Comparison of Droplet Digital PCR and qPCR for the Quantification of Shiga Toxin-Producing Escherichia coli in Bovine Feces
Abstract
:1. Introduction
2. Results
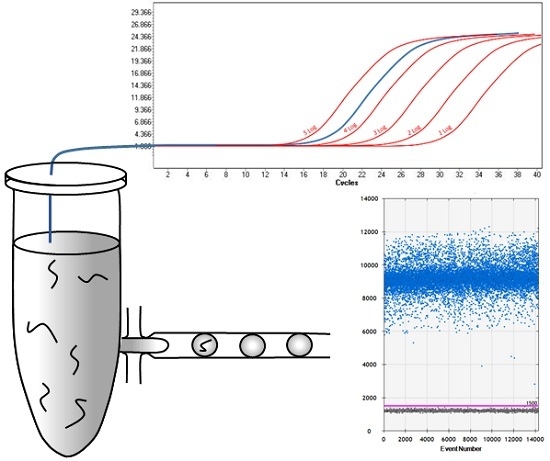
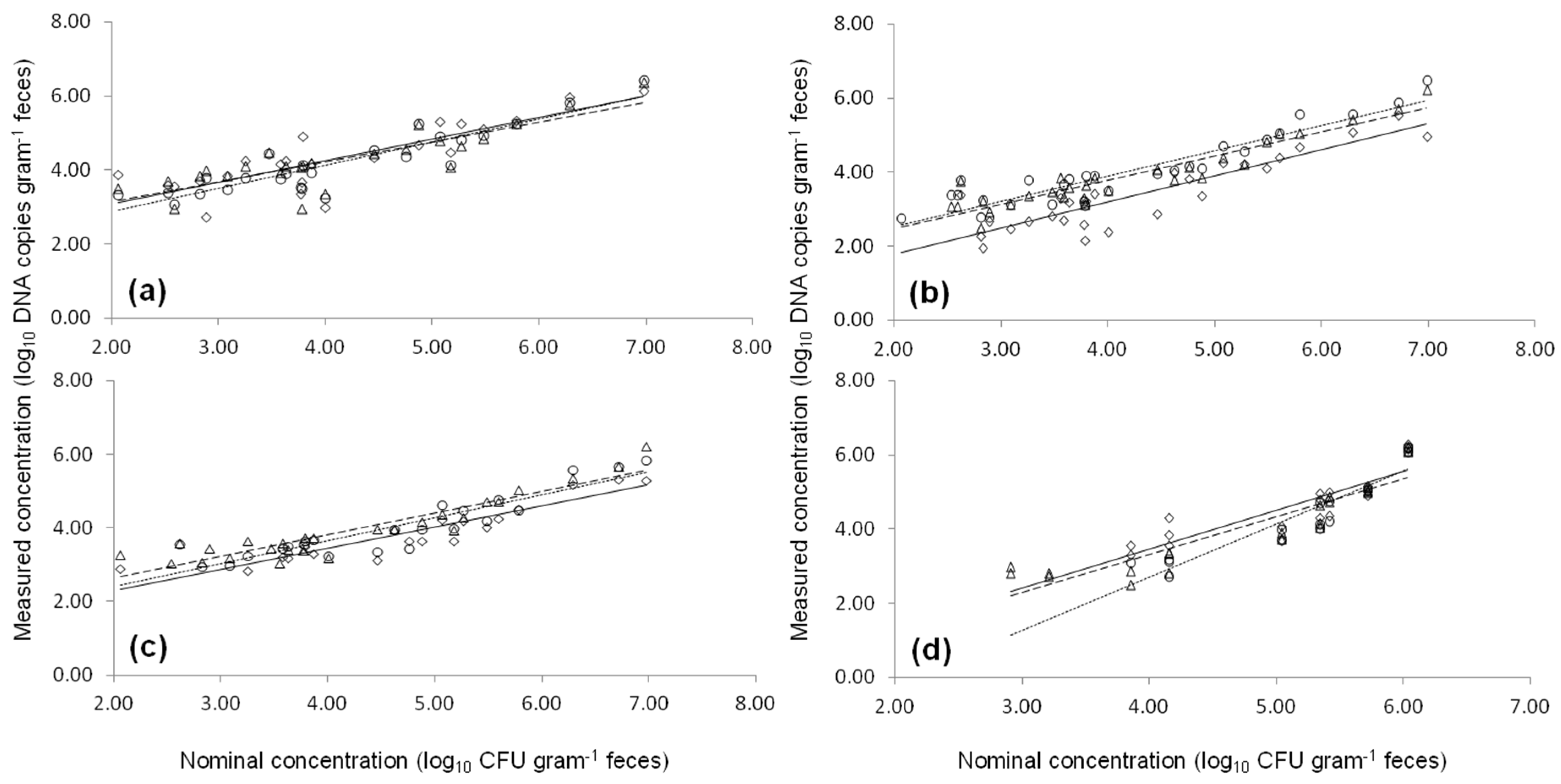
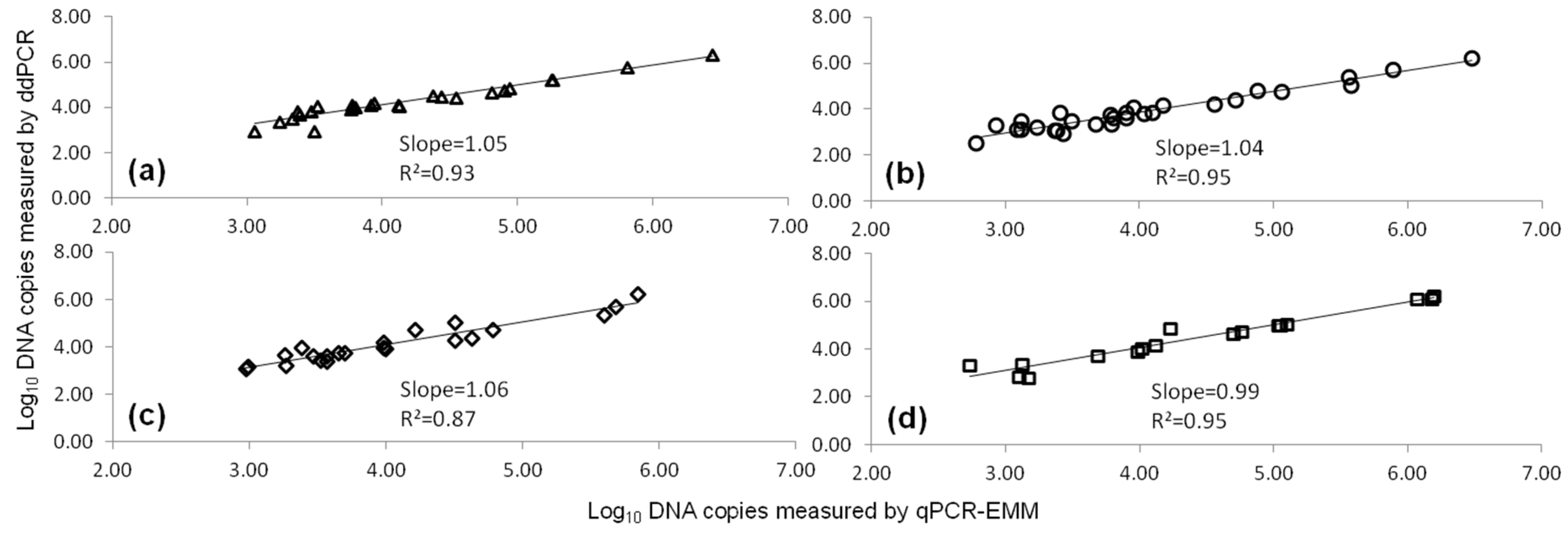
2.1. Comparison of qPCR and ddPCR Standard Curves
2.2. Inhibition
2.3. Artificial Contaminated Fecal Sample
2.4. Natural Contaminated Fecal Samples
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Strains
5.2. qPCR Assays
5.3. ddPCR Assays
5.4. Inhibition
5.5. Internal Amplification Control (IAC)
5.6. Artificial STEC Contaminated Fecal Samples
5.7. Natural STEC Contaminated Fecal Samples
5.8. Statistical Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| qPCR: | real-time quantitative PCR |
| ddPCR: | Droplet digital PCR |
| EMM: | TaqMan® Environmental Master Mix 2.0 |
| UMM: | TaqMan® Universal PCR Master Mix |
| STEC: | Shiga toxin-producing Escherichia coli |
| VTEC: | Verocytotoxin-producing Escherichia coli |
| R2: | degree of linearity |
| IAC: | Internal Amplification Control |
| TSB: | Tryptone Soy Broth |
| UNG: | Uracil-DNA Glycosylase |
| RAMS: | recto-anal-mucosal swab |
| LOQ: | limit of quantification |
References
- Nataro, J.P.; Kaper, J.B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev. 1998, 11, 142–201. [Google Scholar] [PubMed]
- Jinneman, K.C.; Waite-Cusic, J.G.; Yoshitomi, K.J. Evaluation of shiga toxin-producing Escherichia coli (STEC) method for the detection and identification of STEC O104 strains from sprouts. Food Microbiol. 2012, 30, 321–328. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, S.C.; Son, I.; Maounounen-Laasri, A.; Lin, A.; Fischer, M.; Kase, J.A. Prevalence of Hemolysin Genes and Comparison of ehxA Subtype Patterns in Shiga toxin-producing Escherichia coli (STEC) and non-STEC Strains from Clinical, Food, and Animal Sources. Appl. Environ. Microbiol. 2013, 79, 6301–6311. [Google Scholar] [CrossRef] [PubMed]
