Hydrotalcite Intercalated siRNA: Computational Characterization of the Interlayer Environment
Abstract
1. Introduction
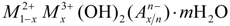 , where M2+ and M3+ are divalent and trivalent metallic cations, respectively, and A is an anion of valence n. The most studied class of LDHs is Mg6Al2(OH)16CO3·4H2O, a popular pharmaceutical antacid talcid for ulcers. LDH structure is closely related to brucite: Mg(OH)2. In a brucite layer, each Mg2+ ion is octahedrally surrounded by six OH− ions and the different octahedrons share edges to form an - two-dimensional layer. Partial replacements of Mg2+ ions by Al3+ give the “brucite-like” layers a permanent positive charge, which is balanced by negatively charged anions located in the interlayer region. This gallery also contains water molecules, hydrogen bonded to layer hydroxide and/or to the interlayer anions. Through electrostatic interactions and hydrogen bonds, the layers may be stabilized in a crystalline form. The charge density in the hydroxide layer, represented by x in the general formula, is normally in the range 0.2–0.33 [6].
, where M2+ and M3+ are divalent and trivalent metallic cations, respectively, and A is an anion of valence n. The most studied class of LDHs is Mg6Al2(OH)16CO3·4H2O, a popular pharmaceutical antacid talcid for ulcers. LDH structure is closely related to brucite: Mg(OH)2. In a brucite layer, each Mg2+ ion is octahedrally surrounded by six OH− ions and the different octahedrons share edges to form an - two-dimensional layer. Partial replacements of Mg2+ ions by Al3+ give the “brucite-like” layers a permanent positive charge, which is balanced by negatively charged anions located in the interlayer region. This gallery also contains water molecules, hydrogen bonded to layer hydroxide and/or to the interlayer anions. Through electrostatic interactions and hydrogen bonds, the layers may be stabilized in a crystalline form. The charge density in the hydroxide layer, represented by x in the general formula, is normally in the range 0.2–0.33 [6]. 2. Simulation Methods
2.1. Model Construction
 (this symmetry is the most commonly found in nature). The rhombohedral lattice parameters are a = 3.0460 Å, c = 22.772 Å, α = 90°, β = 90° and γ = 120°. The initial interlayer carbonate anions CO32− and water molecules were then removed, and a supercell of 24a× 10a× 2c was set up with lattice parameters 24a = 73.1042 Å, 10b = 30.460 Å, and 2c = 45.5444 Å. The three layer repeat was built with a central layer spacing of 30.363 Å and the other two layer spacing of 7.591 Å. Each hydroxide layer contains 77 Al3+ and 163 Mg2+ ions, with the latter being arranged in such a way that they are not located in adjacent octahedrals. Thus, the final Mg:Al ratio in the LDH layer is roughly 2:1. The simulation model thus consists of three host layers and three guest layers.
