyednejad, H.; Ghassemi, A.H.; van Nostrum, C.F.; Vermonden, T.; Hennink, W.E. Functional aliphatic polyesters for biomedical and pharmaceutical applications. J. Control. Release 2011, 152, 168–176. [Google Scholar] [CrossRef]
- Nampoothiri, K.M.; Nair, N.R.; John, R.P. An overview of the recent developments in polylactide (PLA) research. Bioresour. Technol. 2010, 101, 8493–8501. [Google Scholar] [CrossRef]
- Kataoka, M.; Miyakawa, T.; Shimizu, S.; Tanokura, M. Enzymes useful for chiral compound synthesis: Structural biology, directed evolution, and protein engineering for industrial use. Appl. Microbiol. Biotechnol. 2016, 100, 5747–5757. [Google Scholar] [CrossRef]
- Kobayashi, S. Enzymatic polymerization: A new method of polymer synthesis. J. Polym. Sci. Part A Polym. Chem. 1999, 37, 3041–3056. [Google Scholar] [CrossRef]
- Duchiron, S.W.; Pollet, E.; Givry, S.; Avérous, L. Enzymatic synthesis of amino acids endcapped polycaprolactone: A green route towards functional polyesters. Molecules 2018, 23, 290. [Google Scholar] [CrossRef]
- Gross, R.A.; Kumar, A.; Kalra, B. Polymer synthesis by in vitro enzyme catalysis. Chem. Rev. 2001, 101, 2097–2124. [Google Scholar] [CrossRef] [PubMed]
- Ajima, A.; Yoshimoto, T.; Takahashi, K.; Tamaura, Y.; Saito, Y.; Inada, Y. Polymerization of 10-hydroxydecanoic acid in benzene with polyethylene glycol-modified lipase. Biotechnol. Lett. 1985, 7, 303–306. [Google Scholar] [CrossRef]
- Okumura, S.; Iwai, M.; Tominaga, Y. Synthesis of ester oligomer by Aspergillus niger lipase. Agric. Biol. Chem. 1984, 48, 2805–2808. [Google Scholar] [CrossRef]
- Ohara, H.; Onogi, A.; Yamamoto, M.; Kobayashi, S. Lipase-catalyzed oligomerization and hydrolysis of alkyl lactates: Direct evidence in the catalysis mechanism that enantioselection is governed by a deacylation step. Biomacromolecules 2010, 11, 2008–2015. [Google Scholar] [CrossRef]
- Mahapatro, A.; Kumar, A.; Kalra, B.; Gross, R.A. Solvent-Free Adipic Acid/1, 8-Octanediol Condensation Polymerizations Catalyzed by Candida a ntartica Lipase B. Macromolecules 2004, 37, 35–40. [Google Scholar] [CrossRef]
- Mahapatro, A.; Kalra, B.; Kumar, A.; Gross, R.A. Lipase-catalyzed polycondensations: Effect of substrates and solvent on chain formation, dispersity, and end-group structure. Biomacromolecules 2003, 4, 544–551. [Google Scholar] [CrossRef]
- Arrieta-Baez, D.; Hernández Ortíz, J.V.; Campos Terán, J.; Torres, E.; Gómez-Patiño, M.B. Aliphatic diacidic long-chain C16 polyesters from 10, 16-Dihydroxyhexadecanoic acid obtained from tomato residual wastes. Molecules 2019, 24, 1524. [Google Scholar] [CrossRef]
- Douka, A.; Vouyiouka, S.; Papaspyridi, L.-M.; Papaspyrides, C.D. A review on enzymatic polymerization to produce polycondensation polymers: The case of aliphatic polyesters, polyamides and polyesteramides. Prog. Polym. Sci. 2018, 79, 1–25. [Google Scholar] [CrossRef]
- Fabbri, F.; Bertolini, F.A.; Guebitz, G.M.; Pellis, A. Biocatalyzed Synthesis of Flavor Esters and Polyesters: A Design of Experiments (DoE) Approach. Int. J. Mol. Sci. 2021, 22, 8493. [Google Scholar] [CrossRef]
- Yagihara, T.; Matsumura, S. Enzymatic synthesis and chemical recycling of novel polyester-type thermoplastic elastomers. Polymers 2012, 4, 1259–1277. [Google Scholar] [CrossRef]
- Bazin, A.; Avérous, L.; Pollet, E. Ferulic Acid as Building Block for the Lipase-Catalyzed Synthesis of Biobased Aromatic Polyesters. Polymers 2021, 13, 3693. [Google Scholar] [CrossRef]
- Bilal, M.H.; Prehm, M.; Njau, A.E.; Samiullah, M.H.; Meister, A.; Kressler, J. Enzymatic synthesis and characterization of hydrophilic sugar based polyesters and their modification with stearic acid. Polymers 2016, 8, 80. [Google Scholar] [CrossRef]
- Azim, H.; Dekhterman, A.; Jiang, Z.; Gross, R.A. Candida a ntarctica Lipase B-Catalyzed Synthesis of Poly (butylene succinate): Shorter Chain Building Blocks Also Work. Biomacromolecules 2006, 7, 3093–3097. [Google Scholar] [CrossRef]
- Jiang, Y.; Woortman, A.J.; Alberda van Ekenstein, G.O.; Loos, K. Enzyme-catalyzed synthesis of unsaturated aliphatic polyesters based on green monomers from renewable resources. Biomolecules 2013, 3, 461–480. [Google Scholar] [CrossRef]
- Lang, K.; Bhattacharya, S.; Ning, Z.; Sánchez-Leija, R.J.; Bramson, M.T.; Centore, R.; Corr, D.T.; Linhardt, R.J.; Gross, R.A. Enzymatic polymerization of poly (glycerol-1, 8-octanediol-sebacate): Versatile poly (glycerol sebacate) analogues that form monocomponent biodegradable fiber scaffolds. Biomacromolecules 2020, 21, 3197–3206. [Google Scholar] [CrossRef]
- Villavicencio, J.; Orozco, F.; Benitez, R.; Martin, J.; Rojas, G. A Greener Enzymatic Oligoesterification of Biobased Renewable Synthons. J. Braz. Chem. Soc. 2021, 32, 1294–1299. [Google Scholar] [CrossRef]
- Gustini, L.; Lavilla, C.; Finzel, L.; Noordover, B.A.; Hendrix, M.M.; Koning, C.E. Sustainable coatings from bio-based, enzymatically synthesized polyesters with enhanced functionalities. Polym. Chem. 2016, 7, 6586–6597. [Google Scholar] [CrossRef]
- Lang, K.; Sánchez-Leija, R.J.; Gross, R.A.; Linhardt, R.J. Review on the impact of polyols on the properties of bio-based polyesters. Polymers 2020, 12, 2969. [Google Scholar] [CrossRef]