- Thomas, K.M.; McCann, M.S.; Collery, M.M.; Logan, A.; Whyte, P.; McDowell, D.A.; Duffy, G. Tracking verocytotoxigenic Escherichia coli O157, O26, O111, O103 and O145 in Irish cattle. Int. J. Food Microbiol. 2012, 153, 288–296. [Google Scholar] [CrossRef] [PubMed]
- EFSA. The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2014. EFSA J. 2016, 13, 4329. [Google Scholar]
- Munns, K.D.; Selinger, L.B.; Stanford, K.; Guan, L.; Callaway, T.R.; McAllister, T.A. Perspectives on Super-Shedding of Escherichia coli O157:H7 by Cattle. Foodborne Pathog. Dis. 2015, 12, 89–103. [Google Scholar] [CrossRef] [PubMed]
- Matthews, L.; McKendrick, I.J.; Ternent, H.; Gunn, G.J.; Synge, B.; Woolhouse, M.E.J. Super-shedding cattle and the transmission dynamics of Escherichia coli O157. Epidemiol. Infect. 2006, 134, 131–142. [Google Scholar] [CrossRef] [PubMed]
- Duffy, G.; Burgess, C.M.; Bolton, D.J. A review of factors that affect transmission and survival of verocytotoxigenic Escherichia coli in the European farm to fork beef chain. Meat Sci. 2014, 97, 375–383. [Google Scholar] [CrossRef] [PubMed]
- Lawal, D.; Burgess, C.; McCabe, E.; Whyte, P.; Duffy, G. Development of a quantitative real time PCR assay to detect and enumerate Escherichia coli O157 and O26 serogroups in bovine recto-anal swabs. J. Microbiol. Methods 2015, 114, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Verhaegen, B.; De Reu, K.; Heyndrickx, M.; De Zutter, L. Comparison of Six Chromogenic Agar Media for the Isolation of a Broad Variety of Non-O157 Shiga toxin-Producing Escherichia coli (STEC) Serogroups. Int. J. Environ. Res. Public Health 2015, 12, 6965–6978. [Google Scholar] [CrossRef] [PubMed]
- Verstraete, K.; Van Coillie, E.; Werbrouck, H.; Van Weyenberg, S.; Herman, L.; Del-Favero, J.; De Rijk, P.; De Zutter, L.; Joris, M.-A.; Heyndrickx, M.; et al. A qPCR assay to detect and quantify Shiga toxin-producing E. coli (STEC) in cattle and on farms: A potential predictive tool for STEC culture-positive farms. Toxins (Basel) 2014, 6, 1201–1221. [Google Scholar] [CrossRef] [PubMed]
