An Improved Method for Named Entity Recognition and Its Application to CEMR †
Abstract
:1. Introduction
2. Related Work
3. Method
3.1. Pretreatment
3.1.1. Data Formatting
3.1.2. Tagging Scheme
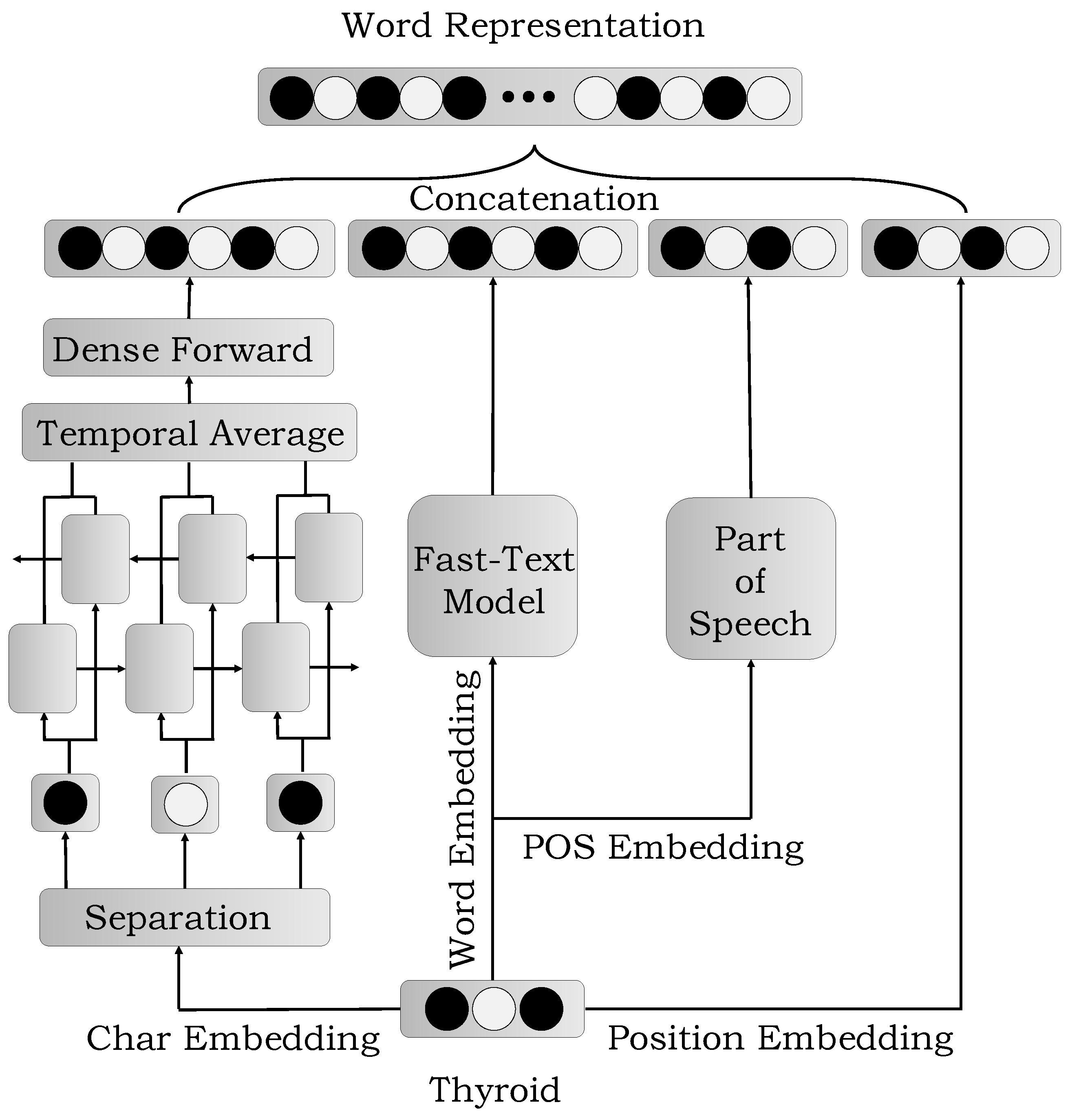
3.2. The Construction of Features
3.2.1. Word Embedding
3.2.2. Char Embedding
3.2.3. POS Embedding
3.2.4. Position Embedding
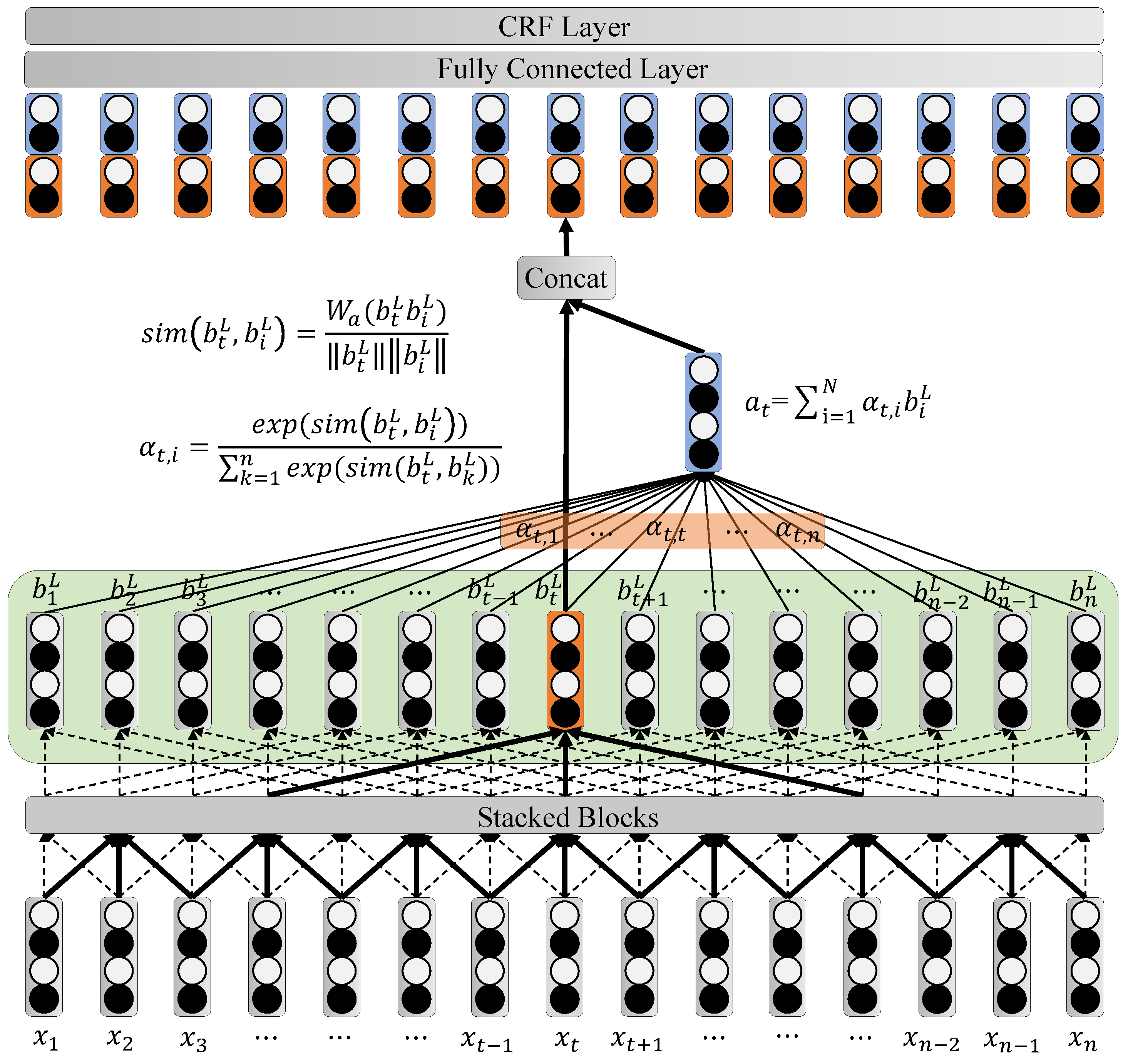
3.3. Attention-Based ID-CNNs-CRF Model
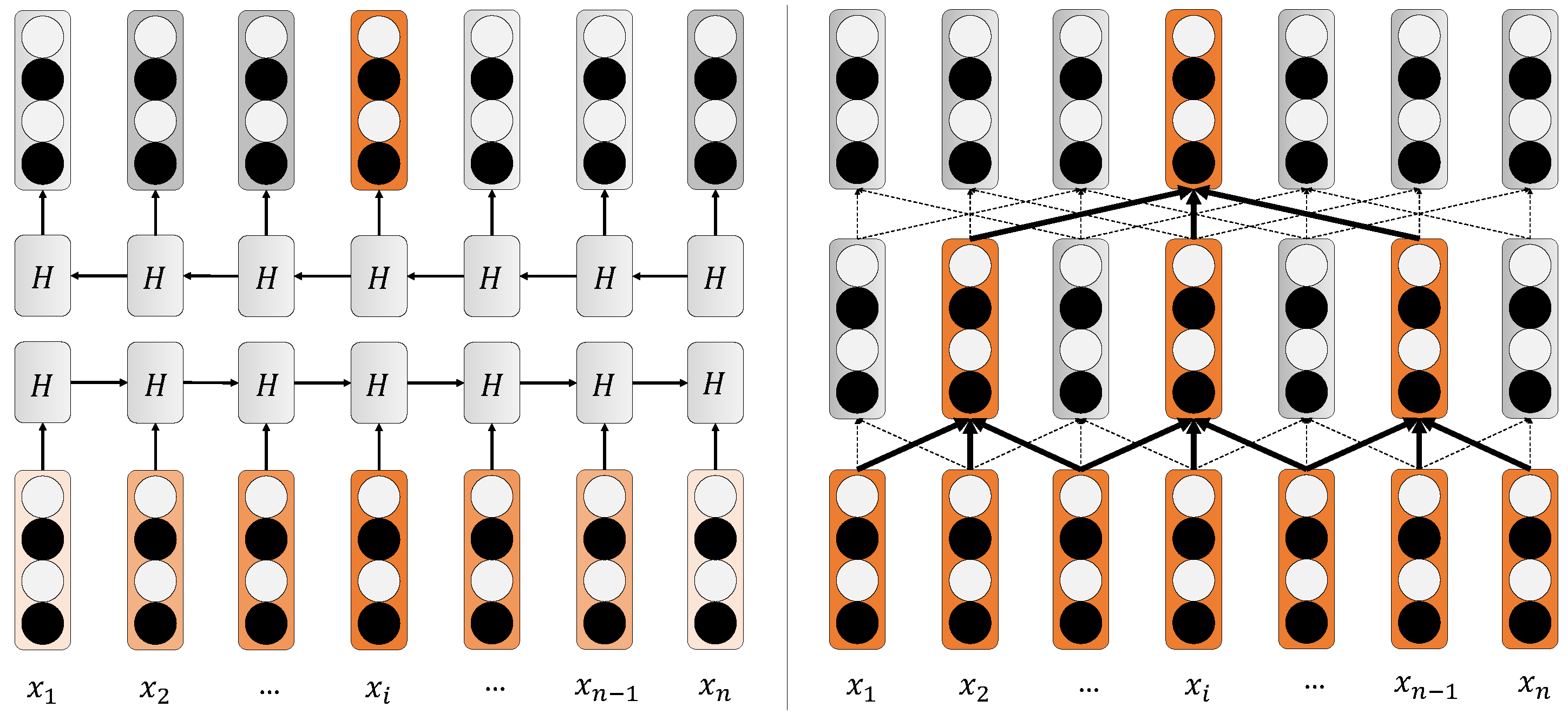
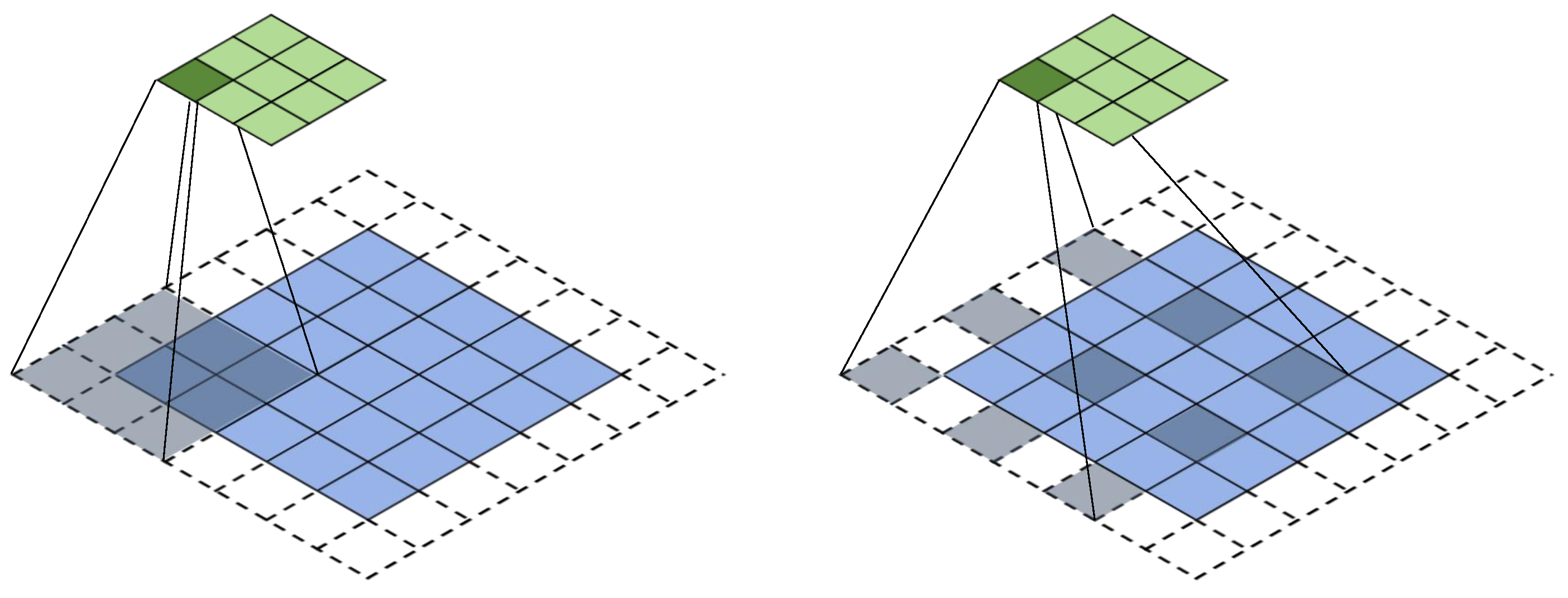
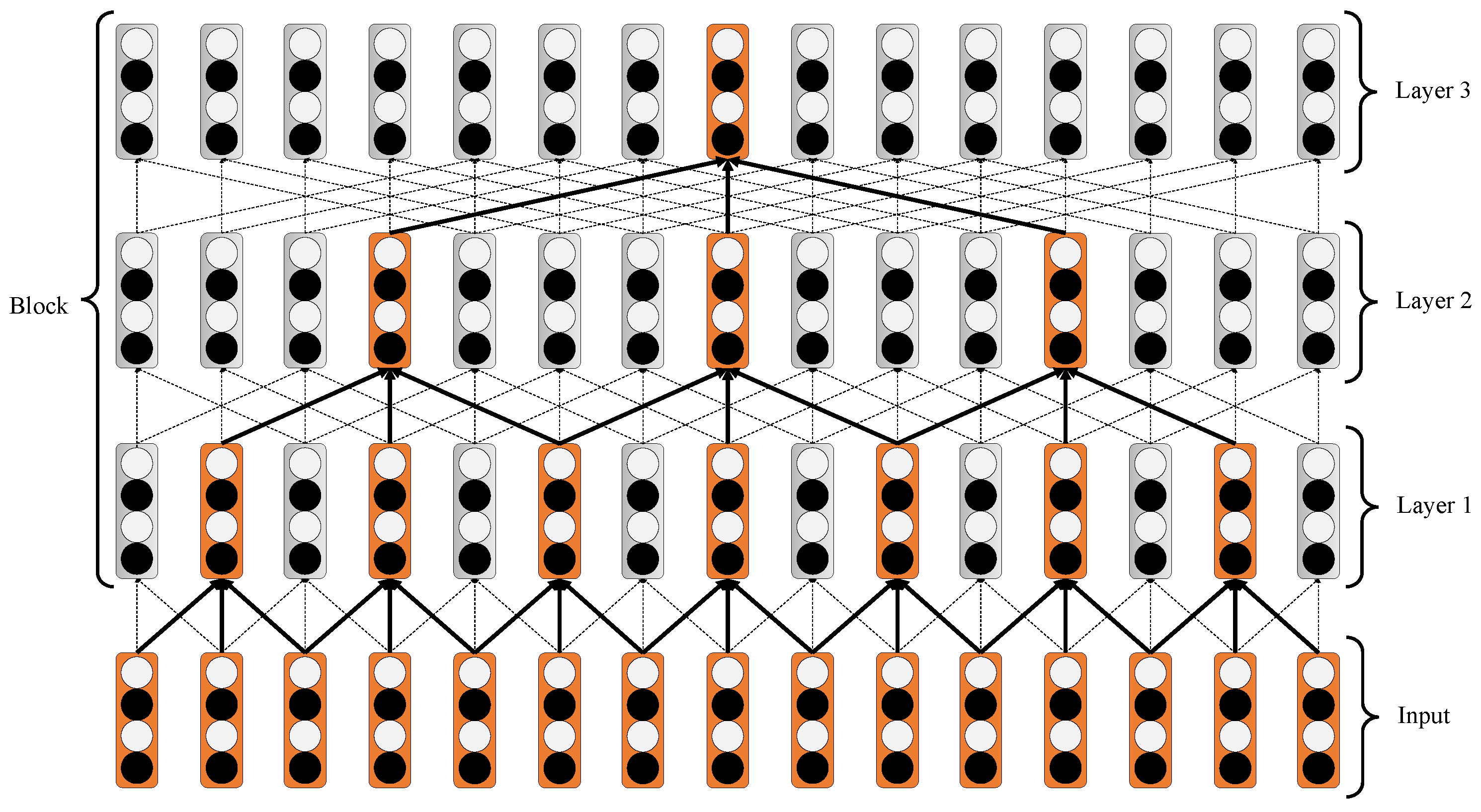
3.3.1. Iterated Dilated CNNs
3.3.2. Attention Mechanism
3.3.3. Linear Chain CRF
4. Experiments and Analysis
4.1. Datasets
4.2. Parameter Setting
4.3. Evaluation Metrics
4.4. Comparison with Other Methods
4.5. Comparison of Entity Category
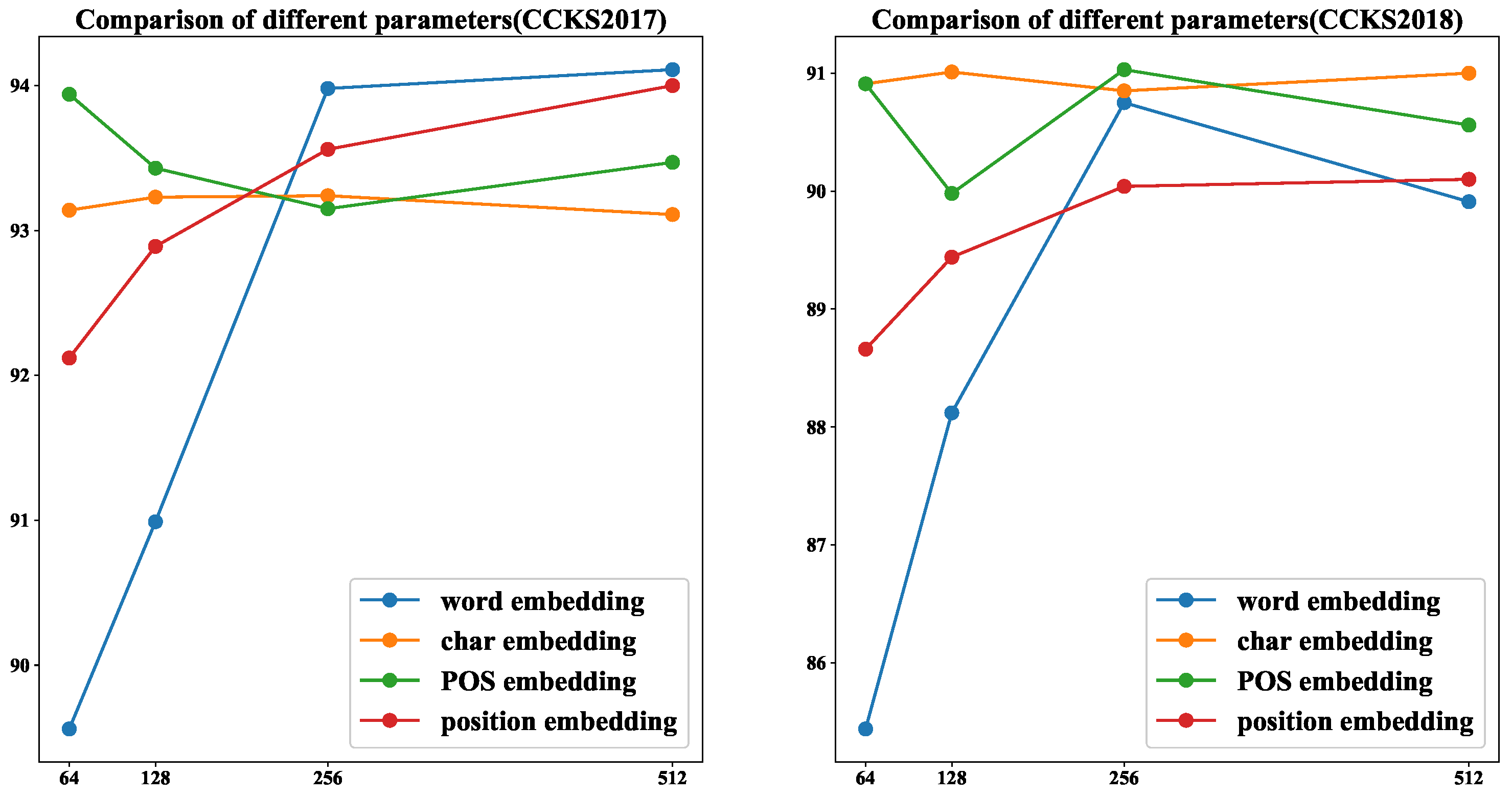
4.6. Comparison of Different Parameters
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lample, G.; Ballesteros, M.; Subramanian, S.; Kawakami, K.; Dyer, C. Neural architectures for named entity recognition. arXiv 2004, arXiv:1603.01360. [Google Scholar]
- Ma, X.; Hovy, E. End-to-end sequence labeling via bi-directional lstm-cnns-crf. arXiv 2016, arXiv:1603.01354. [Google Scholar]
- Rondeau, M.A.; Su, Y. LSTM-Based NeuroCRFs for Named Entity Recognition. In Proceedings of the Interspeech, San Francisco, SF, USA, 8–12 September 2016; pp. 665–669. [Google Scholar]
