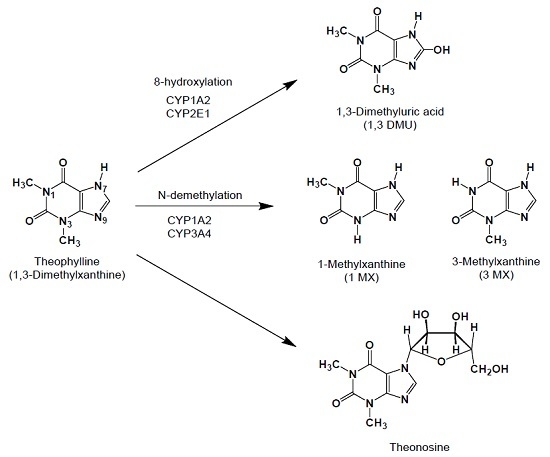
Theophylline-7β-d-Ribofuranoside (Theonosine), a New Theophylline Metabolite Generated in Human and Animal Lung Tissue
Abstract
:1. Introduction
2. Experimental Section
2.1. Tissue Sources and Collection
2.2. Methods
2.2.1. Preparation of Microsomes
2.2.2. In Vitro Metabolism Studies
2.2.3. Synthesis of Theophylline-7β-d-Ribofuranoside
2.2.4. Chemicals
2.2.5. Data Analyses
3. Results and Discussion
3.1. Microsomal Preparation
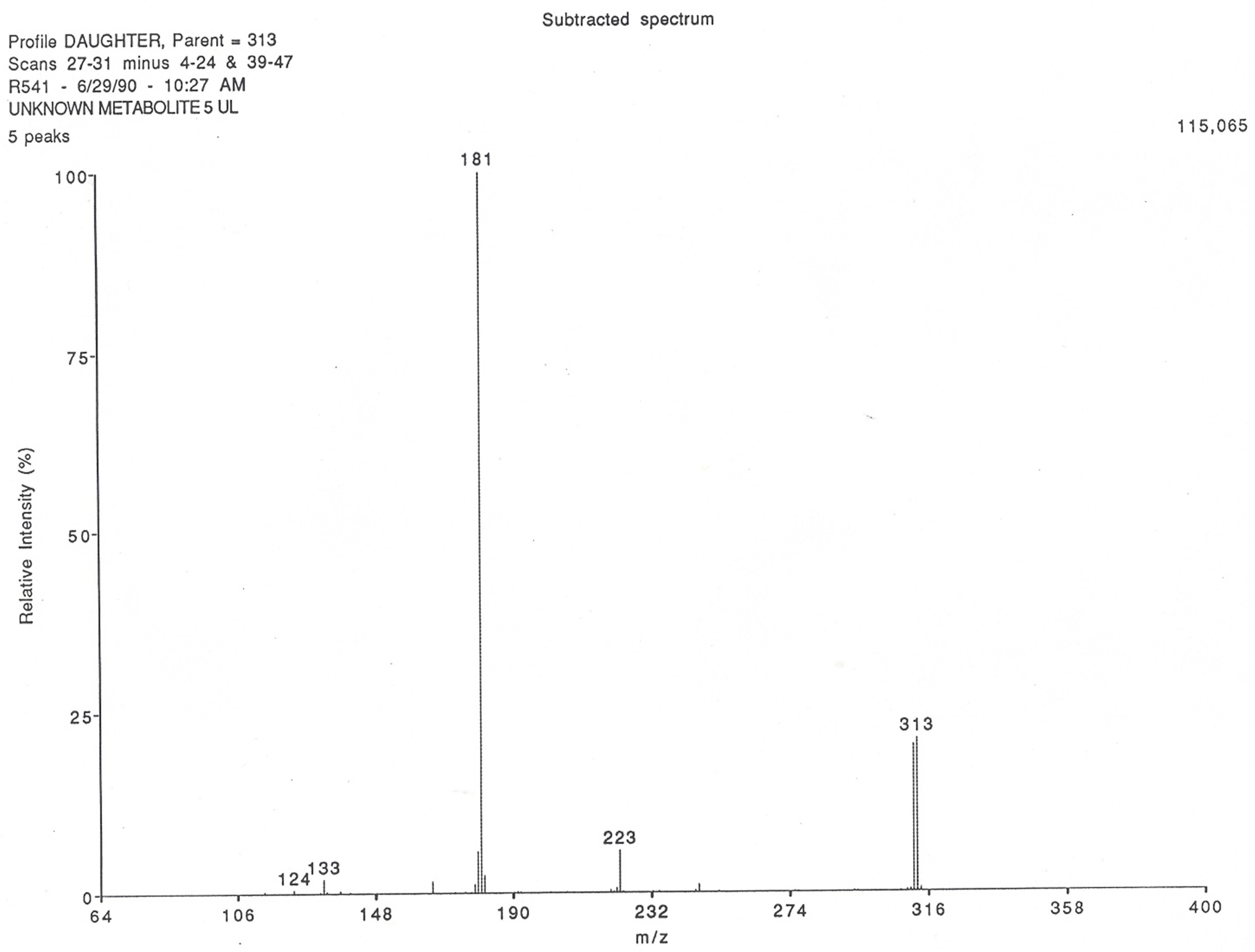
3.2. Identification and Characterization of the Production of the New Theophylline Metabolite
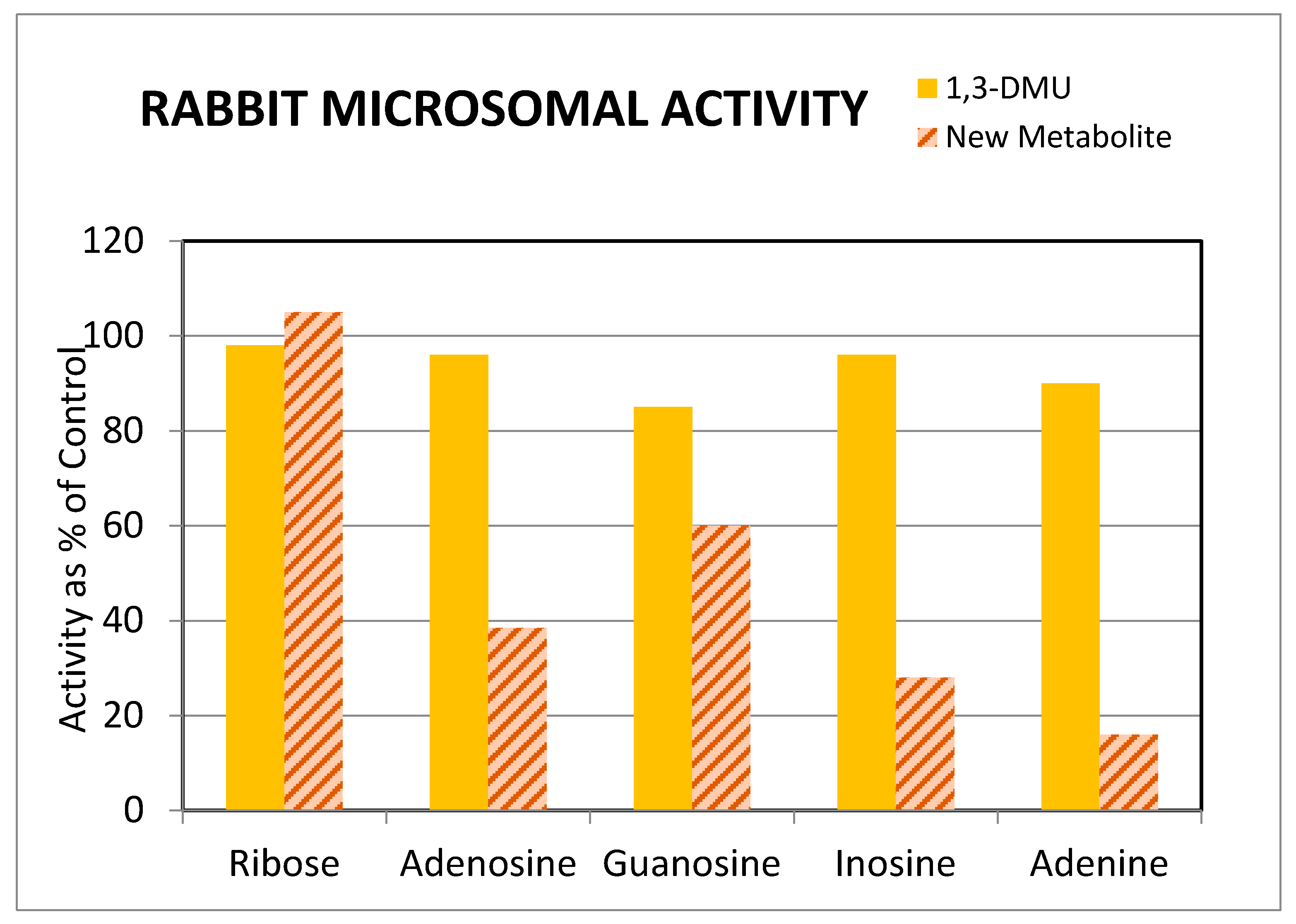
3.3. Potential for Pharmacological Activity of Theonosine
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bend, J.R.; Serabjit-Singh, C.J.; Philpot, R.M. The pulmonary uptake, accumulation, and metabolism of xenobiotics. Ann. Rev. Pharmacol. Toxicol. 1985, 25, 97–125. [Google Scholar] [CrossRef] [PubMed]
- Krishna, D.R.; Klotz, U. Extrahepatic metabolism of drugs in humans. Clin. Pharmacokinet. 1994, 24, 144–160. [Google Scholar] [CrossRef] [PubMed]
- Hukkanen, J.; Pelkonen, O.; Hakkola, J.; Raunio, H. Expression and regulation of xenobiotic-metabolizing cytochrome P450 (CYP) enzymes in human lung. Crit. Rev. Toxicol. 2002, 32, 391–411. [Google Scholar] [CrossRef] [PubMed]
- Nhamburo, P.T.; Gonzalez, F.J.; McBride, G.W.; Gelboin, H.V.; Kimura, S. Identification of a new P450 expressed in human lung: Complete cDNA sequence, cDNA-directed expression, and chromosome mapping. Biochemistry 1989, 28, 8060–8066. [Google Scholar] [CrossRef] [PubMed]
