A Method for Evaluating Robustness of Limited Sampling Strategies—Exemplified by Serum Iohexol Clearance for Determination of Measured Glomerular Filtration Rate
Abstract
1. Introduction
2. Materials and Methods
2.1. Population Pharmacokinetics Model and Limited Sampling Strategy of Iohexol
2.2. Semi-Parametric Simulation from Support Points
2.3. Deviation from Optimal Sample Times
2.4. Optimal Sample Windows
3. Results
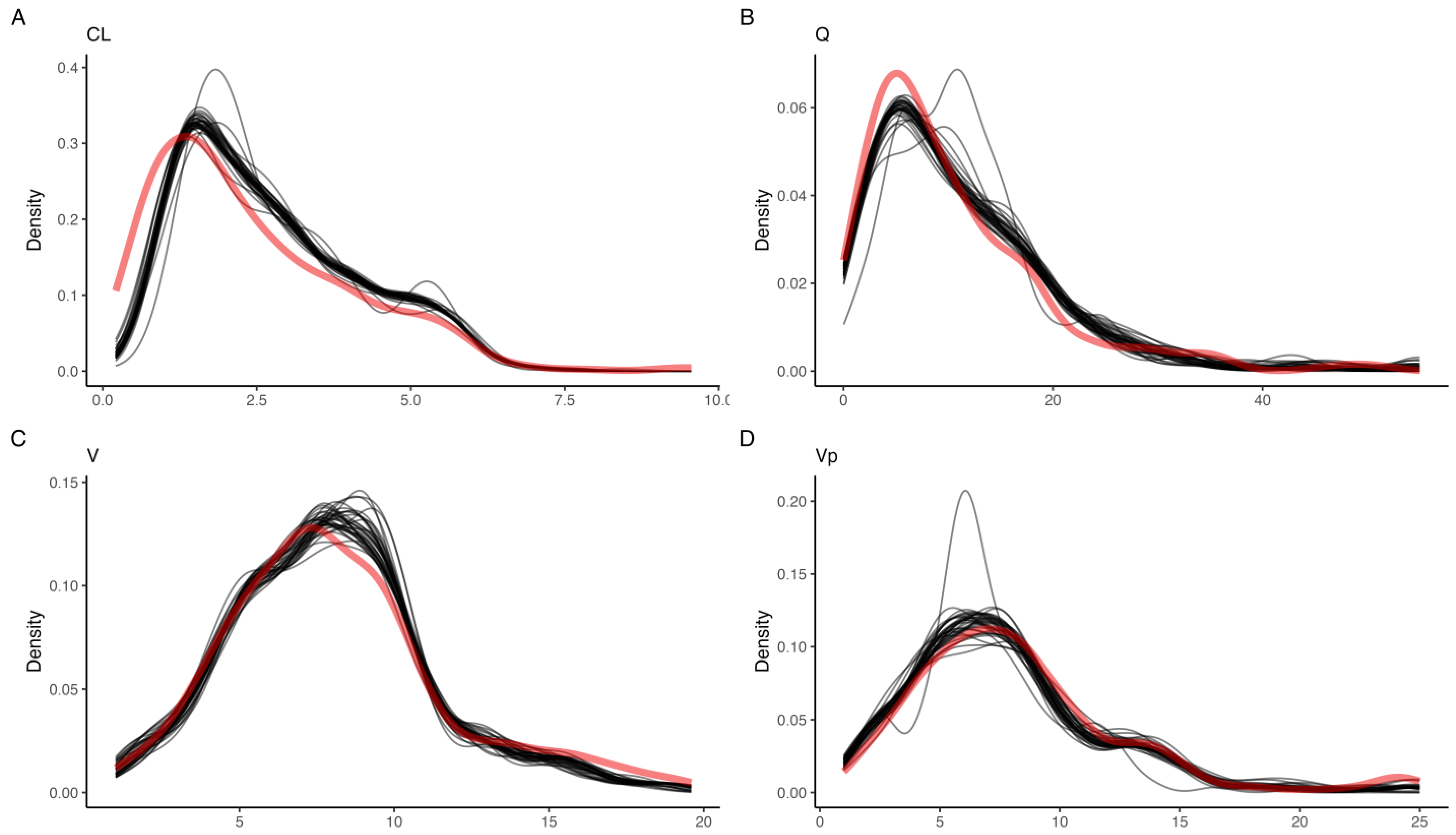
3.1. Simulated Profiles
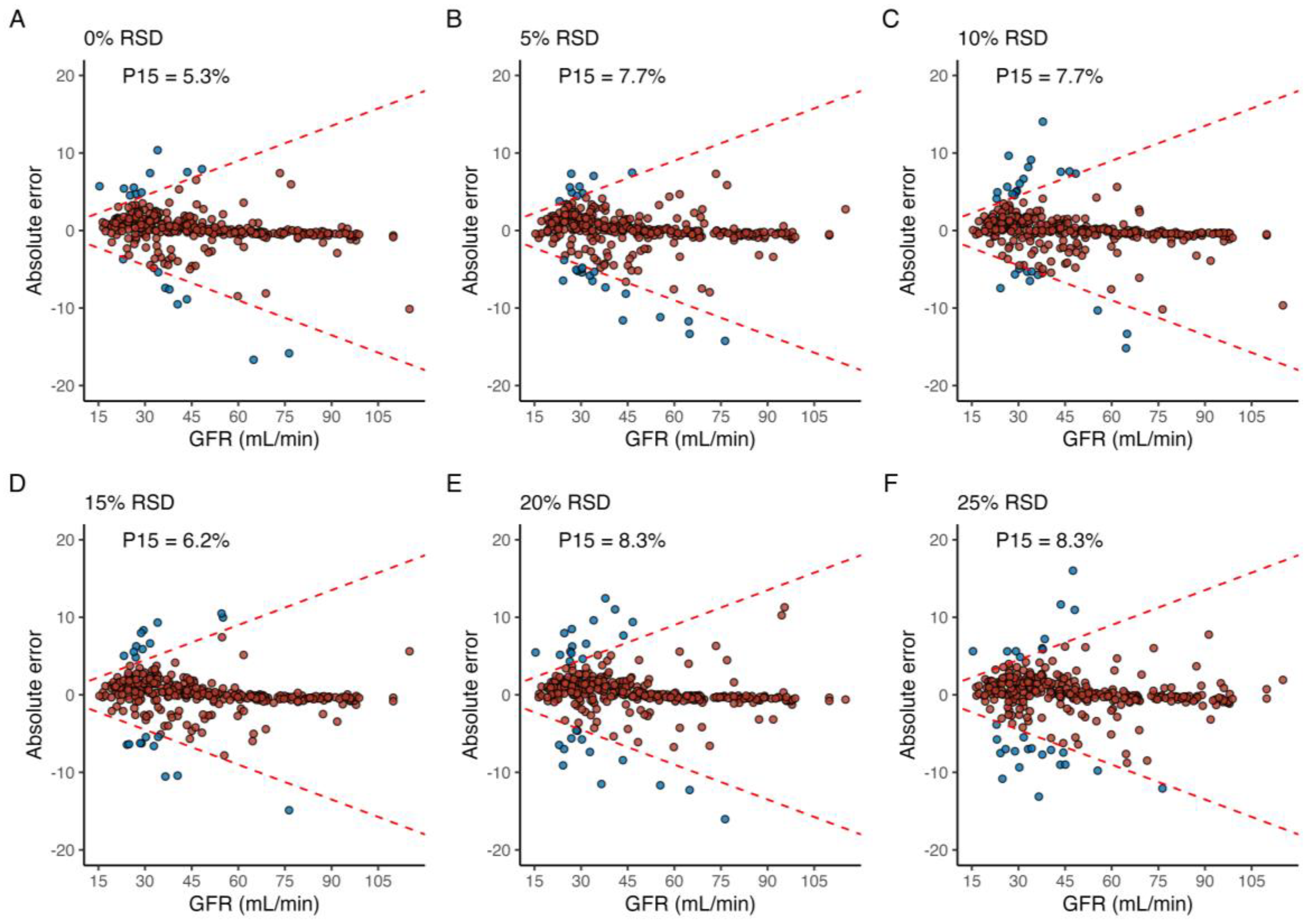
3.2. LSS Performance on Simulated Profiles
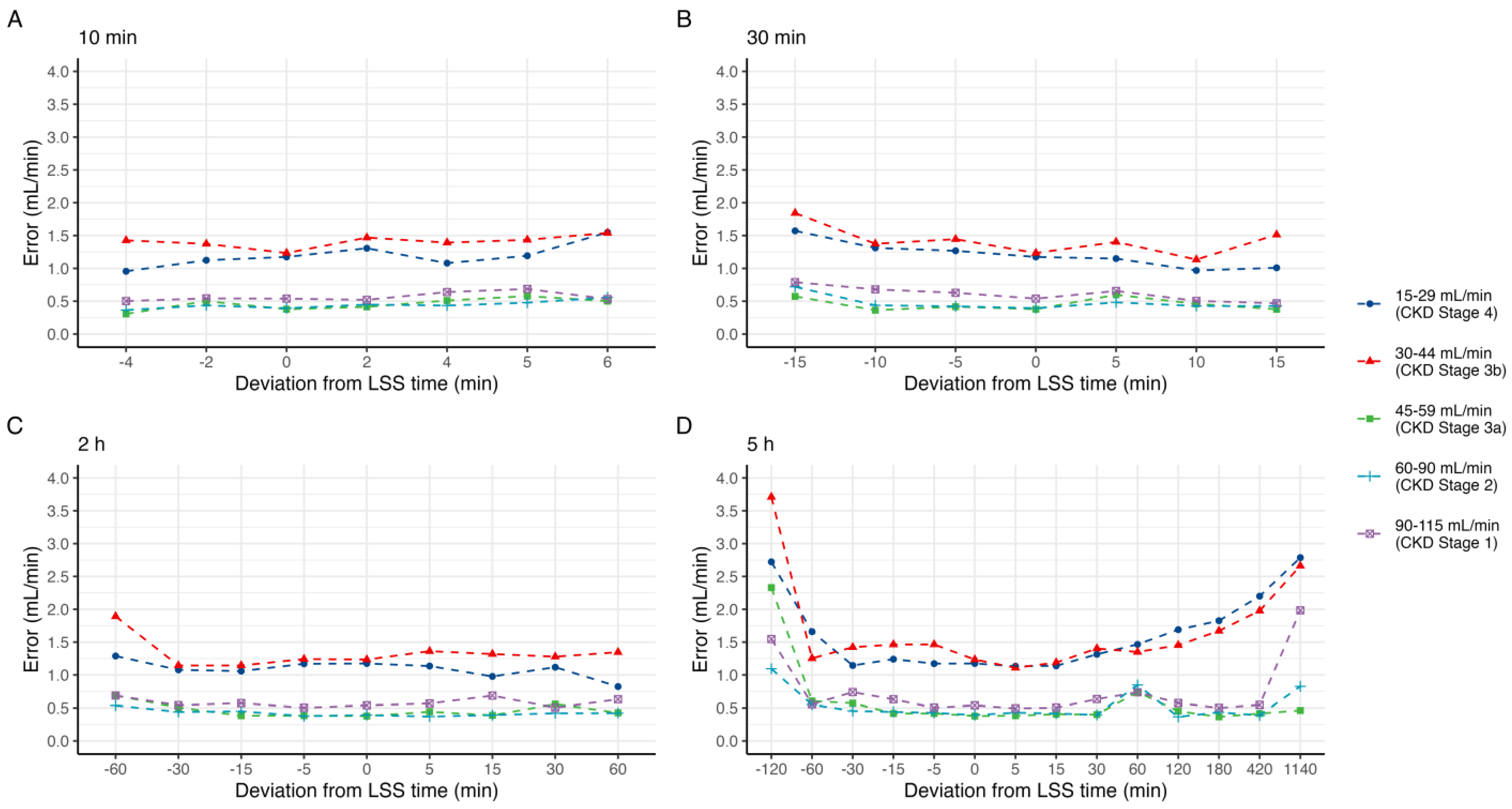
3.3. Effect of Shifts in Sample Times on Estimated GFR
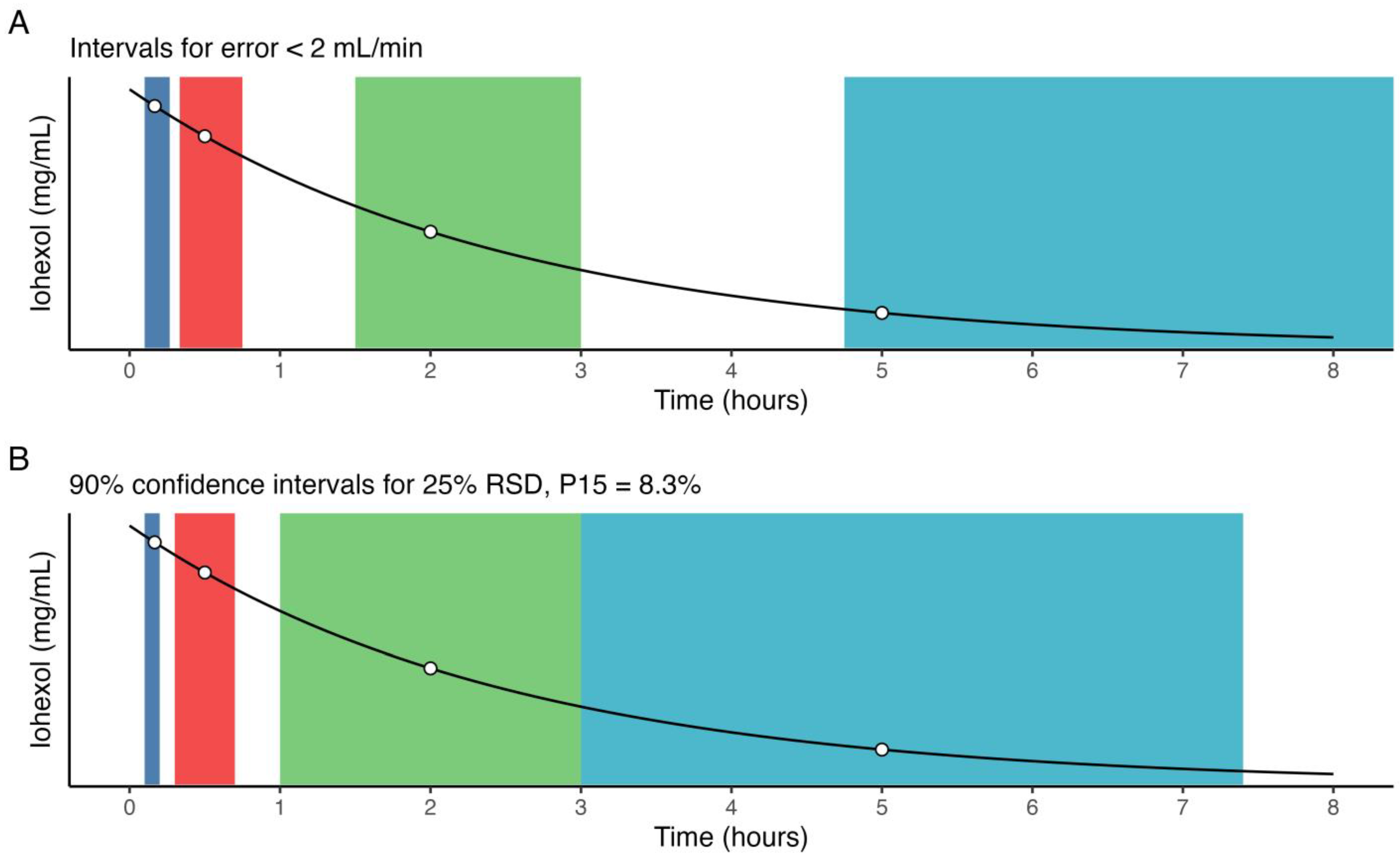
3.4. Optimal Sample Windows
3.5. Effect of Shifts in Sample Times on Model Parameters
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Lim, A.S.; Foo, S.H.W.; Benjamin Seng, J.J.; Magdeline Ng, T.T.; Chng, H.T.; Han, Z. Area-Under-Curve–Guided Versus Trough-Guided Monitoring of Vancomycin and Its Impact on Nephrotoxicity: A Systematic Review and Meta-Analysis. Ther. Drug Monit. 2023. [Google Scholar] [CrossRef] [PubMed]
