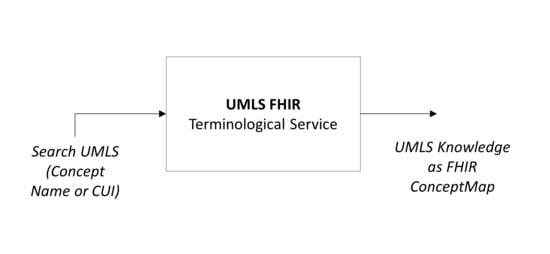
An Interoperable UMLS Terminology Service Using FHIR
Abstract
1. Introduction
2. Literature Review
3. Background
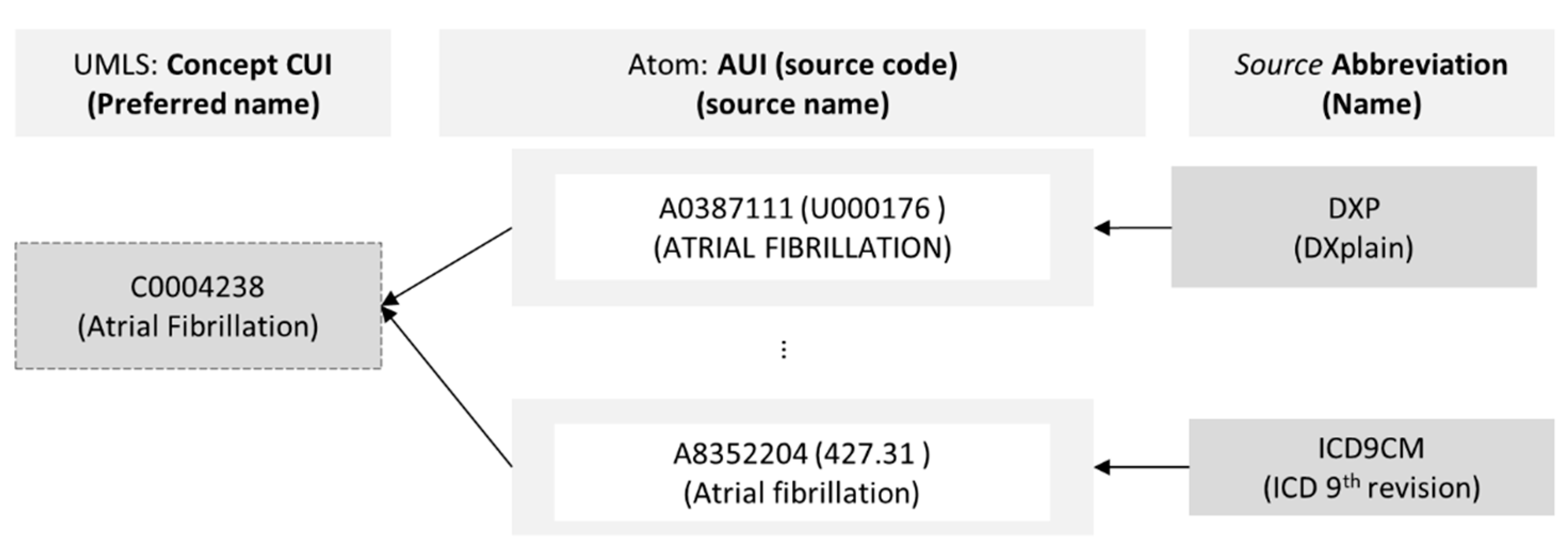
3.1. UMLS Knowledge Structure
3.2. FHIR Standard
3.2.1. FHIR Resources
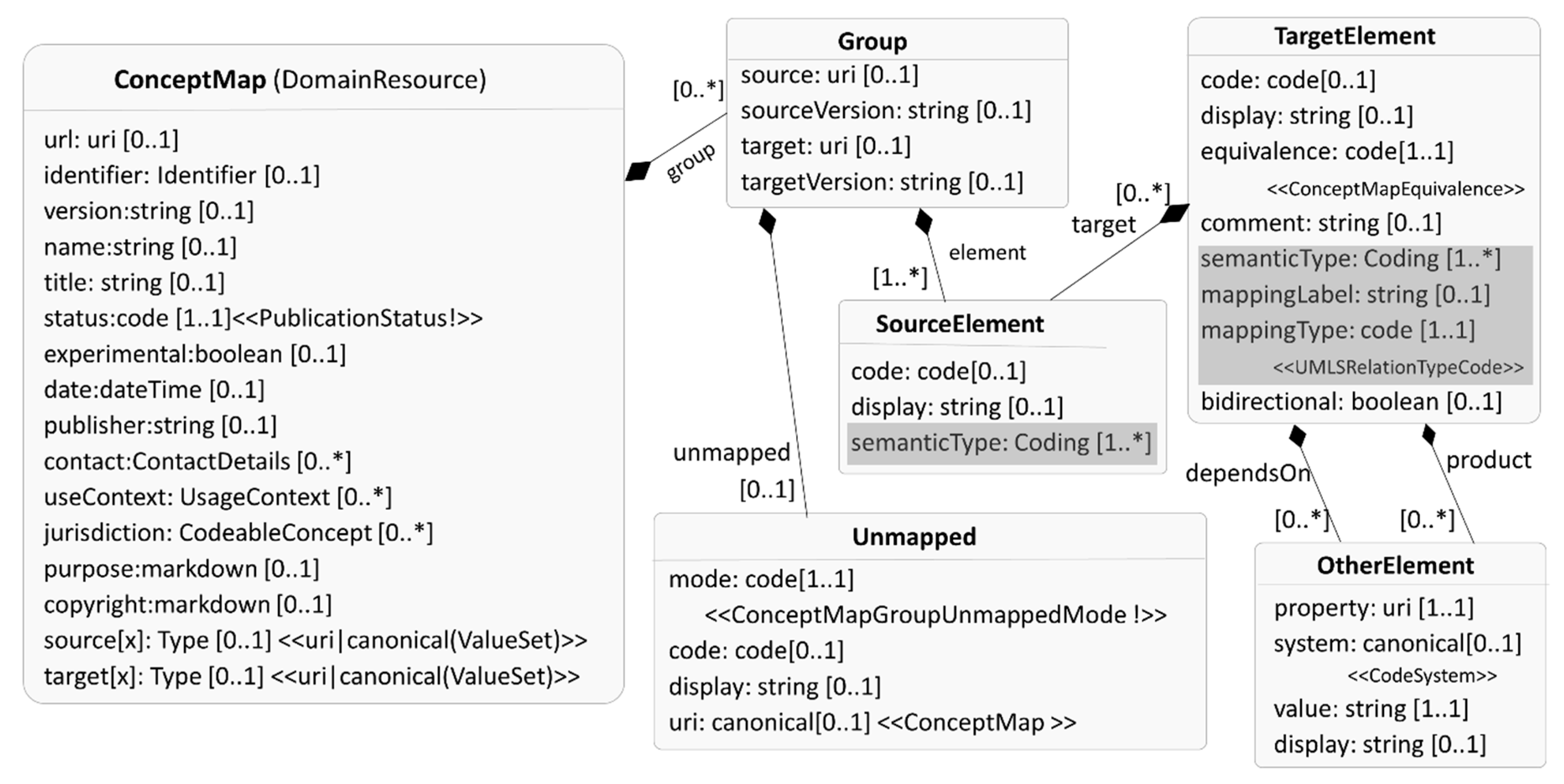
3.2.2. FHIR ConeptMap
3.2.3. FHIR CodeSystem
3.2.4. FHIR ValueSet
3.3. Comparing UMLS Knowledge Structure and FHIR ConceptMap
3.3.1. Similarities
- The equivalence attribute of TargetElement in ConceptMap is bound to ConceptMapEquivalence valueset, i.e., the equivalence attribute will only accept values defined in this ConceptMapEquivalence valueset. The ConceptMapEquivalence valueset defines ten values (e.g., equal, specializes, subsumes, wider, etc.), and these values describe the nature of the mapping between SourceElement and TargetElement. For example, when the equivalence attribute takes the value “equal”, the code captured by TargetElement is equal to the code captured by SourceElement;
3.3.2. Differences
- The ConceptMap cannot capture the label attribute of the relationship (Relationship class, Figure 3). For example, Atrial Fibrillation (source with CUI C0004238) is related to Digoxin (target with CUI C0012265), where the attribute’s type has the value related, and the label has the value may_be_treated_by.