- Upton, R.N.; Doolette, D.J. Kinetic aspects of drug disposition in the lungs. Clin. Exp. Pharmacol. Physiol. 1999, 26, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Vane, J.R. The release and fate of vaso-active hormones in the circulation. Br. J. Pharmacol. 1969, 35, 209–242. [Google Scholar] [CrossRef] [PubMed]
- Bakhle, Y.S. Pharmacokinetic and metabolic properties of lung. Br. J. Anaesth. 1990, 65, 79–93. [Google Scholar] [CrossRef] [PubMed]
- Hook, G.E.R.; Bend, J.R. Pulmonary metabolism of xenobiotics. Life Sci. 1976, 18, 279–290. [Google Scholar] [CrossRef]
- Wolf, C.R.; Szutowski, M.M.; Ball, L.M.; Philpot, R.M. The rabbit pulmonary monooxygenase system: Characteristics and activities of two forms of pulmonary cytochrome P-450. Chem. Biol. Interact. 1978, 21, 29–43. [Google Scholar] [CrossRef]
- Petruzzelli, S.; Camus, A.-M.; Carrozzi, L.; Ghelarducci, L.; Rindi, M.; Menconi, G.; Angeletti, C.A.; Ahotupa, M.; Hietanen, E.; Aitio, A.; et al. Long-lasting effects of tobacco smoking on pulmonary drug-metabolizing enzymes: A case-control study on lung cancer patients. Cancer Res. 1988, 48, 4695–4700. [Google Scholar] [PubMed]
- Hermann, G.; Aynesworth, M.B. Successful treatment of persistent extreme dyspnea “status asthmaticus”: Use of theophylline ethylene diamine (aminophylline USP) intravenously. J. Lab. Clin. Med. 1937, 23, 135–148. [Google Scholar]
- Brodie, B.B.; Axelrod, J.; Reichenthal, A. Metabolism of theophylline (1,3-dimethylxantine) in man. J. Biol. Chem. 1952, 194, 215–222. [Google Scholar] [PubMed]
