Unequal Behaviour between Hydrolysable Functions of Nirmatrelvir under Stress Conditions: Structural and Theoretical Approaches in Support of Preformulation Studies
Abstract
:1. Introduction
2. Materials and Methods
2.1. Reagents and Sample Preparation
2.2. Instrumental
2.3. Computational Conditions
3. Results
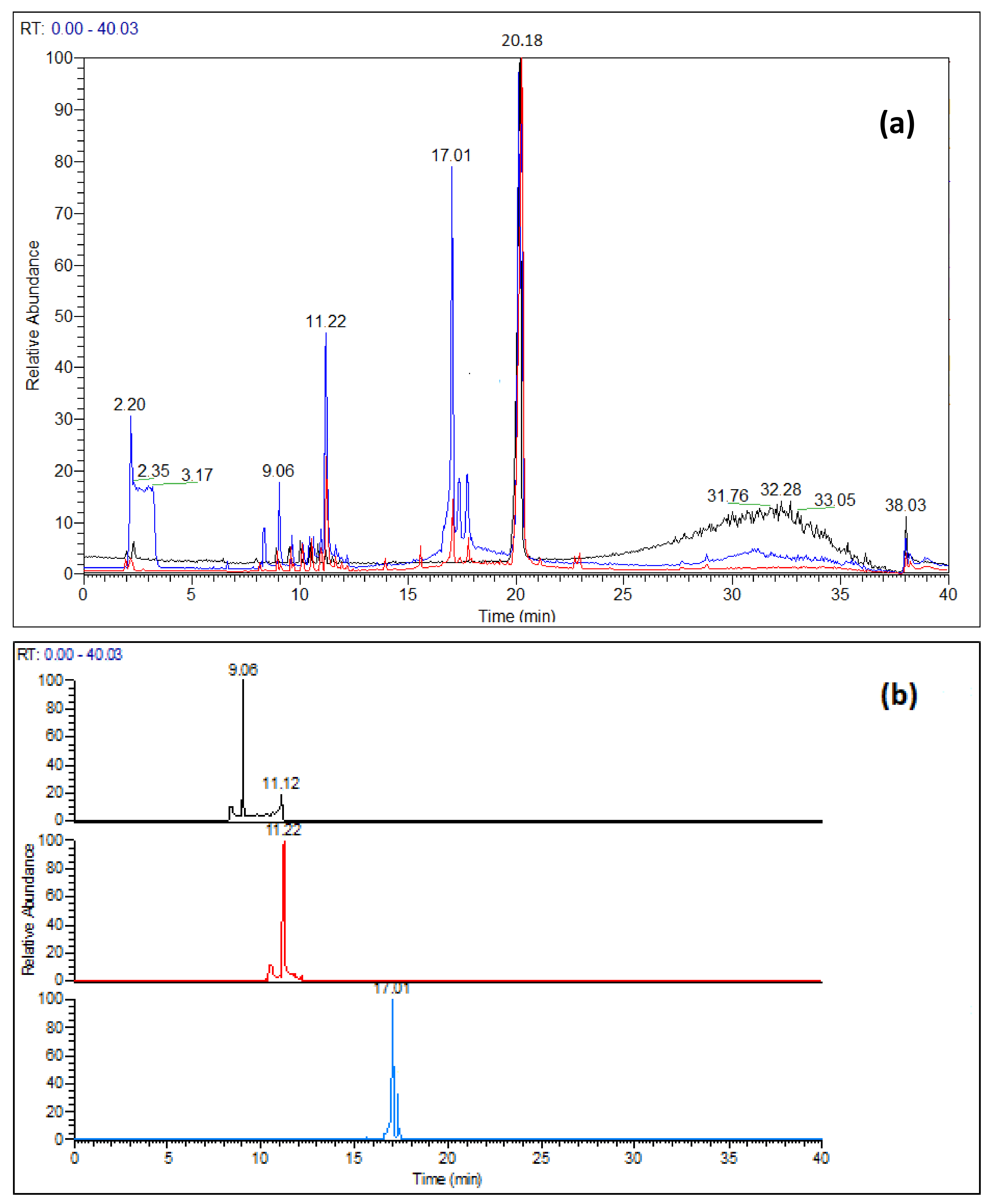
3.1. Degradation Products Detected under Hydrolytic Conditions
3.2. Structural Characterization of the DPs
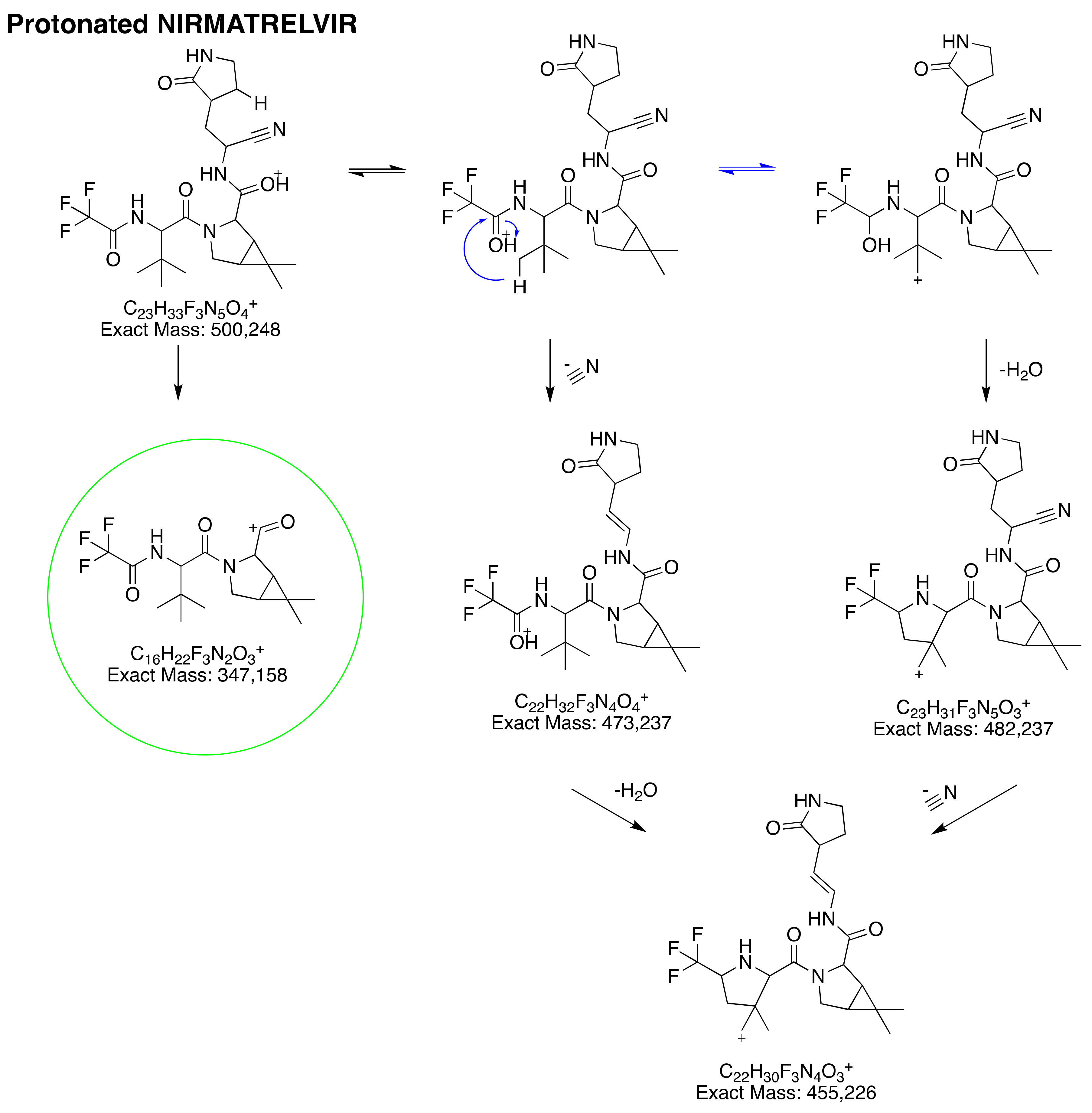
3.2.1. Protonated Nirmatrelvir’s Fragmentation Pattern
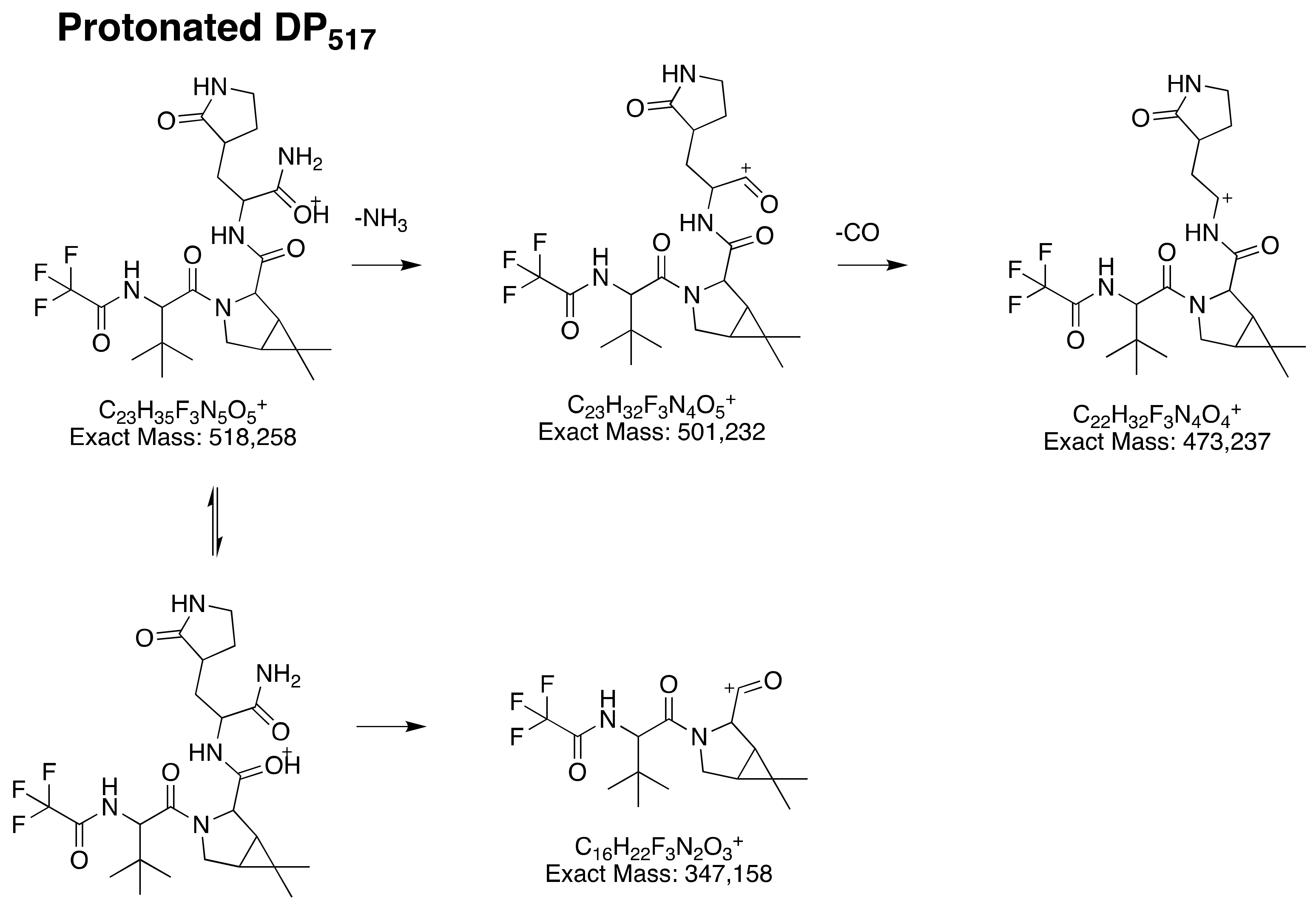
3.2.2. Structural Elucidation of DP517
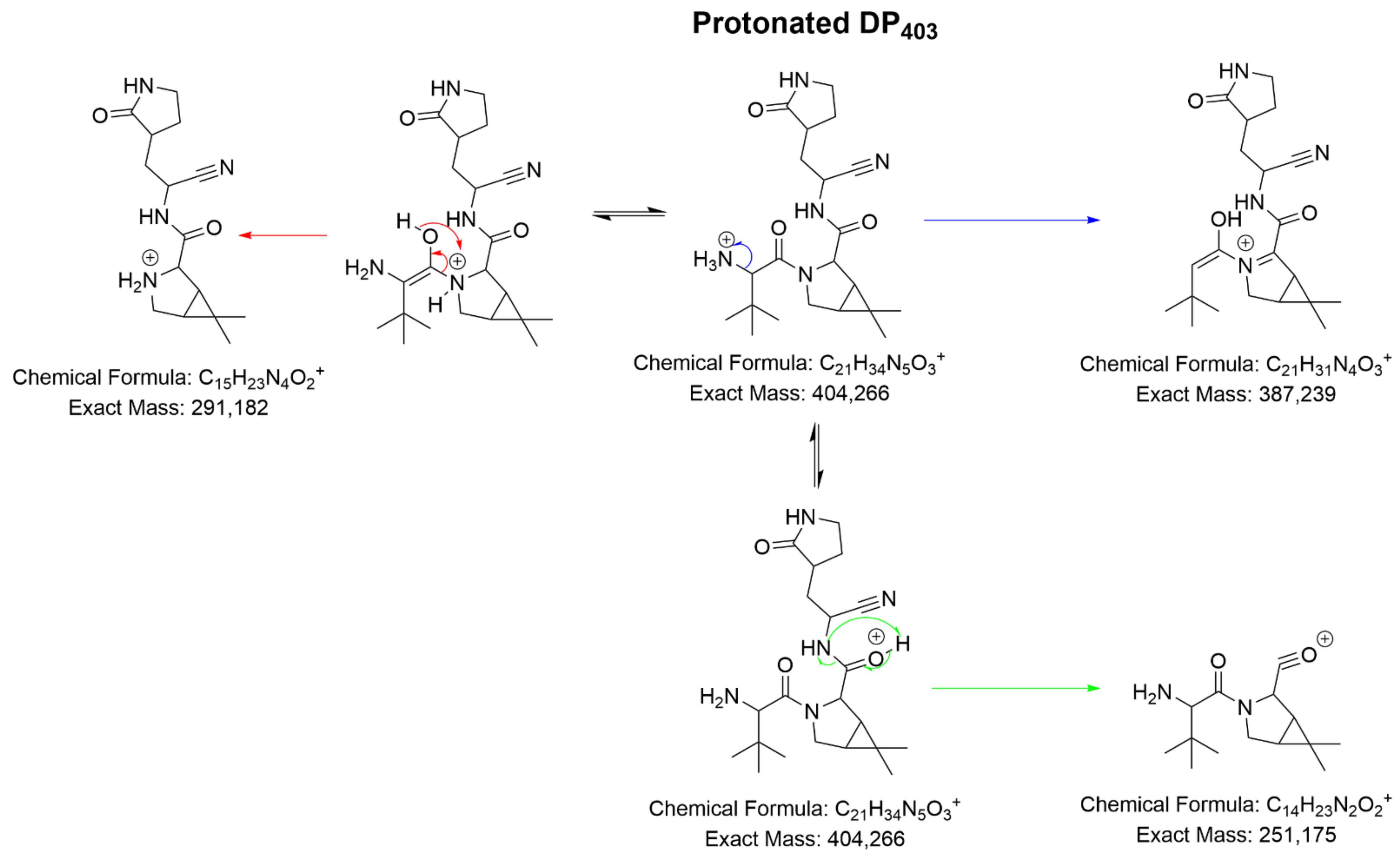
3.2.3. Structural Elucidation of DP403
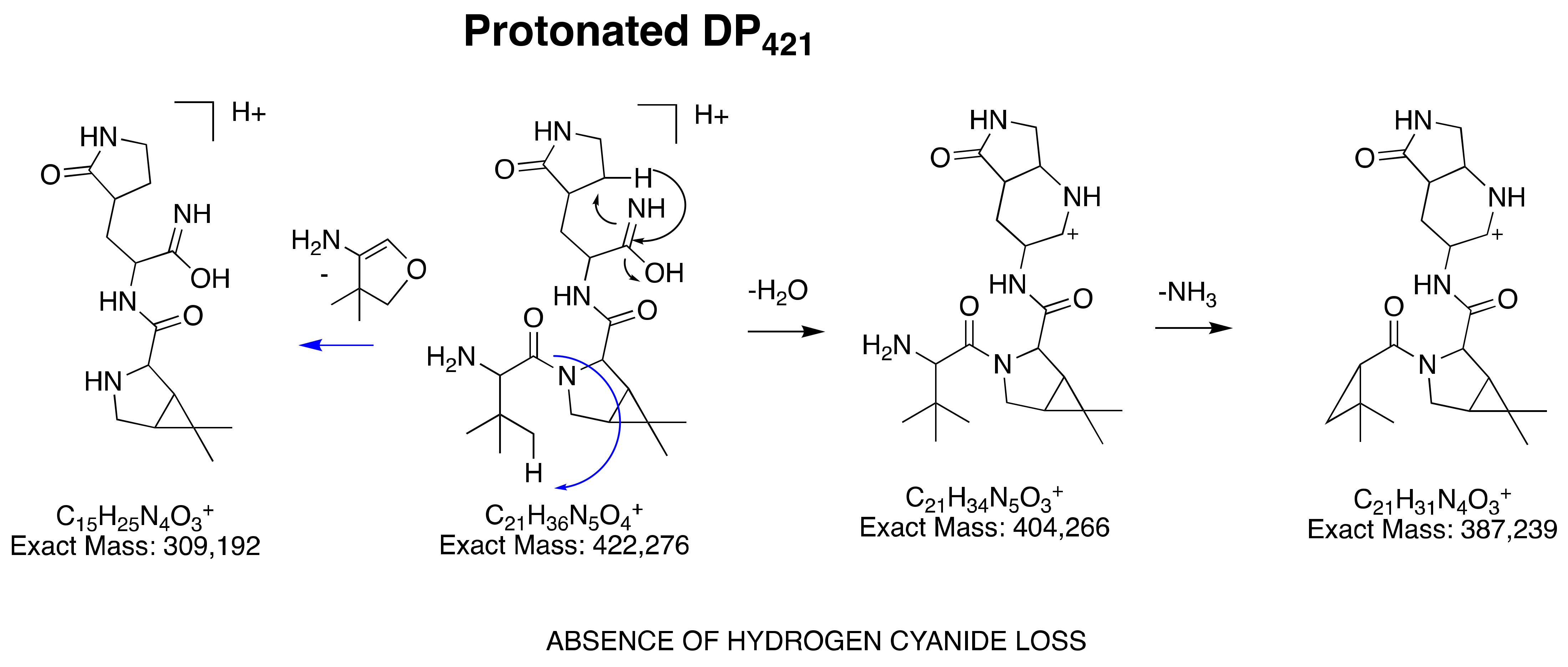
3.2.4. Structural Elucidation of DP421