- Luedtke, B.E.; Bono, J.L.; Bosilevac, J.M. Evaluation of real time PCR assays for the detection and enumeration of enterohemorrhagic Escherichia coli directly from cattle feces. J. Microbiol. Methods 2014, 105, 72–79. [Google Scholar] [CrossRef] [PubMed]
- Noll, L.W.; Shridhar, P.B.; Shi, X.; An, B.; Cernicchiaro, N.; Renter, D.G.; Nagaraja, T.G.; Bai, J. A Four-Plex Real-Time PCR Assay, Based on rfbE, stx 1, stx 2, and eae Genes, for the Detection and Quantification of Shiga Toxin–Producing Escherichia coli O157 in Cattle Feces. Foodborne Pathog. Dis. 2015, 12, 787–794. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Gyawali, P.; Toze, S. Quantitative PCR measurements of Escherichia coli including shiga toxin-producing E. coli (STEC) in animal feces and environmental waters. Environ. Sci. Technol. 2015, 49, 3084–3090. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Nolan, T. Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J. Biomol. Tech. 2004, 15, 155–166. [Google Scholar] [PubMed]
- Baker, M. Digital PCR hits its stride. Nat. Methods 2012, 9, 541–544. [Google Scholar] [CrossRef]
- Hindson, C.M.; Chevillet, J.R.; Briggs, H.A.; Gallichotte, E.N.; Ruf, I.K.; Hindson, B.J.; Vessella, R.L.; Tewari, M. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat. Methods 2013, 10, 1003–1005. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, L.B.; Coleman, V.A.; Hindson, C.M.; Herrmann, J.; Hindson, B.J.; Bhat, S.; Emslie, K.R. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal. Chem. 2012, 84, 1003–1011. [Google Scholar] [CrossRef] [PubMed]