- Tang-Liu, D.D.-S.; Williams, R.L.; Riegelman, S. Nonlinear theophylline elimination. Clin. Pharmacol. Ther. 1982, 31, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Knoppert, D.C.; Spino, M.; Beck, R.; Thiessen, J.J.; MacLeod, S.M. Cystic fibrosis: Enhanced theophylline metabolism may be linked to the disease. Clin. Pharmacol. Ther. 1988, 44, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, T.; Prough, R.A.; Burke, M.D.; Estabrook, R.W. The preparation of microsomal fractions of rodent respiratory tract and their characterization. Cancer Res. 1974, 34, 2196–2203. [Google Scholar] [PubMed]
- Lowry, O.H.; Rosebrough, N.J.; Farr, A.L.; Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 1951, 193, 265–275. [Google Scholar] [PubMed]
- St. Pierre, M.V.; Tesoro, A.; Spino, M.; MacLeod, S.M. An HPLC method for the determination of theophylline and its metabolites in serum and urine. J. Liquid Chromatogr. 1984, 7, 1593–1608. [Google Scholar] [CrossRef]
- Dyroff, M.C.; Neal, R.A. Studies of the mechanism of metabolism of thioacetamide S-oxide by rat liver microsomes. Mol. Pharmacol. 1983, 23, 219–227. [Google Scholar] [PubMed]
- Miyaki, M.; Saito, A.; Shimizu, B. N→N Alkyl and glycosyl migrations of purines and pyrimidines. IV. trans-Glycosylation from pyrimidines to purines. (A novel synthetic method of purine nucleosides and nucleotides). Chem. Pharm. Bull. 1970, 18, 2459–2468. [Google Scholar] [CrossRef]
- Kröll, F.; Karlsson, J.-A.; Nilsson, E.; Ryrfeldt, Å.; Persson, C.G.A. Rapid clearance of xanthines from airway and pulmonary tissues. Am. Rev. Respir. Dis. 1990, 141, 1167–1171. [Google Scholar] [CrossRef] [PubMed]