- Rei, M.; Crichton, G.K.; Pyysalo, S. Attending to characters in neural sequence labeling models. arXiv 2016, arXiv:1611.04361. [Google Scholar]
- Yin, W.; Kann, K.; Yu, M.; Schütze, H. Comparative study of CNN and RNN for natural language processing. arXiv 2017, arXiv:1702.01923. [Google Scholar]
- Minh, D.L.; Sadeghi-Niaraki, A.; Huy, H.D.; Min, K.; Moon, H. Deep learning approach for short-term stock trends prediction based on two-stream gated recurrent unit network. IEEE Access 2018, 6, 55392–55404. [Google Scholar] [CrossRef]
- Collobert, R.; Weston, J.; Bottou, L.; Karlen, M.; Kavukcuoglu, K.; Kuksa, P. Natural language processing (almost) from scratch. J. Mach. Learn. Res. 2011, 12, 2493–2537. [Google Scholar]
- Wang, C.; Chen, W.; Xu, B. Named entity recognition with gated convolutional neural networks. In Chinese Computational Linguistics and Natural Language Processing Based on Naturally Annotated Big Data; Springer: Cham, Germany; Basel, Switzerland, 2017; pp. 110–121. [Google Scholar]
- Strubell, E.; Verga, P.; Belanger, D.; McCallum, A. Fast and accurate sequence labeling with iterated dilated convolutions. arXiv 2017, arXiv:1702.02098, 138. [Google Scholar]
- Hirschman, L.; Morgan, A.A.; Yeh, A.S. Rutabaga by any other name: extracting biological names. J. Biomed. Inf. 2002, 35, 247–259. [Google Scholar] [CrossRef] [Green Version]