(this symmetry is the most commonly found in nature). The rhombohedral lattice parameters are a = 3.0460 Å, c = 22.772 Å, α = 90°, β = 90° and γ = 120°. The initial interlayer carbonate anions CO32− and water molecules were then removed, and a supercell of 24a× 10a× 2c was set up with lattice parameters 24a = 73.1042 Å, 10b = 30.460 Å, and 2c = 45.5444 Å. The three layer repeat was built with a central layer spacing of 30.363 Å and the other two layer spacing of 7.591 Å. Each hydroxide layer contains 77 Al3+ and 163 Mg2+ ions, with the latter being arranged in such a way that they are not located in adjacent octahedrals. Thus, the final Mg:Al ratio in the LDH layer is roughly 2:1. The simulation model thus consists of three host layers and three guest layers. 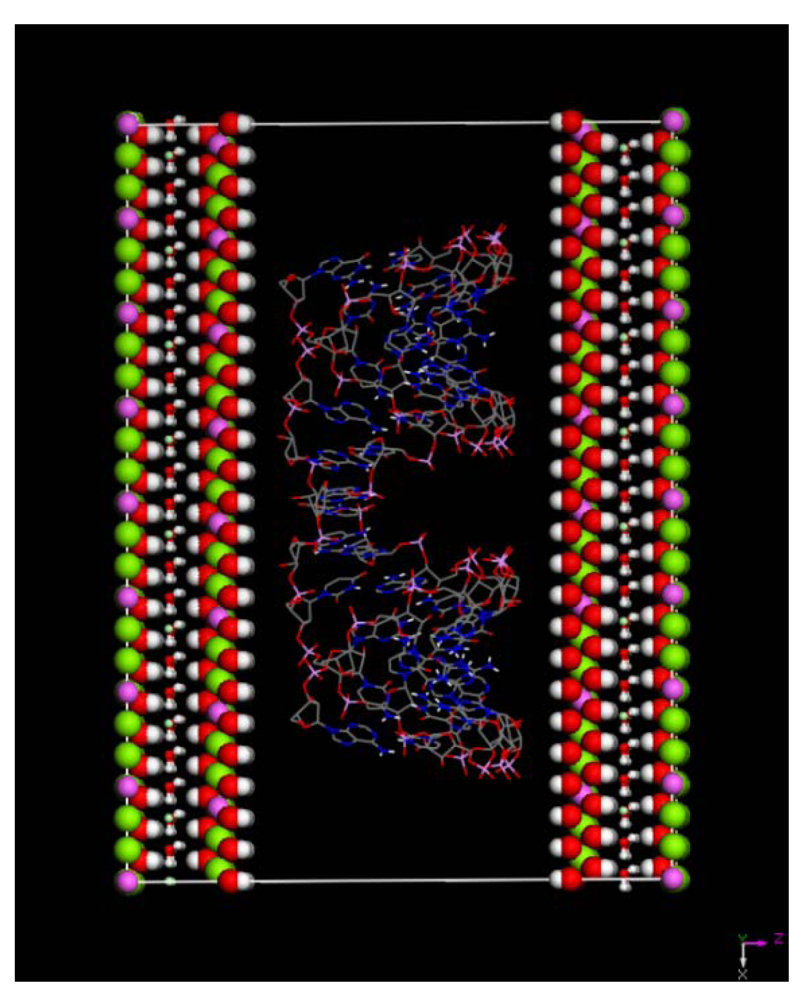
2.2. Minimization
2.3. Molecular Dynamics Techniques
3. Results
3.1. Simulation Results
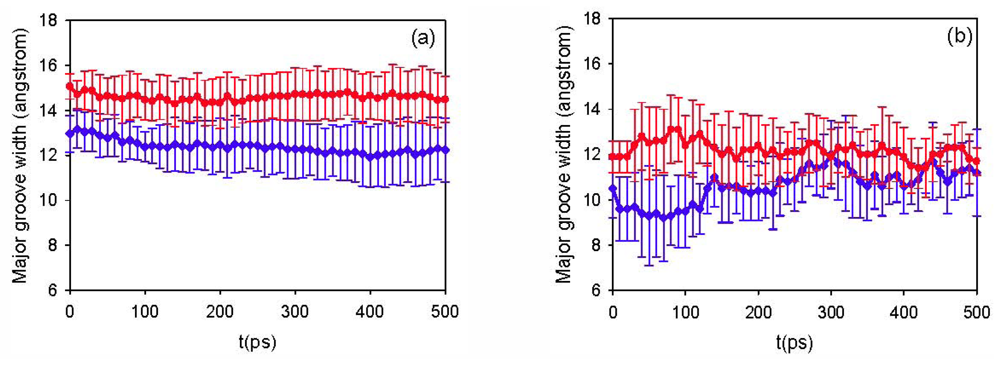

3.2. Diffusion Coefficients
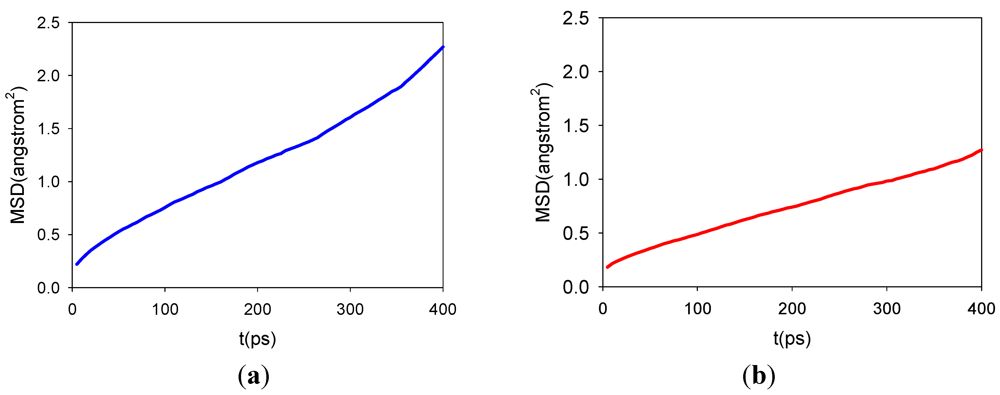
3.3. Comparisons with Experiments
 m rhombohedral symmetry generally observed in LDH systems. The peak occurring at a low 2θ angle (at about 2°) was attributed to the reflections from the (003) family of crystallographic planes for the central layer. It corresponds to the interlayer repeat distance d including a contribution from the metal hydroxide sheet (approximately 0.48 nm) and the interlayer space that contains the guest siRNA anions surrounded by water molecules. The intercalation of siRNA into the LDH was clearly evidenced by the net increase of the interlayer distance, from roughly 0.7 nm (for the two edge layers) to roughly 3.0 nm, for the siRNA layer. The presence of several harmonics of relatively high intensity (especially from the partially relaxed simulations) is evidence of generally well-defined MgAl hybrid materials. Figure 6a also reveals the apparent peaks from two Cl− containing edge layers, which displays a sharp (003) peak at larger angles (at about 10°). In all the cases, the peaks for the central layer at the low angle are the strongest and dominate in the calculated XRD pattern. In the fully relaxed simulations, the corresponding peaks are much lower due to the deformations of the two edge layers (there is a lesser degree of ordering). Since the COMPASS force field is a general ab initio force field (i.e., not specifically designed for LDH system), the precise degree of deformation of the layers that is observed may vary somewhat from the true situation, but nevertheless the observation is indicative and in accord with findings for other intercalated LDH systems (e.g., [41]). Furthermore, it must be borne in mind that the current periodic simulations correspond to an effective loading of siRNA which may induce greater distortions than the experimental case.
