New Drug Design Avenues Targeting Alzheimer’s Disease by Pharmacoinformatics-Aided Tools
Abstract
:1. Introduction
1.1. Main Hypotheses Currently Approved for AD
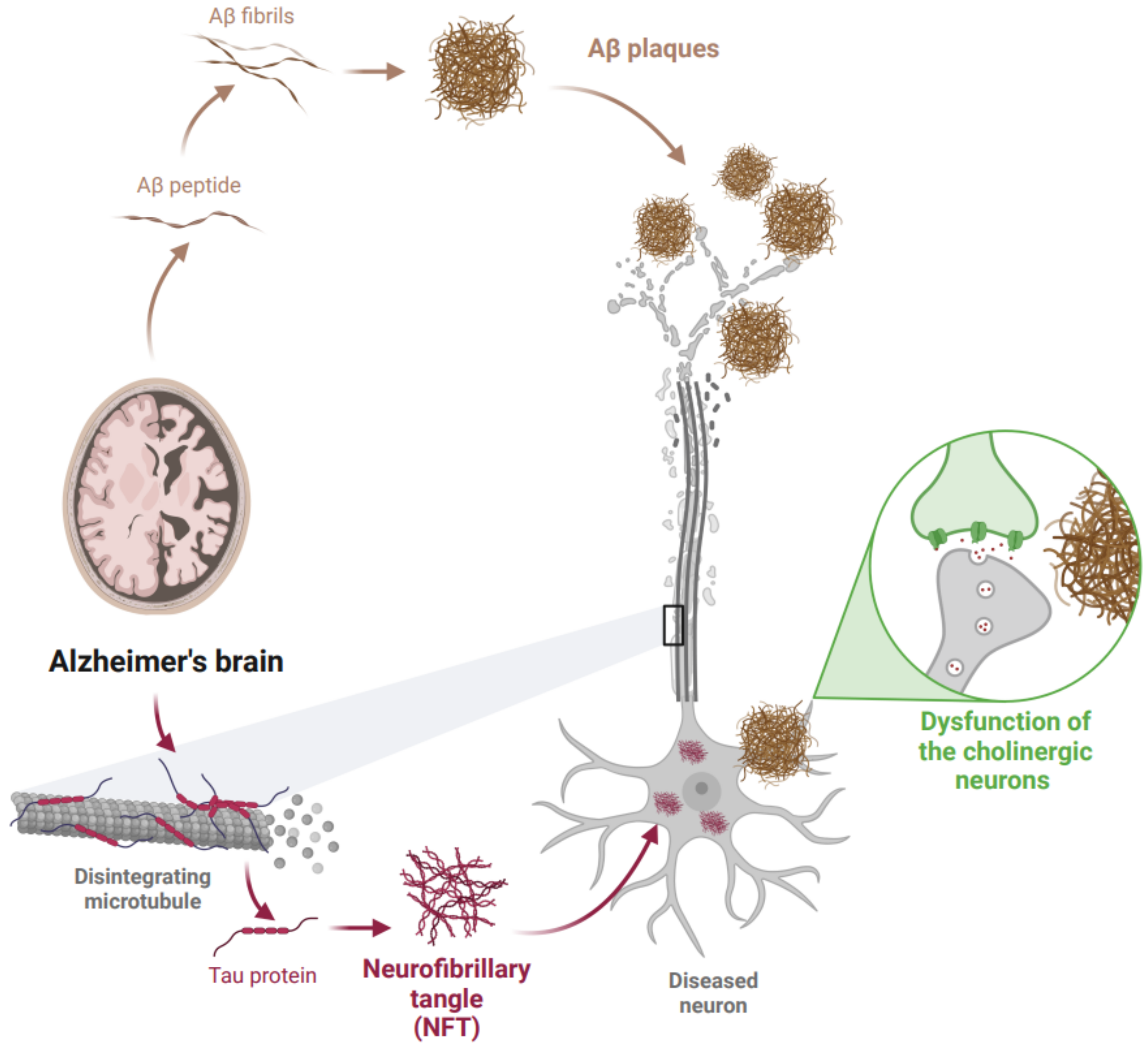
1.1.1. Cholinergic Hypothesis of AD
1.1.2. Amyloid Hypothesis of AD
1.1.3. Tau Hypothesis of AD
1.2. Other Hypotheses and New State of the Art in Pathophysiology of AD
2. Advances Achieved by Bioinformatics Tools in the Diagnosis of AD
3. Current Therapeutic Strategies against AD
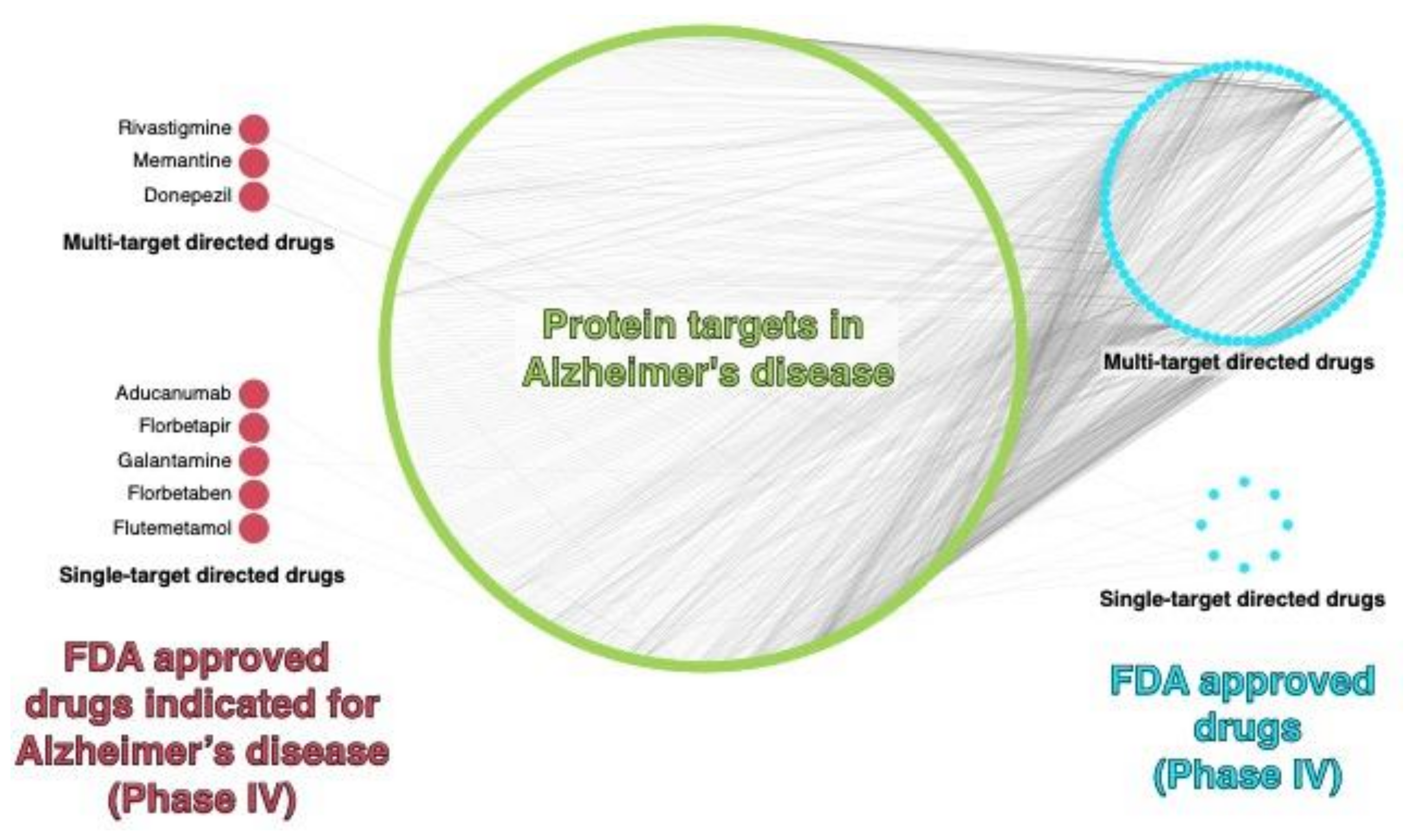
4. Computational Polypharmacology Applied to Multitarget Drug Design in AD
Multi-Target Directed Ligands (MTDLs) for AD
5. Pharmacoinformatics Tools in Drug Design against AD
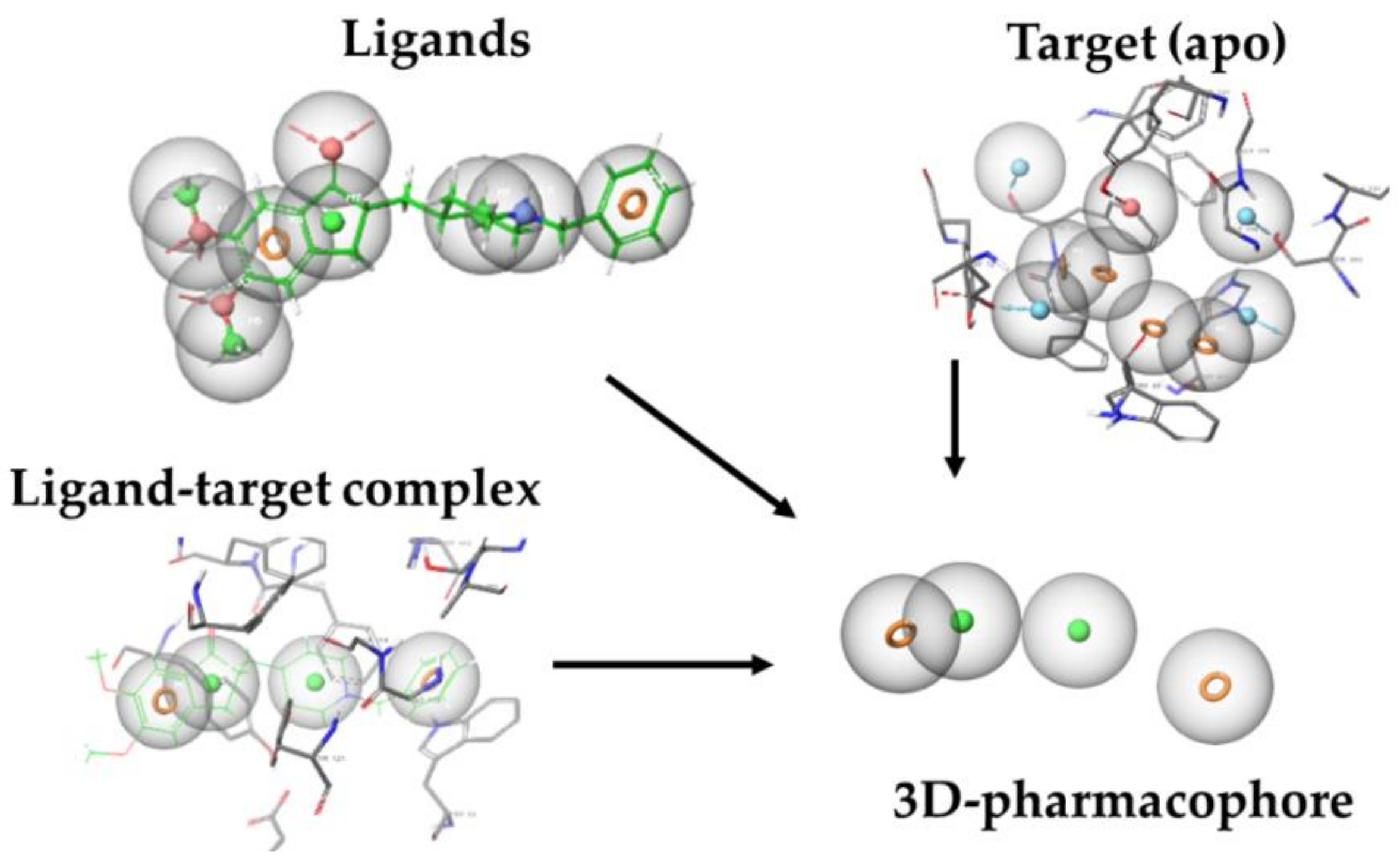
5.1. New Opportunities in Drug Discovery—Pharmacophore Modeling
Pharmacophore Modeling Classification
5.2. Machine Learning and Artificial Intelligence to Enhance Drug Design against AD
6. Drug Repurposing Strategies
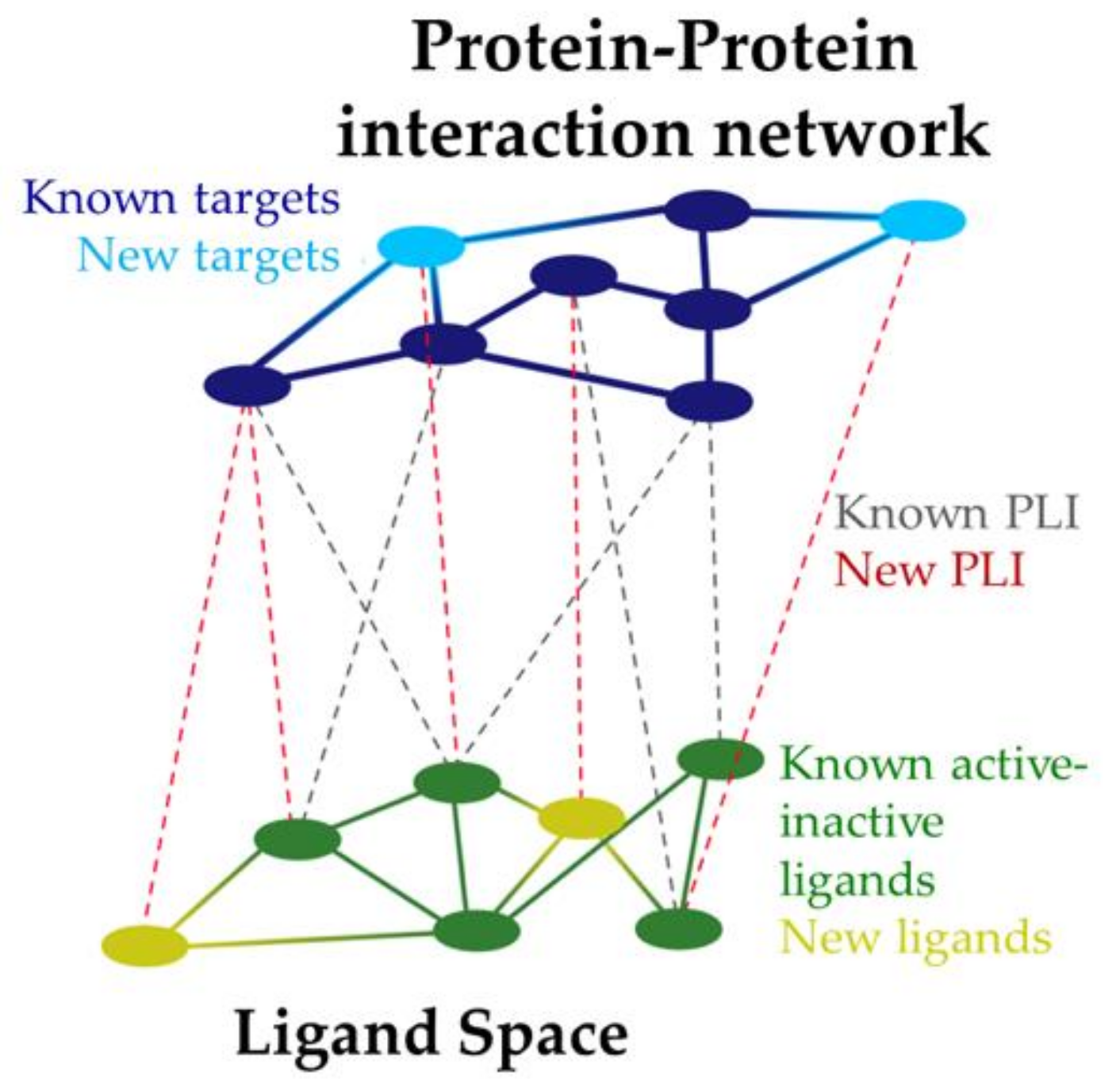
7. Applications of System Pharmacology in Drug Design against AD
8. Challenges and Future Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Niu, H.; Álvarez-Álvarez, I.; Guillén-Grima, F.; Aguinaga-Ontoso, I. Prevalencia e Incidencia de La Enfermedad de Alzheimer En Europa: Metaanálisis. Neurología 2017, 32, 523–532. [Google Scholar] [CrossRef] [PubMed]
- Islam, J.; Zhang, Y. Brain MRI Analysis for Alzheimer’s Disease Diagnosis Using an Ensemble System of Deep Convolutional Neural Networks. Brain Inform. 2018, 5, 2. [Google Scholar] [CrossRef] [PubMed]
- Benek, O.; Korabecny, J.; Soukup, O. A Perspective on Multi-Target Drugs for Alzheimer’s Disease. Trends Pharmacol. Sci. 2020, 41, 434–445. [Google Scholar] [CrossRef]
- Dumery, L.; Bourdel, F.; Soussan, Y.; Fialkowsky, A.; Viale, S.; Nicolas, P.; Reboud-Ravaux, M. β-Amyloid Protein Aggregation: Its Implication in the Physiopathology of Alzheimer’s Disease. Pathol. Biol. 2001, 49, 72–85. [Google Scholar] [CrossRef]
- Brion, J.P.; Anderton, B.H.; Authelet, M.; Dayanandan, R.; Leroy, K.; Lovestone, S.; Octave, J.N.; Pradier, L.; Touchet, N.; Tremp, G. Neurofibrillary Tangles and Tau Phosphorylation. Biochem. Soc. Symp. 2001, 67, 81–88. [Google Scholar] [CrossRef]
- Ramirez, D.; Arrue, L.; Reyes-parada, M. Targeting Alzheimer’ s Disease Through Pharmacoinformatics: New Challenges in Drug Design. Alzheimer’s Dis. Treat. 2021, 4, 1–5. [Google Scholar]
- Ower, A.K.; Hadjichrysanthou, C.; Gras, L.; Goudsmit, J.; Anderson, R.M.; de Wolf, F. Temporal Association Patterns and Dynamics of Amyloid-β and Tau in Alzheimer’s Disease. Eur. J. Epidemiol. 2018, 33, 657–666. [Google Scholar] [CrossRef]
- Aisen, P.S.; Cummings, J.; Jack, C.R.; Morris, J.C.; Sperling, R.; Frölich, L.; Jones, R.W.; Dowsett, S.A.; Matthews, B.R.; Raskin, J.; et al. On the Path to 2025: Understanding the Alzheimer’s Disease Continuum. Alzheimer’s Res. Ther. 2017, 9, 60. [Google Scholar] [CrossRef]
- Neubig, R.R.; Spedding, M.; Kenakin, T.; Christopoulos, A. Update on Terms and Symbols in Quantitative Pharmacology. Pharmacol. Rev. 2003, 55, 597–606. [Google Scholar] [CrossRef]
- Puzzo, D.; Arancio, O. Amyloid-β Peptide: Dr. Jekyll or Mr. Hyde? J. Alzheimer’s Dis. 2012, 33, S111–S120. [Google Scholar] [CrossRef]
- Driscoll, I.; Troncoso, J. Asymptomatic Alzheimer’s Disease: A Prodrome or a State of Resilience? Curr. Alzheimer Res. 2011, 8, 330–335. [Google Scholar] [CrossRef] [PubMed]
- Roher, A.E.; Chaney, M.O.; Kuo, Y.-M.; Webster, S.D.; Stine, W.B.; Haverkamp, L.J.; Woods, A.S.; Cotter, R.J.; Tuohy, J.M.; Krafft, G.A.; et al. Morphology and Toxicity of Aβ-(1-42) Dimer Derived from Neuritic and Vascular Amyloid Deposits of Alzheimer’s Disease. J. Biol. Chem. 1996, 271, 20631–20635. [Google Scholar] [CrossRef] [PubMed]
- Lesné, S.; Ming, T.K.; Kotilinek, L.; Kayed, R.; Glabe, C.G.; Yang, A.; Gallagher, M.; Ashe, K.H. A Specific Amyloid-β Protein Assembly in the Brain Impairs Memory. Nature 2006, 440, 352–357. [Google Scholar] [CrossRef] [PubMed]
- Piller, C. Blots on a Field? Science 2022, 377, 358–363. [Google Scholar] [CrossRef]
- Weller, J.; Budson, A. Current Understanding of Alzheimer’s Disease Diagnosis and Treatment. F1000Research 2018, 7, 1–9. [Google Scholar] [CrossRef]
- Kehoe, P.G. The Coming of Age of the Angiotensin Hypothesis in Alzheimer’s Disease: Progress Toward Disease Prevention and Treatment? J. Alzheimer’s Dis. 2018, 62, 1443–1466. [Google Scholar] [CrossRef]
- De la Torre, J.C. The Vascular Hypothesis of Alzheimer’s Disease: Bench to Bedside and Beyond. Neurodegener. Dis. 2010, 7, 116–121. [Google Scholar] [CrossRef] [PubMed]
- Markesbery, W.R. Oxidative Stress Hypothesis in Alzheimer’s Disease. Free Radic. Biol. Med. 1997, 23, 134–147. [Google Scholar] [CrossRef]
- Siblerud, R.; Mutter, J.; Moore, E.; Naumann, J.; Walach, H. A Hypothesis and Evidence That Mercury May Be an Etiological Factor in Alzheimer’s Disease. Int. J. Environ. Res. Public Health 2019, 16, 5152. [Google Scholar] [CrossRef]
