Pathway for Biodegrading Nodularin (NOD) by Sphingopyxis sp. USTB-05
Abstract
:1. Introduction
2. Results
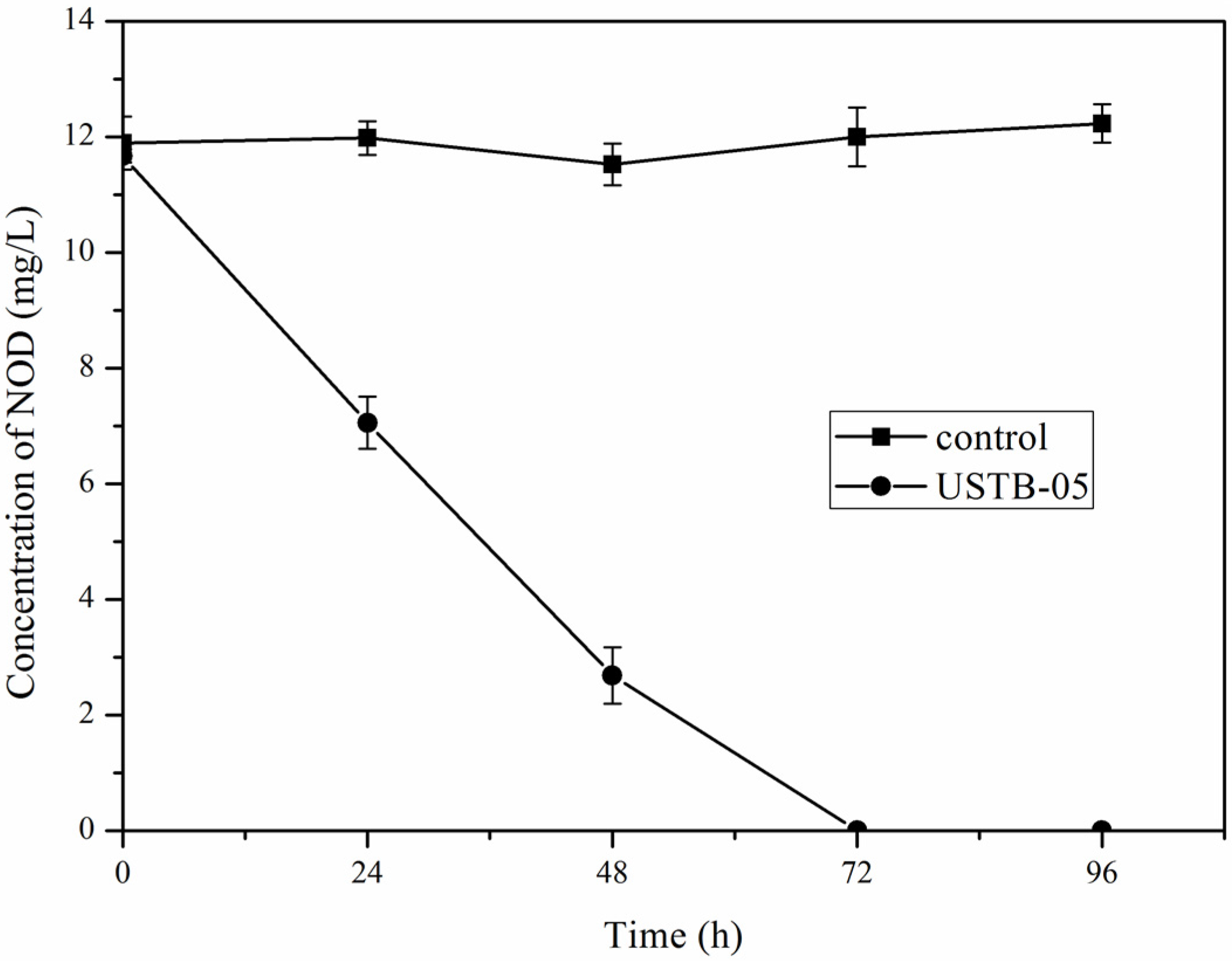
2.1. Biodegradation of NOD by Sphingopyxis sp. USTB-05
2.2. Enzymatic Kinetics of the NOD Biodegradation by CEs of USTB-05
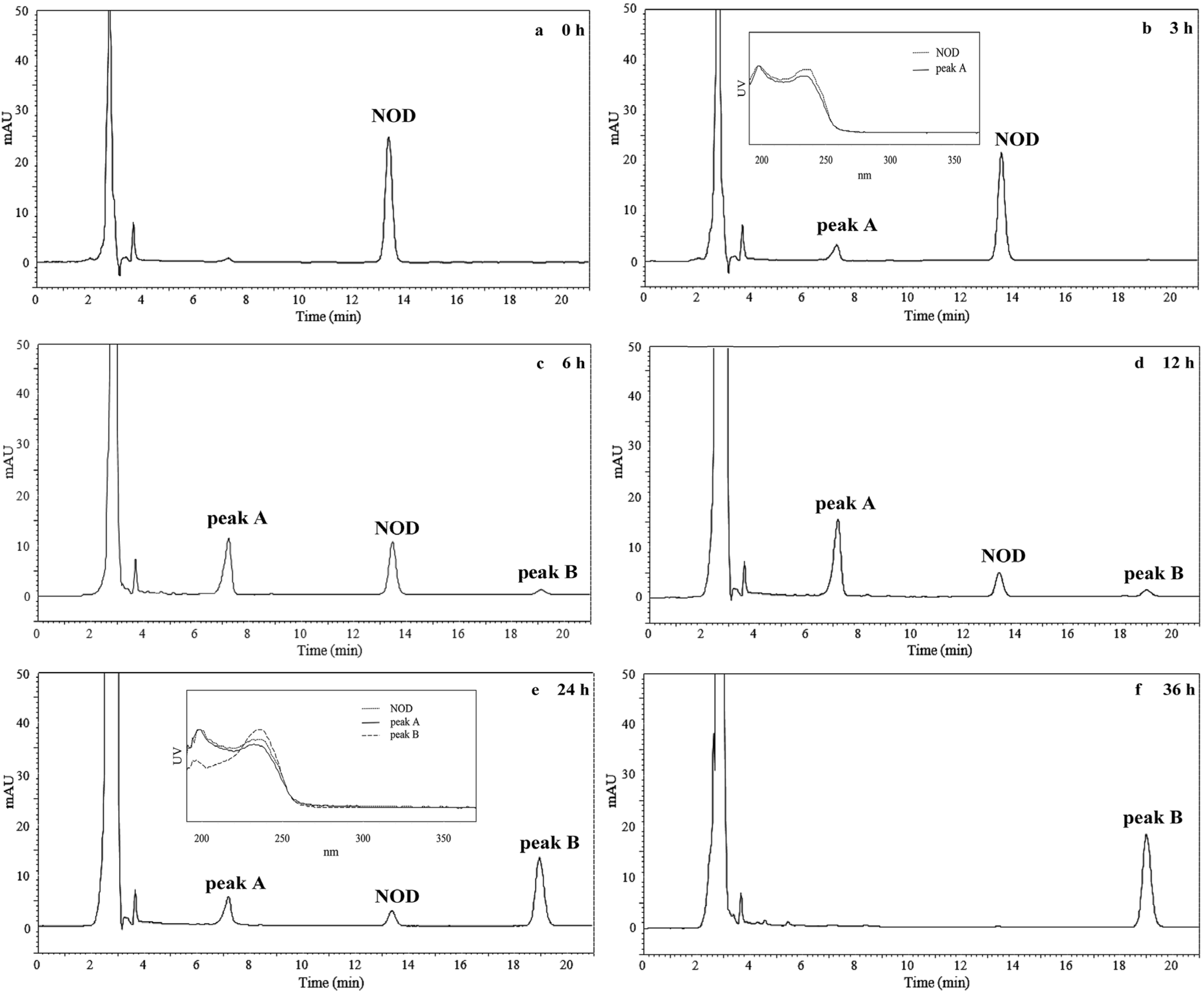
2.3. Products of NOD Detected at the Wavelength of 238 nm on HPLC
2.4. RRLC-MS Analysis of NOD and Its Biodegradation Products
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Chemicals
5.2. Bacterial Strain and Cultural Conditions
5.3. Preparation for the Crude Enzymes (CEs) of Sphingopyxis sp. USTB-05
5.4. NOD-Biodegrading Activity of Sphingopyxis sp. USTB-05
5.5. Enzymatic Biodegradation of NOD
5.6. Analysis of NOD and Its Biodegradation Products
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Svrcek, C.; Smith, D.W. Cyanobacteria toxins and the current state of knowledge on water treatment options: A review. J. Environ. Eng. Sci. 2004, 3, 155–185. [Google Scholar] [CrossRef]
- Repka, S.; Meyerhofer, M.; von Brockel, K.; Sivonen, K. Associations of cyanobacterial toxin, nodularin, with environmental factors and zooplankton in the Baltic Sea. Microb. Ecol. 2004, 47, 350–358. [Google Scholar] [CrossRef] [PubMed]
- Heresztyn, T.; Nicholson, B.C. Nodularin concentrations in Lakes Alexandrina and Albert, South Australia, during a bloom of the cyanobacterium (blue-green alga) Nodularia spumigena and degradation of the toxin. Environ. Toxicol. Water Qual. 1997, 12, 273–282. [Google Scholar] [CrossRef]
- Horne, A.J.; Galat, D.L. Nitrogen fixation in an oligotrophic, saline desert lake: Pyramid Lake, Nevada. Limnol. Oceanogr. 1985, 30, 1229–1239. [Google Scholar] [CrossRef]
- Nehring, S. Mortality of dogs associated with a mass development of Nodularia spumigena (Cyanophyceae) in a brackish lake at the German North Sea coast. J. Plankton Res. 1993, 15, 867–872. [Google Scholar] [CrossRef]