m rhombohedral symmetry generally observed in LDH systems. The peak occurring at a low 2θ angle (at about 2°) was attributed to the reflections from the (003) family of crystallographic planes for the central layer. It corresponds to the interlayer repeat distance d including a contribution from the metal hydroxide sheet (approximately 0.48 nm) and the interlayer space that contains the guest siRNA anions surrounded by water molecules. The intercalation of siRNA into the LDH was clearly evidenced by the net increase of the interlayer distance, from roughly 0.7 nm (for the two edge layers) to roughly 3.0 nm, for the siRNA layer. The presence of several harmonics of relatively high intensity (especially from the partially relaxed simulations) is evidence of generally well-defined MgAl hybrid materials. Figure 6a also reveals the apparent peaks from two Cl− containing edge layers, which displays a sharp (003) peak at larger angles (at about 10°). In all the cases, the peaks for the central layer at the low angle are the strongest and dominate in the calculated XRD pattern. In the fully relaxed simulations, the corresponding peaks are much lower due to the deformations of the two edge layers (there is a lesser degree of ordering). Since the COMPASS force field is a general ab initio force field (i.e., not specifically designed for LDH system), the precise degree of deformation of the layers that is observed may vary somewhat from the true situation, but nevertheless the observation is indicative and in accord with findings for other intercalated LDH systems (e.g., [41]). Furthermore, it must be borne in mind that the current periodic simulations correspond to an effective loading of siRNA which may induce greater distortions than the experimental case.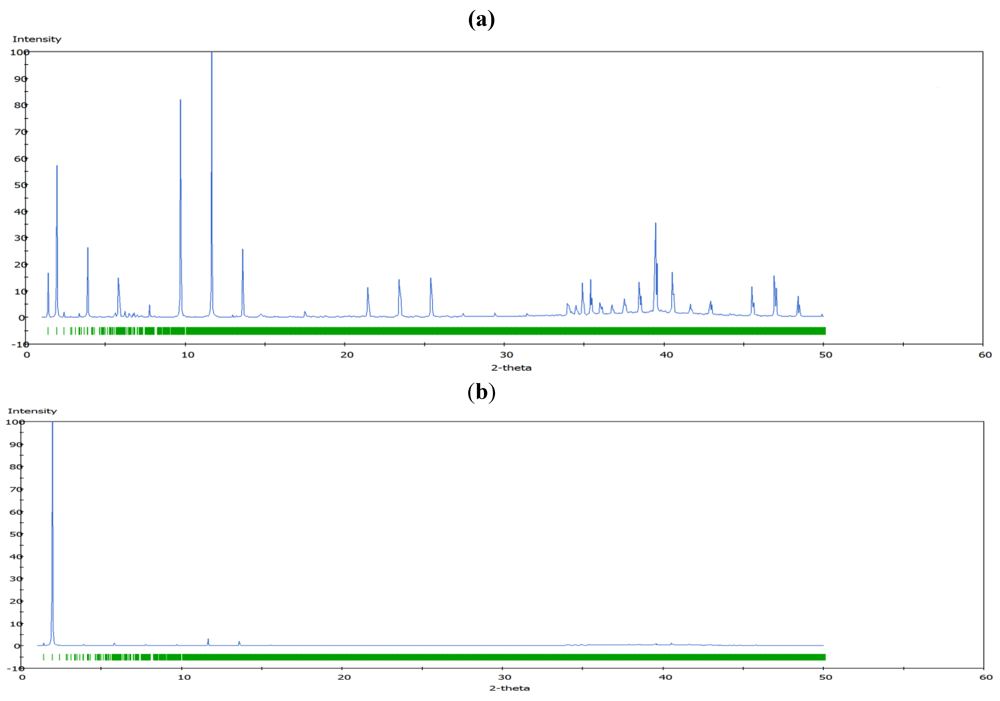
4. Conclusions
Acknowledgements
References
- Cavani, F.; Trifiro, F.; Vaccari, A. Hydrotalcite-type anionic clays: Preparation, properties and applications. Catal. Today 1991, 11, 173–301. [Google Scholar]
- Tichit, D.; Coq, B. Catalysis by hydrotalcites and related materials. Cattech 2003, 7, 206–217. [Google Scholar] [CrossRef]
- van der Ven, L.; van Gemert, M.L.M.; Batenburg, L.F.; Keern, J.J.; Gielgens, L.H.; Koster, T.P.M.; Fischer, H.R. On the action of hydrotalcite-like clay materials as stabilizers in polyvinylchloride. Appl. Clay Sci. 2000, 17, 25–34. [Google Scholar] [CrossRef]