- Wood, W.G.; Li, L.; Müller, W.E.; Eckert, G.P. Cholesterol as a Causative Factor in Alzheimer’s Disease: A Debatable Hypothesis. J. Neurochem. 2014, 129, 559–572. [Google Scholar] [CrossRef]
- Gezen-Ak, D.; Yılmazer, S.; Dursun, E. Why Vitamin D in Alzheimer’s Disease? The Hypothesis. J. Alzheimer’s Dis. 2014, 40, 257–269. [Google Scholar] [CrossRef] [PubMed]
- Berridge, M.J. Calcium Hypothesis of Alzheimer’s Disease. Pflügers Arch. Eur. J. Physiol. 2010, 459, 441–449. [Google Scholar] [CrossRef]
- Area-Gomez, E.; Schon, E.A. On the Pathogenesis of Alzheimer’s Disease: The MAM Hypothesis. FASEB J. 2017, 31, 864–867. [Google Scholar] [CrossRef] [PubMed]
- Devanand, D.P. Viral Hypothesis and Antiviral Treatment in Alzheimer’s Disease. Curr. Neurol. Neurosci. Rep. 2018, 18, 55. [Google Scholar] [CrossRef] [PubMed]
- Adelusi, T.I.; Oyedele, A.Q.K.; Boyenle, I.D.; Ogunlana, A.T.; Adeyemi, R.O.; Ukachi, C.D.; Idris, M.O.; Olaoba, O.T.; Adedotun, I.O.; Kolawole, O.E.; et al. Molecular Modeling in Drug Discovery. Inform. Med. Unlocked 2022, 29, 100880. [Google Scholar] [CrossRef]
- Schaduangrat, N.; Lampa, S.; Simeon, S.; Gleeson, M.P.; Spjuth, O.; Nantasenamat, C. Towards Reproducible Computational Drug Discovery. J. Cheminform. 2020, 12, 9. [Google Scholar] [CrossRef]
- Rossi, M.; Freschi, M.; De Camargo Nascente, L.; Salerno, A.; De Melo Viana Teixeira, S.; Nachon, F.; Chantegreil, F.; Soukup, O.; Prchal, L.; Malaguti, M.; et al. Sustainable Drug Discovery of Multi-Target-Directed Ligands for Alzheimer’s Disease. J. Med. Chem. 2021, 64, 4972–4990. [Google Scholar] [CrossRef]
- Jarada, T.N.; Rokne, J.G.; Alhajj, R. A Review of Computational Drug Repositioning: Strategies, Approaches, Opportunities, Challenges, and Directions. J. Cheminform. 2020, 12, 46. [Google Scholar] [CrossRef]
- Azer, K.; Kaddi, C.D.; Barrett, J.S.; Bai, J.P.F.; McQuade, S.T.; Merrill, N.J.; Piccoli, B.; Neves-Zaph, S.; Marchetti, L.; Lombardo, R.; et al. History and Future Perspectives on the Discipline of Quantitative Systems Pharmacology Modeling and Its Applications. Front. Physiol. 2021, 12, 637999. [Google Scholar] [CrossRef]
- Aghamiri, S.S.; Amin, R.; Helikar, T. Recent Applications of Quantitative Systems Pharmacology and Machine Learning Models across Diseases. J. Pharmacokinet. Pharmacodyn. 2022, 49, 19–37. [Google Scholar] [CrossRef]
- Salmaso, V.; Moro, S. Bridging Molecular Docking to Molecular Dynamics in Exploring Ligand-Protein Recognition Process: An Overview. Front. Pharmacol. 2018, 9, 923. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pinzi, L.; Rastelli, G. Molecular Docking: Shifting Paradigms in Drug Discovery. Int. J. Mol. Sci. 2019, 20, 4331. [Google Scholar] [CrossRef] [PubMed]
- Gimeno, A.; Ojeda-Montes, M.J.; Tomás-Hernández, S.; Cereto-Massagué, A.; Beltrán-Debón, R.; Mulero, M.; Pujadas, G.; Garcia-Vallvé, S. The Light and Dark Sides of Virtual Screening: What Is There to Know? Int. J. Mol. Sci. 2019, 20, 1375. [Google Scholar] [CrossRef]
- Maia, E.H.B.; Assis, L.C.; de Oliveira, T.A.; da Silva, A.M.; Taranto, A.G. Structure-Based Virtual Screening: From Classical to Artificial Intelligence. Front. Chem. 2020, 8, 343. [Google Scholar] [CrossRef] [PubMed]
- Walters, W.P.; Wang, R. New Trends in Virtual Screening. J. Chem. Inf. Model. 2020, 60, 4109–4111. [Google Scholar] [CrossRef] [PubMed]
- Schaller, D.; Šribar, D.; Noonan, T.; Deng, L.; Nguyen, T.N.; Pach, S.; Machalz, D.; Bermudez, M.; Wolber, G. Next Generation 3D Pharmacophore Modeling. WIREs Comput. Mol. Sci. 2020, 10, e1468. [Google Scholar] [CrossRef]
- Wolber, G.; Langer, T. LigandScout: 3-D Pharmacophores Derived from Protein-Bound Ligands and Their Use as Virtual Screening Filters. J. Chem. Inf. Model. 2005, 45, 160–169. [Google Scholar] [CrossRef]
- Saurabh, S.; Sivakumar, P.M.; Perumal, V.; Khosravi, A.; Sugumaran, A.; Prabhawathi, V. Molecular Dynamics Simulations in Drug Discovery and Drug Delivery. In Integrative Nanomedicine for New Therapies; Springer: Berlin/Heidelberg, Germany, 2020; pp. 275–301. [Google Scholar] [CrossRef]
- Liu, X.; Shi, D.; Zhou, S.; Liu, H.; Liu, H.; Yao, X. Molecular Dynamics Simulations and Novel Drug Discovery. Expert Opin. Drug Discov. 2018, 13, 23–37. [Google Scholar] [CrossRef]
- Fang, J.; Wang, L.; Li, Y.; Lian, W.; Pang, X.; Wang, H.; Yuan, D.; Wang, Q.; Liu, A.-L.; Du, G.-H. AlzhCPI: A Knowledge Base for Predicting Chemical-Protein Interactions towards Alzheimer’s Disease. PLoS ONE 2017, 12, e0178347. [Google Scholar] [CrossRef]
- Liu, H.; Wang, L.; Lv, M.; Pei, R.; Li, P.; Pei, Z.; Wang, Y.; Su, W.; Xie, X.-Q. AlzPlatform: An Alzheimer’s Disease Domain-Specific Chemogenomics Knowledgebase for Polypharmacology and Target Identification Research. J. Chem. Inf. Model. 2014, 54, 1050–1060. [Google Scholar] [CrossRef]