- van den Elsen, S.H.J.; Sturkenboom, M.G.G.; Akkerman, O.W.; Manika, K.; Kioumis, I.P.; van der Werf, T.S.; Johnson, J.L.; Peloquin, C.; Touw, D.J.; Alffenaar, J.-W.C. Limited Sampling Strategies Using Linear Regression and the Bayesian Approach for Therapeutic Drug Monitoring of Moxifloxacin in Tuberculosis Patients. Antimicrob. Agents Chemother. 2019, 63, e00384-19. [Google Scholar] [CrossRef] [PubMed]
- Teitelbaum, Z.; Nassar, L.; Scherb, I.; Fink, D.; Ring, G.; Lurie, Y.; Krivoy, N.; Bentur, Y.; Efrati, E.; Kurnik, D. Limited Sampling Strategies Supporting Individualized Dose Adjustment of Intravenous Busulfan in Children and Young Adults. Ther. Drug Monit. 2020, 42, 427–434. [Google Scholar] [CrossRef] [PubMed]
- van der Meer, A.F.; Marcus, M.A.; Touw, D.J.; Proost, J.H.; Neef, C. Optimal sampling strategy development methodology using maximum a posteriori Bayesian estimation. Ther. Drug Monit. 2011, 33, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Porrini, E.; Ruggenenti, P.; Luis-Lima, S.; Carrara, F.; Jiménez, A.; de Vries, A.P.J.; Torres, A.; Gaspari, F.; Remuzzi, G. Estimated GFR: Time for a critical appraisal. Nat. Rev. Nephrol. 2019, 15, 177–190. [Google Scholar] [CrossRef] [PubMed]
- Dubourg, L.; Lemoine, S.; Joannard, B.; Chardon, L.; de Souza, V.; Cochat, P.; Iwaz, J.; Rabilloud, M.; Selistre, L. Comparison of iohexol plasma clearance formulas vs. inulin urinary clearance for measuring glomerular filtration rate. Clin. Chem. Lab. Med. 2021, 59, 571–579. [Google Scholar] [CrossRef] [PubMed]
- Delanaye, P.; Ebert, N.; Melsom, T.; Gaspari, F.; Mariat, C.; Cavalier, E.; Björk, J.; Christensson, A.; Nyman, U.; Porrini, E.; et al. Iohexol plasma clearance for measuring glomerular filtration rate in clinical practice and research: A review. Part 1: How to measure glomerular filtration rate with iohexol? Clin. Kidney J. 2016, 9, 682–699. [Google Scholar] [CrossRef]