- As mentioned previously, the equivalence attribute of TargetElement in ConceptMap is bound up with ConceptMapEquivalence valueset, and the values of the valueset describe the nature of the mapping between SourceElement and TargetElement. The equivalence attribute of the TargetElement class can be mapped to the type attribute of the UMLS relationship (Relationship class, Figure 3). A relationship type attribute can only take a value from the UMLS REL set. The mapping between the ConceptMapEquivalence valueset’s values and the REL set’s values is not exactly one to one. There is one to one mapping between a few values across the sets without loss of semantics, but for some values, there is no mapping. The lack of the mapping is either due to the unavailability of an equivalent value in the other set, or the fact that the official description of the value, particularly the values in the UMLS REL set, is vague, making it hard to determine a mapping. For example,
- ○
- The value PAR or CHD (representing hierarchy between UMLS concepts) from the REL set has an equivalent mapping to Subsumes or Specializes in ConceptMapEquivalence;
- ○
- The values AQ or QB or Empty from the REL set do not map to any value in the ConceptMapEquivalence valueset;
- ○
- The value RU in the UMLS REL set, described as “Related, unspecified”, is not clear, making it hard to determine a mapping to ConceptMapEquivalence values, and is hence unmapped.
- ConceptMap cannot capture the semantic type of a concept or atom.