- Valerio, O.; Misra, M.; Mohanty, A.K. Poly (glycerol-co-diacids) polyesters: From glycerol biorefinery to sustainable engineering applications, a review. ACS Sustain. Chem. Eng. 2018, 6, 5681–5693. [Google Scholar] [CrossRef]
- Kulshrestha, A.S.; Gao, W.; Gross, R.A. Glycerol copolyesters: Control of branching and molecular weight using a lipase catalyst. Macromolecules 2005, 38, 3193–3204. [Google Scholar] [CrossRef]
- Taresco, V.; Creasey, R.; Kennon, J.; Mantovani, G.; Alexander, C.; Burley, J.C.; Garnett, M.C. Variation in structure and properties of poly (glycerol adipate) via control of chain branching during enzymatic synthesis. Polymer 2016, 89, 41–49. [Google Scholar] [CrossRef]
- Perin, G.B.; Felisberti, M.I. Enzymatic synthesis of poly (glycerol sebacate): Kinetics, chain growth, and branching behavior. Macromolecules 2020, 53, 7925–7935. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.T.; Leferink, N.G.; Scrutton, N.S. Chemo-enzymatic routes towards the synthesis of bio-based monomers and polymers. Mol. Catal. 2019, 467, 95–110. [Google Scholar] [CrossRef]
- Kobayashi, S.; Uyama, H. Enzymatic polymerization of cyclic acid anhydrides and glycols by a lipase catalyst. Die Makromol. Chem. Rapid Commun. 1993, 14, 841–844. [Google Scholar] [CrossRef]
- Kobayashi, S.; Uyama, H. Synthesis of Polyesters I: Hydrolase as Catalyst for Polycondensation (Condensation Polymerization). In Enzymatic Polymerization towards Green Polymer Chemistry; Kobayashi, S., Uyama, H., Kadokawa, J.-I., Eds.; Springer: Singapore, 2019; pp. 105–163. [Google Scholar] [CrossRef]
- Debuissy, T.; Pollet, E.; Avérous, L. Biotic and abiotic synthesis of renewable aliphatic polyesters from short building blocks obtained from biotechnology. ChemSusChem 2018, 11, 3836–3870. [Google Scholar] [CrossRef]
- Jiang, Z. Lipase-catalyzed synthesis of aliphatic polyesters via copolymerization of lactone, dialkyl diester, and diol. Biomacromolecules 2008, 9, 3246–3251. [Google Scholar] [CrossRef]
- Uyama, H.; Kobayashi, S. Enzyme-catalyzed polymerization to functional polymers. J. Mol. Catal. B Enzym. 2002, 19, 117–127. [Google Scholar] [CrossRef]
- Kobayashi, S. Lipase-catalyzed polyester synthesis–a green polymer chemistry. Proc. Japan Acad. Ser. B 2010, 86, 338–365. [Google Scholar] [CrossRef]
- Liu, Y.; Song, L.; Feng, N.; Jiang, W.; Jin, Y.; Li, X. Recent advances in the synthesis of biodegradable polyesters by sustainable polymerization: Lipase-catalyzed polymerization. RSC Adv. 2020, 10, 36230–36240. [Google Scholar] [CrossRef]
- Weinberger, S.; Pellis, A.; Comerford, J.W.; Farmer, T.J.; Guebitz, G.M. Efficient physisorption of Candida Antarctica Lipase B on Polypropylene Beads and application for polyester synthesis. Catalysts 2018, 8, 369. [Google Scholar] [CrossRef]
- Magnusson, A.O.; Rotticci-Mulder, J.C.; Santagostino, A.; Hult, K. Creating space for large secondary alcohols by rational redesign of Candida antarctica lipase B. ChemBioChem 2005, 6, 1051–1056. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Gross, R.A. Candida antartica Lipase B Catalyzed Polycaprolactone Synthesis: Effects of Organic Media and Temperature. Biomacromolecules 2000, 1, 133–138. [Google Scholar] [CrossRef]
- Poojari, Y.; Clarson, S.J. Thermal stability of Candida antarctica lipase B immobilized on macroporous acrylic resin particles in organic media. Biocatal. Agric. Biotechnol. 2013, 2, 7–11. [Google Scholar] [CrossRef]
- Kundys, A.; Białecka-Florjańczyk, E.; Fabiszewska, A.; Małajowicz, J. Candida antarctica lipase B as catalyst for cyclic esters synthesis, their polymerization and degradation of aliphatic polyesters. J. Polym. Environ. 2018, 26, 396–407. [Google Scholar] [CrossRef]
- Reis, P.; Holmberg, K.; Watzke, H.; Leser, M.E.; Miller, R. Lipases at interfaces: A review. Adv. Colloid Interface Sci. 2009, 147, 237–250. [Google Scholar] [CrossRef] [PubMed]
- Brocca, S.; Secundo, F.; Ossola, M.; Alberghina, L.; Carrea, G.; Lotti, M. Sequence of the lid affects activity and specificity of Candida rugosa lipase isoenzymes. Protein Sci. 2009, 12, 2312–2319. [Google Scholar] [CrossRef] [PubMed]
- Khan, F.I.; Lan, D.; Durrani, R.; Huan, W.; Zhao, Z.; Wang, Y. The lid domain in lipases: Structural and functional determinant of enzymatic properties. Front. Bioeng. Biotechnol. 2017, 5, 16. [Google Scholar] [CrossRef]
- Xu, T.; Liu, L.; Hou, S.; Xu, J.; Yang, B.; Wang, Y.; Liu, J. Crystal structure of a mono- and diacylglycerol lipase from Malassezia globosa reveals a novel lid conformation and insights into the substrate specificity. J. Struct. Biol. 2012, 178, 363–369. [Google Scholar] [CrossRef]
- Brzozowski, A.M.; Derewenda, Z.S.; Dodson, E.J.; Dodson, G.G.; Turkenburg, J.P. Structure and molecular model refinement of Rhizomucor miehei triacyglyceride lipase: A case study of the use of simulated annealing in partial model refinement. Acta Crystallogr. Sect. B Struct. Sci. 1992, 48, 307–319. [Google Scholar] [CrossRef]