- Tsai, R.T.H.; Sung, C.L.; Dai, H.J.; Hung, H.C.; Sung, T.Y.; Hsu, W.L. NERBio: Using selected word conjunctions, term normalization, and global patterns to improve biomedical named entity recognition. In Proceedings of the Fifth International Conference on Bioinformatics, New Delhi, India, 18–20 December 2006. [Google Scholar]
- Tsuruoka, Y.; Tsujii, J.I. Boosting precision and recall of dictionary-based protein name recognition. In Proceedings of the ACL 2003 workshop on Natural language processing in biomedicine, Sapporo, Japan, 11 July 2003; pp. 41–48. [Google Scholar]
- Yang, Z.; Lin, H.; Li, Y. Exploiting the performance of dictionary-based bio-entity name recognition in biomedical literature. Comput. Biol. Chem. 2008, 32, 287–291. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Ruonan, R. The method of medical named entity recognition based on semantic model and improved svm-knn algorithm. In Proceedings of the 2011 Seventh International Conference on Semantics, Knowledge and Grids, Beijing, China, 24–26 October 2011; pp. 21–27. [Google Scholar]
- Collier, N.; Nobata, C.; Tsujii, J.I. Extracting the names of genes and gene products with a hidden markov model. In Proceedings of the 18th conference on Computational linguistics, Saarbrücken, Germany, 31 July–4 August 2000; pp. 201–207. [Google Scholar]