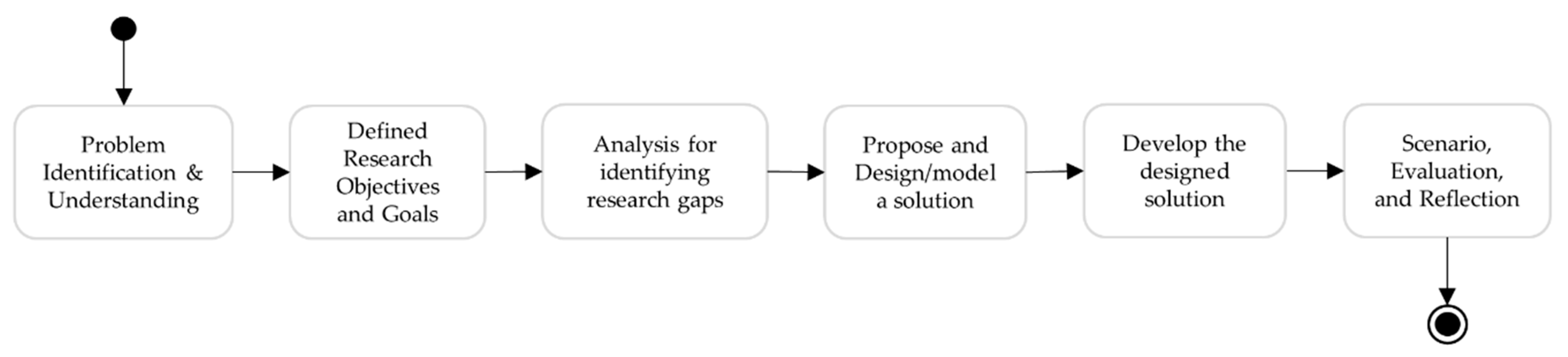
4. Method
- Step 1: Problem identification and understanding, and defining research objectives and goals, discussed in Section 1.
- Step 2: An in-depth analysis to understand the fundamental similarities and differences between the UMLS knowledge structure and ConceptMap, discussed in Section 3.3.
- Step 4: Implement the designed FHIR extensions and profiled ConceptMap using the HAPI library. Develop the UMLS FHIR terminology service using HAPI library, profiled ConceptMap and other FHIR terminological resources.
- Step 5: Demonstrate the terminology server, evaluate the results, and reflect on the research.
5. UMLS FHIR Terminology Service: Design and Implementation
5.1. Design: FHIR Extensions to ConceptMap
- An extension to capture the semantic type(s) of the SourceElement (UMLS concept) and TargetElement (Atom).
- An extension to capture the label attribute associated with the UMLS relationship (Relationship class, Figure 3).
- An extension to capture the type attribute of the UMLS relationship (Relationship class, Figure 3). As discussed, the type attribute only takes values from the UMLS REL set. Thus, the REL set is translated into an FHIR valuleset, named UMLSRelationTypeCode. This extension is bound to the UMLSRelationTypeCode valueset. This design is similar to the equivalence attribute of the TargetElement class in ConceptMap that is bound to the ConceptMapEquivalence valueset.
5.2. Implementation: FHIR Extensions and Profiled ConceptMap
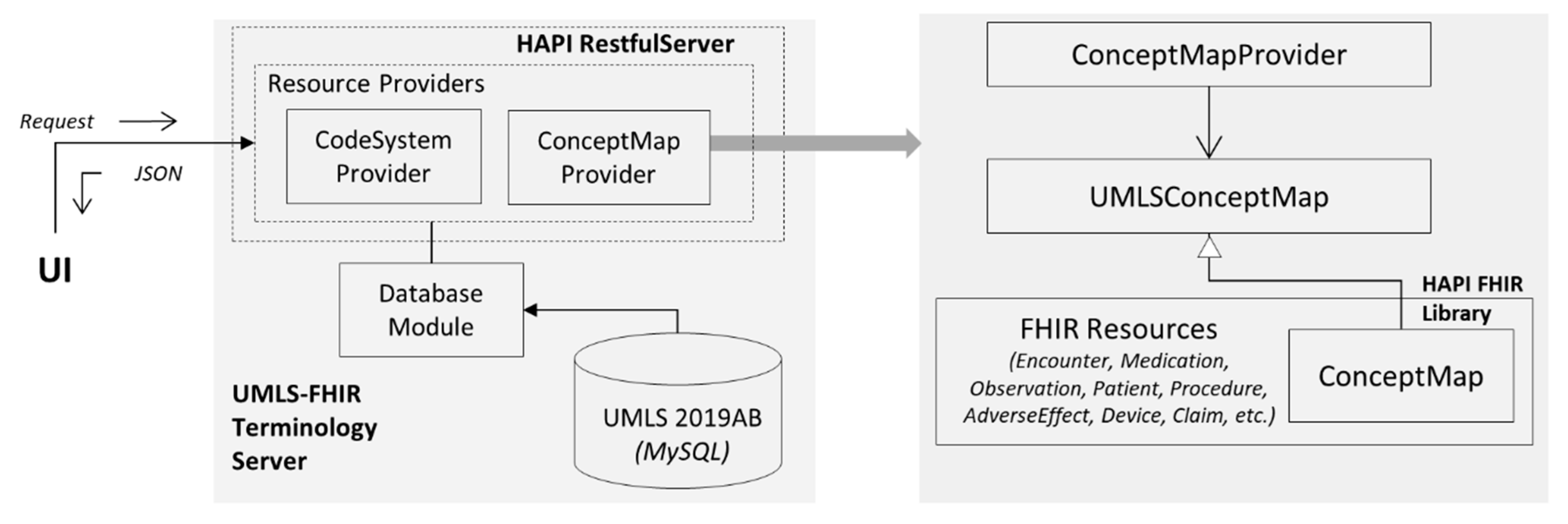
5.3. Architecture and Impementation: UMLS FHIR Terminology Service
- A user-facing client application written in HTML and JavaScript;
- UMLS FHIR API developed using HAPI REST Server, running on the Apache Tomcat web server;
- A MySQL database for UMLS.