- Sügis, E.; Dauvillier, J.; Leontjeva, A.; Adler, P.; Hindie, V.; Moncion, T.; Collura, V.; Daudin, R.; Loe-Mie, Y.; Herault, Y.; et al. HENA, Heterogeneous Network-Based Data Set for Alzheimer’s Disease. Sci. Data 2019, 6, 151. [Google Scholar] [CrossRef] [PubMed]
- Kuzma, A.; Valladares, O.; Cweibel, R.; Greenfest-Allen, E.; Childress, D.M.; Malamon, J.; Gangadharan, P.; Zhao, Y.; Qu, L.; Leung, Y.Y.; et al. NIAGADS: The NIA Genetics of Alzheimer’s Disease Data Storage Site. Alzheimer’s Dement. 2016, 12, 1200–1203. [Google Scholar] [CrossRef]
- Kaur, S.; DasGupta, G.; Singh, S. Altered Neurochemistry in Alzheimer’s Disease: Targeting Neurotransmitter Receptor Mechanisms and Therapeutic Strategy. Neurophysiology 2019, 51, 293–309. [Google Scholar] [CrossRef]
- Guzman-Martinez, L.; Calfío, C.; Farias, G.A.; Vilches, C.; Prieto, R.; Maccioni, R.B. New Frontiers in the Prevention, Diagnosis, and Treatment of Alzheimer’s Disease. J. Alzheimer’s Dis. 2021, 82, S51–S63. [Google Scholar] [CrossRef] [PubMed]
- Frisoni, G.B.; Fox, N.C.; Jack, C.R.; Scheltens, P.; Thompson, P.M. The Clinical Use of Structural MRI in Alzheimer Disease. Nat. Rev. Neurol. 2010, 6, 67–77. [Google Scholar] [CrossRef]
- Yu, W.; Yu, W.; Yang, Y.; Lü, Y. Exploring the Key Genes and Identification of Potential Diagnosis Biomarkers in Alzheimer’s Disease Using Bioinformatics Analysis. Front. Aging Neurosci. 2021, 13, 276. [Google Scholar] [CrossRef]
- Wang, X.; Wang, L. Screening and Identification of Potential Peripheral Blood Biomarkers for Alzheimer’s Disease Based on Bioinformatics Analysis. Med. Sci. Monit. 2020, 26, e924263. [Google Scholar] [CrossRef]
- Drew, K.; Lee, C.; Huizar, R.L.; Tu, F.; Borgeson, B.; McWhite, C.D.; Ma, Y.; Wallingford, J.B.; Marcotte, E.M. Integration of over 9,000 Mass Spectrometry Experiments Builds a Global Map of Human Protein Complexes. Mol. Syst. Biol. 2017, 13, 932. [Google Scholar] [CrossRef]
- Petersen, R.C.; Aisen, P.S.; Beckett, L.A.; Donohue, M.C.; Gamst, A.C.; Harvey, D.J.; Jack, C.R.; Jagust, W.J.; Shaw, L.M.; Toga, A.W.; et al. Alzheimer’s Disease Neuroimaging Initiative (ADNI): Clinical Characterization. Neurology 2010, 74, 201–209. [Google Scholar] [CrossRef]
- He, S.; Dou, L.; Li, X.; Zhang, Y. Review of Bioinformatics in Azheimer’s Disease Research. Comput. Biol. Med. 2022, 143, 105269. [Google Scholar] [CrossRef]
- Tan, M.S.; Cheah, P.-L.; Chin, A.-V.; Looi, L.-M.; Chang, S.-W. A Review on Omics-Based Biomarkers Discovery for Alzheimer’s Disease from the Bioinformatics Perspectives: Statistical Approach vs Machine Learning Approach. Comput. Biol. Med. 2021, 139, 104947. [Google Scholar] [CrossRef] [PubMed]
- Beach, T.G.; Monsell, S.E.; Phillips, L.E.; Kukull, W. Accuracy of the Clinical Diagnosis of Alzheimer Disease at National Institute on Aging Alzheimer Disease Centers, 2005–2010. J. Neuropathol. Exp. Neurol. 2012, 71, 266–273. [Google Scholar] [CrossRef] [PubMed]
- Jack, C.R.; Bennett, D.A.; Blennow, K.; Carrillo, M.C.; Feldman, H.H.; Frisoni, G.B.; Hampel, H.; Jagust, W.J.; Johnson, K.A.; Knopman, D.S.; et al. A/T/N: An Unbiased Descriptive Classification Scheme for Alzheimer Disease Biomarkers. Neurology 2016, 87, 539–547. [Google Scholar] [CrossRef] [PubMed]
- Maccioni, R.B.; Muñoz, J.P.; Barbeito, L. The Molecular Bases of Alzheimer’s Disease and Other Neurodegenerative Disorders. Arch. Med. Res. 2001, 32, 367–381. [Google Scholar] [CrossRef]
- Mullane, K.; Williams, M. Alzheimer’s Disease (AD) Therapeutics--1: Repeated Clinical Failures Continue to Question the Amyloid Hypothesis of AD and the Current Understanding of AD Causality. Biochem. Pharmacol. 2018, 158, 359–375. [Google Scholar] [CrossRef] [PubMed]
- Montanari, S.; Bartolini, M.; Neviani, P.; Belluti, F.; Gobbi, S.; Pruccoli, L.; Tarozzi, A.; Falchi, F.; Andrisano, V.; Miszta, P.; et al. Multitarget Strategy to Address Alzheimer’s Disease: Design, Synthesis, Biological Evaluation, and Computational Studies of Coumarin-Based Derivatives. ChemMedChem 2016, 11, 1296–1308. [Google Scholar] [CrossRef]
- Husna Ibrahim, N.; Yahaya, M.F.; Mohamed, W.; Teoh, S.L.; Hui, C.K.; Kumar, J. Pharmacotherapy of Alzheimer’s Disease: Seeking Clarity in a Time of Uncertainty. Front. Pharmacol. 2020, 11, 261. [Google Scholar] [CrossRef]
- Kevadiya, B.D.; Ottemann, B.M.; Thomas, M.B.; Mukadam, I.; Nigam, S.; McMillan, J.; Gorantla, S.; Bronich, T.K.; Edagwa, B.; Gendelman, H.E. Neurotheranostics as Personalized Medicines. Adv. Drug Deliv. Rev. 2019, 148, 252–289. [Google Scholar] [CrossRef] [PubMed]
- Jeremic, D.; Jiménez-Díaz, L.; Navarro-López, J.D. Past, Present and Future of Therapeutic Strategies against Amyloid-β Peptides in Alzheimer’s Disease: A Systematic Review. Ageing Res. Rev. 2021, 72, 101496. [Google Scholar] [CrossRef]
- Crismon, M.L. Tacrine: First Drug Approved for Alzheimer’s Disease. Ann. Pharmacother. 1994, 28, 744–751. [Google Scholar] [CrossRef]
- Knopman, D. Tacrine for Alzheimer’s Disease. Pharmacoeconomics 1995, 7, 275–279. [Google Scholar] [CrossRef] [PubMed]