- GuoDong, Z.; Jian, S. Exploring deep knowledge resources in biomedical name recognition. In Proceedings of the International Joint Workshop on Natural Language Processing in Biomedicine and its Applications, Geneva, Switzerland, 28–29 August 2004; pp. 96–99. [Google Scholar]
- Chieu, H.L.; Ng, H.T. Named entity recognition with a maximum entropy approach. In Proceedings of the Seventh Conference on Natural Language Learning at HLT-NAACL 2003, Edmonton, AB, Canada, 27 May–1 June 2003; pp. 160–163. [Google Scholar]
- Leaman, R.; Islamaj Doğan, R.; Lu, Z. Dnorm: disease name normalization with pairwise learning to rank. Bioinformatics 2013, 29, 2909–2917. [Google Scholar] [CrossRef] [PubMed]
- Kaewphan, S.; Van Landeghem, S.; Ohta, T.; Van de Peer, Y.; Ginter, F.; Pyysalo, S. Cell line name recognition in support of the identification of synthetic lethality in cancer from text. Bioinformatics 2015, 32, 276–282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, Q.; Li, X.; Conesa, A.; Pereira, C. Gram-cnn: A deep learning approach with local context for named entity recognition in biomedical text. Bioinformatics 2015, 34, 1547–1554. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Zhou, Z.; Gong, T.; Hao, T.; Liu, W. Sblc: a hybrid model for disease named entity recognition based on semantic bidirectional lstms and conditional random fields. In Proceedings of the 2018 Sino-US Conference on Health Informatics, Guangzhou, China, 28 June–1 July 2018. [Google Scholar]