- Matsumura, H.; Yamamoto, T.; Leow, T.C.; Mori, T.; Salleh, A.B.; Basri, M.; Inoue, T.; Kai, Y.; Rahman, R.N.Z.R.A. Novel cation-π interaction revealed by crystal structure of thermoalkalophilic lipase. Proteins Struct. Funct. Bioinform. 2008, 70, 592–598. [Google Scholar] [CrossRef]
- Nordwald, E.M.; Plaks, J.G.; Snell, J.R.; Sousa, M.C.; Kaar, J.L. Crystallographic investigation of imidazolium ionic liquid effects on enzyme structure. ChemBioChem 2015, 16, 2456–2459. [Google Scholar] [CrossRef]
- Rengachari, S.; Aschauer, P.; Schittmayer, M.; Mayer, N.; Gruber, K.; Breinbauer, R.; Birner-Gruenberger, R.; Dreveny, I.; Oberer, M. Conformational plasticity and ligand binding of bacterial monoacylglycerol lipase. J. Biol. Chem. 2013, 288, 31093–31104. [Google Scholar] [CrossRef]
- Bordes, F.; Barbe, S.; Escalier, P.; Mourey, L.; André, I.; Marty, A.; Tranier, S. Exploring the conformational states and rearrangements of Yarrowia lipolytica lipase. Biophys. J. 2010, 99, 2225–2234. [Google Scholar] [CrossRef]
- Carrasco-López, C.; Godoy, C.; de Las Rivas, B.; Fernández-Lorente, G.; Palomo, J.M.; Guisán, J.M.; Fernández-Lafuente, R.; Martínez-Ripoll, M.; Hermoso, J.A. Activation of bacterial thermoalkalophilic lipases is spurred by dramatic structural rearrangements. J. Biol. Chem. 2009, 284, 4365–4372. [Google Scholar] [CrossRef]
- Hæffner, F.; Norin, T.; Hult, K. Molecular modeling of the enantioselectivity in lipase-catalyzed transesterification reactions. Biophys. J. 1998, 74, 1251–1262. [Google Scholar] [CrossRef]
- Li, C.; Tan, T.; Zhang, H.; Feng, W. Analysis of the conformational stability and activity of Candida antarctica lipase B in organic solvents: Insight from molecular dynamics and quantum mechanics/simulations. J. Biol. Chem. 2010, 285, 28434–28441. [Google Scholar] [CrossRef]
- Jaeger, K.E.; Dijkstra, B.W.; Reetz, M.T. Bacterial Biocatalysts: Molecular Biology, Three-Dimensional Structures, and Biotechnological Applications of Lipases. Annu. Rev. Microbiol. 1999, 53, 315–351. [Google Scholar] [CrossRef]
- Strzelczyk, P.; Bujacz, G.D.; Kiełbasiński, P.; Błaszczyk, J. Crystal and molecular structure of hexagonal form of lipase B from Candida antarctica. Acta Biochim. Pol. 2016, 63, 103–109. [Google Scholar] [CrossRef]
- Yang, J.; Liu, Y.; Liang, X.; Yang, Y.; Li, Q. Enantio-, Regio-, and Chemoselective Lipase-Catalyzed Polymer Synthesis. Macromol. Biosci. 2018, 18, 1800131. [Google Scholar] [CrossRef]
- Hevilla, V.; Sonseca, A.; Echeverría, C.; Muñoz-Bonilla, A.; Fernández-García, M. Enzymatic Synthesis of Polyesters and Their Bioapplications: Recent Advances and Perspectives. Macromol. Biosci. 2021, 21, e2100156. [Google Scholar] [CrossRef]
- Binns, F.; Roberts, S.M.; Taylor, A.; Williams, C.F. Enzymic polymerisation of an unactivated diol/diacid system. J. Chem. Soc. Perkin Trans. 1 1993, 899–904. [Google Scholar] [CrossRef]
- Chaplin, M. Do we underestimate the importance of water in cell biology? Nat. Rev. Mol. Cell Biol. 2006, 7, 861–866. [Google Scholar] [CrossRef]
- Zaks, A.; Klibanov, A.M. The effect of water on enzyme action in organic media. J. Biol. Chem. 1988, 263, 8017–8021. [Google Scholar] [CrossRef]
- Zaks, A.; Klibanov, A.M. Enzymatic catalysis in nonaqueous solvents. J. Biol. Chem. 1988, 263, 3194–3201. [Google Scholar] [CrossRef]
- Klibanov, A.M. Asymmetric transformations catalyzed by enzymes in organic solvents. Acc. Chem. Res. 1990, 23, 114–120. [Google Scholar] [CrossRef]
- Nawani, N.; Dosanjh, N.S.; Kaur, J. A novel thermostable lipase from a thermophilic Bacillus sp.: Characterization and esterification studies. Biotechnol. Lett. 1998, 20, 997–1000. [Google Scholar] [CrossRef]
- Kumar, A.; Dhar, K.; Kanwar, S.S.; Arora, P.K. Lipase catalysis in organic solvents: Advantages and applications. Biol. Proced. Online 2016, 18, 2. [Google Scholar] [CrossRef]
- Kamal, M.Z.; Yedavalli, P.; Deshmukh, M.V.; Rao, N.M. Lipase in aqueous-polar organic solvents: Activity, structure, and stability. Protein Sci. 2013, 22, 904–915. [Google Scholar] [CrossRef]
- Trodler, P.; Pleiss, J. Modeling structure and flexibility of Candida antarctica lipase B in organic solvents. BMC Struct. Biol. 2008, 8, 9. [Google Scholar] [CrossRef]
- Norin, M.; Haeffner, F.; Hult, K.; Edholm, O. Molecular dynamics simulations of an enzyme surrounded by vacuum, water, or a hydrophobic solvent. Biophys. J. 1994, 67, 548–559. [Google Scholar] [CrossRef]
- Rehm, S.; Trodler, P.; Pleiss, J. Solvent-induced lid opening in lipases: A molecular dynamics study. Protein Sci. 2010, 19, 2122–2130. [Google Scholar] [CrossRef] [PubMed]
- van Tol, J.B.A.; Stevens, R.M.; Veldhuizen, W.J.; Jongejan, J.A.; Duine, J.A. Do organic solvents affect the catalytic properties of lipase? Intrinsic kinetic parameters of lipases in ester hydrolysis and formation in various organic solvents. Biotechnol. Bioeng. 1995, 47, 71–81. [Google Scholar] [CrossRef]
- Gotor-Fernández, V.; Brieva, R.; Gotor, V. Lipases: Useful biocatalysts for the preparation of pharmaceuticals. J. Mol. Catal. B Enzym. 2006, 40, 111–120. [Google Scholar] [CrossRef]