- Bautista-Aguilera, Ó.M.; Ismaili, L.; Iriepa, I.; Diez-Iriepa, D.; Chabchoub, F.; Marco-Contelles, J.; Pérez, M. Tacrines as Therapeutic Agents for Alzheimer’s Disease. V. Recent Developments. Chem. Rec. 2021, 21, 162–174. [Google Scholar] [CrossRef] [PubMed]
- Reddy, E.K.; Remya, C.; Mantosh, K.; Sajith, A.M.; Omkumar, R.V.; Sadasivan, C.; Anwar, S. Novel Tacrine Derivatives Exhibiting Improved Acetylcholinesterase Inhibition: Design, Synthesis and Biological Evaluation. Eur. J. Med. Chem. 2017, 139, 367–377. [Google Scholar] [CrossRef]
- Zha, X.; Lamba, D.; Zhang, L.; Lou, Y.; Xu, C.; Kang, D.; Chen, L.; Xu, Y.; Zhang, L.; De Simone, A.; et al. Novel Tacrine-Benzofuran Hybrids as Potent Multitarget-Directed Ligands for the Treatment of Alzheimers Disease: Design, Synthesis, Biological Evaluation, and X-Ray Crystallography. J. Med. Chem. 2016, 59, 114–131. [Google Scholar] [CrossRef] [PubMed]
- Sameem, B.; Saeedi, M.; Mahdavi, M.; Shafiee, A. A Review on Tacrine-Based Scaffolds as Multi-Target Drugs (MTDLs) for Alzheimer’s Disease. Eur. J. Med. Chem. 2017, 128, 332–345. [Google Scholar] [CrossRef]
- Tumiatti, V.; Minarini, A.; Bolognesi, M.L.; Milelli, A.; Rosini, M.; Melchiorre, C. Tacrine Derivatives and Alzheimers Disease. Curr. Med. Chem. 2010, 17, 1825–1838. [Google Scholar] [CrossRef]
- Arndt, J.W.; Qian, F.; Smith, B.A.; Quan, C.; Kilambi, K.P.; Bush, M.W.; Walz, T.; Pepinsky, R.B.; Bussière, T.; Hamann, S.; et al. Structural and Kinetic Basis for the Selectivity of Aducanumab for Aggregated Forms of Amyloid-β. Sci. Rep. 2018, 8, 6412. [Google Scholar] [CrossRef]
- Harilal, S.; Jose, J.; Parambi, D.G.T.; Kumar, R.; Mathew, G.E.; Uddin, M.S.; Kim, H.; Mathew, B. Advancements in Nanotherapeutics for Alzheimer’s Disease: Current Perspectives. J. Pharm. Pharmacol. 2019, 71, 1370–1383. [Google Scholar] [CrossRef]
- Ross, C.; Taylor, M.; Fullwood, N.; Allsop, D. Liposome Delivery Systems for the Treatment of Alzheimer’s Disease. Int. J. Nanomed. 2018, 13, 8507–8522. [Google Scholar] [CrossRef]
- Yiannopoulou, K.G.; Papageorgiou, S.G. Current and Future Treatments in Alzheimer Disease: An Update. J. Cent. Nerv. Syst. Dis. 2020, 12, 117957352090739. [Google Scholar] [CrossRef] [Green Version]
- Nisticò, R.; Borg, J.J. Aducanumab for Alzheimer’s Disease: A Regulatory Perspective. Pharmacol. Res. 2021, 171, 105754. [Google Scholar] [CrossRef] [PubMed]
- Hollmann, P. Update: FDA Approval of Biogen’s Aducanumab. Geriatr. Nurs. 2022, 43, 318–319. [Google Scholar] [CrossRef] [PubMed]
- FDA. FDA Grants Accelerated Approval 986 for Alzheimer’s Drug. Available online: https://www.fda.gov/news-events/press-announcements/fda-grants-accelerated-approval-alzheimers-drug (accessed on 1 July 2022).
- Becker, R.E.; Greig, N.H.; Giacobini, E. Why Do So Many Drugs for Alzheimer’s Disease Fail in Development? Time for New Methods and New Practices? Bone 2008, 15, 303–325. [Google Scholar] [CrossRef] [PubMed]
- Guest, F.L.; Rahmoune, H.; Guest, P.C. Advances in experimental medicine and biology. In Reviews on New Drug Targets in Age-Related Disorders; Guest, P.C., Ed.; Springer International Publishing: Cham, Switzerland, 2021; Volume 1286, ISBN 978-3-030-55034-9. [Google Scholar]
- Cummings, J. Lessons Learned from Alzheimer Disease: Clinical Trials with Negative Outcomes. Clin. Transl. Sci. 2018, 11, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Mesh. Polypharmacology. Available online: https://www.ncbi.nlm.nih.gov/mesh/?term=polypharmacology (accessed on 1 July 2022).
- Rastelli, G.; Pinzi, L. Computational Polypharmacology Comes of Age. Front. Pharmacol. 2015, 6, 157. [Google Scholar] [CrossRef]
- Oddsson, S.; Kowal, N.M.; Ahring, P.K.; Olafsdottir, E.S.; Balle, T. Structure-Based Discovery of Dual-Target Hits for Acetylcholinesterase and the A7 Nicotinic Acetylcholine Receptors: In Silico Studies and In Vitro Confirmation. Molecules 2020, 25, 2872. [Google Scholar] [CrossRef]
- Oyinloye, B.E.; Iwaloye, O.; Ajiboye, B.O. Polypharmacology of Gongronema Latifolium Leaf Secondary Metabolites against Protein Kinases Implicated in Parkinson’s Disease and Alzheimer’s Disease. Sci. Afr. 2021, 12, e00826. [Google Scholar] [CrossRef]
- Nozal, V.; García-Rubia, A.; Cuevas, E.P.; Pérez, C.; Tosat-Bitrián, C.; Bartolomé, F.; Carro, E.; Ramírez, D.; Palomo, V.; Martínez, A. From Kinase Inhibitors to Multitarget Ligands as Powerful Drug Leads for Alzheimer’s Disease Using Protein-Templated Synthesis. Angew. Chem. Int. Ed. 2021, 35, 19344–19354. [Google Scholar] [CrossRef]
- Rosini, M.; Simoni, E.; Minarini, A.; Melchiorre, C. Multi-Target Design Strategies in the Context of Alzheimer’s Disease: Acetylcholinesterase Inhibition and NMDA Receptor Antagonism as the Driving Forces. Neurochem. Res. 2014, 39, 1914–1923. [Google Scholar] [CrossRef] [PubMed]