- Chowdhury, S.; Dong, X.; Qian, L.; Li, X.; Guan, Y.; Yang, J.; Yu, Q. A multitask bi-directional rnn model for named entity recognition on chinese electronic medical records. BMC Bioinform. 2018, 19. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Yang, Z.; Yang, P.; Zhang, Y.; Wang, L.; Lin, H.; Wang, J. An attention-based bilstm-crf approach to document-level chemical named entity recognition. Bioinformatics 2017, 34, 1381–1388. [Google Scholar] [CrossRef] [PubMed]
- Sang, E.F.; Veenstra, J. Representing text chunks. In Proceedings of the Conference on European Chapter of the Association for Computational Linguistics, Bergen, Norway, 8–12 June 1999. [Google Scholar]
- Lai, S.; Liu, K.; He, S.; Zhao, J. How to generate a good word embedding. IEEE Intell. Syst. 2016, 31, 5–14. [Google Scholar] [CrossRef]
- Joulin, A.; Grave, E.; Bojanowski, P.; Mikolov, T. Bag of tricks for efficient text classification. arXiv 2016, arXiv:1607.01759. [Google Scholar]
- Le, Q.; Mikolov, T. Distributed representations of sentences and documents. In Proceedings of the International Conference on International Conference on Machine Learning, Beijing, China, 21–26 June 2014. [Google Scholar]
- Available online: http://case.medlive.cn/all/case-case/index.html?ver=branch (accessed on 21 August 2019). (In Chinese).
- Available online: https://github.com/fxsjy/jieba (accessed on 21 August 2019).