6. Results
- GET/CodingSystem?search={searchString}
- ○
- Description: Searches for all UMLS concepts whose name (Figure 3) begins with the user input string, captured by the parameter “searchString” and returns the result as an FHIR Bundle resource. A Bundle is a container for the collection of resources, and a search operation on any FHIR resource returns a Bundle [10]. In this case, the bundle contains a collection of CodeSystem.
- ○
- Example: http://umls.it.ilstu.edu/umlsfhir/fhir/CodeSystem?search=malaria returns all the concepts in whose name starts with “malaria”.
- ○
- Responses: Table 3 lists the response codes, its description, and resources for this endpoint.
- GET/ConceptMap/{concept_cui}
- ○
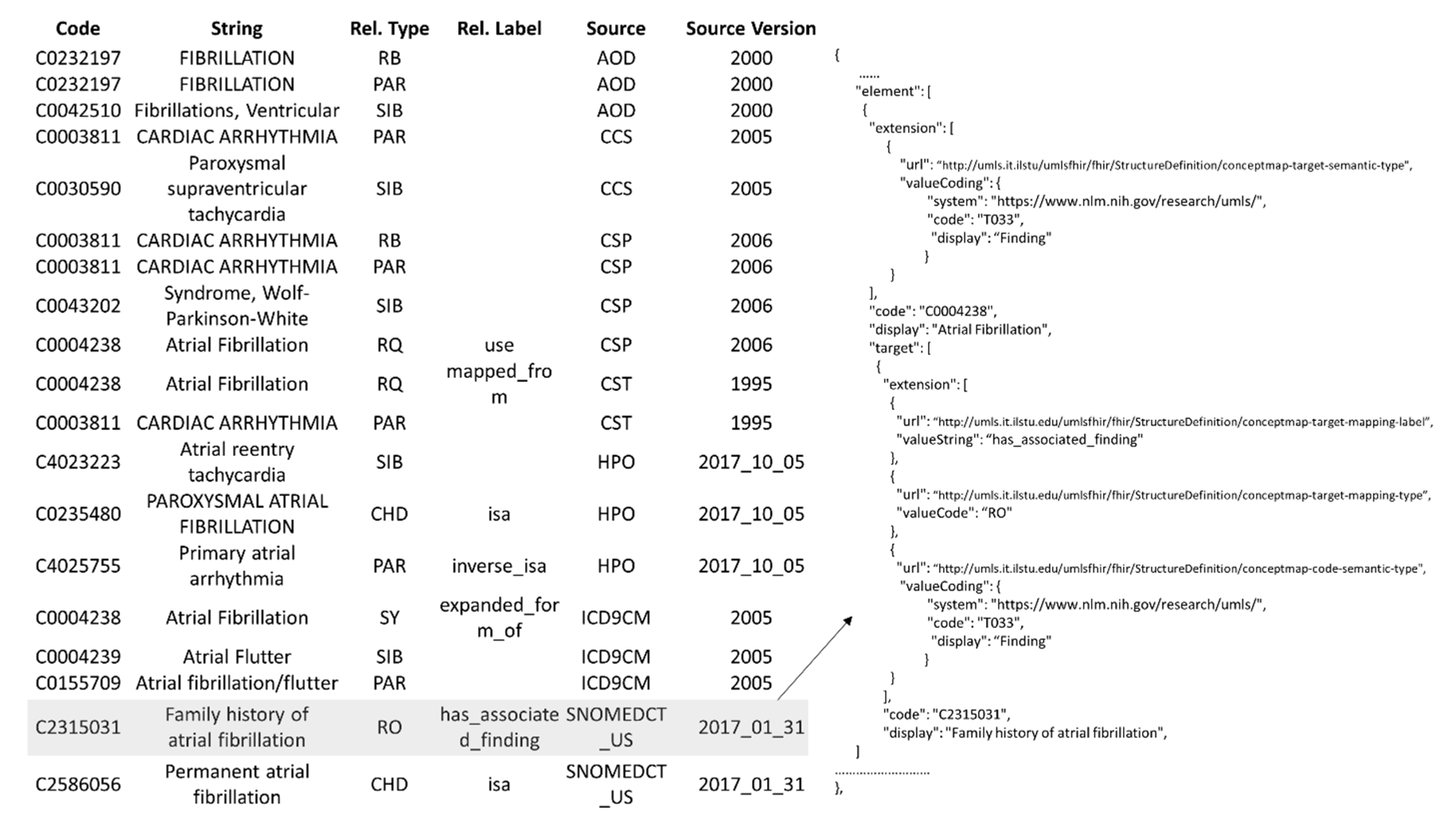
- Description: Searches for the UMLS concept using the user-provided identifier, captured by the “concept_cui” parameter, and returns the concept knowledge as a ConceptMap (Figure 7).