- Miyata, S. Anion-exchange properties of hydrotalcite-like compounds. Clays Clay Miner. 1983, 31, 305–311. [Google Scholar]
- Choy, J.H.; Kwak, S.-Y.; Park, J.-S.; Jeong, Y.-J. Cellular uptake behavior of [γ-32P] labeled ATP-LDH nanohybrids. J. Mater. Chem. 2001, 11, 1671–1674. [Google Scholar] [CrossRef]
- Bergaya, F.; Theng, B.K.G.; Lagaly, G. Handbook of Clay Science. In Developments in Clay Science; Elsevier Ltd.: Amsterdam, The Netherlands, 2006; p. 1021. [Google Scholar]
- Krishnamoorti, R.; Vaia, R.A.; Giannelis, E.P. Structure and dynamics of polymer-layered silicate nanocomposites. Chem. Mater. 1996, 8, 1728–1734. [Google Scholar] [CrossRef]
- Kovář, P.; Melánová, K.; Zima, V.; Beneš, L.; Čapková, P. Layered double hydroxide intercalated with p-methylbenzoate and p-bromobenzoate: Molecular simulations and XRD analysis. J. Colloid Interface Sci. 2008, 319, 19–24. [Google Scholar] [CrossRef]
- Xu, Z.P.; Braterman, P.S. Competitive Intercalation of Sulfonates into Layered Double Hydroxides (LDHs): The key role of hydrophobic interactions. J. Phys. Chem. C 2007, 111, 4021–4026. [Google Scholar] [CrossRef]
- Costa, F.R.; Leuteritz, A.; Wagenknecht, U.; Jehnichen, D.; Häußler, L.; Heinrich, G. Intercalation of Mg–Al layered double hydroxide by anionic surfactants: Preparation and characterization. Appl. Clay Sci. 2008, 38, 153–164. [Google Scholar] [CrossRef]
- Nakayama, H.; Wada, N.; Tsuhako, M. Intercalation of amino acids and peptides into Mg-Al-layered double hydroxide by reconstruction method. Int. J. Pharm. 2004, 269, 469–478. [Google Scholar] [CrossRef]
- Choy, J.H.; Kwak, S.-Y.; Park, J.-S.; Jeong, Y.-J.; Portier, J. Intercalative nanohybrids of nucleotide monophosphates and DNA in layered metal hydroxide. J. Am. Chem. Soc. 1999, 121, 1399–1400. [Google Scholar]
- Xu, Z.P.; Niebert, M.; Porazik, K.; Walker, T.L.; Cooper, H.M.; Middelberg, A.P.; Gray, P.P.; Bartlett, P.F.; Lu, G.Q. Subcellular compartment targeting of layered double hydroxide nanoparticles. J. Control. Release 2008, 130, 86–94. [Google Scholar] [CrossRef]
- Kwak, S.Y.; Jeong, Y.-J.; Park, J.-S.; Choy, J.H. Bio-LDH nanohybrid for gene therapy. Solid State Ion. 2002, 151, 229–234. [Google Scholar] [CrossRef]
- bin Hussein, M.Z.; Yahaya, A.H.; Zainal, Z.; Kian, L.H. Nanocomposite-based controlled release formulation of an herbicide, 2,4-dichlorophenoxyacetate incapsulated in zinc-aluminium-layered double hydroxide. Sci. Technol. Adv. Mater. 2005, 6, 956–962. [Google Scholar] [CrossRef]