- Huggett, J.F.; Novak, T.; Garson, J.A.; Green, C.; Morris-jones, S.D.; Miller, R.F.; Zumla, A. Differential susceptibility of PCR reactions to inhibitors: An important and unrecognised phenomenon. BMC Res. Notes 2008, 1, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Doi, H.; Takahara, T.; Minamoto, T.; Matsuhashi, S.; Uchii, K.; Yamanaka, H. Droplet digital polymerase chain reaction (PCR) outperforms real-time PCR in the detection of environmental DNA from an invasive fish species. Environ. Sci. Technol. 2015, 49, 5601–5608. [Google Scholar] [CrossRef] [PubMed]
- Strain, M.C.; Lada, S.M.; Luong, T.; Rought, S.E.; Gianella, S.; Terry, V.H.; Spina, C.A.; Woelk, C.H.; Richman, D.D. Highly Precise Measurement of HIV DNA by Droplet Digital PCR. PLoS ONE 2013, 8, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sze, M.A.; Abbasi, M.; Hogg, J.C.; Sin, D.D. A comparison between droplet digital and quantitative PCR in the analysis of bacterial 16S load in lung tissue samples from control and COPD GOLD 2. PLoS ONE 2014, 9, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Beer, N.R.; Hindson, B.J.; Wheeler, E.K.; Hall, S.B.; Rose, K.A.; Kennedy, I.M.; Colston, B.W. Reaction in Picoliter Droplets. Anal. Chem. 2007, 79, 8471–8475. [Google Scholar] [CrossRef] [PubMed]
- Davis, M.A.; Rice, D.H.; Sheng, H.; Hancock, D.D.; Thomas, E.B.; Cobbold, R.; Hovde, C.J.; Besser, T.E. Comparison of cultures from rectoanal-junction mucosal swabs and feces for detection of Escherichia coli O157 in dairy heifers. Appl. Environ. Microbiol. 2006, 72, 3766–3770. [Google Scholar] [CrossRef] [PubMed]
- Perna, N.T.; Plunkett, G.; Burland, V.; Mau, B.; Glasner, J.D.; Rose, D.J.; Mayhew, G.F.; Evans, P.S.; Gregor, J.; Kirkpatrick, H.A. Genome sequence of enterohaemorrhagic Escherichia coli O157: H7. Nature 2001, 409, 529–533. [Google Scholar] [CrossRef] [PubMed]
- Thompson, R.E.; Duncan, G.; McCord, B.R. An Investigation of PCR Inhibition Using Plexor-Based Quantitative PCR and Short Tandem Repeat Amplification. J. Forensic Sci. 2014, 59, 1517–1529. [Google Scholar] [CrossRef] [PubMed]
- The R Project for Statistical Computing. Available online: http://www.R-project.org/ (accessed on 7 September 2015).