- ○
- Example: http://umls.it.ilstu.edu/umlsfhir/fhir/ConceptMap/C0004238 returns knowledge related to the concept identified by CUI C0004238 as ConceptMap.
- ○
- Response: Table 4 lists the response codes, its description, and resources for this endpoint.
7. Discussion
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A


References
- Schriml, L.M.; Arze, C.; Nadendla, S.; Chang, Y.W.W.; Mazaitis, M.; Felix, V.; Feng, G.; Kibbe, W.A. Disease Ontology: A backbone for disease semantic integration. Nucleic Acids Res. 2012, 40, 940. [Google Scholar] [CrossRef] [PubMed]
- Amos, L.; Anderson, D.; Brody, S.; Ripple, A.; Humphreys, B.L. UMLS users and uses: A current overview. J. Am. Med. Informatics Assoc. 2020, 27, 1606–1611. [Google Scholar] [CrossRef] [PubMed]
- Bodenreider, O. The Unified Medical Language System (UMLS): Integrating biomedical terminology. Nucleic Acids Res. 2004, 32, D26–D270. [Google Scholar] [CrossRef] [PubMed]
- Humphreys, B.L.; Lindberg, D.A.B.; Schoolman, H.M.; Barnett, G.O. The Unified Medical Language System: An Informatics Research Collaboration. J. Am. Med. Inform. Assoc. 1997, 5, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Medicine, N.L. Of UMLS® Reference Manual; National Library of Medicine: Bethesda, MD, USA, 2009. [Google Scholar]
- NLM UMLS REST API. Available online: https://documentation.uts.nlm.nih.gov/rest/home.html (accessed on 12 November 2020).
- Saripalle, R.K. Fast health interoperability resources (FHIR): Current status in the healthcare system. Int. J. E-Health Med. Commun. 2019, 10, 76–93. [Google Scholar] [CrossRef]
- Benson, T.; Grieve, G. Principles of Health Interoperability: SNOMED CT, HL7 and FHIR, 3rd ed.; Springer: Berlin, Germany, 2016. [Google Scholar]
- HL7 Fast Healthcare Interoperability Resources Documentation. Available online: https://www.hl7.org/fhir/documentation.html (accessed on 12 November 2020).
- HL HL7 FHIR Restful API. Available online: https://www.hl7.org/fhir/http.html (accessed on 5 February 2019).
- Boone, K.W. The CDATM Book, 2011th ed.; Springer: Berlin, Germany, 2011. [Google Scholar]
- Beale, T.; Heard, S. openEHR Architecture—Architecture Overview; openEHR Foundation, 2008; Available online: https://specifications.openehr.org/releases/1.0.2/architecture/overview.pdf (accessed on 12 November 2020).
- HAPI FHIR 2020. Available online: https://hapifhir.io/hapi-fhir/ (accessed on 12 November 2020).
- Tran, L.T.; Divita, G.; Carter, M.E.; Judd, J.; Samore, M.H.; Gundlapalli, A.V. Exploiting the UMLS Metathesaurus for extracting and categorizing concepts representing signs and symptoms to anatomically related organ systems. J. Biomed. Inform. 2015, 58, 19–27. [Google Scholar] [CrossRef]
- Plaza, L.; Diaz, A. Retrieval of Similar Electronic Health Records Using UMLS Concept Graphs. In Lecture Notes in Computer Science; Hopfe, C., Rezgui, Y., Metais, E., Preece, A., Li, H., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; Volume 6177, pp. 296–303. [Google Scholar] [CrossRef]
- Becker, M.; Bockmann, B. Extraction of UMLS Concepts Using Apache cTAKESTM for German Language. Stud. Health Technol. Inform. 2016, 223, 71–76. [Google Scholar]
- Shivade, C.; Malewadkar, P.; Fosler-Lussier, E.; Lai, A.M. Comparison of UMLS terminologies to identify risk of heart disease using clinical notes. J. Biomed. Inform. 2015, 58, S103–S110. [Google Scholar] [CrossRef]
- Divita, G.; Zeng, Q.T.; Gundlapalli, A.V.; Duvall, S.; Nebeker, J.; Samore, M.H. Sophia: A Expedient UMLS Concept Extraction Annotator. AMIA Annu. Symp. Proc. 2014, 2014, 467–476. [Google Scholar]
- Morrey, C.P.; Perl, Y.; Halper, M.; Chen, L.; Gu, H.H. A chemical specialty semantic network for the Unified Medical Language System. J. Cheminform. 2012, 4, 9. [Google Scholar] [CrossRef]