- Beattie, K.A.; Kaya, K.; Codd, G.A. The cyanobacterium Nodularia PCC 7804, of freshwater origin, produces [L-Har2] nodularin. Phytochemistry 2000, 54, 57–61. [Google Scholar] [CrossRef]
- Rinehart, K.L.; Harada, K.; Namikoshi, M.; Chen, C.; Harvis, C.A.; Munro, M.H.G.; Blunt, J.W.; Mulligan, P.E.; Beasley, V.R.; Al, E. Nodularin, microcystin, and the configuration of Adda. J. Am. Chem. Soc. 1988, 110. [Google Scholar] [CrossRef]
- Ohta, T.; Sueoka, E.; Iida, N.; Komori, A.; Suganuma, M.; Nishiwaki, R.; Tatematsu, M.; Kim, S.J.; Carmichael, W.W.; Fujiki, H. Nodularin, a potent inhibitor of protein phosphatases 1 and 2A, is a new environmental carcinogen in male F344 rat liver. Cancer Res. 1994, 54, 6402–6406. [Google Scholar] [PubMed]
- Carmichael, W.W.; Eschedor, J.T.; Patterson, G.M.; Moore, R.E. Toxicity and partial structure of a hepatotoxic peptide produced by the cyanobacterium Nodularia spumigena Mertens emend. L575 from New Zealand. Appl. Environ. Microbiol. 1988, 54, 2257–2263. [Google Scholar] [PubMed]
- Codd, G.A.; Bell, S.G.; Kaya, K.; Ward, C.J.; Tie, K.A.; Metcalf, J.S. Cyanobacteria toxins, exposure routes and human health. Eur. J. Phycol. 1999, 34, 405–415. [Google Scholar] [CrossRef]
- Stewart, I.; Seawright, A.A.; Shaw, G.R. Cyanobacterial poisoning in livestock, wild mammals and birds—An overview. In Cyanobacterial Harmful Algal Blooms: State of the Science and Research Needs; Hudnell, H.K., Ed.; Publisher: New York, NY, USA, 2008; Volume 619, pp. 613–637. [Google Scholar]
- Van, H.A.; Harding, W.R.; Wessels, J.C.; Schneider, D.J.; Heine, E.W.; Van, D.M.J.; Fourie, J.M. Cyanobacterial (blue-green algae) poisoning of livestock in the western Cape Province of South Africa. J. S. Afr. Vet. Assoc. 1995, 66, 260–264. [Google Scholar]
- Pearson, L.; Mihali, T.; Moffitt, M.; Kellmann, R.; Neilan, B. On the chemistry, toxicology and genetics of the cyanobacterial toxins, microcystin, nodularin, saxitoxin and cylindrospermopsin. Mar. Drugs 2010, 8, 1650–1680. [Google Scholar] [CrossRef] [PubMed]