- Yu, F.; Koltun, V. Multi-scale context aggregation by dilated convolutions. arXiv 2015, arXiv:1511.07122. [Google Scholar]
- Bharadwaj, A.; Mortensen, D.; Dyer, C.; Carbonell, J. Phonologically aware neural model for named entity recognition in low resource transfer settings. In Proceedings of the 2016 Conference on Empirical Methods in Natural Language Processing, Austin, TX, USA, 1–5 November 2016; pp. 1462–1472. [Google Scholar]
- Li, J.; Zhou, M.; Qi, G.; Lao, N.; Ruan, T.; Du, J. Knowledge Graph and Semantic Computing. Language, Knowledge, and Intelligence; Springer: Singapore, 2018. [Google Scholar]
- Zhao, J.; Van Harmelen, F.; Tang, J.; Han, X.; Wang, Q.; Li, X. Knowledge Graph and Semantic Computing. Knowledge Computing and Language Understanding; Springer: Singapore, 2019. [Google Scholar]
| CCKS2017 | CCKS2018 | ||||||
|---|---|---|---|---|---|---|---|
| Entity | Count | Train | Test | Entity | Count | Train | Test |
| Symptom | 7831 | 6846 | 1345 | Symptom | 3055 | 2199 | 856 |
| Check | 9546 | 7887 | 1659 | Description | 2066 | 1529 | 537 |
| Treatment | 1048 | 853 | 195 | Operation | 1116 | 924 | 192 |
| Disease | 722 | 515 | 207 | Drug | 1005 | 884 | 121 |
| Body | 10,719 | 8942 | 1777 | Body | 7838 | 6448 | 1390 |
| Total | 29,866 | 24,683 | 5183 | Total | 15,080 | 11,984 | 3096 |
| Module | Parameter Name | Value |
|---|---|---|
| Word Representation | word embedding dim | 256 |