- Chen, Y.; Gu, H.; Perl, Y.; Geller, J. Overcoming an obstacle in expanding a UMLS semantic type extent. J. Biomed. Inform. 2012, 45, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Gu, H.; Perl, Y.; Halper, M.; Xu, J. Expanding the Extent of a UMLS Semantic Type via Group Neighborhood Auditing. J. Am. Med. Inform. Assoc. 2009, 16, 746–757. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Uzuner, O. Semi-Supervised Learning to Identify UMLS Semantic Relations. AMIA Summits Transl. Sci. Proc. 2014, 2014, 67–75. [Google Scholar] [PubMed]
- Gu, H.; He, Z.; Wei, D.; Elhanan, G.; Chen, Y. Validating UMLS semantic type assignments using SNOMED CT semantic tags. Methods Inf. Med. 2018, 57, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Detmer, W.M.; Barnett, G.O.; Hersh, W.R. MedWeaver: Integrating decision support, literature searching, and Web exploration using the UMLS Metathesaurus. In Proceedings of the AMIA Annual Fall Symposium; American Medical Informatics Association: Bethesda, MD, USA, 1997; pp. 490–494. [Google Scholar]
- Atal, I.; Zeitoun, J.-D.; Neveol, A.; Ravaud, P.; Porcher, R.; Trinquart, L. Automatic classification of registered clinical trials towards the Global Burden of Diseases taxonomy of diseases and injuries. BMC Bioinform. 2016, 17, 392. [Google Scholar] [CrossRef] [PubMed]
- Saripalle, R.; Runyan, C.; Russell, M. Using HL7 FHIR to achieve interoperability in patient health record. J. Biomed. Inform. 2019, 94. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Kiefer, R.; Prud’hommeaux, E.; Solbrig, H.R. Building Interoperable FHIR-Based Vocabulary Mapping Services: A Case Study of OHDSI Vocabularies and Mappings. Stud. Health Technol. Inform. 2017, 245, 1327. [Google Scholar]
- Metke-Jimenez, A.; Steel, J.; Hansen, D.; Lawley, M. Ontoserver: A syndicated terminology server. J. Biomed. Semant. 2018, 9. [Google Scholar] [CrossRef]
- LONIC FHIR Terminology Server. Available online: https://loinc.org/fhir/ (accessed on 12 November 2020).
- Zong, N.; Wen, A.; Stone, D.J.; Sharma, D.K.; Wang, C.; Yu, Y.; Liu, H.; Shi, Q.; Jiang, G. Developing an FHIR-Based Computational Pipeline for Automatic Population of Case Report Forms for Colorectal Cancer Clinical Trials Using Electronic Health Records. JCO Clin. Cancer Informatics 2020, 4, 201–209. [Google Scholar] [CrossRef]
- Zong, N.; Sharma, D.K.; Yu, Y.; Egan, J.B.; Davila, J.I.; Wang, C.; Jiang, G. Developing a FHIR-based Framework for Phenome Wide Association Studies: A Case Study with A Pan-Cancer Cohort. AMIA Jt. Summits Transl. Sci. Proc. AMIA Jt. Summits Transl. Sci. 2020, 2020, 750–759. [Google Scholar]
- Bangalore, A.; Thorn, K.E.; Tilley, C.; Peters, L. The UMLS knowledge source server: An object model for delivering UMLS data. AMIA Annu. Symp. Proc. 2003, 2003, 51–55. [Google Scholar]
- UMLS Database Query Diagrams: How to Find All Information Associated with a Particular UMLS Concept. Available online: https://www.nlm.nih.gov/research/umls/implementation_resources/query_diagrams/er1.html (accessed on 12 November 2020).
- HL7 ConceptMap. Available online: http://hl7.org/implement/standards/fhir/conceptmap.html (accessed on 12 November 2020).
- HAPI Custom Structures. Available online: https://hapifhir.io/hapi-fhir/docs/model/custom_structures.html (accessed on 12 November 2020).
- HAPI REST Server Type. Available online: https://hapifhir.io/hapi-fhir/docs/server_plain/server_types.html (accessed on 12 November 2020).
- Resource Providers and Plan Providers—HAPI FHIR Documentation. Available online: https://hapifhir.io/hapi-fhir/docs/server_plain/resource_providers.html (accessed on 7 November 2020).