- Sharma, S.; Kanwar, S.S. Organic solvent tolerant lipases and applications. Sci. World J. 2014, 2014, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Priyanka, P.; Tan, Y.; Kinsella, G.K.; Henehan, G.T.; Ryan, B.J. Solvent stable microbial lipases: Current understanding and biotechnological applications. Biotechnol. Lett. 2019, 41, 203–220. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, T.; Yamada, R.; Ogino, H. Chemical treatments for modification and immobilization to improve the solvent-stability of lipase. World J. Microbiol. Biotechnol. 2019, 35, 1–8. [Google Scholar] [CrossRef]
- Cavalcante, F.T.T.; Neto, F.S.; de Aguiar Falcão, I.R.; da Silva Souza, J.E.; de Moura Junior, L.S.; da Silva Sousa, P.; Rocha, T.G.; de Sousa, I.G.; de Lima Gomes, P.H.; de Souza, M.C.M. Opportunities for improving biodiesel production via lipase catalysis. Fuel 2021, 288, 119577. [Google Scholar] [CrossRef]
- Priyanka, P.; Kinsella, G.K.; Henehan, G.T.; Ryan, B.J. Listeria monocytogenes is a solvent tolerant organism secreting a solvent stable lipase: Potential biotechnological applications. Biotechnol. Lett. 2022, 44, 1–9. [Google Scholar] [CrossRef]
- Ali, Y.; Ahmad, B.; Alghamdi, K.M.; Kamal, T.; Ali, H.; Anwar, Y.; Hussain, A.; Jogezai, N.U. Characterization of recombinant cold active lipase from a novel Pseudomonas spp. MG687270. Int. J. Agric. Biol. 2019, 22, 855–865. [Google Scholar] [CrossRef]
- Somero, G.N. Proteins and temperature. Annu. Rev. Physiol. 1995, 57, 43–68. [Google Scholar] [CrossRef]
- Pace, C.; Hermans, J. The stability of globular protein. CRC Crit. Rev. Biochem. 1975, 3, 1–43. [Google Scholar] [CrossRef] [PubMed]
- Bhanuramanand, K.; Ahmad, S.; Rao, N. Engineering deamidation-susceptible asparagines leads to improved stability to thermal cycling in a lipase. Protein Sci. 2014, 23, 1479–1490. [Google Scholar] [CrossRef] [PubMed]
- Mansfeld, J.; Ulbrich-Hofmann, R. The stability of engineered thermostable neutral proteases from Bacillus stearothermophilus in organic solvents and detergents. Biotechnol. Bioeng. 2007, 97, 672–679. [Google Scholar] [CrossRef] [PubMed]
- Hamdan, S.H.; Maiangwa, J.; Ali, M.S.M.; Normi, Y.M.; Sabri, S.; Leow, T.C. Thermostable lipases and their dynamics of improved enzymatic properties. Appl. Microbiol. Biotechnol. 2021, 105, 7069–7094. [Google Scholar] [CrossRef] [PubMed]
- Wyss, M.; Pasamontes, L.; Rémy, R.; Kohler, J.; Kusznir, E.; Gadient, M.; Müller, F.; van Loon, A.P. Comparison of the thermostability properties of three acid phosphatases from molds: Aspergillus fumigatus phytase, A. niger phytase, and A. niger pH 2.5 acid phosphatase. Appl. Environ. Microbiol. 1998, 64, 4446–4451. [Google Scholar] [CrossRef] [PubMed]
- Castro-Ochoa, L.D.; Rodríguez-Gómez, C.; Valerio-Alfaro, G.; Ros, R.O. Screening, purification and characterization of the thermoalkalophilic lipase produced by Bacillus thermoleovorans CCR11. Enzym. Microb. Technol. 2005, 37, 648–654. [Google Scholar] [CrossRef]
- Kamal, M.Z.; Ahmad, S.; Molugu, T.R.; Vijayalakshmi, A.; Deshmukh, M.V.; Sankaranarayanan, R.; Rao, N.M. In vitro evolved non-aggregating and thermostable lipase: Structural and thermodynamic investigation. J. Mol. Biol. 2011, 413, 726–741. [Google Scholar] [CrossRef]
- Castilla, A.; Panizza, P.; Rodríguez, D.; Bonino, L.; Giordano, S.R. A novel thermophilic and halophilic esterase from Janibacter sp. R02, the first member of a new lipase family (Family XVII). Enzym. Microb. Technol. 2016, 98, 86–95. [Google Scholar] [CrossRef]
- Szczęsna-Antczak, M.; Kamińska, J.; Florczak, T.; Turkiewicz, M. Cold-active yeast lipases: Recent issues and future prospects. In Cold-Adapted Yeasts; Springer: Berlin/Heidelberg, Germany, 2014; pp. 353–375. [Google Scholar] [CrossRef]
- Cai, H.; Zhang, T.; Dumba, T.; Zhao, M.; Li, Y.; Tong, Z.; Mao, J.; Zhang, H.; Feng, F. High-efficient Expression and Properties Characterization of Thermomyces lanuginosus Lipase in Pichia pastoris. J. Chin. Inst. Food Sci. Technol. 2018, 18, 262–271. [Google Scholar] [CrossRef]
- Zhang, J.; Tian, M.; Lv, P.; Luo, W.; Wang, Z.; Xu, J.; Wang, Z. High-efficiency expression of the thermophilic lipase from Geobacillus thermocatenulatus in Escherichia coli and its application in the enzymatic hydrolysis of rapeseed oil. 3 Biotech 2020, 10, 1–9. [Google Scholar] [CrossRef]
- Fang, Y.; Zhou, Y.; Xin, Y.; Shi, Y.; Guo, Z.; Li, Y.; Gu, Z.; Ding, Z.; Shi, G.; Zhang, L. Preparation and characterization of a novel thermostable lipase from Thermomicrobium roseum. Catal. Sci. Technol. 2021, 11, 7386–7397. [Google Scholar] [CrossRef]
- Chaudhary, A.K.; Kline, B.J.; Beckman, E.J.; Russell, A.J. Biocatalytic solvent-free polymerization to produce high molecular weight polyesters. Am. Chem. Soc. Polym. Prepr. Div. Polym. Chem. 1997, 38, 396–397. [Google Scholar] [CrossRef]
- Sheldon, R.A. Green solvents for sustainable organic synthesis: State of the art. Green Chem. 2005, 7, 267–278. [Google Scholar] [CrossRef]
- Linko, Y.Y.; Lms, M.; Wu, X.; Uosukainen, E.; Seppl, J.; Linko, P. Biodegradable products by lipase biocatalysis. J. Biotechnol. 1998, 66, 41. [Google Scholar] [CrossRef]
- Hun, C.J.; Rahman, R.; Salleh, A.B.; Basri, M. A newly isolated organic solvent tolerant Bacillus sphaericus 205y producing organic solvent-stable lipase. Biochem. Eng. J. 2003, 15, 147–151. [Google Scholar] [CrossRef]