- Annila, A.; Lehtimaki, J.; Mattila, K.; Eriksson, J.E.; Sivonen, K.; Rantala, T.T.; Drakenberg, T. Solution structure of nodularin. An inhibitor of serine/threonine-specific protein phosphatases. J. Biol. Chem. 1996, 271, 16695–16702. [Google Scholar] [PubMed]
- Merel, S.; Clement, M.; Thomas, O. State of the art on cyanotoxins in water and their behaviour towards chlorine. Toxicon 2010, 55, 677–691. [Google Scholar] [CrossRef] [PubMed]
- Rapala, J.; Berg, K.A.; Lyra, C.; Niemi, R.M.; Manz, W.; Suomalainen, S.; Paulin, L.; Lahti, K. Paucibacter toxinivorans gen. nov., sp nov., a bacterium that degrades cyclic cyanobacterial hepatotoxins microcystins and nodularin. Int. J. Syst. Evol. Microbiol. 2005, 55, 1563–1568. [Google Scholar] [CrossRef] [PubMed]
- Imanishi, S.; Kato, H.; Mizuno, M.; Tsuji, K.; Harada, K. Bacterial degradation of microcystins and nodularin. Chem. Res. Toxicol. 2005, 18, 591–598. [Google Scholar] [CrossRef] [PubMed]
- Lawton, L.A.; Welgamage, A.; Manage, P.M.; Edwards, C. Novel bacterial strains for the removal of microcystins from drinking water. Water Sci. Technol. 2011, 63, 1137–1142. [Google Scholar] [CrossRef] [PubMed]
- Bourne, D.G.; Jones, G.J.; Blakeley, R.L.; Jones, A.; Negri, A.P.; Riddles, P. Enzymatic pathway for the bacterial degradation of the cyanobacterial cyclic peptide toxin microcystin LR. Appl. Environ. Microbiol. 1996, 62, 4086–4094. [Google Scholar] [PubMed]
- Hai, Y.; Wang, H.; Wang, J.; Yin, C.; Song, M.; Liu, X.; Yin, X. Cloning and expression of the first gene for biodegrading microcystin LR by Sphingopyxis sp. USTB-05. J. Environ. Sci. 2012, 10, 1816–1822. [Google Scholar]
- Shimizu, K.; Maseda, H.; Okano, K.; Kurashima, T.; Kawauchi, Y.; Qiang, X.; Utsumi, M.; Zhang, Z.; Sugiura, N. Enzymatic pathway for biodegrading microcystin LR in Sphingopyxis sp. C-1. J. Biosci. Bioeng. 2012, 114, 630–634. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wu, P.; Chen, J.; Yan, H. Biodegradation of microcystin-RR by a new isolated Sphingopyxis sp. USTB-05. Chin. J. Chem. Eng. 2010, 18, 108–112. [Google Scholar] [CrossRef]