- Choy, J.H.; Choi, S.-J.; Oh, J.-M.; Park, T. Clay minerals and layered double hydroxides for novel biological applications. Appl. Clay Sci. 2007, 36, 122–132. [Google Scholar] [CrossRef]
- Ladewig, K.; Zhi, P.X.; Gao, Q.L. Layered double hydroxide nanoparticles in gene and drug delivery. Expert Opin. Drug Deliv. 2009, 6, 907–922. [Google Scholar] [CrossRef]
- Choy, J.H.; Oh, J.-M.; Park, M.; Sohn, K.-M.; Kim, J.-W. Inorganic-biomolecular hybrid nanomaterials as a genetic molecular code system. Adv. Mater. 2004, 16, 1181–1184. [Google Scholar] [CrossRef]
- Choy, J.H.; Kwak, S.-Y.; Jeong, Y.-J.; Park, J.-S. Inorganic layered double hydroxides as nonviral vectors. Angew. Chem.-Int. Ed. 2000, 39, 4042–4045. [Google Scholar]
- Wong, Y.; Markham, K.; Xu, Z.P.; Chen, M.; Max, L.G.Q.; Bartlett, P.F.; Cooper, H.M. Efficient delivery of siRNA to cortical neurons using layered double hydroxide nanoparticles. Biomaterials 2010, 31, 8770–8779. [Google Scholar]
- Voet, D.; Voet, J.G.; Pratt, C.W. Foundamentals of Biochemistry: Life at the Molecular Level, 3rd ed; Wiley: Hoboken, NJ, USA, 2008. [Google Scholar]
- Theimer, C.A.; Finger, L.D.; Feigon, J. YNMG tetraloop formation by a dyskeratosiscongenita mutation in human telomerase RNA. RNA 2003, 9, 1446–1455. [Google Scholar] [CrossRef]
- Popenda, L.; Adamiak, R.W.; Gdaniec, Z. Bulged adenosine influence on the RNA duplex conformation in solution. Biochemistry 2008, 47, 5059–5067. [Google Scholar]
- Schmitz, M.; Tinoco, I. Solution structure and metal-ion binding of the P4 element from bacterial RNase P RNA. RNA 2000, 6, 1212–1225. [Google Scholar] [CrossRef]
- Nomura, Y.; Kajikawa, M.; Baba, S.; Nakazato, S.; Imai, T.; Sakamoto, T.; Okada, N.; Kawai, G. Solution structure and functional importance of a conserved RNA hairpin of eel LINE UnaL2. Nucleic Acids Res. 2006, 34, 5184–5193. [Google Scholar] [Green Version]
- Arnott, S.; Fuller, W.; Hodgson, A.; Prutton, I. Molecular conformations and structure transitions of RNA complementary helices and their possible biological significance. Nature 1968, 220, 561–564. [Google Scholar]
- Arnott, S.; Hukins, D.W.L.; Dover, S.D.; Fuller, W.; Hodgson, A.R. Structures of synthetic polynucleotides in the A-RNA and A'-RNA conformations: X-ray diffraction analyses of the molecular conformations of polyadenylic acid-polyuridylic acid and polyinosinic acid-polycytidylic acid. J. Mol. Biol. 1973, 81, 107–122. [Google Scholar] [CrossRef]
- Neidle, S. Principles of Nucleic Acid Structure; Elsevier: Amsterdam, The Netherlands, 2008. [Google Scholar]