- Núñez-Vivanco, G.; Fierro, A.; Moya, P.; Iturriaga-Vásquez, P.; Reyes-Parada, M. 3D Similarities between the Binding Sites of Monoaminergic Target Proteins. PLoS ONE 2018, 13, e0200637. [Google Scholar] [CrossRef]
- Wenzel, T.J.; Klegeris, A. Novel Multi-Target Directed Ligand-Based Strategies for Reducing Neuroinflammation in Alzheimer’s Disease. Life Sci. 2018, 207, 314–322. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Franco, M.I.; Fernández-Bachiller, M.I.; Pérez, C.; Hernández-Ledesma, B.; Bartolomé, B. Novel Tacrine- Melatonin Hybrids as Dual-Acting Drugs for Alzheimer Disease, with Improved Acetylcholinesterase Inhibitory and Antioxidant Properties. J. Med. Chem. 2006, 49, 459–462. [Google Scholar] [CrossRef]
- Fernández-Bachiller, M.I.; Pérez, C.; Monjas, L.; Rademann, J.; Rodríguez-Franco, M.I. New Tacrine–4-Oxo-4 H-Chromene Hybrids as Multifunctional Agents for the Treatment of Alzheimer’s Disease, with Cholinergic, Antioxidant, and β-Amyloid-Reducing Properties. J. Med. Chem. 2012, 55, 1303–1317. [Google Scholar] [CrossRef]
- Zhang, P.; Wang, Z.; Mou, C.; Zou, J.; Xie, Y.; Liu, Z.; Benjamin Naman, C.; Mao, Y.; Wei, J.; Huang, X.; et al. Design and Synthesis of Novel Tacrine-Dipicolylamine Dimers That Are Multiple-Target-Directed Ligands with Potential to Treat Alzheimer’s Disease. Bioorg. Chem. 2021, 116, 105387. [Google Scholar] [CrossRef]
- Benek, O.; Soukup, O.; Pasdiorova, M.; Hroch, L.; Sepsova, V.; Jost, P.; Hrabinova, M.; Jun, D.; Kuca, K.; Zala, D.; et al. Design, Synthesis and in Vitro Evaluation of Indolotacrine Analogues as Multitarget-Directed Ligands for the Treatment of Alzheimer’s Disease. Chem. Med. Chem. 2016, 1264–1269. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, R.R.; Popovic-Nikolic, M.R.; Nikolic, K.; Uliassi, E.; Bolognesi, M.L. A Perspective on Multi-Target Drug Discovery and Design for Complex Diseases. Clin. Transl. Med. 2018, 7, 3. [Google Scholar] [CrossRef]
- Domínguez, J.L.; Fernández-Nieto, F.; Castro, M.; Catto, M.; Paleo, M.R.; Porto, S.; Sardina, F.J.; Brea, J.M.; Carotti, A.; Villaverde, M.C.; et al. Computer-Aided Structure-Based Design of Multitarget Leads for Alzheimer’s Disease. J. Chem. Inf. Model. 2015, 55, 135–148. [Google Scholar] [CrossRef]
- Núñez-Vivanco, G.; Valdés-Jiménez, A.; Besoaín, F.; Reyes-Parada, M. Geomfinder: A Multi-Feature Identifier of Similar Three-Dimensional Protein Patterns: A Ligand-Independent Approach. J. Cheminform. 2016, 8, 19. [Google Scholar] [CrossRef]
- Valdés-Jiménez, A.; Larriba-Pey, J.; Reyes-Parada, M.; Nuñez-Vivanco, G. 3D-PP: A Tool for Discovering Conserved 3D Protein Patterns. In Proceedings of the International Conference on Multidisciplinary Sciences, Ebene, Mauritius, 22–23 June 2018. [Google Scholar]
- Konc, J.; Janežič, D. ProBiS: A Web Server for Detection of Structurally Similar Protein Binding Sites. Nucleic Acids Res. 2010, 38, W436–W440. [Google Scholar] [CrossRef]
- Eguida, M.; Rognan, D. A Computer Vision Approach to Align and Compare Protein Cavities: Application to Fragment-Based Drug Design. J. Med. Chem. 2020, 63, 7127–7142. [Google Scholar] [CrossRef]
- Yeturu, K.; Chandra, N. PocketMatch: A New Algorithm to Compare Binding Sites in Protein Structures. BMC Bioinform. 2008, 9, 543. [Google Scholar] [CrossRef] [PubMed]
- Duran-Frigola, M.; Siragusa, L.; Ruppin, E.; Barril, X.; Cruciani, G.; Aloy, P. Detecting Similar Binding Pockets to Enable Systems Polypharmacology. PLoS Comput. Biol. 2017, 13, e1005522. [Google Scholar] [CrossRef]
- Ehrt, C.; Brinkjost, T.; Koch, O. A Benchmark Driven Guide to Binding Site Comparison: An Exhaustive Evaluation Using Tailor-Made Data Sets (ProSPECCTs). PLoS Comput. Biol. 2018, 14, e1006483. [Google Scholar] [CrossRef]
- Naderi, M.; Lemoine, J.M.; Govindaraj, R.G.; Kana, O.Z.; Feinstein, W.P.; Brylinski, M. Binding Site Matching in Rational Drug Design: Algorithms and Applications. Brief. Bioinform. 2019, 20, 2167–2184. [Google Scholar] [CrossRef] [PubMed]
- Ehrt, C.; Brinkjost, T.; Koch, O. Binding Site Characterization-Similarity, Promiscuity, and Druggability. Medchemcomm 2019, 10, 1145–1159. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Chen, M.; Liu, Z.; Fang, X.; Gou, S.; Chen, L. Ferulic Acid-Carbazole Hybrid Compounds: Combination of Cholinesterase Inhibition, Antioxidant and Neuroprotection as Multifunctional Anti-Alzheimer Agents. Bioorganic Med. Chem. 2016, 24, 886–893. [Google Scholar] [CrossRef]
- Chen, Z.; Digiacomo, M.; Tu, Y.; Gu, Q.; Wang, S.; Yang, X.; Chu, J.; Chen, Q.; Han, Y.; Chen, J.; et al. Discovery of Novel Rivastigmine-Hydroxycinnamic Acid Hybrids as Multi-Targeted Agents for Alzheimer’s Disease. Eur. J. Med. Chem. 2017, 125, 784–792. [Google Scholar] [CrossRef]
- Dias, K.S.T.; de Paula, C.T.; dos Santos, T.; Souza, I.N.O.; Boni, M.S.; Guimarães, M.J.R.; da Silva, F.M.R.; Castro, N.G.; Neves, G.A.; Veloso, C.C.; et al. Design, Synthesis and Evaluation of Novel Feruloyl-Donepezil Hybrids as Potential Multitarget Drugs for the Treatment of Alzheimer’s Disease. Eur. J. Med. Chem. 2017, 130, 440–457. [Google Scholar] [CrossRef]