- Rocha, R.A.; Huff, S.M.; Haug, P.J.; Warner, H.R. Designing acontrolled medical vocabulary server: The VOSER project. Comput. Biomed. Res. 1994, 27, 472–507. [Google Scholar] [CrossRef] [PubMed]
- Rector, A.L.; Solomon, W.D.; Nowlan, W.A.; Rush, T.W.; Zanstra, P.E.; Claassen, W.M. A Terminology Server for medical language and medical information systems. Methods Inf. Med. 1995, 34, 147–157. [Google Scholar]


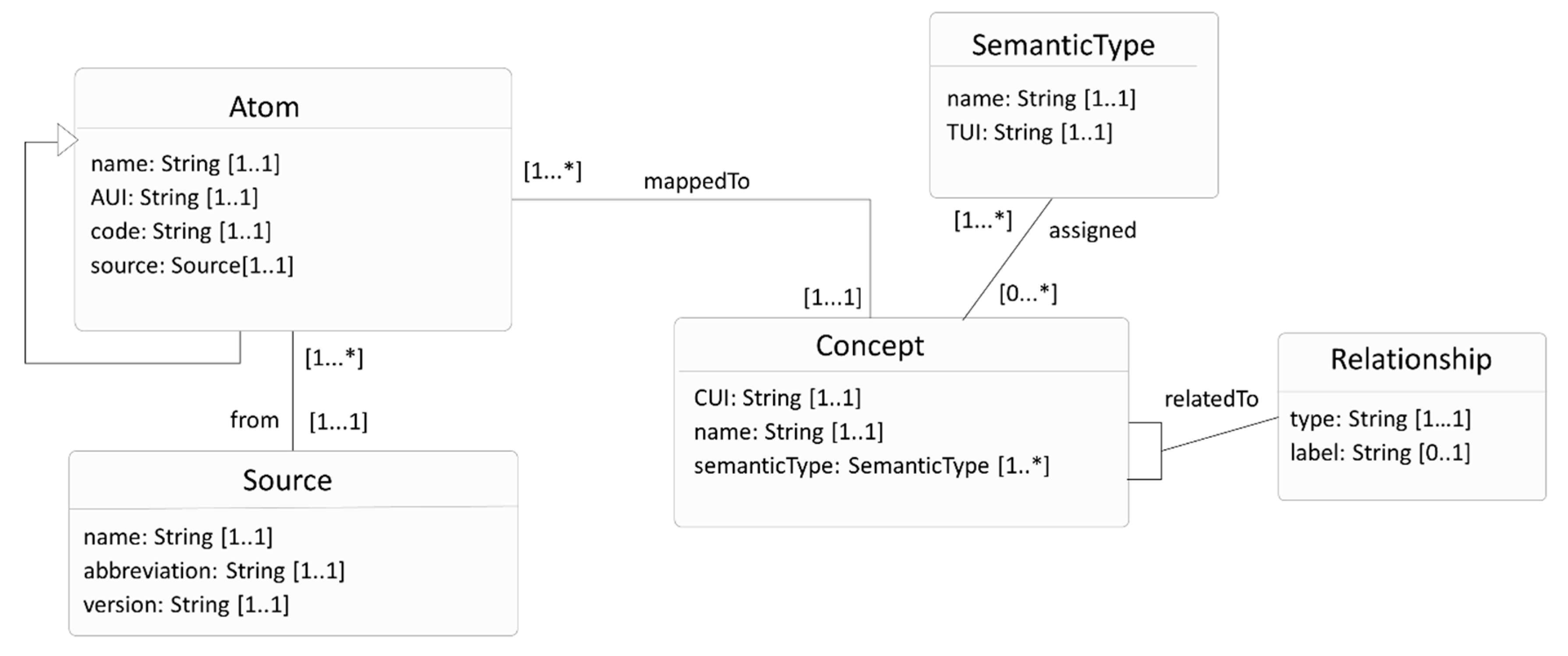
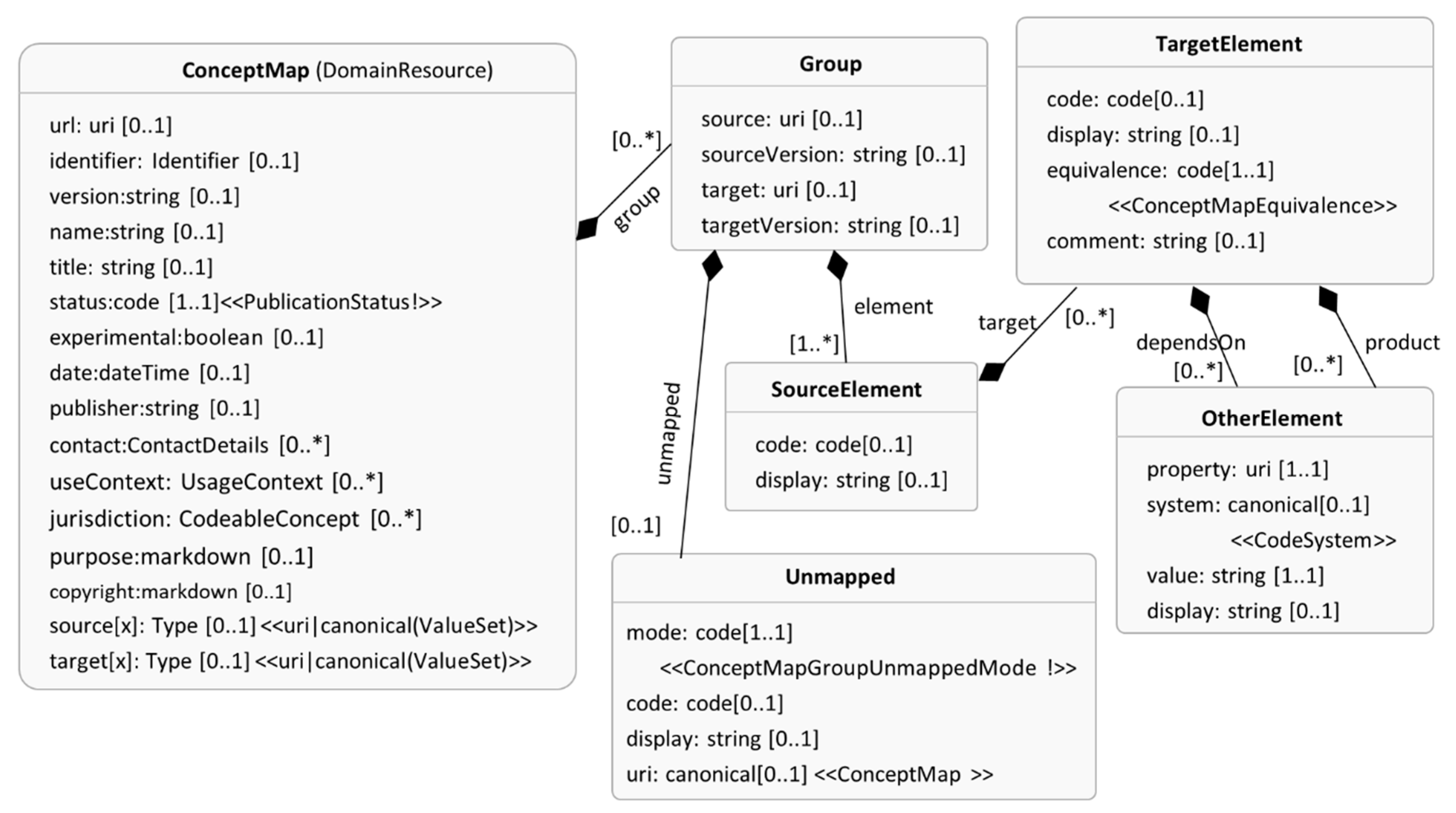



| UMLS Knowledge Structure (Figure 3) | FHIR ConceptMap (Figure 4) |
|---|---|
| Concept.CUI | ConceptMap.Group.SourceElement.code |
| Concept.name | ConceptMap.Group.SourceElement.display |
| Concept.semanticType | ---- |
| Atom.AUI | ConeptMap.Group.TargetElement.code |
| Atom.name | ConceptMap.Group.TargetElement.display |
| Mapping of atoms to a CUI (mapped_to) | ConceptMap.Group.TargetElement.equivalence |
| Relationship.type (Relationship between CUI) | ---- |
| Relationship.label | ---- |
| Source.name (Source standard) | ---- |
| Source.uri | ConceptMap.Group.target |
| Source.version | ConceptMap.Group.targetVersion |
| UMLS REL Values (Description) | ConceptMapEquivalence Values (Description) |
|---|---|
| REL set value with equivalence in ConceptMapEquivalence | |
| PAR (has parent relationship) | subsumes (The target mapping subsumes the meaning of the source concept) |
| CHD (has child relationship) | specializes (The target mapping specializes the meaning of the source concept) |
| SIB (has sibling relationship) | specializes |
| RB (has broader relationship) | wider (The target mapping is wider in meaning than the source concept) |
| RN (has narrower relationship) | narrower (The target mapping is narrower in meaning than the source concept) |
| XR (Not related, no mapping) | unmatched (There is no match for this concept in the target code system.) |
| SY (source asserted synonymy) | equivalent (The definitions of the concepts mean the same thing (including when structural implications of meaning are considered) (i.e., extensionally identical). |