| char embedding dim | 64 | |
| POS embedding dim | 64 | |
| position embedding dim | 128 | |
| Iterated Dilated CNNs | filter width | 3 |
| numfilter | 256 | |
| dilation | [1,1,2] | |
| block number | 4 | |
| Other | learning rate | 1e-3 |
| dropout | 0.5 | |
| gradient clipping | 5 | |
| batch size | 64 | |
| epoch | 40 |
| Word Feature | POS Feature | Description |
|---|---|---|
| :%W[−1,0] | :%P[−1,0] | previous word (POS) |
| :%W[0,0] | :%P[0,0] | current word (POS) |
| :%W[1,0] | :%P[1,0] | next word (POS) |
| :%W[0,0] %W[−1,0] | :%P[0,0] %P[−1,0] | current word and previous word (POS) |
| :%W[0,0] %W[1,0] | :%P[0,0] %P[1,0] | current word and next word (POS) |
| Model | CCKS2017 | CCKS2018 | ||||
|---|---|---|---|---|---|---|
| Precision | Recall | F1-Score | Precision | Recall | F1-Score | |
| % | % | % | % | % | % | |
| CRF [19] | 89.18 | 81.60 | 85.11 | 92.67 | 72.10 | 77.58 |
| LSTM-CRF [1] | 87.73 | 87.00 | 87.24 | 82.61 | 81.70 | 82.08 |
| Bi-LSTM-CRF [2] | 94.73 | 93.29 | 93.97 | 90.43 | 90.49 | 90.44 |
| ID-CNNs-CRF [9] | 88.20 | 87.15 | 87.47 | 81.27 | 81.42 | 81.34 |
| Attention-ID-CNNs-CRF(ours) | 94.15 | 94.63 | 94.55 | 91.11 | 91.25 | 91.17 |
| Model (512 Test Data) | Time (s) | Speed |
|---|---|---|
| Bi-LSTM-CRF [2] | 15.62 | 1.0× |
| ID-CNNs-CRF [9] | 11.96 | 1.31× |
| Attention-ID-CNNs-CRF (ours) | 12.81 | 1.22× |
| Model | CCKS2017 | |||||
