Insect Cells for High-Yield Production of SARS-CoV-2 Spike Protein: Building a Virosome-Based COVID-19 Vaccine Candidate
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Lines and Culture Media
2.2. Recombinant Baculovirus
2.2.1. Expression Vectors
2.2.2. Baculovirus Generation
2.3. Protein Production and Purification
2.4. Virosomes Preparation
2.5. Analytics
2.5.1. Cell Concentration and Viability
2.5.2. Baculovirus Titration
2.5.3. Protein Concentration
2.5.4. Western Blot
2.5.5. Differential Scanning Fluorimetry
2.5.6. HPLC-SEC
2.5.7. Site-Specific Glycosylation Analysis
2.5.8. ELISA
2.6. Statistical Analysis
3. Results
3.1. SARS-CoV-2 Spike Protein Production in Insect Cells
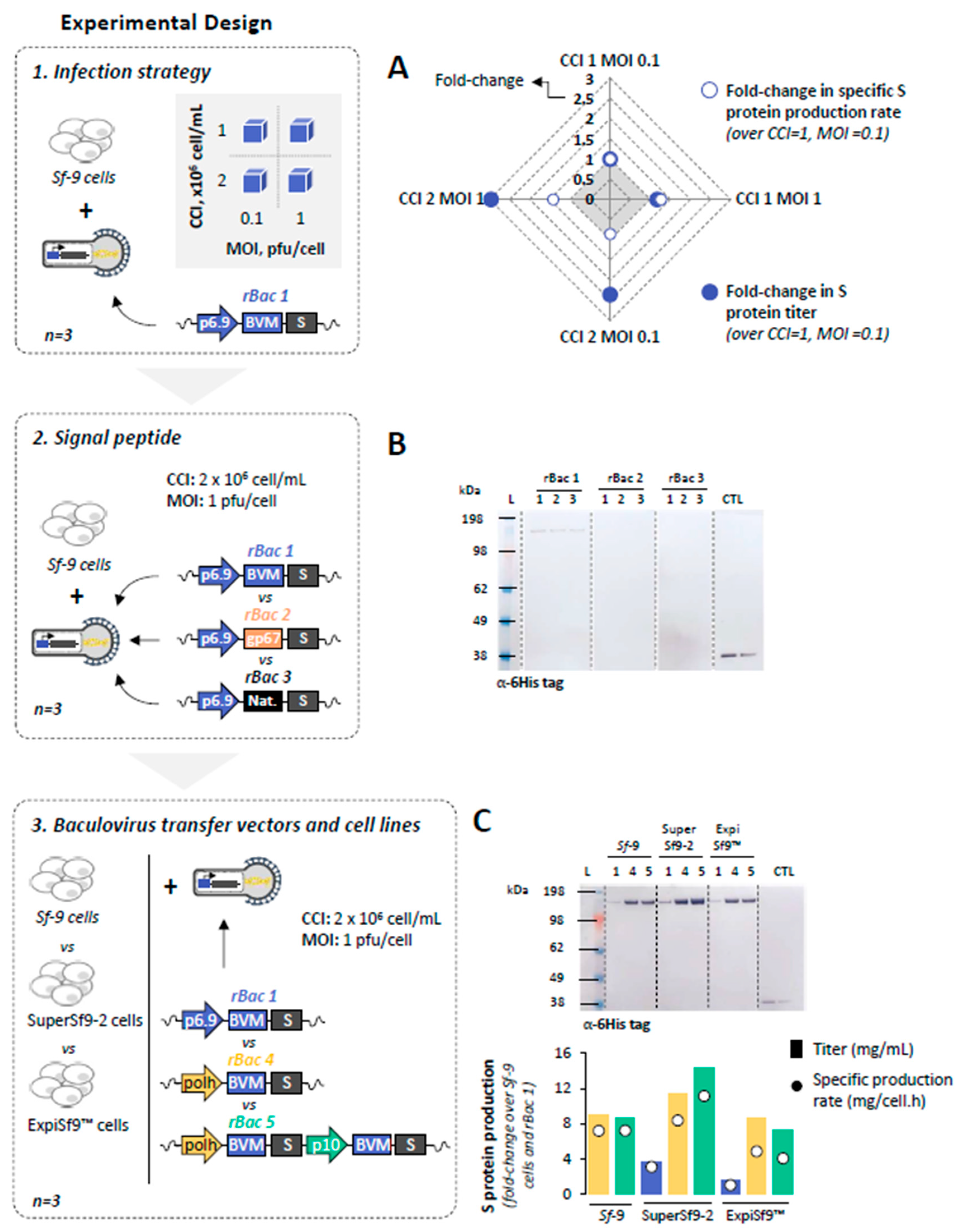
3.1.1. Infection Strategy
3.1.2. Signal Peptide
3.1.3. Baculovirus Transfer Vectors and Cell Lines
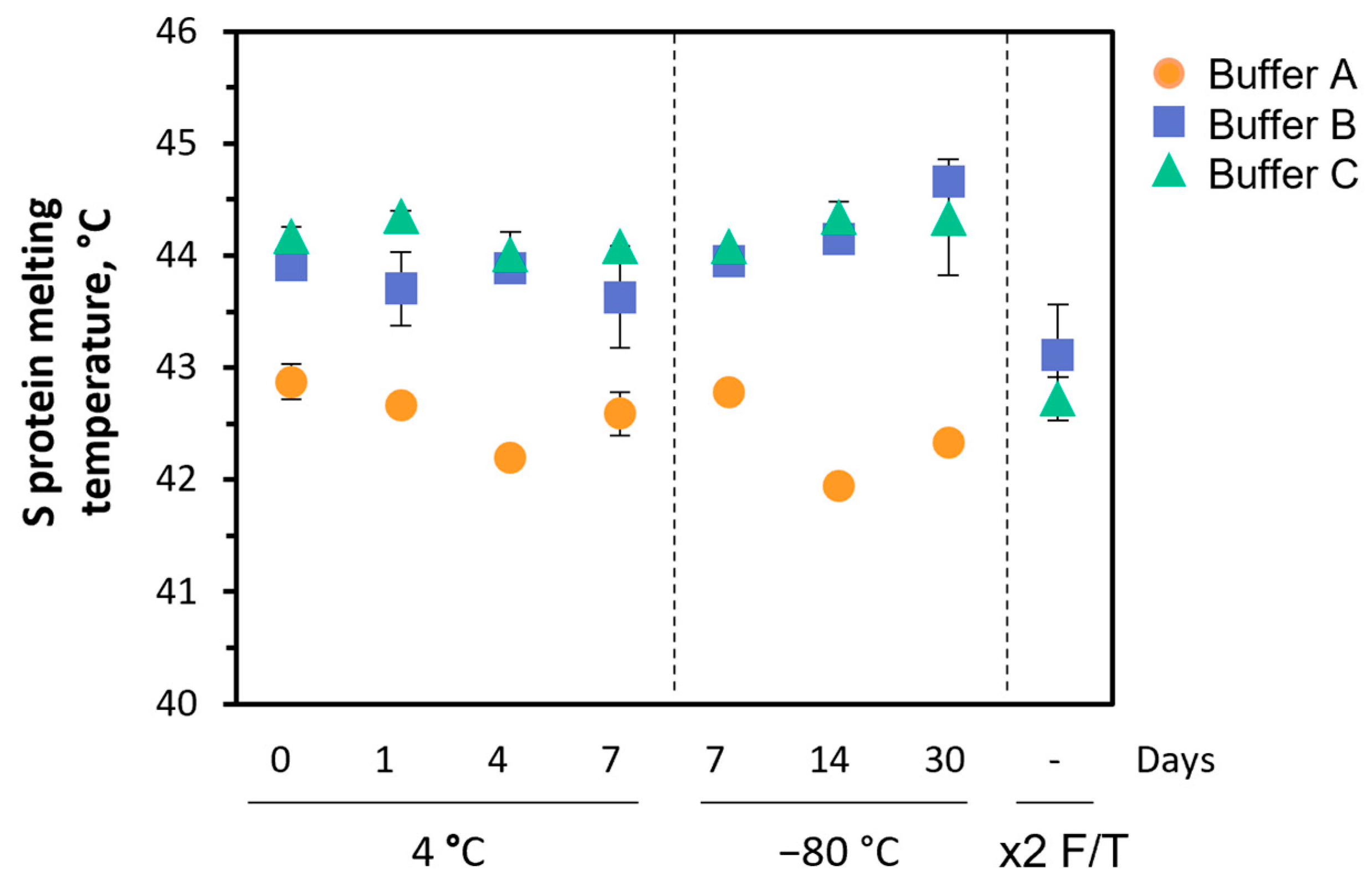
3.1.4. Formulation Buffer
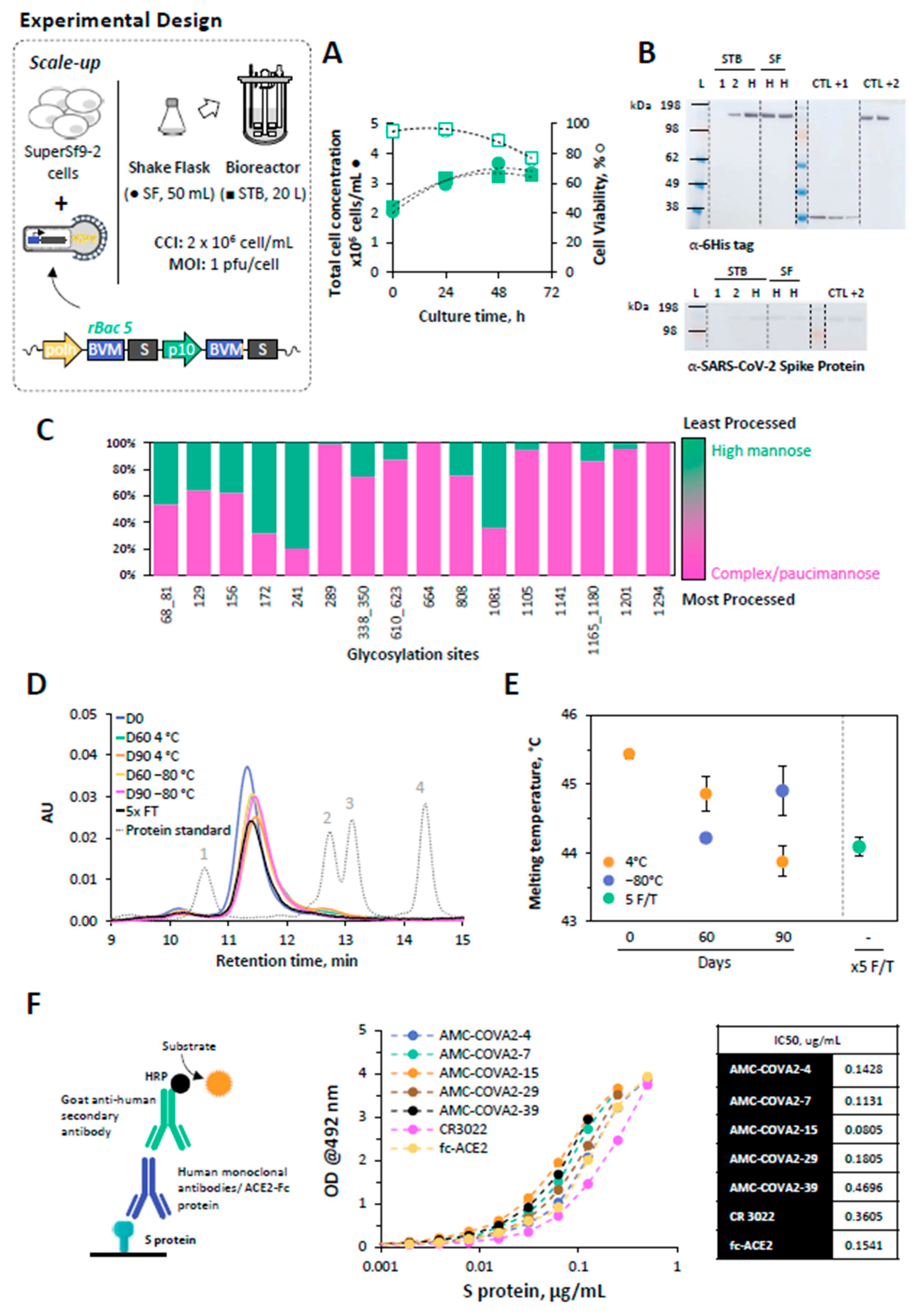
3.2. Scale-Up SARS-CoV-2 Spike Protein Production
3.2.1. Glycosylation Pattern of S Protein
3.2.2. Mid-Term Storage Stability of S Protein
3.2.3. Epitope Mapping of S Protein
3.3. Conjugation of SARS-CoV-2 Spike Protein to Virosomes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Coronavirus Graphs: Worldwide Cases and Deaths—Worldometer. Available online: https://www.worldometers.info/coronavirus/worldwide-graphs/#total-deaths (accessed on 11 August 2021).