- Lee, H.-J.; Kang, Y.-S.; Kim, C.-Y.; Seo, E.-J.; Pyo, S.-H. Multi-Step Enzymatic Synthesis of 1,9-Nonanedioic Acid from a Renewable Fatty Acid and Its Application for the Enzymatic Production of Biopolyesters. Polymers 2019, 11, 1690. [Google Scholar] [CrossRef] [PubMed]
- Simon, L.M.; László, K.; Vértesi, A.; Bagi, K.; Szajáni, B. Stability of hydrolytic enzymes in water-organic solvent systems. J. Mol. Catal. B Enzym. 1998, 4, 41–45. [Google Scholar] [CrossRef]
- Debuissy, T.; Pollet, E.; Avérous, L. Enzymatic synthesis of biobased poly(1,4-butylene succinate-ran-2,3-butylene succinate) copolyesters and characterization. Influence of 1,4- and 2,3-butanediol contents. Eur. Polym. J. 2017, 93, 103–115. [Google Scholar] [CrossRef]
- Inoue, A.; Horikoshi, K. Estimation of Solvent-Tolerance of Bacteria by the Solvent Parameter Log-P. J. Ferment. Bioeng. 1991, 71, 194–196. [Google Scholar] [CrossRef]
- Iemhoff, A.; Sherwood, J.; McElroy, C.R.; Hunt, A.J. Towards sustainable kinetic resolution, a combination of bio-catalysis, flow chemistry and bio-based solvents. Green Chem. 2018, 20, 136–140. [Google Scholar] [CrossRef]
- Sprunger, L.M.; Proctor, A.; Acree, W.E.; Abraham, M.H.; Benjelloun-Dakhama, N. Correlation and prediction of partition coefficient between the gas phase and water, and the solvents dry methyl acetate, dry and wet ethyl acetate, and dry and wet butyl acetate. Fluid Phase Equilibria 2008, 270, 30–44. [Google Scholar] [CrossRef]
- Cruz-Izquierdo, Á.; Van, D.; Serra, J.L.; Llama, M.J.; Boeriu, C.G. Lipase-catalyzed synthesis of oligoesters of 2,5-furandicarboxylic acid with aliphatic diols. Pure Appl. Chem. 2015, 87, 59–69. [Google Scholar] [CrossRef]
- Combes, J.; Clavijo, E.; Clément, T.; Fojcik, C.; Allais, F. Solvent selection strategy for an ISPR (In Situ/In Stream Product Recovery) process: The case of microbial production of p-coumaric acid coupled with a liquid-liquid extraction. Sep. Purif. Technol. 2020, 259, 118170. [Google Scholar] [CrossRef]
- Saeed, N.H.M.; Abbas, A.M. Kinetics and mechanism of tetrahydrofuran oxidation by chloraminet in acidic media. Period. Tche Quim. 2020, 17, 449–461. [Google Scholar] [CrossRef]
- Völgyi, G.; Deák, K.; Vámos, J.; Valkó, K.; Takács-Novák, K. RPTLC determination of log P of structurally diverse neutral compounds. J. Planar Chromatogr. Modern TLC 2008, 21, 143–149. [Google Scholar] [CrossRef]
- Yao, D.; Li, G.; Kuila, T.; Peng, L.; Kim, N.H.; Kim, S.I.; Lee, J.H. Lipase-catalyzed synthesis and characterization of biodegradable polyester containing l-malic acid unit in solvent system. J. Appl. Polym. Sci. 2011, 120, 1114–1120. [Google Scholar] [CrossRef]
- Juais, D.; Naves, A.F.; Li, C.; Gross, R.A.; Catalani, L.H. Isosorbide Polyesters from Enzymatic Catalysis. Macromolecules 2010, 43, 10315–10319. [Google Scholar] [CrossRef]
- Linko, Y.Y.; Wang, Z.L.; Seppaelae, J. Lipase-catalyzed synthesis of poly(1,4-butyl sebacate) from sebacic acid or its derivatives with 1,4-butanediol. J. Biotechnol. 1995, 40, 133–138. [Google Scholar] [CrossRef]
- Feder, D.; Gross, R.A. Exploring chain length selectivity in HIC-catalyzed polycondensation reactions. Biomacromolecules 2010, 11, 690. [Google Scholar] [CrossRef]
- Yang, Y.; Lu, W.; Zhang, X.; Xie, W.; Cai, M.; Gross, R.A. Two-step biocatalytic route to biobased functional polyesters from omega-carboxy fatty acids and diols. Biomacromolecules 2010, 11, 259–268. [Google Scholar] [CrossRef]
- Pellis, A.; Byrne, F.P.; Sherwood, J.; Vastano, M.; Comerford, J.W.; Farmer, T.J. Safer bio-based solvents to replace toluene and tetrahydrofuran for the biocatalyzed synthesis of polyesters. Green Chem. 2019, 21, 1686–1694. [Google Scholar] [CrossRef] [PubMed]
- Pellis, A.; Weinberger, S.; Gigli, M.; Guebitz, G.M.; Farmer, T. Enzymatic synthesis of biobased polyesters utilizing aromatic diols as the rigid component. Eur. Polym. J. 2020, 130, 109680. [Google Scholar] [CrossRef]
- Stebbins, N.D.; Yu, W.; Uhrich, K.E. Enzymatic Polymerization of an Ibuprofen-Containing Monomer and Subsequent Drug Release. Macromol. Biosci. 2015, 15, 1115–1124. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Uyama, H.; Wada, S.; Fukui, T.; Kobayashi, S. Lipase-catalyzed synthesis of polyesters from anhydride derivatives involving dehydration. Biochem. Eng. J. 2003, 16, 145–152. [Google Scholar] [CrossRef]
- Jiang, Y.; Woortman, A.J.J.; Alberda van Ekenstein, G.O.R.; Loos, K. Environmentally benign synthesis of saturated and unsaturated aliphatic polyesters via enzymatic polymerization of biobased monomers derived from renewable resources. Polym. Chem. 2015, 6, 5451–5463. [Google Scholar] [CrossRef]
- Nasr, K.; Favrelle-Huret, A.; Mincheva, R.; Stoclet, G.; Bria, M.; Raquez, J.-M.; Zinck, P. The Impact of Diethyl Furan-2, 5-dicarboxylate as an Aromatic Biobased Monomer toward Lipase-Catalyzed Synthesis of Semiaromatic Copolyesters. ACS Appl. Polym. Mater. 2022, 4, 1387–1400. [Google Scholar] [CrossRef]
- Sokołowska, M.; Nowak-Grzebyta, J.; Stachowska, E.; El Fray, M. Enzymatic Catalysis in Favor of Blocky Structure and Higher Crystallinity of Poly (Butylene Succinate)-Co-(Dilinoleic Succinate)(PBS-DLS) Copolymers of Variable Segmental Composition. Materials 2022, 15, 1132. [Google Scholar] [CrossRef]