- Destere, A.; Salmon Gandonnière, C.; Åsberg, A.; Loustaud-Ratti, V.; Carrier, P.; Ehrmann, S.; Barin-Le Guellec, C.; Marquet, P.; Woillard, J.B. A single Bayesian estimator for iohexol clearance estimation in ICU, liver failure and renal transplant patients. Br. J. Clin. Pharmacol. 2022, 88, 2793–2801. [Google Scholar] [CrossRef]
- Asberg, A.; Bjerre, A.; Almaas, R.; Luis-Lima, S.; Robertsen, I.; Salvador, C.L.; Porrini, E.; Schwartz, G.J.; Hartmann, A.; Bergan, S. Measured GFR by Utilizing Population Pharmacokinetic Methods to Determine Iohexol Clearance. Kidney Int. Rep. 2020, 5, 189–198. [Google Scholar] [CrossRef]
- Sarem, S.; Nekka, F.; Ahmed, I.S.; Litalien, C.; Li, J. Impact of sampling time deviations on the prediction of the area under the curve using regression limited sampling strategies. Biopharm. Drug Dispos. 2015, 36, 417–428. [Google Scholar] [CrossRef] [PubMed]
- Neely, M.N.; van Guilder, M.G.; Yamada, W.M.; Schumitzky, A.; Jelliffe, R.W. Accurate detection of outliers and subpopulations with Pmetrics, a nonparametric and parametric pharmacometric modeling and simulation package for R. Ther. Drug Monit. 2012, 34, 467–476. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Yamada, W.M.; Neely, M.N.; Bartroff, J.; Bayard, D.S.; Burke, J.V.; Guilder, M.V.; Jelliffe, R.W.; Kryshchenko, A.; Leary, R.; Tatarinova, T.; et al. An Algorithm for Nonparametric Estimation of a Multivariate Mixing Distribution with Applications to Population Pharmacokinetics. Pharmaceutics 2020, 13, 42. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, S.; Manjunath, B.G. tmvtnorm: A Package for the Truncated Multivariate Normal Distribution. R J. 2010, 2, 25–29. [Google Scholar] [CrossRef]
- Pastore, M.; Calcagni, A. Measuring Distribution Similarities Between Samples: A Distribution-Free Overlapping Index. Front. Psychol. 2019, 10, 1089. [Google Scholar] [CrossRef] [PubMed]
- Baron, K.T. Mrgsolve: Simulate from ODE-Based Models. R package version 1.0.6. 2022. Available online: https://CRAN.R-project.org/package=mrgsolve (accessed on 1 February 2023).