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhao, S.; Ou, J.; Zhang, J.; Lan, W.; Guan, W.; Wu, X.; Yan, Y.; Zhao, W.; Wu, J.; et al. COVID-19: Coronavirus Vaccine Development Updates. Front. Immunol. 2020, 11, 602256. [Google Scholar] [CrossRef] [PubMed]
- Mahase, E. COVID-19: Novavax vaccine efficacy is 86% against UK variant and 60% against South African variant. BMJ 2021, 372, n296. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Interim Recommendations for Use of the Novavax NVX-CoV2373 Vaccine against COVID-19: Interim Guidance, 20 December 2021. Available online: https://apps.who.int/iris/handle/10665/350881 (accessed on 11 August 2021).
- Bovier, P.A. Epaxal: A virosomal vaccine to prevent hepatitis A infection. Expert Rev. Vaccines 2008, 7, 1141–1150. [Google Scholar] [CrossRef]
- Herzog, C.; Hartmann, K.; Künzi, V.; Kürsteiner, O.; Mischler, R.; Lazar, H.; Glück, R. Eleven years of Inflexal® V—A virosomal adjuvanted influenza vaccine. Vaccine 2009, 27, 4381–4387. [Google Scholar] [CrossRef] [PubMed]
- Moser, C.; Amacker, M.; Kammer, A.R.; Rasi, S.; Westerfeld, N.; Zurbriggen, R. Influenza virosomes as a combined vaccine carrier and adjuvant system for prophylactic and therapeutic immunizations. Expert Rev. Vaccines 2007, 6, 711–721. [Google Scholar] [CrossRef]
- Moser, C.; Müller, M.; Kaeser, M.D.; Weydemann, U.; Amacker, M. Influenza virosomes as vaccine adjuvant and carrier system. Expert Rev. Vaccines 2013, 12, 779–791. [Google Scholar] [CrossRef]
- Tamborrini, M.; Hauser, J.; Schäfer, A.; Amacker, M.; Favuzza, P.; Kyungtak, K.; Fleury, S.; Pluschke, G. Vaccination with virosomally formulated recombinant CyRPA elicits protective antibodies against Plasmodium falciparum parasites in preclinical in vitro and in vivo models. NPJ Vaccines 2020, 5, 9. [Google Scholar] [CrossRef] [Green Version]
- Van der Velden, Y.U.; Grobben, M.; Caniels, T.G.; Burger, J.A.; Poniman, M.; Oomen, M.; Rijnstra, E.S.; Tejjani, K.; Guerra, D.; Kempers, R.; et al. A SARS-CoV-2 Wuhan spike virosome vaccine induces superior neutralization breadth compared to one using the Beta spike. Sci. Rep. 2022, 12, 3884. [Google Scholar] [CrossRef]
- Amanat, F.; Stadlbauer, D.; Strohmeier, S.; Nguyen, T.H.O.; Chromikova, V.; McMahon, M.; Jiang, K.; Arunkumar, G.A.; Jurczyszak, D.; Polanco, J.; et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 2020, 26, 1033–1036. [Google Scholar] [CrossRef]
- Letko, M.; Marzi, A.; Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microbiol. 2020, 5, 562–569. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bangaru, S.; Ozorowski, G.; Turner, H.L.; Antanasijevic, A.; Huang, D.; Wang, X.; Torres, J.L.; Diedrich, J.K.; Tian, J.-H.; Portnoff, A.D.; et al. Structural analysis of full-length SARS-CoV-2 spike protein from an advanced vaccine candidate. Science 2020, 370, 1089–1094. [Google Scholar] [CrossRef] [PubMed]