- Pfluck, A.C.; de Barros, D.P.; Oliva, A.; Fonseca, L.P. Enzymatic Poly (octamethylene suberate) Synthesis by a Two-Step Polymerization Method Based on the New Greener Polymer-5B Technology. Processes 2022, 10, 221. [Google Scholar] [CrossRef]
- Adhami, W.; Bakkour, Y.; Rolando, C. Polylactones synthesis by enzymatic ring opening polymerization in flow. Polymer 2021, 230, 124040. [Google Scholar] [CrossRef]
- Maniar, D.; Fodor, C.; Karno Adi, I.; Woortman, A.J.; van Dijken, J.; Loos, K. Enzymatic synthesis of muconic acid-based polymers: Trans, trans-dimethyl muconate and trans, β-dimethyl hydromuconate. Polymers 2021, 13, 2498. [Google Scholar] [CrossRef]
- Esquivel Guzmán, J.A.; Zhang, H.; Rivera, E.; Lavertu, M.; Zhu, X.-X. Porphyrin-Based Polyesters Synthesized by Enzymatic Catalysis. ACS Appl. Polym. Mater. 2021, 3, 3659–3665. [Google Scholar] [CrossRef]
- Avila-Ortiz, C.G.; Juaristi, E. Novel Methodologies for Chemical Activation in Organic Synthesis under Solvent-Free Reaction Conditions. Molecules 2020, 25, 3579. [Google Scholar] [CrossRef] [PubMed]
- Zangade, S.; Patil, P. A review on solvent-free methods in organic synthesis. Curr. Org. Chem. 2019, 23, 2295–2318. [Google Scholar] [CrossRef]
- Loupy, A. Solvent-free reactions. In Modern Solvents in Organic Synthesis; Springer: Berlin/Heidelberg, Germany, 1999; pp. 153–207. [Google Scholar] [CrossRef]
- Zhao, H. Enzymatic ring-opening polymerization (ROP) of polylactones: Roles of non-aqueous solvents. J. Chem. Technol. Biotechnol. 2018, 93, 9–19. [Google Scholar] [CrossRef]
- Kato, M.; Toshima, K.; Matsumura, S. Direct enzymatic synthesis of a polyester with free pendant mercapto groups. Biomacromolecules 2009, 10, 366–373. [Google Scholar] [CrossRef]
- Campisano, I.; Erika, D.; Cláudia, D.; Dias, M.L.; Castro, A.D.; Langone, M. Solvent-free lipase-catalyzed synthesis of linear and thermally stable polyesters obtained from diacids and diols. Braz. J. Chem. Eng. 2021, 38, 549–562. [Google Scholar] [CrossRef]
- Zhao, X.; Li, Y.; Fu, J.; Wang, H.; Hong, J. Insight into the in-situ solvent-free lipase-catalyzed coating on cotton with polyesters. Process Biochem. 2021, 102, 82–91. [Google Scholar] [CrossRef]
- Nguyen, H.D.; Löf, D.; Hvilsted, S.; Daugaard, A.E. Highly branched bio-based unsaturated polyesters by enzymatic polymerization. Polymers 2016, 8, 363. [Google Scholar] [CrossRef]
- Fehling, E.; Bergander, K.; Klein, E.; Weber, N.; Vosmann, K. Thiol-functionalized copolymeric polyesters by lipase-catalyzed esterification and transesterification of 1,12-dodecanedioic acid and its diethyl ester, respectively, with 1-thioglycerol. Biotechnol. Lett. 2010, 32, 1463–1471. [Google Scholar] [CrossRef]
- Yang, Y.; Lu, W.; Cai, J.; Hou, Y.; Ouyang, S.; Xie, W.; Gross, R.A. Poly (oleic diacid-co-glycerol): Comparison of polymer structure resulting from chemical and lipase catalysis. Macromolecules 2011, 44, 1977–1985. [Google Scholar] [CrossRef]
- Alessandro, P.; Comerford, J.W.; Maneffa, A.J.; Sipponen, M.H.; Clark, J.H.; Farmer, T.J. Elucidating enzymatic polymerisations: Chain-length selectivity of Candida antarctica lipase B towards various aliphatic diols and dicarboxylic acid diesters. Eur. Polym. J. 2018, 106, 79–84. [Google Scholar] [CrossRef]
- Comerford, J.W.; Byrne, F.P.; Weinberger, S.; Farmer, T.J.; Pellis, A. Thermal Upgrade of Enzymatically Synthesized Aliphatic and Aromatic Oligoesters. Materials 2020, 13, 368. [Google Scholar] [CrossRef]
- Liu, W.; Chen, B.; Fang, W.; Tan, T.; Li, D. Lipase-catalyzed synthesis of aliphatic polyesters and properties characterization. Process Biochem. 2011, 46, 1993–2000. [Google Scholar] [CrossRef]
- Finnveden, M.; Martinelle, M.; Malmström], E. Itaconate based polyesters: Selectivity and performance of esterification catalysts. Eur. Polym. J. 2018, 103, 370–377. [Google Scholar] [CrossRef]
- Pellis, A.; Hanson, P.A.; Comerford, J.; Clark, J.H.; Farmer, T.J. Enzymatic synthesis of unsaturated polyesters: Functionalization and reversibility of the aza-Michael addition of pendants. Polym. Chem. 2019, 10, 843–851. [Google Scholar] [CrossRef]
- Guarneri, A.; Cutifani, V.; Cespugli, M.; Pellis, A.; Vassallo, R.; Asaro, F.; Ebert, C.; Gardossi, L. Functionalization of Enzymatically Synthesized Rigid Poly (itaconate) s via Post-Polymerization Aza-Michael Addition of Primary Amines. Adv. Synth. Catal. 2019, 361, 2559–2573. [Google Scholar] [CrossRef]
- Barrett, D.G.; Merkel, T.J.; Luft, J.C.; Yousaf, M.N. One-step syntheses of photocurable polyesters based on a renewable resource. Macromolecules 2010, 43, 9660–9667. [Google Scholar] [CrossRef]
- Baheti, P.; Bonneaud, C.; Bouilhac, C.; Joly-Duhamel, C.; Howdle, S.; Lacroix-Desmazes, P. Novel green route towards polyesters-based resin by photopolymerization of star polymers. Express Polym. Lett. 2019, 13, 1104–1115. [Google Scholar] [CrossRef]
- Núñez, M.; Muñoz-Guerra, S.; Martínez de Ilarduya, A. Poly (butylene succinate-co-ε-caprolactone) Copolyesters: Enzymatic Synthesis in Bulk and Thermal Properties. Polymers 2021, 13, 2679. [Google Scholar] [CrossRef]