- Prati, F.; Bottegoni, G.; Bolognesi, M.L.; Cavalli, A. BACE-1 Inhibitors: From Recent Single-Target Molecules to Multitarget Compounds for Alzheimer’s Disease. J. Med. Chem. 2018, 61, 619–637. [Google Scholar] [CrossRef]
- Prati, F.; Cavalli, A.; Bolognesi, M.L. Navigating the Chemical Space of Multitarget-Directed Ligands: From Hybrids to Fragments in Alzheimer’s Disease. Molecules 2016, 21, 466. [Google Scholar] [CrossRef] [Green Version]
- Seong, S.H.; Ali, M.Y.; Kim, H.R.; Jung, H.A.; Choi, J.S. BACE1 Inhibitory Activity and Molecular Docking Analysis of Meroterpenoids from Sargassum Serratifolium. Bioorganic Med. Chem. 2017, 25, 3964–3970. [Google Scholar] [CrossRef] [PubMed]
- Rao, V.S.; Srinivas, K.; Kumar, G.S.; Sujin, G. Protein Interaction Network for Alzheimer’s Disease Using Computational Approach. Bioinformation 2013, 9, 968–972. [Google Scholar] [CrossRef]
- Caberlotto, L.; Nguyen, T.-P. A Systems Biology Investigation of Neurodegenerative Dementia Reveals a Pivotal Role of Autophagy. BMC Syst. Biol. 2014, 8, 65. [Google Scholar] [CrossRef]
- Pang, X.C.; Kang, D.; Fang, J.S.; Zhao, Y.; Xu, L.J.; Lian, W.W.; Liu, A.L.; Du, G.H. Network Pharmacology-Based Analysis of Chinese Herbal Naodesheng Formula for Application to Alzheimer’s Disease. Chin. J. Nat. Med. 2018, 16, 53–62. [Google Scholar] [CrossRef]
- Fang, J.; Li, Y.; Liu, R.; Pang, X.; Li, C.; Yang, R.; He, Y.; Lian, W.; Liu, A.-L.; Du, G.-H. Discovery of Multitarget-Directed Ligands against Alzheimer’s Disease through Systematic Prediction of Chemical–Protein Interactions. J. Chem. Inf. Model. 2015, 55, 149–164. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, A.; Younesi, E.; Sahadevan, S.; Zimmermann, J.; Hofmann-Apitius, M. Exploring Novel Mechanistic Insights in Alzheimer’s Disease by Assessing Reliability of Protein Interactions. Sci. Rep. 2015, 5, 13634. [Google Scholar] [CrossRef] [PubMed]
- Güner, O.F.; Bowen, J.P. Setting the Record Straight: The Origin of the Pharmacophore Concept. J. Chem. Inf. Model. 2014, 54, 1269–1283. [Google Scholar] [CrossRef]
- Seidel, T.; Schuetz, D.A.; Garon, A.; Langer, T. The Pharmacophore Concept and Its Applications in Computer-Aided Drug Design. In Progress in the Chemistry of Organic Natural Products; Springer: Cham, Switzerland, 2019; Volume 110, pp. 99–141. ISBN 9783030146313. [Google Scholar]
- Pradeepkiran, J.A.; Reddy, A.P.; Reddy, P.H. Pharmacophore-Based Models for Therapeutic Drugs against Phosphorylated Tau in Alzheimer’s Disease. Drug Discov. Today 2019, 24, 616–623. [Google Scholar] [CrossRef]
- Van Drie, J.H. Generation of Three-Dimensional Pharmacophore Models. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2013, 3, 449–464. [Google Scholar] [CrossRef]
- Seidel, T.; Wieder, O.; Garon, A.; Langer, T. Applications of the Pharmacophore Concept in Natural Product Inspired Drug Design. Mol. Inform. 2020, 39, 2000059. [Google Scholar] [CrossRef] [PubMed]
- Schuster, D.; Wolber, G. Identification of Bioactive Natural Products by Pharmacophore-Based Virtual Screening. Curr. Pharm. Des. 2010, 16, 1666–1681. [Google Scholar] [CrossRef] [PubMed]
- Dhanjal, J.K.; Sharma, S.; Grover, A.; Das, A. Use of Ligand-Based Pharmacophore Modeling and Docking Approach to Find Novel Acetylcholinesterase Inhibitors for Treating Alzheimer’s. Biomed. Pharmacother. 2015, 71, 146–152. [Google Scholar] [CrossRef] [PubMed]
- Goodford, P.J. A Computational Procedure for Determining Energetically Favorable Binding Sites on Biologically Important Macromolecules. J. Med. Chem. 1985, 28, 849–857. [Google Scholar] [CrossRef]
- Von Itzstein, M.; Wu, W.Y.; Kok, G.B.; Pegg, M.S.; Dyason, J.C.; Jin, B.; Van Phan, T.; Smythe, M.L.; White, H.F.; Oliver, S.W.; et al. Rational Design of Potent Sialidase-Based Inhibitors of Influenza Virus Replication. Nature 1993, 363, 418–423. [Google Scholar] [CrossRef] [PubMed]
- Mortier, J.; Dhakal, P.; Volkamer, A. Truly Target-Focused Pharmacophore Modeling: A Novel Tool for Mapping Intermolecular Surfaces. Molecules 2018, 23, 1959. [Google Scholar] [CrossRef]
- Wolber, G.; Dornhofer, A.A.; Langer, T. Efficient Overlay of Small Organic Molecules Using 3D Pharmacophores. J. Comput. Aided. Mol. Des. 2006, 20, 773–788. [Google Scholar] [CrossRef]
- Baroni, M.; Cruciani, G.; Sciabola, S.; Perruccio, F.; Mason, J.S. A Common Reference Framework for Analyzing/Comparing Proteins and Ligands. Fingerprints for Ligands and Proteins (FLAP): Theory and Application. J. Chem. Inf. Model. 2007, 47, 279–294. [Google Scholar] [CrossRef]
- Fabrizio, C.; Termine, A.; Caltagirone, C.; Sancesario, G. Artificial Intelligence for Alzheimer’s Disease: Promise or Challenge? Diagnostics 2021, 11, 1473. [Google Scholar] [CrossRef]