- IJzerman, A.P.; Van der Wenden, R.A.A.; Mathôt, M.; Danhof, M.; Borea, P.A.; Varani, K. Partial agonism of theophylline-7-riboside on adenosine receptors. Naunyn Schmiedebergs Arch. Pharmacol. 1994, 350, 638–645. [Google Scholar] [CrossRef] [PubMed]
- Elion, G.B.; Hitchings, G.H. Metabolic basis for the actions of analogs of purines and pyrimidines. Adv. Chemother. 1965, 2, 91–117. [Google Scholar] [PubMed]
- Krenitsky, T.A.; Elion, G.B.; Strelitz, R.A.; Hitchings, G.H. Ribonucleosides of allopurinol and oxallopurinol. Isolation from human urine, enzymatic synthesis, and characterization. J. Biol. Chem. 1967, 242, 2675–2682. [Google Scholar] [PubMed]
- Nelson, D.J.; Buggé, C.J.L.; Krasny, H.C.; Elion, G.B. Formation of nucleotides of [6-14C] allopurinol and [6-14C] oxipurinol in rat tissues and effects on uridine nucleotide pools. Biochem. Pharmacol. 1973, 22, 2003–2022. [Google Scholar] [CrossRef]
- Reiter, S.; Simmonds, H.A.; Webster, D.R.; Watson, A.R. On the metabolism of allopurinol: Formation of allopurinol-1-riboside in purine nucleoside phosphorylase deficiency. Biochem. Pharmacol. 1983, 32, 2167–2174. [Google Scholar] [CrossRef]
- Van Galen, P.J.M.; Uzerman, A.P. Xanthine-7-ribosides as adenosine A1 receptor antagonists: Further evidence for adenosine’s anti mode of binding. Nucleosides Nucleotides 1990, 9, 275–291. [Google Scholar] [CrossRef]