- Reis, C.A.; Tauber, R.; Blanchard, V. Glycosylation is a key in SARS-CoV-2 infection. J. Mol. 2021, 99, 1023–1031. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, Y.; Allen, J.D.; Wrapp, D.; McLellan, J.S.; Crispin, M. Site-specific glycan analysis of the SARS-CoV-2 spike. Science 2020, 369, 330–333. [Google Scholar] [CrossRef]
- Stuible, M.; Gervais, C.; Lord-Dufour, S.; Perret, S.; L’Abbé, D.; Schrag, J.; St-Laurent, G.; Durocher, Y. Rapid, high-yield production of full-length SARS-CoV-2 spike ectodomain by transient gene expression in CHO cells. J. Biotechnol. 2021, 326, 21–27. [Google Scholar] [CrossRef]
- Buckland, B.; Boulanger, R.; Fino, M.; Srivastava, I.; Holtz, K.; Khramtsov, N.; McPherson, C.; Meghrous, J.; Kubera, P.; Cox, M.M.J. Technology transfer and scale-up of the Flublok® recombinant hemagglutinin (HA) influenza vaccine manufacturing process. Vaccine 2014, 32, 5496–5502. [Google Scholar] [CrossRef]
- Van Oers, M.M.; Pijlman, G.P.; Vlak, J.M. Thirty years of baculovirus–insect cell protein expression: From dark horse to mainstream technology. J. Gen. Virol. 2015, 96, 6–23. [Google Scholar] [CrossRef]
- Shi, X.; Jarvis, D.L. Protein N-glycosylation in the baculovirus-insect cell system. Curr. Drug Targets 2007, 8, 1116–1125. [Google Scholar] [CrossRef] [Green Version]
- Van Oosten, L.; Altenburg, J.J.; Fougeroux, C.; Geertsema, C.; van den End, F.; Evers, W.A.C.; Westphal, A.H.; Lindhoud, S.; van den Berg, W.; Swarts, D.C.; et al. Two-Component Nanoparticle Vaccine Displaying Glycosylated Spike S1 Domain Induces Neutralizing Antibody Response against SARS-CoV-2 Variants. MBio 2021, 12, e01813-21. [Google Scholar] [CrossRef]
- Vieira, H.L.A.; Estêvão, C.; Roldão, A.; Peixoto, C.C.; Sousa, M.F.Q.; Cruz, P.E.; Carrondo, M.J.T.; Alves, P.M. Triple layered rotavirus VLP production: Kinetics of vector replication, mRNA stability and recombinant protein production. J. Biotechnol. 2005, 120, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Pais, D.A.M.; Portela, R.M.C.; Carrondo, M.J.T.; Isidro, I.A.; Alves, P.M. Enabling PAT in insect cell bioprocesses: In situ monitoring of recombinant adeno-associated virus production by fluorescence spectroscopy. Biotechnol. Bioeng. 2019, 116, 2803–2814. [Google Scholar] [CrossRef] [PubMed]
- Amacker, M.; Smardon, C.; Mason, L.; Sorrell, J.; Jeffery, K.; Adler, M.; Bhoelan, F.; Belova, O.; Spengler, M.; Punnamoottil, B.; et al. New GMP manufacturing processes to obtain thermostable HIV-1 gp41 virosomes under solid forms for various mucosal vaccination routes. NPJ Vaccines 2020, 5, 41. [Google Scholar] [CrossRef] [PubMed]
- Tennant, J.R. Evaluation of the trypan blue technique for determination of cell viability. Transplantation 1964, 2, 685–694. [Google Scholar] [CrossRef] [PubMed]
- Mena, J.A.; Ramírez, O.T.; Palomares, L.A. Titration of Non-Occluded Baculovirus Using a Cell Viability Assay. BioTechniques 2003, 34, 260–264. [Google Scholar] [CrossRef] [PubMed]