 , qPCR-UMM
, qPCR-UMM  and ddPCR
and ddPCR  in cattle fecal samples and recto-anal-mucosal swabs (RAMS) of ten animals (1–10) of a STEC-positive farm. * Cattle fecal samples that showed PCR inhibition for the internal amplification control.
in cattle fecal samples and recto-anal-mucosal swabs (RAMS) of ten animals (1–10) of a STEC-positive farm. * Cattle fecal samples that showed PCR inhibition for the internal amplification control.
 , qPCR-UMM
, qPCR-UMM  and ddPCR
and ddPCR  in cattle fecal samples and recto-anal-mucosal swabs (RAMS) of ten animals (1–10) of a STEC-positive farm. * Cattle fecal samples that showed PCR inhibition for the internal amplification control.
in cattle fecal samples and recto-anal-mucosal swabs (RAMS) of ten animals (1–10) of a STEC-positive farm. * Cattle fecal samples that showed PCR inhibition for the internal amplification control.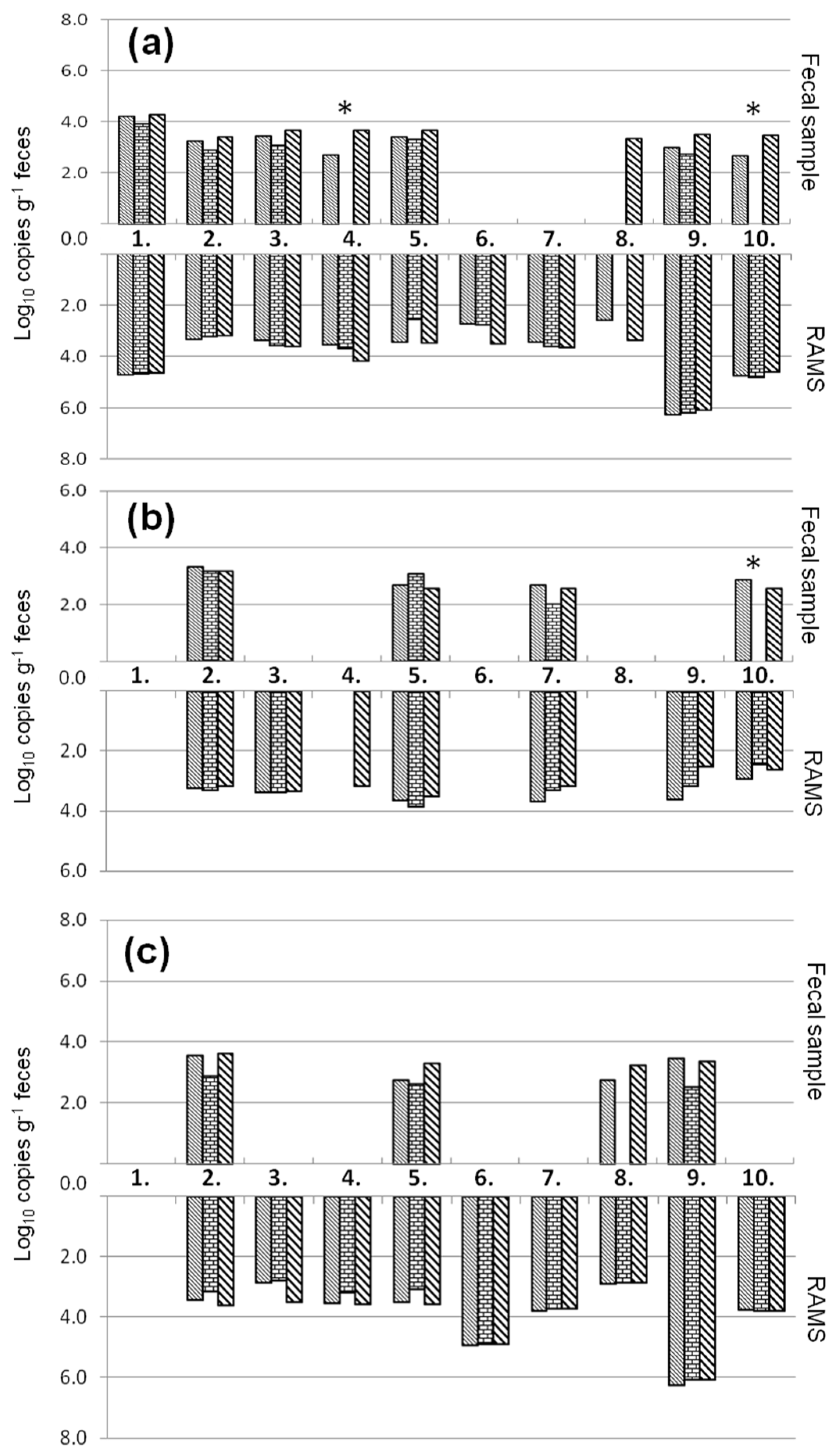
| Concentration (Copies per µL Template) | Gene | |||
|---|---|---|---|---|
| eae | stx1 | stx2 (Set a) | stx2 (Set b) | |
| 2 × 105 | 100% | 100% | 100% | 100% |
| 2 × 104 | 87% | 87% | 87% | 88% |
| 2 × 103 | 19% | 19% | 19% | 21% |
| 2 × 102 | 2.03% | 1.96% | 2.06% | 2.11% |
| 2 × 101 | 0.22% | 0.26% | 0.20% | 0.18% |
| 101 | 0.12% | 0.13% | 0.13% | 0.17% |
| Gene | Primer or Probe * | Sequence (5′-3′) | Labeling (5′-3′) | |
|---|---|---|---|---|
| qPCR | ddPCR | |||
| eae | eae-F | GGA AGC CAA AGC GCA CAA | - | - |
| eae-R | GGC ICG AGC IGT CAC TTT ATA A | - | - | |
| eae-P | TAC CAG GCT ATT TTG CCI GCT TAT GTG C | FAM–BHQ-1 | FAM–ZEN–IBFQ | |
| stx1 | stx1-F | GAC GCA GTC TGT IGC AAG AG | - | - |
| stx1-R | CGA AAA CGI AAA GCT TCA GCT G | - | - | |