- Tsuji, S.; Hase, T.; Yachie-Kinoshita, A.; Nishino, T.; Ghosh, S.; Kikuchi, M.; Shimokawa, K.; Aburatani, H.; Kitano, H.; Tanaka, H. Artificial Intelligence-Based Computational Framework for Drug-Target Prioritization and Inference of Novel Repositionable Drugs for Alzheimer’s Disease. Alzheimers. Res. Ther. 2021, 13, 92. [Google Scholar] [CrossRef]
- Rodriguez, S.; Hug, C.; Todorov, P.; Moret, N.; Boswell, S.A.; Evans, K.; Zhou, G.; Johnson, N.T.; Hyman, B.T.; Sorger, P.K.; et al. Machine Learning Identifies Candidates for Drug Repurposing in Alzheimer’s Disease. Nat. Commun. 2021, 12, 1033. [Google Scholar] [CrossRef]
- Vamathevan, J.; Clark, D.; Czodrowski, P.; Dunham, I.; Ferran, E.; Lee, G.; Li, B.; Madabhushi, A.; Shah, P.; Spitzer, M.; et al. Applications of Machine Learning in Drug Discovery and Development. Nat. Rev. Drug Discov. 2019, 18, 463–477. [Google Scholar] [CrossRef]
- Ochoa, D.; Hercules, A.; Carmona, M.; Suveges, D.; Gonzalez-Uriarte, A.; Malangone, C.; Miranda, A.; Fumis, L.; Carvalho-Silva, D.; Spitzer, M.; et al. Open Targets Platform: Supporting Systematic Drug–Target Identification and Prioritisation. Nucleic Acids Res. 2021, 49, D1302–D1310. [Google Scholar] [CrossRef]
- Chen, X. TTD: Therapeutic Target Database. Nucleic Acids Res. 2002, 30, 412–415. [Google Scholar] [CrossRef]
- Saratxaga, C.L.; Moya, I.; Picón, A.; Acosta, M.; Moreno-Fernandez-de-Leceta, A.; Garrote, E.; Bereciartua-Perez, A. MRI Deep Learning-Based Solution for Alzheimer’s Disease Prediction. J. Pers. Med. 2021, 11, 902. [Google Scholar] [CrossRef]
- Basheer, S.; Bhatia, S.; Sakri, S.B. Computational Modeling of Dementia Prediction Using Deep Neural Network: Analysis on OASIS Dataset. IEEE Access 2021, 9, 42449–42462. [Google Scholar] [CrossRef]
- Sudharsan, M.; Thailambal, G. Alzheimer’s Disease Prediction Using Machine Learning Techniques and Principal Component Analysis (PCA). Mater. Today Proc. 2021. [Google Scholar] [CrossRef]
- Helaly, H.A.; Badawy, M.; Haikal, A.Y. Deep Learning Approach for Early Detection of Alzheimer’s Disease. Cognit. Comput. 2021, 24, 17–26. [Google Scholar] [CrossRef]
- Prajapati, R.; Khatri, U.; Kwon, G.R. An Efficient Deep Neural Network Binary Classifier for Alzheimer’s Disease Classification. In Proceedings of the 2021 International Conference on Artificial Intelligence in Information and Communication (ICAIIC), Jeju Island, Korea, 13–16 April 2021; IEEE: Piscataway, NJ, USA; pp. 231–234.
- Martinez-Murcia, F.J.; Ortiz, A.; Gorriz, J.M.; Ramirez, J.; Castillo-Barnes, D. Studying the Manifold Structure of Alzheimer’s Disease: A Deep Learning Approach Using Convolutional Autoencoders. IEEE J. Biomed. Heal. Inform. 2020, 24, 17–26. [Google Scholar] [CrossRef]
- Arabi, A.A. Artificial Intelligence in Drug Design: Algorithms, Applications, Challenges and Ethics. Futur. Drug Discov. 2021, 3, 2167–2180. [Google Scholar] [CrossRef]
- Yang, X.-G.; Lv, W.; Chen, Y.-Z.; Xue, Y. In Silico Prediction and Screening of γ-Secretase Inhibitors by Molecular Descriptors and Machine Learning Methods. J. Comput. Chem. 2009, 32, 1249–1258. [Google Scholar] [CrossRef]
- Silva-Spínola, A.; Baldeiras, I.; Arrais, J.P.; Santana, I. The Road to Personalized Medicine in Alzheimer’s Disease: The Use of Artificial Intelligence. Biomedicines 2022, 10, 315. [Google Scholar] [CrossRef] [PubMed]
- Sadeghi, S.; Lu, J.; Ngom, A. A Network-Based Drug Repurposing Method via Non-Negative Matrix Factorization. Bioinformatics 2022, 38, 1369–1377. [Google Scholar] [CrossRef]
- Kumar, S.; Chowdhury, S.; Kumar, S. In Silico Repurposing of Antipsychotic Drugs for Alzheimer’s Disease. BMC Neurosci. 2017, 18, 76. [Google Scholar] [CrossRef] [PubMed]
- Oprea, T.I.; Bauman, J.E.; Bologa, C.G.; Buranda, T.; Chigaev, A.; Edwards, B.S.; Jarvik, J.W.; Gresham, H.D.; Haynes, M.K.; Hjelle, B.; et al. Drug Repurposing from an Academic Perspective. Drug Discov. Today Ther. Strateg. 2011, 8, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Yu, R.; Chen, L.; Lan, R.; Shen, R.; Li, P. Computational Screening of Antagonists against the SARS-CoV-2 (COVID-19) Coronavirus by Molecular Docking. Int. J. Antimicrob. Agents 2020, 2, 106012. [Google Scholar] [CrossRef]
- Pan, X.; Lin, X.; Cao, D.; Zeng, X.; Yu, P.S.; He, L.; Nussinov, R.; Cheng, F. Deep Learning for Drug Repurposing: Methods, Databases, and Applications. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2022, 12, e1597. [Google Scholar] [CrossRef]
- Siavelis, J.C.; Bourdakou, M.M.; Athanasiadis, E.I.; Spyrou, G.M.; Nikita, K.S. Bioinformatics Methods in Drug Repurposing for Alzheimer’s Disease. Brief. Bioinform. 2016, 17, 322–335. [Google Scholar] [CrossRef]
- Clausznitzer, D.; Pichardo-Almarza, C.; Relo, A.L.; van Bergeijk, J.; van der Kam, E.; Laplanche, L.; Benson, N.; Nijsen, M. Quantitative Systems Pharmacology Model for Alzheimer Disease Indicates Targeting Sphingolipid Dysregulation as Potential Treatment Option. CPT Pharmacomet. Syst. Pharmacol. 2018, 7, 759–770. [Google Scholar] [CrossRef]