| Body | Check | Disease | Signs | Treatment | Average | |
| Bi-LSTM-CRF [2] | 94.81 | 96.40 | 87.79 | 95.82 | 88.10 | 93.97 |
| ID-CNNs-CRF [9] | 90.89 | 88.28 | 79.67 | 91.04 | 81.27 | 87.47 |
| Attention-ID-CNNs-CRF (ours) | 95.38 | 97.79 | 86.55 | 96.91 | 87.64 | 94.55 |
| Model | CCKS2018 | |||||
| Body | Symptom | Operation | Drug | Description | Average | |
| Bi-LSTM-CRF [2] | 92.59 | 93.12 | 87.43 | 82.86 | 86.33 | 90.44 |
| ID-CNNs-CRF [9] | 84.37 | 85.81 | 76.59 | 80.42 | 84.61 | 81.34 |
| Attention-ID-CNNs-CRF (ours) | 95.18 | 94.47 | 85.86 | 80.06 | 91.99 | 91.17 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, M.; Xiao, Q.; Wu, S.; Deng, K. An Improved Method for Named Entity Recognition and Its Application to CEMR. Future Internet 2019, 11, 185. https://doi.org/10.3390/fi11090185
Gao M, Xiao Q, Wu S, Deng K. An Improved Method for Named Entity Recognition and Its Application to CEMR. Future Internet. 2019; 11(9):185. https://doi.org/10.3390/fi11090185
Chicago/Turabian StyleGao, Ming, Qifeng Xiao, Shaochun Wu, and Kun Deng. 2019. "An Improved Method for Named Entity Recognition and Its Application to CEMR" Future Internet 11, no. 9: 185. https://doi.org/10.3390/fi11090185
APA StyleGao, M., Xiao, Q., Wu, S., & Deng, K. (2019). An Improved Method for Named Entity Recognition and Its Application to CEMR. Future Internet, 11(9), 185. https://doi.org/10.3390/fi11090185