- Xiao, C.; Yan, H.; Wang, J.; Wei, W.; Ning, J.; Pan, G. Microcystin-LR biodegradation by Sphingopyxis sp. USTB-05. Front. Environ. Sci. Eng. China 2011, 5, 526–532. [Google Scholar] [CrossRef]
- Xu, H.; Wang, H.; Xu, Q.; Lv, L.; Yin, C.; Liu, X.; Du, H.; Yan, H. Pathway for Biodegrading Microcystin-YR by Sphingopyxis sp. USTB-05. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Namikoshi, M.; Rinehart, K.L.; Sakai, R.; Stotts, R.R.; Dahlem, A.M.; Beasley, V.R.; Carmichael, W.W.; Evans, W.R. Identification of 12 hepatotoxins from a Homer Lake bloom of the cyanobacteria Microcystis aeruginosa, Microcystis viridis, and Microcystis wesenbergii: Nine new microcystins. J. Organ. Chem. 1992, 57. [Google Scholar] [CrossRef]
- Mazur-Marzec, H.; Torunska, A.; Blonska, M.J.; Moskot, M.; Plinski, M.; Jakobkiewicz-Banecka, J.; Wegrzyn, G. Biodegradation of nodularin and effects of the toxin on bacterial isolates from the Gulf of Gdansk. Water Res. 2009, 43, 2801–2810. [Google Scholar] [CrossRef] [PubMed]
- Torunska, A.; Bolalek, J.; Plinski, M.; Mazur-Marzec, H. Biodegradation and sorption of nodularin (NOD) in fine-grained sediments. Chemosphere 2008, 70, 2039–2046. [Google Scholar] [CrossRef] [PubMed]
- Holst, T.; Jorgensen, N.O.G.; Jorgensen, C.; Johansen, A. Degradation of microcystin in sediments at oxic and anoxic, denitrifying conditions. Water Res. 2003, 37, 4748–4760. [Google Scholar] [CrossRef]
- Saitou, T.; Sugiura, N.; Itayama, T.; Inamori, Y.; Matsumura, M. Degradation characteristics of microcystins by isolated bacteria from Lake Kasumigaura. J. Water Supply Res. Technol. AQUA 2003, 52, 13–18. [Google Scholar]
- Yang, F.; Zhou, Y.; Sun, R.; Wei, H.; Li, Y.; Yin, L.; Pu, Y. Biodegradation of microcystin-LR and-RR by a novel microcystin-degrading bacterium isolated from Lake Taihu. Biodegradation 2014, 25, 447–457. [Google Scholar] [CrossRef] [PubMed]