- Pospiech, D.; Choińska, R.; Flugrat, D.; Sahre, K.; Jehnichen, D.; Korwitz, A.; Friedel, P.; Werner, A.; Voit, B. Enzymatic Synthesis of Poly (alkylene succinate) s: Influence of Reaction Conditions. Processes 2021, 9, 411. [Google Scholar] [CrossRef]
- Gkountela, C.; Rigopoulou, M.; Barampouti, E.M.; Vouyiouka, S. Enzymatic prepolymerization combined with bulk post-polymerization towards the production of bio-based polyesters: The case of poly (butylene succinate). Eur. Polym. J. 2021, 143, 110197. [Google Scholar] [CrossRef]
- Nasr, K.; Meimoun, J.; Favrelle-Huret, A.; Winter, J.D.; Raquez, J.-M.; Zinck, P. Enzymatic polycondensation of 1, 6-hexanediol and diethyl adipate: A statistical approach predicting the key-parameters in solution and in bulk. Polymers 2020, 12, 1907. [Google Scholar] [CrossRef]
- Kobayashi, S.; Uyama, H.; Namekawa, S. In vitro biosynthesis of polyesters with isolated enzymes in aqueous systems and organic solvents. Polym. Degrad. Stab. 1998, 59, 195–201. [Google Scholar] [CrossRef]
- Namekawa, S.; Uyama, H.; Kobayashi, S. Lipase-catalyzed ring-opening polymerization of lactones in water. Polym. J. 1998, 30, 269–271. [Google Scholar] [CrossRef]
- Pfluck, A.; Barros, D.; Fonseca, L.P. Biodegradable Polyester Synthesis in Renewed Aqueous Polycondensation Media: The Core of the New Greener Polymer-5B Technology. Processes 2021, 9, 365. [Google Scholar] [CrossRef]
- Jennings, J.; Beija, M.; Richez, A.P.; Cooper, S.D.; Mignot, P.E.; Thurecht, K.J.; Jack, K.S.; Howdle, S.M. One-Pot Synthesis of Block Copolymers in Supercritical Carbon Dioxide: A Simple Versatile Route to Nanostructured Microparticles. J. Am. Chem. Soc. 2012, 134, 4772–4781. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, M.L.; Yates, M.Z.; Smith, K.; Wilkinson, S.P. Dispersion Polymerization in Supercritical CO2 with a Siloxane-Based Macromonomer: 1. The Particle Growth Regime. Macromolecules 1998, 31, 2838–2847. [Google Scholar] [CrossRef]
- Lepilleur, C.; Beckman, E.J. Dispersion Polymerization of Methyl Methacrylate in Supercritical CO2. Macromolecules 1997, 30, 745–756. [Google Scholar] [CrossRef]
- Polloni, A.E.; Veneral, J.G.; Rebelatto, E.A.; de Oliveira, D.; Oliveira, J.V.; Araújo, P.H.; Sayer, C. Enzymatic ring opening polymerization of omega-pentadecalactone using supercritical carbon dioxide. J. Supercrit. Fluids 2017, 119, 221–228. [Google Scholar] [CrossRef]
- Sagnelli, D.; Vestri, A.; Curia, S.; Taresco, V.; Santagata, G.; Johansson, M.K.; Howdle, S.M. Green enzymatic synthesis and processing of poly (cis -9,10-epoxy-18-hydroxyoctadecanoic acid) in supercritical carbon dioxide (scCO2). Eur. Polym. J. 2021, 161, 110827. [Google Scholar] [CrossRef]
- Tsai, W.-C.; Wang, Y. Progress of supercritical fluid technology in polymerization and its applications in biomedical engineering. Prog. Polym. Sci. 2019, 98, 101161. [Google Scholar] [CrossRef]
- Sun, S.; Chen, X. Kinetics of enzymatic synthesis of monoferuloyl glycerol and diferuloyl glycerol by transesterification in [BMIM]PF6. Biochem. Eng. J. 2015, 97, 25–31. [Google Scholar] [CrossRef]
- Elgharbawy, A.A.M.; Moniruzzaman, M.; Goto, M. Recent advances of enzymatic reactions in ionic liquids: Part II. Biochem. Eng. J. 2020, 154, 107426. [Google Scholar] [CrossRef]
- Piotrowska, U.; Sobczak, M.; Oledzka, E. Characterization of aliphatic polyesters synthesized via enzymatic ring-opening polymerization in ionic liquids. Molecules 2017, 22, 923. [Google Scholar] [CrossRef] [PubMed]
- Curie, C.A.; Darmawan, M.A.; Dianursanti, D.; Budhijanto, W.; Gozan, M. The Effect of Solvent Hydrophilicity on the Enzymatic Ring-Opening Polymerization of L-Lactide by Candida rugosa Lipase. Polymers 2022, 14, 3856. [Google Scholar] [CrossRef] [PubMed]
- Wasserscheid, P.; Welton, T. Ionic Liquids in Synthesis, 2nd ed.; Wiley-VCH: Weinheim, Germany, 2008. [Google Scholar] [CrossRef]
- Zhao, H.; Afriyie, L.O.; Larm, N.E.; Baker, G.A. Glycol-functionalized ionic liquids for high-temperature enzymatic ring-opening polymerization. RSC Adv. 2018, 8, 36025–36033. [Google Scholar] [CrossRef] [PubMed]
- You, K.W.; Arumugasamy, S.K. Deep learning techniques for polycaprolactone molecular weight prediction via enzymatic polymerization process. J. Taiwan Inst. Chem. Eng. 2020, 116, 238–255. [Google Scholar] [CrossRef]
- Zhao, X.; Bansode, S.R.; Ribeiro, A.; Abreu, A.S.; Oliveira, C.; Parpot, P.; Gogate, P.R.; Rathod, V.K.; Cavaco-Paulo, A. Ultrasound enhances lipase-catalyzed synthesis of poly (ethylene glutarate). Ultrason. Sonochemistry 2016, 31, 506–511. [Google Scholar] [CrossRef]
- Idris, A.; Bukhari, A. Immobilized Candida antarctica lipase B: Hydration, stripping off and application in ring opening polyester synthesis. Biotechnol. Adv. 2012, 30, 550–563. [Google Scholar] [CrossRef]
- Montanier, C.Y.; Chabot, N.; Emond, S.; Guieysse, D.; Remaud-Siméon, M.; Peruch, F.; André, I. Engineering of Candida antarctica lipase B for poly (ε-caprolactone) synthesis. Eur. Polym. J. 2017, 95, 809–819. [Google Scholar] [CrossRef]

| Solvent | Log P | Boiling Point (°C) | Reference |
|---|---|---|---|