- Roldão, A.; Oliveira, R.; Carrondo, M.J.T.; Alves, P.M. Error assessment in recombinant baculovirus titration: Evaluation of different methods. J. Virol. Methods 2009, 159, 69–80. [Google Scholar] [CrossRef]
- Correia, R.; Fernandes, B.; Alves, P.M.; Carrondo, M.J.T.; Roldão, A. Improving Influenza HA-Vlps Production in Insect High Five Cells via Adaptive Laboratory Evolution. Vaccines 2020, 8, 589. [Google Scholar] [CrossRef]
- Castro, R.; Nobre, L.S.; Eleutério, R.P.; Thomaz, M.; Pires, A.; Monteiro, S.M.; Mendes, S.; Gomes, R.A.; Clemente, J.J.; Sousa, M.F.Q.; et al. Production of high-quality SARS-CoV-2 antigens: Impact of bioprocess and storage on glycosylation, biophysical attributes, and ELISA serologic tests performance. Biotechnol. Bioeng. 2021, 118, 2202–2219. [Google Scholar] [CrossRef]
- Brouwer, P.J.M.; Caniels, T.G.; van der Straten, K.; Snitselaar, J.L.; Aldon, Y.; Bangaru, S.; Torres, J.L.; Okba, N.M.A.; Claireaux, M.; Kerster, G.; et al. Potent neutralizing antibodies from COVID-19 patients define multiple targets of vulnerability. Science 2020, 369, 643–650. [Google Scholar] [CrossRef]
- Yuan, M.; Wu, N.C.; Zhu, X.; Lee, C.-C.D.; So, R.T.Y.; Lv, H.; Mok, C.K.P.; Wilson, I.A. A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science 2020, 368, 630–633. [Google Scholar] [CrossRef] [Green Version]
- Esposito, D.; Mehalko, J.; Drew, M.; Snead, K.; Wall, V.; Taylor, T.; Frank, P.; Denson, J.-P.; Hong, M.; Gulten, G.; et al. Optimizing high-yield production of SARS-CoV-2 soluble spike trimers for serology assays. Protein Expr. Purif. 2020, 174, 105686. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.; Qu, K.; Ciazynska, K.A.; Hosmillo, M.; Carter, A.P.; Ebrahimi, S.; Ke, Z.; Scheres, S.H.W.; Bergamaschi, L.; Grice, G.L.; et al. A thermostable, closed SARS-CoV-2 spike protein trimer. Nat. Struct. Mol. Biol. 2020, 27, 934–941. [Google Scholar] [CrossRef] [PubMed]
- Kollewe, C.; Vilcinskas, A. Production of recombinant proteins in insect cells. Am. J. Biochem. Biotechnol. 2013, 9, 255–271. [Google Scholar] [CrossRef]
- Sanders, R.W.; Moore, J.P. Virus vaccines: Proteins prefer prolines. Cell Host Microbe 2021, 29, 327–333. [Google Scholar] [CrossRef] [PubMed]
- Watterson, D.; Wijesundara, D.K.; Modhiran, N.; Mordant, F.L.; Li, Z.; Avumegah, M.S.; McMillan, C.L.; Lackenby, J.; Guilfoyle, K.; Amerongen, G.; et al. Preclinical development of a molecular clamp-stabilised subunit vaccine for severe acute respiratory syndrome coronavirus 2. Clin. Transl. Immunol. 2021, 10, e1269. [Google Scholar] [CrossRef] [PubMed]
- Oshima, H.; Kinoshita, M. Effects of sugars on the thermal stability of a protein. J. Chem. Phys. 2013, 138, 245101. [Google Scholar] [CrossRef] [PubMed]
- Vagenende, V.; Yap, M.G.S.; Trout, B.L. Mechanisms of Protein Stabilization and Prevention of Protein Aggregation by Glycerol. Biochemistry 2009, 48, 11084–11096. [Google Scholar] [CrossRef]
- Pegg, D.E. Principles of cryopreservation. Methods Mol Biol. 2007, 368, 39–57. [Google Scholar] [CrossRef]