- Edwards, C.; Graham, D.; Fowler, N.; Lawton, L.A. Biodegradation of microcystins and nodularin in freshwaters. Chemosphere 2008, 73, 1315–1321. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Imanishi, S.Y.; Tsuji, K.; Harada, K.-I. Microbial degradation of cyanobacterial cyclic peptides. Water Res. 2007, 41, 1754–1762. [Google Scholar] [CrossRef] [PubMed]
- Ishii, H.; Nishijima, M.; Abe, T. Characterization of degradation process of cyanobacterial hepatotoxins by a gram-negative aerobic bacterium. Water Res. 2004, 38, 2667–2676. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Yan, H.; Ma, S.; Liu, X.; Yin, C.; Wang, H.; Xu, Q.; Lv, L. Characterization of the second and third steps in the enzymatic pathway for microcystin-RR biodegradation by Sphingopyxis sp. USTB-05. Ann. Microbiol. 2015, 65, 495–502. [Google Scholar] [CrossRef]
- Yan, H.; Wang, J.; Chen, J.; Wei, W.; Wang, H.; Wang, H. Characterization of the first step involved in enzymatic pathway for microcystin-RR biodegraded by Sphingopyxis sp. USTB-05. Chemosphere 2012, 87, 12–18. [Google Scholar] [CrossRef] [PubMed]



© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feng, N.; Yang, F.; Yan, H.; Yin, C.; Liu, X.; Zhang, H.; Xu, Q.; Lv, L.; Wang, H. Pathway for Biodegrading Nodularin (NOD) by Sphingopyxis sp. USTB-05. Toxins 2016, 8, 116. https://doi.org/10.3390/toxins8050116
Feng N, Yang F, Yan H, Yin C, Liu X, Zhang H, Xu Q, Lv L, Wang H. Pathway for Biodegrading Nodularin (NOD) by Sphingopyxis sp. USTB-05. Toxins. 2016; 8(5):116. https://doi.org/10.3390/toxins8050116
Chicago/Turabian StyleFeng, Nan, Fan Yang, Hai Yan, Chunhua Yin, Xiaolu Liu, Haiyang Zhang, Qianqian Xu, Le Lv, and Huasheng Wang. 2016. "Pathway for Biodegrading Nodularin (NOD) by Sphingopyxis sp. USTB-05" Toxins 8, no. 5: 116. https://doi.org/10.3390/toxins8050116
APA StyleFeng, N., Yang, F., Yan, H., Yin, C., Liu, X., Zhang, H., Xu, Q., Lv, L., & Wang, H. (2016). Pathway for Biodegrading Nodularin (NOD) by Sphingopyxis sp. USTB-05. Toxins, 8(5), 116. https://doi.org/10.3390/toxins8050116