| stx1-P | ATG TTA CGG TTT GTT ACT GTG | FAM–MGBNFQ | FAM–ZEN–IBFQ | |
| stx2 | stx2-F | TCA GGC AIA TAC AGA GAG AAT TTC G | - | - |
| stx2-R (set a) | CCG GIG TCA TCG TAT ACA CAG | - | - | |
| stx2-R (set b) | CCG GIG TCA TCG TAT AAA CAG | - | - | |
| stx2-P | CAC TGT CTG AAA CTG CT | FAM–MGBNFQ | FAM–ZEN–IBFQ | |
| IAC | stx1-F | GAC GCA GTC TGT IGC AAG AG | - | - |
| stx1-R | CGA AAA CGI AAA GCT TCA GCT G | - | - | |
| eae-P | TAC CAG GCT ATT TTG CCI GCT TAT GTG C | HEX–BHQ-1 | HEX–ZEN–IBFQ | |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Verhaegen, B.; De Reu, K.; De Zutter, L.; Verstraete, K.; Heyndrickx, M.; Van Coillie, E. Comparison of Droplet Digital PCR and qPCR for the Quantification of Shiga Toxin-Producing Escherichia coli in Bovine Feces. Toxins 2016, 8, 157. https://doi.org/10.3390/toxins8050157
Verhaegen B, De Reu K, De Zutter L, Verstraete K, Heyndrickx M, Van Coillie E. Comparison of Droplet Digital PCR and qPCR for the Quantification of Shiga Toxin-Producing Escherichia coli in Bovine Feces. Toxins. 2016; 8(5):157. https://doi.org/10.3390/toxins8050157
Chicago/Turabian StyleVerhaegen, Bavo, Koen De Reu, Lieven De Zutter, Karen Verstraete, Marc Heyndrickx, and Els Van Coillie. 2016. "Comparison of Droplet Digital PCR and qPCR for the Quantification of Shiga Toxin-Producing Escherichia coli in Bovine Feces" Toxins 8, no. 5: 157. https://doi.org/10.3390/toxins8050157
APA StyleVerhaegen, B., De Reu, K., De Zutter, L., Verstraete, K., Heyndrickx, M., & Van Coillie, E. (2016). Comparison of Droplet Digital PCR and qPCR for the Quantification of Shiga Toxin-Producing Escherichia coli in Bovine Feces. Toxins, 8(5), 157. https://doi.org/10.3390/toxins8050157