3.3. Main Degradation Process of Nirmatrelvir Characterized by Mass Spectrometry
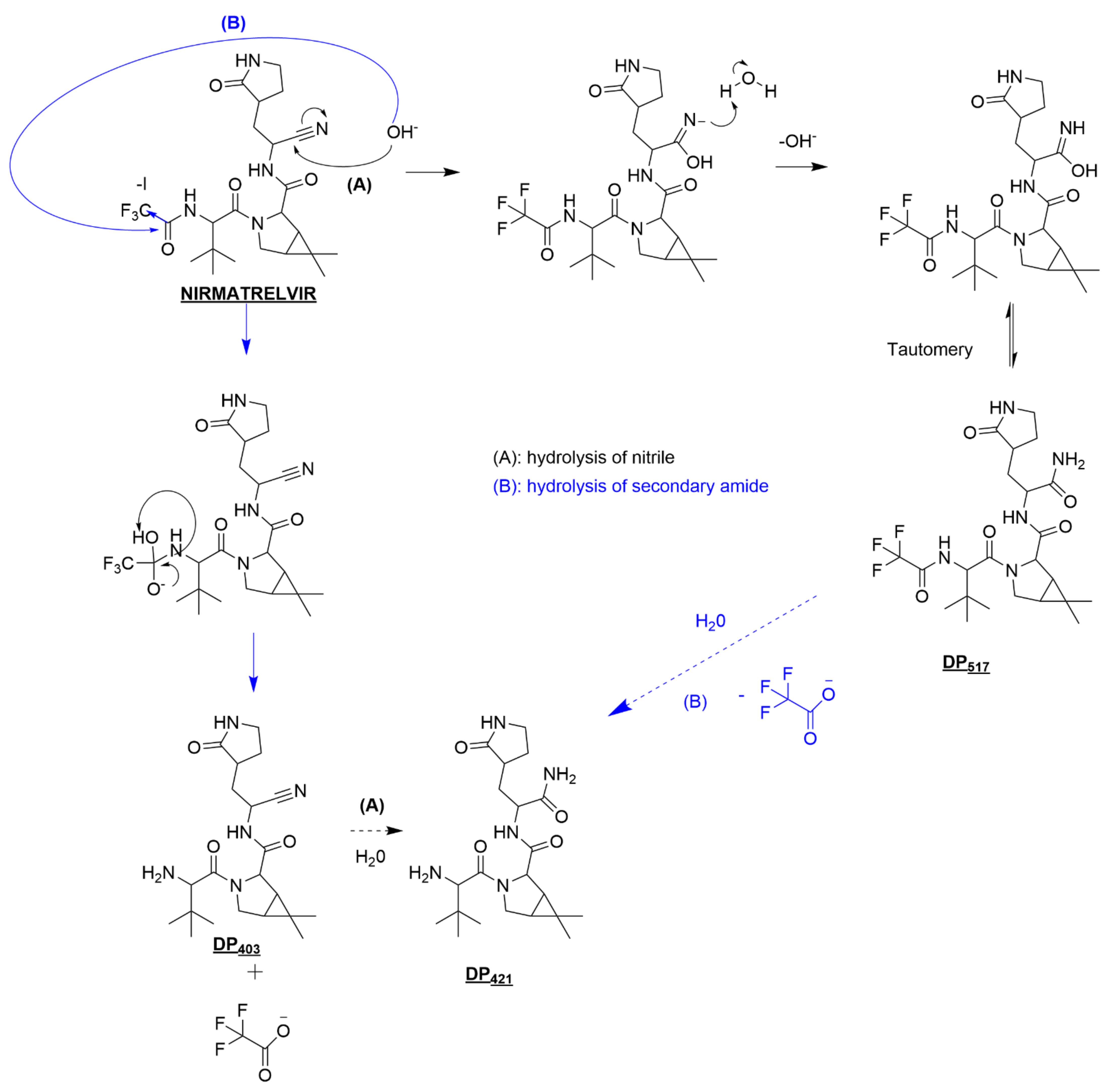
3.3.1. Nirmatrelvir Degradation under Basic Conditions
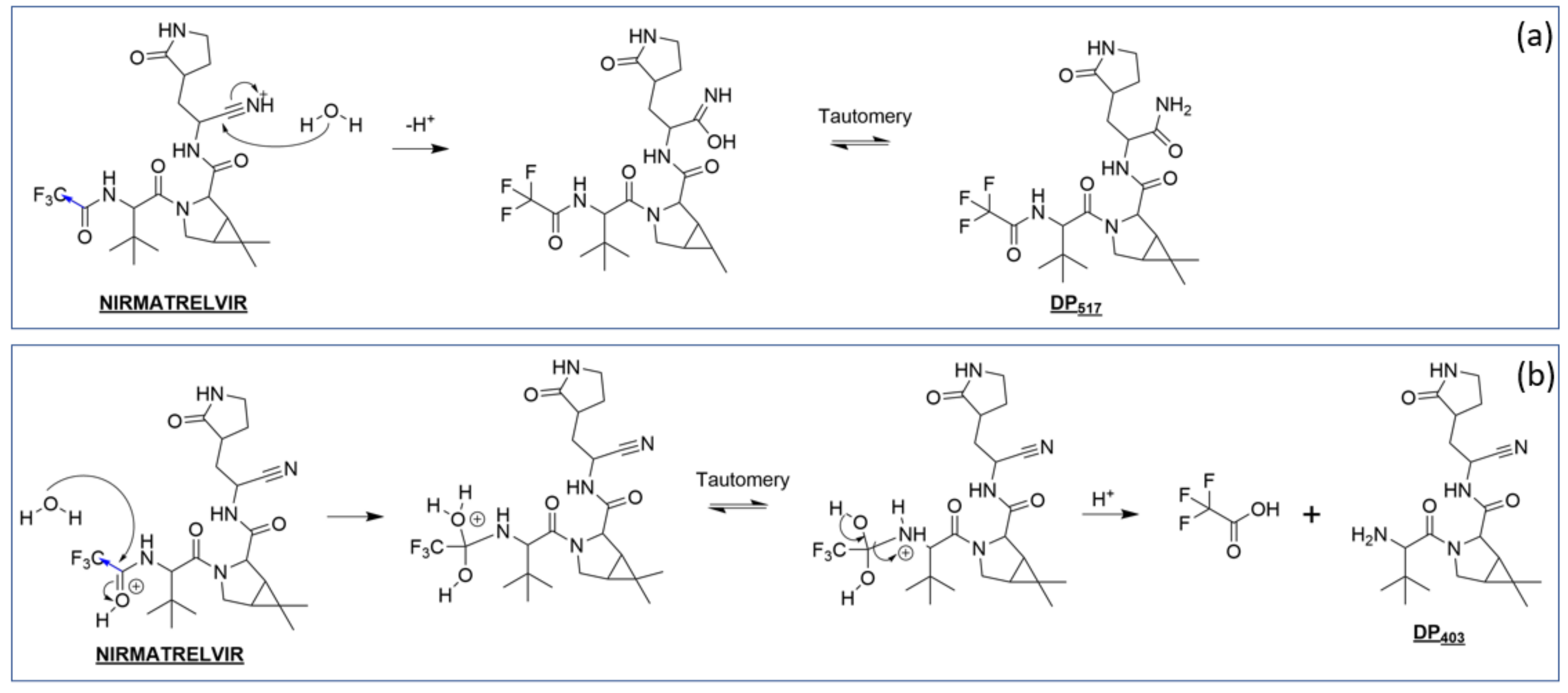
3.3.2. Nirmatrelvir Degradation under Acidic Conditions
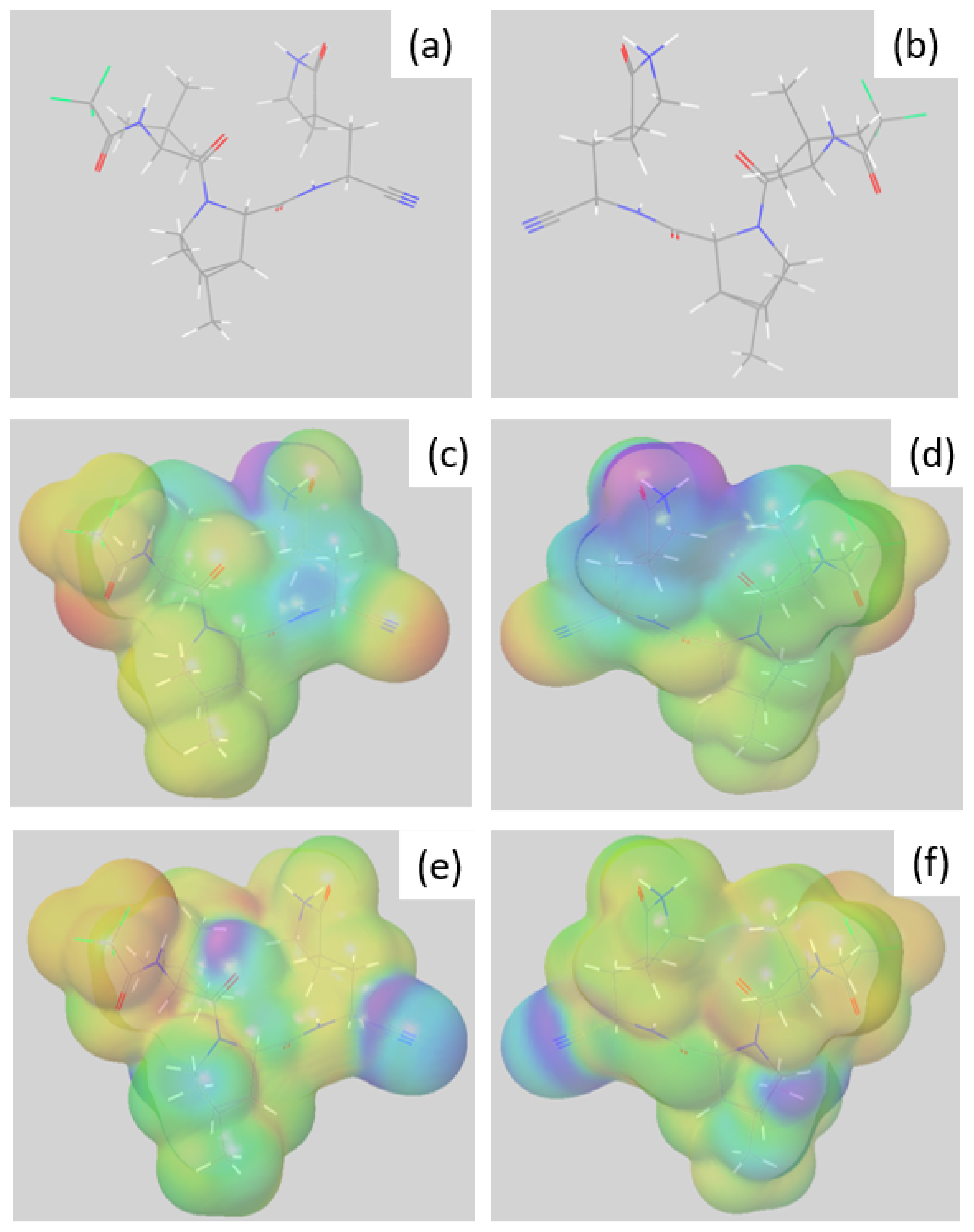
3.4. Preferential Hydrolysis Sites of Nimatrelvir, an Investigation by DFT
3.4.1. Reactivity of Nirmatrelvir to Acidic pH
3.4.2. Reactivity of Nirmatrelvir to Basic pH
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Saravolatz, L.D.; Depcinski, S.; Sharma, M. Molnupiravir and Nirmatrelvir-Ritonavir: Oral COVID Antiviral Drugs. Clin. Infect. Dis. 2022, ciac180. [Google Scholar] [CrossRef] [PubMed]