- Van der Wenden, E.M.; Hartog-Witte, H.R.; Roelen, H.C.P.F.; von Frijtag Drabbe Kunzel, J.K.; Pirovano, I.M.; Mathôt, R.A.A.; Danhof, M.; Van Aerschot, A.; Lidaks, M.J.; Ijzerman, A.P.; et al. 8-Substituted adenosine and theophylline-7-riboside analogues as potential partial agonists for the adenosine A1 receptor. Eur. J. Pharmacol. 1995, 290, 189–199. [Google Scholar] [CrossRef]


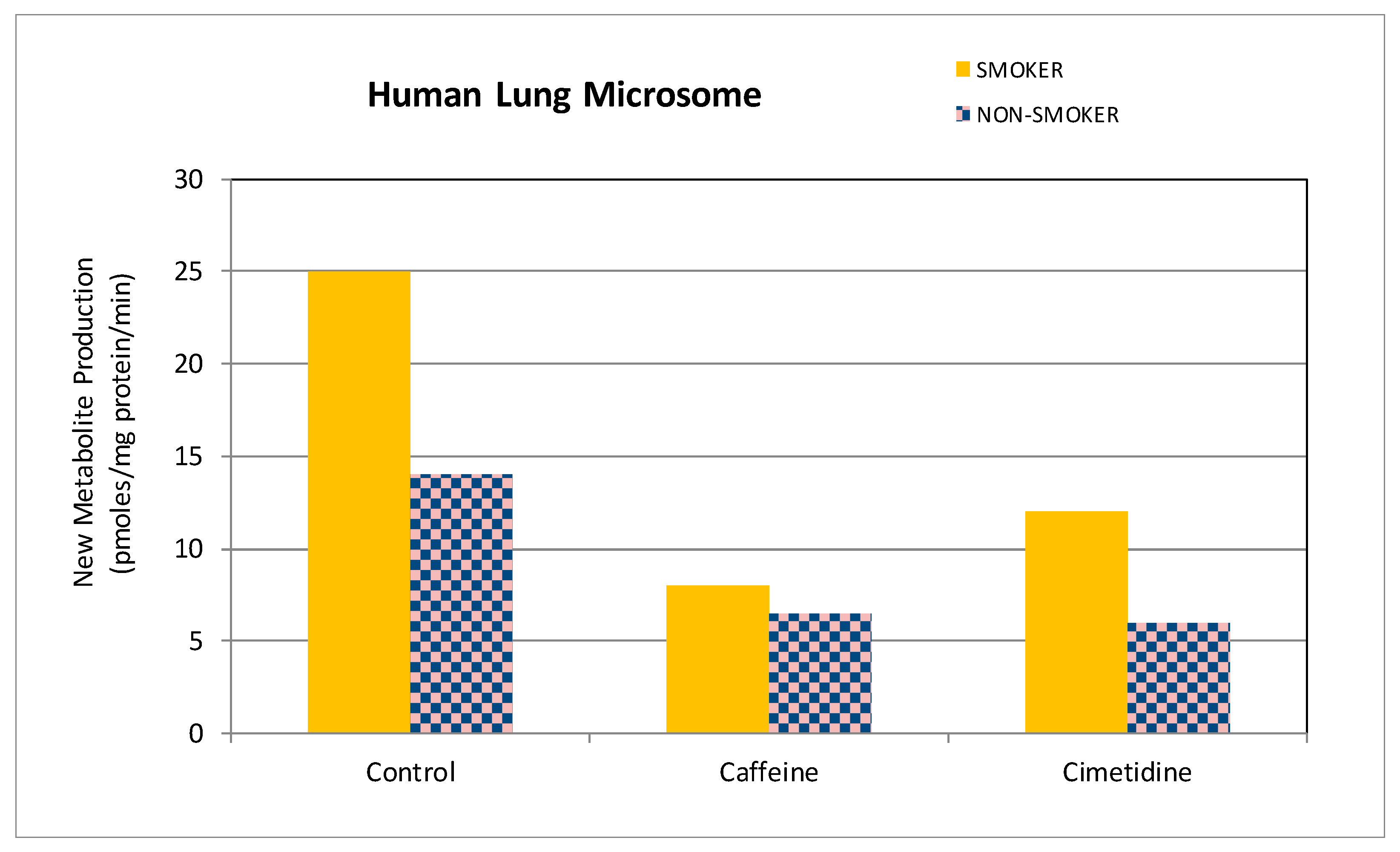
| Species | Theonosine Production (pmol/mg protein/min) |
|---|---|
| Dog (n = 5) | 31 ± 15 |
| Sheep (n = 1) | 32 |
| Rabbit (n = 8) | 11 ± 5 |
| Rat (n = 7) | 41 ± 10 |
| Pig (n = 3) | 40 ± 19 |
| Human (n = 16) | 18 ± 11 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sitar, D.S.; Bowen, J.M.; He, J.; Tesoro, A.; Spino, M. Theophylline-7β-d-Ribofuranoside (Theonosine), a New Theophylline Metabolite Generated in Human and Animal Lung Tissue. Pharmaceutics 2017, 9, 28. https://doi.org/10.3390/pharmaceutics9030028
Sitar DS, Bowen JM, He J, Tesoro A, Spino M. Theophylline-7β-d-Ribofuranoside (Theonosine), a New Theophylline Metabolite Generated in Human and Animal Lung Tissue. Pharmaceutics. 2017; 9(3):28. https://doi.org/10.3390/pharmaceutics9030028
Chicago/Turabian StyleSitar, Daniel S., James M. Bowen, Juan He, Angelo Tesoro, and Michael Spino. 2017. "Theophylline-7β-d-Ribofuranoside (Theonosine), a New Theophylline Metabolite Generated in Human and Animal Lung Tissue" Pharmaceutics 9, no. 3: 28. https://doi.org/10.3390/pharmaceutics9030028
APA StyleSitar, D. S., Bowen, J. M., He, J., Tesoro, A., & Spino, M. (2017). Theophylline-7β-d-Ribofuranoside (Theonosine), a New Theophylline Metabolite Generated in Human and Animal Lung Tissue. Pharmaceutics, 9(3), 28. https://doi.org/10.3390/pharmaceutics9030028