- Blackburn, G.M.E. Nucleic Acids in Chemistry and Biology, 3rd ed; RSC: Cambridge, UK, 2006; pp. 57–58. [Google Scholar]
- Neidle, S. Oxford Handbook of Nucleic Acid Structure; Oxford University Press: Oxford, UK, 1999. [Google Scholar]
- Ferris, J.P. Montmorillonite-catalysed formation of RNA oligomers: The possible role of catalysis in the origins of life. Philos. Trans. R. Soc. B-Biol. Sci. 2006, 361, 1777–1786. [Google Scholar] [CrossRef]
- Arrhenius, G.; Sales, B.; Mojzsis, S.; Lee, T. Entropy and charge in molecular evolution-the case of phosphate. J. Theor. Biol. 1997, 187, 503–522. [Google Scholar] [CrossRef]
- Wattis, J.A.D.; Coveney, P.V. Chiral polymerization and the RNA world. Int. J. Astrobiol. 2005, 4, 66–73. [Google Scholar]
- Newman, S.P.; Cristina, T.D.; Coveney, P.V. Molecular dynamics simulation of cationic and anionic clays containing amino acids. Langmuir 2002, 18, 2933–2939. [Google Scholar]
- Greenwell, H.C.; Jones, W.; Newman, S.P.; Coveney, P.V. Computer simulation of interlayer arrangement in cinnamate intercalated layered double hydroxides. J. Mol. Struct. 2003, 647, 75–83. [Google Scholar] [CrossRef]
- Mohanambe, L.; Vasudevan, S. Anionic clays containing anti-inflammatory drug molecules: Comparison of molecular dynamics simulation and measurements. J. Phys. Chem. B 2005, 109, 15651–15658. [Google Scholar] [CrossRef]
- Kim, N.; Kim, Y.; Tsotsis, T.T.; Sahimi, M. Atomistic simulation of nanoporous layered double hydroxide materials and their properties. I. Structural modeling. J. Chem Phys. 2005, 122, 214713:1–214713:12. [Google Scholar]
- Wang, J.; Kalinichev, A.G.; Amonette, J.E.; Kirkpatrick, R.J. Interlayer structure and dynamics of Cl-bearing hydrotalcite: Far infrared spectroscopy and molecular dynamics modeling. Am. Mineral. 2003, 88, 398–409. [Google Scholar]
- Newman, S.P.; Williams, S.J.; Coveney, P.V.; Jones, W. Interlayer arrangement of hydrated MgAl LDHs containing guest terephthalate anions: Comparison of simulation and measurement. J. Phys. Chem. B 1998, 102, 6710–6719. [Google Scholar]
- Kumar, P.P.; Kalinichev, A.G.; Kirkpatrick, R.J. Molecular dynamics simulation of the energetics and structure of layered double hydroxides intercalated with carboxylic acids. J. Phys. Chem. C 2007, 111, 13517–13523. [Google Scholar]
- Thyveetil, M.-A.; Coveney, P.V.; Greenwell, H.C.; Suter, J.L. Computer simulation study of the structural stability and materials properties of DNA-intercalated layered double hydroxides. J. Am. Chem. Soc. 2008, 130, 4742–4756. [Google Scholar]