- Menéndez, J.C. Approaches to the Potential Therapy of COVID-19: A General Overview from the Medicinal Chemistry Perspective. Molecules 2022, 27, 658. [Google Scholar] [CrossRef] [PubMed]
- Blessy, M.; Patel, R.D.; Prajapati, P.N.; Agrawal, Y.K. Development of Forced Degradation and Stability Indicating Studies of Drugs—A Review. J. Pharm. Anal. 2014, 4, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Baertschi, S.W.; Alsante, K.M.; Reed, R.A. (Eds.) Stress Testing: A Predictive Tool. In Pharmaceutical Stress Testing; CRC Press: Boca Raton, FL, USA, 2016; pp. 22–60. ISBN 978-0-429-13608-5. [Google Scholar]
- Bakowies, D.; Kollman, P.A. Theoretical Study of Base-Catalyzed Amide Hydrolysis: Gas- and Aqueous-Phase Hydrolysis of Formamide. J. Am. Chem. Soc. 1999, 121, 5712–5726. [Google Scholar] [CrossRef]
- Zahn, D. Theoretical Study of the Mechanisms of Acid-Catalyzed Amide Hydrolysis in Aqueous Solution. J. Phys. Chem. B 2003, 107, 12303–12306. [Google Scholar] [CrossRef]
- Mitchell, S.M.; Ullman, J.L.; Teel, A.L.; Watts, R.J. PH and Temperature Effects on the Hydrolysis of Three β-Lactam Antibiotics: Ampicillin, Cefalotin and Cefoxitin. Sci. Total Environ. 2014, 466, 547–555. [Google Scholar] [CrossRef] [PubMed]
- Secrétan, P.-H.; Sadou-Yayé, H.; Aymes-Chodur, C.; Bernard, M.; Solgadi, A.; Amrani, F.; Yagoubi, N.; Do, B. A Comprehensive Study of Apixaban’s Degradation Pathways under Stress Conditions Using Liquid Chromatography Coupled to Multistage Mass Spectrometry. RSC Adv. 2015, 5, 35586–35597. [Google Scholar] [CrossRef]
- Mary, Y.S.; Mary, Y.S.; Bielenica, A.; Armaković, S.; Armaković, S.J.; Chandramohan, V.; Dammalli, M. Investigation of the Reactivity Properties of a Thiourea Derivative with Anticancer Activity by DFT and MD Simulations. J. Mol. Model. 