| RQ (related and possible synonymous) | relatedto (The concepts are related to each other, and have at least some overlap in meaning, but the exact relationship is not known.) |
| RL (The relationships are similar or “alike”. The two concepts are similar or “alike”) | relatedto |
| REL set value with no equivalence in ConceptMapEquivalence and vice versa | |
| RO (has relationship other than synonymous, narrower, or broader) | ---- |
| AQ (Allowed qualifier) | ---- |
| DEL (deleted concept) | ---- |
| QB (can be qualified by) | ---- |
| RU (related, unspecified) | ---- |
| (empty relationship) | ---- |
| ---- | inexact (The target mapping overlaps with the source concept, but both source and target cover additional meaning, or the definitions are imprecise and it is uncertain whether they have the same boundaries to their meaning.) |
| ---- | disjoint (There is no match for this concept in the target code system.) |
| Code | Status | Description | Resource (FHIR Resource) |
|---|---|---|---|
| 200 | OK | Returns all UMLS concepts whose name begins with the search string. | Bundle |
| 400 | Bad Request | Invalid request. The endpoint does not know how to handle the request. | None |
| Code | Status | Description | Resource (FHIR Resource) |
|---|---|---|---|
| 200 | OK | Returns knowledge of the UMLS concept identified by the concept_cui | ConceptMap |
| 400 | Bad Request | Invalid request. The endpoint does not know how to handle the request | None |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saripalle, R.; Sookhak, M.; Haghparast, M. An Interoperable UMLS Terminology Service Using FHIR. Future Internet 2020, 12, 199. https://doi.org/10.3390/fi12110199
Saripalle R, Sookhak M, Haghparast M. An Interoperable UMLS Terminology Service Using FHIR. Future Internet. 2020; 12(11):199. https://doi.org/10.3390/fi12110199
Chicago/Turabian StyleSaripalle, Rishi, Mehdi Sookhak, and Mahboobeh Haghparast. 2020. "An Interoperable UMLS Terminology Service Using FHIR" Future Internet 12, no. 11: 199. https://doi.org/10.3390/fi12110199
APA StyleSaripalle, R., Sookhak, M., & Haghparast, M. (2020). An Interoperable UMLS Terminology Service Using FHIR. Future Internet, 12(11), 199. https://doi.org/10.3390/fi12110199