| diphenyl ether | 4.21 | 258 | [122] |
| p-cymene | 4.1 | 177 | [123] |
| n-heptane | 4 | 98.4 | [124] |
| cyclohexane | 3.71 | 80.74 | [122] |
| n-hexane | 3.5 | 68.73 | [122] |
| toluene | 2.5 | 110.6 | [125] |
| 2-MeTHF | 1.85 | 80.2 | [126] |
| t-butyl methyl ether | 1.43 | 55.2 | [124] |
| tetrahydrofuran | 0.53 | 66 | [127] |
| Cyrene | −1.52 | 226 | [123] |
| dimethyl sulfoxide | −1.35 | 189 | [124] |
| ethyl ether | −0.83 | 34 | [124] |
| acetone | −0.21 | 56 | [128] |
| Enzyme | Monomer | Temperature (°C) | Solvent | Mw (g/mol) | Mn (g/mol) | Ref |
|---|---|---|---|---|---|---|
| immobilized Humicola insolens | brassylic acid, 1,8-octanediol, ω-hydroxyhexadecanoic acid | 70 | diphenyl ether | - | 40,400 | [132] |
| Novozym 435 | unsaturated/epoxidized, diacid (C18, C22), diols | 90 | diphenyl ether | 57,000 | - | [133] |
| Novozym 435 | dimethyl adipate, 1,4-butanediol, 1,8-octanediol | 85 | tetrahydrofuran | 2200 | - | [134] |
| immobilized CALB | diester, aromatic diol | 85 | diphenyl ether | 2000– 4000 | - | [135] |
| immobilized CALB | azelaic acid, 1,8-octanediol | 75 | toluene | 27,000 | 21,000 | [119] |
| Novozym 435 | L-malic acid, 1,3-propanediol, 1,5-pentanediol, or 1,8-octanediol | 40 | dimethylformamide | 4800– 8100 | - | [136] |
| Novozym 435 | adipic acid, 1,8-octanediol | 70 | diphenyl ether | - | 28,500 | [37] |
| Candida antarctica Lipase | polynonadic anhydride, 1,8-octanediol | 60 | toluene | - | 10,000 | [137] |
| Novozym 435 | adipic acid divinyl ester, glycerol | 50 | tetrahydrofuran | - | 5200–13,000 | [52] |
| Novozym 435 | adipic acid, L-malic acid, 1,8-octanediol | 80 | isooctane | 17,400 | - | [129] |
| Novozym 435 | dimethyl itaconate, 1,4-butanediol, diethyl adipate | 80 | diphenyl ether | - | 94,000 | [138] |
| Novozym 435 | dimethyl itaconate, butanedioic acid, butanediol | 80 | diphenyl ether | - | 1948–13,288 | [45] |
| Novozym 435 | diethyl furan-2,5-dicarboxylate dodecane-1,12-diol, diethyl oxalate | 100 | diphenyl ether | - | 8900 | [139] |
| CALB | diethyl succinate, 1,4-butanediol dilinoleic diol | 95 | diphenyl ether | - | 16,300–25,200 | [140] |
| immobilized CALB | suberic acid, 1,8-octanediol | 60– 80 | cyclohexane: tetrahydrofuran 5:1 v/v | 16,000–19,800 | - | [141] |
| CALB | ferulic-based diester, 1,4-butanediol | 90 | diphenyl ether | - | 2000 | [42] |
| Novozym 435 | δ-valerolactone | 85–95– 110 | toluene | - | 2300 | [142] |
| Novozym 435 | trans, β-dimethyl hydromuconate, adipic acid, 1,8-octanediol | 85–95– 110 | diphenyl ether | - | 21,900 | [143] |
| CALB | cis-tetraphenylporphyrin macrocycle | 70 | toluene | - | 9600 | [144] |
| Enzyme | Monomer | Temperature (°C) | Solvent | Mw (g/mol) | Mn (g/mol) | Ref |
|---|---|---|---|---|---|---|
| Novozym 435 | unsaturated/epoxidized, diacid (C18, C22), diols | 90 | no-solvent | 25,000 | - | [133] |
| Novozym 435 | 1,12-dodecanedioic, 1-thioglycerol | 80 | no-solvent | 5380 | - | [153] |
| Novozym 435 | 1,18-cis-9-octadecenedioic, glycerol | 90 | no-solvent | - | 6000–9100 | [154] |
| Novozym 435 | dicarboxylic acid diesters and diols | 85 | no-solvent | - | 1094- 11,549 | [155] |
| immobilized CALB | 1,4-butanediol, dimethyl adipate | 70 | no-solvent | 2200 | 1000 | [156] |
| lipase from Candida sp.99-125 | diethyl sebacate, 1,4-butanediol | 70 | no-solvent | - | 15,800 | [157] |
| Novozym 435 | tall oil fatty acid, glycerol, azelaic acid | 90 | no-solvent | 20,900–39,700 | - | [152] |
| Novozym 435 | dimethyl itaconate, 1,4-butanediol, dimethyl succinate | 60 | no-solvent | - | 840 | [158] |
| Novozym 435 | itaconic acid, diethylamine, 1,8-octanediol | 85 | no-solvent | - | 696 | [159] |
| Novozym 435 | dimethyl itaconate, 1,4-cyclohexanedimethanol | 50 | no-solvent | - | 720–2859 | [160] |
| Novozym 435 | itaconic acid, succinic acid, sorbitol | 90 | no-solvent | - | 1140–11,900 | [161] |
| Novozym 435 | ε-caprolactone | 90 | no-solvent | - | 3100 | [162] |
| Novozym 435 | dimethyl succinateand, 1,4-butanediol, ε-caprolactone. | 90–110 | no-solvent | - | 7100–10,100 | [163] |
| CALB | divinyladipate, 1,4-butanediol | 70 | no-solvent | 31,000 | 7100 | [164] |
| CALB | 1,4-butanediol, diethyl succinate | 40 | no-solvent | - | 2800 | [165] |
| Novozym 435 | 1,6-hexandiol and diethyl adipate | 100 | no-solvent | - | 5000–12,000 | [166] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, H.; Li, H.; Lee, C.K.; Mat Nanyan, N.S.; Tay, G.S. Recent Advances in the Enzymatic Synthesis of Polyester. Polymers 2022, 14, 5059. https://doi.org/10.3390/polym14235059
Wang H, Li H, Lee CK, Mat Nanyan NS, Tay GS. Recent Advances in the Enzymatic Synthesis of Polyester. Polymers. 2022; 14(23):5059. https://doi.org/10.3390/polym14235059
Chicago/Turabian StyleWang, Hong, Hongpeng Li, Chee Keong Lee, Noreen Suliani Mat Nanyan, and Guan Seng Tay. 2022. "Recent Advances in the Enzymatic Synthesis of Polyester" Polymers 14, no. 23: 5059. https://doi.org/10.3390/polym14235059
APA StyleWang, H., Li, H., Lee, C. K., Mat Nanyan, N. S., & Tay, G. S. (2022). Recent Advances in the Enzymatic Synthesis of Polyester. Polymers, 14(23), 5059. https://doi.org/10.3390/polym14235059