2021, 27, 217. [Google Scholar] [CrossRef] [PubMed]
- Sureshkumar, B.; Mary, Y.S.; Panicker, C.Y.; Suma, S.; Armaković, S.; Armaković, S.J.; Van Alsenoy, C.; Narayana, B. Quinoline Derivatives as Possible Lead Compounds for Anti-Malarial Drugs: Spectroscopic, DFT and MD Study. Arab. J. Chem. 2020, 13, 632–648. [Google Scholar] [CrossRef]
- Bochevarov, A.D.; Harder, E.; Hughes, T.F.; Greenwood, J.R.; Braden, D.A.; Philipp, D.M.; Rinaldo, D.; Halls, M.D.; Zhang, J.; Friesner, R.A. Jaguar: A High-performance Quantum Chemistry Software Program with Strengths in Life and Materials Sciences. Int. J. Quantum Chem. 2013, 113, 2110–2142. [Google Scholar] [CrossRef]
- Theilacker, K.; Arbuznikov, A.V.; Bahmann, H.; Kaupp, M. Evaluation of a Combination of Local Hybrid Functionals with DFT-D3 Corrections for the Calculation of Thermochemical and Kinetic Data. J. Phys. Chem. A 2011, 115, 8990–8996. [Google Scholar] [CrossRef] [PubMed]
- Khemalapure, S.S.; Katti, V.S.; Hiremath, C.S.; Basanagouda, M.; Hiremath, S.M.; Armaković, S.J.; Armaković, S. Molecular Structure, Optoelectronic Properties, Spectroscopic (FT-IR, FT-Raman and UV–Vis), H-BDE, NBO and Drug Likeness Investigations on 7, 8-Benzocoumarin-4-Acetic Acid (7BAA). J. Mol. Struct. 2019, 1195, 815–826. [Google Scholar] [CrossRef]
- Li, Y.; Evans, J.N.S. The Fukui Function: A Key Concept Linking Frontier Molecular Orbital Theory and the Hard-Soft-Acid-Base Principle. J. Am. Chem. Soc. 1995, 117, 7756–7759. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Secretan, P.-H.; Annereau, M.; Kini-Matondo, W.; Prost, B.; Prudhomme, J.; Bournane, L.; Paul, M.; Yagoubi, N.; Sadou-Yayé, H.; Do, B. Unequal Behaviour between Hydrolysable Functions of Nirmatrelvir under Stress Conditions: Structural and Theoretical Approaches in Support of Preformulation Studies. Pharmaceutics 2022, 14, 1720. https://doi.org/10.3390/pharmaceutics14081720
Secretan P-H, Annereau M, Kini-Matondo W, Prost B, Prudhomme J, Bournane L, Paul M, Yagoubi N, Sadou-Yayé H, Do B. Unequal Behaviour between Hydrolysable Functions of Nirmatrelvir under Stress Conditions: Structural and Theoretical Approaches in Support of Preformulation Studies. Pharmaceutics. 2022; 14(8):1720. https://doi.org/10.3390/pharmaceutics14081720
Chicago/Turabian StyleSecretan, Philippe-Henri, Maxime Annereau, Willy Kini-Matondo, Bastien Prost, Jade Prudhomme, Lina Bournane, Muriel Paul, Najet Yagoubi, Hassane Sadou-Yayé, and Bernard Do. 2022. "Unequal Behaviour between Hydrolysable Functions of Nirmatrelvir under Stress Conditions: Structural and Theoretical Approaches in Support of Preformulation Studies" Pharmaceutics 14, no. 8: 1720. https://doi.org/10.3390/pharmaceutics14081720
APA StyleSecretan, P.-H., Annereau, M., Kini-Matondo, W., Prost, B., Prudhomme, J., Bournane, L., Paul, M., Yagoubi, N., Sadou-Yayé, H., & Do, B. (2022). Unequal Behaviour between Hydrolysable Functions of Nirmatrelvir under Stress Conditions: Structural and Theoretical Approaches in Support of Preformulation Studies. Pharmaceutics, 14(8), 1720. https://doi.org/10.3390/pharmaceutics14081720