- Bayard, D.S.; Jelliffe, R.W. A Bayesian Approach to Tracking Patients Having Changing Pharmacokinetic Parameters. J. Pharmacokinet. Pharmacodyn. 2004, 31, 75–107. [Google Scholar] [CrossRef] [PubMed]


| Weighted Mean | Weighted Median (95% Credibility Interval) | |||
|---|---|---|---|---|
| Original | Simulated | Original | Simulated | |
| CL (L/h) | 2.89 | 2.84 | 1.95 (1.54–2.60) | 2.42 (2.16–2.72) |
| V (L) | 10.36 | 9.32 | 10.11 (9.19–10.91) | 8.98 (8.25–9.57) |
| Vp (L) | 9.20 | 7.98 | 7.95 (7.23–8.60) | 7.46 (7.06–7.81) |
| Q (L/h) | 10.65 | 11.37 | 8.03 (6.53–9.23) | 8.65 (7.50–9.72) |
| Group | Absolute Error (mL/min) | Relative Error (%) | P15 (%) | n |
|---|---|---|---|---|
| All profiles | 1.5 ± 2.2 | 4.1 ± 5.5 | 5.3 | 339 |
| CKD Stage 4 (15–29 mL/min) | 1.5 ± 1.3 | 6.3 ± 5.8 | 7.8 | 90 |
| CKD Stage 3b (30–44 mL/min) | 1.9 ± 2.2 | 5.4 ± 6.0 | 8.0 | 100 |
| CKD Stage 3a (45–59 mL/min) | 1.0 ± 1.6 | 2.2 ± 3.4 | 1.8 | 57 |
| CKD Stage 2 (60–90 mL/min) | 1.3 ± 3.0 | 1.9 ± 4.4 | 2.7 | 73 |
| CKD Stage 1 (90–115 mL/min) | 1.2 ± 2.2 | 1.2 ± 1.9 | 0.0 | 19 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hovd, M.; Robertsen, I.; Woillard, J.-B.; Åsberg, A. A Method for Evaluating Robustness of Limited Sampling Strategies—Exemplified by Serum Iohexol Clearance for Determination of Measured Glomerular Filtration Rate. Pharmaceutics 2023, 15, 1073. https://doi.org/10.3390/pharmaceutics15041073
Hovd M, Robertsen I, Woillard J-B, Åsberg A. A Method for Evaluating Robustness of Limited Sampling Strategies—Exemplified by Serum Iohexol Clearance for Determination of Measured Glomerular Filtration Rate. Pharmaceutics. 2023; 15(4):1073. https://doi.org/10.3390/pharmaceutics15041073
Chicago/Turabian StyleHovd, Markus, Ida Robertsen, Jean-Baptiste Woillard, and Anders Åsberg. 2023. "A Method for Evaluating Robustness of Limited Sampling Strategies—Exemplified by Serum Iohexol Clearance for Determination of Measured Glomerular Filtration Rate" Pharmaceutics 15, no. 4: 1073. https://doi.org/10.3390/pharmaceutics15041073
APA StyleHovd, M., Robertsen, I., Woillard, J.-B., & Åsberg, A. (2023). A Method for Evaluating Robustness of Limited Sampling Strategies—Exemplified by Serum Iohexol Clearance for Determination of Measured Glomerular Filtration Rate. Pharmaceutics, 15(4), 1073. https://doi.org/10.3390/pharmaceutics15041073