- Mayo, S.L.; Olafson, B.D.; Goddard, W.A. DREIDING: A generic force field for molecular simulations. J. Phys. Chem. 1990, 94, 8897–8909. [Google Scholar]
- Cygan, R.T.; Liang, J.-J.; Kalinichev, A.G. Molecular models of hydroxide, oxyhydroxide, and clay phases and the development of a general force field. J. Phys. Chem. B 2004, 108, 1255–1266. [Google Scholar]
- Kitson, D.H.; Hagler, A.T. Theoretical studies of the structure and molecular dynamics of a peptide crystal. Biochemistry 1988, 27, 5246–5257. [Google Scholar] [CrossRef]
- Zhang, H.; Xu, Z.P.; Lu, G.Q.; Smith, S.C. Intercalation of organic molecules into layered double hydroxide: Comparison of simulation with experiment. J. Phys. Chem. C 2008, 113, 559–566. [Google Scholar]
- Ladewig, K.; Niebert, M.; Xu, Z.P.; Gray, P.P.; Lu, G.Q.M. Efficient siRNA delivery to mammalian cells using layered double hydroxide nanoparticles. Biomaterials 2010, 31, 1821–1829. [Google Scholar]
- Ouyang, D.; Zhang, H.; Herten, D.-P.; Parekh, H.-S.; Smith, S.C. Flexibility of short-strand rna in aqueous solution as revealed by molecular dynamics simulation: Are a-RNA and a-RNA distinct conformational structures? Aust. J. Chem. 2009, 62, 1054–1061. [Google Scholar] [CrossRef]
- Sun, H. COMPASS: An ab initio force-field optimized for condensed-phase applications-Overview with details on alkane and benzene compounds. J. Phys. Chem. B 1998, 102, 7338–7364. [Google Scholar] [CrossRef]
- Bellotto, M.; Rebours, B.; Clause, O.; Lynch, J. A reexamination of hydrotalcite crystal chemistry. J. Phys. Chem. 1996, 100, 8527–8534. [Google Scholar]
- Putral, L.N.; Bywater, M.J.; Gu, W.; Saunders, N.A.; Gabrielli, B.G.; Leggatt, G.R.; McMillan, N.A.J. RNA interference against human papillomavirusoncogenes in cervical cancer cells results in increased sensitivity to cisplatin. Mol. Pharmacol. 2005, 68, 1311–1319. [Google Scholar] [CrossRef]
- Zhang, H.; Xu, Z.P.; Lu, G.Q.; Smith, S.C. Intercalation of sulfonate into layered double hydroxide: Comparison of simulation with experiment. J. Phys. Chem. C 2009, 113, 559–566. [Google Scholar]
- Desigaux, L.; Belkacem, M.B.; Richard, P.; Cellier, J.; Léone, P.; Cario, L.; Leroux, F.; Taviot-Guého, C.; Pitard, B. Self-assembly and characterization of layered double hydroxide/DNA hybrids. NanoLett. 2006, 6, 199–204. [Google Scholar] [CrossRef]
- Masarudin, M.J.; Yusoff1, K.; Rahim, R.A.; Hussein, M.Z. Successful transfer of plasmid DNA into in vitro cells transfected with an inorganic plasmid—Mg/Al-LDH nanobiocomposite material as a vector for gene expression. Nanotechnology 2009, 20, 045602. [Google Scholar]