- Berger, S.I.; Iyengar, R. Network Analyses in Systems Pharmacology. Bioinformatics 2009, 25, 2466–2472. [Google Scholar] [CrossRef]
- Subramanian, N.; Torabi-Parizi, P.; Gottschalk, R.A.; Germain, R.N.; Dutta, B. Network Representations of Immune System Complexity. Wiley Interdiscip. Rev. Syst. Biol. Med. 2015, 7, 13–38. [Google Scholar] [CrossRef]
- Boran, A.D.W.; Iyengar, R. Systems Pharmacology. Mt. Sinai J. Med. A J. Transl. Pers. Med. 2010, 77, 333–344. [Google Scholar] [CrossRef]
- Nguyen, T.-P.; Priami, C.; Caberlotto, L. Novel Drug Target Identification for the Treatment of Dementia Using Multi-Relational Association Mining. Sci. Rep. 2015, 5, 11104. [Google Scholar] [CrossRef]
- Danhof, M. Systems Pharmacology-Towards the Modeling of Network Interactions. Eur. J. Pharm. Sci. 2016, 94, 4–14. [Google Scholar] [CrossRef] [Green Version]

| Software/Platform | Description | Link | Reference |
|---|---|---|---|
| AlzhCPI | With HTML and CSS technology that provides models and important fragments for MTDLs against AD | http://rcidm.org/AlzhCPI | [40] |
| AlzPlatform | AD-specific chemogenomics database based on ligands | http://www.cbligand.org/AD | [41] |
| HENA | Heterogeneous network-based dataset for Alzheimer’s disease | https://github.com/esugis/hena | [42] |
| NIAGADS | National Institute on Aging Genetics of Alzheimer’s Disease Data Storage Site | https://www.niagads.org/ | [43] |
| Treatment Type | Number of Associated Projects | Description |
|---|---|---|
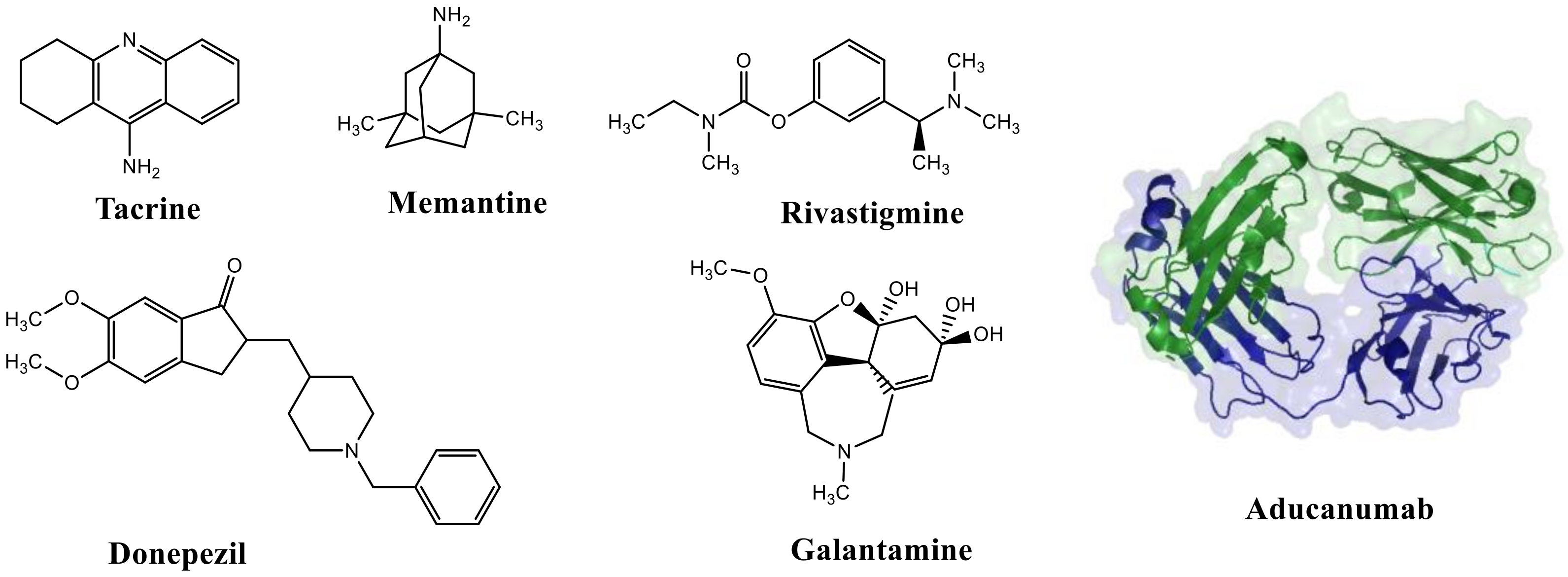
| Drug | 1353 | Analytical/experimental study. The patient is treated with different drugs. In the cases reported, 105 have used donepezil, 4 rivastigmine, and 4 galantamine, either in the absence of or in addition to other drugs and treatments. |
| Behavioral | 425 | Observational study. The patient undergoes therapies, lifestyle changes, sports, and cognitive activities to improve memory. It may or may not be accompanied by other types of therapies. Family therapy and psycho-emotional support are included. |
| Device | 263 | Interventional study where devices such as transcranial alternating current stimulation (tACS) and deep brain stimulation (DBS) are used to evaluate possible improvements in patient responses. |
| Procedure | 112 | The patient undergoes procedures such as yoga, hypnosis, surgery, or acupuncture. |
| Dietary supplement | 65 | New types of diets are implemented for the patient with specific supplements such as vitamin E, curcumin, and omega 3, among others. |
| Compound | Hybrid-Related | Biological Activity IC50 (µM) | Reference |
|---|---|---|---|
 | Carbazole-curcumin | AChE: 6.9 ± 0.9 BuChE: 2.8 ± 0.4 | [101] |
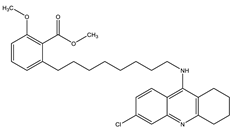 | Tacrine–Anacardic acid | AChE: 2.54 ± 0.07 BuChE: 0.265 ± 0.027 | [27] |
 | Rivastigmine | AchE at 1 µM = 24.43% BuChe at 1 µM = 72.30% At 10 µM 2.2% of Aβ self aggregation | [102] |
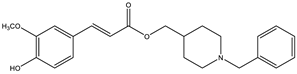 | Donepezil–curcumin | AChE: 0.46 BuChE: 24.97 | [103] |
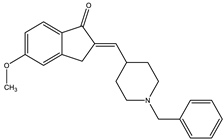 | Donepezil | AChE: 0.029 BACE1: 0.33 | [104] |
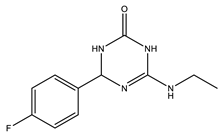 | Cyclic amide group | BACE1: 16.0 GSK-3: 7.1 | [105] |
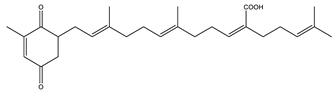 | Sargaquinoic-acid | AChE: 69.3 BuChE: 10.5 BACE1: 12.1 | [106] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arrué, L.; Cigna-Méndez, A.; Barbosa, T.; Borrego-Muñoz, P.; Struve-Villalobos, S.; Oviedo, V.; Martínez-García, C.; Sepúlveda-Lara, A.; Millán, N.; Márquez Montesinos, J.C.E.; et al. New Drug Design Avenues Targeting Alzheimer’s Disease by Pharmacoinformatics-Aided Tools. Pharmaceutics 2022, 14, 1914. https://doi.org/10.3390/pharmaceutics14091914
Arrué L, Cigna-Méndez A, Barbosa T, Borrego-Muñoz P, Struve-Villalobos S, Oviedo V, Martínez-García C, Sepúlveda-Lara A, Millán N, Márquez Montesinos JCE, et al. New Drug Design Avenues Targeting Alzheimer’s Disease by Pharmacoinformatics-Aided Tools. Pharmaceutics. 2022; 14(9):1914. https://doi.org/10.3390/pharmaceutics14091914
Chicago/Turabian StyleArrué, Lily, Alexandra Cigna-Méndez, Tábata Barbosa, Paola Borrego-Muñoz, Silvia Struve-Villalobos, Victoria Oviedo, Claudia Martínez-García, Alexis Sepúlveda-Lara, Natalia Millán, José C. E. Márquez Montesinos, and et al. 2022. "New Drug Design Avenues Targeting Alzheimer’s Disease by Pharmacoinformatics-Aided Tools" Pharmaceutics 14, no. 9: 1914. https://doi.org/10.3390/pharmaceutics14091914
APA StyleArrué, L., Cigna-Méndez, A., Barbosa, T., Borrego-Muñoz, P., Struve-Villalobos, S., Oviedo, V., Martínez-García, C., Sepúlveda-Lara, A., Millán, N., Márquez Montesinos, J. C. E., Muñoz, J., Santana, P. A., Peña-Varas, C., Barreto, G. E., González, J., & Ramírez, D. (2022). New Drug Design Avenues Targeting Alzheimer’s Disease by Pharmacoinformatics-Aided Tools. Pharmaceutics, 14(9), 1914. https://doi.org/10.3390/pharmaceutics14091914