- Herrera, N.G.; Morano, N.C.; Celikgil, A.; Georgiev, G.I.; Malonis, R.J.; Lee, J.H.; Tong, K.; Vergnolle, O.; Massimi, A.B.; Yen, L.Y.; et al. Characterization of the SARS-CoV-2 S Protein: Biophysical, Biochemical, Structural, and Antigenic Analysis. ACS Omega 2021, 6, 85–102. [Google Scholar] [CrossRef]
- Li, T.; Zheng, Q.; Yu, H.; Wu, D.; Xue, W.; Zhang, Y.; Huang, X.; Zhou, L.; Zhang, Z.; Zha, Z.; et al. Characterization of the SARS-CoV-2 Spike in an Early Prefusion Conformation. BioRxiv 2020. [Google Scholar] [CrossRef]
- Ristic, M.; Rosa, N.; Seabrook, S.A.; Newman, J. Formulation screening by differential scanning fluorimetry: How often does it work? Acta Crystallogr. Sect. F Struct. Biol. Commun. 2015, 71, 1359–1364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Den Engelsman, J.; Garidel, P.; Smulders, R.; Koll, H.; Smith, B.; Bassarab, S.; Seidl, A.; Hainzl, O.; Jiskoot, W. Strategies for the Assessment of Protein Aggregates in Pharmaceutical Biotech Product Development. Pharm. Res. 2011, 28, 920–933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wadhwa, M.; Knezevic, I.; Kang, H.-N.; Thorpe, R. Immunogenicity assessment of biotherapeutic products: An overview of assays and their utility. Biologicals 2015, 43, 298–306. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Expression Plasmid | Signal Peptide | rBac Transfer Vector |
|---|---|---|
| rBac 1 | BVM | pOET3 |
| rBac 2 | gp67 | pOET3 |
| rBac 3 | native | pOET3 |
| rBac 4 | BVM | pOET1 |
| rBac 5 | BVM | pOET5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernandes, B.; Castro, R.; Bhoelan, F.; Bemelman, D.; Gomes, R.A.; Costa, J.; Gomes-Alves, P.; Stegmann, T.; Amacker, M.; Alves, P.M.; et al. Insect Cells for High-Yield Production of SARS-CoV-2 Spike Protein: Building a Virosome-Based COVID-19 Vaccine Candidate. Pharmaceutics 2022, 14, 854. https://doi.org/10.3390/pharmaceutics14040854
Fernandes B, Castro R, Bhoelan F, Bemelman D, Gomes RA, Costa J, Gomes-Alves P, Stegmann T, Amacker M, Alves PM, et al. Insect Cells for High-Yield Production of SARS-CoV-2 Spike Protein: Building a Virosome-Based COVID-19 Vaccine Candidate. Pharmaceutics. 2022; 14(4):854. https://doi.org/10.3390/pharmaceutics14040854
Chicago/Turabian StyleFernandes, Bárbara, Rute Castro, Farien Bhoelan, Denzel Bemelman, Ricardo A. Gomes, Júlia Costa, Patrícia Gomes-Alves, Toon Stegmann, Mario Amacker, Paula M. Alves, and et al. 2022. "Insect Cells for High-Yield Production of SARS-CoV-2 Spike Protein: Building a Virosome-Based COVID-19 Vaccine Candidate" Pharmaceutics 14, no. 4: 854. https://doi.org/10.3390/pharmaceutics14040854
APA StyleFernandes, B., Castro, R., Bhoelan, F., Bemelman, D., Gomes, R. A., Costa, J., Gomes-Alves, P., Stegmann, T., Amacker, M., Alves, P. M., Fleury, S., & Roldão, A. (2022). Insect Cells for High-Yield Production of SARS-CoV-2 Spike Protein: Building a Virosome-Based COVID-19 Vaccine Candidate. Pharmaceutics, 14(4), 854. https://doi.org/10.3390/pharmaceutics14040854