- Oh, J.M.; Kwak, S.Y.; Choy, J.H. Intracrystalline structure of DNA molecules stabilized in the layered double hydroxide. J. Phys. Chem. Solids 2006, 67, 1028–1031. [Google Scholar] [CrossRef]
- Arnott, S.; Hukins, D.W.L.; Dover, S.D. Optimized parameters for RNA double-helices. Biochem. Biophys. Res. Commun. 1972, 48, 1392–1399. [Google Scholar] [CrossRef]
- Tanaka, Y.; Fujii, S.; Hiroaki, H.; Sakata, T.; Tanaka, T.; Uesugi, S.; Tomita, K.; Kyogoku, Y. A'-form RNA double helix in the single crystal structure of r(UGAGCUUCGGCUC). Nucleic Acids Res. 1999, 27, 949–955. [Google Scholar]
- Aicken, A.M.; Bell, I.S.; Coveney, P.V.; Jones, W. Simulation of layered double hydroxide intercalates. Adv. Mater. 1997, 9, 496–500. [Google Scholar] [CrossRef]
- Boek, E.S.; Coveney, P.V.; Williams, S.J.; Bains, A.S. A robust water potential parameterisation. Mol. Simul. 1996, 18, 145–154. [Google Scholar] [CrossRef]
- Mills, R. Self-diffusion in normal and heavy-water in range 1–45 degrees. J. Phys. Chem. 1973, 77, 685–688. [Google Scholar] [CrossRef]
- Zabner, J.; Fasbender, A.J.; Moninger, T.; Poellinger, K.A.; Welsh, M.J. Cellular and molecular barriers to gene transfer by a cationic lipid. J. Biol. Chem. 1995, 270, 18997–19007. [Google Scholar]
- Schaffer, D.V.; Fidelman, N.A.; Dan, N.; Lauffenburger, D.A. Vector unpacking as a potential barrier for receptor-mediated polyplex gene delivery. Biotechnol. Bioeng. 2000, 67, 598–606. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhang, H.; Ouyang, D.; Murthy, V.; Wong, Y.; Xu, Z.; Smith, S.C. Hydrotalcite Intercalated siRNA: Computational Characterization of the Interlayer Environment. Pharmaceutics 2012, 4, 296-313. https://doi.org/10.3390/pharmaceutics4020296
Zhang H, Ouyang D, Murthy V, Wong Y, Xu Z, Smith SC. Hydrotalcite Intercalated siRNA: Computational Characterization of the Interlayer Environment. Pharmaceutics. 2012; 4(2):296-313. https://doi.org/10.3390/pharmaceutics4020296
Chicago/Turabian StyleZhang, Hong, Defang Ouyang, Vinuthaa Murthy, Yunyi Wong, Zhiping Xu, and Sean C. Smith. 2012. "Hydrotalcite Intercalated siRNA: Computational Characterization of the Interlayer Environment" Pharmaceutics 4, no. 2: 296-313. https://doi.org/10.3390/pharmaceutics4020296
APA StyleZhang, H., Ouyang, D., Murthy, V., Wong, Y., Xu, Z., & Smith, S. C. (2012). Hydrotalcite Intercalated siRNA: Computational Characterization of the Interlayer Environment. Pharmaceutics, 4(2), 296-313. https://doi.org/10.3390/pharmaceutics4020296